Please be patient as the page loads
|
SCN7A_HUMAN
|
||||||
| SwissProt Accessions | Q01118 | Gene names | SCN7A, SCN6A | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 7 subunit alpha (Sodium channel protein type VII subunit alpha) (Putative voltage-gated sodium channel alpha subunit Nax) (Sodium channel protein, cardiac and skeletal muscle alpha-subunit). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
SC10A_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.981513 (rank : 11) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9Y5Y9 | Gene names | SCN10A, PN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium channel protein type 10 subunit alpha (Sodium channel protein type X subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.8) (Peripheral nerve sodium channel 3) (hPN3). | |||||
|
SC10A_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.980622 (rank : 12) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q6QIY3, Q62243, Q6EWG7, Q6KCH7, Q703F9 | Gene names | Scn10a, Pn3, Sns | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium channel protein type 10 subunit alpha (Sodium channel protein type X subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.8) (Peripheral nerve sodium channel 3) (Sensory neuron sodium channel). | |||||
|
SC11A_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.983137 (rank : 10) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9UI33, Q68K15, Q8NDX3, Q9UHE0, Q9UHM0 | Gene names | SCN11A, PN5, SCN12A, SNS2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium channel protein type 11 subunit alpha (Sodium channel protein type XI subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.9) (Sensory neuron sodium channel 2) (Peripheral nerve sodium channel 5) (hNaN). | |||||
|
SC11A_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.984827 (rank : 9) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q9R053, Q9JMD4 | Gene names | Scn11a, Nan, Nat, Sns2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium channel protein type 11 subunit alpha (Sodium channel protein type XI subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.9) (Sensory neuron sodium channel 2) (NaN). | |||||
|
SCN1A_HUMAN
|
||||||
| θ value | 0 (rank : 5) | NC score | 0.985313 (rank : 6) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P35498, Q16172, Q96LA3, Q9C008 | Gene names | SCN1A, NAC1, SCN1 | |||
|
Domain Architecture |
|
|||||
| Description | Sodium channel protein type 1 subunit alpha (Sodium channel protein type I subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.1) (Sodium channel protein, brain I subunit alpha). | |||||
|
SCN2A_HUMAN
|
||||||
| θ value | 0 (rank : 6) | NC score | 0.985672 (rank : 4) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q99250, Q14472, Q9BZC9, Q9BZD0 | Gene names | SCN2A, NAC2, SCN2A2 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 2 subunit alpha (Sodium channel protein type II subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.2) (Sodium channel protein, brain II subunit alpha) (HBSC II). | |||||
|
SCN3A_HUMAN
|
||||||
| θ value | 0 (rank : 7) | NC score | 0.986449 (rank : 3) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q9NY46, Q16142, Q9BZB3, Q9C006, Q9NYK2, Q9UPD1, Q9Y6P4 | Gene names | SCN3A, NAC3 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 3 subunit alpha (Sodium channel protein type III subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.3) (Sodium channel protein, brain III subunit alpha) (Voltage- gated sodium channel subtype III). | |||||
|
SCN4A_HUMAN
|
||||||
| θ value | 0 (rank : 8) | NC score | 0.989721 (rank : 2) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | P35499, Q15478, Q16447, Q7Z6B1 | Gene names | SCN4A | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 4 subunit alpha (Sodium channel protein type IV subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.4) (Sodium channel protein, skeletal muscle subunit alpha) (SkM1). | |||||
|
SCN5A_HUMAN
|
||||||
| θ value | 0 (rank : 9) | NC score | 0.978532 (rank : 13) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q14524 | Gene names | SCN5A | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 5 subunit alpha (Sodium channel protein type V subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.5) (Sodium channel protein, cardiac muscle alpha-subunit) (HH1). | |||||
|
SCN7A_HUMAN
|
||||||
| θ value | 0 (rank : 10) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 135 | |
| SwissProt Accessions | Q01118 | Gene names | SCN7A, SCN6A | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 7 subunit alpha (Sodium channel protein type VII subunit alpha) (Putative voltage-gated sodium channel alpha subunit Nax) (Sodium channel protein, cardiac and skeletal muscle alpha-subunit). | |||||
|
SCN8A_HUMAN
|
||||||
| θ value | 0 (rank : 11) | NC score | 0.985606 (rank : 5) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9UQD0, O95788, Q9NYX2, Q9UPB2 | Gene names | SCN8A | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 8 subunit alpha (Sodium channel protein type VIII subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.6). | |||||
|
SCN8A_MOUSE
|
||||||
| θ value | 0 (rank : 12) | NC score | 0.985297 (rank : 7) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q9WTU3, Q60828, Q60858, Q62449 | Gene names | Scn8a, Nbna1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 8 subunit alpha (Sodium channel protein type VIII subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.6). | |||||
|
SCN9A_HUMAN
|
||||||
| θ value | 0 (rank : 13) | NC score | 0.985220 (rank : 8) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q15858, Q6B4R9, Q6B4S0, Q6B4S1, Q70HX1, Q70HX2, Q8WTU1, Q8WWN4 | Gene names | SCN9A, NENA, PN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium channel protein type 9 subunit alpha (Sodium channel protein type IX subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.7) (Neuroendocrine sodium channel) (hNE-Na) (Peripheral sodium channel 1). | |||||
|
CAC1C_MOUSE
|
||||||
| θ value | 5.67233e-95 (rank : 14) | NC score | 0.851684 (rank : 25) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q01815, Q04476, Q61242, Q99242 | Gene names | Cacna1c, Cach2, Cacn2, Cacnl1a1, Cchl1a1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1C (Voltage- gated calcium channel subunit alpha Cav1.2) (Calcium channel, L type, alpha-1 polypeptide, isoform 1, cardiac muscle) (Mouse brain class C) (MBC) (MELC-CC). | |||||
|
CAC1C_HUMAN
|
||||||
| θ value | 1.88829e-90 (rank : 15) | NC score | 0.848490 (rank : 30) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q13936, Q13917, Q13918, Q13919, Q13920, Q13921, Q13922, Q13923, Q13924, Q13925, Q13926, Q13927, Q13928, Q13929, Q13930, Q13932, Q13933, Q14743, Q14744, Q15877, Q99025, Q99241, Q99875 | Gene names | CACNA1C, CACH2, CACN2, CACNL1A1, CCHL1A1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1C (Voltage- gated calcium channel subunit alpha Cav1.2) (Calcium channel, L type, alpha-1 polypeptide, isoform 1, cardiac muscle). | |||||
|
CAC1H_HUMAN
|
||||||
| θ value | 1.31987e-59 (rank : 16) | NC score | 0.869295 (rank : 16) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | O95180, O95802, Q8WWI6, Q96QI6, Q96RZ9, Q9NYY4, Q9NYY5 | Gene names | CACNA1H | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1H (Voltage- gated calcium channel subunit alpha Cav3.2) (Low-voltage-activated calcium channel alpha1 3.2 subunit). | |||||
|
CAC1H_MOUSE
|
||||||
| θ value | 1.7238e-59 (rank : 17) | NC score | 0.876186 (rank : 15) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | O88427, Q80TJ2, Q9JKU5 | Gene names | Cacna1h, Kiaa1120 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1H (Voltage- gated calcium channel subunit alpha Cav3.2). | |||||
|
CAC1I_HUMAN
|
||||||
| θ value | 3.84025e-59 (rank : 18) | NC score | 0.895687 (rank : 14) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9P0X4, O95504, Q5JZ88, Q7Z6S9, Q8NFX6, Q9NZC8, Q9UH15, Q9UH30, Q9ULU9, Q9UNE6 | Gene names | CACNA1I, KIAA1120 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1I (Voltage- gated calcium channel subunit alpha Cav3.3) (Ca(v)3.3). | |||||
|
CAC1E_MOUSE
|
||||||
| θ value | 8.55514e-59 (rank : 19) | NC score | 0.862984 (rank : 17) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q61290 | Gene names | Cacna1e, Cach6, Cacnl1a6, Cchra1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent R-type calcium channel subunit alpha-1E (Voltage- gated calcium channel subunit alpha Cav2.3) (Calcium channel, L type, alpha-1 polypeptide, isoform 6) (Brain calcium channel II) (BII). | |||||
|
CAC1E_HUMAN
|
||||||
| θ value | 1.45929e-58 (rank : 20) | NC score | 0.862604 (rank : 18) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q15878, Q14580, Q14581 | Gene names | CACNA1E, CACH6, CACNL1A6 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent R-type calcium channel subunit alpha-1E (Voltage- gated calcium channel subunit alpha Cav2.3) (Calcium channel, L type, alpha-1 polypeptide, isoform 6) (Brain calcium channel II) (BII). | |||||
|
CAC1A_HUMAN
|
||||||
| θ value | 2.48917e-58 (rank : 21) | NC score | 0.851338 (rank : 26) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | O00555, P78510, P78511, Q16290, Q92690, Q99790, Q99791, Q99792, Q99793 | Gene names | CACNA1A, CACH4, CACN3, CACNL1A4 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent P/Q-type calcium channel subunit alpha-1A (Voltage- gated calcium channel subunit alpha Cav2.1) (Calcium channel, L type, alpha-1 polypeptide isoform 4) (Brain calcium channel I) (BI). | |||||
|
CAC1A_MOUSE
|
||||||
| θ value | 2.48917e-58 (rank : 22) | NC score | 0.859059 (rank : 19) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | P97445 | Gene names | Cacna1a, Cach4, Cacn3, Cacnl1a4, Ccha1a | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent P/Q-type calcium channel subunit alpha-1A (Voltage- gated calcium channel subunit alpha Cav2.1) (Calcium channel, L type, alpha-1 polypeptide isoform 4) (Brain calcium channel I) (BI). | |||||
|
CAC1F_MOUSE
|
||||||
| θ value | 3.59435e-57 (rank : 23) | NC score | 0.850362 (rank : 27) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9JIS7 | Gene names | Cacna1f | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1F (Voltage- gated calcium channel subunit alpha Cav1.4). | |||||
|
CAC1G_HUMAN
|
||||||
| θ value | 8.00737e-57 (rank : 24) | NC score | 0.852488 (rank : 24) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 587 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | O43497, O43498, O94770, Q9NYU4, Q9NYU5, Q9NYU6, Q9NYU7, Q9NYU8, Q9NYU9, Q9NYV0, Q9NYV1, Q9UHN9, Q9UHP0, Q9ULU6, Q9UNG7, Q9Y5T2, Q9Y5T3 | Gene names | CACNA1G, KIAA1123 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1G (Voltage- gated calcium channel subunit alpha Cav3.1) (Cav3.1c) (NBR13). | |||||
|
CAC1D_HUMAN
|
||||||
| θ value | 1.36585e-56 (rank : 25) | NC score | 0.855685 (rank : 21) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q01668, Q13916, Q13931 | Gene names | CACNA1D, CACH3, CACN4, CACNL1A2, CCHL1A2 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1D (Voltage- gated calcium channel subunit alpha Cav1.3) (Calcium channel, L type, alpha-1 polypeptide, isoform 2). | |||||
|
CAC1F_HUMAN
|
||||||
| θ value | 1.78386e-56 (rank : 26) | NC score | 0.849894 (rank : 29) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O60840, O43901 | Gene names | CACNA1F, CACNAF1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1F (Voltage- gated calcium channel subunit alpha Cav1.4). | |||||
|
CAC1B_MOUSE
|
||||||
| θ value | 8.85319e-56 (rank : 27) | NC score | 0.850112 (rank : 28) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | O55017, Q60609 | Gene names | Cacna1b, Cach5, Cacnl1a5, Cchn1a | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent N-type calcium channel subunit alpha-1B (Voltage- gated calcium channel subunit alpha Cav2.2) (Calcium channel, L type, alpha-1 polypeptide isoform 5) (Brain calcium channel III) (BIII). | |||||
|
CAC1S_MOUSE
|
||||||
| θ value | 3.36421e-55 (rank : 28) | NC score | 0.856526 (rank : 20) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q02789, Q99240 | Gene names | Cacna1s, Cach1, Cach1b, Cacn1, Cacnl1a3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1S (Voltage- gated calcium channel subunit alpha Cav1.1) (Calcium channel, L type, alpha-1 polypeptide, isoform 3, skeletal muscle). | |||||
|
CAC1B_HUMAN
|
||||||
| θ value | 1.66964e-54 (rank : 29) | NC score | 0.853346 (rank : 23) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q00975 | Gene names | CACNA1B, CACH5, CACNL1A5 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent N-type calcium channel subunit alpha-1B (Voltage- gated calcium channel subunit alpha Cav2.2) (Calcium channel, L type, alpha-1 polypeptide isoform 5) (Brain calcium channel III) (BIII). | |||||
|
CAC1S_HUMAN
|
||||||
| θ value | 2.18062e-54 (rank : 30) | NC score | 0.855059 (rank : 22) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q13698, Q12896, Q13934 | Gene names | CACNA1S, CACH1, CACN1, CACNL1A3 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1S (Voltage- gated calcium channel subunit alpha Cav1.1) (Calcium channel, L type, alpha-1 polypeptide, isoform 3, skeletal muscle). | |||||
|
KCNF1_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 31) | NC score | 0.217548 (rank : 36) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9H3M0, O43527 | Gene names | KCNF1 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium voltage-gated channel subfamily F member 1 (Voltage-gated potassium channel subunit Kv5.1) (kH1). | |||||
|
KCNC4_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 32) | NC score | 0.114002 (rank : 54) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q03721, Q3MIM4, Q5TBI6 | Gene names | KCNC4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 4 (Voltage-gated potassium channel subunit Kv3.4) (KSHIIIC). | |||||
|
KCNH2_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 33) | NC score | 0.018011 (rank : 101) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q12809, O75418, O75680, Q9BT72, Q9BUT7, Q9H3P0 | Gene names | KCNH2, ERG, ERG1, HERG, HERG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 2 (Voltage-gated potassium channel subunit Kv11.1) (Ether-a-go-go-related gene potassium channel 1) (H-ERG) (Erg1) (Ether-a-go-go-related protein 1) (Eag-related protein 1) (eag homolog). | |||||
|
KCNH2_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 34) | NC score | 0.019201 (rank : 99) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O35219, O35220, O35221, O35989 | Gene names | Kcnh2, Erg, Merg1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 2 (Voltage-gated potassium channel subunit Kv11.1) (Ether-a-go-go-related gene potassium channel 1) (ERG1) (MERG) (Merg1) (Ether-a-go-go-related protein 1) (Eag-related protein 1). | |||||
|
KCNA1_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 35) | NC score | 0.079537 (rank : 63) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q09470 | Gene names | KCNA1 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium voltage-gated channel subfamily A member 1 (Voltage-gated potassium channel subunit Kv1.1) (HUKI) (HBK1). | |||||
|
KCNA1_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 36) | NC score | 0.080894 (rank : 62) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P16388 | Gene names | Kcna1 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium voltage-gated channel subfamily A member 1 (Voltage-gated potassium channel subunit Kv1.1) (MKI) (MBK1). | |||||
|
KCNB2_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 37) | NC score | 0.197786 (rank : 37) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q92953, Q9BXD3 | Gene names | KCNB2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily B member 2 (Voltage-gated potassium channel subunit Kv2.2). | |||||
|
TRPV3_HUMAN
|
||||||
| θ value | 0.125558 (rank : 38) | NC score | 0.086090 (rank : 61) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8NET8, Q8NDW7, Q8NET9, Q8NFH2 | Gene names | TRPV3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 3 (TrpV3) (Vanilloid receptor-like 3) (VRL-3). | |||||
|
KCNC4_MOUSE
|
||||||
| θ value | 0.163984 (rank : 39) | NC score | 0.106469 (rank : 56) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q8R1C0 | Gene names | Kcnc4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium voltage-gated channel subfamily C member 4 (Voltage-gated potassium channel subunit Kv3.4). | |||||
|
KCND3_HUMAN
|
||||||
| θ value | 0.163984 (rank : 40) | NC score | 0.162978 (rank : 44) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9UK17, O60576, O60577, Q5T0M0, Q9UH85, Q9UH86, Q9UK16 | Gene names | KCND3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 3 (Voltage-gated potassium channel subunit Kv4.3). | |||||
|
KCND3_MOUSE
|
||||||
| θ value | 0.163984 (rank : 41) | NC score | 0.163118 (rank : 43) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9Z0V1, Q8CC44, Q9Z0V0 | Gene names | Kcnd3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 3 (Voltage-gated potassium channel subunit Kv4.3). | |||||
|
PK2L1_HUMAN
|
||||||
| θ value | 0.163984 (rank : 42) | NC score | 0.267650 (rank : 32) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9P0L9, O75972, Q9UP35, Q9UPA2 | Gene names | PKD2L1, PKD2L, PKDL | |||
|
Domain Architecture |

