Please be patient as the page loads
|
WWTR1_MOUSE
|
||||||
| SwissProt Accessions | Q9EPK5, Q99KI4 | Gene names | Wwtr1, Taz | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing transcription regulator protein 1 (Transcriptional coactivator with PDZ-binding motif). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
WWTR1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.992829 (rank : 2) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9GZV5, Q8N3P2, Q9Y3W6 | Gene names | WWTR1, TAZ | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing transcription regulator protein 1 (Transcriptional coactivator with PDZ-binding motif). | |||||
|
WWTR1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9EPK5, Q99KI4 | Gene names | Wwtr1, Taz | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing transcription regulator protein 1 (Transcriptional coactivator with PDZ-binding motif). | |||||
|
YAP1_HUMAN
|
||||||
| θ value | 4.36332e-79 (rank : 3) | NC score | 0.922775 (rank : 3) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P46937 | Gene names | YAP1, YAP65 | |||
|
Domain Architecture |

|
|||||
| Description | 65 kDa Yes-associated protein (YAP65). | |||||
|
YAP1_MOUSE
|
||||||
| θ value | 4.68348e-65 (rank : 4) | NC score | 0.904695 (rank : 4) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P46938, Q52KJ5 | Gene names | Yap1, Yap, Yap65 | |||
|
Domain Architecture |

|
|||||
| Description | 65 kDa Yes-associated protein (YAP65). | |||||
|
ITCH_HUMAN
|
||||||
| θ value | 3.19293e-05 (rank : 5) | NC score | 0.379304 (rank : 14) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q96J02, O43584, Q5TEL0, Q96F66, Q9BY75, Q9H451, Q9H4U5 | Gene names | ITCH | |||
|
Domain Architecture |

|
|||||
| Description | Itchy homolog E3 ubiquitin protein ligase (EC 6.3.2.-) (Itch) (Atrophin-1-interacting protein 4) (AIP4) (NFE2-associated polypeptide 1) (NAPP1). | |||||
|
ITCH_MOUSE
|
||||||
| θ value | 3.19293e-05 (rank : 6) | NC score | 0.381258 (rank : 12) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8C863, O54971 | Gene names | Itch | |||
|
Domain Architecture |

|
|||||
| Description | Itchy E3 ubiquitin protein ligase (EC 6.3.2.-). | |||||
|
WWP2_HUMAN
|
||||||
| θ value | 3.19293e-05 (rank : 7) | NC score | 0.382767 (rank : 11) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O00308, Q96CZ2, Q9BWN6 | Gene names | WWP2 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP2 (EC 6.3.2.-) (WW domain- containing protein 2) (Atrophin-1-interacting protein 2) (AIP2). | |||||
|
WWP2_MOUSE
|
||||||
| θ value | 3.19293e-05 (rank : 8) | NC score | 0.383664 (rank : 10) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9DBH0, Q8BTG4, Q923F6 | Gene names | Wwp2 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP2 (EC 6.3.2.-) (WW domain- containing protein 2). | |||||
|
NEDD4_MOUSE
|
||||||
| θ value | 4.1701e-05 (rank : 9) | NC score | 0.353588 (rank : 21) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P46935, O08758, Q3UZI2, Q8BGB3 | Gene names | Nedd4, Kiaa0093, Nedd-4, Nedd4a | |||
|
Domain Architecture |

|
|||||
| Description | E3 ubiquitin-protein ligase NEDD4 (EC 6.3.2.-) (Neural precursor cell expressed developmentally down-regulated protein 4). | |||||
|
SMUF2_HUMAN
|
||||||
| θ value | 4.1701e-05 (rank : 10) | NC score | 0.367195 (rank : 15) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9HAU4, Q9H260 | Gene names | SMURF2 | |||
|
Domain Architecture |

|
|||||
| Description | Smad ubiquitination regulatory factor 2 (EC 6.3.2.-) (Ubiquitin-- protein ligase SMURF2) (Smad-specific E3 ubiquitin ligase 2) (hSMURF2). | |||||
|
NEDD4_HUMAN
|
||||||
| θ value | 5.44631e-05 (rank : 11) | NC score | 0.355080 (rank : 19) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P46934 | Gene names | NEDD4, KIAA0093 | |||
|
Domain Architecture |

|
|||||
| Description | E3 ubiquitin-protein ligase NEDD4 (EC 6.3.2.-). | |||||
|
NED4L_HUMAN
|
||||||
| θ value | 7.1131e-05 (rank : 12) | NC score | 0.352715 (rank : 22) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q96PU5, O43165, Q7Z5F1, Q7Z5F2, Q7Z5N3, Q8N5A7, Q8WUU9, Q9BW58, Q9H2W4, Q9NT88 | Gene names | NEDD4L, KIAA0439, NEDL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin-protein ligase NEDD4-like protein (EC 6.3.2.-) (Nedd4-2) (NEDD4.2). | |||||
|
NED4L_MOUSE
|
||||||
| θ value | 7.1131e-05 (rank : 13) | NC score | 0.354024 (rank : 20) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8CFI0, Q8BRT9, Q8BS42, Q99PK2 | Gene names | Nedd4l, Kiaa0439, Nedd4b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin-protein ligase NEDD4-like protein (EC 6.3.2.-) (Nedd4-2) (NEDD4.2). | |||||
|
WWP1_HUMAN
|
||||||
| θ value | 7.1131e-05 (rank : 14) | NC score | 0.379666 (rank : 13) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9H0M0, O00307, Q96BP4 | Gene names | WWP1 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP1 (EC 6.3.2.-) (WW domain- containing protein 1) (Atropin-1-interacting protein 5) (AIP5). | |||||
|
WWP1_MOUSE
|
||||||
| θ value | 7.1131e-05 (rank : 15) | NC score | 0.387002 (rank : 9) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8BZZ3, Q8BIV9, Q8VDP8 | Gene names | Wwp1 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP1 (EC 6.3.2.-) (WW domain- containing protein 1). | |||||
|
MAGI1_HUMAN
|
||||||
| θ value | 9.29e-05 (rank : 16) | NC score | 0.296401 (rank : 32) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q96QZ7, O00309, O43863, O75085, Q96QZ8, Q96QZ9 | Gene names | MAGI1, BAIAP1, BAP1, TNRC19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 1 (BAI1-associated protein 1) (BAP-1) (Membrane-associated guanylate kinase inverted 1) (MAGI-1) (Atrophin-1-interacting protein 3) (AIP3) (WW domain-containing protein 3) (WWP3) (Trinucleotide repeat-containing gene 19 protein). | |||||
|
MAGI1_MOUSE
|
||||||
| θ value | 9.29e-05 (rank : 17) | NC score | 0.296441 (rank : 31) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q6RHR9, O54893, O54894, O54895 | Gene names | Magi1, Baiap1, Bap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 1 (BAI1-associated protein 1) (BAP-1) (Membrane-associated guanylate kinase inverted 1) (MAGI-1). | |||||
|
MAGI2_MOUSE
|
||||||
| θ value | 9.29e-05 (rank : 18) | NC score | 0.313962 (rank : 29) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 370 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9WVQ1, Q3UH81, Q6GT88, Q8BYT1, Q8CA85 | Gene names | Magi2, Acvrinp1, Aip1, Arip1 | |||
|
Domain Architecture |

