Please be patient as the page loads
|
CNN2_MOUSE
|
||||||
| SwissProt Accessions | Q08093 | Gene names | Cnn2 | |||
|
Domain Architecture |
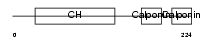
|
|||||
| Description | Calponin-2 (Calponin H2, smooth muscle) (Neutral calponin). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
CNN2_MOUSE
|
||||||
| θ value | 6.14192e-166 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q08093 | Gene names | Cnn2 | |||
|
Domain Architecture |

|
|||||
| Description | Calponin-2 (Calponin H2, smooth muscle) (Neutral calponin). | |||||
|
CNN2_HUMAN
|
||||||
| θ value | 3.37019e-164 (rank : 2) | NC score | 0.998826 (rank : 2) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q99439, Q92578 | Gene names | CNN2 | |||
|
Domain Architecture |
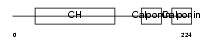
|
|||||
| Description | Calponin-2 (Calponin H2, smooth muscle) (Neutral calponin). | |||||
|
CNN3_HUMAN
|
||||||
| θ value | 4.04623e-109 (rank : 3) | NC score | 0.991115 (rank : 3) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q15417 | Gene names | CNN3 | |||
|
Domain Architecture |
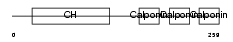
|
|||||
| Description | Calponin-3 (Calponin, acidic isoform). | |||||
|
CNN3_MOUSE
|
||||||
| θ value | 4.47364e-108 (rank : 4) | NC score | 0.990618 (rank : 6) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9DAW9, Q3TW23 | Gene names | Cnn3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calponin-3 (Calponin, acidic isoform). | |||||
|
CNN1_HUMAN
|
||||||
| θ value | 6.91785e-101 (rank : 5) | NC score | 0.990707 (rank : 5) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P51911, O00638, Q15416, Q8IY93, Q99438 | Gene names | CNN1 | |||
|
Domain Architecture |
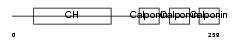
|
|||||
| Description | Calponin-1 (Calponin H1, smooth muscle) (Basic calponin). | |||||
|
CNN1_MOUSE
|
||||||
| θ value | 1.18001e-100 (rank : 6) | NC score | 0.990759 (rank : 4) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q08091 | Gene names | Cnn1 | |||
|
Domain Architecture |
|
|||||
| Description | Calponin-1 (Calponin H1, smooth muscle) (Basic calponin). | |||||
|
TAGL2_HUMAN
|
||||||
| θ value | 2.59387e-31 (rank : 7) | NC score | 0.880814 (rank : 10) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P37802, Q5JRQ6, Q5JRQ7, Q9BUH5, Q9H4P0 | Gene names | TAGLN2, KIAA0120 | |||
|
Domain Architecture |
|
|||||
| Description | Transgelin-2 (SM22-alpha homolog). | |||||
|
TAGL3_HUMAN
|
||||||
| θ value | 5.77852e-31 (rank : 8) | NC score | 0.882021 (rank : 9) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UI15, Q96A74 | Gene names | TAGLN3, NP25 | |||
|
Domain Architecture |
|
|||||
| Description | Transgelin-3 (Neuronal protein NP25) (Neuronal protein 22) (NP22). | |||||
|
TAGL2_MOUSE
|
||||||
| θ value | 7.54701e-31 (rank : 9) | NC score | 0.878459 (rank : 12) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9WVA4 | Gene names | Tagln2 | |||
|
Domain Architecture |
|
|||||
| Description | Transgelin-2. | |||||
|
TAGL3_MOUSE
|
||||||
| θ value | 1.28732e-30 (rank : 10) | NC score | 0.880700 (rank : 11) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9R1Q8 | Gene names | Tagln3, Np25 | |||
|
Domain Architecture |
|
|||||
| Description | Transgelin-3 (Neuronal protein NP25). | |||||
|
TAGL_MOUSE
|
||||||
| θ value | 1.08979e-29 (rank : 11) | NC score | 0.882878 (rank : 8) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P37804, Q545W0 | Gene names | Tagln, Sm22, Sm22a | |||
|
Domain Architecture |
|
|||||
| Description | Transgelin (Smooth muscle protein 22-alpha) (SM22-alpha) (Actin- associated protein p27). | |||||
|
TAGL_HUMAN
|
||||||
| θ value | 2.42779e-29 (rank : 12) | NC score | 0.883676 (rank : 7) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q01995, O15542 | Gene names | TAGLN, SM22, WS3-10 | |||
|
Domain Architecture |
|
|||||