|
|||||
| Description | Polycystic kidney disease 2-like 1 protein (Polycystin-L) (Polycystin- 2 homolog). | |||||
|
KCNA3_MOUSE
|
||||||
| θ value | 0.21417 (rank : 43) | NC score | 0.069688 (rank : 74) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P16390 | Gene names | Kcna3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 3 (Voltage-gated potassium channel subunit Kv1.3) (MK3). | |||||
|
MEPE_HUMAN
|
||||||
| θ value | 0.21417 (rank : 44) | NC score | 0.039495 (rank : 89) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9NQ76 | Gene names | MEPE | |||
|
Domain Architecture |
|
|||||
| Description | Matrix extracellular phosphoglycoprotein precursor (Osteoblast/osteocyte factor 45) (OF45). | |||||
|
KCNA5_MOUSE
|
||||||
| θ value | 0.365318 (rank : 45) | NC score | 0.071493 (rank : 71) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q61762 | Gene names | Kcna5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 5 (Voltage-gated potassium channel subunit Kv1.5) (KV1-5). | |||||
|
KCNA3_HUMAN
|
||||||
| θ value | 0.47712 (rank : 46) | NC score | 0.070266 (rank : 73) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P22001 | Gene names | KCNA3, HGK5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 3 (Voltage-gated potassium channel subunit Kv1.3) (HPCN3) (HGK5) (HuKIII) (HLK3). | |||||
|
PKD2_MOUSE
|
||||||
| θ value | 0.47712 (rank : 47) | NC score | 0.248608 (rank : 34) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O35245, Q9QWP0, Q9Z193, Q9Z194 | Gene names | Pkd2 | |||
|
Domain Architecture |

|
|||||
| Description | Polycystin-2 (Polycystic kidney disease 2 protein homolog). | |||||
|
SEGN_HUMAN
|
||||||
| θ value | 0.47712 (rank : 48) | NC score | 0.026503 (rank : 95) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O76038, Q5VV44, Q96QV7, Q9UJF6 | Gene names | SCGN, SECRET | |||
|
Domain Architecture |
|
|||||
| Description | Secretagogin. | |||||
|
KCND2_HUMAN
|
||||||
| θ value | 0.62314 (rank : 49) | NC score | 0.155114 (rank : 47) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9NZV8, O95012, O95021, Q9UBY7, Q9UN98, Q9UNH9 | Gene names | KCND2, KIAA1044 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 2 (Voltage-gated potassium channel subunit Kv4.2). | |||||
|
KCND2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 50) | NC score | 0.155276 (rank : 46) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9Z0V2, Q8BSK3, Q8CHB7, Q9JJ60 | Gene names | Kcnd2, Kiaa1044 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 2 (Voltage-gated potassium channel subunit Kv4.2). | |||||
|
DZIP1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 51) | NC score | 0.013397 (rank : 104) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 530 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8BMD2, Q6ZQ10, Q9CRK5 | Gene names | Dzip1, Kiaa0996 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein DZIP1 (DAZ-interacting protein 1 homolog). | |||||
|
KCNQ4_HUMAN
|
||||||
| θ value | 0.813845 (rank : 52) | NC score | 0.075006 (rank : 68) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P56696, O96025 | Gene names | KCNQ4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 4 (Voltage-gated potassium channel subunit Kv7.4) (Potassium channel subunit alpha KvLQT4) (KQT-like 4). | |||||
|
RIF1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 53) | NC score | 0.012981 (rank : 105) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q6PR54, Q6T8D9, Q7TPD8, Q8BTU2, Q8C408, Q9CS77 | Gene names | Rif1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Telomere-associated protein RIF1 (Rap1-interacting factor 1 homolog) (mRif1). | |||||
|
ZN318_HUMAN
|
||||||
| θ value | 0.813845 (rank : 54) | NC score | 0.019689 (rank : 98) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 497 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q5VUA4, O94796, Q4G0E4, Q8NEM6, Q9UNU8, Q9Y2W9 | Gene names | ZNF318 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 318 (Endocrine regulatory protein). | |||||
|
KCNA4_MOUSE
|
||||||
| θ value | 1.06291 (rank : 55) | NC score | 0.056192 (rank : 80) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q61423 | Gene names | Kcna4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 4 (Voltage-gated potassium channel subunit Kv1.4). | |||||
|
KCNS2_MOUSE
|
||||||
| θ value | 1.06291 (rank : 56) | NC score | 0.189779 (rank : 39) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | O35174, Q543P3 | Gene names | Kcns2 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium voltage-gated channel subfamily S member 2 (Voltage-gated potassium channel subunit Kv9.2) (Delayed-rectifier K(+) channel alpha subunit 2). | |||||
|
NPM2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 57) | NC score | 0.018418 (rank : 100) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q86SE8 | Gene names | NPM2 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoplasmin-2. | |||||
|
TAXB1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 58) | NC score | 0.009494 (rank : 113) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 829 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q86VP1, O60398, O95770, Q13311, Q9BQG5, Q9UI88 | Gene names | TAX1BP1, T6BP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tax1-binding protein 1 (TRAF6-binding protein). | |||||
|
TMF1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 59) | NC score | 0.008353 (rank : 116) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 987 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P82094 | Gene names | TMF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TATA element modulatory factor (TMF). | |||||
|
KCNA2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 60) | NC score | 0.077179 (rank : 67) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P16389 | Gene names | KCNA2 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium voltage-gated channel subfamily A member 2 (Voltage-gated potassium channel subunit Kv1.2) (HBK5) (NGK1) (HUKIV). | |||||
|
KCNA2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 61) | NC score | 0.078011 (rank : 66) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P63141, P15386, Q02010, Q8C8W4 | Gene names | Kcna2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium voltage-gated channel subfamily A member 2 (Voltage-gated potassium channel subunit Kv1.2) (MK2). | |||||
|
MKL1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 62) | NC score | 0.012028 (rank : 107) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 667 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8K4J6 | Gene names | Mkl1, Bsac | |||
|
Domain Architecture |

|
|||||
| Description | MKL/myocardin-like protein 1 (Myocardin-related transcription factor A) (MRTF-A) (Megakaryoblastic leukemia 1 protein homolog) (Basic SAP coiled-coil transcription activator). | |||||
|
MYH13_HUMAN
|
||||||
| θ value | 1.38821 (rank : 63) | NC score | 0.007861 (rank : 121) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1504 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9UKX3, O95252 | Gene names | MYH13 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-13 (Myosin heavy chain, skeletal muscle, extraocular) (MyHC- eo). | |||||
|
PKD2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 64) | NC score | 0.245715 (rank : 35) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q13563, O60441, Q15764 | Gene names | PKD2 | |||
|
Domain Architecture |

|
|||||
| Description | Polycystin-2 (Polycystic kidney disease 2 protein homolog) (Autosomal dominant polycystic kidney disease type II protein) (Polycystwin) (R48321). | |||||
|
TRPM2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 65) | NC score | 0.016592 (rank : 102) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O94759, Q5KTC2, Q6J3P5, Q96KN6, Q96Q93 | Gene names | TRPM2, EREG1, KNP3, LTRPC2, TRPC7 | |||
|
Domain Architecture |

|
|||||
| Description | Transient receptor potential cation channel subfamily M member 2 (EC 3.6.1.13) (Long transient receptor potential channel 2) (LTrpC2) (LTrpC-2) (Transient receptor potential channel 7) (TrpC7) (Estrogen- responsive element-associated gene 1 protein). | |||||
|
CP135_HUMAN
|
||||||
| θ value | 1.81305 (rank : 66) | NC score | 0.007413 (rank : 125) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1290 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q66GS9, O75130, Q9H8H7 | Gene names | CEP135, CEP4, KIAA0635 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 135 kDa (Cep135 protein) (Centrosomal protein 4). | |||||
|
DDX23_HUMAN
|
||||||
| θ value | 1.81305 (rank : 67) | NC score | 0.003061 (rank : 147) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9BUQ8, O43188 | Gene names | DDX23 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX23 (EC 3.6.1.-) (DEAD box protein 23) (100 kDa U5 snRNP-specific protein) (U5-100kD) (PRP28 homolog). | |||||
|
DRD1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 68) | NC score | -0.000808 (rank : 156) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 733 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61616, Q8C8P8 | Gene names | Drd1, Drd1a, Gpcr15 | |||
|
Domain Architecture |

|
|||||
| Description | D(1A) dopamine receptor. | |||||
|
I17RA_MOUSE
|
||||||
| θ value | 1.81305 (rank : 69) | NC score | 0.010994 (rank : 109) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q60943 | Gene names | IL17ra, Il17r | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interleukin-17 receptor A precursor (IL-17 receptor). | |||||
|
KCNA4_HUMAN
|
||||||
| θ value | 1.81305 (rank : 70) | NC score | 0.055659 (rank : 81) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P22459 | Gene names | KCNA4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 4 (Voltage-gated potassium channel subunit Kv1.4) (HK1) (HPCN2) (HBK4) (HUKII). | |||||
|
KCNA5_HUMAN
|
||||||
| θ value | 1.81305 (rank : 71) | NC score | 0.065813 (rank : 75) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P22460 | Gene names | KCNA5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 5 (Voltage-gated potassium channel subunit Kv1.5) (HK2) (HPCN1). | |||||
|
KCNA6_HUMAN
|
||||||
| θ value | 1.81305 (rank : 72) | NC score | 0.071763 (rank : 70) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P17658 | Gene names | KCNA6 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 6 (Voltage-gated potassium channel subunit Kv1.6) (HBK2). | |||||
|
KCNA6_MOUSE
|
||||||
| θ value | 1.81305 (rank : 73) | NC score | 0.074927 (rank : 69) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q61923 | Gene names | Kcna6 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 6 (Voltage-gated potassium channel subunit Kv1.6) (MK1.6). | |||||
|
KCNC1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 74) | NC score | 0.128919 (rank : 51) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P48547 | Gene names | KCNC1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 1 (Voltage-gated potassium channel subunit Kv3.1) (Kv4) (NGK2). | |||||
|
PK2L2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 75) | NC score | 0.286844 (rank : 31) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9NZM6, Q9UNJ0 | Gene names | PKD2L2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polycystic kidney disease 2-like 2 protein (Polycystin-L2). | |||||
|
TRPA1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 76) | NC score | 0.005546 (rank : 138) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 339 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O75762 | Gene names | TRPA1, ANKTM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily A member 1 (Ankyrin-like with transmembrane domains protein 1) (Transformation sensitive-protein p120). | |||||
|
TRPM8_MOUSE
|
||||||
| θ value | 1.81305 (rank : 77) | NC score | 0.041875 (rank : 88) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8R4D5 | Gene names | Trpm8, Trpp8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily M member 8. | |||||
|
CALB2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 78) | NC score | 0.026840 (rank : 94) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P22676, Q53HD2, Q96BK4 | Gene names | CALB2, CAB29 | |||
|
Domain Architecture |
|
|||||
| Description | Calretinin (CR) (29 kDa calbindin). | |||||
|
CALB2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 79) | NC score | 0.027071 (rank : 93) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q08331, Q60964, Q9JM81 | Gene names | Calb2 | |||
|
Domain Architecture |

