Please be patient as the page loads
|
IQGA3_HUMAN
|
||||||
| SwissProt Accessions | Q86VI3 | Gene names | IQGAP3 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP3. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
IQGA1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.936760 (rank : 4) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | P46940 | Gene names | IQGAP1, KIAA0051 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP1 (p195). | |||||
|
IQGA1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.945776 (rank : 2) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q9JKF1 | Gene names | Iqgap1 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP1. | |||||
|
IQGA2_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.940706 (rank : 3) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q13576 | Gene names | IQGAP2 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP2. | |||||
|
IQGA3_HUMAN
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 125 | |
| SwissProt Accessions | Q86VI3 | Gene names | IQGAP3 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP3. | |||||
|
NF1_HUMAN
|
||||||
| θ value | 1.06045e-08 (rank : 5) | NC score | 0.342416 (rank : 6) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P21359, O00662, Q14284, Q14930, Q9UMK3 | Gene names | NF1 | |||
|
Domain Architecture |

|
|||||
| Description | Neurofibromin (Neurofibromatosis-related protein NF-1) [Contains: Neurofibromin truncated]. | |||||
|
NF1_MOUSE
|
||||||
| θ value | 1.38499e-08 (rank : 6) | NC score | 0.343159 (rank : 5) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q04690, Q61956, Q61957 | Gene names | Nf1 | |||
|
Domain Architecture |

|
|||||
| Description | Neurofibromin (Neurofibromatosis-related protein NF-1). | |||||
|
ASPM_HUMAN
|
||||||
| θ value | 2.61198e-07 (rank : 7) | NC score | 0.221602 (rank : 8) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8IZT6, Q86UX4, Q8IUL2, Q8IZJ7, Q8IZJ8, Q8IZJ9, Q8N4D1, Q9NVS1, Q9NVT6 | Gene names | ASPM, MCPH5 | |||
|
Domain Architecture |

|
|||||
| Description | Abnormal spindle-like microcephaly-associated protein (Abnormal spindle protein homolog) (Asp homolog). | |||||
|
ASPM_MOUSE
|
||||||
| θ value | 5.44631e-05 (rank : 8) | NC score | 0.195385 (rank : 15) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8CJ27, O88482, Q8BJI8, Q8BKT4 | Gene names | Aspm, Calmbp1, Sha1 | |||
|
Domain Architecture |

|
|||||
| Description | Abnormal spindle-like microcephaly-associated protein homolog (Calmodulin-binding protein 1) (Spindle and hydroxyurea checkpoint abnormal protein) (Calmodulin-binding protein Sha1). | |||||
|
RASL1_HUMAN
|
||||||
| θ value | 9.29e-05 (rank : 9) | NC score | 0.215207 (rank : 12) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O95294, Q52M03, Q96CC7 | Gene names | RASAL1, RASAL | |||
|
Domain Architecture |

|
|||||
| Description | RasGAP-activating-like protein 1. | |||||
|
MYO5C_HUMAN
|
||||||
| θ value | 0.00020696 (rank : 10) | NC score | 0.074018 (rank : 34) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1206 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9NQX4 | Gene names | MYO5C | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5C (Myosin Vc). | |||||
|
RASL1_MOUSE
|
||||||
| θ value | 0.00020696 (rank : 11) | NC score | 0.216863 (rank : 11) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Z268 | Gene names | Rasal1, Rasal | |||
|
Domain Architecture |

|
|||||
| Description | RasGAP-activating-like protein 1. | |||||
|
RASA1_HUMAN
|
||||||
| θ value | 0.000270298 (rank : 12) | NC score | 0.211493 (rank : 13) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P20936 | Gene names | RASA1, RASA | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 1 (GTPase-activating protein) (GAP) (Ras p21 protein activator) (p120GAP) (RasGAP). | |||||
|
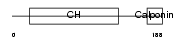
TAGL2_MOUSE
|
||||||
| θ value | 0.00134147 (rank : 13) | NC score | 0.130509 (rank : 19) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9WVA4 | Gene names | Tagln2 | |||
|
Domain Architecture |
|
|||||
| Description | Transgelin-2. | |||||
|
RASA3_HUMAN
|
||||||
| θ value | 0.00298849 (rank : 14) | NC score | 0.166122 (rank : 16) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q14644 | Gene names | RASA3 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 3 (GAP1(IP4BP)) (Ins P4-binding protein). | |||||
|
DAB2P_MOUSE
|
||||||
| θ value | 0.00869519 (rank : 15) | NC score | 0.220180 (rank : 9) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 284 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q3UHC7, Q3TPD5, Q3UH44, Q6JTV1, Q80T97 | Gene names | Dab2ip, Kiaa1743 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Disabled homolog 2-interacting protein (DAB2-interacting protein). | |||||
|
NGAP_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 16) | NC score | 0.222415 (rank : 7) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 517 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9UJF2, O95174, Q5TFU9 | Gene names | RASAL2, NGAP | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein nGAP (RAS protein activator-like 1). | |||||
|
MYO5A_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 17) | NC score | 0.074297 (rank : 33) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1092 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9Y4I1, O60653, Q07902, Q16249, Q9UE30, Q9UE31 | Gene names | MYO5A, MYH12 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5A (Myosin Va) (Dilute myosin heavy chain, non-muscle) (Myosin heavy chain 12) (Myoxin). | |||||
|
DAB2P_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 18) | NC score | 0.217859 (rank : 10) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q5VWQ8, Q8TDL2, Q96SE1, Q9C0C0 | Gene names | DAB2IP, AF9Q34, AIP1, KIAA1743 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Disabled homolog 2-interacting protein (DAB2-interacting protein) (DAB2 interaction protein) (ASK-interacting protein 1). | |||||
|
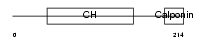
TAGL2_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 19) | NC score | 0.098045 (rank : 25) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P37802, Q5JRQ6, Q5JRQ7, Q9BUH5, Q9H4P0 | Gene names | TAGLN2, KIAA0120 | |||
|
Domain Architecture |
|
|||||
| Description | Transgelin-2 (SM22-alpha homolog). | |||||
|
TACC3_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 20) | NC score | 0.056749 (rank : 77) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9Y6A5, Q9UMQ1 | Gene names | TACC3, ERIC1 | |||
|
Domain Architecture |

|
|||||
| Description | Transforming acidic coiled-coil-containing protein 3 (ERIC-1). | |||||
|
K1683_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 21) | NC score | 0.111005 (rank : 23) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9H0B3, Q8N4G8, Q96M14, Q9C0I0 | Gene names | KIAA1683 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1683. | |||||
|
CK5P2_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 22) | NC score | 0.048607 (rank : 92) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1151 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q8K389, Q6PCN1, Q6ZPL0 | Gene names | Cdk5rap2, Kiaa1633 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CDK5 regulatory subunit-associated protein 2 (CDK5 activator-binding protein C48). | |||||
|
MYO7A_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 23) | NC score | 0.069653 (rank : 39) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P97479 | Gene names | Myo7a, Myo7 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-7A (Myosin VIIa). | |||||
|
MYH11_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 24) | NC score | 0.064292 (rank : 46) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1806 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | P35749, O00396, O94944, P78422 | Gene names | MYH11, KIAA0866 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
MYO5A_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 25) | NC score | 0.075925 (rank : 30) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1117 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q99104 | Gene names | Myo5a, Dilute | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5A (Myosin Va) (Dilute myosin heavy chain, non-muscle). | |||||
|
RASL2_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 26) | NC score | 0.146038 (rank : 18) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O43374, O60286, Q86UW3, Q96QU0 | Gene names | RASA4, CAPRI, GAPL, KIAA0538 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 4 (RasGAP-activating-like protein 2) (Calcium-promoted Ras inactivator). | |||||
|
MYO7A_HUMAN
|
||||||
| θ value | 0.125558 (rank : 27) | NC score | 0.070000 (rank : 38) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q13402, P78427, Q13321, Q14785, Q92821, Q92822 | Gene names | MYO7A, USH1B | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-7A (Myosin VIIa). | |||||
|
RASA3_MOUSE
|
||||||
| θ value | 0.125558 (rank : 28) | NC score | 0.129556 (rank : 20) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q60790 | Gene names | Rasa3 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 3 (GAP1(IP4BP)) (Ins P4-binding protein) (GapIII). | |||||
|
CCD41_HUMAN
|
||||||
| θ value | 0.163984 (rank : 29) | NC score | 0.047521 (rank : 95) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1139 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9Y592 | Gene names | CCDC41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 41 (NY-REN-58 antigen). | |||||
|
LMO7_HUMAN
|
||||||
| θ value | 0.163984 (rank : 30) | NC score | 0.067070 (rank : 41) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8WWI1, O15462, O95346, Q9UKC1, Q9UQM5, Q9Y6A7 | Gene names | LMO7, FBX20, FBXO20, KIAA0858 | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain only protein 7 (LOMP) (F-box only protein 20). | |||||
|
MYH11_MOUSE
|
||||||
| θ value | 0.163984 (rank : 31) | NC score | 0.065581 (rank : 45) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1777 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | O08638, O08639, Q62462, Q64195 | Gene names | Myh11 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
PRP40_HUMAN
|
||||||
| θ value | 0.21417 (rank : 32) | NC score | 0.085033 (rank : 28) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 442 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O75400, O43856, O75404, Q8TBQ1, Q9H782, Q9NWU9, Q9P0Q2, Q9Y5A8 | Gene names | PRPF40A, FLAF1, FNBP3, HYPA | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Huntingtin yeast partner A) (Huntingtin-interacting protein HYPA/FBP11) (Fas ligand-associated factor 1) (NY-REN-6 antigen). | |||||
|
PRP40_MOUSE
|
||||||
| θ value | 0.21417 (rank : 33) | NC score | 0.089894 (rank : 27) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9R1C7, Q61049, Q8BQ76, Q8BRW4, Q8C2U1 | Gene names | Prpf40a, Fbp11, Fnbp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Formin-binding protein 11) (FBP-11). | |||||
|
VAV2_HUMAN
|
||||||
| θ value | 0.21417 (rank : 34) | NC score | 0.054355 (rank : 83) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 453 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P52735 | Gene names | VAV2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-2. | |||||
|
VAV2_MOUSE
|
||||||
| θ value | 0.21417 (rank : 35) | NC score | 0.053977 (rank : 84) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 456 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q60992 | Gene names | Vav2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-2. | |||||
|
CP250_HUMAN
|
||||||
| θ value | 0.279714 (rank : 36) | NC score | 0.047715 (rank : 94) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1805 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9BV73, O14812, O60588, Q9H450 | Gene names | CEP250, CEP2, CNAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Centrosome-associated protein CEP250 (Centrosomal protein 2) (Centrosomal Nek2-associated protein 1) (C-Nap1). | |||||
|
TCRG1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 37) | NC score | 0.057329 (rank : 74) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 929 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O14776 | Gene names | TCERG1, CA150, TAF2S | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150). | |||||
|
TXLNB_MOUSE
|
||||||
| θ value | 0.279714 (rank : 38) | NC score | 0.044680 (rank : 97) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 640 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q8VBT1, Q3UVB8, Q8BUK2 | Gene names | Txlnb, Mdp77 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-taxilin (Muscle-derived protein 77). | |||||
|
SYGP1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 39) | NC score | 0.197607 (rank : 14) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q96PV0, Q8TCS2, Q9UGE2 | Gene names | SYNGAP1, KIAA1938 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein SynGAP (Synaptic Ras-GTPase-activating protein 1) (Synaptic Ras-GAP 1) (Neuronal RasGAP). | |||||
|
TCRG1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 40) | NC score | 0.056715 (rank : 78) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8CGF7, Q61051, Q8C490, Q8CHT8, Q9R0R5 | Gene names | Tcerg1, Taf2s | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150) (p144) (Formin- binding protein 28) (FBP 28). | |||||
|
CROCC_MOUSE
|
||||||
| θ value | 0.47712 (rank : 41) | NC score | 0.047737 (rank : 93) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1460 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q8CJ40, Q7TQL2, Q80U01, Q8R0B9 | Gene names | Crocc, Kiaa0445 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rootletin (Ciliary rootlet coiled-coil protein). | |||||
|
MCF2L_MOUSE
|
||||||
| θ value | 0.47712 (rank : 42) | NC score | 0.043461 (rank : 100) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q64096 | Gene names | Mcf2l, Dbs | |||
|
Domain Architecture |

