Please be patient as the page loads
|
WBP4_HUMAN
|
||||||
| SwissProt Accessions | O75554 | Gene names | WBP4, FBP21, FNBP21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 4 (WBP-4) (Formin-binding protein 21). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
WBP4_HUMAN
|
||||||
| θ value | 3.47952e-169 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 86 | |
| SwissProt Accessions | O75554 | Gene names | WBP4, FBP21, FNBP21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 4 (WBP-4) (Formin-binding protein 21). | |||||
|
WBP4_MOUSE
|
||||||
| θ value | 2.52267e-127 (rank : 2) | NC score | 0.942255 (rank : 2) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q61048, Q8K1Z9, Q9CS45 | Gene names | Wbp4, Fbp21, Fnbp21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 4 (WBP-4) (Formin-binding protein 21). | |||||
|
PRP40_HUMAN
|
||||||
| θ value | 7.34386e-10 (rank : 3) | NC score | 0.405093 (rank : 4) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 442 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O75400, O43856, O75404, Q8TBQ1, Q9H782, Q9NWU9, Q9P0Q2, Q9Y5A8 | Gene names | PRPF40A, FLAF1, FNBP3, HYPA | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Huntingtin yeast partner A) (Huntingtin-interacting protein HYPA/FBP11) (Fas ligand-associated factor 1) (NY-REN-6 antigen). | |||||
|
PRP40_MOUSE
|
||||||
| θ value | 7.34386e-10 (rank : 4) | NC score | 0.406196 (rank : 3) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9R1C7, Q61049, Q8BQ76, Q8BRW4, Q8C2U1 | Gene names | Prpf40a, Fbp11, Fnbp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Formin-binding protein 11) (FBP-11). | |||||
|
TCRG1_HUMAN
|
||||||
| θ value | 0.00228821 (rank : 5) | NC score | 0.184246 (rank : 7) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 929 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O14776 | Gene names | TCERG1, CA150, TAF2S | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150). | |||||
|
TCRG1_MOUSE
|
||||||
| θ value | 0.00228821 (rank : 6) | NC score | 0.181522 (rank : 8) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8CGF7, Q61051, Q8C490, Q8CHT8, Q9R0R5 | Gene names | Tcerg1, Taf2s | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150) (p144) (Formin- binding protein 28) (FBP 28). | |||||
|
IQGA1_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 7) | NC score | 0.156523 (rank : 9) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P46940 | Gene names | IQGAP1, KIAA0051 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP1 (p195). | |||||
|
RU1C_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 8) | NC score | 0.230562 (rank : 5) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P09234 | Gene names | SNRPC | |||
|
Domain Architecture |

|
|||||
| Description | U1 small nuclear ribonucleoprotein C (U1 snRNP protein C) (U1C protein) (U1-C). | |||||
|
RU1C_MOUSE
|
||||||
| θ value | 0.0148317 (rank : 9) | NC score | 0.228862 (rank : 6) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q62241 | Gene names | Snrp1c, Snrpc | |||
|
Domain Architecture |

|
|||||
| Description | U1 small nuclear ribonucleoprotein C (U1 snRNP protein C) (U1C protein) (U1-C). | |||||
|
IQGA1_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 10) | NC score | 0.151935 (rank : 10) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9JKF1 | Gene names | Iqgap1 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP1. | |||||
|
GAS7_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 11) | NC score | 0.147562 (rank : 12) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q60780, Q9QY25, Q9QY26, Q9QY34 | Gene names | Gas7 | |||
|
Domain Architecture |

|
|||||
| Description | Growth-arrest-specific protein 7 (GAS-7). | |||||
|
YTDC1_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 12) | NC score | 0.095236 (rank : 18) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q96MU7, Q7Z622, Q8TF35 | Gene names | YTHDC1, KIAA1966, YT521 | |||
|
Domain Architecture |

|
|||||
| Description | YTH domain-containing protein 1 (Putative splicing factor YT521). | |||||
|
K1688_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 13) | NC score | 0.098450 (rank : 17) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9C0H5 | Gene names | KIAA1688 | |||
|
Domain Architecture |

|
|||||
| Description | Protein KIAA1688. | |||||
|
NKTR_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 14) | NC score | 0.064874 (rank : 28) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 658 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P30414 | Gene names | NKTR | |||
|
Domain Architecture |

|
|||||
| Description | NK-tumor recognition protein (Natural-killer cells cyclophilin-related protein) (NK-TR protein). | |||||
|
GAS7_HUMAN
|
||||||
| θ value | 0.21417 (rank : 15) | NC score | 0.138185 (rank : 14) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O60861, O43144 | Gene names | GAS7, KIAA0394 | |||
|
Domain Architecture |

|
|||||
| Description | Growth-arrest-specific protein 7 (GAS-7). | |||||
|
GP179_HUMAN
|
||||||
| θ value | 0.21417 (rank : 16) | NC score | 0.056607 (rank : 38) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 372 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q6PRD1 | Gene names | GPR179, GPR158L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 179 precursor (Probable G-protein coupled receptor 158-like 1). | |||||
|
IQGA2_HUMAN
|
||||||
| θ value | 0.21417 (rank : 17) | NC score | 0.144408 (rank : 13) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q13576 | Gene names | IQGAP2 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP2. | |||||
|
OXR1_MOUSE
|
||||||
| θ value | 0.21417 (rank : 18) | NC score | 0.072979 (rank : 26) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q4KMM3, Q5FWW1, Q99L06, Q99MK1, Q99MP4 | Gene names | Oxr1, C7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oxidation resistance protein 1 (Protein C7). | |||||
|
SEC63_MOUSE
|
||||||
| θ value | 0.365318 (rank : 19) | NC score | 0.088375 (rank : 19) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 601 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8VHE0, Q8VEB9 | Gene names | Sec63, Sec63l | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
HSP7C_MOUSE
|
||||||
| θ value | 0.47712 (rank : 20) | NC score | 0.030029 (rank : 67) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P63017, P08109, P12225, Q62373, Q62374, Q62375 | Gene names | Hspa8, Hsc70, Hsc73 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heat shock cognate 71 kDa protein (Heat shock 70 kDa protein 8). | |||||
|
IQGA3_HUMAN
|
||||||
| θ value | 0.47712 (rank : 21) | NC score | 0.150241 (rank : 11) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q86VI3 | Gene names | IQGAP3 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP3. | |||||
|
RPTN_MOUSE
|
||||||
| θ value | 0.62314 (rank : 22) | NC score | 0.045930 (rank : 56) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P97347 | Gene names | Rptn | |||
|
Domain Architecture |