| Description | Transgelin (Smooth muscle protein 22-alpha) (SM22-alpha) (WS3-10) (22 kDa actin-binding protein). | |||||
|
LMO7_HUMAN
|
||||||
| θ value | 8.11959e-09 (rank : 13) | NC score | 0.425135 (rank : 13) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8WWI1, O15462, O95346, Q9UKC1, Q9UQM5, Q9Y6A7 | Gene names | LMO7, FBX20, FBXO20, KIAA0858 | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain only protein 7 (LOMP) (F-box only protein 20). | |||||
|
ARHG6_MOUSE
|
||||||
| θ value | 4.92598e-06 (rank : 14) | NC score | 0.253831 (rank : 14) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8K4I3, Q8C9V4 | Gene names | Arhgef6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 6 (Rac/Cdc42 guanine nucleotide exchange factor 6). | |||||
|
ARHG6_HUMAN
|
||||||
| θ value | 1.09739e-05 (rank : 15) | NC score | 0.240907 (rank : 15) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 376 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q15052, Q15396, Q5JQ66, Q86XH0 | Gene names | ARHGEF6, COOL2, KIAA0006, PIXA | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 6 (Rac/Cdc42 guanine nucleotide exchange factor 6) (PAK-interacting exchange factor alpha) (Alpha-Pix) (COOL-2). | |||||
|
VAV2_MOUSE
|
||||||
| θ value | 1.43324e-05 (rank : 16) | NC score | 0.227726 (rank : 19) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 456 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q60992 | Gene names | Vav2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-2. | |||||
|
VAV2_HUMAN
|
||||||
| θ value | 1.87187e-05 (rank : 17) | NC score | 0.227889 (rank : 18) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 453 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P52735 | Gene names | VAV2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-2. | |||||
|
VAV3_MOUSE
|
||||||
| θ value | 4.1701e-05 (rank : 18) | NC score | 0.237098 (rank : 16) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 404 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9R0C8, Q7TS85 | Gene names | Vav3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-3. | |||||
|
LRCH3_HUMAN
|
||||||
| θ value | 0.000121331 (rank : 19) | NC score | 0.212784 (rank : 24) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 270 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q96II8, Q96FP9, Q9NT52 | Gene names | LRCH3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 3 precursor. | |||||
|
ARHG7_HUMAN
|
||||||
| θ value | 0.00035302 (rank : 20) | NC score | 0.163339 (rank : 30) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q14155, Q6P9G3, Q6PII2, Q86W63, Q8N3M1 | Gene names | ARHGEF7, COOL1, KIAA0142, P85SPR, PAK3BP, PIXB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 7 (PAK-interacting exchange factor beta) (Beta-Pix) (COOL-1) (p85). | |||||
|
LRCH4_HUMAN
|
||||||
| θ value | 0.00035302 (rank : 21) | NC score | 0.184891 (rank : 29) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O75427, Q8WV85, Q96ID0 | Gene names | LRCH4, LRN, LRRN4 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 4 (Leucine-rich neuronal protein). | |||||
|
VAV3_HUMAN
|
||||||
| θ value | 0.00035302 (rank : 22) | NC score | 0.223285 (rank : 23) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 398 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9UKW4, O95230, Q9Y5X8 | Gene names | VAV3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-3. | |||||
|
VAV_MOUSE
|
||||||
| θ value | 0.00035302 (rank : 23) | NC score | 0.229732 (rank : 17) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 430 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P27870, Q8BTV7 | Gene names | Vav1, Vav | |||
|
Domain Architecture |

|
|||||
| Description | Proto-oncogene vav (p95vav). | |||||
|
VAV_HUMAN
|
||||||
| θ value | 0.000602161 (rank : 24) | NC score | 0.223375 (rank : 22) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P15498, Q15860 | Gene names | VAV1, VAV | |||
|
Domain Architecture |