|
|||||
| Description | Guanine nucleotide exchange factor DBS (DBL's big sister) (MCF2- transforming sequence-like protein). | |||||
|
WBP4_HUMAN
|
||||||
| θ value | 0.47712 (rank : 43) | NC score | 0.150241 (rank : 17) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O75554 | Gene names | WBP4, FBP21, FNBP21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 4 (WBP-4) (Formin-binding protein 21). | |||||
|
AB1IP_MOUSE
|
||||||
| θ value | 0.62314 (rank : 44) | NC score | 0.024087 (rank : 130) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8R5A3, O35329, Q8BRU0, Q99KV8 | Gene names | Apbb1ip, Prel1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 1- interacting protein (APBB1-interacting protein 1) (Proline-rich EVH1 ligand 1) (PREL-1) (Proline-rich protein 48). | |||||
|
IFT74_HUMAN
|
||||||
| θ value | 0.813845 (rank : 45) | NC score | 0.049677 (rank : 90) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 949 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q96LB3, Q6PGQ8, Q9H643, Q9H8G7 | Gene names | IFT74, CCDC2, CMG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intraflagellar transport 74 homolog (Coiled-coil domain-containing protein 2) (Capillary morphogenesis protein 1) (CMG-1). | |||||
|
LRCH4_MOUSE
|
||||||
| θ value | 0.813845 (rank : 46) | NC score | 0.018871 (rank : 136) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q921G6 | Gene names | Lrch4 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 4. | |||||
|
MYO9B_MOUSE
|
||||||
| θ value | 0.813845 (rank : 47) | NC score | 0.073255 (rank : 35) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9QY06, Q9QY07, Q9QY08, Q9QY09 | Gene names | Myo9b, Myr5 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9B (Myosin IXb) (Unconventional myosin-9b). | |||||
|
RGNEF_MOUSE
|
||||||
| θ value | 0.813845 (rank : 48) | NC score | 0.031127 (rank : 125) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 641 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P97433 | Gene names | Rgnef, Rhoip2 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-guanine nucleotide exchange factor (Rho-interacting protein 2) (RhoGEF) (RIP2). | |||||
|
TPR_HUMAN
|
||||||
| θ value | 0.813845 (rank : 49) | NC score | 0.042514 (rank : 103) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
VAV_MOUSE
|
||||||
| θ value | 0.813845 (rank : 50) | NC score | 0.059606 (rank : 63) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 430 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P27870, Q8BTV7 | Gene names | Vav1, Vav | |||
|
Domain Architecture |

|
|||||
| Description | Proto-oncogene vav (p95vav). | |||||
|
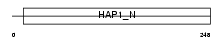
BICD2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 51) | NC score | 0.039784 (rank : 109) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 856 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q8TD16, O75181, Q5TBQ2, Q5TBQ3, Q96LH2, Q9BT84, Q9H561 | Gene names | BICD2, KIAA0699 | |||
|
Domain Architecture |
|
|||||
| Description | Cytoskeleton-like bicaudal D protein homolog 2. | |||||
|
MYH2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 52) | NC score | 0.065643 (rank : 43) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1491 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9UKX2, Q14322, Q16229 | Gene names | MYH2, MYHSA2 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, skeletal muscle, adult 2 (Myosin heavy chain IIa) (MyHC-IIa). | |||||
|
VAV_HUMAN
|
||||||
| θ value | 1.06291 (rank : 53) | NC score | 0.057716 (rank : 70) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P15498, Q15860 | Gene names | VAV1, VAV | |||
|
Domain Architecture |

|
|||||
| Description | Proto-oncogene vav. | |||||
|
WBP4_MOUSE
|
||||||
| θ value | 1.06291 (rank : 54) | NC score | 0.123995 (rank : 21) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q61048, Q8K1Z9, Q9CS45 | Gene names | Wbp4, Fbp21, Fnbp21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 4 (WBP-4) (Formin-binding protein 21). | |||||
|
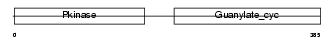
ANPRA_HUMAN
|
||||||
| θ value | 1.38821 (rank : 55) | NC score | 0.006329 (rank : 153) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 612 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P16066, Q6P4Q3 | Gene names | NPR1, ANPRA | |||
|
Domain Architecture |
|
|||||
| Description | Atrial natriuretic peptide receptor A precursor (ANP-A) (ANPRA) (GC-A) (Guanylate cyclase) (EC 4.6.1.2) (NPR-A) (Atrial natriuretic peptide A-type receptor). | |||||
|
VAV3_MOUSE
|
||||||
| θ value | 1.38821 (rank : 56) | NC score | 0.042498 (rank : 104) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 404 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9R0C8, Q7TS85 | Gene names | Vav3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-3. | |||||
|
BAZ2B_HUMAN
|
||||||
| θ value | 1.81305 (rank : 57) | NC score | 0.024107 (rank : 129) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 803 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9UIF8, Q96EA1, Q96SQ8, Q9P252, Q9Y4N8 | Gene names | BAZ2B, KIAA1476 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2B (hWALp4). | |||||
|
CCD1A_HUMAN
|
||||||
| θ value | 1.81305 (rank : 58) | NC score | 0.030434 (rank : 126) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q6P1N0, Q7Z435, Q86XV0, Q8NF89, Q9H603, Q9NXI1 | Gene names | CC2D1A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil and C2 domain-containing protein 1A (Five repressor element under dual repression-binding protein 1) (FRE under dual repression-binding protein 1) (Freud-1) (Putative NF-kappa-B- activating protein 023N). | |||||
|
FOXP4_HUMAN
|
||||||
| θ value | 1.81305 (rank : 59) | NC score | 0.013040 (rank : 143) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8IVH2, Q96E19 | Gene names | FOXP4, FKHLA | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P4 (Fork head-related protein-like A). | |||||
|
FOXP4_MOUSE
|
||||||
| θ value | 1.81305 (rank : 60) | NC score | 0.013418 (rank : 142) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9DBY0, Q80V92, Q8CG10, Q8CIS1 | Gene names | Foxp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein P4 (Fork head-related protein like A) (mFKHLA). | |||||
|
MYH8_MOUSE
|
||||||
| θ value | 1.81305 (rank : 61) | NC score | 0.060165 (rank : 61) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1600 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | P13542, Q5SX36 | Gene names | Myh8, Myhsp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-8 (Myosin heavy chain, skeletal muscle, perinatal) (MyHC- perinatal). | |||||
|
MYO9B_HUMAN
|
||||||
| θ value | 1.81305 (rank : 62) | NC score | 0.075083 (rank : 31) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 641 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q13459, O75314, Q9NUJ2, Q9UHN0 | Gene names | MYO9B, MYR5 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9B (Myosin IXb) (Unconventional myosin-9b). | |||||
|
PERT_MOUSE
|
||||||
| θ value | 1.81305 (rank : 63) | NC score | 0.011297 (rank : 145) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P35419, Q8C8B1 | Gene names | Tpo | |||
|
Domain Architecture |
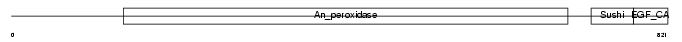
|
|||||
| Description | Thyroid peroxidase precursor (EC 1.11.1.8) (TPO). | |||||
|
FA5_HUMAN
|
||||||
| θ value | 2.36792 (rank : 64) | NC score | 0.010678 (rank : 147) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P12259, Q14285, Q6UPU6 | Gene names | F5 | |||
|
Domain Architecture |

|
|||||
| Description | Coagulation factor V precursor (Activated protein C cofactor) [Contains: Coagulation factor V heavy chain; Coagulation factor V light chain]. | |||||
|
HOOK3_HUMAN
|
||||||
| θ value | 2.36792 (rank : 65) | NC score | 0.038047 (rank : 116) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1070 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q86VS8, Q8NBH0, Q9BY13 | Gene names | HOOK3 | |||
|
Domain Architecture |

|
|||||
| Description | Hook homolog 3 (hHK3). | |||||
|
SESN1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 66) | NC score | 0.013852 (rank : 140) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P58006, Q7TNF3 | Gene names | Sesn1, Pa26, Sest1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sestrin-1 (p53-regulated protein PA26). | |||||
|
ANPRA_MOUSE
|
||||||
| θ value | 3.0926 (rank : 67) | NC score | 0.005359 (rank : 154) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 600 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P18293 | Gene names | Npr1, Npra | |||
|
Domain Architecture |

|
|||||
| Description | Atrial natriuretic peptide receptor A precursor (ANP-A) (ANPRA) (GC-A) (Guanylate cyclase) (EC 4.6.1.2) (NPR-A) (Atrial natriuretic peptide A-type receptor). | |||||
|
CP135_HUMAN
|
||||||
| θ value | 3.0926 (rank : 68) | NC score | 0.040378 (rank : 108) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1290 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q66GS9, O75130, Q9H8H7 | Gene names | CEP135, CEP4, KIAA0635 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 135 kDa (Cep135 protein) (Centrosomal protein 4). | |||||
|
IFIT5_HUMAN
|
||||||
| θ value | 3.0926 (rank : 69) | NC score | 0.012965 (rank : 144) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q13325 | Gene names | IFIT5, RI58 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon-induced protein with tetratricopeptide repeats 5 (IFIT-5) (Retinoic acid- and interferon-inducible 58 kDa protein). | |||||
|
TAGL3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 70) | NC score | 0.058685 (rank : 68) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UI15, Q96A74 | Gene names | TAGLN3, NP25 | |||
|
Domain Architecture |
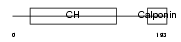
|
|||||
| Description | Transgelin-3 (Neuronal protein NP25) (Neuronal protein 22) (NP22). | |||||
|
APBB2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 71) | NC score | 0.011148 (rank : 146) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9DBR4, Q6DFX8 | Gene names | Apbb2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 2. | |||||
|
ASCC1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 72) | NC score | 0.018888 (rank : 135) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9D8Z1, Q3TAC2 | Gene names | Ascc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Activating signal cointegrator 1 complex subunit 1 (ASC-1 complex subunit p50) (Trip4 complex subunit p50). | |||||
|
CCD1A_MOUSE
|
||||||
| θ value | 4.03905 (rank : 73) | NC score | 0.027862 (rank : 127) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8K1A6, Q2MJB5, Q8R3Z4 | Gene names | Cc2d1a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil and C2 domain-containing protein 1A (Five repressor element under dual repression-binding protein 1) (FRE under dual repression-binding protein 1) (Freud-1). | |||||
|
CNN3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 74) | NC score | 0.079781 (rank : 29) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q15417 | Gene names | CNN3 | |||
|
Domain Architecture |
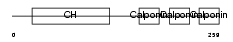
|
|||||
| Description | Calponin-3 (Calponin, acidic isoform). | |||||
|
DHX29_MOUSE
|
||||||
| θ value | 4.03905 (rank : 75) | NC score | 0.007916 (rank : 149) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6PGC1, Q8BQJ4, Q8BT01, Q8C9B7, Q8C9H9 | Gene names | Dhx29 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative ATP-dependent RNA helicase DHX29 (EC 3.6.1.-) (DEAH box protein 29). | |||||
|
ERC2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 76) | NC score | 0.039459 (rank : 112) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1156 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | O15083, Q86TK4 | Gene names | ERC2, KIAA0378 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ERC protein 2. | |||||
|
GOGA4_MOUSE
|
||||||
| θ value | 4.03905 (rank : 77) | NC score | 0.056261 (rank : 79) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1939 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q91VW5, O70365, Q8CGH6 | Gene names | Golga4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (tGolgin-1). | |||||
|
HOOK3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 78) | NC score | 0.036256 (rank : 117) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1007 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q8BUK6, Q8BV48, Q8BW57, Q8BY41 | Gene names | Hook3 | |||
|
Domain Architecture |
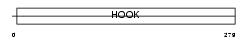
|
|||||
| Description | Hook homolog 3. | |||||
|
PPRB_HUMAN
|
||||||
| θ value | 4.03905 (rank : 79) | NC score | 0.017058 (rank : 137) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q15648, O43810, O75447, Q9HD39 | Gene names | PPARBP, ARC205, DRIP205, TRAP220, TRIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor-binding protein (PBP) (PPAR-binding protein) (Thyroid hormone receptor-associated protein complex 220 kDa component) (Trap220) (Thyroid receptor-interacting protein 2) (TRIP-2) (p53 regulatory protein RB18A) (Vitamin D receptor-interacting protein complex component DRIP205) (Activator- recruited cofactor 205 kDa component) (ARC205). | |||||
|
SPTB2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 80) | NC score | 0.032501 (rank : 123) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 748 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q01082, O60837, Q16057 | Gene names | SPTBN1, SPTB2 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin beta chain, brain 1 (Spectrin, non-erythroid beta chain 1) (Beta-II spectrin) (Fodrin beta chain). | |||||
|
SPTB2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 81) | NC score | 0.032423 (rank : 124) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 770 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q62261, Q5SQL8 | Gene names | Sptbn1, Spnb-2, Spnb2, Sptb2 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin beta chain, brain 1 (Spectrin, non-erythroid beta chain 1) (Beta-II spectrin) (Fodrin beta chain). | |||||
|
TACC3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 82) | NC score | 0.041980 (rank : 105) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 607 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9JJ11, Q9WVK9 | Gene names | Tacc3, Aint | |||
|
Domain Architecture |