|
|||||
| Description | Repetin. | |||||
|
SEC63_HUMAN
|
||||||
| θ value | 0.62314 (rank : 23) | NC score | 0.082591 (rank : 21) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UGP8, O95380, Q86VS9, Q8IWL0, Q9NTE0 | Gene names | SEC63, SEC63L | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
MYRIP_HUMAN
|
||||||
| θ value | 0.813845 (rank : 24) | NC score | 0.058460 (rank : 36) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8NFW9, Q8IUF5, Q9Y3V4 | Gene names | MYRIP, SLAC2C | |||
|
Domain Architecture |
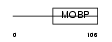
|
|||||
| Description | Rab effector MyRIP (Myosin-VIIa- and Rab-interacting protein) (Exophilin-8) (Slp homolog lacking C2 domains c). | |||||
|
PSIP1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 25) | NC score | 0.062825 (rank : 30) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q99JF8, Q3TEJ7, Q3TTD7, Q3UTA1, Q80WQ7, Q99JF7, Q99LR4, Q9CT03 | Gene names | Psip1, Ledgf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PC4 and SFRS1-interacting protein (Lens epithelium-derived growth factor) (mLEDGF). | |||||
|
TCF8_MOUSE
|
||||||
| θ value | 0.813845 (rank : 26) | NC score | 0.011618 (rank : 89) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1067 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q64318, Q62519 | Gene names | Tcf8, Zfhx1a, Zfx1a, Zfx1ha | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 8 (Zinc finger homeobox protein 1a) (MEB1) (Delta EF1). | |||||
|
IWS1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 27) | NC score | 0.077766 (rank : 25) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
RHG12_MOUSE
|
||||||
| θ value | 1.06291 (rank : 28) | NC score | 0.078215 (rank : 24) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8C0D4, Q8BVP8 | Gene names | Arhgap12 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 12. | |||||
|
SIA7A_HUMAN
|
||||||
| θ value | 1.38821 (rank : 29) | NC score | 0.029984 (rank : 68) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9NSC7, Q6UW90, Q9NSC6 | Gene names | ST6GALNAC1, SIAT7A | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-N-acetylgalactosaminide alpha-2,6-sialyltransferase 1 (EC 2.4.99.3) (GalNAc alpha-2,6-sialyltransferase I) (ST6GalNAc I) (Sialyltransferase 7A). | |||||
|
CEP63_HUMAN
|
||||||
| θ value | 1.81305 (rank : 30) | NC score | 0.021259 (rank : 79) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1005 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q96MT8, Q96CR0, Q9H8F5, Q9H8N0 | Gene names | CEP63 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 63 kDa (Cep63 protein). | |||||
|
HORN_MOUSE
|
||||||
| θ value | 1.81305 (rank : 31) | NC score | 0.042910 (rank : 57) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8VHD8 | Gene names | Hrnr | |||
|
Domain Architecture |

|
|||||
| Description | Hornerin. | |||||
|
NFM_HUMAN
|
||||||
| θ value | 1.81305 (rank : 32) | NC score | 0.029209 (rank : 70) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1328 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P07197 | Gene names | NEF3, NEFM, NFM | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet M protein (160 kDa neurofilament protein) (Neurofilament medium polypeptide) (NF-M) (Neurofilament 3). | |||||
|
SDPR_HUMAN
|
||||||
| θ value | 1.81305 (rank : 33) | NC score | 0.035212 (rank : 63) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O95810 | Gene names | SDPR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serum deprivation-response protein (Phosphatidylserine-binding protein) (PS-p68). | |||||
|
ITCH_HUMAN
|
||||||
| θ value | 2.36792 (rank : 34) | NC score | 0.054943 (rank : 43) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q96J02, O43584, Q5TEL0, Q96F66, Q9BY75, Q9H451, Q9H4U5 | Gene names | ITCH | |||
|
Domain Architecture |

|
|||||
| Description | Itchy homolog E3 ubiquitin protein ligase (EC 6.3.2.-) (Itch) (Atrophin-1-interacting protein 4) (AIP4) (NFE2-associated polypeptide 1) (NAPP1). | |||||
|
ITCH_MOUSE
|
||||||
| θ value | 2.36792 (rank : 35) | NC score | 0.055423 (rank : 41) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8C863, O54971 | Gene names | Itch | |||
|
Domain Architecture |

|
|||||
| Description | Itchy E3 ubiquitin protein ligase (EC 6.3.2.-). | |||||
|
IWS1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 36) | NC score | 0.082115 (rank : 22) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
MYRIP_MOUSE
|
||||||
| θ value | 2.36792 (rank : 37) | NC score | 0.049152 (rank : 53) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8K3I4, Q8CFC0, Q8K4H5 | Gene names | Myrip, Slac2c | |||
|
Domain Architecture |
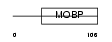
|
|||||
| Description | Rab effector MyRIP (Myosin-VIIa- and Rab-interacting protein) (Exophilin-8) (Slp homolog lacking C2 domains c). | |||||
|
PHF10_MOUSE
|
||||||
| θ value | 2.36792 (rank : 38) | NC score | 0.021491 (rank : 78) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9D8M7, Q99LV5 | Gene names | Phf10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 10. | |||||
|
RHG18_MOUSE
|
||||||
| θ value | 2.36792 (rank : 39) | NC score | 0.035709 (rank : 61) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8K0Q5, Q8BP03, Q8R196 | Gene names | Arhgap18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho-GTPase-activating protein 18. | |||||
|
IRX3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 40) | NC score | 0.023783 (rank : 74) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P78415, Q7Z4A4, Q7Z4A5, Q8IVC6 | Gene names | IRX3, IRXB1 | |||
|
Domain Architecture |

|
|||||
| Description | Iroquois-class homeodomain protein IRX-3 (Iroquois homeobox protein 3) (Homeodomain protein IRXB1). | |||||
|
MINT_MOUSE
|
||||||
| θ value | 3.0926 (rank : 41) | NC score | 0.026773 (rank : 71) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
NPBL_MOUSE
|
||||||
| θ value | 3.0926 (rank : 42) | NC score | 0.055927 (rank : 39) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
PIN1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 43) | NC score | 0.100121 (rank : 16) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q13526 | Gene names | PIN1 | |||
|
Domain Architecture |
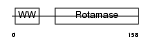
|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase NIMA-interacting 1 (EC 5.2.1.8) (Rotamase Pin1) (PPIase Pin1). | |||||
|
PIN1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 44) | NC score | 0.114665 (rank : 15) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9QUR7, Q543B3 | Gene names | Pin1 | |||
|
Domain Architecture |
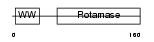
|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase NIMA-interacting 1 (EC 5.2.1.8) (Rotamase Pin1) (PPIase Pin1). | |||||
|
RHG12_HUMAN
|
||||||
| θ value | 3.0926 (rank : 45) | NC score | 0.068825 (rank : 27) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8IWW6, Q86UB3, Q8IWW7, Q8N3L1, Q9NT76 | Gene names | ARHGAP12 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 12. | |||||
|
ANR11_HUMAN
|
||||||
| θ value | 4.03905 (rank : 46) | NC score | 0.031096 (rank : 66) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1289 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q6UB99, Q6NTG1, Q6QMF8 | Gene names | ANKRD11, ANCO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 11 (Ankyrin repeat-containing cofactor 1). | |||||
|
BARH2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 47) | NC score | 0.010028 (rank : 94) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NY43, Q7Z4N7 | Gene names | BARHL2 | |||
|
Domain Architecture |

|
|||||
| Description | BarH-like 2 homeobox protein. | |||||
|
DGC14_MOUSE
|
||||||
| θ value | 4.03905 (rank : 48) | NC score | 0.055874 (rank : 40) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O70279, Q91YX1 | Gene names | Dgcr14, Dgsi, Es2, Es2el | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DGCR14 protein (DiGeorge syndrome critical region 14 homolog) (ES2 protein) (Expressed sequence 2 embryonic lethal). | |||||
|
DMP1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 49) | NC score | 0.058531 (rank : 35) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O55188 | Gene names | Dmp1, Dmp | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1) (AG1). | |||||
|
HSP7C_HUMAN
|
||||||
| θ value | 4.03905 (rank : 50) | NC score | 0.024882 (rank : 73) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P11142, Q9H3R6 | Gene names | HSPA8, HSC70, HSP73, HSPA10 | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock cognate 71 kDa protein (Heat shock 70 kDa protein 8). | |||||
|
IRX3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 51) | NC score | 0.022104 (rank : 76) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P81067 | Gene names | Irx3, Irxb1 | |||
|
Domain Architecture |

|
|||||
| Description | Iroquois-class homeodomain protein IRX-3 (Iroquois homeobox protein 3) (Homeodomain protein IRXB1). | |||||
|
LRCH2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 52) | NC score | 0.010336 (rank : 93) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q5VUJ6, Q9HA88, Q9P233 | Gene names | LRCH2, KIAA1495 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 2. | |||||
|
MYT1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 53) | NC score | 0.038372 (rank : 58) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 475 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q01538, O94922, Q9UPV2 | Gene names | MYT1, KIAA0835, KIAA1050, MTF1, MYTI, PLPB1 | |||
|
Domain Architecture |