|
|||||
| Description | Proto-oncogene vav. | |||||
|
LRCH3_MOUSE
|
||||||
| θ value | 0.000786445 (rank : 25) | NC score | 0.202383 (rank : 26) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8BVU0, Q3U222, Q3UZ74 | Gene names | Lrch3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 3 precursor. | |||||
|
LRCH1_HUMAN
|
||||||
| θ value | 0.00102713 (rank : 26) | NC score | 0.224680 (rank : 20) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9Y2L9, Q7Z5F6, Q7Z5F7 | Gene names | LRCH1, CHDC1, KIAA1016 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 1 (Calponin homology domain-containing protein 1) (Neuronal protein 81) (NP81). | |||||
|
LRCH4_MOUSE
|
||||||
| θ value | 0.00102713 (rank : 27) | NC score | 0.190491 (rank : 28) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q921G6 | Gene names | Lrch4 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 4. | |||||
|
LRCH2_HUMAN
|
||||||
| θ value | 0.00134147 (rank : 28) | NC score | 0.204082 (rank : 25) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q5VUJ6, Q9HA88, Q9P233 | Gene names | LRCH2, KIAA1495 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 2. | |||||
|
LRCH2_MOUSE
|
||||||
| θ value | 0.00228821 (rank : 29) | NC score | 0.201261 (rank : 27) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q3UMG5, Q7TNP9, Q80TC9 | Gene names | Lrch2, Kiaa1495 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 2. | |||||
|
LRCH1_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 30) | NC score | 0.224479 (rank : 21) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P62046 | Gene names | Lrch1, Chdc1, Kiaa1016 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 1 (Calponin homology domain-containing protein 1). | |||||
|
GA2L3_HUMAN
|
||||||
| θ value | 0.163984 (rank : 31) | NC score | 0.098726 (rank : 31) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q86XJ1 | Gene names | GAS2L3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GAS2-like protein 3 (Growth arrest-specific 2-like 3). | |||||
|
IQGA1_HUMAN
|
||||||
| θ value | 0.21417 (rank : 32) | NC score | 0.072434 (rank : 35) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P46940 | Gene names | IQGAP1, KIAA0051 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP1 (p195). | |||||
|
IQGA1_MOUSE
|
||||||
| θ value | 0.21417 (rank : 33) | NC score | 0.073462 (rank : 34) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9JKF1 | Gene names | Iqgap1 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP1. | |||||
|
GAS2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 34) | NC score | 0.063161 (rank : 40) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O43903, Q6ICV8 | Gene names | GAS2 | |||
|
Domain Architecture |
|
|||||
| Description | Growth-arrest-specific protein 2 (GAS-2). | |||||
|
GAS2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 35) | NC score | 0.049660 (rank : 48) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P11862 | Gene names | Gas2, Gas-2 | |||
|
Domain Architecture |
|
|||||
| Description | Growth-arrest-specific protein 2 (GAS-2). | |||||
|
WNT11_HUMAN
|
||||||
| θ value | 5.27518 (rank : 36) | NC score | 0.009763 (rank : 51) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O96014, Q8WZ98 | Gene names | WNT11 | |||
|
Domain Architecture |
|
|||||
| Description | Protein Wnt-11 precursor. | |||||
|
WNT11_MOUSE
|
||||||
| θ value | 5.27518 (rank : 37) | NC score | 0.009719 (rank : 52) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P48615 | Gene names | Wnt11, Wnt-11 | |||
|
Domain Architecture |
|
|||||
| Description | Protein Wnt-11 precursor. | |||||
|
GPAA1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 38) | NC score | 0.016018 (rank : 49) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9WTK3, Q3TNU9, Q9R1U8 | Gene names | Gpaa1, Gaa1 | |||
|
Domain Architecture |

|
|||||
| Description | Glycosylphosphatidylinositol anchor attachment 1 protein (GPI anchor attachment protein 1) (GAA1 protein homolog) (mGAA1). | |||||
|
MICA1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 39) | NC score | 0.012317 (rank : 50) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 400 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8VDP3 | Gene names | Mical1, Mical, Nical | |||
|
Domain Architecture |