|
|||||
| Description | Transforming acidic coiled-coil-containing protein 3 (ARNT-interacting protein). | |||||
|
TAGL3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 83) | NC score | 0.057635 (rank : 71) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9R1Q8 | Gene names | Tagln3, Np25 | |||
|
Domain Architecture |
|
|||||
| Description | Transgelin-3 (Neuronal protein NP25). | |||||
|
TAGL_MOUSE
|
||||||
| θ value | 4.03905 (rank : 84) | NC score | 0.090366 (rank : 26) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P37804, Q545W0 | Gene names | Tagln, Sm22, Sm22a | |||
|
Domain Architecture |
|
|||||
| Description | Transgelin (Smooth muscle protein 22-alpha) (SM22-alpha) (Actin- associated protein p27). | |||||
|
AN30A_HUMAN
|
||||||
| θ value | 5.27518 (rank : 85) | NC score | 0.024488 (rank : 128) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1119 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9BXX3, Q5W025 | Gene names | ANKRD30A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 30A (NY-BR-1 antigen). | |||||
|
BICD2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 86) | NC score | 0.038492 (rank : 115) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 864 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q921C5, Q80TU1, Q8BTE3, Q9DCL3 | Gene names | Bicd2, Kiaa0699 | |||
|
Domain Architecture |
|
|||||
| Description | Cytoskeleton-like bicaudal D protein homolog 2. | |||||
|
CCD41_MOUSE
|
||||||
| θ value | 5.27518 (rank : 87) | NC score | 0.039515 (rank : 111) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q9D5R3, Q3U7X7, Q3UX57, Q80VF0 | Gene names | Ccdc41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 41. | |||||
|
CNN3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 88) | NC score | 0.070492 (rank : 36) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9DAW9, Q3TW23 | Gene names | Cnn3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calponin-3 (Calponin, acidic isoform). | |||||
|
ERC2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 89) | NC score | 0.039610 (rank : 110) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1166 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q6PH08, Q80U20, Q8CCP1 | Gene names | Erc2, Cast1, D14Ertd171e, Kiaa0378 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ERC protein 2 (CAZ-associated structural protein 1) (CAST1). | |||||
|
GOGA5_MOUSE
|
||||||
| θ value | 5.27518 (rank : 90) | NC score | 0.036143 (rank : 118) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1434 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9QYE6, O88317, Q3TGE7, Q3U6S5, Q3UUF9 | Gene names | Golga5, Retii | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 5 (Golgin-84) (Sumiko protein) (Ret-II protein). | |||||
|
WWP2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 91) | NC score | 0.007181 (rank : 150) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O00308, Q96CZ2, Q9BWN6 | Gene names | WWP2 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP2 (EC 6.3.2.-) (WW domain- containing protein 2) (Atrophin-1-interacting protein 2) (AIP2). | |||||
|
AKAP9_HUMAN
|
||||||
| θ value | 6.88961 (rank : 92) | NC score | 0.039160 (rank : 113) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1710 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q99996, O14869, O43355, O94895, Q9UQH3, Q9UQQ4, Q9Y6B8, Q9Y6Y2 | Gene names | AKAP9, AKAP350, AKAP450, KIAA0803 | |||
|
Domain Architecture |

|
|||||
| Description | A-kinase anchor protein 9 (Protein kinase A-anchoring protein 9) (PRKA9) (A-kinase anchor protein 450 kDa) (AKAP 450) (A-kinase anchor protein 350 kDa) (AKAP 350) (hgAKAP 350) (AKAP 120-like protein) (Protein hyperion) (Protein yotiao) (Centrosome- and Golgi-localized PKN-associated protein) (CG-NAP). | |||||
|
DOCK2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 93) | NC score | 0.006490 (rank : 151) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8C3J5, Q99M79 | Gene names | Dock2 | |||
|
Domain Architecture |

|
|||||
| Description | Dedicator of cytokinesis protein 2 (Hch protein). | |||||
|
ES8L1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 94) | NC score | 0.009181 (rank : 148) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8TE68, Q71RE2, Q8NC10, Q96BB7, Q9BSQ2, Q9GZQ2, Q9NXH0 | Gene names | EPS8L1, DRC3, EPS8R1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Epidermal growth factor receptor kinase substrate 8-like protein 1 (Epidermal growth factor receptor pathway substrate 8-related protein 1) (EPS8-like protein 1). | |||||
|
FANCC_MOUSE
|
||||||
| θ value | 6.88961 (rank : 95) | NC score | 0.021481 (rank : 132) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P50652 | Gene names | Fancc, Fac, Facc | |||
|
Domain Architecture |

|
|||||
| Description | Fanconi anemia group C protein homolog (Protein FACC). | |||||
|
GOGB1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 96) | NC score | 0.052305 (rank : 86) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 2297 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q14789, Q14398 | Gene names | GOLGB1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily B member 1 (Giantin) (Macrogolgin) (372 kDa Golgi complex-associated protein) (GCP372). | |||||
|
IQCB1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 97) | NC score | 0.061344 (rank : 58) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q15051, Q5DKQ7, Q8NI79, Q9BS08 | Gene names | IQCB1, KIAA0036, NPHP5 | |||
|
Domain Architecture |

|
|||||
| Description | IQ calmodulin-binding motif-containing protein 1 (Nephrocystin-5). | |||||
|
MRCKB_MOUSE
|
||||||
| θ value | 6.88961 (rank : 98) | NC score | 0.015044 (rank : 139) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 2180 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q7TT50, Q69ZR5, Q80W33 | Gene names | Cdc42bpb, Kiaa1124 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK beta (EC 2.7.11.1) (CDC42-binding protein kinase beta) (Myotonic dystrophy kinase-related CDC42-binding kinase beta) (Myotonic dystrophy protein kinase-like beta) (MRCK beta) (DMPK-like beta). | |||||
|
MYT1L_HUMAN
|
||||||
| θ value | 6.88961 (rank : 99) | NC score | 0.021629 (rank : 131) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9UL68, Q6IQ17, Q9UPP6 | Gene names | MYT1L, KIAA1106 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1-like protein (MyT1L protein) (MyT1-L). | |||||
|
MYT1L_MOUSE
|
||||||
| θ value | 6.88961 (rank : 100) | NC score | 0.021378 (rank : 133) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 461 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P97500, O08996, Q8C643, Q8C7L4, Q8CHB4 | Gene names | Myt1l, Kiaa1106, Nzf1, Png1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1-like protein (MyT1L protein) (Postmitotic neural gene 1 protein) (Zinc finger protein Png-1) (Neural zinc finger factor 1) (NZF-1). | |||||
|
PEPL_MOUSE
|
||||||
| θ value | 6.88961 (rank : 101) | NC score | 0.035815 (rank : 120) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1105 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9R269, O70231, Q9CUT1, Q9JLZ7 | Gene names | Ppl | |||
|
Domain Architecture |

|
|||||
| Description | Periplakin. | |||||
|
RAI14_MOUSE
|
||||||
| θ value | 6.88961 (rank : 102) | NC score | 0.041002 (rank : 107) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1505 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9EP71, Q3URT3, Q6ZPT6 | Gene names | Rai14, Kiaa1334, Norpeg | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankycorbin (Ankyrin repeat and coiled-coil structure-containing protein) (Retinoic acid-induced protein 14) (Novel retinal pigment epithelial cell protein) (p125). | |||||
|
RASA2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 103) | NC score | 0.123587 (rank : 22) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q15283, O00695, Q15284, Q92594, Q99577, Q9UEQ2 | Gene names | RASA2, GAP1M, RASGAP | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 2 (GAP1m). | |||||
|
RBCC1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 104) | NC score | 0.035208 (rank : 121) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1023 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q8TDY2, Q8WVU9, Q92601 | Gene names | RB1CC1, KIAA0203, RBICC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RB1-inducible coiled-coil protein 1. | |||||
|
RT26_HUMAN
|
||||||
| θ value | 6.88961 (rank : 105) | NC score | 0.048674 (rank : 91) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9BYN8, Q96Q58 | Gene names | MRPS26, RPMS13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 28S ribosomal protein S26, mitochondrial precursor (MRP-S26) (MRP- S13). | |||||
|
SNUT1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 106) | NC score | 0.018939 (rank : 134) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9Z315, Q8K155, Q9R1I9, Q9R270 | Gene names | Sart1, Haf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U4/U6.U5 tri-snRNP-associated protein 1 (Squamous cell carcinoma antigen recognized by T-cells 1) (SART-1) (mSART-1) (Hypoxia- associated factor). | |||||
|
SYCP1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 107) | NC score | 0.042873 (rank : 101) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1313 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q15431, O14963 | Gene names | SYCP1, SCP1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptonemal complex protein 1 (SCP-1). | |||||
|
ANXA9_MOUSE
|
||||||
| θ value | 8.99809 (rank : 108) | NC score | 0.004184 (rank : 158) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JHQ0, Q9CQS1 | Gene names | Anxa9 | |||
|
Domain Architecture |
|
|||||
| Description | Annexin A9 (Annexin-31) (Annexin XXXI). | |||||
|
CE290_HUMAN
|
||||||
| θ value | 8.99809 (rank : 109) | NC score | 0.041547 (rank : 106) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1553 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | O15078, Q1PSK5, Q66GS8, Q9H2G6, Q9H6Q7, Q9H8I0 | Gene names | CEP290, KIAA0373, NPHP6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein Cep290 (Nephrocystin-6) (Tumor antigen se2-2). | |||||
|
CNN1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 110) | NC score | 0.046534 (rank : 96) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P51911, O00638, Q15416, Q8IY93, Q99438 | Gene names | CNN1 | |||
|
Domain Architecture |
|
|||||
| Description | Calponin-1 (Calponin H1, smooth muscle) (Basic calponin). | |||||
|
GA2L2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 111) | NC score | 0.051834 (rank : 87) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8NHY3, Q8NHY4 | Gene names | GAS2L2, GAR17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GAS2-like protein 2 (Growth arrest-specific 2-like 2) (GAS2-related protein on chromosome 17) (GAR17 protein). | |||||
|
GA2L2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 112) | NC score | 0.042791 (rank : 102) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q5SSG4 | Gene names | Gas2l2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GAS2-like protein 2 (Growth arrest-specific 2-like 2). | |||||
|
GOGA4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 113) | NC score | 0.065697 (rank : 42) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 2034 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q13439, Q13270, Q13654, Q14436 | Gene names | GOLGA4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (Trans-Golgi p230) (256 kDa golgin) (Golgin-245) (Protein 72.1). | |||||
|
HS105_MOUSE
|
||||||
| θ value | 8.99809 (rank : 114) | NC score | 0.004250 (rank : 157) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q61699, Q3TNS2, Q3UIY8, Q62578, Q62579, Q6A0A5, Q8C430, Q8VCW6 | Gene names | Hsph1, Hsp105, Hsp110, Kiaa0201 | |||
|
Domain Architecture |