|
|||||
| Description | Myelin transcription factor 1 (MyT1) (MyTI) (Proteolipid protein- binding protein) (PLPB1). | |||||
|
NED4L_HUMAN
|
||||||
| θ value | 4.03905 (rank : 54) | NC score | 0.046219 (rank : 55) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q96PU5, O43165, Q7Z5F1, Q7Z5F2, Q7Z5N3, Q8N5A7, Q8WUU9, Q9BW58, Q9H2W4, Q9NT88 | Gene names | NEDD4L, KIAA0439, NEDL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin-protein ligase NEDD4-like protein (EC 6.3.2.-) (Nedd4-2) (NEDD4.2). | |||||
|
NIPBL_HUMAN
|
||||||
| θ value | 4.03905 (rank : 55) | NC score | 0.055026 (rank : 42) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
OGFR_MOUSE
|
||||||
| θ value | 4.03905 (rank : 56) | NC score | 0.029402 (rank : 69) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 584 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q99PG2, Q99L33 | Gene names | Ogfr | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor). | |||||
|
PDE10_HUMAN
|
||||||
| θ value | 4.03905 (rank : 57) | NC score | 0.012562 (rank : 87) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y233, Q9Y5T1 | Gene names | PDE10A | |||
|
Domain Architecture |

|
|||||
| Description | cAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A (EC 3.1.4.17). | |||||
|
SETD2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 58) | NC score | 0.087522 (rank : 20) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9BYW2, O75397, O75405, Q17RW8, Q5BKS9, Q5QGN2, Q69YI5, Q6IN64, Q6ZN53, Q6ZS25, Q8N3R0, Q8TCN0, Q9C0D1, Q9H696, Q9NZW9 | Gene names | SETD2, HIF1, HYPB, KIAA1732, SET2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36-specific (EC 2.1.1.43) (SET domain-containing protein 2) (hSET2) (Huntingtin- interacting protein HYPB) (Huntingtin-interacting protein 1) (HIF-1) (p231HBP). | |||||
|
ADA17_HUMAN
|
||||||
| θ value | 5.27518 (rank : 59) | NC score | 0.011223 (rank : 90) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P78536, O60226 | Gene names | ADAM17, CSVP, TACE | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 17 precursor (EC 3.4.24.86) (A disintegrin and metalloproteinase domain 17) (TNF-alpha-converting enzyme) (TNF-alpha convertase) (Snake venom-like protease) (CD156b antigen). | |||||
|
GANAB_MOUSE
|
||||||
| θ value | 5.27518 (rank : 60) | NC score | 0.014199 (rank : 84) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BHN3, O08794, Q80U81, Q9CS53 | Gene names | Ganab, G2an, Kiaa0088 | |||
|
Domain Architecture |

|
|||||
| Description | Neutral alpha-glucosidase AB precursor (EC 3.2.1.84) (Glucosidase II subunit alpha) (Alpha glucosidase 2). | |||||
|
GAS8_HUMAN
|
||||||
| θ value | 5.27518 (rank : 61) | NC score | 0.014550 (rank : 81) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 819 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O95995 | Gene names | GAS8, GAS11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Growth-arrest-specific protein 8 (Growth arrest-specific 11). | |||||
|
LEO1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 62) | NC score | 0.059542 (rank : 32) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
LRRF1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 63) | NC score | 0.032370 (rank : 64) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 625 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q32MZ4, O75766, O75799, Q32MZ5, Q53T49, Q6PKG2 | Gene names | LRRFIP1, GCF2, TRIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat flightless-interacting protein 1 (LRR FLII- interacting protein 1) (TAR RNA-interacting protein) (GC-binding factor 2). | |||||
|
LSP1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 64) | NC score | 0.036889 (rank : 59) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P33241, Q16096, Q9BUY8 | Gene names | LSP1, WP34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lymphocyte-specific protein 1 (Protein pp52) (52 kDa phosphoprotein) (Lymphocyte-specific antigen WP34) (47 kDa actin-binding protein). | |||||
|
MOR2A_MOUSE
|
||||||
| θ value | 5.27518 (rank : 65) | NC score | 0.014809 (rank : 80) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q69ZX6, Q5QNQ7, Q6P547 | Gene names | Morc2a, Kiaa0852, Zcwcc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORC family CW-type zinc finger protein 2A (Zinc finger CW-type coiled-coil domain protein 1). | |||||
|
NCKX1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 66) | NC score | 0.036634 (rank : 60) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O60721, O43485, O75184 | Gene names | SLC24A1, KIAA0702, NCKX1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium/calcium exchanger 1 (Na(+)/K(+)/Ca(2+)-exchange protein 1) (Retinal rod Na-Ca+K exchanger). | |||||
|
PCIF1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 67) | NC score | 0.035309 (rank : 62) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9H4Z3, Q9NT85 | Gene names | PCIF1, C20orf67 | |||
|
Domain Architecture |
|
|||||
| Description | Phosphorylated CTD-interacting factor 1. | |||||
|
TCF8_HUMAN
|
||||||
| θ value | 5.27518 (rank : 68) | NC score | 0.012750 (rank : 86) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1057 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P37275, Q12924, Q13800 | Gene names | TCF8, AREB6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 8 (NIL-2-A zinc finger protein) (Negative regulator of IL2). | |||||
|
WWP1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 69) | NC score | 0.058735 (rank : 34) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8BZZ3, Q8BIV9, Q8VDP8 | Gene names | Wwp1 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP1 (EC 6.3.2.-) (WW domain- containing protein 1). | |||||
|
DYNA_HUMAN
|
||||||
| θ value | 6.88961 (rank : 70) | NC score | 0.021827 (rank : 77) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1194 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q14203, O95296, Q9BRM9, Q9UIU1, Q9UIU2 | Gene names | DCTN1 | |||
|
Domain Architecture |

|
|||||
| Description | Dynactin-1 (150 kDa dynein-associated polypeptide) (DP-150) (DAP-150) (p150-glued) (p135). | |||||
|
DYNA_MOUSE
|
||||||
| θ value | 6.88961 (rank : 71) | NC score | 0.022919 (rank : 75) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1084 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O08788 | Gene names | Dctn1 | |||
|
Domain Architecture |

|
|||||
| Description | Dynactin-1 (150 kDa dynein-associated polypeptide) (DP-150) (DAP-150) (p150-glued). | |||||
|
LEO1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 72) | NC score | 0.063982 (rank : 29) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
NEDD4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 73) | NC score | 0.046356 (rank : 54) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P46935, O08758, Q3UZI2, Q8BGB3 | Gene names | Nedd4, Kiaa0093, Nedd-4, Nedd4a | |||
|
Domain Architecture |

|
|||||
| Description | E3 ubiquitin-protein ligase NEDD4 (EC 6.3.2.-) (Neural precursor cell expressed developmentally down-regulated protein 4). | |||||
|
NRX1A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 74) | NC score | 0.007889 (rank : 95) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9ULB1, O60323, Q53TJ9, Q53TQ1, Q9C079, Q9C080, Q9C081, Q9H3M2, Q9UDM6 | Gene names | NRXN1, KIAA0578 | |||
|
Domain Architecture |