|
|||||
| Description | NEDD9-interacting protein with calponin homology and LIM domains (Molecule interacting with CasL protein 1). | |||||
|
ARHG4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 40) | NC score | 0.060656 (rank : 41) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NR80, Q9HDC6, Q9UPP0 | Gene names | ARHGEF4, KIAA1112 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 4 (APC-stimulated guanine nucleotide exchange factor) (Asef). | |||||
|
ARHG4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 41) | NC score | 0.063987 (rank : 39) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q7TNR9, Q80TJ6 | Gene names | Arhgef4, Kiaa1112 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 4 (APC-stimulated guanine nucleotide exchange factor) (Asef). | |||||
|
ARHG7_MOUSE
|
||||||
| θ value | θ > 10 (rank : 42) | NC score | 0.078735 (rank : 32) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 396 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9ES28, O08757, Q9ES27 | Gene names | Arhgef7, Pak3bp | |||
|
Domain Architecture |
|
|||||
| Description | Rho guanine nucleotide exchange factor 7 (PAK-interacting exchange factor beta) (Beta-Pix) (p85SPR). | |||||
|
ARHG9_HUMAN
|
||||||
| θ value | θ > 10 (rank : 43) | NC score | 0.059064 (rank : 43) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O43307, Q5JSL6 | Gene names | ARHGEF9, KIAA0424 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 9 (Rac/Cdc42 guanine nucleotide exchange factor 9) (Collybistin) (PEM-2 homolog). | |||||
|
ARHG9_MOUSE
|
||||||
| θ value | θ > 10 (rank : 44) | NC score | 0.059064 (rank : 44) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q3UTH8, Q3TQ60, Q80U06, Q8CAF9 | Gene names | Arhgef9, Kiaa0424 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 9 (Rac/Cdc42 guanine nucleotide exchange factor 9) (Collybistin). | |||||
|
GA2L2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 45) | NC score | 0.074321 (rank : 33) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8NHY3, Q8NHY4 | Gene names | GAS2L2, GAR17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GAS2-like protein 2 (Growth arrest-specific 2-like 2) (GAS2-related protein on chromosome 17) (GAR17 protein). | |||||
|
GA2L2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 46) | NC score | 0.067742 (rank : 38) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5SSG4 | Gene names | Gas2l2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GAS2-like protein 2 (Growth arrest-specific 2-like 2). | |||||
|
IQGA3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 47) | NC score | 0.068031 (rank : 37) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q86VI3 | Gene names | IQGAP3 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP3. | |||||
|
MCF2L_MOUSE
|
||||||
| θ value | θ > 10 (rank : 48) | NC score | 0.052925 (rank : 47) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q64096 | Gene names | Mcf2l, Dbs | |||
|
Domain Architecture |

|
|||||
| Description | Guanine nucleotide exchange factor DBS (DBL's big sister) (MCF2- transforming sequence-like protein). | |||||
|
PKHG1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 49) | NC score | 0.072074 (rank : 36) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9ULL1, Q5T1F2 | Gene names | PLEKHG1, KIAA1209 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family G member 1. | |||||
|
PREX1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 50) | NC score | 0.059987 (rank : 42) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8TCU6, Q5JS95, Q5JS96, Q69YL2, Q7Z2L9, Q9BQH0, Q9BX55, Q9H4Q6, Q9P2D2, Q9UGQ4 | Gene names | PREX1, KIAA1415 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 3,4,5-trisphosphate-dependent Rac exchanger 1 protein (P-Rex1 protein). | |||||
|
TIAM1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 51) | NC score | 0.054302 (rank : 46) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q13009 | Gene names | TIAM1 | |||
|
Domain Architecture |

|
|||||
| Description | T-lymphoma invasion and metastasis-inducing protein 1 (TIAM-1 protein). | |||||
|
TIAM1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 52) | NC score | 0.054989 (rank : 45) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q60610 | Gene names | Tiam1, Tiam-1 | |||
|
Domain Architecture |

|
|||||
| Description | T-lymphoma invasion and metastasis-inducing protein 1 (TIAM-1 protein). | |||||
|
CNN2_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 6.14192e-166 (rank : 1) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q08093 | Gene names | Cnn2 | |||
|
Domain Architecture |