|
|||||
| Description | Heat-shock protein 105 kDa (Heat shock-related 100 kDa protein E7I) (HSP-E7I) (Heat shock 110 kDa protein) (42 degrees C-HSP). | |||||
|
KI21B_HUMAN
|
||||||
| θ value | 8.99809 (rank : 115) | NC score | 0.015907 (rank : 138) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1017 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | O75037, Q5T4J3 | Gene names | KIF21B, KIAA0449 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 21B. | |||||
|
MACF1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 116) | NC score | 0.043519 (rank : 99) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1341 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9UPN3, O75053, Q5VW20, Q8WXY2, Q9H540, Q9UKP0, Q9ULG9 | Gene names | MACF1, ABP620, ACF7, KIAA0465, KIAA1251 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1, isoforms 1/2/3/5 (Actin cross-linking family protein 7) (Macrophin-1) (Trabeculin-alpha) (620 kDa actin-binding protein) (ABP620). | |||||
|
MACF4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 117) | NC score | 0.044422 (rank : 98) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1321 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q96PK2, Q8WXY1 | Gene names | MACF1, ABP620, ACF7, KIAA0465, KIAA1251 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1, isoform 4. | |||||
|
NFKB2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 118) | NC score | 0.002520 (rank : 159) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q00653, Q04860, Q9BU75, Q9H471, Q9H472 | Gene names | NFKB2, LYT10 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor NF-kappa-B p100 subunit (DNA-binding factor KBF2) (H2TF1) (Lymphocyte translocation chromosome 10) (Oncogene Lyt-10) (Lyt10) [Contains: Nuclear factor NF-kappa-B p52 subunit]. | |||||
|
PEPL_HUMAN
|
||||||
| θ value | 8.99809 (rank : 119) | NC score | 0.035932 (rank : 119) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1141 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | O60437, O60314, O60454 | Gene names | PPL, KIAA0568 | |||
|
Domain Architecture |
|
|||||
| Description | Periplakin (195 kDa cornified envelope precursor protein) (190 kDa paraneoplastic pemphigus antigen). | |||||
|
PPRB_MOUSE
|
||||||
| θ value | 8.99809 (rank : 120) | NC score | 0.013704 (rank : 141) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q925J9, O88323, Q3UHV0, Q6AXD5, Q8BW37, Q8BX19, Q8VDQ7, Q925K0 | Gene names | Pparbp, Trip2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor-binding protein (PBP) (PPAR-binding protein) (Thyroid hormone receptor-associated protein complex 220 kDa component) (Trap220) (Thyroid receptor-interacting protein 2) (TRIP-2). | |||||
|
PRS8_HUMAN
|
||||||
| θ value | 8.99809 (rank : 121) | NC score | 0.004419 (rank : 155) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P62195, O35051, O43208, P47210, P52915, P52916 | Gene names | PSMC5, SUG1 | |||
|
Domain Architecture |
|
|||||
| Description | 26S protease regulatory subunit 8 (Proteasome 26S subunit ATPase 5) (Proteasome subunit p45) (p45/SUG) (Thyroid hormone receptor- interacting protein 1) (TRIP1). | |||||
|
PRS8_MOUSE
|
||||||
| θ value | 8.99809 (rank : 122) | NC score | 0.004419 (rank : 156) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P62196, O35051, P47210, P52915, P52916, Q3UL51, Q9CWN5 | Gene names | Psmc5, Sug1 | |||
|
Domain Architecture |
|
|||||
| Description | 26S protease regulatory subunit 8 (Proteasome 26S subunit ATPase 5) (Proteasome subunit p45) (p45/SUG) (mSUG1). | |||||
|
S4A11_HUMAN
|
||||||
| θ value | 8.99809 (rank : 123) | NC score | 0.006365 (rank : 152) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NBS3, Q9BXF4, Q9NTW9 | Gene names | SLC4A11, BTR1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium bicarbonate transporter-like protein 11 (Bicarbonate transporter-related protein 1) (Solute carrier family 4 member 11). | |||||
|
SDCG8_HUMAN
|
||||||
| θ value | 8.99809 (rank : 124) | NC score | 0.032729 (rank : 122) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1002 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q86SQ7, O60527, Q3ZCR6, Q8N5F2, Q9P0F1 | Gene names | SDCCAG8, CCCAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serologically defined colon cancer antigen 8 (Centrosomal colon cancer autoantigen protein) (hCCCAP) (Antigen NY-CO-8). | |||||
|
VAV3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 125) | NC score | 0.038779 (rank : 114) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 398 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9UKW4, O95230, Q9Y5X8 | Gene names | VAV3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-3. | |||||
|
CKAP4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 126) | NC score | 0.055191 (rank : 82) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 596 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8BMK4, Q8BTK8, Q8R3F2 | Gene names | Ckap4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-associated protein 4 (63 kDa membrane protein) (p63). | |||||
|
CNN2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 127) | NC score | 0.068031 (rank : 40) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q08093 | Gene names | Cnn2 | |||
|
Domain Architecture |
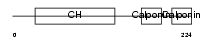
|
|||||
| Description | Calponin-2 (Calponin H2, smooth muscle) (Neutral calponin). | |||||
|
IFT74_MOUSE
|
||||||
| θ value | θ > 10 (rank : 128) | NC score | 0.051079 (rank : 89) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 939 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8BKE9, Q80ZI3, Q9CSY1, Q9CUS0, Q9D9T5 | Gene names | Ift74, Ccdc2, Cmg1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intraflagellar transport 74 homolog (Coiled-coil domain-containing protein 2) (Capillary morphogenesis protein 1) (CMG-1). | |||||
|
IQCB1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 129) | NC score | 0.051200 (rank : 88) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8BP00, Q3TNK4, Q8K306 | Gene names | Iqcb1, Kiaa0036 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ calmodulin-binding motif-containing protein 1. | |||||
|
MAT1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 130) | NC score | 0.060165 (rank : 62) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 179 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P51948, Q15817 | Gene names | MNAT1, CAP35, MAT1, RNF66 | |||
|
Domain Architecture |
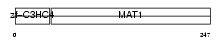
|
|||||
| Description | CDK-activating kinase assembly factor MAT1 (RING finger protein MAT1) (Menage a trois) (CDK7/cyclin H assembly factor) (p36) (p35) (Cyclin G1-interacting protein) (RING finger protein 66). | |||||
|
MAT1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 131) | NC score | 0.065621 (rank : 44) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P51949, Q3TZP0, Q9D8A0, Q9D8D2 | Gene names | Mnat1, Mat1 | |||
|
Domain Architecture |
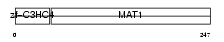
|
|||||
| Description | CDK-activating kinase assembly factor MAT1 (RING finger protein MAT1) (Menage a trois) (CDK7/cyclin H assembly factor) (p36) (p35). | |||||
|
MY18A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 132) | NC score | 0.061630 (rank : 55) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1929 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q92614, Q8IXP8 | Gene names | MYO18A, KIAA0216, MYSPDZ | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
MY18A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 133) | NC score | 0.060238 (rank : 60) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1989 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9JMH9, Q811D7 | Gene names | Myo18a, Myspdz | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
MYH10_HUMAN
|
||||||
| θ value | θ > 10 (rank : 134) | NC score | 0.061375 (rank : 57) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | P35580, Q16087 | Gene names | MYH10 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
MYH10_MOUSE
|
||||||
| θ value | θ > 10 (rank : 135) | NC score | 0.062403 (rank : 51) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1612 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q61879, Q5SV63 | Gene names | Myh10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
MYH13_HUMAN
|
||||||
| θ value | θ > 10 (rank : 136) | NC score | 0.058667 (rank : 69) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1504 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9UKX3, O95252 | Gene names | MYH13 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-13 (Myosin heavy chain, skeletal muscle, extraocular) (MyHC- eo). | |||||
|
MYH14_MOUSE
|
||||||
| θ value | θ > 10 (rank : 137) | NC score | 0.056014 (rank : 80) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1225 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q6URW6, Q80V64, Q80ZE6 | Gene names | Myh14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-14 (Myosin heavy chain, nonmuscle IIc) (Nonmuscle myosin heavy chain IIc) (NMHC II-C). | |||||
|
MYH1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 138) | NC score | 0.064169 (rank : 47) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1478 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | P12882, Q9Y622 | Gene names | MYH1 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-1 (Myosin heavy chain, skeletal muscle, adult 1) (Myosin heavy chain IIx/d) (MyHC-IIx/d). | |||||
|
MYH1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 139) | NC score | 0.064094 (rank : 48) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1461 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q5SX40 | Gene names | Myh1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-1 (Myosin heavy chain, skeletal muscle, adult 1). | |||||
|
MYH3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 140) | NC score | 0.058740 (rank : 67) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1660 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | P11055, Q15492 | Gene names | MYH3 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, fast skeletal muscle, embryonic (Muscle embryonic myosin heavy chain) (SMHCE). | |||||
|
MYH4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 141) | NC score | 0.062908 (rank : 50) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1377 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9Y623 | Gene names | MYH4 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-4 (Myosin heavy chain, skeletal muscle, fetal) (Myosin heavy chain IIb) (MyHC-IIb). | |||||
|
MYH4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 142) | NC score | 0.061240 (rank : 59) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1476 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q5SX39 | Gene names | Myh4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-4 (Myosin heavy chain, skeletal muscle, fetal). | |||||
|
MYH6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 143) | NC score | 0.059544 (rank : 64) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1426 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P13533, Q13943, Q14906, Q14907 | Gene names | MYH6, MYHCA | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
MYH6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 144) | NC score | 0.059451 (rank : 65) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1501 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q02566, Q64258, Q64738 | Gene names | Myh6, Myhca | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
MYH7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 145) | NC score | 0.056800 (rank : 75) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | P12883, Q14836, Q14837, Q14904, Q16579, Q92679, Q9H1D5, Q9UMM8 | Gene names | MYH7, MYHCB | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle beta isoform (MyHC-beta). | |||||
|
MYH8_HUMAN
|
||||||
| θ value | θ > 10 (rank : 146) | NC score | 0.063116 (rank : 49) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1610 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | P13535, Q14910 | Gene names | MYH8 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-8 (Myosin heavy chain, skeletal muscle, perinatal) (MyHC- perinatal). | |||||
|
MYH9_HUMAN
|
||||||
| θ value | θ > 10 (rank : 147) | NC score | 0.061829 (rank : 53) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | P35579, O60805 | Gene names | MYH9 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
MYH9_MOUSE
|
||||||
| θ value | θ > 10 (rank : 148) | NC score | 0.061459 (rank : 56) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1829 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q8VDD5 | Gene names | Myh9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
MYO10_HUMAN
|
||||||
| θ value | θ > 10 (rank : 149) | NC score | 0.055853 (rank : 81) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 920 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9HD67, Q9NYM7, Q9P110, Q9P111, Q9UHF6 | Gene names | MYO10 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-10 (Myosin X). | |||||
|
MYO15_HUMAN
|
||||||
| θ value | θ > 10 (rank : 150) | NC score | 0.058876 (rank : 66) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UKN7 | Gene names | MYO15A, MYO15 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-15 (Myosin XV) (Unconventional myosin-15). | |||||
|
MYO1A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 151) | NC score | 0.057424 (rank : 73) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UBC5, Q9UQD7 | Gene names | MYO1A, MYHL | |||
|
Domain Architecture |

|
|||||
| Description | Myosin Ia (Brush border myosin I) (BBM-I) (BBMI) (Myosin I heavy chain) (MIHC). | |||||
|
MYO1C_HUMAN
|
||||||
| θ value | θ > 10 (rank : 152) | NC score | 0.056765 (rank : 76) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O00159 | Gene names | MYO1C | |||
|
Domain Architecture |

|
|||||
| Description | Myosin Ic (Myosin I beta) (MMI-beta) (MMIb). | |||||
|
MYO1C_MOUSE
|
||||||
| θ value | θ > 10 (rank : 153) | NC score | 0.057477 (rank : 72) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9WTI7, O08571, O08834, Q9QW54 | Gene names | Myo1c | |||
|
Domain Architecture |

|
|||||
| Description | Myosin Ic (Myosin I beta) (MMIb). | |||||
|
MYO5B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 154) | NC score | 0.070299 (rank : 37) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 911 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9ULV0 | Gene names | MYO5B, KIAA1119 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5B (Myosin Vb). | |||||
|
NUF2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 155) | NC score | 0.052570 (rank : 85) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 661 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9BZD4, Q8WU69, Q96HJ4, Q96Q78 | Gene names | CDCA1, NUF2, NUF2R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinetochore protein Nuf2 (hNuf2) (hNuf2R) (hsNuf2) (Cell division cycle-associated protein 1). | |||||
|
PGEA1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 156) | NC score | 0.062395 (rank : 52) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y3M2, Q9UIK9 | Gene names | PGEA1, C22orf2, CBY | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Chibby (PKD2 interactor, Golgi and endoplasmic reticulum associated 1) (PIGEA-14) (Cytosolic leucine-rich protein). | |||||
|
PHLB3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 157) | NC score | 0.061794 (rank : 54) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 304 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q6NSJ2, Q8N7Z4 | Gene names | PHLDB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family B member 3. | |||||
|
RASA2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 158) | NC score | 0.109300 (rank : 24) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P58069 | Gene names | Rasa2 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 2 (GAP1m). | |||||
|
TAGL_HUMAN
|
||||||
| θ value | θ > 10 (rank : 159) | NC score | 0.074754 (rank : 32) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q01995, O15542 | Gene names | TAGLN, SM22, WS3-10 | |||
|
Domain Architecture |
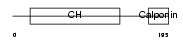
|
|||||
| Description | Transgelin (Smooth muscle protein 22-alpha) (SM22-alpha) (WS3-10) (22 kDa actin-binding protein). | |||||
|
IQGA3_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 125 | |
| SwissProt Accessions | Q86VI3 | Gene names | IQGAP3 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP3. | |||||
|
IQGA1_MOUSE
|
||||||
| NC score | 0.945776 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q9JKF1 | Gene names | Iqgap1 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP1. | |||||
|
IQGA2_HUMAN
|
||||||
| NC score | 0.940706 (rank : 3) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q13576 | Gene names | IQGAP2 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP2. | |||||
|
IQGA1_HUMAN
|
||||||
| NC score | 0.936760 (rank : 4) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | P46940 | Gene names | IQGAP1, KIAA0051 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP1 (p195). | |||||
|
NF1_MOUSE
|
||||||
| NC score | 0.343159 (rank : 5) | θ value | 1.38499e-08 (rank : 6) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q04690, Q61956, Q61957 | Gene names | Nf1 | |||
|
Domain Architecture |

|
|||||
| Description | Neurofibromin (Neurofibromatosis-related protein NF-1). | |||||
|
NF1_HUMAN
|
||||||
| NC score | 0.342416 (rank : 6) | θ value | 1.06045e-08 (rank : 5) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P21359, O00662, Q14284, Q14930, Q9UMK3 | Gene names | NF1 | |||
|
Domain Architecture |

|
|||||
| Description | Neurofibromin (Neurofibromatosis-related protein NF-1) [Contains: Neurofibromin truncated]. | |||||
|
NGAP_HUMAN
|
||||||
| NC score | 0.222415 (rank : 7) | θ value | 0.0113563 (rank : 16) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 517 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9UJF2, O95174, Q5TFU9 | Gene names | RASAL2, NGAP | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein nGAP (RAS protein activator-like 1). | |||||
|
ASPM_HUMAN
|
||||||
| NC score | 0.221602 (rank : 8) | θ value | 2.61198e-07 (rank : 7) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8IZT6, Q86UX4, Q8IUL2, Q8IZJ7, Q8IZJ8, Q8IZJ9, Q8N4D1, Q9NVS1, Q9NVT6 | Gene names | ASPM, MCPH5 | |||
|
Domain Architecture |

