Please be patient as the page loads
|
WWP1_MOUSE
|
||||||
| SwissProt Accessions | Q8BZZ3, Q8BIV9, Q8VDP8 | Gene names | Wwp1 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP1 (EC 6.3.2.-) (WW domain- containing protein 1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
ITCH_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.984252 (rank : 6) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | Q96J02, O43584, Q5TEL0, Q96F66, Q9BY75, Q9H451, Q9H4U5 | Gene names | ITCH | |||
|
Domain Architecture |

|
|||||
| Description | Itchy homolog E3 ubiquitin protein ligase (EC 6.3.2.-) (Itch) (Atrophin-1-interacting protein 4) (AIP4) (NFE2-associated polypeptide 1) (NAPP1). | |||||
|
ITCH_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.991751 (rank : 2) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | Q8C863, O54971 | Gene names | Itch | |||
|
Domain Architecture |

|
|||||
| Description | Itchy E3 ubiquitin protein ligase (EC 6.3.2.-). | |||||
|
WWP1_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.988077 (rank : 5) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 100 | |
| SwissProt Accessions | Q9H0M0, O00307, Q96BP4 | Gene names | WWP1 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP1 (EC 6.3.2.-) (WW domain- containing protein 1) (Atropin-1-interacting protein 5) (AIP5). | |||||
|
WWP1_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 138 | |
| SwissProt Accessions | Q8BZZ3, Q8BIV9, Q8VDP8 | Gene names | Wwp1 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP1 (EC 6.3.2.-) (WW domain- containing protein 1). | |||||
|
WWP2_HUMAN
|
||||||
| θ value | 0 (rank : 5) | NC score | 0.989701 (rank : 4) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | O00308, Q96CZ2, Q9BWN6 | Gene names | WWP2 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP2 (EC 6.3.2.-) (WW domain- containing protein 2) (Atrophin-1-interacting protein 2) (AIP2). | |||||
|
WWP2_MOUSE
|
||||||
| θ value | 0 (rank : 6) | NC score | 0.991677 (rank : 3) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | Q9DBH0, Q8BTG4, Q923F6 | Gene names | Wwp2 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP2 (EC 6.3.2.-) (WW domain- containing protein 2). | |||||
|
NEDD4_HUMAN
|
||||||
| θ value | 3.87392e-144 (rank : 7) | NC score | 0.914846 (rank : 11) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 86 | |
| SwissProt Accessions | P46934 | Gene names | NEDD4, KIAA0093 | |||
|
Domain Architecture |

|
|||||
| Description | E3 ubiquitin-protein ligase NEDD4 (EC 6.3.2.-). | |||||
|
NED4L_HUMAN
|
||||||
| θ value | 2.12569e-142 (rank : 8) | NC score | 0.899753 (rank : 13) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 91 | |
| SwissProt Accessions | Q96PU5, O43165, Q7Z5F1, Q7Z5F2, Q7Z5N3, Q8N5A7, Q8WUU9, Q9BW58, Q9H2W4, Q9NT88 | Gene names | NEDD4L, KIAA0439, NEDL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin-protein ligase NEDD4-like protein (EC 6.3.2.-) (Nedd4-2) (NEDD4.2). | |||||
|
NED4L_MOUSE
|
||||||
| θ value | 8.0776e-142 (rank : 9) | NC score | 0.900556 (rank : 12) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 90 | |
| SwissProt Accessions | Q8CFI0, Q8BRT9, Q8BS42, Q99PK2 | Gene names | Nedd4l, Kiaa0439, Nedd4b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin-protein ligase NEDD4-like protein (EC 6.3.2.-) (Nedd4-2) (NEDD4.2). | |||||
|
NEDD4_MOUSE
|
||||||
| θ value | 1.74296e-136 (rank : 10) | NC score | 0.918255 (rank : 10) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 90 | |
| SwissProt Accessions | P46935, O08758, Q3UZI2, Q8BGB3 | Gene names | Nedd4, Kiaa0093, Nedd-4, Nedd4a | |||
|
Domain Architecture |

|
|||||
| Description | E3 ubiquitin-protein ligase NEDD4 (EC 6.3.2.-) (Neural precursor cell expressed developmentally down-regulated protein 4). | |||||
|
SMUF2_HUMAN
|
||||||
| θ value | 3.88293e-136 (rank : 11) | NC score | 0.960771 (rank : 7) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | Q9HAU4, Q9H260 | Gene names | SMURF2 | |||
|
Domain Architecture |

|
|||||
| Description | Smad ubiquitination regulatory factor 2 (EC 6.3.2.-) (Ubiquitin-- protein ligase SMURF2) (Smad-specific E3 ubiquitin ligase 2) (hSMURF2). | |||||
|
SMUF1_MOUSE
|
||||||
| θ value | 7.33985e-127 (rank : 12) | NC score | 0.953280 (rank : 8) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9CUN6, Q3U412, Q8K300 | Gene names | Smurf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Smad ubiquitination regulatory factor 1 (EC 6.3.2.-) (Ubiquitin-- protein ligase SMURF1) (Smad-specific E3 ubiquitin ligase 1). | |||||
|
SMUF1_HUMAN
|
||||||
| θ value | 2.13557e-126 (rank : 13) | NC score | 0.950211 (rank : 9) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9HCE7, O75853, Q9UJT8 | Gene names | SMURF1, KIAA1625 | |||
|
Domain Architecture |

|
|||||
| Description | Smad ubiquitination regulatory factor 1 (EC 6.3.2.-) (Ubiquitin-- protein ligase SMURF1) (Smad-specific E3 ubiquitin ligase 1) (hSMURF1). | |||||
|
HUWE1_HUMAN
|
||||||
| θ value | 7.93343e-89 (rank : 14) | NC score | 0.866793 (rank : 21) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q7Z6Z7, O15029, Q4G2Z2, Q5H961, Q6P4D0, Q8NG67, Q9BUI0, Q9HCJ4, Q9NSL6, Q9P0A9 | Gene names | HUWE1, KIAA0312, KIAA1578, UREB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (Mcl-1 ubiquitin ligase E3) (Mule) (ARF-binding protein 1) (ARF-BP1). | |||||
|
HUWE1_MOUSE
|
||||||
| θ value | 7.93343e-89 (rank : 15) | NC score | 0.867673 (rank : 20) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q7TMY8, Q4G2Z1, Q5BMM7, Q6NS61, Q8BNJ7, Q8CFH2, Q8VD14, Q921M5, Q9R0P2 | Gene names | Huwe1, Kiaa0312, Ureb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (E3Histone). | |||||
|
HECD2_MOUSE
|
||||||
| θ value | 9.16162e-53 (rank : 16) | NC score | 0.882143 (rank : 14) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8CDU6, Q8CBQ9 | Gene names | Hectd2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase HECTD2 (HECT domain-containing protein 2). | |||||
|
UBE3A_HUMAN
|
||||||
| θ value | 1.32293e-51 (rank : 17) | NC score | 0.874812 (rank : 16) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q05086, P78355, Q93066, Q9UEP4, Q9UEP5, Q9UEP6, Q9UEP7, Q9UEP8, Q9UEP9 | Gene names | UBE3A, E6AP | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase E3A (EC 6.3.2.-) (E6AP ubiquitin-protein ligase) (Oncogenic protein-associated protein E6-AP) (Human papillomavirus E6-associated protein) (NY-REN-54 antigen). | |||||
|
HECD2_HUMAN
|
||||||
| θ value | 6.56568e-51 (rank : 18) | NC score | 0.879269 (rank : 15) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q5U5R9, Q5VZ97, Q5VZ99, Q8TCP5 | Gene names | HECTD2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase HECTD2 (HECT domain-containing protein 2). | |||||
|
UBE3A_MOUSE
|
||||||
| θ value | 6.35935e-46 (rank : 19) | NC score | 0.868105 (rank : 19) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O08759, P97482 | Gene names | Ube3a | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase E3A (EC 6.3.2.-) (Oncogenic protein- associated protein E6-AP). | |||||
|
K0317_HUMAN
|
||||||
| θ value | 1.08474e-45 (rank : 20) | NC score | 0.874331 (rank : 17) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O15033, Q7LDY1, Q8IYY9 | Gene names | KIAA0317 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0317. | |||||
|
K0317_MOUSE
|
||||||
| θ value | 4.122e-45 (rank : 21) | NC score | 0.873727 (rank : 18) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8CHG5, Q6P9Q1, Q80YC4, Q8C5W5 | Gene names | Kiaa0317 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0317. | |||||
|
UBE3C_HUMAN
|
||||||
| θ value | 4.2656e-42 (rank : 22) | NC score | 0.856585 (rank : 22) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q15386, Q8TC15, Q96CR4, Q9UDU3 | Gene names | UBE3C, KIAA0010 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase E3C (EC 6.3.2.-). | |||||
|
UBE3C_MOUSE
|
||||||
| θ value | 3.61106e-41 (rank : 23) | NC score | 0.854985 (rank : 23) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q80U95, Q8BQZ6, Q8C7W6, Q8CDJ1, Q8VDL5 | Gene names | Ube3c, Kiaa0010 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase E3C (EC 6.3.2.-). | |||||
|
HERC3_HUMAN
|
||||||
| θ value | 6.81017e-40 (rank : 24) | NC score | 0.724747 (rank : 26) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q15034 | Gene names | HERC3, KIAA0032 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT domain and RCC1-like domain protein 3. | |||||
|
HERC2_MOUSE
|
||||||
| θ value | 5.22648e-32 (rank : 25) | NC score | 0.620034 (rank : 31) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q4U2R1, O88473, Q3TRJ8, Q3TS47, Q3TST2, Q3UFQ6, Q3URH7, Q5DU32, Q7TPR5, Q80VV7, Q9QYT1, Q9Z168, Q9Z171 | Gene names | Herc2, jdf2, Kiaa0393, rjs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT domain and RCC1-like domain-containing protein 2. | |||||
|
TRIPC_HUMAN
|
||||||
| θ value | 1.52067e-31 (rank : 26) | NC score | 0.793240 (rank : 24) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 319 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q14669, Q15644 | Gene names | TRIP12, KIAA0045 | |||
|
Domain Architecture |

|
|||||
| Description | Thyroid receptor-interacting protein 12 (TRIP12). | |||||
|
HERC2_HUMAN
|
||||||
| θ value | 2.59387e-31 (rank : 27) | NC score | 0.615863 (rank : 32) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O95714, Q86SV7, Q86SV8, Q86SV9, Q86YY3, Q86YY4, Q86YY5, Q86YY6, Q86YY7, Q86YY8, Q86YY9, Q86YZ0, Q86YZ1 | Gene names | HERC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT domain and RCC1-like domain-containing protein 2. | |||||
|
HERC5_HUMAN
|
||||||
| θ value | 7.54701e-31 (rank : 28) | NC score | 0.728197 (rank : 25) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UII4 | Gene names | HERC5, CEB1, CEBP1 | |||
|
Domain Architecture |

|
|||||
| Description | HECT domain and RCC1-like domain-containing protein 5 (Cyclin-E- binding protein 1). | |||||
|
HECD1_HUMAN
|
||||||
| θ value | 8.08199e-25 (rank : 29) | NC score | 0.563312 (rank : 34) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 371 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9ULT8, Q6P445, Q86VJ1, Q96F34, Q9UFZ7 | Gene names | HECTD1, KIAA1131 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin-protein ligase HECTD1 (HECT domain-containing protein 1) (E3 ligase for inhibin receptor) (EULIR). | |||||
|
YAP1_MOUSE
|
||||||
| θ value | 2.60593e-15 (rank : 30) | NC score | 0.563416 (rank : 33) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P46938, Q52KJ5 | Gene names | Yap1, Yap, Yap65 | |||
|
Domain Architecture |

|
|||||
| Description | 65 kDa Yes-associated protein (YAP65). | |||||
|
HECD3_MOUSE
|
||||||
| θ value | 9.90251e-15 (rank : 31) | NC score | 0.689897 (rank : 27) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q3U487, Q3TN76, Q641P3, Q8BQ74, Q8R1L6 | Gene names | Hectd3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase HECTD3 (HECT domain-containing protein 3). | |||||
|
HECD3_HUMAN
|
||||||
| θ value | 4.91457e-14 (rank : 32) | NC score | 0.684451 (rank : 28) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q5T447, Q5T448, Q9H783 | Gene names | HECTD3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase HECTD3 (HECT domain-containing protein 3). | |||||
|
EDD1_HUMAN
|
||||||
| θ value | 1.09485e-13 (rank : 33) | NC score | 0.647820 (rank : 30) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O95071, O94970, Q9NPL3 | Gene names | EDD1, EDD, HYD, KIAA0896 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase EDD1 (EC 6.3.2.-) (Hyperplastic discs protein homolog) (hHYD) (Progestin-induced protein). | |||||
|
EDD1_MOUSE
|
||||||
| θ value | 1.09485e-13 (rank : 34) | NC score | 0.649368 (rank : 29) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q80TP3, Q698K9, Q6PEQ8, Q6PFQ9, Q80VL4, Q810V6, Q9CXE9 | Gene names | Edd1, Edd, Kiaa0896 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase EDD1 (EC 6.3.2.-) (Hyperplastic discs protein homolog). | |||||
|
WWC1_HUMAN
|
||||||
| θ value | 9.26847e-13 (rank : 35) | NC score | 0.445986 (rank : 38) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8IX03, O94946, Q6MZX4, Q6Y2F8, Q7Z4G8, Q8WVM4, Q9BT29 | Gene names | WWC1, KIAA0869 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing protein 1 (Kidney and brain protein) (KIBRA) (HBeAg-binding protein 3). | |||||
|
WWC1_MOUSE
|
||||||
| θ value | 9.26847e-13 (rank : 36) | NC score | 0.445407 (rank : 39) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q5SXA9, Q571D0, Q8K1Y3, Q8VD17, Q922W3 | Gene names | Wwc1, Kiaa0869 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing protein 1 (Kidney and brain protein) (KIBRA). | |||||
|
MAGI2_MOUSE
|
||||||
| θ value | 2.0648e-12 (rank : 37) | NC score | 0.325348 (rank : 48) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 370 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9WVQ1, Q3UH81, Q6GT88, Q8BYT1, Q8CA85 | Gene names | Magi2, Acvrinp1, Aip1, Arip1 | |||
|
Domain Architecture |

|
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 2 (Membrane-associated guanylate kinase inverted 2) (MAGI-2) (Atrophin-1-interacting protein 1) (Activin receptor-interacting protein 1) (Acvrip1). | |||||
|
SAV1_HUMAN
|
||||||
| θ value | 1.74796e-11 (rank : 38) | NC score | 0.534162 (rank : 35) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9H4B6, Q6IA58, Q9H949, Q9HAK9 | Gene names | SAV1, WW45 | |||
|
Domain Architecture |

|
|||||
| Description | Salvador homolog 1 protein (45 kDa WW domain protein) (hWW45). | |||||
|
SAV1_MOUSE
|
||||||
| θ value | 1.74796e-11 (rank : 39) | NC score | 0.533664 (rank : 36) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8VEB2, Q9D9N9, Q9ER46 | Gene names | Sav1, Ww45, Wwp3 | |||
|
Domain Architecture |