|
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 2 (Membrane-associated guanylate kinase inverted 2) (MAGI-2) (Atrophin-1-interacting protein 1) (Activin receptor-interacting protein 1) (Acvrip1). | |||||
|
MAGI2_HUMAN
|
||||||
| θ value | 0.000158464 (rank : 19) | NC score | 0.306625 (rank : 30) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 373 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q86UL8, O60434, O60510, Q86UI7, Q9NP44, Q9UDQ5, Q9UDU1 | Gene names | MAGI2, ACVRINP1, AIP1, KIAA0705 | |||
|
Domain Architecture |

|
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 2 (Membrane-associated guanylate kinase inverted 2) (MAGI-2) (Atrophin-1-interacting protein 1) (Atrophin-1-interacting protein A). | |||||
|
SMUF1_HUMAN
|
||||||
| θ value | 0.000158464 (rank : 20) | NC score | 0.357175 (rank : 18) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9HCE7, O75853, Q9UJT8 | Gene names | SMURF1, KIAA1625 | |||
|
Domain Architecture |

|
|||||
| Description | Smad ubiquitination regulatory factor 1 (EC 6.3.2.-) (Ubiquitin-- protein ligase SMURF1) (Smad-specific E3 ubiquitin ligase 1) (hSMURF1). | |||||
|
SMUF1_MOUSE
|
||||||
| θ value | 0.000158464 (rank : 21) | NC score | 0.361418 (rank : 16) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9CUN6, Q3U412, Q8K300 | Gene names | Smurf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Smad ubiquitination regulatory factor 1 (EC 6.3.2.-) (Ubiquitin-- protein ligase SMURF1) (Smad-specific E3 ubiquitin ligase 1). | |||||
|
BAG3_HUMAN
|
||||||
| θ value | 0.000270298 (rank : 22) | NC score | 0.361164 (rank : 17) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 314 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O95817, Q9NT20, Q9P120 | Gene names | BAG3, BIS | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 3 (BCL-2-binding athanogene- 3) (BAG-3) (Bcl-2-binding protein Bis) (Docking protein CAIR-1). | |||||
|
WWC1_HUMAN
|
||||||
| θ value | 0.00035302 (rank : 23) | NC score | 0.392885 (rank : 7) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8IX03, O94946, Q6MZX4, Q6Y2F8, Q7Z4G8, Q8WVM4, Q9BT29 | Gene names | WWC1, KIAA0869 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing protein 1 (Kidney and brain protein) (KIBRA) (HBeAg-binding protein 3). | |||||
|
WWC1_MOUSE
|
||||||
| θ value | 0.000602161 (rank : 24) | NC score | 0.388151 (rank : 8) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q5SXA9, Q571D0, Q8K1Y3, Q8VD17, Q922W3 | Gene names | Wwc1, Kiaa0869 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing protein 1 (Kidney and brain protein) (KIBRA). | |||||
|
SAV1_HUMAN
|
||||||
| θ value | 0.000786445 (rank : 25) | NC score | 0.467394 (rank : 5) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9H4B6, Q6IA58, Q9H949, Q9HAK9 | Gene names | SAV1, WW45 | |||
|
Domain Architecture |

|
|||||
| Description | Salvador homolog 1 protein (45 kDa WW domain protein) (hWW45). | |||||
|
SAV1_MOUSE
|
||||||
| θ value | 0.000786445 (rank : 26) | NC score | 0.466323 (rank : 6) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8VEB2, Q9D9N9, Q9ER46 | Gene names | Sav1, Ww45, Wwp3 | |||
|
Domain Architecture |