|
|||||
| Description | Neurexin-1-alpha precursor (Neurexin I-alpha). | |||||
|
PQBP1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 75) | NC score | 0.049540 (rank : 52) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q91VJ5, Q80WW2, Q9ER43, Q9QYY2 | Gene names | Pqbp1, Npw38 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyglutamine-binding protein 1 (Polyglutamine tract-binding protein 1) (PQBP-1) (38 kDa nuclear protein containing a WW domain) (Npw38). | |||||
|
RAD50_MOUSE
|
||||||
| θ value | 6.88961 (rank : 76) | NC score | 0.014449 (rank : 82) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1366 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P70388, Q6PEB0, Q8C2T7, Q9CU59 | Gene names | Rad50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (mRad50). | |||||
|
SUHW4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 77) | NC score | 0.005693 (rank : 97) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 737 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q68FE8, Q3T9B1, Q8BI82 | Gene names | Suhw4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppressor of hairy wing homolog 4. | |||||
|
YO11_MOUSE
|
||||||
| θ value | 6.88961 (rank : 78) | NC score | 0.026342 (rank : 72) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 272 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P11260 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | Hypothetical protein ORF-1137. | |||||
|
AMD_MOUSE
|
||||||
| θ value | 8.99809 (rank : 79) | NC score | 0.014423 (rank : 83) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P97467 | Gene names | Pam | |||
|
Domain Architecture |

|
|||||
| Description | Peptidyl-glycine alpha-amidating monooxygenase precursor (PAM) [Includes: Peptidylglycine alpha-hydroxylating monooxygenase (EC 1.14.17.3) (PHM); Peptidyl-alpha-hydroxyglycine alpha-amidating lyase (EC 4.3.2.5) (Peptidylamidoglycolate lyase) (PAL)]. | |||||
|
EVPL_HUMAN
|
||||||
| θ value | 8.99809 (rank : 80) | NC score | 0.012909 (rank : 85) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 895 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q92817 | Gene names | EVPL | |||
|
Domain Architecture |

|
|||||
| Description | Envoplakin (210 kDa paraneoplastic pemphigus antigen) (p210) (210 kDa cornified envelope precursor protein). | |||||
|
KIF5C_HUMAN
|
||||||
| θ value | 8.99809 (rank : 81) | NC score | 0.010943 (rank : 92) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 974 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O60282, O95079 | Gene names | KIF5C, KIAA0531, NKHC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin heavy chain isoform 5C (Kinesin heavy chain neuron-specific 2). | |||||
|
KIF5C_MOUSE
|
||||||
| θ value | 8.99809 (rank : 82) | NC score | 0.011002 (rank : 91) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1040 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P28738, Q9Z2F8 | Gene names | Kif5c, Nkhc2 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin heavy chain isoform 5C (Kinesin heavy chain neuron-specific 2). | |||||
|
LRC50_HUMAN
|
||||||
| θ value | 8.99809 (rank : 83) | NC score | 0.012402 (rank : 88) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8NEP3, Q69YI8, Q69YJ0, Q96LP3 | Gene names | LRRC50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 50. | |||||
|
MYT1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 84) | NC score | 0.031298 (rank : 65) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8CFC2, O08995, Q8CFH1 | Gene names | Myt1, Kiaa0835, Nzf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1 (MyT1) (Neural zinc finger factor 2) (NZF-2). | |||||
|
ROCK2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 85) | NC score | 0.003429 (rank : 98) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 2073 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O75116, Q9UQN5 | Gene names | ROCK2, KIAA0619 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 2 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 2) (p164 ROCK-2) (Rho kinase 2). | |||||
|
WDFY3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 86) | NC score | 0.007264 (rank : 96) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6VNB8, Q8C8H7 | Gene names | Wdfy3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat and FYVE domain-containing protein 3 (Beach domain, WD repeat and FYVE domain-containing protein 1) (BWF1). | |||||
|
CCD55_MOUSE
|
||||||
| θ value | θ > 10 (rank : 87) | NC score | 0.054432 (rank : 45) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q5NCR9 | Gene names | Ccdc55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 55. | |||||
|
CYLC2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 88) | NC score | 0.062315 (rank : 31) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q14093 | Gene names | CYLC2, CYL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-2 (Cylicin II) (Multiple-band polypeptide II). | |||||
|
DMP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 89) | NC score | 0.054773 (rank : 44) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q13316, O43265 | Gene names | DMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1). | |||||
|
K1688_MOUSE
|
||||||
| θ value | θ > 10 (rank : 90) | NC score | 0.057153 (rank : 37) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P59281, Q69ZD4, Q6P9R6 | Gene names | Kiaa1688, D15Wsu169e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1688. | |||||
|
NUCKS_HUMAN
|
||||||
| θ value | θ > 10 (rank : 91) | NC score | 0.050873 (rank : 51) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9H1E3, Q9H1D6, Q9H723 | Gene names | NUCKS1, NUCKS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (P1). | |||||
|
NUCKS_MOUSE
|
||||||
| θ value | θ > 10 (rank : 92) | NC score | 0.059107 (rank : 33) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q80XU3, Q8BVD8 | Gene names | Nucks1, Nucks | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (JC7). | |||||
|
PINL_HUMAN
|
||||||
| θ value | θ > 10 (rank : 93) | NC score | 0.080085 (rank : 23) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O15428 | Gene names | PIN1L | |||
|
Domain Architecture |
|
|||||
| Description | PIN1-like protein. | |||||
|
PKCB1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 94) | NC score | 0.052536 (rank : 50) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 488 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9ULU4, Q13517, Q4JJ94, Q4JJ95, Q5TH09, Q6MZM1, Q8WXC5, Q9H1F3, Q9H1F4, Q9H1F5, Q9H1L8, Q9H1L9, Q9H2G5, Q9NYN3, Q9UIX6 | Gene names | PRKCBP1, KIAA1125, RACK7, ZMYND8 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C-binding protein 1 (Rack7) (Cutaneous T-cell lymphoma- associated antigen se14-3) (CTCL tumor antigen se14-3) (Zinc finger MYND domain-containing protein 8). | |||||
|
WWP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 95) | NC score | 0.052899 (rank : 49) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9H0M0, O00307, Q96BP4 | Gene names | WWP1 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP1 (EC 6.3.2.-) (WW domain- containing protein 1) (Atropin-1-interacting protein 5) (AIP5). | |||||
|
WWP2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 96) | NC score | 0.053656 (rank : 46) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O00308, Q96CZ2, Q9BWN6 | Gene names | WWP2 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP2 (EC 6.3.2.-) (WW domain- containing protein 2) (Atrophin-1-interacting protein 2) (AIP2). | |||||
|
WWP2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 97) | NC score | 0.053483 (rank : 47) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9DBH0, Q8BTG4, Q923F6 | Gene names | Wwp2 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP2 (EC 6.3.2.-) (WW domain- containing protein 2). | |||||
|
YAP1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 98) | NC score | 0.053303 (rank : 48) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P46938, Q52KJ5 | Gene names | Yap1, Yap, Yap65 | |||
|
Domain Architecture |

|
|||||
| Description | 65 kDa Yes-associated protein (YAP65). | |||||
|
WBP4_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 3.47952e-169 (rank : 1) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 86 | |
| SwissProt Accessions | O75554 | Gene names | WBP4, FBP21, FNBP21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 4 (WBP-4) (Formin-binding protein 21). | |||||
|
WBP4_MOUSE
|
||||||
| NC score | 0.942255 (rank : 2) | θ value | 2.52267e-127 (rank : 2) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q61048, Q8K1Z9, Q9CS45 | Gene names | Wbp4, Fbp21, Fnbp21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 4 (WBP-4) (Formin-binding protein 21). | |||||
|
PRP40_MOUSE
|
||||||
| NC score | 0.406196 (rank : 3) | θ value | 7.34386e-10 (rank : 4) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9R1C7, Q61049, Q8BQ76, Q8BRW4, Q8C2U1 | Gene names | Prpf40a, Fbp11, Fnbp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Formin-binding protein 11) (FBP-11). | |||||
|
PRP40_HUMAN
|
||||||
| NC score | 0.405093 (rank : 4) | θ value | 7.34386e-10 (rank : 3) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 442 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O75400, O43856, O75404, Q8TBQ1, Q9H782, Q9NWU9, Q9P0Q2, Q9Y5A8 | Gene names | PRPF40A, FLAF1, FNBP3, HYPA | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Huntingtin yeast partner A) (Huntingtin-interacting protein HYPA/FBP11) (Fas ligand-associated factor 1) (NY-REN-6 antigen). | |||||
|
RU1C_HUMAN
|
||||||
| NC score | 0.230562 (rank : 5) | θ value | 0.0148317 (rank : 8) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P09234 | Gene names | SNRPC | |||
|
Domain Architecture |