|
|||||
| Description | Salvador homolog 1 protein (45 kDa WW domain protein) (mWW45). | |||||
|
WWOX_HUMAN
|
||||||
| θ value | 1.74796e-11 (rank : 40) | NC score | 0.396980 (rank : 40) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9NZC7, Q5MYT5, Q96KM3, Q96RF2, Q9BTT8, Q9NPC9, Q9NRF4, Q9NRF5, Q9NRF6, Q9NRK1, Q9NZC5 | Gene names | WWOX, FOR, WOX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing oxidoreductase (EC 1.1.1.-) (Fragile site FRA16D oxidoreductase). | |||||
|
WWOX_MOUSE
|
||||||
| θ value | 1.74796e-11 (rank : 41) | NC score | 0.393572 (rank : 41) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q91WL8, Q8C8J6, Q920Y2, Q9D2B3, Q9D339, Q9JLF5 | Gene names | Wwox, Wox1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing oxidoreductase (EC 1.1.1.-). | |||||
|
MAGI1_MOUSE
|
||||||
| θ value | 1.47974e-10 (rank : 42) | NC score | 0.303524 (rank : 52) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q6RHR9, O54893, O54894, O54895 | Gene names | Magi1, Baiap1, Bap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 1 (BAI1-associated protein 1) (BAP-1) (Membrane-associated guanylate kinase inverted 1) (MAGI-1). | |||||
|
MAGI1_HUMAN
|
||||||
| θ value | 1.9326e-10 (rank : 43) | NC score | 0.304720 (rank : 51) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q96QZ7, O00309, O43863, O75085, Q96QZ8, Q96QZ9 | Gene names | MAGI1, BAIAP1, BAP1, TNRC19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 1 (BAI1-associated protein 1) (BAP-1) (Membrane-associated guanylate kinase inverted 1) (MAGI-1) (Atrophin-1-interacting protein 3) (AIP3) (WW domain-containing protein 3) (WWP3) (Trinucleotide repeat-containing gene 19 protein). | |||||
|
MAGI2_HUMAN
|
||||||
| θ value | 4.30538e-10 (rank : 44) | NC score | 0.309937 (rank : 50) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 373 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q86UL8, O60434, O60510, Q86UI7, Q9NP44, Q9UDQ5, Q9UDU1 | Gene names | MAGI2, ACVRINP1, AIP1, KIAA0705 | |||
|
Domain Architecture |

|
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 2 (Membrane-associated guanylate kinase inverted 2) (MAGI-2) (Atrophin-1-interacting protein 1) (Atrophin-1-interacting protein A). | |||||
|
PKHA5_HUMAN
|
||||||
| θ value | 3.64472e-09 (rank : 45) | NC score | 0.286133 (rank : 53) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9HAU0, Q8N3K6, Q96DY9, Q9BVR4, Q9C0H7, Q9H924, Q9NVK8 | Gene names | PLEKHA5, KIAA1686, PEPP2 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 5 (Phosphoinositol 3-phosphate-binding protein 2) (PEPP-2). | |||||
|
PRP40_HUMAN
|
||||||
| θ value | 7.59969e-07 (rank : 46) | NC score | 0.231679 (rank : 56) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 442 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O75400, O43856, O75404, Q8TBQ1, Q9H782, Q9NWU9, Q9P0Q2, Q9Y5A8 | Gene names | PRPF40A, FLAF1, FNBP3, HYPA | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Huntingtin yeast partner A) (Huntingtin-interacting protein HYPA/FBP11) (Fas ligand-associated factor 1) (NY-REN-6 antigen). | |||||
|
PRP40_MOUSE
|
||||||
| θ value | 7.59969e-07 (rank : 47) | NC score | 0.230368 (rank : 57) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9R1C7, Q61049, Q8BQ76, Q8BRW4, Q8C2U1 | Gene names | Prpf40a, Fbp11, Fnbp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Formin-binding protein 11) (FBP-11). | |||||
|
BAG3_HUMAN
|
||||||
| θ value | 2.88788e-06 (rank : 48) | NC score | 0.356893 (rank : 47) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 314 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O95817, Q9NT20, Q9P120 | Gene names | BAG3, BIS | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 3 (BCL-2-binding athanogene- 3) (BAG-3) (Bcl-2-binding protein Bis) (Docking protein CAIR-1). | |||||
|
YAP1_HUMAN
|
||||||
| θ value | 8.40245e-06 (rank : 49) | NC score | 0.384374 (rank : 45) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P46937 | Gene names | YAP1, YAP65 | |||
|
Domain Architecture |

|
|||||
| Description | 65 kDa Yes-associated protein (YAP65). | |||||
|
GAS7_MOUSE
|
||||||
| θ value | 1.43324e-05 (rank : 50) | NC score | 0.271894 (rank : 55) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q60780, Q9QY25, Q9QY26, Q9QY34 | Gene names | Gas7 | |||
|
Domain Architecture |

|
|||||
| Description | Growth-arrest-specific protein 7 (GAS-7). | |||||
|
BAG3_MOUSE
|
||||||
| θ value | 2.44474e-05 (rank : 51) | NC score | 0.310464 (rank : 49) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9JLV1, Q9JJC7 | Gene names | Bag3, Bis | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 3 (BCL-2-binding athanogene- 3) (BAG-3) (Bcl-2-binding protein Bis). | |||||
|
PINL_HUMAN
|
||||||
| θ value | 2.44474e-05 (rank : 52) | NC score | 0.450748 (rank : 37) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O15428 | Gene names | PIN1L | |||
|
Domain Architecture |
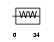
|
|||||
| Description | PIN1-like protein. | |||||
|
GAS7_HUMAN
|
||||||
| θ value | 3.19293e-05 (rank : 53) | NC score | 0.272592 (rank : 54) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O60861, O43144 | Gene names | GAS7, KIAA0394 | |||
|
Domain Architecture |

|
|||||
| Description | Growth-arrest-specific protein 7 (GAS-7). | |||||
|
WWTR1_HUMAN
|
||||||
| θ value | 7.1131e-05 (rank : 54) | NC score | 0.383641 (rank : 46) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9GZV5, Q8N3P2, Q9Y3W6 | Gene names | WWTR1, TAZ | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing transcription regulator protein 1 (Transcriptional coactivator with PDZ-binding motif). | |||||
|
WWTR1_MOUSE
|
||||||
| θ value | 7.1131e-05 (rank : 55) | NC score | 0.387002 (rank : 42) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9EPK5, Q99KI4 | Gene names | Wwtr1, Taz | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing transcription regulator protein 1 (Transcriptional coactivator with PDZ-binding motif). | |||||
|
PIN1_HUMAN
|
||||||
| θ value | 0.000270298 (rank : 56) | NC score | 0.384680 (rank : 44) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q13526 | Gene names | PIN1 | |||
|
Domain Architecture |
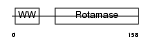
|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase NIMA-interacting 1 (EC 5.2.1.8) (Rotamase Pin1) (PPIase Pin1). | |||||
|
PIN1_MOUSE
|
||||||
| θ value | 0.000270298 (rank : 57) | NC score | 0.386618 (rank : 43) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9QUR7, Q543B3 | Gene names | Pin1 | |||
|
Domain Architecture |
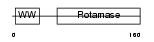
|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase NIMA-interacting 1 (EC 5.2.1.8) (Rotamase Pin1) (PPIase Pin1). | |||||
|
RHG12_HUMAN
|
||||||
| θ value | 0.000602161 (rank : 58) | NC score | 0.111169 (rank : 67) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8IWW6, Q86UB3, Q8IWW7, Q8N3L1, Q9NT76 | Gene names | ARHGAP12 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 12. | |||||
|
RHG12_MOUSE
|
||||||
| θ value | 0.00102713 (rank : 59) | NC score | 0.125458 (rank : 65) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8C0D4, Q8BVP8 | Gene names | Arhgap12 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 12. | |||||
|
STXB4_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 60) | NC score | 0.178053 (rank : 59) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 616 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q6ZWJ1, Q8IVZ5 | Gene names | STXBP4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntaxin-binding protein 4 (Syntaxin 4-interacting protein) (Synip). | |||||
|
STXB4_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 61) | NC score | 0.178963 (rank : 58) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 629 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9WV89, Q5SUA6, Q5SUA8, Q5SUA9, Q8CFL1 | Gene names | Stxbp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntaxin-binding protein 4 (Syntaxin 4-interacting protein) (Synip). | |||||
|
DMD_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 62) | NC score | 0.055173 (rank : 126) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P11532, Q02295, Q14169, Q14170, Q5JYU0, Q7KZ48 | Gene names | DMD | |||
|
Domain Architecture |

|
|||||
| Description | Dystrophin. | |||||
|
DMD_MOUSE
|
||||||
| θ value | 0.00509761 (rank : 63) | NC score | 0.050562 (rank : 139) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1111 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P11531, O35653, Q60703 | Gene names | Dmd | |||
|
Domain Architecture |

|
|||||
| Description | Dystrophin. | |||||
|
MYOF_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 64) | NC score | 0.073017 (rank : 93) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9NZM1, Q9HBU3, Q9NZM0, Q9ULL3, Q9Y4U4 | Gene names | FER1L3, KIAA1207, MYOF | |||
|
Domain Architecture |

|
|||||
| Description | Myoferlin (Fer-1-like protein 3). | |||||
|
SETD2_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 65) | NC score | 0.139144 (rank : 62) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q9BYW2, O75397, O75405, Q17RW8, Q5BKS9, Q5QGN2, Q69YI5, Q6IN64, Q6ZN53, Q6ZS25, Q8N3R0, Q8TCN0, Q9C0D1, Q9H696, Q9NZW9 | Gene names | SETD2, HIF1, HYPB, KIAA1732, SET2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36-specific (EC 2.1.1.43) (SET domain-containing protein 2) (hSET2) (Huntingtin- interacting protein HYPB) (Huntingtin-interacting protein 1) (HIF-1) (p231HBP). | |||||
|
APBB2_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 66) | NC score | 0.161005 (rank : 61) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9DBR4, Q6DFX8 | Gene names | Apbb2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 2. | |||||
|
TCRG1_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 67) | NC score | 0.090031 (rank : 74) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q8CGF7, Q61051, Q8C490, Q8CHT8, Q9R0R5 | Gene names | Tcerg1, Taf2s | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150) (p144) (Formin- binding protein 28) (FBP 28). | |||||
|
APBB2_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 68) | NC score | 0.171308 (rank : 60) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q92870, Q8IUI6 | Gene names | APBB2, FE65L, FE65L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 2 (Fe65-like protein). | |||||
|
TCRG1_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 69) | NC score | 0.090370 (rank : 72) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 929 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O14776 | Gene names | TCERG1, CA150, TAF2S | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150). | |||||
|
MDC1_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 70) | NC score | 0.030067 (rank : 149) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
SYTL4_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 71) | NC score | 0.046738 (rank : 143) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q96C24, Q5H9J3, Q5JPG8, Q8N9P4, Q9H4R0, Q9H4R1 | Gene names | SYTL4 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 4 (Exophilin-2) (Granuphilin). | |||||
|
SYTL4_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 72) | NC score | 0.044530 (rank : 145) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9R0Q1, Q8R321, Q9R0Q0 | Gene names | Sytl4, Slp4 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 4 (Exophilin-2) (Granuphilin). | |||||
|
DGCR8_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 73) | NC score | 0.093082 (rank : 70) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8WYQ5, Q6DCB2, Q6MZE9, Q6Y2L0, Q96G39, Q96GP8, Q9H6L8, Q9H6T7, Q9NRW2 | Gene names | DGCR8, C22orf12, DGCRK6 | |||
|
Domain Architecture |
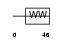
|
|||||
| Description | DGCR8 protein (DiGeorge syndrome critical region 8). | |||||
|
DGCR8_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 74) | NC score | 0.091070 (rank : 71) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9EQM6 | Gene names | Dgcr8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DGCR8 protein (DiGeorge syndrome critical region 8 homolog) (Gy1). | |||||
|
DRP2_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 75) | NC score | 0.105234 (rank : 68) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q13474 | Gene names | DRP2 | |||
|
Domain Architecture |

|
|||||
| Description | Dystrophin-related protein 2. | |||||
|
ZN217_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 76) | NC score | 0.001454 (rank : 186) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 905 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O75362 | Gene names | ZNF217, ZABC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 217. | |||||
|
APBB3_HUMAN
|
||||||
| θ value | 0.125558 (rank : 77) | NC score | 0.128743 (rank : 63) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O95704, Q9NYX6, Q9NYX7, Q9NYX8 | Gene names | APBB3, FE65L2 | |||
|
Domain Architecture |
|
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 3 (Fe65-like protein 2) (Fe65L2). | |||||
|
APBB3_MOUSE
|
||||||
| θ value | 0.163984 (rank : 78) | NC score | 0.127837 (rank : 64) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8R1C9 | Gene names | Apbb3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 3. | |||||
|
EPN3_MOUSE
|
||||||
| θ value | 0.163984 (rank : 79) | NC score | 0.013909 (rank : 164) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q91W69, Q9CV55 | Gene names | Epn3 | |||
|
Domain Architecture |
|
|||||
| Description | Epsin-3 (EPS-15-interacting protein 3). | |||||
|
UTRO_HUMAN
|
||||||
| θ value | 0.21417 (rank : 80) | NC score | 0.046023 (rank : 144) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P46939 | Gene names | UTRN, DMDL | |||
|
Domain Architecture |

|
|||||
| Description | Utrophin (Dystrophin-related protein 1) (DRP1) (DRP). | |||||
|
AIM1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 81) | NC score | 0.016966 (rank : 161) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Y4K1, O00296, Q5VWJ2 | Gene names | AIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Absent in melanoma 1 protein. | |||||
|
4ET_HUMAN
|
||||||
| θ value | 0.813845 (rank : 82) | NC score | 0.018148 (rank : 159) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NRA8, Q8NCF2, Q9H708 | Gene names | EIF4ENIF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4E transporter (eIF4E transporter) (4E-T) (Eukaryotic translation initiation factor 4E nuclear import factor 1). | |||||
|
CSF1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 83) | NC score | 0.019882 (rank : 155) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P09603, Q13130, Q14086, Q14806, Q9UQR8 | Gene names | CSF1 | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage colony-stimulating factor 1 precursor (CSF-1) (MCSF) (M- CSF) (Lanimostim). | |||||
|
ANK2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 84) | NC score | 0.013597 (rank : 165) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
APBB1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 85) | NC score | 0.090203 (rank : 73) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O00213, Q96A93 | Gene names | APBB1, FE65 | |||
|
Domain Architecture |

|
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 1 (Fe65 protein). | |||||
|
APBB1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 86) | NC score | 0.089608 (rank : 75) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9QXJ1, O08642, Q3TPU0, Q8BNF4, Q8BSR9 | Gene names | Apbb1, Fe65 | |||
|
Domain Architecture |

|
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 1 (Fe65 protein). | |||||
|
DYSF_HUMAN
|
||||||
| θ value | 1.06291 (rank : 87) | NC score | 0.062360 (rank : 106) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O75923, O75696, Q9UEN7 | Gene names | DYSF, FER1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Dysferlin (Dystrophy-associated fer-1-like protein) (Fer-1-like protein 1). | |||||
|
MLH3_HUMAN
|
||||||
| θ value | 1.06291 (rank : 88) | NC score | 0.024248 (rank : 152) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UHC1, P49751, Q56DK9, Q9P292, Q9UHC0 | Gene names | MLH3 | |||
|
Domain Architecture |

|
|||||
| Description | DNA mismatch repair protein Mlh3 (MutL protein homolog 3). | |||||
|
UBP2L_HUMAN
|
||||||
| θ value | 1.06291 (rank : 89) | NC score | 0.018271 (rank : 158) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q14157, Q5VU75, Q5VU76, Q9BTU3, Q9UGL5 | Gene names | UBAP2L, KIAA0144, NICE4 | |||
|
Domain Architecture |
|
|||||
| Description | Ubiquitin-associated protein 2-like (Protein NICE-4). | |||||
|
CDKL2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 90) | NC score | -0.004645 (rank : 194) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 838 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q92772 | Gene names | CDKL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-dependent kinase-like 2 (EC 2.7.11.22) (Serine/threonine- protein kinase KKIAMRE) (Protein kinase p56 KKIAMRE). | |||||
|
DLG7_HUMAN
|
||||||
| θ value | 1.81305 (rank : 91) | NC score | 0.025607 (rank : 151) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q15398, Q8NG58 | Gene names | DLG7, KIAA0008 | |||
|
Domain Architecture |