|
|||||
| Description | Calponin-2 (Calponin H2, smooth muscle) (Neutral calponin). | |||||
|
CNN2_HUMAN
|
||||||
| NC score | 0.998826 (rank : 2) | θ value | 3.37019e-164 (rank : 2) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q99439, Q92578 | Gene names | CNN2 | |||
|
Domain Architecture |

|
|||||
| Description | Calponin-2 (Calponin H2, smooth muscle) (Neutral calponin). | |||||
|
CNN3_HUMAN
|
||||||
| NC score | 0.991115 (rank : 3) | θ value | 4.04623e-109 (rank : 3) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q15417 | Gene names | CNN3 | |||
|
Domain Architecture |

|
|||||
| Description | Calponin-3 (Calponin, acidic isoform). | |||||
|
CNN1_MOUSE
|
||||||
| NC score | 0.990759 (rank : 4) | θ value | 1.18001e-100 (rank : 6) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q08091 | Gene names | Cnn1 | |||
|
Domain Architecture |

|
|||||
| Description | Calponin-1 (Calponin H1, smooth muscle) (Basic calponin). | |||||
|
CNN1_HUMAN
|
||||||
| NC score | 0.990707 (rank : 5) | θ value | 6.91785e-101 (rank : 5) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P51911, O00638, Q15416, Q8IY93, Q99438 | Gene names | CNN1 | |||
|
Domain Architecture |

|
|||||
| Description | Calponin-1 (Calponin H1, smooth muscle) (Basic calponin). | |||||
|
CNN3_MOUSE
|
||||||
| NC score | 0.990618 (rank : 6) | θ value | 4.47364e-108 (rank : 4) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9DAW9, Q3TW23 | Gene names | Cnn3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calponin-3 (Calponin, acidic isoform). | |||||
|
TAGL_HUMAN
|
||||||
| NC score | 0.883676 (rank : 7) | θ value | 2.42779e-29 (rank : 12) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q01995, O15542 | Gene names | TAGLN, SM22, WS3-10 | |||
|
Domain Architecture |

|
|||||
| Description | Transgelin (Smooth muscle protein 22-alpha) (SM22-alpha) (WS3-10) (22 kDa actin-binding protein). | |||||
|
TAGL_MOUSE
|
||||||
| NC score | 0.882878 (rank : 8) | θ value | 1.08979e-29 (rank : 11) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P37804, Q545W0 | Gene names | Tagln, Sm22, Sm22a | |||
|
Domain Architecture |

|
|||||
| Description | Transgelin (Smooth muscle protein 22-alpha) (SM22-alpha) (Actin- associated protein p27). | |||||
|
TAGL3_HUMAN
|
||||||
| NC score | 0.882021 (rank : 9) | θ value | 5.77852e-31 (rank : 8) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UI15, Q96A74 | Gene names | TAGLN3, NP25 | |||
|
Domain Architecture |

|
|||||
| Description | Transgelin-3 (Neuronal protein NP25) (Neuronal protein 22) (NP22). | |||||
|
TAGL2_HUMAN
|
||||||
| NC score | 0.880814 (rank : 10) | θ value | 2.59387e-31 (rank : 7) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P37802, Q5JRQ6, Q5JRQ7, Q9BUH5, Q9H4P0 | Gene names | TAGLN2, KIAA0120 | |||
|
Domain Architecture |

|
|||||
| Description | Transgelin-2 (SM22-alpha homolog). | |||||
|
TAGL3_MOUSE
|
||||||
| NC score | 0.880700 (rank : 11) | θ value | 1.28732e-30 (rank : 10) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9R1Q8 | Gene names | Tagln3, Np25 | |||
|
Domain Architecture |

|
|||||
| Description | Transgelin-3 (Neuronal protein NP25). | |||||
|
TAGL2_MOUSE
|
||||||
| NC score | 0.878459 (rank : 12) | θ value | 7.54701e-31 (rank : 9) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9WVA4 | Gene names | Tagln2 | |||
|
Domain Architecture |