|
|||||
| Description | U1 small nuclear ribonucleoprotein C (U1 snRNP protein C) (U1C protein) (U1-C). | |||||
|
RU1C_MOUSE
|
||||||
| NC score | 0.228862 (rank : 6) | θ value | 0.0148317 (rank : 9) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q62241 | Gene names | Snrp1c, Snrpc | |||
|
Domain Architecture |

|
|||||
| Description | U1 small nuclear ribonucleoprotein C (U1 snRNP protein C) (U1C protein) (U1-C). | |||||
|
TCRG1_HUMAN
|
||||||
| NC score | 0.184246 (rank : 7) | θ value | 0.00228821 (rank : 5) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 929 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O14776 | Gene names | TCERG1, CA150, TAF2S | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150). | |||||
|
TCRG1_MOUSE
|
||||||
| NC score | 0.181522 (rank : 8) | θ value | 0.00228821 (rank : 6) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8CGF7, Q61051, Q8C490, Q8CHT8, Q9R0R5 | Gene names | Tcerg1, Taf2s | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150) (p144) (Formin- binding protein 28) (FBP 28). | |||||
|
IQGA1_HUMAN
|
||||||
| NC score | 0.156523 (rank : 9) | θ value | 0.0113563 (rank : 7) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P46940 | Gene names | IQGAP1, KIAA0051 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP1 (p195). | |||||
|
IQGA1_MOUSE
|
||||||
| NC score | 0.151935 (rank : 10) | θ value | 0.0252991 (rank : 10) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9JKF1 | Gene names | Iqgap1 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP1. | |||||
|
IQGA3_HUMAN
|
||||||
| NC score | 0.150241 (rank : 11) | θ value | 0.47712 (rank : 21) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q86VI3 | Gene names | IQGAP3 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP3. | |||||
|
GAS7_MOUSE
|
||||||
| NC score | 0.147562 (rank : 12) | θ value | 0.0330416 (rank : 11) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q60780, Q9QY25, Q9QY26, Q9QY34 | Gene names | Gas7 | |||
|
Domain Architecture |

|
|||||
| Description | Growth-arrest-specific protein 7 (GAS-7). | |||||
|
IQGA2_HUMAN
|
||||||
| NC score | 0.144408 (rank : 13) | θ value | 0.21417 (rank : 17) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q13576 | Gene names | IQGAP2 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP2. | |||||
|
GAS7_HUMAN
|
||||||
| NC score | 0.138185 (rank : 14) | θ value | 0.21417 (rank : 15) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O60861, O43144 | Gene names | GAS7, KIAA0394 | |||
|
Domain Architecture |

|
|||||
| Description | Growth-arrest-specific protein 7 (GAS-7). | |||||
|
PIN1_MOUSE
|
||||||
| NC score | 0.114665 (rank : 15) | θ value | 3.0926 (rank : 44) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9QUR7, Q543B3 | Gene names | Pin1 | |||
|
Domain Architecture |

|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase NIMA-interacting 1 (EC 5.2.1.8) (Rotamase Pin1) (PPIase Pin1). | |||||
|
PIN1_HUMAN
|
||||||
| NC score | 0.100121 (rank : 16) | θ value | 3.0926 (rank : 43) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q13526 | Gene names | PIN1 | |||
|
Domain Architecture |

|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase NIMA-interacting 1 (EC 5.2.1.8) (Rotamase Pin1) (PPIase Pin1). | |||||
|
K1688_HUMAN
|
||||||
| NC score | 0.098450 (rank : 17) | θ value | 0.0961366 (rank : 13) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9C0H5 | Gene names | KIAA1688 | |||
|
Domain Architecture |

|
|||||
| Description | Protein KIAA1688. | |||||
|
YTDC1_HUMAN
|
||||||
| NC score | 0.095236 (rank : 18) | θ value | 0.0736092 (rank : 12) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q96MU7, Q7Z622, Q8TF35 | Gene names | YTHDC1, KIAA1966, YT521 | |||
|
Domain Architecture |

|
|||||
| Description | YTH domain-containing protein 1 (Putative splicing factor YT521). | |||||
|
SEC63_MOUSE
|
||||||
| NC score | 0.088375 (rank : 19) | θ value | 0.365318 (rank : 19) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 601 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8VHE0, Q8VEB9 | Gene names | Sec63, Sec63l | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
SETD2_HUMAN
|
||||||
| NC score | 0.087522 (rank : 20) | θ value | 4.03905 (rank : 58) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9BYW2, O75397, O75405, Q17RW8, Q5BKS9, Q5QGN2, Q69YI5, Q6IN64, Q6ZN53, Q6ZS25, Q8N3R0, Q8TCN0, Q9C0D1, Q9H696, Q9NZW9 | Gene names | SETD2, HIF1, HYPB, KIAA1732, SET2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36-specific (EC 2.1.1.43) (SET domain-containing protein 2) (hSET2) (Huntingtin- interacting protein HYPB) (Huntingtin-interacting protein 1) (HIF-1) (p231HBP). | |||||
|
SEC63_HUMAN
|
||||||
| NC score | 0.082591 (rank : 21) | θ value | 0.62314 (rank : 23) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UGP8, O95380, Q86VS9, Q8IWL0, Q9NTE0 | Gene names | SEC63, SEC63L | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.082115 (rank : 22) | θ value | 2.36792 (rank : 36) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
PINL_HUMAN
|
||||||
| NC score | 0.080085 (rank : 23) | θ value | θ > 10 (rank : 93) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O15428 | Gene names | PIN1L | |||
|
Domain Architecture |

|
|||||
| Description | PIN1-like protein. | |||||
|
RHG12_MOUSE
|
||||||
| NC score | 0.078215 (rank : 24) | θ value | 1.06291 (rank : 28) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8C0D4, Q8BVP8 | Gene names | Arhgap12 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 12. | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.077766 (rank : 25) | θ value | 1.06291 (rank : 27) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
OXR1_MOUSE
|
||||||
| NC score | 0.072979 (rank : 26) | θ value | 0.21417 (rank : 18) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q4KMM3, Q5FWW1, Q99L06, Q99MK1, Q99MP4 | Gene names | Oxr1, C7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oxidation resistance protein 1 (Protein C7). | |||||
|
RHG12_HUMAN
|
||||||
| NC score | 0.068825 (rank : 27) | θ value | 3.0926 (rank : 45) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8IWW6, Q86UB3, Q8IWW7, Q8N3L1, Q9NT76 | Gene names | ARHGAP12 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 12. | |||||
|
NKTR_HUMAN
|
||||||
| NC score | 0.064874 (rank : 28) | θ value | 0.0961366 (rank : 14) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 658 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P30414 | Gene names | NKTR | |||
|
Domain Architecture |