|
|||||
| Description | Discs large homolog 7 (Hepatoma up-regulated protein) (HURP). | |||||
|
DLG7_MOUSE
|
||||||
| θ value | 1.81305 (rank : 92) | NC score | 0.022937 (rank : 153) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8K4R9, Q6ZQL3, Q80VS7, Q8BUC8, Q8BZ79 | Gene names | Dlg7, Kiaa0008 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Discs large homolog 7 (Hepatoma up-regulated protein homolog) (HURP). | |||||
|
PAK7_HUMAN
|
||||||
| θ value | 1.81305 (rank : 93) | NC score | -0.004660 (rank : 195) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 875 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9P286, Q5W115, Q9BX09, Q9ULF6 | Gene names | PAK7, KIAA1264, PAK5 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase PAK 7 (EC 2.7.11.1) (p21-activated kinase 7) (PAK-7) (PAK-5). | |||||
|
PCLO_HUMAN
|
||||||
| θ value | 1.81305 (rank : 94) | NC score | 0.061484 (rank : 109) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
BMX_HUMAN
|
||||||
| θ value | 2.36792 (rank : 95) | NC score | 0.000225 (rank : 189) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 942 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P51813, O60564, Q12871 | Gene names | BMX | |||
|
Domain Architecture |

|
|||||
| Description | Cytoplasmic tyrosine-protein kinase BMX (EC 2.7.10.2) (Bone marrow tyrosine kinase gene in chromosome X protein) (Epithelial and endothelial tyrosine kinase) (ETK) (NTK38). | |||||
|
CKAP2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 96) | NC score | 0.012559 (rank : 168) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8WWK9, Q3KRA5, Q5VXB4, Q8IWV5, Q8IWV6, Q96FH9, Q9H012, Q9H0D0, Q9H988, Q9HC49, Q9NVG4 | Gene names | CKAP2, LB1, TMAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-associated protein 2 (Tumor-associated microtubule- associated protein) (CTCL tumor antigen se20-10). | |||||
|
MAP4_HUMAN
|
||||||
| θ value | 2.36792 (rank : 97) | NC score | 0.012177 (rank : 169) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 428 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P27816, Q13082, Q96A76 | Gene names | MAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 4 (MAP 4). | |||||
|
RBBP6_HUMAN
|
||||||
| θ value | 2.36792 (rank : 98) | NC score | 0.018792 (rank : 157) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
MILK1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 99) | NC score | 0.014965 (rank : 162) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 788 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BGT6, Q8BJ60 | Gene names | Mirab13, D15Mit260, Kiaa1668 | |||
|
Domain Architecture |

|
|||||
| Description | Molecule interacting with Rab13 (MIRab13) (MICAL-like protein 1). | |||||
|
NCOR1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 100) | NC score | 0.009506 (rank : 171) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 521 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q60974, Q60812 | Gene names | Ncor1, Rxrip13 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR) (Retinoid X receptor- interacting protein 13) (RIP13). | |||||
|
PCLO_MOUSE
|
||||||
| θ value | 3.0926 (rank : 101) | NC score | 0.062943 (rank : 105) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
PECI_MOUSE
|
||||||
| θ value | 3.0926 (rank : 102) | NC score | 0.006388 (rank : 178) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9WUR2 | Gene names | Peci | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisomal 3,2-trans-enoyl-CoA isomerase (EC 5.3.3.8) (Dodecenoyl-CoA isomerase) (Delta(3),delta(2)-enoyl-CoA isomerase) (D3,D2-enoyl-CoA isomerase). | |||||
|
PRLR_MOUSE
|
||||||
| θ value | 3.0926 (rank : 103) | NC score | 0.006590 (rank : 177) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q08501, P15212, P15213, Q62099 | Gene names | Prlr | |||
|
Domain Architecture |
|
|||||
| Description | Prolactin receptor precursor (PRL-R). | |||||
|
RFIP1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 104) | NC score | 0.021632 (rank : 154) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9D620, Q3UBC2, Q8BN24 | Gene names | Rab11fip1, Rcp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab11 family-interacting protein 1 (Rab11-FIP1) (Rab-coupling protein). | |||||
|
AKA12_HUMAN
|
||||||
| θ value | 4.03905 (rank : 105) | NC score | 0.033134 (rank : 148) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q02952, O00310, O00498, Q99970 | Gene names | AKAP12, AKAP250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 12 (A-kinase anchor protein 250 kDa) (AKAP 250) (Myasthenia gravis autoantigen gravin). | |||||
|
AKAP1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 106) | NC score | 0.009324 (rank : 172) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O08715, O08714, P97488 | Gene names | Akap1, Akap | |||
|
Domain Architecture |

|
|||||
| Description | A kinase anchor protein 1, mitochondrial precursor (Protein kinase A- anchoring protein 1) (PRKA1) (Dual specificity A-kinase-anchoring protein 1) (D-AKAP-1) (Spermatid A-kinase anchor protein) (S-AKAP). | |||||
|
FOXN3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 107) | NC score | 0.001765 (rank : 185) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O00409, Q96II7, Q9UIE7 | Gene names | CHES1, FOXN3 | |||
|
Domain Architecture |
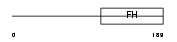
|
|||||
| Description | Checkpoint suppressor 1 (Forkhead box protein N3). | |||||
|
ITSN1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 108) | NC score | 0.038542 (rank : 146) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1757 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q15811, O95216, Q9UET5, Q9UK60, Q9UNK1, Q9UNK2, Q9UQ92 | Gene names | ITSN1, ITSN, SH3D1A | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-1 (SH3 domain-containing protein 1A) (SH3P17). | |||||
|
ITSN1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 109) | NC score | 0.037742 (rank : 147) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1669 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9Z0R4, Q9R143 | Gene names | Itsn1, Ese1, Itsn | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-1 (EH and SH3 domains protein 1). | |||||
|
OTOF_HUMAN
|
||||||
| θ value | 4.03905 (rank : 110) | NC score | 0.050757 (rank : 138) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9HC10, Q9HC08, Q9HC09, Q9Y650 | Gene names | OTOF, FER1L2 | |||
|
Domain Architecture |

|
|||||
| Description | Otoferlin (Fer-1-like protein 2). | |||||
|
OTOF_MOUSE
|
||||||
| θ value | 4.03905 (rank : 111) | NC score | 0.052683 (rank : 131) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9ESF1, Q9ESF2 | Gene names | Otof, Fer1l2 | |||
|
Domain Architecture |

|
|||||
| Description | Otoferlin (Fer-1-like protein 2). | |||||
|
FA62A_MOUSE
|
||||||
| θ value | 5.27518 (rank : 112) | NC score | 0.101037 (rank : 69) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q3U7R1, Q8C8R1, Q91X62, Q9CVH0, Q9Z1X5, Q9Z1X6 | Gene names | Fam62a, Mbc2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM62A (Membrane-bound C2 domain-containing protein). | |||||
|
KSR2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 113) | NC score | -0.004243 (rank : 193) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 920 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6VAB6, Q8N775 | Gene names | KSR2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinase suppressor of ras-2 (hKSR2). | |||||
|
NUMB_HUMAN
|
||||||
| θ value | 5.27518 (rank : 114) | NC score | 0.008409 (rank : 174) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P49757, Q6NUQ7, Q86SY1, Q8WW73, Q9UBG1, Q9UEQ4, Q9UKE8, Q9UKE9, Q9UKF0, Q9UQJ4 | Gene names | NUMB | |||
|
Domain Architecture |
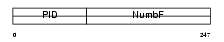
|
|||||
| Description | Protein numb homolog (h-Numb) (Protein S171). | |||||
|
WBP4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 115) | NC score | 0.058735 (rank : 119) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O75554 | Gene names | WBP4, FBP21, FNBP21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 4 (WBP-4) (Formin-binding protein 21). | |||||
|
WBP4_MOUSE
|
||||||
| θ value | 5.27518 (rank : 116) | NC score | 0.119079 (rank : 66) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q61048, Q8K1Z9, Q9CS45 | Gene names | Wbp4, Fbp21, Fnbp21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 4 (WBP-4) (Formin-binding protein 21). | |||||
|
CN103_MOUSE
|
||||||
| θ value | 6.88961 (rank : 117) | NC score | 0.006218 (rank : 179) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q80XK6, Q8R1H5, Q9CWK6 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C14orf103 homolog. | |||||
|
DSG2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 118) | NC score | -0.001079 (rank : 191) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14126 | Gene names | DSG2 | |||
|
Domain Architecture |

|
|||||
| Description | Desmoglein-2 precursor (HDGC). | |||||
|
DYSF_MOUSE
|
||||||
| θ value | 6.88961 (rank : 119) | NC score | 0.052556 (rank : 132) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9ESD7, Q9QXC0 | Gene names | Dysf, Fer1l1 | |||
|
Domain Architecture |

|
|||||
| Description | Dysferlin (Dystrophy-associated fer-1-like protein) (Fer-1-like protein 1). | |||||
|
HA26_HUMAN
|
||||||
| θ value | 6.88961 (rank : 120) | NC score | 0.000717 (rank : 188) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P01906, O19789 | Gene names | HLA-DQA2 | |||
|
Domain Architecture |
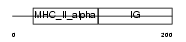
|
|||||
| Description | HLA class II histocompatibility antigen, DQ(6) alpha chain precursor (DX alpha chain) (HLA-DQA1). | |||||
|
LIMA1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 121) | NC score | 0.013356 (rank : 166) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9ERG0, Q9ERG1 | Gene names | Lima1, D15Ertd366e, Eplin | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain and actin-binding protein 1 (Epithelial protein lost in neoplasm) (mEPLIN). | |||||
|
PDLI4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 122) | NC score | 0.010521 (rank : 170) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P70271, Q8K0W4 | Gene names | Pdlim4, Ril | |||
|
Domain Architecture |

|
|||||
| Description | PDZ and LIM domain protein 4 (LIM protein RIL) (Reversion-induced LIM protein). | |||||
|
RBBP6_MOUSE
|
||||||
| θ value | 6.88961 (rank : 123) | NC score | 0.019241 (rank : 156) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
RM49_HUMAN
|
||||||
| θ value | 6.88961 (rank : 124) | NC score | 0.008530 (rank : 173) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q13405 | Gene names | MRPL49, C11orf4, NOF1 | |||
|
Domain Architecture |
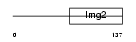
|
|||||
| Description | Mitochondrial 39S ribosomal protein L49 (L49mt) (MRP-L49) (Protein NOF1) (Neighbor of FAU) (NOF). | |||||
|
TARA_HUMAN
|
||||||
| θ value | 6.88961 (rank : 125) | NC score | 0.028023 (rank : 150) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9H2D6, O94797, Q2PZW8, Q2Q3Z9, Q2Q400, Q5R3M6, Q96DW1, Q9BT77, Q9BTL7, Q9BY98, Q9Y3L4 | Gene names | TRIOBP, KIAA1662, TARA | |||
|
Domain Architecture |

|
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
ZAN_MOUSE
|
||||||
| θ value | 6.88961 (rank : 126) | NC score | 0.005190 (rank : 180) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 808 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O88799, O08647 | Gene names | Zan | |||
|
Domain Architecture |

|
|||||
| Description | Zonadhesin precursor. | |||||
|
4ET_MOUSE
|
||||||
| θ value | 8.99809 (rank : 127) | NC score | 0.012995 (rank : 167) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9EST3, Q9CSS3 | Gene names | Eif4enif1, Clast4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4E transporter (eIF4E transporter) (4E-T) (Eukaryotic translation initiation factor 4E nuclear import factor 1). | |||||
|
BCAR1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 128) | NC score | 0.001107 (rank : 187) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 554 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P56945 | Gene names | BCAR1, CAS, CRKAS | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer anti-estrogen resistance protein 1 (CRK-associated substrate) (p130cas). | |||||
|
CF060_HUMAN
|
||||||
| θ value | 8.99809 (rank : 129) | NC score | 0.001948 (rank : 184) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1270 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8NB25, Q96GY8, Q9H851 | Gene names | C6orf60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C6orf60. | |||||
|
E2AK3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 130) | NC score | -0.000894 (rank : 190) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 859 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NZJ5, O95846 | Gene names | EIF2AK3, PEK, PERK | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2-alpha kinase 3 precursor (EC 2.7.11.1) (PRKR-like endoplasmic reticulum kinase) (Pancreatic eIF2-alpha kinase) (HsPEK). | |||||
|
MINT_MOUSE
|
||||||
| θ value | 8.99809 (rank : 131) | NC score | 0.007330 (rank : 176) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
MLL3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 132) | NC score | 0.004652 (rank : 182) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
ORC3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 133) | NC score | 0.008018 (rank : 175) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UBD5, Q13565, Q6IUY7, Q9UG44, Q9UNT6 | Gene names | ORC3L, LATHEO | |||
|
Domain Architecture |
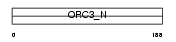
|
|||||
| Description | Origin recognition complex subunit 3 (Origin recognition complex subunit Latheo). | |||||
|
PHC1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 134) | NC score | 0.004818 (rank : 181) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P78364, Q8WVM3, Q9BU63 | Gene names | PHC1, EDR1, PH1 | |||
|
Domain Architecture |

|
|||||
| Description | Polyhomeotic-like protein 1 (hPH1) (Early development regulatory protein 1). | |||||
|
RFIP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 135) | NC score | 0.017957 (rank : 160) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q6WKZ4, Q6AZK4, Q6WKZ2, Q6WKZ6, Q86YV4, Q8TDL1, Q9H642 | Gene names | RAB11FIP1, RCP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab11 family-interacting protein 1 (Rab11-FIP1) (Rab-coupling protein). | |||||
|
RGS3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 136) | NC score | 0.014319 (rank : 163) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
SCN7A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 137) | NC score | 0.001973 (rank : 183) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q01118 | Gene names | SCN7A, SCN6A | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 7 subunit alpha (Sodium channel protein type VII subunit alpha) (Putative voltage-gated sodium channel alpha subunit Nax) (Sodium channel protein, cardiac and skeletal muscle alpha-subunit). | |||||
|
SUHW4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 138) | NC score | -0.002583 (rank : 192) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 737 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q68FE8, Q3T9B1, Q8BI82 | Gene names | Suhw4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppressor of hairy wing homolog 4. | |||||
|
ALS2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 139) | NC score | 0.052745 (rank : 130) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96Q42, Q8N1E0, Q96PC4, Q96Q41, Q9H973, Q9HCK9 | Gene names | ALS2, KIAA1563 | |||
|
Domain Architecture |

|
|||||
| Description | Alsin (Amyotrophic lateral sclerosis protein 2). | |||||
|
ALS2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 140) | NC score | 0.050949 (rank : 136) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q920R0, Q8JZR1, Q9CXJ3 | Gene names | Als2 | |||
|
Domain Architecture |

|
|||||
| Description | Alsin (Amyotrophic lateral sclerosis protein 2 homolog). | |||||
|
BAIP3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 141) | NC score | 0.052058 (rank : 133) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q80TT2, Q6RUT4 | Gene names | Baiap3, Gm937, Kiaa0734 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BAI1-associated protein 3 (BAP3). | |||||
|
CUL7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 142) | NC score | 0.056596 (rank : 123) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q14999, Q5T654 | Gene names | CUL7, KIAA0076 | |||
|
Domain Architecture |