|
|||||
| Description | Calretinin (CR). | |||||
|
KCNC1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 80) | NC score | 0.126614 (rank : 52) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P15388 | Gene names | Kcnc1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 1 (Voltage-gated potassium channel subunit Kv3.1) (Kv4) (NGK2). | |||||
|
KCNS2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 81) | NC score | 0.184566 (rank : 40) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9ULS6 | Gene names | KCNS2, KIAA1144 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily S member 2 (Voltage-gated potassium channel subunit Kv9.2) (Delayed-rectifier K(+) channel alpha subunit 2). | |||||
|
MAP1B_MOUSE
|
||||||
| θ value | 2.36792 (rank : 82) | NC score | 0.010640 (rank : 110) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 977 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P14873 | Gene names | Map1b, Mtap1b, Mtap5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) (MAP1.2) (MAP1(X)) [Contains: MAP1 light chain LC1]. | |||||
|
MYH6_HUMAN
|
||||||
| θ value | 2.36792 (rank : 83) | NC score | 0.007047 (rank : 126) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1426 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P13533, Q13943, Q14906, Q14907 | Gene names | MYH6, MYHCA | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
MYH6_MOUSE
|
||||||
| θ value | 2.36792 (rank : 84) | NC score | 0.007043 (rank : 127) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1501 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q02566, Q64258, Q64738 | Gene names | Myh6, Myhca | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
MYH7_HUMAN
|
||||||
| θ value | 2.36792 (rank : 85) | NC score | 0.006763 (rank : 131) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P12883, Q14836, Q14837, Q14904, Q16579, Q92679, Q9H1D5, Q9UMM8 | Gene names | MYH7, MYHCB | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle beta isoform (MyHC-beta). | |||||
|
TPR_HUMAN
|
||||||
| θ value | 2.36792 (rank : 86) | NC score | 0.008205 (rank : 117) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
ZC11A_HUMAN
|
||||||
| θ value | 2.36792 (rank : 87) | NC score | 0.012284 (rank : 106) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O75152, Q6AHY4, Q6AHY9, Q6AW79, Q6AWA1, Q6PJK4, Q86XZ7 | Gene names | ZC3H11A, KIAA0663 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 11A. | |||||
|
DRD1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 88) | NC score | -0.001323 (rank : 157) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 750 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P21728, Q4QRJ0 | Gene names | DRD1 | |||
|
Domain Architecture |
|
|||||
| Description | D(1A) dopamine receptor. | |||||
|
KIF3A_HUMAN
|
||||||
| θ value | 3.0926 (rank : 89) | NC score | 0.002446 (rank : 150) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9Y496, Q86XE9, Q9Y6V4 | Gene names | KIF3A, KIF3 | |||
|
Domain Architecture |
|
|||||
| Description | Kinesin-like protein KIF3A (Microtubule plus end-directed kinesin motor 3A). | |||||
|
NCKX1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 90) | NC score | 0.006104 (rank : 136) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O60721, O43485, O75184 | Gene names | SLC24A1, KIAA0702, NCKX1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium/calcium exchanger 1 (Na(+)/K(+)/Ca(2+)-exchange protein 1) (Retinal rod Na-Ca+K exchanger). | |||||
|
RXINP_MOUSE
|
||||||
| θ value | 3.0926 (rank : 91) | NC score | 0.010458 (rank : 111) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q5U5Q9, Q60811 | Gene names | Rxrip110, Rip110 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoid X receptor-interacting protein 110. | |||||
|
CP135_MOUSE
|
||||||
| θ value | 4.03905 (rank : 92) | NC score | 0.006995 (rank : 128) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1278 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q6P5D4, Q6A030 | Gene names | Cep135, Cep4, Kiaa0635 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 135 kDa (Cep135 protein) (Centrosomal protein 4). | |||||
|
CROCC_MOUSE
|
||||||
| θ value | 4.03905 (rank : 93) | NC score | 0.007946 (rank : 119) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1460 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8CJ40, Q7TQL2, Q80U01, Q8R0B9 | Gene names | Crocc, Kiaa0445 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rootletin (Ciliary rootlet coiled-coil protein). | |||||
|
CT2NL_HUMAN
|
||||||
| θ value | 4.03905 (rank : 94) | NC score | 0.011657 (rank : 108) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1059 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9P2B4, Q96B40 | Gene names | CTTNBP2NL, KIAA1433 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CTTNBP2 N-terminal-like protein. | |||||
|
ERC2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 95) | NC score | 0.006711 (rank : 133) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1156 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O15083, Q86TK4 | Gene names | ERC2, KIAA0378 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ERC protein 2. | |||||
|
ERC2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 96) | NC score | 0.006729 (rank : 132) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1166 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q6PH08, Q80U20, Q8CCP1 | Gene names | Erc2, Cast1, D14Ertd171e, Kiaa0378 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ERC protein 2 (CAZ-associated structural protein 1) (CAST1). | |||||
|
GPR64_HUMAN
|
||||||
| θ value | 4.03905 (rank : 97) | NC score | 0.002406 (rank : 152) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8IZP9, O00406, Q8IWT2, Q8IZE4, Q8IZE5, Q8IZE6, Q8IZE7, Q8IZP3, Q8IZP4 | Gene names | GPR64, HE6, TM7LN2 | |||
|
Domain Architecture |

|
|||||
| Description | G-protein coupled receptor 64 precursor (Epididymis-specific protein 6) (He6 receptor). | |||||
|
OAT6_MOUSE
|
||||||
| θ value | 4.03905 (rank : 98) | NC score | 0.004112 (rank : 144) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q80UJ1 | Gene names | Slc22a20, Gm963, Oat6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Solute carrier family 22 member 20 (Organic anion transporter 6). | |||||
|
PIGW_HUMAN
|
||||||
| θ value | 4.03905 (rank : 99) | NC score | 0.009703 (rank : 112) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7Z7B1, Q8N9G3 | Gene names | PIGW | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol-glycan biosynthesis class W protein (EC 2.3.-.-) (PIG-W). | |||||
|
PLEC1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 100) | NC score | 0.003399 (rank : 146) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1326 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q15149, Q15148, Q16640 | Gene names | PLEC1 | |||
|
Domain Architecture |

|
|||||
| Description | Plectin-1 (PLTN) (PCN) (Hemidesmosomal protein 1) (HD1). | |||||
|
SMC1A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 101) | NC score | 0.006939 (rank : 130) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1346 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q14683, O14995, Q16351 | Gene names | SMC1L1, DXS423E, KIAA0178, SB1.8, SMC1 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Sb1.8). | |||||
|
SMC1A_MOUSE
|
||||||
| θ value | 4.03905 (rank : 102) | NC score | 0.006657 (rank : 134) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1384 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9CU62, Q3V480, Q9CUX9, Q9D959, Q9WTU1 | Gene names | Smc1l1, Sb1.8, Smc1, Smcb | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Chromosome segregation protein SmcB) (Sb1.8). | |||||
|
TRPV1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 103) | NC score | 0.021245 (rank : 96) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q704Y3, Q5SSE1, Q5SSE2, Q5SSE4, Q5WPV5, Q68SW0 | Gene names | Trpv1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 1 (TrpV1) (osm-9-like TRP channel 1) (OTRPC1) (Vanilloid receptor 1). | |||||
|
CALM_HUMAN
|
||||||
| θ value | 5.27518 (rank : 104) | NC score | 0.002530 (rank : 148) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P62158, P02593, P70667, P99014, Q13942, Q53S29, Q61379, Q61380, Q96HK3 | Gene names | CALM1, CALM, CAM, CAM1 | |||
|
Domain Architecture |
|
|||||
| Description | Calmodulin (CaM). | |||||
|
CALM_MOUSE
|
||||||
| θ value | 5.27518 (rank : 105) | NC score | 0.002530 (rank : 149) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P62204, P02593, P70667, P99014, Q61379, Q61380 | Gene names | Calm1, Calm, Cam, Cam1 | |||
|
Domain Architecture |
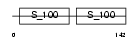
|
|||||
| Description | Calmodulin (CaM). | |||||
|
CKAP4_MOUSE
|
||||||
| θ value | 5.27518 (rank : 106) | NC score | 0.007957 (rank : 118) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 596 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8BMK4, Q8BTK8, Q8R3F2 | Gene names | Ckap4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-associated protein 4 (63 kDa membrane protein) (p63). | |||||
|
ITPR1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 107) | NC score | 0.009390 (rank : 114) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14643, Q14660, Q99897 | Gene names | ITPR1, INSP3R1 | |||
|
Domain Architecture |

|
|||||
| Description | Inositol 1,4,5-trisphosphate receptor type 1 (Type 1 inositol 1,4,5- trisphosphate receptor) (Type 1 InsP3 receptor) (IP3 receptor isoform 1) (InsP3R1) (IP3R). | |||||
|
ITPR1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 108) | NC score | 0.009375 (rank : 115) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P11881, P20943, Q99LG5 | Gene names | Itpr1, Insp3r, Pcd6, Pcp1 | |||
|
Domain Architecture |

|
|||||
| Description | Inositol 1,4,5-trisphosphate receptor type 1 (Type 1 inositol 1,4,5- trisphosphate receptor) (Type 1 InsP3 receptor) (IP3 receptor isoform 1) (InsP3R1) (Inositol 1,4,5-trisphosphate-binding protein P400) (Purkinje cell protein 1) (Protein PCD-6). | |||||
|
KCMA1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 109) | NC score | 0.031710 (rank : 92) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q12791, Q12886, Q12917, Q12921, Q12960, Q13150, Q96LG8, Q9UBB0, Q9UQK6 | Gene names | KCNMA1, KCNMA | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-activated potassium channel subunit alpha 1 (Calcium-activated potassium channel, subfamily M subunit alpha 1) (Maxi K channel) (MaxiK) (BK channel) (K(VCA)alpha) (BKCA alpha) (KCa1.1) (Slowpoke homolog) (Slo homolog) (Slo-alpha) (Slo1) (hSlo). | |||||
|
KCMA1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 110) | NC score | 0.031766 (rank : 91) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q08460, Q64703, Q8VHF1, Q9R196 | Gene names | Kcnma1, Kcnma | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-activated potassium channel subunit alpha 1 (Calcium-activated potassium channel, subfamily M subunit alpha 1) (Maxi K channel) (MaxiK) (BK channel) (K(VCA)alpha) (BKCA alpha) (KCa1.1) (Slowpoke homolog) (Slo homolog) (Slo-alpha) (Slo1) (mSlo) (mSlo1). | |||||
|
KCNS1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 111) | NC score | 0.106632 (rank : 55) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | O35173 | Gene names | Kcns1 | |||
|
Domain Architecture |
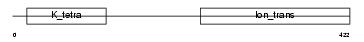
|
|||||
| Description | Potassium voltage-gated channel subfamily S member 1 (Voltage-gated potassium channel subunit Kv9.1) (Delayed-rectifier K(+) channel alpha subunit 1). | |||||
|
MYH4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 112) | NC score | 0.007869 (rank : 120) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1377 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9Y623 | Gene names | MYH4 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-4 (Myosin heavy chain, skeletal muscle, fetal) (Myosin heavy chain IIb) (MyHC-IIb). | |||||
|
MYH9_HUMAN
|
||||||
| θ value | 5.27518 (rank : 113) | NC score | 0.005688 (rank : 137) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P35579, O60805 | Gene names | MYH9 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
SEGN_MOUSE
|
||||||
| θ value | 5.27518 (rank : 114) | NC score | 0.016543 (rank : 103) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q91WD9 | Gene names | Scgn | |||
|
Domain Architecture |

|
|||||
| Description | Secretagogin. | |||||
|
SMC4_MOUSE
|
||||||
| θ value | 5.27518 (rank : 115) | NC score | 0.007608 (rank : 124) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 830 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8CG47, Q8BTS7, Q8BTY9, Q99K21 | Gene names | Smc4l1, Capc, Smc4 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosomes 4-like 1 protein (Chromosome- associated polypeptide C) (XCAP-C homolog). | |||||
|
DSPP_HUMAN
|
||||||
| θ value | 6.88961 (rank : 116) | NC score | 0.002415 (rank : 151) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
KCNC3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 117) | NC score | 0.104609 (rank : 57) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q14003 | Gene names | KCNC3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 3 (Voltage-gated potassium channel subunit Kv3.3) (KSHIIID). | |||||
|
MAP1B_HUMAN
|
||||||
| θ value | 6.88961 (rank : 118) | NC score | 0.007780 (rank : 122) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P46821 | Gene names | MAP1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) [Contains: MAP1 light chain LC1]. | |||||
|
NKTR_HUMAN
|
||||||
| θ value | 6.88961 (rank : 119) | NC score | 0.004460 (rank : 142) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 658 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P30414 | Gene names | NKTR | |||
|
Domain Architecture |