|
|||||
| Description | Salvador homolog 1 protein (45 kDa WW domain protein) (mWW45). | |||||
|
BAG3_MOUSE
|
||||||
| θ value | 0.00102713 (rank : 27) | NC score | 0.319069 (rank : 28) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9JLV1, Q9JJC7 | Gene names | Bag3, Bis | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 3 (BCL-2-binding athanogene- 3) (BAG-3) (Bcl-2-binding protein Bis). | |||||
|
PKHA5_HUMAN
|
||||||
| θ value | 0.00102713 (rank : 28) | NC score | 0.246808 (rank : 33) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9HAU0, Q8N3K6, Q96DY9, Q9BVR4, Q9C0H7, Q9H924, Q9NVK8 | Gene names | PLEKHA5, KIAA1686, PEPP2 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 5 (Phosphoinositol 3-phosphate-binding protein 2) (PEPP-2). | |||||
|
WWOX_HUMAN
|
||||||
| θ value | 0.00102713 (rank : 29) | NC score | 0.337108 (rank : 24) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9NZC7, Q5MYT5, Q96KM3, Q96RF2, Q9BTT8, Q9NPC9, Q9NRF4, Q9NRF5, Q9NRF6, Q9NRK1, Q9NZC5 | Gene names | WWOX, FOR, WOX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing oxidoreductase (EC 1.1.1.-) (Fragile site FRA16D oxidoreductase). | |||||
|
WWOX_MOUSE
|
||||||
| θ value | 0.00102713 (rank : 30) | NC score | 0.334222 (rank : 25) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q91WL8, Q8C8J6, Q920Y2, Q9D2B3, Q9D339, Q9JLF5 | Gene names | Wwox, Wox1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing oxidoreductase (EC 1.1.1.-). | |||||
|
STXB4_HUMAN
|
||||||
| θ value | 0.00298849 (rank : 31) | NC score | 0.222685 (rank : 37) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 616 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q6ZWJ1, Q8IVZ5 | Gene names | STXBP4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntaxin-binding protein 4 (Syntaxin 4-interacting protein) (Synip). | |||||
|
STXB4_MOUSE
|
||||||
| θ value | 0.00298849 (rank : 32) | NC score | 0.225954 (rank : 36) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 629 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9WV89, Q5SUA6, Q5SUA8, Q5SUA9, Q8CFL1 | Gene names | Stxbp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntaxin-binding protein 4 (Syntaxin 4-interacting protein) (Synip). | |||||
|
GAS7_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 33) | NC score | 0.244379 (rank : 34) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q60780, Q9QY25, Q9QY26, Q9QY34 | Gene names | Gas7 | |||
|
Domain Architecture |

|
|||||
| Description | Growth-arrest-specific protein 7 (GAS-7). | |||||
|
PIN1_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 34) | NC score | 0.326469 (rank : 27) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q13526 | Gene names | PIN1 | |||
|
Domain Architecture |
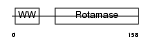
|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase NIMA-interacting 1 (EC 5.2.1.8) (Rotamase Pin1) (PPIase Pin1). | |||||
|
PIN1_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 35) | NC score | 0.331726 (rank : 26) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9QUR7, Q543B3 | Gene names | Pin1 | |||
|
Domain Architecture |
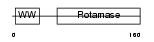
|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase NIMA-interacting 1 (EC 5.2.1.8) (Rotamase Pin1) (PPIase Pin1). | |||||
|
GAS7_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 36) | NC score | 0.230005 (rank : 35) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O60861, O43144 | Gene names | GAS7, KIAA0394 | |||
|
Domain Architecture |

|
|||||
| Description | Growth-arrest-specific protein 7 (GAS-7). | |||||
|
PINL_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 37) | NC score | 0.351807 (rank : 23) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O15428 | Gene names | PIN1L | |||
|
Domain Architecture |
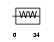
|
|||||
| Description | PIN1-like protein. | |||||
|
DGCR8_HUMAN
|
||||||
| θ value | 0.47712 (rank : 38) | NC score | 0.111621 (rank : 58) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8WYQ5, Q6DCB2, Q6MZE9, Q6Y2L0, Q96G39, Q96GP8, Q9H6L8, Q9H6T7, Q9NRW2 | Gene names | DGCR8, C22orf12, DGCRK6 | |||
|
Domain Architecture |
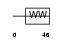
|
|||||
| Description | DGCR8 protein (DiGeorge syndrome critical region 8). | |||||
|
DGCR8_MOUSE
|
||||||
| θ value | 0.47712 (rank : 39) | NC score | 0.110836 (rank : 59) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9EQM6 | Gene names | Dgcr8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DGCR8 protein (DiGeorge syndrome critical region 8 homolog) (Gy1). | |||||
|
DMD_HUMAN
|
||||||
| θ value | 0.813845 (rank : 40) | NC score | 0.045075 (rank : 70) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P11532, Q02295, Q14169, Q14170, Q5JYU0, Q7KZ48 | Gene names | DMD | |||
|
Domain Architecture |

|
|||||
| Description | Dystrophin. | |||||
|
DMD_MOUSE
|
||||||
| θ value | 0.813845 (rank : 41) | NC score | 0.042302 (rank : 71) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1111 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P11531, O35653, Q60703 | Gene names | Dmd | |||
|
Domain Architecture |

|
|||||
| Description | Dystrophin. | |||||
|
BTBDC_HUMAN
|
||||||
| θ value | 1.38821 (rank : 42) | NC score | 0.049062 (rank : 69) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8IY92, Q96JP1 | Gene names | BTBD12, KIAA1784 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein 12. | |||||
|
TLK2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 43) | NC score | 0.003410 (rank : 82) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1088 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O55047, Q9D5Y5 | Gene names | Tlk2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase tousled-like 2 (EC 2.7.11.1) (Tousled- like kinase 2). | |||||
|
MYH7_HUMAN
|
||||||
| θ value | 3.0926 (rank : 44) | NC score | 0.008653 (rank : 77) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P12883, Q14836, Q14837, Q14904, Q16579, Q92679, Q9H1D5, Q9UMM8 | Gene names | MYH7, MYHCB | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle beta isoform (MyHC-beta). | |||||
|
USF2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 45) | NC score | 0.029898 (rank : 72) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q64705, Q3UIL2, Q8VE59 | Gene names | Usf2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (Major late transcription factor 2). | |||||
|
GABR2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 46) | NC score | 0.014648 (rank : 75) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O75899, O75974, O75975, Q5VXZ2, Q8WX04, Q9P1R2, Q9UNR1, Q9UNS9 | Gene names | GABBR2, GPR51, GPRC3B | |||
|
Domain Architecture |