|
|||||
| Description | Cullin-7 (CUL-7). | |||||
|
DOC2A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 143) | NC score | 0.069665 (rank : 97) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q14183, Q6P4G4, Q7Z5G0, Q8IVX0 | Gene names | DOC2A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double C2-like domain-containing protein alpha (Doc2-alpha) (Doc2). | |||||
|
DOC2A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 144) | NC score | 0.073844 (rank : 91) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q7TNF0 | Gene names | Doc2a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double C2-like domain-containing protein alpha (Doc2-alpha). | |||||
|
DOC2B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 145) | NC score | 0.071328 (rank : 94) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q14184 | Gene names | DOC2B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double C2-like domain-containing protein beta (Doc2-beta). | |||||
|
DOC2B_MOUSE
|
||||||
| θ value | θ > 10 (rank : 146) | NC score | 0.070229 (rank : 96) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P70169, Q6NXK3 | Gene names | Doc2b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double C2-like domain-containing protein beta (Doc2-beta). | |||||
|
DOC2G_MOUSE
|
||||||
| θ value | θ > 10 (rank : 147) | NC score | 0.058143 (rank : 120) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9ESN1, Q6P8R8 | Gene names | Doc2g | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double C2-like domain-containing protein gamma (Doc2-gamma). | |||||
|
FA62A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 148) | NC score | 0.084881 (rank : 77) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9BSJ8, O94848, Q6PJN4, Q9H6J1, Q9H6W2, Q9Y416 | Gene names | FAM62A, KIAA0747, MBC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM62A (Membrane-bound C2 domain-containing protein). | |||||
|
K1333_MOUSE
|
||||||
| θ value | θ > 10 (rank : 149) | NC score | 0.063216 (rank : 104) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5RJY2, Q6ZPT7, Q8BNA4 | Gene names | Kiaa1333 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1333 (EC 6.3.2.-). | |||||
|
PA24B_MOUSE
|
||||||
| θ value | θ > 10 (rank : 150) | NC score | 0.059023 (rank : 116) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q4QQM1, Q80VV8, Q91W88 | Gene names | Pla2g4b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytosolic phospholipase A2 beta (EC 3.1.1.4) (cPLA2-beta) (Phospholipase A2 group IVB). | |||||
|
RASL1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 151) | NC score | 0.077622 (rank : 89) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O95294, Q52M03, Q96CC7 | Gene names | RASAL1, RASAL | |||
|
Domain Architecture |

|
|||||
| Description | RasGAP-activating-like protein 1. | |||||
|
RASL1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 152) | NC score | 0.073191 (rank : 92) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9Z268 | Gene names | Rasal1, Rasal | |||
|
Domain Architecture |

|
|||||
| Description | RasGAP-activating-like protein 1. | |||||
|
RASL2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 153) | NC score | 0.070452 (rank : 95) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O43374, O60286, Q86UW3, Q96QU0 | Gene names | RASA4, CAPRI, GAPL, KIAA0538 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 4 (RasGAP-activating-like protein 2) (Calcium-promoted Ras inactivator). | |||||
|
RCBT1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 154) | NC score | 0.060145 (rank : 113) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NDN9, Q969U9 | Gene names | RCBTB1, CLLD7, E4.5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RCC1 and BTB domain-containing protein 1 (Regulator of chromosome condensation and BTB domain-containing protein 1) (Chronic lymphocytic leukemia deletion region gene 7 protein) (CLL deletion region gene 7 protein). | |||||
|
RCBT1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 155) | NC score | 0.061754 (rank : 108) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6NXM2, Q8BTZ6, Q8BZV0 | Gene names | Rcbtb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RCC1 and BTB domain-containing protein 1 (Regulator of chromosome condensation and BTB domain-containing protein 1). | |||||
|
RCBT2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 156) | NC score | 0.066156 (rank : 102) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O95199 | Gene names | RCBTB2, CHC1L, RLG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RCC1 and BTB domain-containing protein 2 (Regulator of chromosome condensation and BTB domain-containing protein 2) (Chromosome condensation 1-like) (CHC1-L) (RCC1-like G exchanging factor). | |||||
|
RCBT2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 157) | NC score | 0.067083 (rank : 101) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99LJ7, Q3TUA3, Q8BMG2 | Gene names | Rcbtb2, Chc1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RCC1 and BTB domain-containing protein 2 (Regulator of chromosome condensation and BTB domain-containing protein 2) (Chromosome condensation 1-like). | |||||
|
RCC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 158) | NC score | 0.081059 (rank : 85) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P18754 | Gene names | RCC1, CHC1 | |||
|
Domain Architecture |
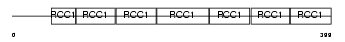
|
|||||
| Description | Regulator of chromosome condensation (Chromosome condensation protein 1) (Cell cycle regulatory protein). | |||||
|
RCC1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 159) | NC score | 0.078949 (rank : 87) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8VE37, Q3UDB6 | Gene names | Rcc1, Chc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of chromosome condensation (Chromosome condensation protein 1). | |||||
|
RCC2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 160) | NC score | 0.083536 (rank : 82) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9P258, Q8IVL9, Q9BSN6, Q9NPV8 | Gene names | RCC2, KIAA1470, TD60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein RCC2 (Telophase disk protein of 60 kDa) (RCC1-like protein TD- 60). | |||||
|
RCC2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 161) | NC score | 0.083060 (rank : 83) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BK67, Q6ZPQ0 | Gene names | Rcc2, Kiaa1470 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein RCC2. | |||||
|
RIMS2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 162) | NC score | 0.068285 (rank : 99) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UQ26, O43413, Q86XL9, Q8IWV9, Q8IWW1 | Gene names | RIMS2, KIAA0751, RIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 2 (Rab3-interacting molecule 2) (RIM 2). | |||||
|
RIMS2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 163) | NC score | 0.067671 (rank : 100) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 376 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9EQZ7, Q8C433, Q8CCK2 | Gene names | Rims2, Rab3ip2, Rim2 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 2 (Rab3-interacting molecule 2) (RIM 2) (Rab3-interacting protein 2). | |||||
|
RP3A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 164) | NC score | 0.059396 (rank : 115) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9Y2J0, Q96AE0 | Gene names | RPH3A, KIAA0985 | |||
|
Domain Architecture |

|
|||||
| Description | Rabphilin-3A (Exophilin-1). | |||||
|
RP3A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 165) | NC score | 0.055123 (rank : 127) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 510 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P47708 | Gene names | Rph3a | |||
|
Domain Architecture |

|
|||||
| Description | Rabphilin-3A (Exophilin-1). | |||||
|
RPGR_HUMAN
|
||||||
| θ value | θ > 10 (rank : 166) | NC score | 0.083741 (rank : 81) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q92834, O00702, O00737, Q8N5T6, Q93039, Q9HD29, Q9UMR1 | Gene names | RPGR, RP3, XLRP3 | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator. | |||||
|
RPGR_MOUSE
|
||||||
| θ value | θ > 10 (rank : 167) | NC score | 0.080381 (rank : 86) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9R0X5, O88408, Q9CU92 | Gene names | Rpgr | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator (mRpgr). | |||||
|
SRGEF_HUMAN
|
||||||
| θ value | θ > 10 (rank : 168) | NC score | 0.086761 (rank : 76) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UGK8, Q9UGK9 | Gene names | SERGEF, DELGEF, GNEFR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Secretion-regulating guanine nucleotide exchange factor (Guanine nucleotide exchange factor-related protein) (Deafness locus-associated putative guanine nucleotide exchange factor) (DelGEF). | |||||
|
SRGEF_MOUSE
|
||||||
| θ value | θ > 10 (rank : 169) | NC score | 0.084679 (rank : 78) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q80YD6, Q9QXB7 | Gene names | Sergef, Delgef, Gnefr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Secretion-regulating guanine nucleotide exchange factor (Guanine nucleotide exchange factor-related protein) (Deafness locus-associated putative guanine nucleotide exchange factor) (DelGEF). | |||||
|
SYT10_HUMAN
|
||||||
| θ value | θ > 10 (rank : 170) | NC score | 0.058836 (rank : 118) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q6XYQ8 | Gene names | SYT10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptotagmin-10 (Synaptotagmin X) (SytX). | |||||
|
SYT10_MOUSE
|
||||||
| θ value | θ > 10 (rank : 171) | NC score | 0.057472 (rank : 121) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9R0N4 | Gene names | Syt10 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-10 (Synaptotagmin X) (SytX). | |||||
|
SYT1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 172) | NC score | 0.060334 (rank : 111) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P21579 | Gene names | SYT1, SYT | |||
|
Domain Architecture |
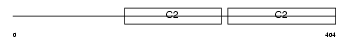
|
|||||
| Description | Synaptotagmin-1 (Synaptotagmin I) (SytI) (p65). | |||||
|
SYT1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 173) | NC score | 0.060333 (rank : 112) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P46096 | Gene names | Syt1 | |||
|
Domain Architecture |
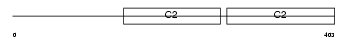
|
|||||
| Description | Synaptotagmin-1 (Synaptotagmin I) (SytI) (p65). | |||||
|
SYT2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 174) | NC score | 0.056188 (rank : 125) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8N9I0, Q496K5, Q8NBE5 | Gene names | SYT2 | |||
|
Domain Architecture |
|
|||||
| Description | Synaptotagmin-2 (Synaptotagmin II) (SytII). | |||||
|
SYT2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 175) | NC score | 0.056289 (rank : 124) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P46097, Q8R0E1 | Gene names | Syt2 | |||
|
Domain Architecture |
|
|||||
| Description | Synaptotagmin-2 (Synaptotagmin II) (SytII). | |||||
|
SYT3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 176) | NC score | 0.050262 (rank : 141) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9BQG1, Q8N5Z1, Q8N640 | Gene names | SYT3 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-3 (Synaptotagmin III) (SytIII). | |||||
|
SYT3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 177) | NC score | 0.050918 (rank : 137) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O35681, P97791, Q80WV1 | Gene names | Syt3 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-3 (Synaptotagmin III) (SytIII). | |||||
|
SYT5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 178) | NC score | 0.062350 (rank : 107) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O00445, Q86X72 | Gene names | SYT5 | |||
|
Domain Architecture |
|
|||||
| Description | Synaptotagmin-5 (Synaptotagmin V) (SytV). | |||||
|
SYT5_MOUSE
|
||||||
| θ value | θ > 10 (rank : 179) | NC score | 0.056735 (rank : 122) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9R0N5 | Gene names | Syt5, Syt9 | |||
|
Domain Architecture |
|
|||||
| Description | Synaptotagmin-5 (Synaptotagmin V) (SytV) (Synaptotagmin IX). | |||||
|
SYT6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 180) | NC score | 0.050264 (rank : 140) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q5T7P8 | Gene names | SYT6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptotagmin-6 (Synaptotagmin VI) (SytVI). | |||||
|
SYT6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 181) | NC score | 0.051813 (rank : 134) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9R0N8 | Gene names | Syt6 | |||
|
Domain Architecture |
|
|||||
| Description | Synaptotagmin-6 (Synaptotagmin VI) (SytVI). | |||||
|
SYT7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 182) | NC score | 0.059794 (rank : 114) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O43581 | Gene names | SYT7 | |||
|
Domain Architecture |
|
|||||
| Description | Synaptotagmin-7 (Synaptotagmin VII) (SytVII). | |||||
|
SYT7_MOUSE
|
||||||
| θ value | θ > 10 (rank : 183) | NC score | 0.058854 (rank : 117) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9R0N7 | Gene names | Syt7 | |||
|
Domain Architecture |
|
|||||
| Description | Synaptotagmin-7 (Synaptotagmin VII) (SytVII). | |||||
|
SYT9_HUMAN
|
||||||
| θ value | θ > 10 (rank : 184) | NC score | 0.053341 (rank : 129) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q86SS6 | Gene names | SYT9 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-9 (Synaptotagmin IX) (SytIX). | |||||
|
SYT9_MOUSE
|
||||||
| θ value | θ > 10 (rank : 185) | NC score | 0.054535 (rank : 128) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9R0N9, Q8C280 | Gene names | Syt9, Syt5 | |||
|
Domain Architecture |
|
|||||
| Description | Synaptotagmin-9 (Synaptotagmin IX) (SytIX) (Synaptotagmin V). | |||||
|
SYTL2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 186) | NC score | 0.050050 (rank : 142) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 253 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q99N50, Q8BT37, Q99J89, Q99J90, Q99N51, Q99N52, Q99N55, Q99N56 | Gene names | Sytl2, Slp2 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 2 (Exophilin-4). | |||||
|
TOLIP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 187) | NC score | 0.064334 (rank : 103) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9H0E2, Q9H9E6, Q9UJ69 | Gene names | TOLLIP | |||
|
Domain Architecture |
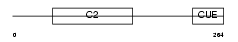
|
|||||
| Description | Toll-interacting protein. | |||||
|
TOLIP_MOUSE
|
||||||
| θ value | θ > 10 (rank : 188) | NC score | 0.069350 (rank : 98) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9QZ06, Q543N1, Q9D5P5 | Gene names | Tollip | |||
|
Domain Architecture |
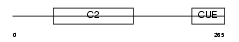
|
|||||
| Description | Toll-interacting protein. | |||||
|
UN13A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 189) | NC score | 0.081656 (rank : 84) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9UPW8 | Gene names | UNC13A, KIAA1032 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Unc-13 homolog A (Munc13-1). | |||||
|
UN13B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 190) | NC score | 0.084320 (rank : 80) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O14795 | Gene names | UNC13B, UNC13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Unc-13 homolog B (Munc13-2) (munc13). | |||||
|
UN13B_MOUSE
|
||||||
| θ value | θ > 10 (rank : 191) | NC score | 0.084550 (rank : 79) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9Z1N9 | Gene names | Unc13b, Unc13a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Unc-13 homolog B (Munc13-2) (munc13). | |||||
|
UN13D_HUMAN
|
||||||
| θ value | θ > 10 (rank : 192) | NC score | 0.060982 (rank : 110) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q70J99, Q9H7K5 | Gene names | UNC13D | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Unc-13 homolog D (Munc13-4). | |||||
|
WBS16_HUMAN
|
||||||
| θ value | θ > 10 (rank : 193) | NC score | 0.076668 (rank : 90) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96I51, Q548B1, Q9H0G7 | Gene names | WBSCR16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Williams-Beuren syndrome chromosome region 16 protein (RCC1-like G exchanging factor-like protein). | |||||
|
WBS16_MOUSE
|
||||||
| θ value | θ > 10 (rank : 194) | NC score | 0.078301 (rank : 88) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9CYF5 | Gene names | Wbscr16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Williams-Beuren syndrome chromosome region 16 protein homolog. | |||||
|
WWC3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 195) | NC score | 0.051282 (rank : 135) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9ULE0, Q659C1, Q9BTQ1 | Gene names | WWC3, KIAA1280 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WWC family member 3. | |||||
|
WWP1_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 138 | |
| SwissProt Accessions | Q8BZZ3, Q8BIV9, Q8VDP8 | Gene names | Wwp1 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP1 (EC 6.3.2.-) (WW domain- containing protein 1). | |||||
|
ITCH_MOUSE
|
||||||
| NC score | 0.991751 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | Q8C863, O54971 | Gene names | Itch | |||
|
Domain Architecture |

|
|||||
| Description | Itchy E3 ubiquitin protein ligase (EC 6.3.2.-). | |||||
|
WWP2_MOUSE
|
||||||
| NC score | 0.991677 (rank : 3) | θ value | 0 (rank : 6) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | Q9DBH0, Q8BTG4, Q923F6 | Gene names | Wwp2 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP2 (EC 6.3.2.-) (WW domain- containing protein 2). | |||||
|
WWP2_HUMAN
|
||||||
| NC score | 0.989701 (rank : 4) | θ value | 0 (rank : 5) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | O00308, Q96CZ2, Q9BWN6 | Gene names | WWP2 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP2 (EC 6.3.2.-) (WW domain- containing protein 2) (Atrophin-1-interacting protein 2) (AIP2). | |||||
|
WWP1_HUMAN
|
||||||
| NC score | 0.988077 (rank : 5) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 100 | |
| SwissProt Accessions | Q9H0M0, O00307, Q96BP4 | Gene names | WWP1 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP1 (EC 6.3.2.-) (WW domain- containing protein 1) (Atropin-1-interacting protein 5) (AIP5). | |||||
|
ITCH_HUMAN
|
||||||
| NC score | 0.984252 (rank : 6) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | Q96J02, O43584, Q5TEL0, Q96F66, Q9BY75, Q9H451, Q9H4U5 | Gene names | ITCH | |||
|
Domain Architecture |