|
|||||
| Description | Transgelin-2. | |||||
|
LMO7_HUMAN
|
||||||
| NC score | 0.425135 (rank : 13) | θ value | 8.11959e-09 (rank : 13) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8WWI1, O15462, O95346, Q9UKC1, Q9UQM5, Q9Y6A7 | Gene names | LMO7, FBX20, FBXO20, KIAA0858 | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain only protein 7 (LOMP) (F-box only protein 20). | |||||
|
ARHG6_MOUSE
|
||||||
| NC score | 0.253831 (rank : 14) | θ value | 4.92598e-06 (rank : 14) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8K4I3, Q8C9V4 | Gene names | Arhgef6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 6 (Rac/Cdc42 guanine nucleotide exchange factor 6). | |||||
|
ARHG6_HUMAN
|
||||||
| NC score | 0.240907 (rank : 15) | θ value | 1.09739e-05 (rank : 15) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 376 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q15052, Q15396, Q5JQ66, Q86XH0 | Gene names | ARHGEF6, COOL2, KIAA0006, PIXA | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 6 (Rac/Cdc42 guanine nucleotide exchange factor 6) (PAK-interacting exchange factor alpha) (Alpha-Pix) (COOL-2). | |||||
|
VAV3_MOUSE
|
||||||
| NC score | 0.237098 (rank : 16) | θ value | 4.1701e-05 (rank : 18) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 404 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9R0C8, Q7TS85 | Gene names | Vav3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-3. | |||||
|
VAV_MOUSE
|
||||||
| NC score | 0.229732 (rank : 17) | θ value | 0.00035302 (rank : 23) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 430 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P27870, Q8BTV7 | Gene names | Vav1, Vav | |||
|
Domain Architecture |

|
|||||
| Description | Proto-oncogene vav (p95vav). | |||||
|
VAV2_HUMAN
|
||||||
| NC score | 0.227889 (rank : 18) | θ value | 1.87187e-05 (rank : 17) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 453 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P52735 | Gene names | VAV2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-2. | |||||
|
VAV2_MOUSE
|
||||||
| NC score | 0.227726 (rank : 19) | θ value | 1.43324e-05 (rank : 16) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 456 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q60992 | Gene names | Vav2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-2. | |||||
|
LRCH1_HUMAN
|
||||||
| NC score | 0.224680 (rank : 20) | θ value | 0.00102713 (rank : 26) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9Y2L9, Q7Z5F6, Q7Z5F7 | Gene names | LRCH1, CHDC1, KIAA1016 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 1 (Calponin homology domain-containing protein 1) (Neuronal protein 81) (NP81). | |||||
|
LRCH1_MOUSE
|
||||||
| NC score | 0.224479 (rank : 21) | θ value | 0.00390308 (rank : 30) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P62046 | Gene names | Lrch1, Chdc1, Kiaa1016 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 1 (Calponin homology domain-containing protein 1). | |||||
|
VAV_HUMAN
|
||||||
| NC score | 0.223375 (rank : 22) | θ value | 0.000602161 (rank : 24) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P15498, Q15860 | Gene names | VAV1, VAV | |||
|
Domain Architecture |

|
|||||
| Description | Proto-oncogene vav. | |||||
|
VAV3_HUMAN
|
||||||
| NC score | 0.223285 (rank : 23) | θ value | 0.00035302 (rank : 22) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 398 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9UKW4, O95230, Q9Y5X8 | Gene names | VAV3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-3. | |||||
|
LRCH3_HUMAN
|
||||||
| NC score | 0.212784 (rank : 24) | θ value | 0.000121331 (rank : 19) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 270 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q96II8, Q96FP9, Q9NT52 | Gene names | LRCH3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 3 precursor. | |||||
|
LRCH2_HUMAN
|
||||||
| NC score | 0.204082 (rank : 25) | θ value | 0.00134147 (rank : 28) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q5VUJ6, Q9HA88, Q9P233 | Gene names | LRCH2, KIAA1495 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 2. | |||||
|
LRCH3_MOUSE
|
||||||
| NC score | 0.202383 (rank : 26) | θ value | 0.000786445 (rank : 25) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8BVU0, Q3U222, Q3UZ74 | Gene names | Lrch3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 3 precursor. | |||||
|
LRCH2_MOUSE
|
||||||
| NC score | 0.201261 (rank : 27) | θ value | 0.00228821 (rank : 29) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q3UMG5, Q7TNP9, Q80TC9 | Gene names | Lrch2, Kiaa1495 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 2. | |||||
|
LRCH4_MOUSE
|
||||||
| NC score | 0.190491 (rank : 28) | θ value | 0.00102713 (rank : 27) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q921G6 | Gene names | Lrch4 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 4. | |||||
|
LRCH4_HUMAN
|
||||||
| NC score | 0.184891 (rank : 29) | θ value | 0.00035302 (rank : 21) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O75427, Q8WV85, Q96ID0 | Gene names | LRCH4, LRN, LRRN4 | |||
|
Domain Architecture |