|
|||||
| Description | NK-tumor recognition protein (Natural-killer cells cyclophilin-related protein) (NK-TR protein). | |||||
|
RB6I2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 120) | NC score | 0.007743 (rank : 123) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1584 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8IUD2, Q6NVK2, Q8IUD3, Q8IUD4, Q8IUD5, Q8NAS1, Q9NXN5, Q9UIK7, Q9UPS1 | Gene names | ERC1, ELKS, KIAA1081, RAB6IP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ELKS/RAB6-interacting/CAST family member 1 (RAB6-interacting protein 2) (ERC protein 1). | |||||
|
TRPM2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 121) | NC score | 0.019774 (rank : 97) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q91YD4 | Gene names | Trpm2, Ltrpc2, Trpc7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily M member 2 (EC 3.6.1.13) (Long transient receptor potential channel 2) (LTrpC2) (LTrpC-2) (Transient receptor potential channel 7) (TrpC7). | |||||
|
TRPM8_HUMAN
|
||||||
| θ value | 6.88961 (rank : 122) | NC score | 0.035930 (rank : 90) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q7Z2W7, Q6QNH9, Q8TAC3, Q8TDX8, Q9BVK1 | Gene names | TRPM8, LTRPC6, TRPP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily M member 8 (Transient receptor potential-p8) (Trp-p8) (Long transient receptor potential channel 6) (LTrpC6). | |||||
|
CENPF_HUMAN
|
||||||
| θ value | 8.99809 (rank : 123) | NC score | 0.003758 (rank : 145) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1932 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P49454, Q13171, Q13246 | Gene names | CENPF | |||
|
Domain Architecture |

|
|||||
| Description | Centromere protein F (Kinetochore protein CENP-F) (Mitosin) (AH antigen). | |||||
|
CF152_HUMAN
|
||||||
| θ value | 8.99809 (rank : 124) | NC score | 0.006983 (rank : 129) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q86VQ0, Q9BWX7 | Gene names | C6orf152 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf152. | |||||
|
HUWE1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 125) | NC score | 0.002016 (rank : 154) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q7Z6Z7, O15029, Q4G2Z2, Q5H961, Q6P4D0, Q8NG67, Q9BUI0, Q9HCJ4, Q9NSL6, Q9P0A9 | Gene names | HUWE1, KIAA0312, KIAA1578, UREB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (Mcl-1 ubiquitin ligase E3) (Mule) (ARF-binding protein 1) (ARF-BP1). | |||||
|
KCNB1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 126) | NC score | 0.176887 (rank : 42) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q14721, Q14193 | Gene names | KCNB1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily B member 1 (Voltage-gated potassium channel subunit Kv2.1) (h-DRK1). | |||||
|
KCNB1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 127) | NC score | 0.177025 (rank : 41) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q03717 | Gene names | Kcnb1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily B member 1 (Voltage-gated potassium channel subunit Kv2.1) (mShab). | |||||
|
KCNC3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 128) | NC score | 0.102305 (rank : 58) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q63959, Q62088 | Gene names | Kcnc3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 3 (Voltage-gated potassium channel subunit Kv3.3) (KSHIIID). | |||||
|
MOR2A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 129) | NC score | 0.002304 (rank : 153) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q69ZX6, Q5QNQ7, Q6P547 | Gene names | Morc2a, Kiaa0852, Zcwcc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORC family CW-type zinc finger protein 2A (Zinc finger CW-type coiled-coil domain protein 1). | |||||
|
MYH10_HUMAN
|
||||||
| θ value | 8.99809 (rank : 130) | NC score | 0.005413 (rank : 139) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P35580, Q16087 | Gene names | MYH10 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
MYH10_MOUSE
|
||||||
| θ value | 8.99809 (rank : 131) | NC score | 0.005279 (rank : 140) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1612 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q61879, Q5SV63 | Gene names | Myh10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
MYH9_MOUSE
|
||||||
| θ value | 8.99809 (rank : 132) | NC score | 0.005206 (rank : 141) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1829 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8VDD5 | Gene names | Myh9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
TMCC3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 133) | NC score | 0.006352 (rank : 135) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9ULS5, Q8IWB2 | Gene names | TMCC3, KIAA1145 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane and coiled-coil domains protein 3. | |||||
|
WWP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 134) | NC score | 0.004304 (rank : 143) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9H0M0, O00307, Q96BP4 | Gene names | WWP1 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP1 (EC 6.3.2.-) (WW domain- containing protein 1) (Atropin-1-interacting protein 5) (AIP5). | |||||
|
WWP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 135) | NC score | 0.001973 (rank : 155) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BZZ3, Q8BIV9, Q8VDP8 | Gene names | Wwp1 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP1 (EC 6.3.2.-) (WW domain- containing protein 1). | |||||
|
CABP5_MOUSE
|
||||||
| θ value | θ > 10 (rank : 136) | NC score | 0.052512 (rank : 85) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9JLK3 | Gene names | Cabp5 | |||
|
Domain Architecture |
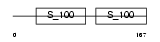
|
|||||
| Description | Calcium-binding protein 5 (CaBP5). | |||||
|
KCND1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 137) | NC score | 0.134626 (rank : 49) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9NSA2, O75671 | Gene names | KCND1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 1 (Voltage-gated potassium channel subunit Kv4.1). | |||||
|
KCND1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 138) | NC score | 0.136555 (rank : 48) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q03719, Q8CC68 | Gene names | Kcnd1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 1 (Voltage-gated potassium channel subunit Kv4.1) (mShal). | |||||
|
KCNG1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 139) | NC score | 0.053167 (rank : 83) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9UIX4, O43528, Q9BRC1 | Gene names | KCNG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily G member 1 (Voltage-gated potassium channel subunit Kv6.1) (kH2). | |||||
|
KCNG3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 140) | NC score | 0.078435 (rank : 65) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q8TAE7 | Gene names | KCNG3 | |||
|
Domain Architecture |
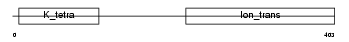
|
|||||
| Description | Potassium voltage-gated channel subfamily G member 3 (Voltage-gated potassium channel subunit Kv6.3) (Kv10.1). | |||||
|
KCNG3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 141) | NC score | 0.078657 (rank : 64) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P59053 | Gene names | Kcng3 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium voltage-gated channel subfamily G member 3 (Voltage-gated potassium channel subunit Kv6.3) (Kv10.1). | |||||
|
KCNQ1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 142) | NC score | 0.055557 (rank : 82) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P51787, O00347, O60607, O94787, Q7Z6G9, Q92960, Q9UMN8, Q9UMN9 | Gene names | KCNQ1, KCNA8, KCNA9, KVLQT1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 1 (Voltage-gated potassium channel subunit Kv7.1) (IKs producing slow voltage-gated potassium channel subunit alpha KvLQT1) (KQT-like 1). | |||||
|
KCNQ1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 143) | NC score | 0.064037 (rank : 76) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P97414, O88702, Q7TNZ1, Q7TPL7, Q80VR7 | Gene names | Kcnq1, Kcna9, Kvlqt1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 1 (Voltage-gated potassium channel subunit Kv7.1) (IKs producing slow voltage-gated potassium channel subunit alpha KvLQT1) (KQT-like 1). | |||||
|
KCNQ5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 144) | NC score | 0.051970 (rank : 86) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9NR82, Q9NRN0, Q9NYA6 | Gene names | KCNQ5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 5 (Voltage-gated potassium channel subunit Kv7.5) (Potassium channel subunit alpha KvLQT5) (KQT-like 5). | |||||
|
KCNS1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 145) | NC score | 0.123835 (rank : 53) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q96KK3, O43652, Q6DJU6 | Gene names | KCNS1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily S member 1 (Voltage-gated potassium channel subunit Kv9.1) (Delayed-rectifier K(+) channel alpha subunit 1). | |||||
|
KCNS3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 146) | NC score | 0.156037 (rank : 45) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9BQ31, O43651, Q96B56 | Gene names | KCNS3 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium voltage-gated channel subfamily S member 3 (Voltage-gated potassium channel subunit Kv9.3) (Delayed-rectifier K(+) channel alpha subunit 3). | |||||
|
KCNV2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 147) | NC score | 0.197135 (rank : 38) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q8TDN2 | Gene names | KCNV2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily V member 2 (Voltage-gated potassium channel subunit Kv8.2). | |||||
|
PK2L2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 148) | NC score | 0.250235 (rank : 33) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9JLG4 | Gene names | Pkd2l2 | |||
|
Domain Architecture |

|
|||||
| Description | Polycystic kidney disease 2-like 2 protein (Polycystin-L2). | |||||
|
PKDRE_HUMAN
|
||||||
| θ value | θ > 10 (rank : 149) | NC score | 0.097757 (rank : 59) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9NTG1, O95850 | Gene names | PKDREJ | |||
|
Domain Architecture |

|
|||||
| Description | Polycystic kidney disease and receptor for egg jelly-related protein precursor (PKD and REJ homolog). | |||||
|
PKDRE_MOUSE
|
||||||
| θ value | θ > 10 (rank : 150) | NC score | 0.070325 (rank : 72) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9Z0T6 | Gene names | Pkdrej | |||
|
Domain Architecture |

|
|||||
| Description | Polycystic kidney disease and receptor for egg jelly-related protein precursor (PKD and REJ homolog). | |||||
|
TPTE2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 151) | NC score | 0.129866 (rank : 50) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6XPS3, Q5VUH2, Q8WWL4, Q8WWL5 | Gene names | TPTE2, TPIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol-3,4,5-trisphosphate 3-phosphatase TPTE2 (EC 3.1.3.67) (TPTE and PTEN homologous inositol lipid phosphatase) (Lipid phosphatase TPIP). | |||||
|
TPTE_HUMAN
|
||||||
| θ value | θ > 10 (rank : 152) | NC score | 0.086284 (rank : 60) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P56180, Q6XPS4, Q6XPS5, Q71JA8, Q8NCS8 | Gene names | TPTE | |||
|
Domain Architecture |

|
|||||
| Description | Putative tyrosine-protein phosphatase TPTE (EC 3.1.3.48) (Transmembrane phosphatase with tensin homology) (Protein BJ-HCC-5). | |||||
|
TRPC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 153) | NC score | 0.057140 (rank : 78) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P48995 | Gene names | TRPC1, TRP1 | |||
|
Domain Architecture |

|
|||||
| Description | Short transient receptor potential channel 1 (TrpC1) (TRP-1 protein). | |||||
|
TRPC1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 154) | NC score | 0.057228 (rank : 77) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q61056, O35722 | Gene names | Trpc1, Trp1, Trrp1 | |||
|
Domain Architecture |