|
|||||
| Description | Itchy homolog E3 ubiquitin protein ligase (EC 6.3.2.-) (Itch) (Atrophin-1-interacting protein 4) (AIP4) (NFE2-associated polypeptide 1) (NAPP1). | |||||
|
SMUF2_HUMAN
|
||||||
| NC score | 0.960771 (rank : 7) | θ value | 3.88293e-136 (rank : 11) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | Q9HAU4, Q9H260 | Gene names | SMURF2 | |||
|
Domain Architecture |

|
|||||
| Description | Smad ubiquitination regulatory factor 2 (EC 6.3.2.-) (Ubiquitin-- protein ligase SMURF2) (Smad-specific E3 ubiquitin ligase 2) (hSMURF2). | |||||
|
SMUF1_MOUSE
|
||||||
| NC score | 0.953280 (rank : 8) | θ value | 7.33985e-127 (rank : 12) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9CUN6, Q3U412, Q8K300 | Gene names | Smurf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Smad ubiquitination regulatory factor 1 (EC 6.3.2.-) (Ubiquitin-- protein ligase SMURF1) (Smad-specific E3 ubiquitin ligase 1). | |||||
|
SMUF1_HUMAN
|
||||||
| NC score | 0.950211 (rank : 9) | θ value | 2.13557e-126 (rank : 13) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9HCE7, O75853, Q9UJT8 | Gene names | SMURF1, KIAA1625 | |||
|
Domain Architecture |

|
|||||
| Description | Smad ubiquitination regulatory factor 1 (EC 6.3.2.-) (Ubiquitin-- protein ligase SMURF1) (Smad-specific E3 ubiquitin ligase 1) (hSMURF1). | |||||
|
NEDD4_MOUSE
|
||||||
| NC score | 0.918255 (rank : 10) | θ value | 1.74296e-136 (rank : 10) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 90 | |
| SwissProt Accessions | P46935, O08758, Q3UZI2, Q8BGB3 | Gene names | Nedd4, Kiaa0093, Nedd-4, Nedd4a | |||
|
Domain Architecture |

|
|||||
| Description | E3 ubiquitin-protein ligase NEDD4 (EC 6.3.2.-) (Neural precursor cell expressed developmentally down-regulated protein 4). | |||||
|
NEDD4_HUMAN
|
||||||
| NC score | 0.914846 (rank : 11) | θ value | 3.87392e-144 (rank : 7) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 86 | |
| SwissProt Accessions | P46934 | Gene names | NEDD4, KIAA0093 | |||
|
Domain Architecture |

|
|||||
| Description | E3 ubiquitin-protein ligase NEDD4 (EC 6.3.2.-). | |||||
|
NED4L_MOUSE
|
||||||
| NC score | 0.900556 (rank : 12) | θ value | 8.0776e-142 (rank : 9) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 90 | |
| SwissProt Accessions | Q8CFI0, Q8BRT9, Q8BS42, Q99PK2 | Gene names | Nedd4l, Kiaa0439, Nedd4b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin-protein ligase NEDD4-like protein (EC 6.3.2.-) (Nedd4-2) (NEDD4.2). | |||||
|
NED4L_HUMAN
|
||||||
| NC score | 0.899753 (rank : 13) | θ value | 2.12569e-142 (rank : 8) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 91 | |
| SwissProt Accessions | Q96PU5, O43165, Q7Z5F1, Q7Z5F2, Q7Z5N3, Q8N5A7, Q8WUU9, Q9BW58, Q9H2W4, Q9NT88 | Gene names | NEDD4L, KIAA0439, NEDL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin-protein ligase NEDD4-like protein (EC 6.3.2.-) (Nedd4-2) (NEDD4.2). | |||||
|
HECD2_MOUSE
|
||||||
| NC score | 0.882143 (rank : 14) | θ value | 9.16162e-53 (rank : 16) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8CDU6, Q8CBQ9 | Gene names | Hectd2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase HECTD2 (HECT domain-containing protein 2). | |||||
|
HECD2_HUMAN
|
||||||
| NC score | 0.879269 (rank : 15) | θ value | 6.56568e-51 (rank : 18) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q5U5R9, Q5VZ97, Q5VZ99, Q8TCP5 | Gene names | HECTD2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase HECTD2 (HECT domain-containing protein 2). | |||||
|
UBE3A_HUMAN
|
||||||
| NC score | 0.874812 (rank : 16) | θ value | 1.32293e-51 (rank : 17) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q05086, P78355, Q93066, Q9UEP4, Q9UEP5, Q9UEP6, Q9UEP7, Q9UEP8, Q9UEP9 | Gene names | UBE3A, E6AP | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase E3A (EC 6.3.2.-) (E6AP ubiquitin-protein ligase) (Oncogenic protein-associated protein E6-AP) (Human papillomavirus E6-associated protein) (NY-REN-54 antigen). | |||||
|
K0317_HUMAN
|
||||||
| NC score | 0.874331 (rank : 17) | θ value | 1.08474e-45 (rank : 20) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O15033, Q7LDY1, Q8IYY9 | Gene names | KIAA0317 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0317. | |||||
|
K0317_MOUSE
|
||||||
| NC score | 0.873727 (rank : 18) | θ value | 4.122e-45 (rank : 21) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8CHG5, Q6P9Q1, Q80YC4, Q8C5W5 | Gene names | Kiaa0317 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0317. | |||||
|
UBE3A_MOUSE
|
||||||
| NC score | 0.868105 (rank : 19) | θ value | 6.35935e-46 (rank : 19) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O08759, P97482 | Gene names | Ube3a | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase E3A (EC 6.3.2.-) (Oncogenic protein- associated protein E6-AP). | |||||
|
HUWE1_MOUSE
|
||||||
| NC score | 0.867673 (rank : 20) | θ value | 7.93343e-89 (rank : 15) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q7TMY8, Q4G2Z1, Q5BMM7, Q6NS61, Q8BNJ7, Q8CFH2, Q8VD14, Q921M5, Q9R0P2 | Gene names | Huwe1, Kiaa0312, Ureb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (E3Histone). | |||||
|
HUWE1_HUMAN
|
||||||
| NC score | 0.866793 (rank : 21) | θ value | 7.93343e-89 (rank : 14) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q7Z6Z7, O15029, Q4G2Z2, Q5H961, Q6P4D0, Q8NG67, Q9BUI0, Q9HCJ4, Q9NSL6, Q9P0A9 | Gene names | HUWE1, KIAA0312, KIAA1578, UREB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (Mcl-1 ubiquitin ligase E3) (Mule) (ARF-binding protein 1) (ARF-BP1). | |||||
|
UBE3C_HUMAN
|
||||||
| NC score | 0.856585 (rank : 22) | θ value | 4.2656e-42 (rank : 22) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q15386, Q8TC15, Q96CR4, Q9UDU3 | Gene names | UBE3C, KIAA0010 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase E3C (EC 6.3.2.-). | |||||
|
UBE3C_MOUSE
|
||||||
| NC score | 0.854985 (rank : 23) | θ value | 3.61106e-41 (rank : 23) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q80U95, Q8BQZ6, Q8C7W6, Q8CDJ1, Q8VDL5 | Gene names | Ube3c, Kiaa0010 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase E3C (EC 6.3.2.-). | |||||
|
TRIPC_HUMAN
|
||||||
| NC score | 0.793240 (rank : 24) | θ value | 1.52067e-31 (rank : 26) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 319 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q14669, Q15644 | Gene names | TRIP12, KIAA0045 | |||
|
Domain Architecture |

|
|||||
| Description | Thyroid receptor-interacting protein 12 (TRIP12). | |||||
|
HERC5_HUMAN
|
||||||
| NC score | 0.728197 (rank : 25) | θ value | 7.54701e-31 (rank : 28) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UII4 | Gene names | HERC5, CEB1, CEBP1 | |||
|
Domain Architecture |

|
|||||
| Description | HECT domain and RCC1-like domain-containing protein 5 (Cyclin-E- binding protein 1). | |||||
|
HERC3_HUMAN
|
||||||
| NC score | 0.724747 (rank : 26) | θ value | 6.81017e-40 (rank : 24) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q15034 | Gene names | HERC3, KIAA0032 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT domain and RCC1-like domain protein 3. | |||||
|
HECD3_MOUSE
|
||||||
| NC score | 0.689897 (rank : 27) | θ value | 9.90251e-15 (rank : 31) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q3U487, Q3TN76, Q641P3, Q8BQ74, Q8R1L6 | Gene names | Hectd3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase HECTD3 (HECT domain-containing protein 3). | |||||
|
HECD3_HUMAN
|
||||||
| NC score | 0.684451 (rank : 28) | θ value | 4.91457e-14 (rank : 32) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q5T447, Q5T448, Q9H783 | Gene names | HECTD3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase HECTD3 (HECT domain-containing protein 3). | |||||
|
EDD1_MOUSE
|
||||||
| NC score | 0.649368 (rank : 29) | θ value | 1.09485e-13 (rank : 34) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q80TP3, Q698K9, Q6PEQ8, Q6PFQ9, Q80VL4, Q810V6, Q9CXE9 | Gene names | Edd1, Edd, Kiaa0896 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase EDD1 (EC 6.3.2.-) (Hyperplastic discs protein homolog). | |||||
|
EDD1_HUMAN
|
||||||
| NC score | 0.647820 (rank : 30) | θ value | 1.09485e-13 (rank : 33) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O95071, O94970, Q9NPL3 | Gene names | EDD1, EDD, HYD, KIAA0896 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase EDD1 (EC 6.3.2.-) (Hyperplastic discs protein homolog) (hHYD) (Progestin-induced protein). | |||||
|
HERC2_MOUSE
|
||||||
| NC score | 0.620034 (rank : 31) | θ value | 5.22648e-32 (rank : 25) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q4U2R1, O88473, Q3TRJ8, Q3TS47, Q3TST2, Q3UFQ6, Q3URH7, Q5DU32, Q7TPR5, Q80VV7, Q9QYT1, Q9Z168, Q9Z171 | Gene names | Herc2, jdf2, Kiaa0393, rjs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT domain and RCC1-like domain-containing protein 2. | |||||
|
HERC2_HUMAN
|
||||||
| NC score | 0.615863 (rank : 32) | θ value | 2.59387e-31 (rank : 27) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O95714, Q86SV7, Q86SV8, Q86SV9, Q86YY3, Q86YY4, Q86YY5, Q86YY6, Q86YY7, Q86YY8, Q86YY9, Q86YZ0, Q86YZ1 | Gene names | HERC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT domain and RCC1-like domain-containing protein 2. | |||||
|
YAP1_MOUSE
|
||||||
| NC score | 0.563416 (rank : 33) | θ value | 2.60593e-15 (rank : 30) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P46938, Q52KJ5 | Gene names | Yap1, Yap, Yap65 | |||
|
Domain Architecture |

|
|||||
| Description | 65 kDa Yes-associated protein (YAP65). | |||||
|
HECD1_HUMAN
|
||||||
| NC score | 0.563312 (rank : 34) | θ value | 8.08199e-25 (rank : 29) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 371 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9ULT8, Q6P445, Q86VJ1, Q96F34, Q9UFZ7 | Gene names | HECTD1, KIAA1131 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin-protein ligase HECTD1 (HECT domain-containing protein 1) (E3 ligase for inhibin receptor) (EULIR). | |||||
|
SAV1_HUMAN
|
||||||
| NC score | 0.534162 (rank : 35) | θ value | 1.74796e-11 (rank : 38) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9H4B6, Q6IA58, Q9H949, Q9HAK9 | Gene names | SAV1, WW45 | |||
|
Domain Architecture |

|
|||||
| Description | Salvador homolog 1 protein (45 kDa WW domain protein) (hWW45). | |||||
|
SAV1_MOUSE
|
||||||
| NC score | 0.533664 (rank : 36) | θ value | 1.74796e-11 (rank : 39) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8VEB2, Q9D9N9, Q9ER46 | Gene names | Sav1, Ww45, Wwp3 | |||
|
Domain Architecture |

|
|||||
| Description | Salvador homolog 1 protein (45 kDa WW domain protein) (mWW45). | |||||
|
PINL_HUMAN
|
||||||
| NC score | 0.450748 (rank : 37) | θ value | 2.44474e-05 (rank : 52) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O15428 | Gene names | PIN1L | |||
|
Domain Architecture |

|
|||||
| Description | PIN1-like protein. | |||||
|
WWC1_HUMAN
|
||||||
| NC score | 0.445986 (rank : 38) | θ value | 9.26847e-13 (rank : 35) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8IX03, O94946, Q6MZX4, Q6Y2F8, Q7Z4G8, Q8WVM4, Q9BT29 | Gene names | WWC1, KIAA0869 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing protein 1 (Kidney and brain protein) (KIBRA) (HBeAg-binding protein 3). | |||||
|
WWC1_MOUSE
|
||||||
| NC score | 0.445407 (rank : 39) | θ value | 9.26847e-13 (rank : 36) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q5SXA9, Q571D0, Q8K1Y3, Q8VD17, Q922W3 | Gene names | Wwc1, Kiaa0869 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing protein 1 (Kidney and brain protein) (KIBRA). | |||||
|
WWOX_HUMAN
|
||||||
| NC score | 0.396980 (rank : 40) | θ value | 1.74796e-11 (rank : 40) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9NZC7, Q5MYT5, Q96KM3, Q96RF2, Q9BTT8, Q9NPC9, Q9NRF4, Q9NRF5, Q9NRF6, Q9NRK1, Q9NZC5 | Gene names | WWOX, FOR, WOX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing oxidoreductase (EC 1.1.1.-) (Fragile site FRA16D oxidoreductase). | |||||
|
WWOX_MOUSE
|
||||||
| NC score | 0.393572 (rank : 41) | θ value | 1.74796e-11 (rank : 41) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q91WL8, Q8C8J6, Q920Y2, Q9D2B3, Q9D339, Q9JLF5 | Gene names | Wwox, Wox1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing oxidoreductase (EC 1.1.1.-). | |||||
|
WWTR1_MOUSE
|
||||||
| NC score | 0.387002 (rank : 42) | θ value | 7.1131e-05 (rank : 55) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9EPK5, Q99KI4 | Gene names | Wwtr1, Taz | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing transcription regulator protein 1 (Transcriptional coactivator with PDZ-binding motif). | |||||
|
PIN1_MOUSE
|
||||||
| NC score | 0.386618 (rank : 43) | θ value | 0.000270298 (rank : 57) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9QUR7, Q543B3 | Gene names | Pin1 | |||
|
Domain Architecture |

|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase NIMA-interacting 1 (EC 5.2.1.8) (Rotamase Pin1) (PPIase Pin1). | |||||
|
PIN1_HUMAN
|
||||||
| NC score | 0.384680 (rank : 44) | θ value | 0.000270298 (rank : 56) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q13526 | Gene names | PIN1 | |||
|
Domain Architecture |

|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase NIMA-interacting 1 (EC 5.2.1.8) (Rotamase Pin1) (PPIase Pin1). | |||||
|
YAP1_HUMAN
|
||||||
| NC score | 0.384374 (rank : 45) | θ value | 8.40245e-06 (rank : 49) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P46937 | Gene names | YAP1, YAP65 | |||
|
Domain Architecture |

|
|||||
| Description | 65 kDa Yes-associated protein (YAP65). | |||||
|
WWTR1_HUMAN
|
||||||
| NC score | 0.383641 (rank : 46) | θ value | 7.1131e-05 (rank : 54) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9GZV5, Q8N3P2, Q9Y3W6 | Gene names | WWTR1, TAZ | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing transcription regulator protein 1 (Transcriptional coactivator with PDZ-binding motif). | |||||
|
BAG3_HUMAN
|
||||||
| NC score | 0.356893 (rank : 47) | θ value | 2.88788e-06 (rank : 48) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 314 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O95817, Q9NT20, Q9P120 | Gene names | BAG3, BIS | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 3 (BCL-2-binding athanogene- 3) (BAG-3) (Bcl-2-binding protein Bis) (Docking protein CAIR-1). | |||||
|
MAGI2_MOUSE
|
||||||
| NC score | 0.325348 (rank : 48) | θ value | 2.0648e-12 (rank : 37) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 370 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9WVQ1, Q3UH81, Q6GT88, Q8BYT1, Q8CA85 | Gene names | Magi2, Acvrinp1, Aip1, Arip1 | |||
|
Domain Architecture |