|
|||||
| Description | Abnormal spindle-like microcephaly-associated protein (Abnormal spindle protein homolog) (Asp homolog). | |||||
|
DAB2P_MOUSE
|
||||||
| NC score | 0.220180 (rank : 9) | θ value | 0.00869519 (rank : 15) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 284 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q3UHC7, Q3TPD5, Q3UH44, Q6JTV1, Q80T97 | Gene names | Dab2ip, Kiaa1743 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Disabled homolog 2-interacting protein (DAB2-interacting protein). | |||||
|
DAB2P_HUMAN
|
||||||
| NC score | 0.217859 (rank : 10) | θ value | 0.0193708 (rank : 18) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q5VWQ8, Q8TDL2, Q96SE1, Q9C0C0 | Gene names | DAB2IP, AF9Q34, AIP1, KIAA1743 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Disabled homolog 2-interacting protein (DAB2-interacting protein) (DAB2 interaction protein) (ASK-interacting protein 1). | |||||
|
RASL1_MOUSE
|
||||||
| NC score | 0.216863 (rank : 11) | θ value | 0.00020696 (rank : 11) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Z268 | Gene names | Rasal1, Rasal | |||
|
Domain Architecture |

|
|||||
| Description | RasGAP-activating-like protein 1. | |||||
|
RASL1_HUMAN
|
||||||
| NC score | 0.215207 (rank : 12) | θ value | 9.29e-05 (rank : 9) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O95294, Q52M03, Q96CC7 | Gene names | RASAL1, RASAL | |||
|
Domain Architecture |

|
|||||
| Description | RasGAP-activating-like protein 1. | |||||
|
RASA1_HUMAN
|
||||||
| NC score | 0.211493 (rank : 13) | θ value | 0.000270298 (rank : 12) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P20936 | Gene names | RASA1, RASA | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 1 (GTPase-activating protein) (GAP) (Ras p21 protein activator) (p120GAP) (RasGAP). | |||||
|
SYGP1_HUMAN
|
||||||
| NC score | 0.197607 (rank : 14) | θ value | 0.365318 (rank : 39) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q96PV0, Q8TCS2, Q9UGE2 | Gene names | SYNGAP1, KIAA1938 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein SynGAP (Synaptic Ras-GTPase-activating protein 1) (Synaptic Ras-GAP 1) (Neuronal RasGAP). | |||||
|
ASPM_MOUSE
|
||||||
| NC score | 0.195385 (rank : 15) | θ value | 5.44631e-05 (rank : 8) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8CJ27, O88482, Q8BJI8, Q8BKT4 | Gene names | Aspm, Calmbp1, Sha1 | |||
|
Domain Architecture |

|
|||||
| Description | Abnormal spindle-like microcephaly-associated protein homolog (Calmodulin-binding protein 1) (Spindle and hydroxyurea checkpoint abnormal protein) (Calmodulin-binding protein Sha1). | |||||
|
RASA3_HUMAN
|
||||||
| NC score | 0.166122 (rank : 16) | θ value | 0.00298849 (rank : 14) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q14644 | Gene names | RASA3 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 3 (GAP1(IP4BP)) (Ins P4-binding protein). | |||||
|
WBP4_HUMAN
|
||||||
| NC score | 0.150241 (rank : 17) | θ value | 0.47712 (rank : 43) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O75554 | Gene names | WBP4, FBP21, FNBP21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 4 (WBP-4) (Formin-binding protein 21). | |||||
|
RASL2_HUMAN
|
||||||
| NC score | 0.146038 (rank : 18) | θ value | 0.0736092 (rank : 26) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O43374, O60286, Q86UW3, Q96QU0 | Gene names | RASA4, CAPRI, GAPL, KIAA0538 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 4 (RasGAP-activating-like protein 2) (Calcium-promoted Ras inactivator). | |||||
|
TAGL2_MOUSE
|
||||||
| NC score | 0.130509 (rank : 19) | θ value | 0.00134147 (rank : 13) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9WVA4 | Gene names | Tagln2 | |||
|
Domain Architecture |

|
|||||
| Description | Transgelin-2. | |||||
|
RASA3_MOUSE
|
||||||
| NC score | 0.129556 (rank : 20) | θ value | 0.125558 (rank : 28) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q60790 | Gene names | Rasa3 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 3 (GAP1(IP4BP)) (Ins P4-binding protein) (GapIII). | |||||
|
WBP4_MOUSE
|
||||||
| NC score | 0.123995 (rank : 21) | θ value | 1.06291 (rank : 54) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q61048, Q8K1Z9, Q9CS45 | Gene names | Wbp4, Fbp21, Fnbp21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 4 (WBP-4) (Formin-binding protein 21). | |||||
|
RASA2_HUMAN
|
||||||
| NC score | 0.123587 (rank : 22) | θ value | 6.88961 (rank : 103) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q15283, O00695, Q15284, Q92594, Q99577, Q9UEQ2 | Gene names | RASA2, GAP1M, RASGAP | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 2 (GAP1m). | |||||
|
K1683_HUMAN
|
||||||
| NC score | 0.111005 (rank : 23) | θ value | 0.0330416 (rank : 21) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9H0B3, Q8N4G8, Q96M14, Q9C0I0 | Gene names | KIAA1683 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1683. | |||||
|
RASA2_MOUSE
|
||||||
| NC score | 0.109300 (rank : 24) | θ value | θ > 10 (rank : 158) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P58069 | Gene names | Rasa2 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 2 (GAP1m). | |||||
|
TAGL2_HUMAN
|
||||||
| NC score | 0.098045 (rank : 25) | θ value | 0.0193708 (rank : 19) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P37802, Q5JRQ6, Q5JRQ7, Q9BUH5, Q9H4P0 | Gene names | TAGLN2, KIAA0120 | |||
|
Domain Architecture |

|
|||||
| Description | Transgelin-2 (SM22-alpha homolog). | |||||
|
TAGL_MOUSE
|
||||||
| NC score | 0.090366 (rank : 26) | θ value | 4.03905 (rank : 84) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P37804, Q545W0 | Gene names | Tagln, Sm22, Sm22a | |||
|
Domain Architecture |

|
|||||
| Description | Transgelin (Smooth muscle protein 22-alpha) (SM22-alpha) (Actin- associated protein p27). | |||||
|
PRP40_MOUSE
|
||||||
| NC score | 0.089894 (rank : 27) | θ value | 0.21417 (rank : 33) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9R1C7, Q61049, Q8BQ76, Q8BRW4, Q8C2U1 | Gene names | Prpf40a, Fbp11, Fnbp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Formin-binding protein 11) (FBP-11). | |||||
|
PRP40_HUMAN
|
||||||
| NC score | 0.085033 (rank : 28) | θ value | 0.21417 (rank : 32) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 442 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O75400, O43856, O75404, Q8TBQ1, Q9H782, Q9NWU9, Q9P0Q2, Q9Y5A8 | Gene names | PRPF40A, FLAF1, FNBP3, HYPA | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Huntingtin yeast partner A) (Huntingtin-interacting protein HYPA/FBP11) (Fas ligand-associated factor 1) (NY-REN-6 antigen). | |||||
|
CNN3_HUMAN
|
||||||
| NC score | 0.079781 (rank : 29) | θ value | 4.03905 (rank : 74) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q15417 | Gene names | CNN3 | |||
|
Domain Architecture |

|
|||||
| Description | Calponin-3 (Calponin, acidic isoform). | |||||
|
MYO5A_MOUSE
|
||||||
| NC score | 0.075925 (rank : 30) | θ value | 0.0563607 (rank : 25) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1117 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q99104 | Gene names | Myo5a, Dilute | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5A (Myosin Va) (Dilute myosin heavy chain, non-muscle). | |||||
|
MYO9B_HUMAN
|
||||||
| NC score | 0.075083 (rank : 31) | θ value | 1.81305 (rank : 62) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 641 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q13459, O75314, Q9NUJ2, Q9UHN0 | Gene names | MYO9B, MYR5 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9B (Myosin IXb) (Unconventional myosin-9b). | |||||
|
TAGL_HUMAN
|
||||||
| NC score | 0.074754 (rank : 32) | θ value | θ > 10 (rank : 159) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q01995, O15542 | Gene names | TAGLN, SM22, WS3-10 | |||
|
Domain Architecture |

|
|||||
| Description | Transgelin (Smooth muscle protein 22-alpha) (SM22-alpha) (WS3-10) (22 kDa actin-binding protein). | |||||
|
MYO5A_HUMAN
|
||||||
| NC score | 0.074297 (rank : 33) | θ value | 0.0148317 (rank : 17) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1092 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9Y4I1, O60653, Q07902, Q16249, Q9UE30, Q9UE31 | Gene names | MYO5A, MYH12 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5A (Myosin Va) (Dilute myosin heavy chain, non-muscle) (Myosin heavy chain 12) (Myoxin). | |||||
|
MYO5C_HUMAN
|
||||||
| NC score | 0.074018 (rank : 34) | θ value | 0.00020696 (rank : 10) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1206 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9NQX4 | Gene names | MYO5C | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5C (Myosin Vc). | |||||
|
MYO9B_MOUSE
|
||||||
| NC score | 0.073255 (rank : 35) | θ value | 0.813845 (rank : 47) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9QY06, Q9QY07, Q9QY08, Q9QY09 | Gene names | Myo9b, Myr5 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9B (Myosin IXb) (Unconventional myosin-9b). | |||||
|
CNN3_MOUSE
|
||||||
| NC score | 0.070492 (rank : 36) | θ value | 5.27518 (rank : 88) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9DAW9, Q3TW23 | Gene names | Cnn3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calponin-3 (Calponin, acidic isoform). | |||||
|
MYO5B_HUMAN
|
||||||
| NC score | 0.070299 (rank : 37) | θ value | θ > 10 (rank : 154) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 911 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9ULV0 | Gene names | MYO5B, KIAA1119 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5B (Myosin Vb). | |||||
|
MYO7A_HUMAN
|
||||||
| NC score | 0.070000 (rank : 38) | θ value | 0.125558 (rank : 27) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q13402, P78427, Q13321, Q14785, Q92821, Q92822 | Gene names | MYO7A, USH1B | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-7A (Myosin VIIa). | |||||
|
MYO7A_MOUSE
|
||||||
| NC score | 0.069653 (rank : 39) | θ value | 0.0431538 (rank : 23) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P97479 | Gene names | Myo7a, Myo7 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-7A (Myosin VIIa). | |||||
|
CNN2_MOUSE
|
||||||
| NC score | 0.068031 (rank : 40) | θ value | θ > 10 (rank : 127) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q08093 | Gene names | Cnn2 | |||
|
Domain Architecture |

|
|||||
| Description | Calponin-2 (Calponin H2, smooth muscle) (Neutral calponin). | |||||
|
LMO7_HUMAN
|
||||||
| NC score | 0.067070 (rank : 41) | θ value | 0.163984 (rank : 30) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8WWI1, O15462, O95346, Q9UKC1, Q9UQM5, Q9Y6A7 | Gene names | LMO7, FBX20, FBXO20, KIAA0858 | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain only protein 7 (LOMP) (F-box only protein 20). | |||||
|
GOGA4_HUMAN
|
||||||
| NC score | 0.065697 (rank : 42) | θ value | 8.99809 (rank : 113) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 2034 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q13439, Q13270, Q13654, Q14436 | Gene names | GOLGA4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (Trans-Golgi p230) (256 kDa golgin) (Golgin-245) (Protein 72.1). | |||||
|
MYH2_HUMAN
|
||||||
| NC score | 0.065643 (rank : 43) | θ value | 1.06291 (rank : 52) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1491 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9UKX2, Q14322, Q16229 | Gene names | MYH2, MYHSA2 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, skeletal muscle, adult 2 (Myosin heavy chain IIa) (MyHC-IIa). | |||||
|
MAT1_MOUSE
|
||||||
| NC score | 0.065621 (rank : 44) | θ value | θ > 10 (rank : 131) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P51949, Q3TZP0, Q9D8A0, Q9D8D2 | Gene names | Mnat1, Mat1 | |||
|
Domain Architecture |