|
|||||
| Description | Gamma-aminobutyric acid type B receptor, subunit 2 precursor (GABA-B receptor 2) (GABA-B-R2) (Gb2) (GABABR2) (G-protein coupled receptor 51) (HG20). | |||||
|
USF2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 47) | NC score | 0.028958 (rank : 73) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15853, O00671, O00709, Q05750, Q07952, Q15851, Q15852 | Gene names | USF2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (FOS- interacting protein) (FIP) (Major late transcription factor 2). | |||||
|
ADA29_HUMAN
|
||||||
| θ value | 5.27518 (rank : 48) | NC score | 0.004386 (rank : 81) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UKF5, Q9UHP1, Q9UKF3, Q9UKF4 | Gene names | ADAM29 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 29 precursor (A disintegrin and metalloproteinase domain 29). | |||||
|
CIZ1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 49) | NC score | 0.016654 (rank : 74) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9ULV3, Q9NYM8, Q9UHK4, Q9Y3F9, Q9Y3G0 | Gene names | CIZ1, LSFR1, NP94 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cip1-interacting zinc finger protein (Nuclear protein NP94). | |||||
|
INSR_MOUSE
|
||||||
| θ value | 5.27518 (rank : 50) | NC score | -0.001131 (rank : 83) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 968 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P15208 | Gene names | Insr | |||
|
Domain Architecture |

|
|||||
| Description | Insulin receptor precursor (EC 2.7.10.1) (IR) (CD220 antigen) [Contains: Insulin receptor subunit alpha; Insulin receptor subunit beta]. | |||||
|
ANKY1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 51) | NC score | 0.008405 (rank : 78) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9P2S6, Q8IYX5, Q8NDK5, Q9NX10 | Gene names | ANKMY1, TSAL1, ZMYND13 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin repeat and MYND domain-containing protein 1 (Testis-specific ankyrin-like protein 1) (Zinc-finger MYND domain-containing protein 13). | |||||
|
BRCA1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 52) | NC score | 0.006082 (rank : 80) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P38398 | Gene names | BRCA1, RNF53 | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer type 1 susceptibility protein (RING finger protein 53). | |||||
|
K0157_MOUSE
|
||||||
| θ value | 8.99809 (rank : 53) | NC score | 0.011495 (rank : 76) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q3TCJ1, Q3TB08, Q8K0R4 | Gene names | Kiaa0157 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0157. | |||||
|
TRPM6_MOUSE
|
||||||
| θ value | 8.99809 (rank : 54) | NC score | 0.006862 (rank : 79) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CIR4 | Gene names | Trpm6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily M member 6 (EC 2.7.11.1) (Channel kinase 2) (Melastatin-related TRP cation channel 6). | |||||
|
APBB2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 55) | NC score | 0.057252 (rank : 67) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q92870, Q8IUI6 | Gene names | APBB2, FE65L, FE65L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 2 (Fe65-like protein). | |||||
|
DRP2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 56) | NC score | 0.078995 (rank : 62) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q13474 | Gene names | DRP2 | |||
|
Domain Architecture |

|
|||||
| Description | Dystrophin-related protein 2. | |||||
|
EDD1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 57) | NC score | 0.130800 (rank : 53) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O95071, O94970, Q9NPL3 | Gene names | EDD1, EDD, HYD, KIAA0896 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase EDD1 (EC 6.3.2.-) (Hyperplastic discs protein homolog) (hHYD) (Progestin-induced protein). | |||||
|
EDD1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 58) | NC score | 0.130963 (rank : 52) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q80TP3, Q698K9, Q6PEQ8, Q6PFQ9, Q80VL4, Q810V6, Q9CXE9 | Gene names | Edd1, Edd, Kiaa0896 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase EDD1 (EC 6.3.2.-) (Hyperplastic discs protein homolog). | |||||
|
HECD1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 59) | NC score | 0.115877 (rank : 57) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 371 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9ULT8, Q6P445, Q86VJ1, Q96F34, Q9UFZ7 | Gene names | HECTD1, KIAA1131 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin-protein ligase HECTD1 (HECT domain-containing protein 1) (E3 ligase for inhibin receptor) (EULIR). | |||||
|
HECD2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 60) | NC score | 0.187316 (rank : 43) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q5U5R9, Q5VZ97, Q5VZ99, Q8TCP5 | Gene names | HECTD2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase HECTD2 (HECT domain-containing protein 2). | |||||
|
HECD2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 61) | NC score | 0.188357 (rank : 42) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8CDU6, Q8CBQ9 | Gene names | Hectd2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase HECTD2 (HECT domain-containing protein 2). | |||||
|
HECD3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 62) | NC score | 0.129792 (rank : 54) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q5T447, Q5T448, Q9H783 | Gene names | HECTD3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase HECTD3 (HECT domain-containing protein 3). | |||||
|
HECD3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 63) | NC score | 0.132098 (rank : 51) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q3U487, Q3TN76, Q641P3, Q8BQ74, Q8R1L6 | Gene names | Hectd3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase HECTD3 (HECT domain-containing protein 3). | |||||
|
HERC2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 64) | NC score | 0.122090 (rank : 56) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O95714, Q86SV7, Q86SV8, Q86SV9, Q86YY3, Q86YY4, Q86YY5, Q86YY6, Q86YY7, Q86YY8, Q86YY9, Q86YZ0, Q86YZ1 | Gene names | HERC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT domain and RCC1-like domain-containing protein 2. | |||||
|
HERC2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 65) | NC score | 0.123484 (rank : 55) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q4U2R1, O88473, Q3TRJ8, Q3TS47, Q3TST2, Q3UFQ6, Q3URH7, Q5DU32, Q7TPR5, Q80VV7, Q9QYT1, Q9Z168, Q9Z171 | Gene names | Herc2, jdf2, Kiaa0393, rjs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT domain and RCC1-like domain-containing protein 2. | |||||
|
HERC3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 66) | NC score | 0.152482 (rank : 50) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q15034 | Gene names | HERC3, KIAA0032 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT domain and RCC1-like domain protein 3. | |||||
|
HERC5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 67) | NC score | 0.156958 (rank : 49) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UII4 | Gene names | HERC5, CEB1, CEBP1 | |||
|
Domain Architecture |