|
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 2 (Membrane-associated guanylate kinase inverted 2) (MAGI-2) (Atrophin-1-interacting protein 1) (Activin receptor-interacting protein 1) (Acvrip1). | |||||
|
BAG3_MOUSE
|
||||||
| NC score | 0.310464 (rank : 49) | θ value | 2.44474e-05 (rank : 51) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9JLV1, Q9JJC7 | Gene names | Bag3, Bis | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 3 (BCL-2-binding athanogene- 3) (BAG-3) (Bcl-2-binding protein Bis). | |||||
|
MAGI2_HUMAN
|
||||||
| NC score | 0.309937 (rank : 50) | θ value | 4.30538e-10 (rank : 44) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 373 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q86UL8, O60434, O60510, Q86UI7, Q9NP44, Q9UDQ5, Q9UDU1 | Gene names | MAGI2, ACVRINP1, AIP1, KIAA0705 | |||
|
Domain Architecture |

|
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 2 (Membrane-associated guanylate kinase inverted 2) (MAGI-2) (Atrophin-1-interacting protein 1) (Atrophin-1-interacting protein A). | |||||
|
MAGI1_HUMAN
|
||||||
| NC score | 0.304720 (rank : 51) | θ value | 1.9326e-10 (rank : 43) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q96QZ7, O00309, O43863, O75085, Q96QZ8, Q96QZ9 | Gene names | MAGI1, BAIAP1, BAP1, TNRC19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 1 (BAI1-associated protein 1) (BAP-1) (Membrane-associated guanylate kinase inverted 1) (MAGI-1) (Atrophin-1-interacting protein 3) (AIP3) (WW domain-containing protein 3) (WWP3) (Trinucleotide repeat-containing gene 19 protein). | |||||
|
MAGI1_MOUSE
|
||||||
| NC score | 0.303524 (rank : 52) | θ value | 1.47974e-10 (rank : 42) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q6RHR9, O54893, O54894, O54895 | Gene names | Magi1, Baiap1, Bap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 1 (BAI1-associated protein 1) (BAP-1) (Membrane-associated guanylate kinase inverted 1) (MAGI-1). | |||||
|
PKHA5_HUMAN
|
||||||
| NC score | 0.286133 (rank : 53) | θ value | 3.64472e-09 (rank : 45) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9HAU0, Q8N3K6, Q96DY9, Q9BVR4, Q9C0H7, Q9H924, Q9NVK8 | Gene names | PLEKHA5, KIAA1686, PEPP2 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 5 (Phosphoinositol 3-phosphate-binding protein 2) (PEPP-2). | |||||
|
GAS7_HUMAN
|
||||||
| NC score | 0.272592 (rank : 54) | θ value | 3.19293e-05 (rank : 53) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O60861, O43144 | Gene names | GAS7, KIAA0394 | |||
|
Domain Architecture |

|
|||||
| Description | Growth-arrest-specific protein 7 (GAS-7). | |||||
|
GAS7_MOUSE
|
||||||
| NC score | 0.271894 (rank : 55) | θ value | 1.43324e-05 (rank : 50) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q60780, Q9QY25, Q9QY26, Q9QY34 | Gene names | Gas7 | |||
|
Domain Architecture |

|
|||||
| Description | Growth-arrest-specific protein 7 (GAS-7). | |||||
|
PRP40_HUMAN
|
||||||
| NC score | 0.231679 (rank : 56) | θ value | 7.59969e-07 (rank : 46) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 442 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O75400, O43856, O75404, Q8TBQ1, Q9H782, Q9NWU9, Q9P0Q2, Q9Y5A8 | Gene names | PRPF40A, FLAF1, FNBP3, HYPA | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Huntingtin yeast partner A) (Huntingtin-interacting protein HYPA/FBP11) (Fas ligand-associated factor 1) (NY-REN-6 antigen). | |||||
|
PRP40_MOUSE
|
||||||
| NC score | 0.230368 (rank : 57) | θ value | 7.59969e-07 (rank : 47) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9R1C7, Q61049, Q8BQ76, Q8BRW4, Q8C2U1 | Gene names | Prpf40a, Fbp11, Fnbp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Formin-binding protein 11) (FBP-11). | |||||
|
STXB4_MOUSE
|
||||||
| NC score | 0.178963 (rank : 58) | θ value | 0.00390308 (rank : 61) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 629 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9WV89, Q5SUA6, Q5SUA8, Q5SUA9, Q8CFL1 | Gene names | Stxbp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntaxin-binding protein 4 (Syntaxin 4-interacting protein) (Synip). | |||||
|
STXB4_HUMAN
|
||||||
| NC score | 0.178053 (rank : 59) | θ value | 0.00390308 (rank : 60) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 616 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q6ZWJ1, Q8IVZ5 | Gene names | STXBP4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntaxin-binding protein 4 (Syntaxin 4-interacting protein) (Synip). | |||||
|
APBB2_HUMAN
|
||||||
| NC score | 0.171308 (rank : 60) | θ value | 0.0148317 (rank : 68) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q92870, Q8IUI6 | Gene names | APBB2, FE65L, FE65L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 2 (Fe65-like protein). | |||||
|
APBB2_MOUSE
|
||||||
| NC score | 0.161005 (rank : 61) | θ value | 0.0113563 (rank : 66) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9DBR4, Q6DFX8 | Gene names | Apbb2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 2. | |||||
|
SETD2_HUMAN
|
||||||
| NC score | 0.139144 (rank : 62) | θ value | 0.00869519 (rank : 65) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q9BYW2, O75397, O75405, Q17RW8, Q5BKS9, Q5QGN2, Q69YI5, Q6IN64, Q6ZN53, Q6ZS25, Q8N3R0, Q8TCN0, Q9C0D1, Q9H696, Q9NZW9 | Gene names | SETD2, HIF1, HYPB, KIAA1732, SET2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36-specific (EC 2.1.1.43) (SET domain-containing protein 2) (hSET2) (Huntingtin- interacting protein HYPB) (Huntingtin-interacting protein 1) (HIF-1) (p231HBP). | |||||
|
APBB3_HUMAN
|
||||||
| NC score | 0.128743 (rank : 63) | θ value | 0.125558 (rank : 77) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O95704, Q9NYX6, Q9NYX7, Q9NYX8 | Gene names | APBB3, FE65L2 | |||
|
Domain Architecture |

|
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 3 (Fe65-like protein 2) (Fe65L2). | |||||
|
APBB3_MOUSE
|
||||||
| NC score | 0.127837 (rank : 64) | θ value | 0.163984 (rank : 78) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8R1C9 | Gene names | Apbb3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 3. | |||||
|
RHG12_MOUSE
|
||||||
| NC score | 0.125458 (rank : 65) | θ value | 0.00102713 (rank : 59) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8C0D4, Q8BVP8 | Gene names | Arhgap12 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 12. | |||||
|
WBP4_MOUSE
|
||||||
| NC score | 0.119079 (rank : 66) | θ value | 5.27518 (rank : 116) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q61048, Q8K1Z9, Q9CS45 | Gene names | Wbp4, Fbp21, Fnbp21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 4 (WBP-4) (Formin-binding protein 21). | |||||
|
RHG12_HUMAN
|
||||||
| NC score | 0.111169 (rank : 67) | θ value | 0.000602161 (rank : 58) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8IWW6, Q86UB3, Q8IWW7, Q8N3L1, Q9NT76 | Gene names | ARHGAP12 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 12. | |||||
|
DRP2_HUMAN
|
||||||
| NC score | 0.105234 (rank : 68) | θ value | 0.0961366 (rank : 75) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q13474 | Gene names | DRP2 | |||
|
Domain Architecture |

|
|||||
| Description | Dystrophin-related protein 2. | |||||
|
FA62A_MOUSE
|
||||||
| NC score | 0.101037 (rank : 69) | θ value | 5.27518 (rank : 112) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q3U7R1, Q8C8R1, Q91X62, Q9CVH0, Q9Z1X5, Q9Z1X6 | Gene names | Fam62a, Mbc2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM62A (Membrane-bound C2 domain-containing protein). | |||||
|
DGCR8_HUMAN
|
||||||
| NC score | 0.093082 (rank : 70) | θ value | 0.0563607 (rank : 73) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8WYQ5, Q6DCB2, Q6MZE9, Q6Y2L0, Q96G39, Q96GP8, Q9H6L8, Q9H6T7, Q9NRW2 | Gene names | DGCR8, C22orf12, DGCRK6 | |||
|
Domain Architecture |

|
|||||
| Description | DGCR8 protein (DiGeorge syndrome critical region 8). | |||||
|
DGCR8_MOUSE
|
||||||
| NC score | 0.091070 (rank : 71) | θ value | 0.0563607 (rank : 74) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9EQM6 | Gene names | Dgcr8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DGCR8 protein (DiGeorge syndrome critical region 8 homolog) (Gy1). | |||||
|
TCRG1_HUMAN
|
||||||
| NC score | 0.090370 (rank : 72) | θ value | 0.0148317 (rank : 69) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 929 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O14776 | Gene names | TCERG1, CA150, TAF2S | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150). | |||||
|
APBB1_HUMAN
|
||||||
| NC score | 0.090203 (rank : 73) | θ value | 1.06291 (rank : 85) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O00213, Q96A93 | Gene names | APBB1, FE65 | |||
|
Domain Architecture |

|
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 1 (Fe65 protein). | |||||
|
TCRG1_MOUSE
|
||||||
| NC score | 0.090031 (rank : 74) | θ value | 0.0113563 (rank : 67) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q8CGF7, Q61051, Q8C490, Q8CHT8, Q9R0R5 | Gene names | Tcerg1, Taf2s | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150) (p144) (Formin- binding protein 28) (FBP 28). | |||||
|
APBB1_MOUSE
|
||||||
| NC score | 0.089608 (rank : 75) | θ value | 1.06291 (rank : 86) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9QXJ1, O08642, Q3TPU0, Q8BNF4, Q8BSR9 | Gene names | Apbb1, Fe65 | |||
|
Domain Architecture |

|
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 1 (Fe65 protein). | |||||
|
SRGEF_HUMAN
|
||||||
| NC score | 0.086761 (rank : 76) | θ value | θ > 10 (rank : 168) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UGK8, Q9UGK9 | Gene names | SERGEF, DELGEF, GNEFR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Secretion-regulating guanine nucleotide exchange factor (Guanine nucleotide exchange factor-related protein) (Deafness locus-associated putative guanine nucleotide exchange factor) (DelGEF). | |||||
|
FA62A_HUMAN
|
||||||
| NC score | 0.084881 (rank : 77) | θ value | θ > 10 (rank : 148) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9BSJ8, O94848, Q6PJN4, Q9H6J1, Q9H6W2, Q9Y416 | Gene names | FAM62A, KIAA0747, MBC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM62A (Membrane-bound C2 domain-containing protein). | |||||
|
SRGEF_MOUSE
|
||||||
| NC score | 0.084679 (rank : 78) | θ value | θ > 10 (rank : 169) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q80YD6, Q9QXB7 | Gene names | Sergef, Delgef, Gnefr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Secretion-regulating guanine nucleotide exchange factor (Guanine nucleotide exchange factor-related protein) (Deafness locus-associated putative guanine nucleotide exchange factor) (DelGEF). | |||||
|
UN13B_MOUSE
|
||||||
| NC score | 0.084550 (rank : 79) | θ value | θ > 10 (rank : 191) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9Z1N9 | Gene names | Unc13b, Unc13a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Unc-13 homolog B (Munc13-2) (munc13). | |||||
|
UN13B_HUMAN
|
||||||
| NC score | 0.084320 (rank : 80) | θ value | θ > 10 (rank : 190) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O14795 | Gene names | UNC13B, UNC13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Unc-13 homolog B (Munc13-2) (munc13). | |||||
|
RPGR_HUMAN
|
||||||
| NC score | 0.083741 (rank : 81) | θ value | θ > 10 (rank : 166) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q92834, O00702, O00737, Q8N5T6, Q93039, Q9HD29, Q9UMR1 | Gene names | RPGR, RP3, XLRP3 | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator. | |||||
|
RCC2_HUMAN
|
||||||
| NC score | 0.083536 (rank : 82) | θ value | θ > 10 (rank : 160) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9P258, Q8IVL9, Q9BSN6, Q9NPV8 | Gene names | RCC2, KIAA1470, TD60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein RCC2 (Telophase disk protein of 60 kDa) (RCC1-like protein TD- 60). | |||||
|
RCC2_MOUSE
|
||||||
| NC score | 0.083060 (rank : 83) | θ value | θ > 10 (rank : 161) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BK67, Q6ZPQ0 | Gene names | Rcc2, Kiaa1470 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein RCC2. | |||||
|
UN13A_HUMAN
|
||||||
| NC score | 0.081656 (rank : 84) | θ value | θ > 10 (rank : 189) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9UPW8 | Gene names | UNC13A, KIAA1032 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Unc-13 homolog A (Munc13-1). | |||||
|
RCC1_HUMAN
|
||||||
| NC score | 0.081059 (rank : 85) | θ value | θ > 10 (rank : 158) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P18754 | Gene names | RCC1, CHC1 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of chromosome condensation (Chromosome condensation protein 1) (Cell cycle regulatory protein). | |||||
|
RPGR_MOUSE
|
||||||
| NC score | 0.080381 (rank : 86) | θ value | θ > 10 (rank : 167) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9R0X5, O88408, Q9CU92 | Gene names | Rpgr | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator (mRpgr). | |||||
|
RCC1_MOUSE
|
||||||
| NC score | 0.078949 (rank : 87) | θ value | θ > 10 (rank : 159) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8VE37, Q3UDB6 | Gene names | Rcc1, Chc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of chromosome condensation (Chromosome condensation protein 1). | |||||
|
WBS16_MOUSE
|
||||||
| NC score | 0.078301 (rank : 88) | θ value | θ > 10 (rank : 194) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9CYF5 | Gene names | Wbscr16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Williams-Beuren syndrome chromosome region 16 protein homolog. | |||||
|
RASL1_HUMAN
|
||||||
| NC score | 0.077622 (rank : 89) | θ value | θ > 10 (rank : 151) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O95294, Q52M03, Q96CC7 | Gene names | RASAL1, RASAL | |||
|
Domain Architecture |

|
|||||
| Description | RasGAP-activating-like protein 1. | |||||
|
WBS16_HUMAN
|
||||||
| NC score | 0.076668 (rank : 90) | θ value | θ > 10 (rank : 193) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96I51, Q548B1, Q9H0G7 | Gene names | WBSCR16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Williams-Beuren syndrome chromosome region 16 protein (RCC1-like G exchanging factor-like protein). | |||||
|
DOC2A_MOUSE
|
||||||
| NC score | 0.073844 (rank : 91) | θ value | θ > 10 (rank : 144) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q7TNF0 | Gene names | Doc2a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double C2-like domain-containing protein alpha (Doc2-alpha). | |||||
|
RASL1_MOUSE
|
||||||
| NC score | 0.073191 (rank : 92) | θ value | θ > 10 (rank : 152) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9Z268 | Gene names | Rasal1, Rasal | |||
|
Domain Architecture |

|
|||||
| Description | RasGAP-activating-like protein 1. | |||||
|
MYOF_HUMAN
|
||||||
| NC score | 0.073017 (rank : 93) | θ value | 0.00869519 (rank : 64) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9NZM1, Q9HBU3, Q9NZM0, Q9ULL3, Q9Y4U4 | Gene names | FER1L3, KIAA1207, MYOF | |||
|
Domain Architecture |