|
|||||
| Description | CDK-activating kinase assembly factor MAT1 (RING finger protein MAT1) (Menage a trois) (CDK7/cyclin H assembly factor) (p36) (p35). | |||||
|
MYH11_MOUSE
|
||||||
| NC score | 0.065581 (rank : 45) | θ value | 0.163984 (rank : 31) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1777 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | O08638, O08639, Q62462, Q64195 | Gene names | Myh11 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
MYH11_HUMAN
|
||||||
| NC score | 0.064292 (rank : 46) | θ value | 0.0563607 (rank : 24) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1806 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | P35749, O00396, O94944, P78422 | Gene names | MYH11, KIAA0866 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
MYH1_HUMAN
|
||||||
| NC score | 0.064169 (rank : 47) | θ value | θ > 10 (rank : 138) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1478 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | P12882, Q9Y622 | Gene names | MYH1 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-1 (Myosin heavy chain, skeletal muscle, adult 1) (Myosin heavy chain IIx/d) (MyHC-IIx/d). | |||||
|
MYH1_MOUSE
|
||||||
| NC score | 0.064094 (rank : 48) | θ value | θ > 10 (rank : 139) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1461 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q5SX40 | Gene names | Myh1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-1 (Myosin heavy chain, skeletal muscle, adult 1). | |||||
|
MYH8_HUMAN
|
||||||
| NC score | 0.063116 (rank : 49) | θ value | θ > 10 (rank : 146) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1610 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | P13535, Q14910 | Gene names | MYH8 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-8 (Myosin heavy chain, skeletal muscle, perinatal) (MyHC- perinatal). | |||||
|
MYH4_HUMAN
|
||||||
| NC score | 0.062908 (rank : 50) | θ value | θ > 10 (rank : 141) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1377 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9Y623 | Gene names | MYH4 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-4 (Myosin heavy chain, skeletal muscle, fetal) (Myosin heavy chain IIb) (MyHC-IIb). | |||||
|
MYH10_MOUSE
|
||||||
| NC score | 0.062403 (rank : 51) | θ value | θ > 10 (rank : 135) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1612 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q61879, Q5SV63 | Gene names | Myh10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
PGEA1_HUMAN
|
||||||
| NC score | 0.062395 (rank : 52) | θ value | θ > 10 (rank : 156) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y3M2, Q9UIK9 | Gene names | PGEA1, C22orf2, CBY | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Chibby (PKD2 interactor, Golgi and endoplasmic reticulum associated 1) (PIGEA-14) (Cytosolic leucine-rich protein). | |||||
|
MYH9_HUMAN
|
||||||
| NC score | 0.061829 (rank : 53) | θ value | θ > 10 (rank : 147) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | P35579, O60805 | Gene names | MYH9 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
PHLB3_HUMAN
|
||||||
| NC score | 0.061794 (rank : 54) | θ value | θ > 10 (rank : 157) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 304 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q6NSJ2, Q8N7Z4 | Gene names | PHLDB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family B member 3. | |||||
|
MY18A_HUMAN
|
||||||
| NC score | 0.061630 (rank : 55) | θ value | θ > 10 (rank : 132) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1929 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q92614, Q8IXP8 | Gene names | MYO18A, KIAA0216, MYSPDZ | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
MYH9_MOUSE
|
||||||
| NC score | 0.061459 (rank : 56) | θ value | θ > 10 (rank : 148) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1829 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q8VDD5 | Gene names | Myh9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
MYH10_HUMAN
|
||||||
| NC score | 0.061375 (rank : 57) | θ value | θ > 10 (rank : 134) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | P35580, Q16087 | Gene names | MYH10 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-10 (Myosin heavy chain, nonmuscle IIb) (Nonmuscle myosin heavy chain IIb) (NMMHC II-b) (NMMHC-IIB) (Cellular myosin heavy chain, type B) (Nonmuscle myosin heavy chain-B) (NMMHC-B). | |||||
|
IQCB1_HUMAN
|
||||||
| NC score | 0.061344 (rank : 58) | θ value | 6.88961 (rank : 97) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q15051, Q5DKQ7, Q8NI79, Q9BS08 | Gene names | IQCB1, KIAA0036, NPHP5 | |||
|
Domain Architecture |

|
|||||
| Description | IQ calmodulin-binding motif-containing protein 1 (Nephrocystin-5). | |||||
|
MYH4_MOUSE
|
||||||
| NC score | 0.061240 (rank : 59) | θ value | θ > 10 (rank : 142) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1476 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q5SX39 | Gene names | Myh4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-4 (Myosin heavy chain, skeletal muscle, fetal). | |||||
|
MY18A_MOUSE
|
||||||
| NC score | 0.060238 (rank : 60) | θ value | θ > 10 (rank : 133) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1989 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9JMH9, Q811D7 | Gene names | Myo18a, Myspdz | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
MYH8_MOUSE
|
||||||
| NC score | 0.060165 (rank : 61) | θ value | 1.81305 (rank : 61) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1600 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | P13542, Q5SX36 | Gene names | Myh8, Myhsp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-8 (Myosin heavy chain, skeletal muscle, perinatal) (MyHC- perinatal). | |||||
|
MAT1_HUMAN
|
||||||
| NC score | 0.060165 (rank : 62) | θ value | θ > 10 (rank : 130) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 179 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P51948, Q15817 | Gene names | MNAT1, CAP35, MAT1, RNF66 | |||
|
Domain Architecture |

|
|||||
| Description | CDK-activating kinase assembly factor MAT1 (RING finger protein MAT1) (Menage a trois) (CDK7/cyclin H assembly factor) (p36) (p35) (Cyclin G1-interacting protein) (RING finger protein 66). | |||||
|
VAV_MOUSE
|
||||||
| NC score | 0.059606 (rank : 63) | θ value | 0.813845 (rank : 50) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 430 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P27870, Q8BTV7 | Gene names | Vav1, Vav | |||
|
Domain Architecture |

|
|||||
| Description | Proto-oncogene vav (p95vav). | |||||
|
MYH6_HUMAN
|
||||||
| NC score | 0.059544 (rank : 64) | θ value | θ > 10 (rank : 143) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1426 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P13533, Q13943, Q14906, Q14907 | Gene names | MYH6, MYHCA | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
MYH6_MOUSE
|
||||||
| NC score | 0.059451 (rank : 65) | θ value | θ > 10 (rank : 144) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1501 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q02566, Q64258, Q64738 | Gene names | Myh6, Myhca | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
MYO15_HUMAN
|
||||||
| NC score | 0.058876 (rank : 66) | θ value | θ > 10 (rank : 150) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UKN7 | Gene names | MYO15A, MYO15 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-15 (Myosin XV) (Unconventional myosin-15). | |||||
|
MYH3_HUMAN
|
||||||
| NC score | 0.058740 (rank : 67) | θ value | θ > 10 (rank : 140) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1660 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | P11055, Q15492 | Gene names | MYH3 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, fast skeletal muscle, embryonic (Muscle embryonic myosin heavy chain) (SMHCE). | |||||
|
TAGL3_HUMAN
|
||||||
| NC score | 0.058685 (rank : 68) | θ value | 3.0926 (rank : 70) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UI15, Q96A74 | Gene names | TAGLN3, NP25 | |||
|
Domain Architecture |

|
|||||
| Description | Transgelin-3 (Neuronal protein NP25) (Neuronal protein 22) (NP22). | |||||
|
MYH13_HUMAN
|
||||||
| NC score | 0.058667 (rank : 69) | θ value | θ > 10 (rank : 136) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1504 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9UKX3, O95252 | Gene names | MYH13 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-13 (Myosin heavy chain, skeletal muscle, extraocular) (MyHC- eo). | |||||
|
VAV_HUMAN
|
||||||
| NC score | 0.057716 (rank : 70) | θ value | 1.06291 (rank : 53) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P15498, Q15860 | Gene names | VAV1, VAV | |||
|
Domain Architecture |

|
|||||
| Description | Proto-oncogene vav. | |||||
|
TAGL3_MOUSE
|
||||||
| NC score | 0.057635 (rank : 71) | θ value | 4.03905 (rank : 83) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9R1Q8 | Gene names | Tagln3, Np25 | |||
|
Domain Architecture |

|
|||||
| Description | Transgelin-3 (Neuronal protein NP25). | |||||
|
MYO1C_MOUSE
|
||||||
| NC score | 0.057477 (rank : 72) | θ value | θ > 10 (rank : 153) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9WTI7, O08571, O08834, Q9QW54 | Gene names | Myo1c | |||
|
Domain Architecture |

|
|||||
| Description | Myosin Ic (Myosin I beta) (MMIb). | |||||
|
MYO1A_HUMAN
|
||||||
| NC score | 0.057424 (rank : 73) | θ value | θ > 10 (rank : 151) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UBC5, Q9UQD7 | Gene names | MYO1A, MYHL | |||
|
Domain Architecture |

|
|||||
| Description | Myosin Ia (Brush border myosin I) (BBM-I) (BBMI) (Myosin I heavy chain) (MIHC). | |||||
|
TCRG1_HUMAN
|
||||||
| NC score | 0.057329 (rank : 74) | θ value | 0.279714 (rank : 37) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 929 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O14776 | Gene names | TCERG1, CA150, TAF2S | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150). | |||||
|
MYH7_HUMAN
|
||||||
| NC score | 0.056800 (rank : 75) | θ value | θ > 10 (rank : 145) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | P12883, Q14836, Q14837, Q14904, Q16579, Q92679, Q9H1D5, Q9UMM8 | Gene names | MYH7, MYHCB | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle beta isoform (MyHC-beta). | |||||
|
MYO1C_HUMAN
|
||||||
| NC score | 0.056765 (rank : 76) | θ value | θ > 10 (rank : 152) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O00159 | Gene names | MYO1C | |||
|
Domain Architecture |

|
|||||
| Description | Myosin Ic (Myosin I beta) (MMI-beta) (MMIb). | |||||
|
TACC3_HUMAN
|
||||||
| NC score | 0.056749 (rank : 77) | θ value | 0.0252991 (rank : 20) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9Y6A5, Q9UMQ1 | Gene names | TACC3, ERIC1 | |||
|
Domain Architecture |

|
|||||
| Description | Transforming acidic coiled-coil-containing protein 3 (ERIC-1). | |||||
|
TCRG1_MOUSE
|
||||||
| NC score | 0.056715 (rank : 78) | θ value | 0.365318 (rank : 40) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8CGF7, Q61051, Q8C490, Q8CHT8, Q9R0R5 | Gene names | Tcerg1, Taf2s | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150) (p144) (Formin- binding protein 28) (FBP 28). | |||||
|
GOGA4_MOUSE
|
||||||
| NC score | 0.056261 (rank : 79) | θ value | 4.03905 (rank : 77) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1939 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q91VW5, O70365, Q8CGH6 | Gene names | Golga4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (tGolgin-1). | |||||
|
MYH14_MOUSE
|
||||||
| NC score | 0.056014 (rank : 80) | θ value | θ > 10 (rank : 137) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1225 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q6URW6, Q80V64, Q80ZE6 | Gene names | Myh14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-14 (Myosin heavy chain, nonmuscle IIc) (Nonmuscle myosin heavy chain IIc) (NMHC II-C). | |||||
|
MYO10_HUMAN
|
||||||
| NC score | 0.055853 (rank : 81) | θ value | θ > 10 (rank : 149) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 920 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9HD67, Q9NYM7, Q9P110, Q9P111, Q9UHF6 | Gene names | MYO10 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-10 (Myosin X). | |||||
|
CKAP4_MOUSE
|
||||||
| NC score | 0.055191 (rank : 82) | θ value | θ > 10 (rank : 126) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 596 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8BMK4, Q8BTK8, Q8R3F2 | Gene names | Ckap4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-associated protein 4 (63 kDa membrane protein) (p63). | |||||
|
VAV2_HUMAN
|
||||||
| NC score | 0.054355 (rank : 83) | θ value | 0.21417 (rank : 34) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 453 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P52735 | Gene names | VAV2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-2. | |||||
|
VAV2_MOUSE
|
||||||
| NC score | 0.053977 (rank : 84) | θ value | 0.21417 (rank : 35) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 456 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q60992 | Gene names | Vav2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-2. | |||||
|
NUF2_HUMAN
|
||||||
| NC score | 0.052570 (rank : 85) | θ value | θ > 10 (rank : 155) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 661 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9BZD4, Q8WU69, Q96HJ4, Q96Q78 | Gene names | CDCA1, NUF2, NUF2R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinetochore protein Nuf2 (hNuf2) (hNuf2R) (hsNuf2) (Cell division cycle-associated protein 1). | |||||
|
GOGB1_HUMAN
|
||||||
| NC score | 0.052305 (rank : 86) | θ value | 6.88961 (rank : 96) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 2297 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q14789, Q14398 | Gene names | GOLGB1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily B member 1 (Giantin) (Macrogolgin) (372 kDa Golgi complex-associated protein) (GCP372). | |||||
|
GA2L2_HUMAN
|
||||||
| NC score | 0.051834 (rank : 87) | θ value | 8.99809 (rank : 111) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8NHY3, Q8NHY4 | Gene names | GAS2L2, GAR17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GAS2-like protein 2 (Growth arrest-specific 2-like 2) (GAS2-related protein on chromosome 17) (GAR17 protein). | |||||
|
IQCB1_MOUSE
|
||||||
| NC score | 0.051200 (rank : 88) | θ value | θ > 10 (rank : 129) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8BP00, Q3TNK4, Q8K306 | Gene names | Iqcb1, Kiaa0036 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ calmodulin-binding motif-containing protein 1. | |||||
|
IFT74_MOUSE
|
||||||
| NC score | 0.051079 (rank : 89) | θ value | θ > 10 (rank : 128) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 939 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8BKE9, Q80ZI3, Q9CSY1, Q9CUS0, Q9D9T5 | Gene names | Ift74, Ccdc2, Cmg1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intraflagellar transport 74 homolog (Coiled-coil domain-containing protein 2) (Capillary morphogenesis protein 1) (CMG-1). | |||||
|
IFT74_HUMAN
|
||||||
| NC score | 0.049677 (rank : 90) | θ value | 0.813845 (rank : 45) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 949 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q96LB3, Q6PGQ8, Q9H643, Q9H8G7 | Gene names | IFT74, CCDC2, CMG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intraflagellar transport 74 homolog (Coiled-coil domain-containing protein 2) (Capillary morphogenesis protein 1) (CMG-1). | |||||
|
RT26_HUMAN
|
||||||
| NC score | 0.048674 (rank : 91) | θ value | 6.88961 (rank : 105) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9BYN8, Q96Q58 | Gene names | MRPS26, RPMS13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 28S ribosomal protein S26, mitochondrial precursor (MRP-S26) (MRP- S13). | |||||
|
CK5P2_MOUSE
|
||||||
| NC score | 0.048607 (rank : 92) | θ value | 0.0431538 (rank : 22) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1151 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q8K389, Q6PCN1, Q6ZPL0 | Gene names | Cdk5rap2, Kiaa1633 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CDK5 regulatory subunit-associated protein 2 (CDK5 activator-binding protein C48). | |||||
|
CROCC_MOUSE
|
||||||
| NC score | 0.047737 (rank : 93) | θ value | 0.47712 (rank : 41) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1460 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q8CJ40, Q7TQL2, Q80U01, Q8R0B9 | Gene names | Crocc, Kiaa0445 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rootletin (Ciliary rootlet coiled-coil protein). | |||||
|
CP250_HUMAN
|
||||||
| NC score | 0.047715 (rank : 94) | θ value | 0.279714 (rank : 36) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1805 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9BV73, O14812, O60588, Q9H450 | Gene names | CEP250, CEP2, CNAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Centrosome-associated protein CEP250 (Centrosomal protein 2) (Centrosomal Nek2-associated protein 1) (C-Nap1). | |||||
|
CCD41_HUMAN
|
||||||
| NC score | 0.047521 (rank : 95) | θ value | 0.163984 (rank : 29) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1139 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9Y592 | Gene names | CCDC41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 41 (NY-REN-58 antigen). | |||||
|
CNN1_HUMAN
|
||||||
| NC score | 0.046534 (rank : 96) | θ value | 8.99809 (rank : 110) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P51911, O00638, Q15416, Q8IY93, Q99438 | Gene names | CNN1 | |||
|
Domain Architecture |