|
|||||
| Description | Short transient receptor potential channel 1 (TrpC1) (Transient receptor protein 1) (Mtrp1) (Trp-related protein 1). | |||||
|
TRPV3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 155) | NC score | 0.056818 (rank : 79) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8K424, Q5SV08 | Gene names | Trpv3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 3 (TrpV3). | |||||
|
TRPV4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 156) | NC score | 0.051454 (rank : 87) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9HBA0, Q86YZ6, Q8NDY7, Q8NG64, Q96Q92, Q96RS7, Q9HBC0 | Gene names | TRPV4, VRL2, VROAC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 4 (TrpV4) (osm-9-like TRP channel 4) (OTRPC4) (Vanilloid receptor-like channel 2) (Vanilloid receptor-like protein 2) (VRL-2) (Vanilloid receptor-related osmotically-activated channel) (VR-OAC) (Transient receptor potential protein 12) (TRP12). | |||||
|
TRPV4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 157) | NC score | 0.052805 (rank : 84) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9EPK8, Q91XR5, Q9EQZ4, Q9ERZ7, Q9ES76 | Gene names | Trpv4, Trp12, Vrl2, Vroac | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 4 (TrpV4) (osm-9-like TRP channel 4) (OTRPC4) (Vanilloid receptor-like channel 2) (Vanilloid receptor-like protein 2) (Vanilloid receptor- related osmotically-activated channel) (VR-OAC) (Transient receptor potential protein 12) (TRP12). | |||||
|
SCN7A_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 10) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 135 | |
| SwissProt Accessions | Q01118 | Gene names | SCN7A, SCN6A | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 7 subunit alpha (Sodium channel protein type VII subunit alpha) (Putative voltage-gated sodium channel alpha subunit Nax) (Sodium channel protein, cardiac and skeletal muscle alpha-subunit). | |||||
|
SCN4A_HUMAN
|
||||||
| NC score | 0.989721 (rank : 2) | θ value | 0 (rank : 8) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | P35499, Q15478, Q16447, Q7Z6B1 | Gene names | SCN4A | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 4 subunit alpha (Sodium channel protein type IV subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.4) (Sodium channel protein, skeletal muscle subunit alpha) (SkM1). | |||||
|
SCN3A_HUMAN
|
||||||
| NC score | 0.986449 (rank : 3) | θ value | 0 (rank : 7) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q9NY46, Q16142, Q9BZB3, Q9C006, Q9NYK2, Q9UPD1, Q9Y6P4 | Gene names | SCN3A, NAC3 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 3 subunit alpha (Sodium channel protein type III subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.3) (Sodium channel protein, brain III subunit alpha) (Voltage- gated sodium channel subtype III). | |||||
|
SCN2A_HUMAN
|
||||||
| NC score | 0.985672 (rank : 4) | θ value | 0 (rank : 6) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q99250, Q14472, Q9BZC9, Q9BZD0 | Gene names | SCN2A, NAC2, SCN2A2 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 2 subunit alpha (Sodium channel protein type II subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.2) (Sodium channel protein, brain II subunit alpha) (HBSC II). | |||||
|
SCN8A_HUMAN
|
||||||
| NC score | 0.985606 (rank : 5) | θ value | 0 (rank : 11) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9UQD0, O95788, Q9NYX2, Q9UPB2 | Gene names | SCN8A | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 8 subunit alpha (Sodium channel protein type VIII subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.6). | |||||
|
SCN1A_HUMAN
|
||||||
| NC score | 0.985313 (rank : 6) | θ value | 0 (rank : 5) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P35498, Q16172, Q96LA3, Q9C008 | Gene names | SCN1A, NAC1, SCN1 | |||
|
Domain Architecture |
|
|||||
| Description | Sodium channel protein type 1 subunit alpha (Sodium channel protein type I subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.1) (Sodium channel protein, brain I subunit alpha). | |||||
|
SCN8A_MOUSE
|
||||||
| NC score | 0.985297 (rank : 7) | θ value | 0 (rank : 12) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q9WTU3, Q60828, Q60858, Q62449 | Gene names | Scn8a, Nbna1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 8 subunit alpha (Sodium channel protein type VIII subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.6). | |||||
|
SCN9A_HUMAN
|
||||||
| NC score | 0.985220 (rank : 8) | θ value | 0 (rank : 13) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q15858, Q6B4R9, Q6B4S0, Q6B4S1, Q70HX1, Q70HX2, Q8WTU1, Q8WWN4 | Gene names | SCN9A, NENA, PN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium channel protein type 9 subunit alpha (Sodium channel protein type IX subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.7) (Neuroendocrine sodium channel) (hNE-Na) (Peripheral sodium channel 1). | |||||
|
SC11A_MOUSE
|
||||||
| NC score | 0.984827 (rank : 9) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q9R053, Q9JMD4 | Gene names | Scn11a, Nan, Nat, Sns2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium channel protein type 11 subunit alpha (Sodium channel protein type XI subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.9) (Sensory neuron sodium channel 2) (NaN). | |||||
|
SC11A_HUMAN
|
||||||
| NC score | 0.983137 (rank : 10) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9UI33, Q68K15, Q8NDX3, Q9UHE0, Q9UHM0 | Gene names | SCN11A, PN5, SCN12A, SNS2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium channel protein type 11 subunit alpha (Sodium channel protein type XI subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.9) (Sensory neuron sodium channel 2) (Peripheral nerve sodium channel 5) (hNaN). | |||||
|
SC10A_HUMAN
|
||||||
| NC score | 0.981513 (rank : 11) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9Y5Y9 | Gene names | SCN10A, PN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium channel protein type 10 subunit alpha (Sodium channel protein type X subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.8) (Peripheral nerve sodium channel 3) (hPN3). | |||||
|
SC10A_MOUSE
|
||||||
| NC score | 0.980622 (rank : 12) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q6QIY3, Q62243, Q6EWG7, Q6KCH7, Q703F9 | Gene names | Scn10a, Pn3, Sns | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium channel protein type 10 subunit alpha (Sodium channel protein type X subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.8) (Peripheral nerve sodium channel 3) (Sensory neuron sodium channel). | |||||
|
SCN5A_HUMAN
|
||||||
| NC score | 0.978532 (rank : 13) | θ value | 0 (rank : 9) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q14524 | Gene names | SCN5A | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 5 subunit alpha (Sodium channel protein type V subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.5) (Sodium channel protein, cardiac muscle alpha-subunit) (HH1). | |||||
|
CAC1I_HUMAN
|
||||||
| NC score | 0.895687 (rank : 14) | θ value | 3.84025e-59 (rank : 18) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9P0X4, O95504, Q5JZ88, Q7Z6S9, Q8NFX6, Q9NZC8, Q9UH15, Q9UH30, Q9ULU9, Q9UNE6 | Gene names | CACNA1I, KIAA1120 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1I (Voltage- gated calcium channel subunit alpha Cav3.3) (Ca(v)3.3). | |||||
|
CAC1H_MOUSE
|
||||||
| NC score | 0.876186 (rank : 15) | θ value | 1.7238e-59 (rank : 17) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | O88427, Q80TJ2, Q9JKU5 | Gene names | Cacna1h, Kiaa1120 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1H (Voltage- gated calcium channel subunit alpha Cav3.2). | |||||
|
CAC1H_HUMAN
|
||||||
| NC score | 0.869295 (rank : 16) | θ value | 1.31987e-59 (rank : 16) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | O95180, O95802, Q8WWI6, Q96QI6, Q96RZ9, Q9NYY4, Q9NYY5 | Gene names | CACNA1H | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1H (Voltage- gated calcium channel subunit alpha Cav3.2) (Low-voltage-activated calcium channel alpha1 3.2 subunit). | |||||
|
CAC1E_MOUSE
|
||||||
| NC score | 0.862984 (rank : 17) | θ value | 8.55514e-59 (rank : 19) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q61290 | Gene names | Cacna1e, Cach6, Cacnl1a6, Cchra1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent R-type calcium channel subunit alpha-1E (Voltage- gated calcium channel subunit alpha Cav2.3) (Calcium channel, L type, alpha-1 polypeptide, isoform 6) (Brain calcium channel II) (BII). | |||||
|
CAC1E_HUMAN
|
||||||
| NC score | 0.862604 (rank : 18) | θ value | 1.45929e-58 (rank : 20) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q15878, Q14580, Q14581 | Gene names | CACNA1E, CACH6, CACNL1A6 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent R-type calcium channel subunit alpha-1E (Voltage- gated calcium channel subunit alpha Cav2.3) (Calcium channel, L type, alpha-1 polypeptide, isoform 6) (Brain calcium channel II) (BII). | |||||
|
CAC1A_MOUSE
|
||||||
| NC score | 0.859059 (rank : 19) | θ value | 2.48917e-58 (rank : 22) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | P97445 | Gene names | Cacna1a, Cach4, Cacn3, Cacnl1a4, Ccha1a | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent P/Q-type calcium channel subunit alpha-1A (Voltage- gated calcium channel subunit alpha Cav2.1) (Calcium channel, L type, alpha-1 polypeptide isoform 4) (Brain calcium channel I) (BI). | |||||
|
CAC1S_MOUSE
|
||||||
| NC score | 0.856526 (rank : 20) | θ value | 3.36421e-55 (rank : 28) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q02789, Q99240 | Gene names | Cacna1s, Cach1, Cach1b, Cacn1, Cacnl1a3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1S (Voltage- gated calcium channel subunit alpha Cav1.1) (Calcium channel, L type, alpha-1 polypeptide, isoform 3, skeletal muscle). | |||||
|
CAC1D_HUMAN
|
||||||
| NC score | 0.855685 (rank : 21) | θ value | 1.36585e-56 (rank : 25) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q01668, Q13916, Q13931 | Gene names | CACNA1D, CACH3, CACN4, CACNL1A2, CCHL1A2 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1D (Voltage- gated calcium channel subunit alpha Cav1.3) (Calcium channel, L type, alpha-1 polypeptide, isoform 2). | |||||
|
CAC1S_HUMAN
|
||||||
| NC score | 0.855059 (rank : 22) | θ value | 2.18062e-54 (rank : 30) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q13698, Q12896, Q13934 | Gene names | CACNA1S, CACH1, CACN1, CACNL1A3 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1S (Voltage- gated calcium channel subunit alpha Cav1.1) (Calcium channel, L type, alpha-1 polypeptide, isoform 3, skeletal muscle). | |||||
|
CAC1B_HUMAN
|
||||||
| NC score | 0.853346 (rank : 23) | θ value | 1.66964e-54 (rank : 29) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q00975 | Gene names | CACNA1B, CACH5, CACNL1A5 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent N-type calcium channel subunit alpha-1B (Voltage- gated calcium channel subunit alpha Cav2.2) (Calcium channel, L type, alpha-1 polypeptide isoform 5) (Brain calcium channel III) (BIII). | |||||
|
CAC1G_HUMAN
|
||||||
| NC score | 0.852488 (rank : 24) | θ value | 8.00737e-57 (rank : 24) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 587 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | O43497, O43498, O94770, Q9NYU4, Q9NYU5, Q9NYU6, Q9NYU7, Q9NYU8, Q9NYU9, Q9NYV0, Q9NYV1, Q9UHN9, Q9UHP0, Q9ULU6, Q9UNG7, Q9Y5T2, Q9Y5T3 | Gene names | CACNA1G, KIAA1123 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1G (Voltage- gated calcium channel subunit alpha Cav3.1) (Cav3.1c) (NBR13). | |||||
|
CAC1C_MOUSE
|
||||||
| NC score | 0.851684 (rank : 25) | θ value | 5.67233e-95 (rank : 14) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q01815, Q04476, Q61242, Q99242 | Gene names | Cacna1c, Cach2, Cacn2, Cacnl1a1, Cchl1a1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1C (Voltage- gated calcium channel subunit alpha Cav1.2) (Calcium channel, L type, alpha-1 polypeptide, isoform 1, cardiac muscle) (Mouse brain class C) (MBC) (MELC-CC). | |||||
|
CAC1A_HUMAN
|
||||||
| NC score | 0.851338 (rank : 26) | θ value | 2.48917e-58 (rank : 21) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | O00555, P78510, P78511, Q16290, Q92690, Q99790, Q99791, Q99792, Q99793 | Gene names | CACNA1A, CACH4, CACN3, CACNL1A4 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent P/Q-type calcium channel subunit alpha-1A (Voltage- gated calcium channel subunit alpha Cav2.1) (Calcium channel, L type, alpha-1 polypeptide isoform 4) (Brain calcium channel I) (BI). | |||||
|
CAC1F_MOUSE
|
||||||
| NC score | 0.850362 (rank : 27) | θ value | 3.59435e-57 (rank : 23) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9JIS7 | Gene names | Cacna1f | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1F (Voltage- gated calcium channel subunit alpha Cav1.4). | |||||
|
CAC1B_MOUSE
|
||||||
| NC score | 0.850112 (rank : 28) | θ value | 8.85319e-56 (rank : 27) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | O55017, Q60609 | Gene names | Cacna1b, Cach5, Cacnl1a5, Cchn1a | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent N-type calcium channel subunit alpha-1B (Voltage- gated calcium channel subunit alpha Cav2.2) (Calcium channel, L type, alpha-1 polypeptide isoform 5) (Brain calcium channel III) (BIII). | |||||
|
CAC1F_HUMAN
|
||||||
| NC score | 0.849894 (rank : 29) | θ value | 1.78386e-56 (rank : 26) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O60840, O43901 | Gene names | CACNA1F, CACNAF1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1F (Voltage- gated calcium channel subunit alpha Cav1.4). | |||||
|
CAC1C_HUMAN
|
||||||
| NC score | 0.848490 (rank : 30) | θ value | 1.88829e-90 (rank : 15) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q13936, Q13917, Q13918, Q13919, Q13920, Q13921, Q13922, Q13923, Q13924, Q13925, Q13926, Q13927, Q13928, Q13929, Q13930, Q13932, Q13933, Q14743, Q14744, Q15877, Q99025, Q99241, Q99875 | Gene names | CACNA1C, CACH2, CACN2, CACNL1A1, CCHL1A1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1C (Voltage- gated calcium channel subunit alpha Cav1.2) (Calcium channel, L type, alpha-1 polypeptide, isoform 1, cardiac muscle). | |||||
|
PK2L2_HUMAN
|
||||||
| NC score | 0.286844 (rank : 31) | θ value | 1.81305 (rank : 75) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9NZM6, Q9UNJ0 | Gene names | PKD2L2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polycystic kidney disease 2-like 2 protein (Polycystin-L2). | |||||
|
PK2L1_HUMAN
|
||||||
| NC score | 0.267650 (rank : 32) | θ value | 0.163984 (rank : 42) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9P0L9, O75972, Q9UP35, Q9UPA2 | Gene names | PKD2L1, PKD2L, PKDL | |||
|
Domain Architecture |

|
|||||
| Description | Polycystic kidney disease 2-like 1 protein (Polycystin-L) (Polycystin- 2 homolog). | |||||
|
PK2L2_MOUSE
|
||||||
| NC score | 0.250235 (rank : 33) | θ value | θ > 10 (rank : 148) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9JLG4 | Gene names | Pkd2l2 | |||
|
Domain Architecture |

|
|||||
| Description | Polycystic kidney disease 2-like 2 protein (Polycystin-L2). | |||||
|
PKD2_MOUSE
|
||||||
| NC score | 0.248608 (rank : 34) | θ value | 0.47712 (rank : 47) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O35245, Q9QWP0, Q9Z193, Q9Z194 | Gene names | Pkd2 | |||
|
Domain Architecture |

|
|||||
| Description | Polycystin-2 (Polycystic kidney disease 2 protein homolog). | |||||
|
PKD2_HUMAN
|
||||||
| NC score | 0.245715 (rank : 35) | θ value | 1.38821 (rank : 64) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q13563, O60441, Q15764 | Gene names | PKD2 | |||
|
Domain Architecture |