|
|||||
| Description | Myoferlin (Fer-1-like protein 3). | |||||
|
DOC2B_HUMAN
|
||||||
| NC score | 0.071328 (rank : 94) | θ value | θ > 10 (rank : 145) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q14184 | Gene names | DOC2B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double C2-like domain-containing protein beta (Doc2-beta). | |||||
|
RASL2_HUMAN
|
||||||
| NC score | 0.070452 (rank : 95) | θ value | θ > 10 (rank : 153) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O43374, O60286, Q86UW3, Q96QU0 | Gene names | RASA4, CAPRI, GAPL, KIAA0538 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 4 (RasGAP-activating-like protein 2) (Calcium-promoted Ras inactivator). | |||||
|
DOC2B_MOUSE
|
||||||
| NC score | 0.070229 (rank : 96) | θ value | θ > 10 (rank : 146) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P70169, Q6NXK3 | Gene names | Doc2b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double C2-like domain-containing protein beta (Doc2-beta). | |||||
|
DOC2A_HUMAN
|
||||||
| NC score | 0.069665 (rank : 97) | θ value | θ > 10 (rank : 143) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q14183, Q6P4G4, Q7Z5G0, Q8IVX0 | Gene names | DOC2A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double C2-like domain-containing protein alpha (Doc2-alpha) (Doc2). | |||||
|
TOLIP_MOUSE
|
||||||
| NC score | 0.069350 (rank : 98) | θ value | θ > 10 (rank : 188) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9QZ06, Q543N1, Q9D5P5 | Gene names | Tollip | |||
|
Domain Architecture |

|
|||||
| Description | Toll-interacting protein. | |||||
|
RIMS2_HUMAN
|
||||||
| NC score | 0.068285 (rank : 99) | θ value | θ > 10 (rank : 162) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UQ26, O43413, Q86XL9, Q8IWV9, Q8IWW1 | Gene names | RIMS2, KIAA0751, RIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 2 (Rab3-interacting molecule 2) (RIM 2). | |||||
|
RIMS2_MOUSE
|
||||||
| NC score | 0.067671 (rank : 100) | θ value | θ > 10 (rank : 163) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 376 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9EQZ7, Q8C433, Q8CCK2 | Gene names | Rims2, Rab3ip2, Rim2 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 2 (Rab3-interacting molecule 2) (RIM 2) (Rab3-interacting protein 2). | |||||
|
RCBT2_MOUSE
|
||||||
| NC score | 0.067083 (rank : 101) | θ value | θ > 10 (rank : 157) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99LJ7, Q3TUA3, Q8BMG2 | Gene names | Rcbtb2, Chc1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RCC1 and BTB domain-containing protein 2 (Regulator of chromosome condensation and BTB domain-containing protein 2) (Chromosome condensation 1-like). | |||||
|
RCBT2_HUMAN
|
||||||
| NC score | 0.066156 (rank : 102) | θ value | θ > 10 (rank : 156) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O95199 | Gene names | RCBTB2, CHC1L, RLG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RCC1 and BTB domain-containing protein 2 (Regulator of chromosome condensation and BTB domain-containing protein 2) (Chromosome condensation 1-like) (CHC1-L) (RCC1-like G exchanging factor). | |||||
|
TOLIP_HUMAN
|
||||||
| NC score | 0.064334 (rank : 103) | θ value | θ > 10 (rank : 187) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9H0E2, Q9H9E6, Q9UJ69 | Gene names | TOLLIP | |||
|
Domain Architecture |

|
|||||
| Description | Toll-interacting protein. | |||||
|
K1333_MOUSE
|
||||||
| NC score | 0.063216 (rank : 104) | θ value | θ > 10 (rank : 149) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5RJY2, Q6ZPT7, Q8BNA4 | Gene names | Kiaa1333 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1333 (EC 6.3.2.-). | |||||
|
PCLO_MOUSE
|
||||||
| NC score | 0.062943 (rank : 105) | θ value | 3.0926 (rank : 101) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
DYSF_HUMAN
|
||||||
| NC score | 0.062360 (rank : 106) | θ value | 1.06291 (rank : 87) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O75923, O75696, Q9UEN7 | Gene names | DYSF, FER1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Dysferlin (Dystrophy-associated fer-1-like protein) (Fer-1-like protein 1). | |||||
|
SYT5_HUMAN
|
||||||
| NC score | 0.062350 (rank : 107) | θ value | θ > 10 (rank : 178) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O00445, Q86X72 | Gene names | SYT5 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-5 (Synaptotagmin V) (SytV). | |||||
|
RCBT1_MOUSE
|
||||||
| NC score | 0.061754 (rank : 108) | θ value | θ > 10 (rank : 155) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6NXM2, Q8BTZ6, Q8BZV0 | Gene names | Rcbtb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RCC1 and BTB domain-containing protein 1 (Regulator of chromosome condensation and BTB domain-containing protein 1). | |||||
|
PCLO_HUMAN
|
||||||
| NC score | 0.061484 (rank : 109) | θ value | 1.81305 (rank : 94) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
UN13D_HUMAN
|
||||||
| NC score | 0.060982 (rank : 110) | θ value | θ > 10 (rank : 192) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q70J99, Q9H7K5 | Gene names | UNC13D | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Unc-13 homolog D (Munc13-4). | |||||
|
SYT1_HUMAN
|
||||||
| NC score | 0.060334 (rank : 111) | θ value | θ > 10 (rank : 172) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P21579 | Gene names | SYT1, SYT | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-1 (Synaptotagmin I) (SytI) (p65). | |||||
|
SYT1_MOUSE
|
||||||
| NC score | 0.060333 (rank : 112) | θ value | θ > 10 (rank : 173) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P46096 | Gene names | Syt1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-1 (Synaptotagmin I) (SytI) (p65). | |||||
|
RCBT1_HUMAN
|
||||||
| NC score | 0.060145 (rank : 113) | θ value | θ > 10 (rank : 154) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NDN9, Q969U9 | Gene names | RCBTB1, CLLD7, E4.5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RCC1 and BTB domain-containing protein 1 (Regulator of chromosome condensation and BTB domain-containing protein 1) (Chronic lymphocytic leukemia deletion region gene 7 protein) (CLL deletion region gene 7 protein). | |||||
|
SYT7_HUMAN
|
||||||
| NC score | 0.059794 (rank : 114) | θ value | θ > 10 (rank : 182) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O43581 | Gene names | SYT7 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-7 (Synaptotagmin VII) (SytVII). | |||||
|
RP3A_HUMAN
|
||||||
| NC score | 0.059396 (rank : 115) | θ value | θ > 10 (rank : 164) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9Y2J0, Q96AE0 | Gene names | RPH3A, KIAA0985 | |||
|
Domain Architecture |

|
|||||
| Description | Rabphilin-3A (Exophilin-1). | |||||
|
PA24B_MOUSE
|
||||||
| NC score | 0.059023 (rank : 116) | θ value | θ > 10 (rank : 150) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q4QQM1, Q80VV8, Q91W88 | Gene names | Pla2g4b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytosolic phospholipase A2 beta (EC 3.1.1.4) (cPLA2-beta) (Phospholipase A2 group IVB). | |||||
|
SYT7_MOUSE
|
||||||
| NC score | 0.058854 (rank : 117) | θ value | θ > 10 (rank : 183) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9R0N7 | Gene names | Syt7 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-7 (Synaptotagmin VII) (SytVII). | |||||
|
SYT10_HUMAN
|
||||||
| NC score | 0.058836 (rank : 118) | θ value | θ > 10 (rank : 170) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q6XYQ8 | Gene names | SYT10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptotagmin-10 (Synaptotagmin X) (SytX). | |||||
|
WBP4_HUMAN
|
||||||
| NC score | 0.058735 (rank : 119) | θ value | 5.27518 (rank : 115) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O75554 | Gene names | WBP4, FBP21, FNBP21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 4 (WBP-4) (Formin-binding protein 21). | |||||
|
DOC2G_MOUSE
|
||||||
| NC score | 0.058143 (rank : 120) | θ value | θ > 10 (rank : 147) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9ESN1, Q6P8R8 | Gene names | Doc2g | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Double C2-like domain-containing protein gamma (Doc2-gamma). | |||||
|
SYT10_MOUSE
|
||||||
| NC score | 0.057472 (rank : 121) | θ value | θ > 10 (rank : 171) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9R0N4 | Gene names | Syt10 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-10 (Synaptotagmin X) (SytX). | |||||
|
SYT5_MOUSE
|
||||||
| NC score | 0.056735 (rank : 122) | θ value | θ > 10 (rank : 179) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9R0N5 | Gene names | Syt5, Syt9 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-5 (Synaptotagmin V) (SytV) (Synaptotagmin IX). | |||||
|
CUL7_HUMAN
|
||||||
| NC score | 0.056596 (rank : 123) | θ value | θ > 10 (rank : 142) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q14999, Q5T654 | Gene names | CUL7, KIAA0076 | |||
|
Domain Architecture |

|
|||||
| Description | Cullin-7 (CUL-7). | |||||
|
SYT2_MOUSE
|
||||||
| NC score | 0.056289 (rank : 124) | θ value | θ > 10 (rank : 175) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P46097, Q8R0E1 | Gene names | Syt2 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-2 (Synaptotagmin II) (SytII). | |||||
|
SYT2_HUMAN
|
||||||
| NC score | 0.056188 (rank : 125) | θ value | θ > 10 (rank : 174) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8N9I0, Q496K5, Q8NBE5 | Gene names | SYT2 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-2 (Synaptotagmin II) (SytII). | |||||
|
DMD_HUMAN
|
||||||
| NC score | 0.055173 (rank : 126) | θ value | 0.00509761 (rank : 62) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P11532, Q02295, Q14169, Q14170, Q5JYU0, Q7KZ48 | Gene names | DMD | |||
|
Domain Architecture |

|
|||||
| Description | Dystrophin. | |||||
|
RP3A_MOUSE
|
||||||
| NC score | 0.055123 (rank : 127) | θ value | θ > 10 (rank : 165) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 510 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P47708 | Gene names | Rph3a | |||
|
Domain Architecture |

|
|||||
| Description | Rabphilin-3A (Exophilin-1). | |||||
|
SYT9_MOUSE
|
||||||
| NC score | 0.054535 (rank : 128) | θ value | θ > 10 (rank : 185) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9R0N9, Q8C280 | Gene names | Syt9, Syt5 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-9 (Synaptotagmin IX) (SytIX) (Synaptotagmin V). | |||||
|
SYT9_HUMAN
|
||||||
| NC score | 0.053341 (rank : 129) | θ value | θ > 10 (rank : 184) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q86SS6 | Gene names | SYT9 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-9 (Synaptotagmin IX) (SytIX). | |||||
|
ALS2_HUMAN
|
||||||
| NC score | 0.052745 (rank : 130) | θ value | θ > 10 (rank : 139) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96Q42, Q8N1E0, Q96PC4, Q96Q41, Q9H973, Q9HCK9 | Gene names | ALS2, KIAA1563 | |||
|
Domain Architecture |

|
|||||
| Description | Alsin (Amyotrophic lateral sclerosis protein 2). | |||||
|
OTOF_MOUSE
|
||||||
| NC score | 0.052683 (rank : 131) | θ value | 4.03905 (rank : 111) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9ESF1, Q9ESF2 | Gene names | Otof, Fer1l2 | |||
|
Domain Architecture |

|
|||||
| Description | Otoferlin (Fer-1-like protein 2). | |||||
|
DYSF_MOUSE
|
||||||
| NC score | 0.052556 (rank : 132) | θ value | 6.88961 (rank : 119) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9ESD7, Q9QXC0 | Gene names | Dysf, Fer1l1 | |||
|
Domain Architecture |

|
|||||
| Description | Dysferlin (Dystrophy-associated fer-1-like protein) (Fer-1-like protein 1). | |||||
|
BAIP3_MOUSE
|
||||||
| NC score | 0.052058 (rank : 133) | θ value | θ > 10 (rank : 141) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q80TT2, Q6RUT4 | Gene names | Baiap3, Gm937, Kiaa0734 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BAI1-associated protein 3 (BAP3). | |||||
|
SYT6_MOUSE
|
||||||
| NC score | 0.051813 (rank : 134) | θ value | θ > 10 (rank : 181) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9R0N8 | Gene names | Syt6 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-6 (Synaptotagmin VI) (SytVI). | |||||
|
WWC3_HUMAN
|
||||||
| NC score | 0.051282 (rank : 135) | θ value | θ > 10 (rank : 195) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9ULE0, Q659C1, Q9BTQ1 | Gene names | WWC3, KIAA1280 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WWC family member 3. | |||||
|
ALS2_MOUSE
|
||||||
| NC score | 0.050949 (rank : 136) | θ value | θ > 10 (rank : 140) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q920R0, Q8JZR1, Q9CXJ3 | Gene names | Als2 | |||
|
Domain Architecture |

|
|||||
| Description | Alsin (Amyotrophic lateral sclerosis protein 2 homolog). | |||||
|
SYT3_MOUSE
|
||||||
| NC score | 0.050918 (rank : 137) | θ value | θ > 10 (rank : 177) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O35681, P97791, Q80WV1 | Gene names | Syt3 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-3 (Synaptotagmin III) (SytIII). | |||||
|
OTOF_HUMAN
|
||||||
| NC score | 0.050757 (rank : 138) | θ value | 4.03905 (rank : 110) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9HC10, Q9HC08, Q9HC09, Q9Y650 | Gene names | OTOF, FER1L2 | |||
|
Domain Architecture |

|
|||||
| Description | Otoferlin (Fer-1-like protein 2). | |||||
|
DMD_MOUSE
|
||||||
| NC score | 0.050562 (rank : 139) | θ value | 0.00509761 (rank : 63) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1111 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P11531, O35653, Q60703 | Gene names | Dmd | |||
|
Domain Architecture |

|
|||||
| Description | Dystrophin. | |||||
|
SYT6_HUMAN
|
||||||
| NC score | 0.050264 (rank : 140) | θ value | θ > 10 (rank : 180) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q5T7P8 | Gene names | SYT6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptotagmin-6 (Synaptotagmin VI) (SytVI). | |||||
|
SYT3_HUMAN
|
||||||
| NC score | 0.050262 (rank : 141) | θ value | θ > 10 (rank : 176) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9BQG1, Q8N5Z1, Q8N640 | Gene names | SYT3 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-3 (Synaptotagmin III) (SytIII). | |||||
|
SYTL2_MOUSE
|
||||||
| NC score | 0.050050 (rank : 142) | θ value | θ > 10 (rank : 186) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 253 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q99N50, Q8BT37, Q99J89, Q99J90, Q99N51, Q99N52, Q99N55, Q99N56 | Gene names | Sytl2, Slp2 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 2 (Exophilin-4). | |||||
|
SYTL4_HUMAN
|
||||||
| NC score | 0.046738 (rank : 143) | θ value | 0.0252991 (rank : 71) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q96C24, Q5H9J3, Q5JPG8, Q8N9P4, Q9H4R0, Q9H4R1 | Gene names | SYTL4 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 4 (Exophilin-2) (Granuphilin). | |||||
|
UTRO_HUMAN
|
||||||
| NC score | 0.046023 (rank : 144) | θ value | 0.21417 (rank : 80) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P46939 | Gene names | UTRN, DMDL | |||
|
Domain Architecture |