|
|||||
| Description | HECT domain and RCC1-like domain-containing protein 5 (Cyclin-E- binding protein 1). | |||||
|
HUWE1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 68) | NC score | 0.191052 (rank : 41) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q7Z6Z7, O15029, Q4G2Z2, Q5H961, Q6P4D0, Q8NG67, Q9BUI0, Q9HCJ4, Q9NSL6, Q9P0A9 | Gene names | HUWE1, KIAA0312, KIAA1578, UREB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (Mcl-1 ubiquitin ligase E3) (Mule) (ARF-binding protein 1) (ARF-BP1). | |||||
|
HUWE1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 69) | NC score | 0.191372 (rank : 40) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q7TMY8, Q4G2Z1, Q5BMM7, Q6NS61, Q8BNJ7, Q8CFH2, Q8VD14, Q921M5, Q9R0P2 | Gene names | Huwe1, Kiaa0312, Ureb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (E3Histone). | |||||
|
K0317_HUMAN
|
||||||
| θ value | θ > 10 (rank : 70) | NC score | 0.197508 (rank : 38) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O15033, Q7LDY1, Q8IYY9 | Gene names | KIAA0317 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0317. | |||||
|
K0317_MOUSE
|
||||||
| θ value | θ > 10 (rank : 71) | NC score | 0.197434 (rank : 39) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8CHG5, Q6P9Q1, Q80YC4, Q8C5W5 | Gene names | Kiaa0317 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0317. | |||||
|
PRP40_HUMAN
|
||||||
| θ value | θ > 10 (rank : 72) | NC score | 0.098124 (rank : 60) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 442 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O75400, O43856, O75404, Q8TBQ1, Q9H782, Q9NWU9, Q9P0Q2, Q9Y5A8 | Gene names | PRPF40A, FLAF1, FNBP3, HYPA | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Huntingtin yeast partner A) (Huntingtin-interacting protein HYPA/FBP11) (Fas ligand-associated factor 1) (NY-REN-6 antigen). | |||||
|
PRP40_MOUSE
|
||||||
| θ value | θ > 10 (rank : 73) | NC score | 0.091606 (rank : 61) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9R1C7, Q61049, Q8BQ76, Q8BRW4, Q8C2U1 | Gene names | Prpf40a, Fbp11, Fnbp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Formin-binding protein 11) (FBP-11). | |||||
|
RHG12_HUMAN
|
||||||
| θ value | θ > 10 (rank : 74) | NC score | 0.057968 (rank : 66) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8IWW6, Q86UB3, Q8IWW7, Q8N3L1, Q9NT76 | Gene names | ARHGAP12 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 12. | |||||
|
RHG12_MOUSE
|
||||||
| θ value | θ > 10 (rank : 75) | NC score | 0.068133 (rank : 64) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8C0D4, Q8BVP8 | Gene names | Arhgap12 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 12. | |||||
|
SETD2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 76) | NC score | 0.075424 (rank : 63) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9BYW2, O75397, O75405, Q17RW8, Q5BKS9, Q5QGN2, Q69YI5, Q6IN64, Q6ZN53, Q6ZS25, Q8N3R0, Q8TCN0, Q9C0D1, Q9H696, Q9NZW9 | Gene names | SETD2, HIF1, HYPB, KIAA1732, SET2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36-specific (EC 2.1.1.43) (SET domain-containing protein 2) (hSET2) (Huntingtin- interacting protein HYPB) (Huntingtin-interacting protein 1) (HIF-1) (p231HBP). | |||||
|
TRIPC_HUMAN
|
||||||
| θ value | θ > 10 (rank : 77) | NC score | 0.168209 (rank : 48) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 319 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q14669, Q15644 | Gene names | TRIP12, KIAA0045 | |||
|
Domain Architecture |

|
|||||
| Description | Thyroid receptor-interacting protein 12 (TRIP12). | |||||
|
UBE3A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 78) | NC score | 0.182634 (rank : 44) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q05086, P78355, Q93066, Q9UEP4, Q9UEP5, Q9UEP6, Q9UEP7, Q9UEP8, Q9UEP9 | Gene names | UBE3A, E6AP | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase E3A (EC 6.3.2.-) (E6AP ubiquitin-protein ligase) (Oncogenic protein-associated protein E6-AP) (Human papillomavirus E6-associated protein) (NY-REN-54 antigen). | |||||
|
UBE3A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 79) | NC score | 0.180222 (rank : 46) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O08759, P97482 | Gene names | Ube3a | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase E3A (EC 6.3.2.-) (Oncogenic protein- associated protein E6-AP). | |||||
|
UBE3C_HUMAN
|
||||||
| θ value | θ > 10 (rank : 80) | NC score | 0.180604 (rank : 45) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q15386, Q8TC15, Q96CR4, Q9UDU3 | Gene names | UBE3C, KIAA0010 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase E3C (EC 6.3.2.-). | |||||
|
UBE3C_MOUSE
|
||||||
| θ value | θ > 10 (rank : 81) | NC score | 0.179948 (rank : 47) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q80U95, Q8BQZ6, Q8C7W6, Q8CDJ1, Q8VDL5 | Gene names | Ube3c, Kiaa0010 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase E3C (EC 6.3.2.-). | |||||
|
WBP4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 82) | NC score | 0.056232 (rank : 68) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q61048, Q8K1Z9, Q9CS45 | Gene names | Wbp4, Fbp21, Fnbp21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 4 (WBP-4) (Formin-binding protein 21). | |||||
|
WWC3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 83) | NC score | 0.059776 (rank : 65) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9ULE0, Q659C1, Q9BTQ1 | Gene names | WWC3, KIAA1280 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WWC family member 3. | |||||
|
WWTR1_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9EPK5, Q99KI4 | Gene names | Wwtr1, Taz | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing transcription regulator protein 1 (Transcriptional coactivator with PDZ-binding motif). | |||||
|
WWTR1_HUMAN
|
||||||
| NC score | 0.992829 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9GZV5, Q8N3P2, Q9Y3W6 | Gene names | WWTR1, TAZ | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing transcription regulator protein 1 (Transcriptional coactivator with PDZ-binding motif). | |||||
|
YAP1_HUMAN
|
||||||
| NC score | 0.922775 (rank : 3) | θ value | 4.36332e-79 (rank : 3) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P46937 | Gene names | YAP1, YAP65 | |||
|
Domain Architecture |