|
|||||
| Description | NK-tumor recognition protein (Natural-killer cells cyclophilin-related protein) (NK-TR protein). | |||||
|
LEO1_HUMAN
|
||||||
| NC score | 0.063982 (rank : 29) | θ value | 6.88961 (rank : 72) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
PSIP1_MOUSE
|
||||||
| NC score | 0.062825 (rank : 30) | θ value | 0.813845 (rank : 25) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q99JF8, Q3TEJ7, Q3TTD7, Q3UTA1, Q80WQ7, Q99JF7, Q99LR4, Q9CT03 | Gene names | Psip1, Ledgf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PC4 and SFRS1-interacting protein (Lens epithelium-derived growth factor) (mLEDGF). | |||||
|
CYLC2_HUMAN
|
||||||
| NC score | 0.062315 (rank : 31) | θ value | θ > 10 (rank : 88) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q14093 | Gene names | CYLC2, CYL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-2 (Cylicin II) (Multiple-band polypeptide II). | |||||
|
LEO1_MOUSE
|
||||||
| NC score | 0.059542 (rank : 32) | θ value | 5.27518 (rank : 62) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
NUCKS_MOUSE
|
||||||
| NC score | 0.059107 (rank : 33) | θ value | θ > 10 (rank : 92) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q80XU3, Q8BVD8 | Gene names | Nucks1, Nucks | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (JC7). | |||||
|
WWP1_MOUSE
|
||||||
| NC score | 0.058735 (rank : 34) | θ value | 5.27518 (rank : 69) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8BZZ3, Q8BIV9, Q8VDP8 | Gene names | Wwp1 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP1 (EC 6.3.2.-) (WW domain- containing protein 1). | |||||
|
DMP1_MOUSE
|
||||||
| NC score | 0.058531 (rank : 35) | θ value | 4.03905 (rank : 49) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O55188 | Gene names | Dmp1, Dmp | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1) (AG1). | |||||
|
MYRIP_HUMAN
|
||||||
| NC score | 0.058460 (rank : 36) | θ value | 0.813845 (rank : 24) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8NFW9, Q8IUF5, Q9Y3V4 | Gene names | MYRIP, SLAC2C | |||
|
Domain Architecture |

|
|||||
| Description | Rab effector MyRIP (Myosin-VIIa- and Rab-interacting protein) (Exophilin-8) (Slp homolog lacking C2 domains c). | |||||
|
K1688_MOUSE
|
||||||
| NC score | 0.057153 (rank : 37) | θ value | θ > 10 (rank : 90) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P59281, Q69ZD4, Q6P9R6 | Gene names | Kiaa1688, D15Wsu169e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1688. | |||||
|
GP179_HUMAN
|
||||||
| NC score | 0.056607 (rank : 38) | θ value | 0.21417 (rank : 16) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 372 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q6PRD1 | Gene names | GPR179, GPR158L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 179 precursor (Probable G-protein coupled receptor 158-like 1). | |||||
|
NPBL_MOUSE
|
||||||
| NC score | 0.055927 (rank : 39) | θ value | 3.0926 (rank : 42) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
DGC14_MOUSE
|
||||||
| NC score | 0.055874 (rank : 40) | θ value | 4.03905 (rank : 48) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O70279, Q91YX1 | Gene names | Dgcr14, Dgsi, Es2, Es2el | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DGCR14 protein (DiGeorge syndrome critical region 14 homolog) (ES2 protein) (Expressed sequence 2 embryonic lethal). | |||||
|
ITCH_MOUSE
|
||||||
| NC score | 0.055423 (rank : 41) | θ value | 2.36792 (rank : 35) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8C863, O54971 | Gene names | Itch | |||
|
Domain Architecture |

|
|||||
| Description | Itchy E3 ubiquitin protein ligase (EC 6.3.2.-). | |||||
|
NIPBL_HUMAN
|
||||||
| NC score | 0.055026 (rank : 42) | θ value | 4.03905 (rank : 55) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
ITCH_HUMAN
|
||||||
| NC score | 0.054943 (rank : 43) | θ value | 2.36792 (rank : 34) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q96J02, O43584, Q5TEL0, Q96F66, Q9BY75, Q9H451, Q9H4U5 | Gene names | ITCH | |||
|
Domain Architecture |

|
|||||
| Description | Itchy homolog E3 ubiquitin protein ligase (EC 6.3.2.-) (Itch) (Atrophin-1-interacting protein 4) (AIP4) (NFE2-associated polypeptide 1) (NAPP1). | |||||
|
DMP1_HUMAN
|
||||||
| NC score | 0.054773 (rank : 44) | θ value | θ > 10 (rank : 89) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q13316, O43265 | Gene names | DMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1). | |||||
|
CCD55_MOUSE
|
||||||
| NC score | 0.054432 (rank : 45) | θ value | θ > 10 (rank : 87) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q5NCR9 | Gene names | Ccdc55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 55. | |||||
|
WWP2_HUMAN
|
||||||
| NC score | 0.053656 (rank : 46) | θ value | θ > 10 (rank : 96) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O00308, Q96CZ2, Q9BWN6 | Gene names | WWP2 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP2 (EC 6.3.2.-) (WW domain- containing protein 2) (Atrophin-1-interacting protein 2) (AIP2). | |||||
|
WWP2_MOUSE
|
||||||
| NC score | 0.053483 (rank : 47) | θ value | θ > 10 (rank : 97) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9DBH0, Q8BTG4, Q923F6 | Gene names | Wwp2 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP2 (EC 6.3.2.-) (WW domain- containing protein 2). | |||||
|
YAP1_MOUSE
|
||||||
| NC score | 0.053303 (rank : 48) | θ value | θ > 10 (rank : 98) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P46938, Q52KJ5 | Gene names | Yap1, Yap, Yap65 | |||
|
Domain Architecture |

|
|||||
| Description | 65 kDa Yes-associated protein (YAP65). | |||||
|
WWP1_HUMAN
|
||||||
| NC score | 0.052899 (rank : 49) | θ value | θ > 10 (rank : 95) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9H0M0, O00307, Q96BP4 | Gene names | WWP1 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP1 (EC 6.3.2.-) (WW domain- containing protein 1) (Atropin-1-interacting protein 5) (AIP5). | |||||
|
PKCB1_HUMAN
|
||||||
| NC score | 0.052536 (rank : 50) | θ value | θ > 10 (rank : 94) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 488 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9ULU4, Q13517, Q4JJ94, Q4JJ95, Q5TH09, Q6MZM1, Q8WXC5, Q9H1F3, Q9H1F4, Q9H1F5, Q9H1L8, Q9H1L9, Q9H2G5, Q9NYN3, Q9UIX6 | Gene names | PRKCBP1, KIAA1125, RACK7, ZMYND8 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C-binding protein 1 (Rack7) (Cutaneous T-cell lymphoma- associated antigen se14-3) (CTCL tumor antigen se14-3) (Zinc finger MYND domain-containing protein 8). | |||||
|
NUCKS_HUMAN
|
||||||
| NC score | 0.050873 (rank : 51) | θ value | θ > 10 (rank : 91) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9H1E3, Q9H1D6, Q9H723 | Gene names | NUCKS1, NUCKS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (P1). | |||||
|
PQBP1_MOUSE
|
||||||
| NC score | 0.049540 (rank : 52) | θ value | 6.88961 (rank : 75) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q91VJ5, Q80WW2, Q9ER43, Q9QYY2 | Gene names | Pqbp1, Npw38 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyglutamine-binding protein 1 (Polyglutamine tract-binding protein 1) (PQBP-1) (38 kDa nuclear protein containing a WW domain) (Npw38). | |||||
|
MYRIP_MOUSE
|
||||||
| NC score | 0.049152 (rank : 53) | θ value | 2.36792 (rank : 37) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8K3I4, Q8CFC0, Q8K4H5 | Gene names | Myrip, Slac2c | |||
|
Domain Architecture |

|
|||||
| Description | Rab effector MyRIP (Myosin-VIIa- and Rab-interacting protein) (Exophilin-8) (Slp homolog lacking C2 domains c). | |||||
|
NEDD4_MOUSE
|
||||||
| NC score | 0.046356 (rank : 54) | θ value | 6.88961 (rank : 73) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P46935, O08758, Q3UZI2, Q8BGB3 | Gene names | Nedd4, Kiaa0093, Nedd-4, Nedd4a | |||
|
Domain Architecture |