|
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 4 (Leucine-rich neuronal protein). | |||||
|
ARHG7_HUMAN
|
||||||
| NC score | 0.163339 (rank : 30) | θ value | 0.00035302 (rank : 20) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q14155, Q6P9G3, Q6PII2, Q86W63, Q8N3M1 | Gene names | ARHGEF7, COOL1, KIAA0142, P85SPR, PAK3BP, PIXB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 7 (PAK-interacting exchange factor beta) (Beta-Pix) (COOL-1) (p85). | |||||
|
GA2L3_HUMAN
|
||||||
| NC score | 0.098726 (rank : 31) | θ value | 0.163984 (rank : 31) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q86XJ1 | Gene names | GAS2L3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GAS2-like protein 3 (Growth arrest-specific 2-like 3). | |||||
|
ARHG7_MOUSE
|
||||||
| NC score | 0.078735 (rank : 32) | θ value | θ > 10 (rank : 42) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 396 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9ES28, O08757, Q9ES27 | Gene names | Arhgef7, Pak3bp | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 7 (PAK-interacting exchange factor beta) (Beta-Pix) (p85SPR). | |||||
|
GA2L2_HUMAN
|
||||||
| NC score | 0.074321 (rank : 33) | θ value | θ > 10 (rank : 45) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8NHY3, Q8NHY4 | Gene names | GAS2L2, GAR17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GAS2-like protein 2 (Growth arrest-specific 2-like 2) (GAS2-related protein on chromosome 17) (GAR17 protein). | |||||
|
IQGA1_MOUSE
|
||||||
| NC score | 0.073462 (rank : 34) | θ value | 0.21417 (rank : 33) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9JKF1 | Gene names | Iqgap1 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP1. | |||||
|
IQGA1_HUMAN
|
||||||
| NC score | 0.072434 (rank : 35) | θ value | 0.21417 (rank : 32) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P46940 | Gene names | IQGAP1, KIAA0051 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP1 (p195). | |||||
|
PKHG1_HUMAN
|
||||||
| NC score | 0.072074 (rank : 36) | θ value | θ > 10 (rank : 49) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9ULL1, Q5T1F2 | Gene names | PLEKHG1, KIAA1209 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family G member 1. | |||||
|
IQGA3_HUMAN
|
||||||
| NC score | 0.068031 (rank : 37) | θ value | θ > 10 (rank : 47) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q86VI3 | Gene names | IQGAP3 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP3. | |||||
|
GA2L2_MOUSE
|
||||||
| NC score | 0.067742 (rank : 38) | θ value | θ > 10 (rank : 46) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5SSG4 | Gene names | Gas2l2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GAS2-like protein 2 (Growth arrest-specific 2-like 2). | |||||
|
ARHG4_MOUSE
|
||||||
| NC score | 0.063987 (rank : 39) | θ value | θ > 10 (rank : 41) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q7TNR9, Q80TJ6 | Gene names | Arhgef4, Kiaa1112 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 4 (APC-stimulated guanine nucleotide exchange factor) (Asef). | |||||
|
GAS2_HUMAN
|
||||||
| NC score | 0.063161 (rank : 40) | θ value | 2.36792 (rank : 34) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O43903, Q6ICV8 | Gene names | GAS2 | |||
|
Domain Architecture |