|
|||||
| Description | 65 kDa Yes-associated protein (YAP65). | |||||
|
YAP1_MOUSE
|
||||||
| NC score | 0.904695 (rank : 4) | θ value | 4.68348e-65 (rank : 4) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P46938, Q52KJ5 | Gene names | Yap1, Yap, Yap65 | |||
|
Domain Architecture |

|
|||||
| Description | 65 kDa Yes-associated protein (YAP65). | |||||
|
SAV1_HUMAN
|
||||||
| NC score | 0.467394 (rank : 5) | θ value | 0.000786445 (rank : 25) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9H4B6, Q6IA58, Q9H949, Q9HAK9 | Gene names | SAV1, WW45 | |||
|
Domain Architecture |

|
|||||
| Description | Salvador homolog 1 protein (45 kDa WW domain protein) (hWW45). | |||||
|
SAV1_MOUSE
|
||||||
| NC score | 0.466323 (rank : 6) | θ value | 0.000786445 (rank : 26) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8VEB2, Q9D9N9, Q9ER46 | Gene names | Sav1, Ww45, Wwp3 | |||
|
Domain Architecture |

|
|||||
| Description | Salvador homolog 1 protein (45 kDa WW domain protein) (mWW45). | |||||
|
WWC1_HUMAN
|
||||||
| NC score | 0.392885 (rank : 7) | θ value | 0.00035302 (rank : 23) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8IX03, O94946, Q6MZX4, Q6Y2F8, Q7Z4G8, Q8WVM4, Q9BT29 | Gene names | WWC1, KIAA0869 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing protein 1 (Kidney and brain protein) (KIBRA) (HBeAg-binding protein 3). | |||||
|
WWC1_MOUSE
|
||||||
| NC score | 0.388151 (rank : 8) | θ value | 0.000602161 (rank : 24) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q5SXA9, Q571D0, Q8K1Y3, Q8VD17, Q922W3 | Gene names | Wwc1, Kiaa0869 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing protein 1 (Kidney and brain protein) (KIBRA). | |||||
|
WWP1_MOUSE
|
||||||
| NC score | 0.387002 (rank : 9) | θ value | 7.1131e-05 (rank : 15) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8BZZ3, Q8BIV9, Q8VDP8 | Gene names | Wwp1 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP1 (EC 6.3.2.-) (WW domain- containing protein 1). | |||||
|
WWP2_MOUSE
|
||||||
| NC score | 0.383664 (rank : 10) | θ value | 3.19293e-05 (rank : 8) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9DBH0, Q8BTG4, Q923F6 | Gene names | Wwp2 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP2 (EC 6.3.2.-) (WW domain- containing protein 2). | |||||
|
WWP2_HUMAN
|
||||||
| NC score | 0.382767 (rank : 11) | θ value | 3.19293e-05 (rank : 7) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O00308, Q96CZ2, Q9BWN6 | Gene names | WWP2 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP2 (EC 6.3.2.-) (WW domain- containing protein 2) (Atrophin-1-interacting protein 2) (AIP2). | |||||
|
ITCH_MOUSE
|
||||||
| NC score | 0.381258 (rank : 12) | θ value | 3.19293e-05 (rank : 6) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8C863, O54971 | Gene names | Itch | |||
|
Domain Architecture |

|
|||||
| Description | Itchy E3 ubiquitin protein ligase (EC 6.3.2.-). | |||||
|
WWP1_HUMAN
|
||||||
| NC score | 0.379666 (rank : 13) | θ value | 7.1131e-05 (rank : 14) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9H0M0, O00307, Q96BP4 | Gene names | WWP1 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP1 (EC 6.3.2.-) (WW domain- containing protein 1) (Atropin-1-interacting protein 5) (AIP5). | |||||
|
ITCH_HUMAN
|
||||||
| NC score | 0.379304 (rank : 14) | θ value | 3.19293e-05 (rank : 5) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q96J02, O43584, Q5TEL0, Q96F66, Q9BY75, Q9H451, Q9H4U5 | Gene names | ITCH | |||
|
Domain Architecture |

|
|||||
| Description | Itchy homolog E3 ubiquitin protein ligase (EC 6.3.2.-) (Itch) (Atrophin-1-interacting protein 4) (AIP4) (NFE2-associated polypeptide 1) (NAPP1). | |||||
|
SMUF2_HUMAN
|
||||||
| NC score | 0.367195 (rank : 15) | θ value | 4.1701e-05 (rank : 10) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9HAU4, Q9H260 | Gene names | SMURF2 | |||
|
Domain Architecture |

|
|||||
| Description | Smad ubiquitination regulatory factor 2 (EC 6.3.2.-) (Ubiquitin-- protein ligase SMURF2) (Smad-specific E3 ubiquitin ligase 2) (hSMURF2). | |||||
|
SMUF1_MOUSE
|
||||||
| NC score | 0.361418 (rank : 16) | θ value | 0.000158464 (rank : 21) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9CUN6, Q3U412, Q8K300 | Gene names | Smurf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Smad ubiquitination regulatory factor 1 (EC 6.3.2.-) (Ubiquitin-- protein ligase SMURF1) (Smad-specific E3 ubiquitin ligase 1). | |||||
|
BAG3_HUMAN
|
||||||
| NC score | 0.361164 (rank : 17) | θ value | 0.000270298 (rank : 22) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 314 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O95817, Q9NT20, Q9P120 | Gene names | BAG3, BIS | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 3 (BCL-2-binding athanogene- 3) (BAG-3) (Bcl-2-binding protein Bis) (Docking protein CAIR-1). | |||||
|
SMUF1_HUMAN
|
||||||
| NC score | 0.357175 (rank : 18) | θ value | 0.000158464 (rank : 20) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9HCE7, O75853, Q9UJT8 | Gene names | SMURF1, KIAA1625 | |||
|
Domain Architecture |