|
|||||
| Description | E3 ubiquitin-protein ligase NEDD4 (EC 6.3.2.-) (Neural precursor cell expressed developmentally down-regulated protein 4). | |||||
|
NED4L_HUMAN
|
||||||
| NC score | 0.046219 (rank : 55) | θ value | 4.03905 (rank : 54) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q96PU5, O43165, Q7Z5F1, Q7Z5F2, Q7Z5N3, Q8N5A7, Q8WUU9, Q9BW58, Q9H2W4, Q9NT88 | Gene names | NEDD4L, KIAA0439, NEDL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin-protein ligase NEDD4-like protein (EC 6.3.2.-) (Nedd4-2) (NEDD4.2). | |||||
|
RPTN_MOUSE
|
||||||
| NC score | 0.045930 (rank : 56) | θ value | 0.62314 (rank : 22) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P97347 | Gene names | Rptn | |||
|
Domain Architecture |

|
|||||
| Description | Repetin. | |||||
|
HORN_MOUSE
|
||||||
| NC score | 0.042910 (rank : 57) | θ value | 1.81305 (rank : 31) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8VHD8 | Gene names | Hrnr | |||
|
Domain Architecture |

|
|||||
| Description | Hornerin. | |||||
|
MYT1_HUMAN
|
||||||
| NC score | 0.038372 (rank : 58) | θ value | 4.03905 (rank : 53) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 475 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q01538, O94922, Q9UPV2 | Gene names | MYT1, KIAA0835, KIAA1050, MTF1, MYTI, PLPB1 | |||
|
Domain Architecture |

|
|||||
| Description | Myelin transcription factor 1 (MyT1) (MyTI) (Proteolipid protein- binding protein) (PLPB1). | |||||
|
LSP1_HUMAN
|
||||||
| NC score | 0.036889 (rank : 59) | θ value | 5.27518 (rank : 64) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P33241, Q16096, Q9BUY8 | Gene names | LSP1, WP34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lymphocyte-specific protein 1 (Protein pp52) (52 kDa phosphoprotein) (Lymphocyte-specific antigen WP34) (47 kDa actin-binding protein). | |||||
|
NCKX1_HUMAN
|
||||||
| NC score | 0.036634 (rank : 60) | θ value | 5.27518 (rank : 66) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O60721, O43485, O75184 | Gene names | SLC24A1, KIAA0702, NCKX1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium/calcium exchanger 1 (Na(+)/K(+)/Ca(2+)-exchange protein 1) (Retinal rod Na-Ca+K exchanger). | |||||
|
RHG18_MOUSE
|
||||||
| NC score | 0.035709 (rank : 61) | θ value | 2.36792 (rank : 39) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8K0Q5, Q8BP03, Q8R196 | Gene names | Arhgap18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho-GTPase-activating protein 18. | |||||
|
PCIF1_HUMAN
|
||||||
| NC score | 0.035309 (rank : 62) | θ value | 5.27518 (rank : 67) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9H4Z3, Q9NT85 | Gene names | PCIF1, C20orf67 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphorylated CTD-interacting factor 1. | |||||
|
SDPR_HUMAN
|
||||||
| NC score | 0.035212 (rank : 63) | θ value | 1.81305 (rank : 33) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O95810 | Gene names | SDPR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serum deprivation-response protein (Phosphatidylserine-binding protein) (PS-p68). | |||||
|
LRRF1_HUMAN
|
||||||
| NC score | 0.032370 (rank : 64) | θ value | 5.27518 (rank : 63) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 625 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q32MZ4, O75766, O75799, Q32MZ5, Q53T49, Q6PKG2 | Gene names | LRRFIP1, GCF2, TRIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat flightless-interacting protein 1 (LRR FLII- interacting protein 1) (TAR RNA-interacting protein) (GC-binding factor 2). | |||||
|
MYT1_MOUSE
|
||||||
| NC score | 0.031298 (rank : 65) | θ value | 8.99809 (rank : 84) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8CFC2, O08995, Q8CFH1 | Gene names | Myt1, Kiaa0835, Nzf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1 (MyT1) (Neural zinc finger factor 2) (NZF-2). | |||||
|
ANR11_HUMAN
|
||||||
| NC score | 0.031096 (rank : 66) | θ value | 4.03905 (rank : 46) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1289 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q6UB99, Q6NTG1, Q6QMF8 | Gene names | ANKRD11, ANCO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 11 (Ankyrin repeat-containing cofactor 1). | |||||
|
HSP7C_MOUSE
|
||||||
| NC score | 0.030029 (rank : 67) | θ value | 0.47712 (rank : 20) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P63017, P08109, P12225, Q62373, Q62374, Q62375 | Gene names | Hspa8, Hsc70, Hsc73 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heat shock cognate 71 kDa protein (Heat shock 70 kDa protein 8). | |||||
|
SIA7A_HUMAN
|
||||||
| NC score | 0.029984 (rank : 68) | θ value | 1.38821 (rank : 29) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9NSC7, Q6UW90, Q9NSC6 | Gene names | ST6GALNAC1, SIAT7A | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-N-acetylgalactosaminide alpha-2,6-sialyltransferase 1 (EC 2.4.99.3) (GalNAc alpha-2,6-sialyltransferase I) (ST6GalNAc I) (Sialyltransferase 7A). | |||||
|
OGFR_MOUSE
|
||||||
| NC score | 0.029402 (rank : 69) | θ value | 4.03905 (rank : 56) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 584 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q99PG2, Q99L33 | Gene names | Ogfr | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor). | |||||
|
NFM_HUMAN
|
||||||
| NC score | 0.029209 (rank : 70) | θ value | 1.81305 (rank : 32) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1328 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P07197 | Gene names | NEF3, NEFM, NFM | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet M protein (160 kDa neurofilament protein) (Neurofilament medium polypeptide) (NF-M) (Neurofilament 3). | |||||
|
MINT_MOUSE
|
||||||
| NC score | 0.026773 (rank : 71) | θ value | 3.0926 (rank : 41) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
YO11_MOUSE
|
||||||
| NC score | 0.026342 (rank : 72) | θ value | 6.88961 (rank : 78) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 272 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P11260 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | Hypothetical protein ORF-1137. | |||||
|
HSP7C_HUMAN
|
||||||
| NC score | 0.024882 (rank : 73) | θ value | 4.03905 (rank : 50) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P11142, Q9H3R6 | Gene names | HSPA8, HSC70, HSP73, HSPA10 | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock cognate 71 kDa protein (Heat shock 70 kDa protein 8). | |||||
|
IRX3_HUMAN
|
||||||
| NC score | 0.023783 (rank : 74) | θ value | 3.0926 (rank : 40) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P78415, Q7Z4A4, Q7Z4A5, Q8IVC6 | Gene names | IRX3, IRXB1 | |||
|
Domain Architecture |

|
|||||
| Description | Iroquois-class homeodomain protein IRX-3 (Iroquois homeobox protein 3) (Homeodomain protein IRXB1). | |||||
|
DYNA_MOUSE
|
||||||
| NC score | 0.022919 (rank : 75) | θ value | 6.88961 (rank : 71) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1084 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O08788 | Gene names | Dctn1 | |||
|
Domain Architecture |

|
|||||
| Description | Dynactin-1 (150 kDa dynein-associated polypeptide) (DP-150) (DAP-150) (p150-glued). | |||||
|
IRX3_MOUSE
|
||||||
| NC score | 0.022104 (rank : 76) | θ value | 4.03905 (rank : 51) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P81067 | Gene names | Irx3, Irxb1 | |||
|
Domain Architecture |