|
|||||
| Description | Polycystin-2 (Polycystic kidney disease 2 protein homolog) (Autosomal dominant polycystic kidney disease type II protein) (Polycystwin) (R48321). | |||||
|
KCNF1_HUMAN
|
||||||
| NC score | 0.217548 (rank : 36) | θ value | 0.0113563 (rank : 31) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9H3M0, O43527 | Gene names | KCNF1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily F member 1 (Voltage-gated potassium channel subunit Kv5.1) (kH1). | |||||
|
KCNB2_HUMAN
|
||||||
| NC score | 0.197786 (rank : 37) | θ value | 0.0563607 (rank : 37) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q92953, Q9BXD3 | Gene names | KCNB2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily B member 2 (Voltage-gated potassium channel subunit Kv2.2). | |||||
|
KCNV2_HUMAN
|
||||||
| NC score | 0.197135 (rank : 38) | θ value | θ > 10 (rank : 147) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q8TDN2 | Gene names | KCNV2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily V member 2 (Voltage-gated potassium channel subunit Kv8.2). | |||||
|
KCNS2_MOUSE
|
||||||
| NC score | 0.189779 (rank : 39) | θ value | 1.06291 (rank : 56) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | O35174, Q543P3 | Gene names | Kcns2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily S member 2 (Voltage-gated potassium channel subunit Kv9.2) (Delayed-rectifier K(+) channel alpha subunit 2). | |||||
|
KCNS2_HUMAN
|
||||||
| NC score | 0.184566 (rank : 40) | θ value | 2.36792 (rank : 81) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9ULS6 | Gene names | KCNS2, KIAA1144 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily S member 2 (Voltage-gated potassium channel subunit Kv9.2) (Delayed-rectifier K(+) channel alpha subunit 2). | |||||
|
KCNB1_MOUSE
|
||||||
| NC score | 0.177025 (rank : 41) | θ value | 8.99809 (rank : 127) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q03717 | Gene names | Kcnb1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily B member 1 (Voltage-gated potassium channel subunit Kv2.1) (mShab). | |||||
|
KCNB1_HUMAN
|
||||||
| NC score | 0.176887 (rank : 42) | θ value | 8.99809 (rank : 126) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q14721, Q14193 | Gene names | KCNB1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily B member 1 (Voltage-gated potassium channel subunit Kv2.1) (h-DRK1). | |||||
|
KCND3_MOUSE
|
||||||
| NC score | 0.163118 (rank : 43) | θ value | 0.163984 (rank : 41) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9Z0V1, Q8CC44, Q9Z0V0 | Gene names | Kcnd3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 3 (Voltage-gated potassium channel subunit Kv4.3). | |||||
|
KCND3_HUMAN
|
||||||
| NC score | 0.162978 (rank : 44) | θ value | 0.163984 (rank : 40) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9UK17, O60576, O60577, Q5T0M0, Q9UH85, Q9UH86, Q9UK16 | Gene names | KCND3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 3 (Voltage-gated potassium channel subunit Kv4.3). | |||||
|
KCNS3_HUMAN
|
||||||
| NC score | 0.156037 (rank : 45) | θ value | θ > 10 (rank : 146) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9BQ31, O43651, Q96B56 | Gene names | KCNS3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily S member 3 (Voltage-gated potassium channel subunit Kv9.3) (Delayed-rectifier K(+) channel alpha subunit 3). | |||||
|
KCND2_MOUSE
|
||||||
| NC score | 0.155276 (rank : 46) | θ value | 0.62314 (rank : 50) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9Z0V2, Q8BSK3, Q8CHB7, Q9JJ60 | Gene names | Kcnd2, Kiaa1044 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 2 (Voltage-gated potassium channel subunit Kv4.2). | |||||
|
KCND2_HUMAN
|
||||||
| NC score | 0.155114 (rank : 47) | θ value | 0.62314 (rank : 49) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9NZV8, O95012, O95021, Q9UBY7, Q9UN98, Q9UNH9 | Gene names | KCND2, KIAA1044 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 2 (Voltage-gated potassium channel subunit Kv4.2). | |||||
|
KCND1_MOUSE
|
||||||
| NC score | 0.136555 (rank : 48) | θ value | θ > 10 (rank : 138) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q03719, Q8CC68 | Gene names | Kcnd1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 1 (Voltage-gated potassium channel subunit Kv4.1) (mShal). | |||||
|
KCND1_HUMAN
|
||||||
| NC score | 0.134626 (rank : 49) | θ value | θ > 10 (rank : 137) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9NSA2, O75671 | Gene names | KCND1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 1 (Voltage-gated potassium channel subunit Kv4.1). | |||||
|
TPTE2_HUMAN
|
||||||
| NC score | 0.129866 (rank : 50) | θ value | θ > 10 (rank : 151) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6XPS3, Q5VUH2, Q8WWL4, Q8WWL5 | Gene names | TPTE2, TPIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol-3,4,5-trisphosphate 3-phosphatase TPTE2 (EC 3.1.3.67) (TPTE and PTEN homologous inositol lipid phosphatase) (Lipid phosphatase TPIP). | |||||
|
KCNC1_HUMAN
|
||||||
| NC score | 0.128919 (rank : 51) | θ value | 1.81305 (rank : 74) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P48547 | Gene names | KCNC1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 1 (Voltage-gated potassium channel subunit Kv3.1) (Kv4) (NGK2). | |||||
|
KCNC1_MOUSE
|
||||||
| NC score | 0.126614 (rank : 52) | θ value | 2.36792 (rank : 80) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P15388 | Gene names | Kcnc1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 1 (Voltage-gated potassium channel subunit Kv3.1) (Kv4) (NGK2). | |||||
|
KCNS1_HUMAN
|
||||||
| NC score | 0.123835 (rank : 53) | θ value | θ > 10 (rank : 145) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q96KK3, O43652, Q6DJU6 | Gene names | KCNS1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily S member 1 (Voltage-gated potassium channel subunit Kv9.1) (Delayed-rectifier K(+) channel alpha subunit 1). | |||||
|
KCNC4_HUMAN
|
||||||
| NC score | 0.114002 (rank : 54) | θ value | 0.0252991 (rank : 32) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q03721, Q3MIM4, Q5TBI6 | Gene names | KCNC4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 4 (Voltage-gated potassium channel subunit Kv3.4) (KSHIIIC). | |||||
|
KCNS1_MOUSE
|
||||||
| NC score | 0.106632 (rank : 55) | θ value | 5.27518 (rank : 111) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | O35173 | Gene names | Kcns1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily S member 1 (Voltage-gated potassium channel subunit Kv9.1) (Delayed-rectifier K(+) channel alpha subunit 1). | |||||
|
KCNC4_MOUSE
|
||||||
| NC score | 0.106469 (rank : 56) | θ value | 0.163984 (rank : 39) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q8R1C0 | Gene names | Kcnc4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium voltage-gated channel subfamily C member 4 (Voltage-gated potassium channel subunit Kv3.4). | |||||
|
KCNC3_HUMAN
|
||||||
| NC score | 0.104609 (rank : 57) | θ value | 6.88961 (rank : 117) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q14003 | Gene names | KCNC3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 3 (Voltage-gated potassium channel subunit Kv3.3) (KSHIIID). | |||||
|
KCNC3_MOUSE
|
||||||
| NC score | 0.102305 (rank : 58) | θ value | 8.99809 (rank : 128) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q63959, Q62088 | Gene names | Kcnc3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 3 (Voltage-gated potassium channel subunit Kv3.3) (KSHIIID). | |||||
|
PKDRE_HUMAN
|
||||||
| NC score | 0.097757 (rank : 59) | θ value | θ > 10 (rank : 149) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9NTG1, O95850 | Gene names | PKDREJ | |||
|
Domain Architecture |

|
|||||
| Description | Polycystic kidney disease and receptor for egg jelly-related protein precursor (PKD and REJ homolog). | |||||
|
TPTE_HUMAN
|
||||||
| NC score | 0.086284 (rank : 60) | θ value | θ > 10 (rank : 152) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P56180, Q6XPS4, Q6XPS5, Q71JA8, Q8NCS8 | Gene names | TPTE | |||
|
Domain Architecture |

|
|||||
| Description | Putative tyrosine-protein phosphatase TPTE (EC 3.1.3.48) (Transmembrane phosphatase with tensin homology) (Protein BJ-HCC-5). | |||||
|
TRPV3_HUMAN
|
||||||
| NC score | 0.086090 (rank : 61) | θ value | 0.125558 (rank : 38) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8NET8, Q8NDW7, Q8NET9, Q8NFH2 | Gene names | TRPV3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 3 (TrpV3) (Vanilloid receptor-like 3) (VRL-3). | |||||
|
KCNA1_MOUSE
|
||||||
| NC score | 0.080894 (rank : 62) | θ value | 0.0431538 (rank : 36) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P16388 | Gene names | Kcna1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 1 (Voltage-gated potassium channel subunit Kv1.1) (MKI) (MBK1). | |||||
|
KCNA1_HUMAN
|
||||||
| NC score | 0.079537 (rank : 63) | θ value | 0.0431538 (rank : 35) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q09470 | Gene names | KCNA1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 1 (Voltage-gated potassium channel subunit Kv1.1) (HUKI) (HBK1). | |||||
|
KCNG3_MOUSE
|
||||||
| NC score | 0.078657 (rank : 64) | θ value | θ > 10 (rank : 141) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P59053 | Gene names | Kcng3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily G member 3 (Voltage-gated potassium channel subunit Kv6.3) (Kv10.1). | |||||
|
KCNG3_HUMAN
|
||||||
| NC score | 0.078435 (rank : 65) | θ value | θ > 10 (rank : 140) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q8TAE7 | Gene names | KCNG3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily G member 3 (Voltage-gated potassium channel subunit Kv6.3) (Kv10.1). | |||||
|
KCNA2_MOUSE
|
||||||
| NC score | 0.078011 (rank : 66) | θ value | 1.38821 (rank : 61) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P63141, P15386, Q02010, Q8C8W4 | Gene names | Kcna2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium voltage-gated channel subfamily A member 2 (Voltage-gated potassium channel subunit Kv1.2) (MK2). | |||||
|
KCNA2_HUMAN
|
||||||
| NC score | 0.077179 (rank : 67) | θ value | 1.38821 (rank : 60) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P16389 | Gene names | KCNA2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 2 (Voltage-gated potassium channel subunit Kv1.2) (HBK5) (NGK1) (HUKIV). | |||||
|
KCNQ4_HUMAN
|
||||||
| NC score | 0.075006 (rank : 68) | θ value | 0.813845 (rank : 52) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P56696, O96025 | Gene names | KCNQ4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 4 (Voltage-gated potassium channel subunit Kv7.4) (Potassium channel subunit alpha KvLQT4) (KQT-like 4). | |||||
|
KCNA6_MOUSE
|
||||||
| NC score | 0.074927 (rank : 69) | θ value | 1.81305 (rank : 73) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q61923 | Gene names | Kcna6 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 6 (Voltage-gated potassium channel subunit Kv1.6) (MK1.6). | |||||
|
KCNA6_HUMAN
|
||||||
| NC score | 0.071763 (rank : 70) | θ value | 1.81305 (rank : 72) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P17658 | Gene names | KCNA6 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 6 (Voltage-gated potassium channel subunit Kv1.6) (HBK2). | |||||
|
KCNA5_MOUSE
|
||||||
| NC score | 0.071493 (rank : 71) | θ value | 0.365318 (rank : 45) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q61762 | Gene names | Kcna5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 5 (Voltage-gated potassium channel subunit Kv1.5) (KV1-5). | |||||
|
PKDRE_MOUSE
|
||||||
| NC score | 0.070325 (rank : 72) | θ value | θ > 10 (rank : 150) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9Z0T6 | Gene names | Pkdrej | |||
|
Domain Architecture |

|
|||||
| Description | Polycystic kidney disease and receptor for egg jelly-related protein precursor (PKD and REJ homolog). | |||||
|
KCNA3_HUMAN
|
||||||
| NC score | 0.070266 (rank : 73) | θ value | 0.47712 (rank : 46) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P22001 | Gene names | KCNA3, HGK5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 3 (Voltage-gated potassium channel subunit Kv1.3) (HPCN3) (HGK5) (HuKIII) (HLK3). | |||||
|
KCNA3_MOUSE
|
||||||
| NC score | 0.069688 (rank : 74) | θ value | 0.21417 (rank : 43) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P16390 | Gene names | Kcna3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 3 (Voltage-gated potassium channel subunit Kv1.3) (MK3). | |||||
|
KCNA5_HUMAN
|
||||||
| NC score | 0.065813 (rank : 75) | θ value | 1.81305 (rank : 71) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P22460 | Gene names | KCNA5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 5 (Voltage-gated potassium channel subunit Kv1.5) (HK2) (HPCN1). | |||||
|
KCNQ1_MOUSE
|
||||||
| NC score | 0.064037 (rank : 76) | θ value | θ > 10 (rank : 143) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P97414, O88702, Q7TNZ1, Q7TPL7, Q80VR7 | Gene names | Kcnq1, Kcna9, Kvlqt1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 1 (Voltage-gated potassium channel subunit Kv7.1) (IKs producing slow voltage-gated potassium channel subunit alpha KvLQT1) (KQT-like 1). | |||||
|
TRPC1_MOUSE
|
||||||
| NC score | 0.057228 (rank : 77) | θ value | θ > 10 (rank : 154) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q61056, O35722 | Gene names | Trpc1, Trp1, Trrp1 | |||
|
Domain Architecture |

|
|||||
| Description | Short transient receptor potential channel 1 (TrpC1) (Transient receptor protein 1) (Mtrp1) (Trp-related protein 1). | |||||
|
TRPC1_HUMAN
|
||||||
| NC score | 0.057140 (rank : 78) | θ value | θ > 10 (rank : 153) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P48995 | Gene names | TRPC1, TRP1 | |||
|
Domain Architecture |