|
|||||
| Description | Calponin-1 (Calponin H1, smooth muscle) (Basic calponin). | |||||
|
TXLNB_MOUSE
|
||||||
| NC score | 0.044680 (rank : 97) | θ value | 0.279714 (rank : 38) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 640 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q8VBT1, Q3UVB8, Q8BUK2 | Gene names | Txlnb, Mdp77 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-taxilin (Muscle-derived protein 77). | |||||
|
MACF4_HUMAN
|
||||||
| NC score | 0.044422 (rank : 98) | θ value | 8.99809 (rank : 117) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1321 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q96PK2, Q8WXY1 | Gene names | MACF1, ABP620, ACF7, KIAA0465, KIAA1251 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1, isoform 4. | |||||
|
MACF1_HUMAN
|
||||||
| NC score | 0.043519 (rank : 99) | θ value | 8.99809 (rank : 116) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1341 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9UPN3, O75053, Q5VW20, Q8WXY2, Q9H540, Q9UKP0, Q9ULG9 | Gene names | MACF1, ABP620, ACF7, KIAA0465, KIAA1251 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1, isoforms 1/2/3/5 (Actin cross-linking family protein 7) (Macrophin-1) (Trabeculin-alpha) (620 kDa actin-binding protein) (ABP620). | |||||
|
MCF2L_MOUSE
|
||||||
| NC score | 0.043461 (rank : 100) | θ value | 0.47712 (rank : 42) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q64096 | Gene names | Mcf2l, Dbs | |||
|
Domain Architecture |

|
|||||
| Description | Guanine nucleotide exchange factor DBS (DBL's big sister) (MCF2- transforming sequence-like protein). | |||||
|
SYCP1_HUMAN
|
||||||
| NC score | 0.042873 (rank : 101) | θ value | 6.88961 (rank : 107) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1313 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q15431, O14963 | Gene names | SYCP1, SCP1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptonemal complex protein 1 (SCP-1). | |||||
|
GA2L2_MOUSE
|
||||||
| NC score | 0.042791 (rank : 102) | θ value | 8.99809 (rank : 112) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q5SSG4 | Gene names | Gas2l2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GAS2-like protein 2 (Growth arrest-specific 2-like 2). | |||||
|
TPR_HUMAN
|
||||||
| NC score | 0.042514 (rank : 103) | θ value | 0.813845 (rank : 49) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
VAV3_MOUSE
|
||||||
| NC score | 0.042498 (rank : 104) | θ value | 1.38821 (rank : 56) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 404 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9R0C8, Q7TS85 | Gene names | Vav3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-3. | |||||
|
TACC3_MOUSE
|
||||||
| NC score | 0.041980 (rank : 105) | θ value | 4.03905 (rank : 82) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 607 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9JJ11, Q9WVK9 | Gene names | Tacc3, Aint | |||
|
Domain Architecture |

|
|||||
| Description | Transforming acidic coiled-coil-containing protein 3 (ARNT-interacting protein). | |||||
|
CE290_HUMAN
|
||||||
| NC score | 0.041547 (rank : 106) | θ value | 8.99809 (rank : 109) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1553 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | O15078, Q1PSK5, Q66GS8, Q9H2G6, Q9H6Q7, Q9H8I0 | Gene names | CEP290, KIAA0373, NPHP6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein Cep290 (Nephrocystin-6) (Tumor antigen se2-2). | |||||
|
RAI14_MOUSE
|
||||||
| NC score | 0.041002 (rank : 107) | θ value | 6.88961 (rank : 102) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1505 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9EP71, Q3URT3, Q6ZPT6 | Gene names | Rai14, Kiaa1334, Norpeg | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankycorbin (Ankyrin repeat and coiled-coil structure-containing protein) (Retinoic acid-induced protein 14) (Novel retinal pigment epithelial cell protein) (p125). | |||||
|
CP135_HUMAN
|
||||||
| NC score | 0.040378 (rank : 108) | θ value | 3.0926 (rank : 68) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1290 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q66GS9, O75130, Q9H8H7 | Gene names | CEP135, CEP4, KIAA0635 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 135 kDa (Cep135 protein) (Centrosomal protein 4). | |||||
|
BICD2_HUMAN
|
||||||
| NC score | 0.039784 (rank : 109) | θ value | 1.06291 (rank : 51) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 856 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q8TD16, O75181, Q5TBQ2, Q5TBQ3, Q96LH2, Q9BT84, Q9H561 | Gene names | BICD2, KIAA0699 | |||
|
Domain Architecture |

|
|||||
| Description | Cytoskeleton-like bicaudal D protein homolog 2. | |||||
|
ERC2_MOUSE
|
||||||
| NC score | 0.039610 (rank : 110) | θ value | 5.27518 (rank : 89) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1166 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q6PH08, Q80U20, Q8CCP1 | Gene names | Erc2, Cast1, D14Ertd171e, Kiaa0378 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ERC protein 2 (CAZ-associated structural protein 1) (CAST1). | |||||
|
CCD41_MOUSE
|
||||||
| NC score | 0.039515 (rank : 111) | θ value | 5.27518 (rank : 87) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q9D5R3, Q3U7X7, Q3UX57, Q80VF0 | Gene names | Ccdc41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 41. | |||||
|
ERC2_HUMAN
|
||||||
| NC score | 0.039459 (rank : 112) | θ value | 4.03905 (rank : 76) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1156 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | O15083, Q86TK4 | Gene names | ERC2, KIAA0378 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ERC protein 2. | |||||
|
AKAP9_HUMAN
|
||||||
| NC score | 0.039160 (rank : 113) | θ value | 6.88961 (rank : 92) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1710 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q99996, O14869, O43355, O94895, Q9UQH3, Q9UQQ4, Q9Y6B8, Q9Y6Y2 | Gene names | AKAP9, AKAP350, AKAP450, KIAA0803 | |||
|
Domain Architecture |

|
|||||
| Description | A-kinase anchor protein 9 (Protein kinase A-anchoring protein 9) (PRKA9) (A-kinase anchor protein 450 kDa) (AKAP 450) (A-kinase anchor protein 350 kDa) (AKAP 350) (hgAKAP 350) (AKAP 120-like protein) (Protein hyperion) (Protein yotiao) (Centrosome- and Golgi-localized PKN-associated protein) (CG-NAP). | |||||
|
VAV3_HUMAN
|
||||||
| NC score | 0.038779 (rank : 114) | θ value | 8.99809 (rank : 125) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 398 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9UKW4, O95230, Q9Y5X8 | Gene names | VAV3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-3. | |||||
|
BICD2_MOUSE
|
||||||
| NC score | 0.038492 (rank : 115) | θ value | 5.27518 (rank : 86) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 864 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q921C5, Q80TU1, Q8BTE3, Q9DCL3 | Gene names | Bicd2, Kiaa0699 | |||
|
Domain Architecture |

|
|||||
| Description | Cytoskeleton-like bicaudal D protein homolog 2. | |||||
|
HOOK3_HUMAN
|
||||||
| NC score | 0.038047 (rank : 116) | θ value | 2.36792 (rank : 65) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1070 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q86VS8, Q8NBH0, Q9BY13 | Gene names | HOOK3 | |||
|
Domain Architecture |

|
|||||
| Description | Hook homolog 3 (hHK3). | |||||
|
HOOK3_MOUSE
|
||||||
| NC score | 0.036256 (rank : 117) | θ value | 4.03905 (rank : 78) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1007 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q8BUK6, Q8BV48, Q8BW57, Q8BY41 | Gene names | Hook3 | |||
|
Domain Architecture |

|
|||||
| Description | Hook homolog 3. | |||||
|
GOGA5_MOUSE
|
||||||
| NC score | 0.036143 (rank : 118) | θ value | 5.27518 (rank : 90) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1434 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9QYE6, O88317, Q3TGE7, Q3U6S5, Q3UUF9 | Gene names | Golga5, Retii | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 5 (Golgin-84) (Sumiko protein) (Ret-II protein). | |||||
|
PEPL_HUMAN
|
||||||
| NC score | 0.035932 (rank : 119) | θ value | 8.99809 (rank : 119) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1141 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | O60437, O60314, O60454 | Gene names | PPL, KIAA0568 | |||
|
Domain Architecture |

|
|||||
| Description | Periplakin (195 kDa cornified envelope precursor protein) (190 kDa paraneoplastic pemphigus antigen). | |||||
|
PEPL_MOUSE
|
||||||
| NC score | 0.035815 (rank : 120) | θ value | 6.88961 (rank : 101) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1105 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9R269, O70231, Q9CUT1, Q9JLZ7 | Gene names | Ppl | |||
|
Domain Architecture |

|
|||||
| Description | Periplakin. | |||||
|
RBCC1_HUMAN
|
||||||
| NC score | 0.035208 (rank : 121) | θ value | 6.88961 (rank : 104) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1023 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q8TDY2, Q8WVU9, Q92601 | Gene names | RB1CC1, KIAA0203, RBICC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RB1-inducible coiled-coil protein 1. | |||||
|
SDCG8_HUMAN
|
||||||
| NC score | 0.032729 (rank : 122) | θ value | 8.99809 (rank : 124) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1002 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q86SQ7, O60527, Q3ZCR6, Q8N5F2, Q9P0F1 | Gene names | SDCCAG8, CCCAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serologically defined colon cancer antigen 8 (Centrosomal colon cancer autoantigen protein) (hCCCAP) (Antigen NY-CO-8). | |||||
|
SPTB2_HUMAN
|
||||||
| NC score | 0.032501 (rank : 123) | θ value | 4.03905 (rank : 80) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 748 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q01082, O60837, Q16057 | Gene names | SPTBN1, SPTB2 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin beta chain, brain 1 (Spectrin, non-erythroid beta chain 1) (Beta-II spectrin) (Fodrin beta chain). | |||||
|
SPTB2_MOUSE
|
||||||
| NC score | 0.032423 (rank : 124) | θ value | 4.03905 (rank : 81) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 770 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q62261, Q5SQL8 | Gene names | Sptbn1, Spnb-2, Spnb2, Sptb2 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin beta chain, brain 1 (Spectrin, non-erythroid beta chain 1) (Beta-II spectrin) (Fodrin beta chain). | |||||
|
RGNEF_MOUSE
|
||||||
| NC score | 0.031127 (rank : 125) | θ value | 0.813845 (rank : 48) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 641 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P97433 | Gene names | Rgnef, Rhoip2 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-guanine nucleotide exchange factor (Rho-interacting protein 2) (RhoGEF) (RIP2). | |||||
|
CCD1A_HUMAN
|
||||||
| NC score | 0.030434 (rank : 126) | θ value | 1.81305 (rank : 58) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q6P1N0, Q7Z435, Q86XV0, Q8NF89, Q9H603, Q9NXI1 | Gene names | CC2D1A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil and C2 domain-containing protein 1A (Five repressor element under dual repression-binding protein 1) (FRE under dual repression-binding protein 1) (Freud-1) (Putative NF-kappa-B- activating protein 023N). | |||||
|
CCD1A_MOUSE
|
||||||
| NC score | 0.027862 (rank : 127) | θ value | 4.03905 (rank : 73) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8K1A6, Q2MJB5, Q8R3Z4 | Gene names | Cc2d1a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil and C2 domain-containing protein 1A (Five repressor element under dual repression-binding protein 1) (FRE under dual repression-binding protein 1) (Freud-1). | |||||
|
AN30A_HUMAN
|
||||||
| NC score | 0.024488 (rank : 128) | θ value | 5.27518 (rank : 85) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1119 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9BXX3, Q5W025 | Gene names | ANKRD30A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 30A (NY-BR-1 antigen). | |||||
|
BAZ2B_HUMAN
|
||||||
| NC score | 0.024107 (rank : 129) | θ value | 1.81305 (rank : 57) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 803 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9UIF8, Q96EA1, Q96SQ8, Q9P252, Q9Y4N8 | Gene names | BAZ2B, KIAA1476 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2B (hWALp4). | |||||
|
AB1IP_MOUSE
|
||||||
| NC score | 0.024087 (rank : 130) | θ value | 0.62314 (rank : 44) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8R5A3, O35329, Q8BRU0, Q99KV8 | Gene names | Apbb1ip, Prel1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 1- interacting protein (APBB1-interacting protein 1) (Proline-rich EVH1 ligand 1) (PREL-1) (Proline-rich protein 48). | |||||
|
MYT1L_HUMAN
|
||||||
| NC score | 0.021629 (rank : 131) | θ value | 6.88961 (rank : 99) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9UL68, Q6IQ17, Q9UPP6 | Gene names | MYT1L, KIAA1106 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1-like protein (MyT1L protein) (MyT1-L). | |||||
|
FANCC_MOUSE
|
||||||
| NC score | 0.021481 (rank : 132) | θ value | 6.88961 (rank : 95) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P50652 | Gene names | Fancc, Fac, Facc | |||
|
Domain Architecture |