|
|||||
| Description | Utrophin (Dystrophin-related protein 1) (DRP1) (DRP). | |||||
|
SYTL4_MOUSE
|
||||||
| NC score | 0.044530 (rank : 145) | θ value | 0.0252991 (rank : 72) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9R0Q1, Q8R321, Q9R0Q0 | Gene names | Sytl4, Slp4 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 4 (Exophilin-2) (Granuphilin). | |||||
|
ITSN1_HUMAN
|
||||||
| NC score | 0.038542 (rank : 146) | θ value | 4.03905 (rank : 108) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1757 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q15811, O95216, Q9UET5, Q9UK60, Q9UNK1, Q9UNK2, Q9UQ92 | Gene names | ITSN1, ITSN, SH3D1A | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-1 (SH3 domain-containing protein 1A) (SH3P17). | |||||
|
ITSN1_MOUSE
|
||||||
| NC score | 0.037742 (rank : 147) | θ value | 4.03905 (rank : 109) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1669 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9Z0R4, Q9R143 | Gene names | Itsn1, Ese1, Itsn | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-1 (EH and SH3 domains protein 1). | |||||
|
AKA12_HUMAN
|
||||||
| NC score | 0.033134 (rank : 148) | θ value | 4.03905 (rank : 105) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q02952, O00310, O00498, Q99970 | Gene names | AKAP12, AKAP250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 12 (A-kinase anchor protein 250 kDa) (AKAP 250) (Myasthenia gravis autoantigen gravin). | |||||
|
MDC1_HUMAN
|
||||||
| NC score | 0.030067 (rank : 149) | θ value | 0.0193708 (rank : 70) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
TARA_HUMAN
|
||||||
| NC score | 0.028023 (rank : 150) | θ value | 6.88961 (rank : 125) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9H2D6, O94797, Q2PZW8, Q2Q3Z9, Q2Q400, Q5R3M6, Q96DW1, Q9BT77, Q9BTL7, Q9BY98, Q9Y3L4 | Gene names | TRIOBP, KIAA1662, TARA | |||
|
Domain Architecture |

|
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
DLG7_HUMAN
|
||||||
| NC score | 0.025607 (rank : 151) | θ value | 1.81305 (rank : 91) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q15398, Q8NG58 | Gene names | DLG7, KIAA0008 | |||
|
Domain Architecture |

|
|||||
| Description | Discs large homolog 7 (Hepatoma up-regulated protein) (HURP). | |||||
|
MLH3_HUMAN
|
||||||
| NC score | 0.024248 (rank : 152) | θ value | 1.06291 (rank : 88) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UHC1, P49751, Q56DK9, Q9P292, Q9UHC0 | Gene names | MLH3 | |||
|
Domain Architecture |

|
|||||
| Description | DNA mismatch repair protein Mlh3 (MutL protein homolog 3). | |||||
|
DLG7_MOUSE
|
||||||
| NC score | 0.022937 (rank : 153) | θ value | 1.81305 (rank : 92) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8K4R9, Q6ZQL3, Q80VS7, Q8BUC8, Q8BZ79 | Gene names | Dlg7, Kiaa0008 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Discs large homolog 7 (Hepatoma up-regulated protein homolog) (HURP). | |||||
|
RFIP1_MOUSE
|
||||||
| NC score | 0.021632 (rank : 154) | θ value | 3.0926 (rank : 104) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9D620, Q3UBC2, Q8BN24 | Gene names | Rab11fip1, Rcp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab11 family-interacting protein 1 (Rab11-FIP1) (Rab-coupling protein). | |||||
|
CSF1_HUMAN
|
||||||
| NC score | 0.019882 (rank : 155) | θ value | 0.813845 (rank : 83) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P09603, Q13130, Q14086, Q14806, Q9UQR8 | Gene names | CSF1 | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage colony-stimulating factor 1 precursor (CSF-1) (MCSF) (M- CSF) (Lanimostim). | |||||
|
RBBP6_MOUSE
|
||||||
| NC score | 0.019241 (rank : 156) | θ value | 6.88961 (rank : 123) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
RBBP6_HUMAN
|
||||||
| NC score | 0.018792 (rank : 157) | θ value | 2.36792 (rank : 98) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
UBP2L_HUMAN
|
||||||
| NC score | 0.018271 (rank : 158) | θ value | 1.06291 (rank : 89) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q14157, Q5VU75, Q5VU76, Q9BTU3, Q9UGL5 | Gene names | UBAP2L, KIAA0144, NICE4 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-associated protein 2-like (Protein NICE-4). | |||||
|
4ET_HUMAN
|
||||||
| NC score | 0.018148 (rank : 159) | θ value | 0.813845 (rank : 82) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NRA8, Q8NCF2, Q9H708 | Gene names | EIF4ENIF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4E transporter (eIF4E transporter) (4E-T) (Eukaryotic translation initiation factor 4E nuclear import factor 1). | |||||
|
RFIP1_HUMAN
|
||||||
| NC score | 0.017957 (rank : 160) | θ value | 8.99809 (rank : 135) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q6WKZ4, Q6AZK4, Q6WKZ2, Q6WKZ6, Q86YV4, Q8TDL1, Q9H642 | Gene names | RAB11FIP1, RCP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab11 family-interacting protein 1 (Rab11-FIP1) (Rab-coupling protein). | |||||
|
AIM1_HUMAN
|
||||||
| NC score | 0.016966 (rank : 161) | θ value | 0.279714 (rank : 81) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Y4K1, O00296, Q5VWJ2 | Gene names | AIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Absent in melanoma 1 protein. | |||||
|
MILK1_MOUSE
|
||||||
| NC score | 0.014965 (rank : 162) | θ value | 3.0926 (rank : 99) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 788 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BGT6, Q8BJ60 | Gene names | Mirab13, D15Mit260, Kiaa1668 | |||
|
Domain Architecture |

|
|||||
| Description | Molecule interacting with Rab13 (MIRab13) (MICAL-like protein 1). | |||||
|
RGS3_MOUSE
|
||||||
| NC score | 0.014319 (rank : 163) | θ value | 8.99809 (rank : 136) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
EPN3_MOUSE
|
||||||
| NC score | 0.013909 (rank : 164) | θ value | 0.163984 (rank : 79) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q91W69, Q9CV55 | Gene names | Epn3 | |||
|
Domain Architecture |

|
|||||
| Description | Epsin-3 (EPS-15-interacting protein 3). | |||||
|
ANK2_HUMAN
|
||||||
| NC score | 0.013597 (rank : 165) | θ value | 1.06291 (rank : 84) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
LIMA1_MOUSE
|
||||||
| NC score | 0.013356 (rank : 166) | θ value | 6.88961 (rank : 121) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 298 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9ERG0, Q9ERG1 | Gene names | Lima1, D15Ertd366e, Eplin | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain and actin-binding protein 1 (Epithelial protein lost in neoplasm) (mEPLIN). | |||||
|
4ET_MOUSE
|
||||||
| NC score | 0.012995 (rank : 167) | θ value | 8.99809 (rank : 127) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9EST3, Q9CSS3 | Gene names | Eif4enif1, Clast4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4E transporter (eIF4E transporter) (4E-T) (Eukaryotic translation initiation factor 4E nuclear import factor 1). | |||||
|
CKAP2_HUMAN
|
||||||
| NC score | 0.012559 (rank : 168) | θ value | 2.36792 (rank : 96) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8WWK9, Q3KRA5, Q5VXB4, Q8IWV5, Q8IWV6, Q96FH9, Q9H012, Q9H0D0, Q9H988, Q9HC49, Q9NVG4 | Gene names | CKAP2, LB1, TMAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-associated protein 2 (Tumor-associated microtubule- associated protein) (CTCL tumor antigen se20-10). | |||||
|
MAP4_HUMAN
|
||||||
| NC score | 0.012177 (rank : 169) | θ value | 2.36792 (rank : 97) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 428 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P27816, Q13082, Q96A76 | Gene names | MAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 4 (MAP 4). | |||||
|
PDLI4_MOUSE
|
||||||
| NC score | 0.010521 (rank : 170) | θ value | 6.88961 (rank : 122) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P70271, Q8K0W4 | Gene names | Pdlim4, Ril | |||
|
Domain Architecture |

|
|||||
| Description | PDZ and LIM domain protein 4 (LIM protein RIL) (Reversion-induced LIM protein). | |||||
|
NCOR1_MOUSE
|
||||||
| NC score | 0.009506 (rank : 171) | θ value | 3.0926 (rank : 100) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 521 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q60974, Q60812 | Gene names | Ncor1, Rxrip13 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR) (Retinoid X receptor- interacting protein 13) (RIP13). | |||||
|
AKAP1_MOUSE
|
||||||
| NC score | 0.009324 (rank : 172) | θ value | 4.03905 (rank : 106) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O08715, O08714, P97488 | Gene names | Akap1, Akap | |||
|
Domain Architecture |

|
|||||
| Description | A kinase anchor protein 1, mitochondrial precursor (Protein kinase A- anchoring protein 1) (PRKA1) (Dual specificity A-kinase-anchoring protein 1) (D-AKAP-1) (Spermatid A-kinase anchor protein) (S-AKAP). | |||||
|
RM49_HUMAN
|
||||||
| NC score | 0.008530 (rank : 173) | θ value | 6.88961 (rank : 124) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q13405 | Gene names | MRPL49, C11orf4, NOF1 | |||
|
Domain Architecture |

|
|||||
| Description | Mitochondrial 39S ribosomal protein L49 (L49mt) (MRP-L49) (Protein NOF1) (Neighbor of FAU) (NOF). | |||||
|
NUMB_HUMAN
|
||||||
| NC score | 0.008409 (rank : 174) | θ value | 5.27518 (rank : 114) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P49757, Q6NUQ7, Q86SY1, Q8WW73, Q9UBG1, Q9UEQ4, Q9UKE8, Q9UKE9, Q9UKF0, Q9UQJ4 | Gene names | NUMB | |||
|
Domain Architecture |

|
|||||
| Description | Protein numb homolog (h-Numb) (Protein S171). | |||||
|
ORC3_HUMAN
|
||||||
| NC score | 0.008018 (rank : 175) | θ value | 8.99809 (rank : 133) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UBD5, Q13565, Q6IUY7, Q9UG44, Q9UNT6 | Gene names | ORC3L, LATHEO | |||
|
Domain Architecture |

|
|||||
| Description | Origin recognition complex subunit 3 (Origin recognition complex subunit Latheo). | |||||
|
MINT_MOUSE
|
||||||
| NC score | 0.007330 (rank : 176) | θ value | 8.99809 (rank : 131) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
PRLR_MOUSE
|
||||||
| NC score | 0.006590 (rank : 177) | θ value | 3.0926 (rank : 103) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q08501, P15212, P15213, Q62099 | Gene names | Prlr | |||
|
Domain Architecture |

|
|||||
| Description | Prolactin receptor precursor (PRL-R). | |||||
|
PECI_MOUSE
|
||||||
| NC score | 0.006388 (rank : 178) | θ value | 3.0926 (rank : 102) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9WUR2 | Gene names | Peci | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisomal 3,2-trans-enoyl-CoA isomerase (EC 5.3.3.8) (Dodecenoyl-CoA isomerase) (Delta(3),delta(2)-enoyl-CoA isomerase) (D3,D2-enoyl-CoA isomerase). | |||||
|
CN103_MOUSE
|
||||||
| NC score | 0.006218 (rank : 179) | θ value | 6.88961 (rank : 117) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q80XK6, Q8R1H5, Q9CWK6 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C14orf103 homolog. | |||||
|
ZAN_MOUSE
|
||||||
| NC score | 0.005190 (rank : 180) | θ value | 6.88961 (rank : 126) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 808 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O88799, O08647 | Gene names | Zan | |||
|
Domain Architecture |

|
|||||
| Description | Zonadhesin precursor. | |||||
|
PHC1_HUMAN
|
||||||
| NC score | 0.004818 (rank : 181) | θ value | 8.99809 (rank : 134) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P78364, Q8WVM3, Q9BU63 | Gene names | PHC1, EDR1, PH1 | |||
|
Domain Architecture |

|
|||||
| Description | Polyhomeotic-like protein 1 (hPH1) (Early development regulatory protein 1). | |||||
|
MLL3_HUMAN
|
||||||
| NC score | 0.004652 (rank : 182) | θ value | 8.99809 (rank : 132) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
SCN7A_HUMAN
|
||||||
| NC score | 0.001973 (rank : 183) | θ value | 8.99809 (rank : 137) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q01118 | Gene names | SCN7A, SCN6A | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 7 subunit alpha (Sodium channel protein type VII subunit alpha) (Putative voltage-gated sodium channel alpha subunit Nax) (Sodium channel protein, cardiac and skeletal muscle alpha-subunit). | |||||
|
CF060_HUMAN
|
||||||
| NC score | 0.001948 (rank : 184) | θ value | 8.99809 (rank : 129) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 1270 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8NB25, Q96GY8, Q9H851 | Gene names | C6orf60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C6orf60. | |||||
|
FOXN3_HUMAN
|
||||||
| NC score | 0.001765 (rank : 185) | θ value | 4.03905 (rank : 107) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O00409, Q96II7, Q9UIE7 | Gene names | CHES1, FOXN3 | |||
|
Domain Architecture |

|
|||||
| Description | Checkpoint suppressor 1 (Forkhead box protein N3). | |||||
|
ZN217_HUMAN
|
||||||
| NC score | 0.001454 (rank : 186) | θ value | 0.0961366 (rank : 76) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 905 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O75362 | Gene names | ZNF217, ZABC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 217. | |||||
|
BCAR1_HUMAN
|
||||||
| NC score | 0.001107 (rank : 187) | θ value | 8.99809 (rank : 128) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 554 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P56945 | Gene names | BCAR1, CAS, CRKAS | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer anti-estrogen resistance protein 1 (CRK-associated substrate) (p130cas). | |||||
|
HA26_HUMAN
|
||||||
| NC score | 0.000717 (rank : 188) | θ value | 6.88961 (rank : 120) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P01906, O19789 | Gene names | HLA-DQA2 | |||
|
Domain Architecture |

|
|||||
| Description | HLA class II histocompatibility antigen, DQ(6) alpha chain precursor (DX alpha chain) (HLA-DQA1). | |||||
|
BMX_HUMAN
|
||||||
| NC score | 0.000225 (rank : 189) | θ value | 2.36792 (rank : 95) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 942 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P51813, O60564, Q12871 | Gene names | BMX | |||
|
Domain Architecture |

|
|||||
| Description | Cytoplasmic tyrosine-protein kinase BMX (EC 2.7.10.2) (Bone marrow tyrosine kinase gene in chromosome X protein) (Epithelial and endothelial tyrosine kinase) (ETK) (NTK38). | |||||
|
E2AK3_HUMAN
|
||||||
| NC score | -0.000894 (rank : 190) | θ value | 8.99809 (rank : 130) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 859 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NZJ5, O95846 | Gene names | EIF2AK3, PEK, PERK | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2-alpha kinase 3 precursor (EC 2.7.11.1) (PRKR-like endoplasmic reticulum kinase) (Pancreatic eIF2-alpha kinase) (HsPEK). | |||||
|
DSG2_HUMAN
|
||||||
| NC score | -0.001079 (rank : 191) | θ value | 6.88961 (rank : 118) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14126 | Gene names | DSG2 | |||
|
Domain Architecture |

|
|||||
| Description | Desmoglein-2 precursor (HDGC). | |||||
|
SUHW4_MOUSE
|
||||||
| NC score | -0.002583 (rank : 192) | θ value | 8.99809 (rank : 138) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 737 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q68FE8, Q3T9B1, Q8BI82 | Gene names | Suhw4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppressor of hairy wing homolog 4. | |||||
|
KSR2_HUMAN
|
||||||
| NC score | -0.004243 (rank : 193) | θ value | 5.27518 (rank : 113) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 920 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6VAB6, Q8N775 | Gene names | KSR2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinase suppressor of ras-2 (hKSR2). | |||||
|
CDKL2_HUMAN
|
||||||
| NC score | -0.004645 (rank : 194) | θ value | 1.81305 (rank : 90) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 838 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q92772 | Gene names | CDKL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-dependent kinase-like 2 (EC 2.7.11.22) (Serine/threonine- protein kinase KKIAMRE) (Protein kinase p56 KKIAMRE). | |||||
|
PAK7_HUMAN
|
||||||
| NC score | -0.004660 (rank : 195) | θ value | 1.81305 (rank : 93) | |||
| Query Neighborhood Hits | 138 | Target Neighborhood Hits | 875 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9P286, Q5W115, Q9BX09, Q9ULF6 | Gene names | PAK7, KIAA1264, PAK5 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase PAK 7 (EC 2.7.11.1) (p21-activated kinase 7) (PAK-7) (PAK-5). | |||||