|
|||||
| Description | Smad ubiquitination regulatory factor 1 (EC 6.3.2.-) (Ubiquitin-- protein ligase SMURF1) (Smad-specific E3 ubiquitin ligase 1) (hSMURF1). | |||||
|
NEDD4_HUMAN
|
||||||
| NC score | 0.355080 (rank : 19) | θ value | 5.44631e-05 (rank : 11) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P46934 | Gene names | NEDD4, KIAA0093 | |||
|
Domain Architecture |

|
|||||
| Description | E3 ubiquitin-protein ligase NEDD4 (EC 6.3.2.-). | |||||
|
NED4L_MOUSE
|
||||||
| NC score | 0.354024 (rank : 20) | θ value | 7.1131e-05 (rank : 13) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8CFI0, Q8BRT9, Q8BS42, Q99PK2 | Gene names | Nedd4l, Kiaa0439, Nedd4b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin-protein ligase NEDD4-like protein (EC 6.3.2.-) (Nedd4-2) (NEDD4.2). | |||||
|
NEDD4_MOUSE
|
||||||
| NC score | 0.353588 (rank : 21) | θ value | 4.1701e-05 (rank : 9) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P46935, O08758, Q3UZI2, Q8BGB3 | Gene names | Nedd4, Kiaa0093, Nedd-4, Nedd4a | |||
|
Domain Architecture |

|
|||||
| Description | E3 ubiquitin-protein ligase NEDD4 (EC 6.3.2.-) (Neural precursor cell expressed developmentally down-regulated protein 4). | |||||
|
NED4L_HUMAN
|
||||||
| NC score | 0.352715 (rank : 22) | θ value | 7.1131e-05 (rank : 12) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q96PU5, O43165, Q7Z5F1, Q7Z5F2, Q7Z5N3, Q8N5A7, Q8WUU9, Q9BW58, Q9H2W4, Q9NT88 | Gene names | NEDD4L, KIAA0439, NEDL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin-protein ligase NEDD4-like protein (EC 6.3.2.-) (Nedd4-2) (NEDD4.2). | |||||
|
PINL_HUMAN
|
||||||
| NC score | 0.351807 (rank : 23) | θ value | 0.0961366 (rank : 37) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O15428 | Gene names | PIN1L | |||
|
Domain Architecture |

|
|||||
| Description | PIN1-like protein. | |||||
|
WWOX_HUMAN
|
||||||
| NC score | 0.337108 (rank : 24) | θ value | 0.00102713 (rank : 29) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9NZC7, Q5MYT5, Q96KM3, Q96RF2, Q9BTT8, Q9NPC9, Q9NRF4, Q9NRF5, Q9NRF6, Q9NRK1, Q9NZC5 | Gene names | WWOX, FOR, WOX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing oxidoreductase (EC 1.1.1.-) (Fragile site FRA16D oxidoreductase). | |||||
|
WWOX_MOUSE
|
||||||
| NC score | 0.334222 (rank : 25) | θ value | 0.00102713 (rank : 30) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q91WL8, Q8C8J6, Q920Y2, Q9D2B3, Q9D339, Q9JLF5 | Gene names | Wwox, Wox1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing oxidoreductase (EC 1.1.1.-). | |||||
|
PIN1_MOUSE
|
||||||
| NC score | 0.331726 (rank : 26) | θ value | 0.0431538 (rank : 35) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9QUR7, Q543B3 | Gene names | Pin1 | |||
|
Domain Architecture |

|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase NIMA-interacting 1 (EC 5.2.1.8) (Rotamase Pin1) (PPIase Pin1). | |||||
|
PIN1_HUMAN
|
||||||
| NC score | 0.326469 (rank : 27) | θ value | 0.0431538 (rank : 34) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q13526 | Gene names | PIN1 | |||
|
Domain Architecture |

|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase NIMA-interacting 1 (EC 5.2.1.8) (Rotamase Pin1) (PPIase Pin1). | |||||
|
BAG3_MOUSE
|
||||||
| NC score | 0.319069 (rank : 28) | θ value | 0.00102713 (rank : 27) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9JLV1, Q9JJC7 | Gene names | Bag3, Bis | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 3 (BCL-2-binding athanogene- 3) (BAG-3) (Bcl-2-binding protein Bis). | |||||
|
MAGI2_MOUSE
|
||||||
| NC score | 0.313962 (rank : 29) | θ value | 9.29e-05 (rank : 18) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 370 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9WVQ1, Q3UH81, Q6GT88, Q8BYT1, Q8CA85 | Gene names | Magi2, Acvrinp1, Aip1, Arip1 | |||
|
Domain Architecture |

|
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 2 (Membrane-associated guanylate kinase inverted 2) (MAGI-2) (Atrophin-1-interacting protein 1) (Activin receptor-interacting protein 1) (Acvrip1). | |||||
|
MAGI2_HUMAN
|
||||||
| NC score | 0.306625 (rank : 30) | θ value | 0.000158464 (rank : 19) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 373 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q86UL8, O60434, O60510, Q86UI7, Q9NP44, Q9UDQ5, Q9UDU1 | Gene names | MAGI2, ACVRINP1, AIP1, KIAA0705 | |||
|
Domain Architecture |

|
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 2 (Membrane-associated guanylate kinase inverted 2) (MAGI-2) (Atrophin-1-interacting protein 1) (Atrophin-1-interacting protein A). | |||||
|
MAGI1_MOUSE
|
||||||
| NC score | 0.296441 (rank : 31) | θ value | 9.29e-05 (rank : 17) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q6RHR9, O54893, O54894, O54895 | Gene names | Magi1, Baiap1, Bap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 1 (BAI1-associated protein 1) (BAP-1) (Membrane-associated guanylate kinase inverted 1) (MAGI-1). | |||||
|
MAGI1_HUMAN
|
||||||
| NC score | 0.296401 (rank : 32) | θ value | 9.29e-05 (rank : 16) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q96QZ7, O00309, O43863, O75085, Q96QZ8, Q96QZ9 | Gene names | MAGI1, BAIAP1, BAP1, TNRC19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 1 (BAI1-associated protein 1) (BAP-1) (Membrane-associated guanylate kinase inverted 1) (MAGI-1) (Atrophin-1-interacting protein 3) (AIP3) (WW domain-containing protein 3) (WWP3) (Trinucleotide repeat-containing gene 19 protein). | |||||
|
PKHA5_HUMAN
|
||||||
| NC score | 0.246808 (rank : 33) | θ value | 0.00102713 (rank : 28) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9HAU0, Q8N3K6, Q96DY9, Q9BVR4, Q9C0H7, Q9H924, Q9NVK8 | Gene names | PLEKHA5, KIAA1686, PEPP2 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 5 (Phosphoinositol 3-phosphate-binding protein 2) (PEPP-2). | |||||
|
GAS7_MOUSE
|
||||||
| NC score | 0.244379 (rank : 34) | θ value | 0.0330416 (rank : 33) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q60780, Q9QY25, Q9QY26, Q9QY34 | Gene names | Gas7 | |||
|
Domain Architecture |