|
|||||
| Description | Fanconi anemia group C protein homolog (Protein FACC). | |||||
|
MYT1L_MOUSE
|
||||||
| NC score | 0.021378 (rank : 133) | θ value | 6.88961 (rank : 100) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 461 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P97500, O08996, Q8C643, Q8C7L4, Q8CHB4 | Gene names | Myt1l, Kiaa1106, Nzf1, Png1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1-like protein (MyT1L protein) (Postmitotic neural gene 1 protein) (Zinc finger protein Png-1) (Neural zinc finger factor 1) (NZF-1). | |||||
|
SNUT1_MOUSE
|
||||||
| NC score | 0.018939 (rank : 134) | θ value | 6.88961 (rank : 106) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9Z315, Q8K155, Q9R1I9, Q9R270 | Gene names | Sart1, Haf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U4/U6.U5 tri-snRNP-associated protein 1 (Squamous cell carcinoma antigen recognized by T-cells 1) (SART-1) (mSART-1) (Hypoxia- associated factor). | |||||
|
ASCC1_MOUSE
|
||||||
| NC score | 0.018888 (rank : 135) | θ value | 4.03905 (rank : 72) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9D8Z1, Q3TAC2 | Gene names | Ascc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Activating signal cointegrator 1 complex subunit 1 (ASC-1 complex subunit p50) (Trip4 complex subunit p50). | |||||
|
LRCH4_MOUSE
|
||||||
| NC score | 0.018871 (rank : 136) | θ value | 0.813845 (rank : 46) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q921G6 | Gene names | Lrch4 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 4. | |||||
|
PPRB_HUMAN
|
||||||
| NC score | 0.017058 (rank : 137) | θ value | 4.03905 (rank : 79) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q15648, O43810, O75447, Q9HD39 | Gene names | PPARBP, ARC205, DRIP205, TRAP220, TRIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor-binding protein (PBP) (PPAR-binding protein) (Thyroid hormone receptor-associated protein complex 220 kDa component) (Trap220) (Thyroid receptor-interacting protein 2) (TRIP-2) (p53 regulatory protein RB18A) (Vitamin D receptor-interacting protein complex component DRIP205) (Activator- recruited cofactor 205 kDa component) (ARC205). | |||||
|
KI21B_HUMAN
|
||||||
| NC score | 0.015907 (rank : 138) | θ value | 8.99809 (rank : 115) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1017 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | O75037, Q5T4J3 | Gene names | KIF21B, KIAA0449 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 21B. | |||||
|
MRCKB_MOUSE
|
||||||
| NC score | 0.015044 (rank : 139) | θ value | 6.88961 (rank : 98) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 2180 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q7TT50, Q69ZR5, Q80W33 | Gene names | Cdc42bpb, Kiaa1124 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK beta (EC 2.7.11.1) (CDC42-binding protein kinase beta) (Myotonic dystrophy kinase-related CDC42-binding kinase beta) (Myotonic dystrophy protein kinase-like beta) (MRCK beta) (DMPK-like beta). | |||||
|
SESN1_MOUSE
|
||||||
| NC score | 0.013852 (rank : 140) | θ value | 2.36792 (rank : 66) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P58006, Q7TNF3 | Gene names | Sesn1, Pa26, Sest1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sestrin-1 (p53-regulated protein PA26). | |||||
|
PPRB_MOUSE
|
||||||
| NC score | 0.013704 (rank : 141) | θ value | 8.99809 (rank : 120) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q925J9, O88323, Q3UHV0, Q6AXD5, Q8BW37, Q8BX19, Q8VDQ7, Q925K0 | Gene names | Pparbp, Trip2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor-binding protein (PBP) (PPAR-binding protein) (Thyroid hormone receptor-associated protein complex 220 kDa component) (Trap220) (Thyroid receptor-interacting protein 2) (TRIP-2). | |||||
|
FOXP4_MOUSE
|
||||||
| NC score | 0.013418 (rank : 142) | θ value | 1.81305 (rank : 60) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9DBY0, Q80V92, Q8CG10, Q8CIS1 | Gene names | Foxp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein P4 (Fork head-related protein like A) (mFKHLA). | |||||
|
FOXP4_HUMAN
|
||||||
| NC score | 0.013040 (rank : 143) | θ value | 1.81305 (rank : 59) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8IVH2, Q96E19 | Gene names | FOXP4, FKHLA | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P4 (Fork head-related protein-like A). | |||||
|
IFIT5_HUMAN
|
||||||
| NC score | 0.012965 (rank : 144) | θ value | 3.0926 (rank : 69) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q13325 | Gene names | IFIT5, RI58 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon-induced protein with tetratricopeptide repeats 5 (IFIT-5) (Retinoic acid- and interferon-inducible 58 kDa protein). | |||||
|
PERT_MOUSE
|
||||||
| NC score | 0.011297 (rank : 145) | θ value | 1.81305 (rank : 63) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P35419, Q8C8B1 | Gene names | Tpo | |||
|
Domain Architecture |
|
|||||
| Description | Thyroid peroxidase precursor (EC 1.11.1.8) (TPO). | |||||
|
APBB2_MOUSE
|
||||||
| NC score | 0.011148 (rank : 146) | θ value | 4.03905 (rank : 71) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9DBR4, Q6DFX8 | Gene names | Apbb2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 2. | |||||
|
FA5_HUMAN
|
||||||
| NC score | 0.010678 (rank : 147) | θ value | 2.36792 (rank : 64) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P12259, Q14285, Q6UPU6 | Gene names | F5 | |||
|
Domain Architecture |

|
|||||
| Description | Coagulation factor V precursor (Activated protein C cofactor) [Contains: Coagulation factor V heavy chain; Coagulation factor V light chain]. | |||||
|
ES8L1_HUMAN
|
||||||
| NC score | 0.009181 (rank : 148) | θ value | 6.88961 (rank : 94) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8TE68, Q71RE2, Q8NC10, Q96BB7, Q9BSQ2, Q9GZQ2, Q9NXH0 | Gene names | EPS8L1, DRC3, EPS8R1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Epidermal growth factor receptor kinase substrate 8-like protein 1 (Epidermal growth factor receptor pathway substrate 8-related protein 1) (EPS8-like protein 1). | |||||
|
DHX29_MOUSE
|
||||||
| NC score | 0.007916 (rank : 149) | θ value | 4.03905 (rank : 75) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6PGC1, Q8BQJ4, Q8BT01, Q8C9B7, Q8C9H9 | Gene names | Dhx29 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative ATP-dependent RNA helicase DHX29 (EC 3.6.1.-) (DEAH box protein 29). | |||||
|
WWP2_HUMAN
|
||||||
| NC score | 0.007181 (rank : 150) | θ value | 5.27518 (rank : 91) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O00308, Q96CZ2, Q9BWN6 | Gene names | WWP2 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP2 (EC 6.3.2.-) (WW domain- containing protein 2) (Atrophin-1-interacting protein 2) (AIP2). | |||||
|
DOCK2_MOUSE
|
||||||
| NC score | 0.006490 (rank : 151) | θ value | 6.88961 (rank : 93) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8C3J5, Q99M79 | Gene names | Dock2 | |||
|
Domain Architecture |

|
|||||
| Description | Dedicator of cytokinesis protein 2 (Hch protein). | |||||
|
S4A11_HUMAN
|
||||||
| NC score | 0.006365 (rank : 152) | θ value | 8.99809 (rank : 123) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NBS3, Q9BXF4, Q9NTW9 | Gene names | SLC4A11, BTR1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium bicarbonate transporter-like protein 11 (Bicarbonate transporter-related protein 1) (Solute carrier family 4 member 11). | |||||
|
ANPRA_HUMAN
|
||||||
| NC score | 0.006329 (rank : 153) | θ value | 1.38821 (rank : 55) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 612 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P16066, Q6P4Q3 | Gene names | NPR1, ANPRA | |||
|
Domain Architecture |

|
|||||
| Description | Atrial natriuretic peptide receptor A precursor (ANP-A) (ANPRA) (GC-A) (Guanylate cyclase) (EC 4.6.1.2) (NPR-A) (Atrial natriuretic peptide A-type receptor). | |||||
|
ANPRA_MOUSE
|
||||||
| NC score | 0.005359 (rank : 154) | θ value | 3.0926 (rank : 67) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 600 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P18293 | Gene names | Npr1, Npra | |||
|
Domain Architecture |

|
|||||
| Description | Atrial natriuretic peptide receptor A precursor (ANP-A) (ANPRA) (GC-A) (Guanylate cyclase) (EC 4.6.1.2) (NPR-A) (Atrial natriuretic peptide A-type receptor). | |||||
|
PRS8_HUMAN
|
||||||
| NC score | 0.004419 (rank : 155) | θ value | 8.99809 (rank : 121) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P62195, O35051, O43208, P47210, P52915, P52916 | Gene names | PSMC5, SUG1 | |||
|
Domain Architecture |

|
|||||
| Description | 26S protease regulatory subunit 8 (Proteasome 26S subunit ATPase 5) (Proteasome subunit p45) (p45/SUG) (Thyroid hormone receptor- interacting protein 1) (TRIP1). | |||||
|
PRS8_MOUSE
|
||||||
| NC score | 0.004419 (rank : 156) | θ value | 8.99809 (rank : 122) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P62196, O35051, P47210, P52915, P52916, Q3UL51, Q9CWN5 | Gene names | Psmc5, Sug1 | |||
|
Domain Architecture |

|
|||||
| Description | 26S protease regulatory subunit 8 (Proteasome 26S subunit ATPase 5) (Proteasome subunit p45) (p45/SUG) (mSUG1). | |||||
|
HS105_MOUSE
|
||||||
| NC score | 0.004250 (rank : 157) | θ value | 8.99809 (rank : 114) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q61699, Q3TNS2, Q3UIY8, Q62578, Q62579, Q6A0A5, Q8C430, Q8VCW6 | Gene names | Hsph1, Hsp105, Hsp110, Kiaa0201 | |||
|
Domain Architecture |

|
|||||
| Description | Heat-shock protein 105 kDa (Heat shock-related 100 kDa protein E7I) (HSP-E7I) (Heat shock 110 kDa protein) (42 degrees C-HSP). | |||||
|
ANXA9_MOUSE
|
||||||
| NC score | 0.004184 (rank : 158) | θ value | 8.99809 (rank : 108) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JHQ0, Q9CQS1 | Gene names | Anxa9 | |||
|
Domain Architecture |

|
|||||
| Description | Annexin A9 (Annexin-31) (Annexin XXXI). | |||||
|
NFKB2_HUMAN
|
||||||
| NC score | 0.002520 (rank : 159) | θ value | 8.99809 (rank : 118) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q00653, Q04860, Q9BU75, Q9H471, Q9H472 | Gene names | NFKB2, LYT10 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor NF-kappa-B p100 subunit (DNA-binding factor KBF2) (H2TF1) (Lymphocyte translocation chromosome 10) (Oncogene Lyt-10) (Lyt10) [Contains: Nuclear factor NF-kappa-B p52 subunit]. | |||||