|
|||||
| Description | Iroquois-class homeodomain protein IRX-3 (Iroquois homeobox protein 3) (Homeodomain protein IRXB1). | |||||
|
DYNA_HUMAN
|
||||||
| NC score | 0.021827 (rank : 77) | θ value | 6.88961 (rank : 70) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1194 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q14203, O95296, Q9BRM9, Q9UIU1, Q9UIU2 | Gene names | DCTN1 | |||
|
Domain Architecture |

|
|||||
| Description | Dynactin-1 (150 kDa dynein-associated polypeptide) (DP-150) (DAP-150) (p150-glued) (p135). | |||||
|
PHF10_MOUSE
|
||||||
| NC score | 0.021491 (rank : 78) | θ value | 2.36792 (rank : 38) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9D8M7, Q99LV5 | Gene names | Phf10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 10. | |||||
|
CEP63_HUMAN
|
||||||
| NC score | 0.021259 (rank : 79) | θ value | 1.81305 (rank : 30) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1005 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q96MT8, Q96CR0, Q9H8F5, Q9H8N0 | Gene names | CEP63 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 63 kDa (Cep63 protein). | |||||
|
MOR2A_MOUSE
|
||||||
| NC score | 0.014809 (rank : 80) | θ value | 5.27518 (rank : 65) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q69ZX6, Q5QNQ7, Q6P547 | Gene names | Morc2a, Kiaa0852, Zcwcc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORC family CW-type zinc finger protein 2A (Zinc finger CW-type coiled-coil domain protein 1). | |||||
|
GAS8_HUMAN
|
||||||
| NC score | 0.014550 (rank : 81) | θ value | 5.27518 (rank : 61) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 819 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O95995 | Gene names | GAS8, GAS11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Growth-arrest-specific protein 8 (Growth arrest-specific 11). | |||||
|
RAD50_MOUSE
|
||||||
| NC score | 0.014449 (rank : 82) | θ value | 6.88961 (rank : 76) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1366 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P70388, Q6PEB0, Q8C2T7, Q9CU59 | Gene names | Rad50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (mRad50). | |||||
|
AMD_MOUSE
|
||||||
| NC score | 0.014423 (rank : 83) | θ value | 8.99809 (rank : 79) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P97467 | Gene names | Pam | |||
|
Domain Architecture |

|
|||||
| Description | Peptidyl-glycine alpha-amidating monooxygenase precursor (PAM) [Includes: Peptidylglycine alpha-hydroxylating monooxygenase (EC 1.14.17.3) (PHM); Peptidyl-alpha-hydroxyglycine alpha-amidating lyase (EC 4.3.2.5) (Peptidylamidoglycolate lyase) (PAL)]. | |||||
|
GANAB_MOUSE
|
||||||
| NC score | 0.014199 (rank : 84) | θ value | 5.27518 (rank : 60) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BHN3, O08794, Q80U81, Q9CS53 | Gene names | Ganab, G2an, Kiaa0088 | |||
|
Domain Architecture |

|
|||||
| Description | Neutral alpha-glucosidase AB precursor (EC 3.2.1.84) (Glucosidase II subunit alpha) (Alpha glucosidase 2). | |||||
|
EVPL_HUMAN
|
||||||
| NC score | 0.012909 (rank : 85) | θ value | 8.99809 (rank : 80) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 895 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q92817 | Gene names | EVPL | |||
|
Domain Architecture |

|
|||||
| Description | Envoplakin (210 kDa paraneoplastic pemphigus antigen) (p210) (210 kDa cornified envelope precursor protein). | |||||
|
TCF8_HUMAN
|
||||||
| NC score | 0.012750 (rank : 86) | θ value | 5.27518 (rank : 68) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1057 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P37275, Q12924, Q13800 | Gene names | TCF8, AREB6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 8 (NIL-2-A zinc finger protein) (Negative regulator of IL2). | |||||
|
PDE10_HUMAN
|
||||||
| NC score | 0.012562 (rank : 87) | θ value | 4.03905 (rank : 57) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y233, Q9Y5T1 | Gene names | PDE10A | |||
|
Domain Architecture |

|
|||||
| Description | cAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A (EC 3.1.4.17). | |||||
|
LRC50_HUMAN
|
||||||
| NC score | 0.012402 (rank : 88) | θ value | 8.99809 (rank : 83) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8NEP3, Q69YI8, Q69YJ0, Q96LP3 | Gene names | LRRC50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 50. | |||||
|
TCF8_MOUSE
|
||||||
| NC score | 0.011618 (rank : 89) | θ value | 0.813845 (rank : 26) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1067 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q64318, Q62519 | Gene names | Tcf8, Zfhx1a, Zfx1a, Zfx1ha | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 8 (Zinc finger homeobox protein 1a) (MEB1) (Delta EF1). | |||||
|
ADA17_HUMAN
|
||||||
| NC score | 0.011223 (rank : 90) | θ value | 5.27518 (rank : 59) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P78536, O60226 | Gene names | ADAM17, CSVP, TACE | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 17 precursor (EC 3.4.24.86) (A disintegrin and metalloproteinase domain 17) (TNF-alpha-converting enzyme) (TNF-alpha convertase) (Snake venom-like protease) (CD156b antigen). | |||||
|
KIF5C_MOUSE
|
||||||
| NC score | 0.011002 (rank : 91) | θ value | 8.99809 (rank : 82) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1040 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P28738, Q9Z2F8 | Gene names | Kif5c, Nkhc2 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin heavy chain isoform 5C (Kinesin heavy chain neuron-specific 2). | |||||
|
KIF5C_HUMAN
|
||||||
| NC score | 0.010943 (rank : 92) | θ value | 8.99809 (rank : 81) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 974 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O60282, O95079 | Gene names | KIF5C, KIAA0531, NKHC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin heavy chain isoform 5C (Kinesin heavy chain neuron-specific 2). | |||||
|
LRCH2_HUMAN
|
||||||
| NC score | 0.010336 (rank : 93) | θ value | 4.03905 (rank : 52) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q5VUJ6, Q9HA88, Q9P233 | Gene names | LRCH2, KIAA1495 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 2. | |||||
|
BARH2_HUMAN
|
||||||
| NC score | 0.010028 (rank : 94) | θ value | 4.03905 (rank : 47) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NY43, Q7Z4N7 | Gene names | BARHL2 | |||
|
Domain Architecture |

|
|||||
| Description | BarH-like 2 homeobox protein. | |||||
|
NRX1A_HUMAN
|
||||||
| NC score | 0.007889 (rank : 95) | θ value | 6.88961 (rank : 74) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9ULB1, O60323, Q53TJ9, Q53TQ1, Q9C079, Q9C080, Q9C081, Q9H3M2, Q9UDM6 | Gene names | NRXN1, KIAA0578 | |||
|
Domain Architecture |

|
|||||
| Description | Neurexin-1-alpha precursor (Neurexin I-alpha). | |||||
|
WDFY3_MOUSE
|
||||||
| NC score | 0.007264 (rank : 96) | θ value | 8.99809 (rank : 86) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6VNB8, Q8C8H7 | Gene names | Wdfy3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat and FYVE domain-containing protein 3 (Beach domain, WD repeat and FYVE domain-containing protein 1) (BWF1). | |||||
|
SUHW4_MOUSE
|
||||||
| NC score | 0.005693 (rank : 97) | θ value | 6.88961 (rank : 77) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 737 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q68FE8, Q3T9B1, Q8BI82 | Gene names | Suhw4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppressor of hairy wing homolog 4. | |||||
|
ROCK2_HUMAN
|
||||||
| NC score | 0.003429 (rank : 98) | θ value | 8.99809 (rank : 85) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 2073 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O75116, Q9UQN5 | Gene names | ROCK2, KIAA0619 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 2 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 2) (p164 ROCK-2) (Rho kinase 2). | |||||