|
|||||
| Description | Growth-arrest-specific protein 2 (GAS-2). | |||||
|
ARHG4_HUMAN
|
||||||
| NC score | 0.060656 (rank : 41) | θ value | θ > 10 (rank : 40) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NR80, Q9HDC6, Q9UPP0 | Gene names | ARHGEF4, KIAA1112 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 4 (APC-stimulated guanine nucleotide exchange factor) (Asef). | |||||
|
PREX1_HUMAN
|
||||||
| NC score | 0.059987 (rank : 42) | θ value | θ > 10 (rank : 50) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8TCU6, Q5JS95, Q5JS96, Q69YL2, Q7Z2L9, Q9BQH0, Q9BX55, Q9H4Q6, Q9P2D2, Q9UGQ4 | Gene names | PREX1, KIAA1415 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 3,4,5-trisphosphate-dependent Rac exchanger 1 protein (P-Rex1 protein). | |||||
|
ARHG9_HUMAN
|
||||||
| NC score | 0.059064 (rank : 43) | θ value | θ > 10 (rank : 43) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O43307, Q5JSL6 | Gene names | ARHGEF9, KIAA0424 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 9 (Rac/Cdc42 guanine nucleotide exchange factor 9) (Collybistin) (PEM-2 homolog). | |||||
|
ARHG9_MOUSE
|
||||||
| NC score | 0.059064 (rank : 44) | θ value | θ > 10 (rank : 44) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q3UTH8, Q3TQ60, Q80U06, Q8CAF9 | Gene names | Arhgef9, Kiaa0424 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 9 (Rac/Cdc42 guanine nucleotide exchange factor 9) (Collybistin). | |||||
|
TIAM1_MOUSE
|
||||||
| NC score | 0.054989 (rank : 45) | θ value | θ > 10 (rank : 52) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q60610 | Gene names | Tiam1, Tiam-1 | |||
|
Domain Architecture |

|
|||||
| Description | T-lymphoma invasion and metastasis-inducing protein 1 (TIAM-1 protein). | |||||
|
TIAM1_HUMAN
|
||||||
| NC score | 0.054302 (rank : 46) | θ value | θ > 10 (rank : 51) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q13009 | Gene names | TIAM1 | |||
|
Domain Architecture |

|
|||||
| Description | T-lymphoma invasion and metastasis-inducing protein 1 (TIAM-1 protein). | |||||
|
MCF2L_MOUSE
|
||||||
| NC score | 0.052925 (rank : 47) | θ value | θ > 10 (rank : 48) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q64096 | Gene names | Mcf2l, Dbs | |||
|
Domain Architecture |

|
|||||
| Description | Guanine nucleotide exchange factor DBS (DBL's big sister) (MCF2- transforming sequence-like protein). | |||||
|
GAS2_MOUSE
|
||||||
| NC score | 0.049660 (rank : 48) | θ value | 5.27518 (rank : 35) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P11862 | Gene names | Gas2, Gas-2 | |||
|
Domain Architecture |

|
|||||
| Description | Growth-arrest-specific protein 2 (GAS-2). | |||||
|
GPAA1_MOUSE
|
||||||
| NC score | 0.016018 (rank : 49) | θ value | 8.99809 (rank : 38) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9WTK3, Q3TNU9, Q9R1U8 | Gene names | Gpaa1, Gaa1 | |||
|
Domain Architecture |

|
|||||
| Description | Glycosylphosphatidylinositol anchor attachment 1 protein (GPI anchor attachment protein 1) (GAA1 protein homolog) (mGAA1). | |||||
|
MICA1_MOUSE
|
||||||
| NC score | 0.012317 (rank : 50) | θ value | 8.99809 (rank : 39) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 400 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8VDP3 | Gene names | Mical1, Mical, Nical | |||
|
Domain Architecture |

|
|||||
| Description | NEDD9-interacting protein with calponin homology and LIM domains (Molecule interacting with CasL protein 1). | |||||
|
WNT11_HUMAN
|
||||||
| NC score | 0.009763 (rank : 51) | θ value | 5.27518 (rank : 36) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O96014, Q8WZ98 | Gene names | WNT11 | |||
|
Domain Architecture |

|
|||||
| Description | Protein Wnt-11 precursor. | |||||
|
WNT11_MOUSE
|
||||||
| NC score | 0.009719 (rank : 52) | θ value | 5.27518 (rank : 37) | |||
| Query Neighborhood Hits | 39 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P48615 | Gene names | Wnt11, Wnt-11 | |||
|
Domain Architecture |

|
|||||
| Description | Protein Wnt-11 precursor. | |||||