|
|||||
| Description | Short transient receptor potential channel 1 (TrpC1) (TRP-1 protein). | |||||
|
TRPV3_MOUSE
|
||||||
| NC score | 0.056818 (rank : 79) | θ value | θ > 10 (rank : 155) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8K424, Q5SV08 | Gene names | Trpv3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 3 (TrpV3). | |||||
|
KCNA4_MOUSE
|
||||||
| NC score | 0.056192 (rank : 80) | θ value | 1.06291 (rank : 55) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q61423 | Gene names | Kcna4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 4 (Voltage-gated potassium channel subunit Kv1.4). | |||||
|
KCNA4_HUMAN
|
||||||
| NC score | 0.055659 (rank : 81) | θ value | 1.81305 (rank : 70) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P22459 | Gene names | KCNA4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 4 (Voltage-gated potassium channel subunit Kv1.4) (HK1) (HPCN2) (HBK4) (HUKII). | |||||
|
KCNQ1_HUMAN
|
||||||
| NC score | 0.055557 (rank : 82) | θ value | θ > 10 (rank : 142) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P51787, O00347, O60607, O94787, Q7Z6G9, Q92960, Q9UMN8, Q9UMN9 | Gene names | KCNQ1, KCNA8, KCNA9, KVLQT1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 1 (Voltage-gated potassium channel subunit Kv7.1) (IKs producing slow voltage-gated potassium channel subunit alpha KvLQT1) (KQT-like 1). | |||||
|
KCNG1_HUMAN
|
||||||
| NC score | 0.053167 (rank : 83) | θ value | θ > 10 (rank : 139) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9UIX4, O43528, Q9BRC1 | Gene names | KCNG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily G member 1 (Voltage-gated potassium channel subunit Kv6.1) (kH2). | |||||
|
TRPV4_MOUSE
|
||||||
| NC score | 0.052805 (rank : 84) | θ value | θ > 10 (rank : 157) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9EPK8, Q91XR5, Q9EQZ4, Q9ERZ7, Q9ES76 | Gene names | Trpv4, Trp12, Vrl2, Vroac | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 4 (TrpV4) (osm-9-like TRP channel 4) (OTRPC4) (Vanilloid receptor-like channel 2) (Vanilloid receptor-like protein 2) (Vanilloid receptor- related osmotically-activated channel) (VR-OAC) (Transient receptor potential protein 12) (TRP12). | |||||
|
CABP5_MOUSE
|
||||||
| NC score | 0.052512 (rank : 85) | θ value | θ > 10 (rank : 136) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9JLK3 | Gene names | Cabp5 | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-binding protein 5 (CaBP5). | |||||
|
KCNQ5_HUMAN
|
||||||
| NC score | 0.051970 (rank : 86) | θ value | θ > 10 (rank : 144) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9NR82, Q9NRN0, Q9NYA6 | Gene names | KCNQ5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 5 (Voltage-gated potassium channel subunit Kv7.5) (Potassium channel subunit alpha KvLQT5) (KQT-like 5). | |||||
|
TRPV4_HUMAN
|
||||||
| NC score | 0.051454 (rank : 87) | θ value | θ > 10 (rank : 156) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9HBA0, Q86YZ6, Q8NDY7, Q8NG64, Q96Q92, Q96RS7, Q9HBC0 | Gene names | TRPV4, VRL2, VROAC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 4 (TrpV4) (osm-9-like TRP channel 4) (OTRPC4) (Vanilloid receptor-like channel 2) (Vanilloid receptor-like protein 2) (VRL-2) (Vanilloid receptor-related osmotically-activated channel) (VR-OAC) (Transient receptor potential protein 12) (TRP12). | |||||
|
TRPM8_MOUSE
|
||||||
| NC score | 0.041875 (rank : 88) | θ value | 1.81305 (rank : 77) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8R4D5 | Gene names | Trpm8, Trpp8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily M member 8. | |||||
|
MEPE_HUMAN
|
||||||
| NC score | 0.039495 (rank : 89) | θ value | 0.21417 (rank : 44) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9NQ76 | Gene names | MEPE | |||
|
Domain Architecture |

|
|||||
| Description | Matrix extracellular phosphoglycoprotein precursor (Osteoblast/osteocyte factor 45) (OF45). | |||||
|
TRPM8_HUMAN
|
||||||
| NC score | 0.035930 (rank : 90) | θ value | 6.88961 (rank : 122) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q7Z2W7, Q6QNH9, Q8TAC3, Q8TDX8, Q9BVK1 | Gene names | TRPM8, LTRPC6, TRPP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily M member 8 (Transient receptor potential-p8) (Trp-p8) (Long transient receptor potential channel 6) (LTrpC6). | |||||
|
KCMA1_MOUSE
|
||||||
| NC score | 0.031766 (rank : 91) | θ value | 5.27518 (rank : 110) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q08460, Q64703, Q8VHF1, Q9R196 | Gene names | Kcnma1, Kcnma | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-activated potassium channel subunit alpha 1 (Calcium-activated potassium channel, subfamily M subunit alpha 1) (Maxi K channel) (MaxiK) (BK channel) (K(VCA)alpha) (BKCA alpha) (KCa1.1) (Slowpoke homolog) (Slo homolog) (Slo-alpha) (Slo1) (mSlo) (mSlo1). | |||||
|
KCMA1_HUMAN
|
||||||
| NC score | 0.031710 (rank : 92) | θ value | 5.27518 (rank : 109) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q12791, Q12886, Q12917, Q12921, Q12960, Q13150, Q96LG8, Q9UBB0, Q9UQK6 | Gene names | KCNMA1, KCNMA | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-activated potassium channel subunit alpha 1 (Calcium-activated potassium channel, subfamily M subunit alpha 1) (Maxi K channel) (MaxiK) (BK channel) (K(VCA)alpha) (BKCA alpha) (KCa1.1) (Slowpoke homolog) (Slo homolog) (Slo-alpha) (Slo1) (hSlo). | |||||
|
CALB2_MOUSE
|
||||||
| NC score | 0.027071 (rank : 93) | θ value | 2.36792 (rank : 79) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q08331, Q60964, Q9JM81 | Gene names | Calb2 | |||
|
Domain Architecture |

|
|||||
| Description | Calretinin (CR). | |||||
|
CALB2_HUMAN
|
||||||
| NC score | 0.026840 (rank : 94) | θ value | 2.36792 (rank : 78) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P22676, Q53HD2, Q96BK4 | Gene names | CALB2, CAB29 | |||
|
Domain Architecture |

|
|||||
| Description | Calretinin (CR) (29 kDa calbindin). | |||||
|
SEGN_HUMAN
|
||||||
| NC score | 0.026503 (rank : 95) | θ value | 0.47712 (rank : 48) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O76038, Q5VV44, Q96QV7, Q9UJF6 | Gene names | SCGN, SECRET | |||
|
Domain Architecture |

|
|||||
| Description | Secretagogin. | |||||
|
TRPV1_MOUSE
|
||||||
| NC score | 0.021245 (rank : 96) | θ value | 4.03905 (rank : 103) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q704Y3, Q5SSE1, Q5SSE2, Q5SSE4, Q5WPV5, Q68SW0 | Gene names | Trpv1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily V member 1 (TrpV1) (osm-9-like TRP channel 1) (OTRPC1) (Vanilloid receptor 1). | |||||
|
TRPM2_MOUSE
|
||||||
| NC score | 0.019774 (rank : 97) | θ value | 6.88961 (rank : 121) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q91YD4 | Gene names | Trpm2, Ltrpc2, Trpc7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily M member 2 (EC 3.6.1.13) (Long transient receptor potential channel 2) (LTrpC2) (LTrpC-2) (Transient receptor potential channel 7) (TrpC7). | |||||
|
ZN318_HUMAN
|
||||||
| NC score | 0.019689 (rank : 98) | θ value | 0.813845 (rank : 54) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 497 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q5VUA4, O94796, Q4G0E4, Q8NEM6, Q9UNU8, Q9Y2W9 | Gene names | ZNF318 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 318 (Endocrine regulatory protein). | |||||
|
KCNH2_MOUSE
|
||||||
| NC score | 0.019201 (rank : 99) | θ value | 0.0252991 (rank : 34) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O35219, O35220, O35221, O35989 | Gene names | Kcnh2, Erg, Merg1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 2 (Voltage-gated potassium channel subunit Kv11.1) (Ether-a-go-go-related gene potassium channel 1) (ERG1) (MERG) (Merg1) (Ether-a-go-go-related protein 1) (Eag-related protein 1). | |||||
|
NPM2_HUMAN
|
||||||
| NC score | 0.018418 (rank : 100) | θ value | 1.06291 (rank : 57) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q86SE8 | Gene names | NPM2 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoplasmin-2. | |||||
|
KCNH2_HUMAN
|
||||||
| NC score | 0.018011 (rank : 101) | θ value | 0.0252991 (rank : 33) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q12809, O75418, O75680, Q9BT72, Q9BUT7, Q9H3P0 | Gene names | KCNH2, ERG, ERG1, HERG, HERG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 2 (Voltage-gated potassium channel subunit Kv11.1) (Ether-a-go-go-related gene potassium channel 1) (H-ERG) (Erg1) (Ether-a-go-go-related protein 1) (Eag-related protein 1) (eag homolog). | |||||
|
TRPM2_HUMAN
|
||||||
| NC score | 0.016592 (rank : 102) | θ value | 1.38821 (rank : 65) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O94759, Q5KTC2, Q6J3P5, Q96KN6, Q96Q93 | Gene names | TRPM2, EREG1, KNP3, LTRPC2, TRPC7 | |||
|
Domain Architecture |

|
|||||
| Description | Transient receptor potential cation channel subfamily M member 2 (EC 3.6.1.13) (Long transient receptor potential channel 2) (LTrpC2) (LTrpC-2) (Transient receptor potential channel 7) (TrpC7) (Estrogen- responsive element-associated gene 1 protein). | |||||
|
SEGN_MOUSE
|
||||||
| NC score | 0.016543 (rank : 103) | θ value | 5.27518 (rank : 114) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q91WD9 | Gene names | Scgn | |||
|
Domain Architecture |

|
|||||
| Description | Secretagogin. | |||||
|
DZIP1_MOUSE
|
||||||
| NC score | 0.013397 (rank : 104) | θ value | 0.813845 (rank : 51) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 530 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8BMD2, Q6ZQ10, Q9CRK5 | Gene names | Dzip1, Kiaa0996 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein DZIP1 (DAZ-interacting protein 1 homolog). | |||||
|
RIF1_MOUSE
|
||||||
| NC score | 0.012981 (rank : 105) | θ value | 0.813845 (rank : 53) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q6PR54, Q6T8D9, Q7TPD8, Q8BTU2, Q8C408, Q9CS77 | Gene names | Rif1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Telomere-associated protein RIF1 (Rap1-interacting factor 1 homolog) (mRif1). | |||||
|
ZC11A_HUMAN
|
||||||
| NC score | 0.012284 (rank : 106) | θ value | 2.36792 (rank : 87) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O75152, Q6AHY4, Q6AHY9, Q6AW79, Q6AWA1, Q6PJK4, Q86XZ7 | Gene names | ZC3H11A, KIAA0663 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 11A. | |||||
|
MKL1_MOUSE
|
||||||
| NC score | 0.012028 (rank : 107) | θ value | 1.38821 (rank : 62) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 667 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8K4J6 | Gene names | Mkl1, Bsac | |||
|
Domain Architecture |

|
|||||
| Description | MKL/myocardin-like protein 1 (Myocardin-related transcription factor A) (MRTF-A) (Megakaryoblastic leukemia 1 protein homolog) (Basic SAP coiled-coil transcription activator). | |||||
|
CT2NL_HUMAN
|
||||||
| NC score | 0.011657 (rank : 108) | θ value | 4.03905 (rank : 94) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1059 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9P2B4, Q96B40 | Gene names | CTTNBP2NL, KIAA1433 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CTTNBP2 N-terminal-like protein. | |||||
|
I17RA_MOUSE
|
||||||
| NC score | 0.010994 (rank : 109) | θ value | 1.81305 (rank : 69) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q60943 | Gene names | IL17ra, Il17r | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interleukin-17 receptor A precursor (IL-17 receptor). | |||||
|
MAP1B_MOUSE
|
||||||
| NC score | 0.010640 (rank : 110) | θ value | 2.36792 (rank : 82) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 977 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P14873 | Gene names | Map1b, Mtap1b, Mtap5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) (MAP1.2) (MAP1(X)) [Contains: MAP1 light chain LC1]. | |||||
|
RXINP_MOUSE
|
||||||
| NC score | 0.010458 (rank : 111) | θ value | 3.0926 (rank : 91) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q5U5Q9, Q60811 | Gene names | Rxrip110, Rip110 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoid X receptor-interacting protein 110. | |||||
|
PIGW_HUMAN
|
||||||
| NC score | 0.009703 (rank : 112) | θ value | 4.03905 (rank : 99) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7Z7B1, Q8N9G3 | Gene names | PIGW | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol-glycan biosynthesis class W protein (EC 2.3.-.-) (PIG-W). | |||||
|
TAXB1_HUMAN
|
||||||
| NC score | 0.009494 (rank : 113) | θ value | 1.06291 (rank : 58) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 829 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q86VP1, O60398, O95770, Q13311, Q9BQG5, Q9UI88 | Gene names | TAX1BP1, T6BP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tax1-binding protein 1 (TRAF6-binding protein). | |||||
|
ITPR1_HUMAN
|
||||||
| NC score | 0.009390 (rank : 114) | θ value | 5.27518 (rank : 107) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14643, Q14660, Q99897 | Gene names | ITPR1, INSP3R1 | |||
|
Domain Architecture |

|
|||||
| Description | Inositol 1,4,5-trisphosphate receptor type 1 (Type 1 inositol 1,4,5- trisphosphate receptor) (Type 1 InsP3 receptor) (IP3 receptor isoform 1) (InsP3R1) (IP3R). | |||||
|
ITPR1_MOUSE
|
||||||
| NC score | 0.009375 (rank : 115) | θ value | 5.27518 (rank : 108) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P11881, P20943, Q99LG5 | Gene names | Itpr1, Insp3r, Pcd6, Pcp1 | |||
|
Domain Architecture |

|
|||||
| Description | Inositol 1,4,5-trisphosphate receptor type 1 (Type 1 inositol 1,4,5- trisphosphate receptor) (Type 1 InsP3 receptor) (IP3 receptor isoform 1) (InsP3R1) (Inositol 1,4,5-trisphosphate-binding protein P400) (Purkinje cell protein 1) (Protein PCD-6). | |||||
|
TMF1_HUMAN
|
||||||
| NC score | 0.008353 (rank : 116) | θ value | 1.06291 (rank : 59) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 987 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P82094 | Gene names | TMF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TATA element modulatory factor (TMF). | |||||
|
TPR_HUMAN
|
||||||
| NC score | 0.008205 (rank : 117) | θ value | 2.36792 (rank : 86) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
CKAP4_MOUSE
|
||||||
| NC score | 0.007957 (rank : 118) | θ value | 5.27518 (rank : 106) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 596 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8BMK4, Q8BTK8, Q8R3F2 | Gene names | Ckap4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-associated protein 4 (63 kDa membrane protein) (p63). | |||||
|
CROCC_MOUSE
|
||||||
| NC score | 0.007946 (rank : 119) | θ value | 4.03905 (rank : 93) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1460 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8CJ40, Q7TQL2, Q80U01, Q8R0B9 | Gene names | Crocc, Kiaa0445 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rootletin (Ciliary rootlet coiled-coil protein). | |||||
|
MYH4_HUMAN
|
||||||
| NC score | 0.007869 (rank : 120) | θ value | 5.27518 (rank : 112) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1377 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9Y623 | Gene names | MYH4 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-4 (Myosin heavy chain, skeletal muscle, fetal) (Myosin heavy chain IIb) (MyHC-IIb). | |||||
|
MYH13_HUMAN
|
||||||
| NC score | 0.007861 (rank : 121) | θ value | 1.38821 (rank : 63) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1504 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9UKX3, O95252 | Gene names | MYH13 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-13 (Myosin heavy chain, skeletal muscle, extraocular) (MyHC- eo). | |||||
|
MAP1B_HUMAN
|
||||||
| NC score | 0.007780 (rank : 122) | θ value | 6.88961 (rank : 118) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P46821 | Gene names | MAP1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) [Contains: MAP1 light chain LC1]. | |||||
|
RB6I2_HUMAN
|
||||||
| NC score | 0.007743 (rank : 123) | θ value | 6.88961 (rank : 120) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1584 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8IUD2, Q6NVK2, Q8IUD3, Q8IUD4, Q8IUD5, Q8NAS1, Q9NXN5, Q9UIK7, Q9UPS1 | Gene names | ERC1, ELKS, KIAA1081, RAB6IP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ELKS/RAB6-interacting/CAST family member 1 (RAB6-interacting protein 2) (ERC protein 1). | |||||
|
SMC4_MOUSE
|
||||||
| NC score | 0.007608 (rank : 124) | θ value | 5.27518 (rank : 115) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 830 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8CG47, Q8BTS7, Q8BTY9, Q99K21 | Gene names | Smc4l1, Capc, Smc4 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosomes 4-like 1 protein (Chromosome- associated polypeptide C) (XCAP-C homolog). | |||||
|
CP135_HUMAN
|
||||||
| NC score | 0.007413 (rank : 125) | θ value | 1.81305 (rank : 66) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1290 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q66GS9, O75130, Q9H8H7 | Gene names | CEP135, CEP4, KIAA0635 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 135 kDa (Cep135 protein) (Centrosomal protein 4). | |||||
|
MYH6_HUMAN
|
||||||
| NC score | 0.007047 (rank : 126) | θ value | 2.36792 (rank : 83) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1426 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P13533, Q13943, Q14906, Q14907 | Gene names | MYH6, MYHCA | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
MYH6_MOUSE
|
||||||
| NC score | 0.007043 (rank : 127) | θ value | 2.36792 (rank : 84) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1501 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q02566, Q64258, Q64738 | Gene names | Myh6, Myhca | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
CP135_MOUSE
|
||||||
| NC score | 0.006995 (rank : 128) | θ value | 4.03905 (rank : 92) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1278 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q6P5D4, Q6A030 | Gene names | Cep135, Cep4, Kiaa0635 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 135 kDa (Cep135 protein) (Centrosomal protein 4). | |||||
|
CF152_HUMAN
|
||||||
| NC score | 0.006983 (rank : 129) | θ value | 8.99809 (rank : 124) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q86VQ0, Q9BWX7 | Gene names | C6orf152 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf152. | |||||
|
SMC1A_HUMAN
|
||||||
| NC score | 0.006939 (rank : 130) | θ value | 4.03905 (rank : 101) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1346 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q14683, O14995, Q16351 | Gene names | SMC1L1, DXS423E, KIAA0178, SB1.8, SMC1 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Sb1.8). | |||||
|
MYH7_HUMAN
|
||||||
| NC score | 0.006763 (rank : 131) | θ value | 2.36792 (rank : 85) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P12883, Q14836, Q14837, Q14904, Q16579, Q92679, Q9H1D5, Q9UMM8 | Gene names | MYH7, MYHCB | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle beta isoform (MyHC-beta). | |||||
|
ERC2_MOUSE
|
||||||
| NC score | 0.006729 (rank : 132) | θ value | 4.03905 (rank : 96) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1166 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q6PH08, Q80U20, Q8CCP1 | Gene names | Erc2, Cast1, D14Ertd171e, Kiaa0378 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ERC protein 2 (CAZ-associated structural protein 1) (CAST1). | |||||
|
ERC2_HUMAN
|
||||||
| NC score | 0.006711 (rank : 133) | θ value | 4.03905 (rank : 95) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1156 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O15083, Q86TK4 | Gene names | ERC2, KIAA0378 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ERC protein 2. | |||||
|
SMC1A_MOUSE
|
||||||
| NC score | 0.006657 (rank : 134) | θ value | 4.03905 (rank : 102) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1384 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9CU62, Q3V480, Q9CUX9, Q9D959, Q9WTU1 | Gene names | Smc1l1, Sb1.8, Smc1, Smcb | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Chromosome segregation protein SmcB) (Sb1.8). | |||||
|
TMCC3_HUMAN
|
||||||
| NC score | 0.006352 (rank : 135) | θ value | 8.99809 (rank : 133) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9ULS5, Q8IWB2 | Gene names | TMCC3, KIAA1145 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane and coiled-coil domains protein 3. | |||||
|
NCKX1_HUMAN
|
||||||
| NC score | 0.006104 (rank : 136) | θ value | 3.0926 (rank : 90) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O60721, O43485, O75184 | Gene names | SLC24A1, KIAA0702, NCKX1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium/calcium exchanger 1 (Na(+)/K(+)/Ca(2+)-exchange protein 1) (Retinal rod Na-Ca+K exchanger). | |||||
|
MYH9_HUMAN
|
||||||
| NC score | 0.005688 (rank : 137) | θ value | 5.27518 (rank : 113) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P35579, O60805 | Gene names | MYH9 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
TRPA1_HUMAN
|
||||||
| NC score | 0.005546 (rank : 138) | θ value | 1.81305 (rank : 76) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 339 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O75762 | Gene names | TRPA1, ANKTM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily A member 1 (Ankyrin-like with transmembrane domains protein 1) (Transformation sensitive-protein p120). | |||||
|
MYH10_HUMAN
|
||||||
| NC score | 0.005413 (rank : 139) | θ value | 8.99809 (rank : 130) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P35580, Q16087 | Gene names | MYH10 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
MYH10_MOUSE
|
||||||
| NC score | 0.005279 (rank : 140) | θ value | 8.99809 (rank : 131) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1612 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q61879, Q5SV63 | Gene names | Myh10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
MYH9_MOUSE
|
||||||
| NC score | 0.005206 (rank : 141) | θ value | 8.99809 (rank : 132) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1829 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8VDD5 | Gene names | Myh9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
NKTR_HUMAN
|
||||||
| NC score | 0.004460 (rank : 142) | θ value | 6.88961 (rank : 119) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 658 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P30414 | Gene names | NKTR | |||
|
Domain Architecture |

|
|||||
| Description | NK-tumor recognition protein (Natural-killer cells cyclophilin-related protein) (NK-TR protein). | |||||
|
WWP1_HUMAN
|
||||||
| NC score | 0.004304 (rank : 143) | θ value | 8.99809 (rank : 134) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9H0M0, O00307, Q96BP4 | Gene names | WWP1 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP1 (EC 6.3.2.-) (WW domain- containing protein 1) (Atropin-1-interacting protein 5) (AIP5). | |||||
|
OAT6_MOUSE
|
||||||
| NC score | 0.004112 (rank : 144) | θ value | 4.03905 (rank : 98) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q80UJ1 | Gene names | Slc22a20, Gm963, Oat6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Solute carrier family 22 member 20 (Organic anion transporter 6). | |||||
|
CENPF_HUMAN
|
||||||
| NC score | 0.003758 (rank : 145) | θ value | 8.99809 (rank : 123) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1932 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P49454, Q13171, Q13246 | Gene names | CENPF | |||
|
Domain Architecture |

|
|||||
| Description | Centromere protein F (Kinetochore protein CENP-F) (Mitosin) (AH antigen). | |||||
|
PLEC1_HUMAN
|
||||||
| NC score | 0.003399 (rank : 146) | θ value | 4.03905 (rank : 100) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 1326 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q15149, Q15148, Q16640 | Gene names | PLEC1 | |||
|
Domain Architecture |

|
|||||
| Description | Plectin-1 (PLTN) (PCN) (Hemidesmosomal protein 1) (HD1). | |||||
|
DDX23_HUMAN
|
||||||
| NC score | 0.003061 (rank : 147) | θ value | 1.81305 (rank : 67) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9BUQ8, O43188 | Gene names | DDX23 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX23 (EC 3.6.1.-) (DEAD box protein 23) (100 kDa U5 snRNP-specific protein) (U5-100kD) (PRP28 homolog). | |||||
|
CALM_HUMAN
|
||||||
| NC score | 0.002530 (rank : 148) | θ value | 5.27518 (rank : 104) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P62158, P02593, P70667, P99014, Q13942, Q53S29, Q61379, Q61380, Q96HK3 | Gene names | CALM1, CALM, CAM, CAM1 | |||
|
Domain Architecture |

|
|||||
| Description | Calmodulin (CaM). | |||||
|
CALM_MOUSE
|
||||||
| NC score | 0.002530 (rank : 149) | θ value | 5.27518 (rank : 105) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P62204, P02593, P70667, P99014, Q61379, Q61380 | Gene names | Calm1, Calm, Cam, Cam1 | |||
|
Domain Architecture |

|
|||||
| Description | Calmodulin (CaM). | |||||
|
KIF3A_HUMAN
|
||||||
| NC score | 0.002446 (rank : 150) | θ value | 3.0926 (rank : 89) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9Y496, Q86XE9, Q9Y6V4 | Gene names | KIF3A, KIF3 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF3A (Microtubule plus end-directed kinesin motor 3A). | |||||
|
DSPP_HUMAN
|
||||||
| NC score | 0.002415 (rank : 151) | θ value | 6.88961 (rank : 116) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
GPR64_HUMAN
|
||||||
| NC score | 0.002406 (rank : 152) | θ value | 4.03905 (rank : 97) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8IZP9, O00406, Q8IWT2, Q8IZE4, Q8IZE5, Q8IZE6, Q8IZE7, Q8IZP3, Q8IZP4 | Gene names | GPR64, HE6, TM7LN2 | |||
|
Domain Architecture |

|
|||||
| Description | G-protein coupled receptor 64 precursor (Epididymis-specific protein 6) (He6 receptor). | |||||
|
MOR2A_MOUSE
|
||||||
| NC score | 0.002304 (rank : 153) | θ value | 8.99809 (rank : 129) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q69ZX6, Q5QNQ7, Q6P547 | Gene names | Morc2a, Kiaa0852, Zcwcc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORC family CW-type zinc finger protein 2A (Zinc finger CW-type coiled-coil domain protein 1). | |||||
|
HUWE1_HUMAN
|
||||||
| NC score | 0.002016 (rank : 154) | θ value | 8.99809 (rank : 125) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q7Z6Z7, O15029, Q4G2Z2, Q5H961, Q6P4D0, Q8NG67, Q9BUI0, Q9HCJ4, Q9NSL6, Q9P0A9 | Gene names | HUWE1, KIAA0312, KIAA1578, UREB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (Mcl-1 ubiquitin ligase E3) (Mule) (ARF-binding protein 1) (ARF-BP1). | |||||
|
WWP1_MOUSE
|
||||||
| NC score | 0.001973 (rank : 155) | θ value | 8.99809 (rank : 135) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BZZ3, Q8BIV9, Q8VDP8 | Gene names | Wwp1 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP1 (EC 6.3.2.-) (WW domain- containing protein 1). | |||||
|
DRD1_MOUSE
|
||||||
| NC score | -0.000808 (rank : 156) | θ value | 1.81305 (rank : 68) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 733 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61616, Q8C8P8 | Gene names | Drd1, Drd1a, Gpcr15 | |||
|
Domain Architecture |

|
|||||
| Description | D(1A) dopamine receptor. | |||||
|
DRD1_HUMAN
|
||||||
| NC score | -0.001323 (rank : 157) | θ value | 3.0926 (rank : 88) | |||
| Query Neighborhood Hits | 135 | Target Neighborhood Hits | 750 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P21728, Q4QRJ0 | Gene names | DRD1 | |||
|
Domain Architecture |

|
|||||
| Description | D(1A) dopamine receptor. | |||||