|
|||||
| Description | Growth-arrest-specific protein 7 (GAS-7). | |||||
|
GAS7_HUMAN
|
||||||
| NC score | 0.230005 (rank : 35) | θ value | 0.0736092 (rank : 36) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O60861, O43144 | Gene names | GAS7, KIAA0394 | |||
|
Domain Architecture |

|
|||||
| Description | Growth-arrest-specific protein 7 (GAS-7). | |||||
|
STXB4_MOUSE
|
||||||
| NC score | 0.225954 (rank : 36) | θ value | 0.00298849 (rank : 32) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 629 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9WV89, Q5SUA6, Q5SUA8, Q5SUA9, Q8CFL1 | Gene names | Stxbp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntaxin-binding protein 4 (Syntaxin 4-interacting protein) (Synip). | |||||
|
STXB4_HUMAN
|
||||||
| NC score | 0.222685 (rank : 37) | θ value | 0.00298849 (rank : 31) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 616 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q6ZWJ1, Q8IVZ5 | Gene names | STXBP4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntaxin-binding protein 4 (Syntaxin 4-interacting protein) (Synip). | |||||
|
K0317_HUMAN
|
||||||
| NC score | 0.197508 (rank : 38) | θ value | θ > 10 (rank : 70) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O15033, Q7LDY1, Q8IYY9 | Gene names | KIAA0317 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0317. | |||||
|
K0317_MOUSE
|
||||||
| NC score | 0.197434 (rank : 39) | θ value | θ > 10 (rank : 71) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8CHG5, Q6P9Q1, Q80YC4, Q8C5W5 | Gene names | Kiaa0317 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0317. | |||||
|
HUWE1_MOUSE
|
||||||
| NC score | 0.191372 (rank : 40) | θ value | θ > 10 (rank : 69) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q7TMY8, Q4G2Z1, Q5BMM7, Q6NS61, Q8BNJ7, Q8CFH2, Q8VD14, Q921M5, Q9R0P2 | Gene names | Huwe1, Kiaa0312, Ureb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (E3Histone). | |||||
|
HUWE1_HUMAN
|
||||||
| NC score | 0.191052 (rank : 41) | θ value | θ > 10 (rank : 68) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q7Z6Z7, O15029, Q4G2Z2, Q5H961, Q6P4D0, Q8NG67, Q9BUI0, Q9HCJ4, Q9NSL6, Q9P0A9 | Gene names | HUWE1, KIAA0312, KIAA1578, UREB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (Mcl-1 ubiquitin ligase E3) (Mule) (ARF-binding protein 1) (ARF-BP1). | |||||
|
HECD2_MOUSE
|
||||||
| NC score | 0.188357 (rank : 42) | θ value | θ > 10 (rank : 61) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8CDU6, Q8CBQ9 | Gene names | Hectd2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase HECTD2 (HECT domain-containing protein 2). | |||||
|
HECD2_HUMAN
|
||||||
| NC score | 0.187316 (rank : 43) | θ value | θ > 10 (rank : 60) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q5U5R9, Q5VZ97, Q5VZ99, Q8TCP5 | Gene names | HECTD2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase HECTD2 (HECT domain-containing protein 2). | |||||
|
UBE3A_HUMAN
|
||||||
| NC score | 0.182634 (rank : 44) | θ value | θ > 10 (rank : 78) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q05086, P78355, Q93066, Q9UEP4, Q9UEP5, Q9UEP6, Q9UEP7, Q9UEP8, Q9UEP9 | Gene names | UBE3A, E6AP | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase E3A (EC 6.3.2.-) (E6AP ubiquitin-protein ligase) (Oncogenic protein-associated protein E6-AP) (Human papillomavirus E6-associated protein) (NY-REN-54 antigen). | |||||
|
UBE3C_HUMAN
|
||||||
| NC score | 0.180604 (rank : 45) | θ value | θ > 10 (rank : 80) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q15386, Q8TC15, Q96CR4, Q9UDU3 | Gene names | UBE3C, KIAA0010 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase E3C (EC 6.3.2.-). | |||||
|
UBE3A_MOUSE
|
||||||
| NC score | 0.180222 (rank : 46) | θ value | θ > 10 (rank : 79) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O08759, P97482 | Gene names | Ube3a | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase E3A (EC 6.3.2.-) (Oncogenic protein- associated protein E6-AP). | |||||
|
UBE3C_MOUSE
|
||||||
| NC score | 0.179948 (rank : 47) | θ value | θ > 10 (rank : 81) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q80U95, Q8BQZ6, Q8C7W6, Q8CDJ1, Q8VDL5 | Gene names | Ube3c, Kiaa0010 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase E3C (EC 6.3.2.-). | |||||
|
TRIPC_HUMAN
|
||||||
| NC score | 0.168209 (rank : 48) | θ value | θ > 10 (rank : 77) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 319 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q14669, Q15644 | Gene names | TRIP12, KIAA0045 | |||
|
Domain Architecture |

|
|||||
| Description | Thyroid receptor-interacting protein 12 (TRIP12). | |||||
|
HERC5_HUMAN
|
||||||
| NC score | 0.156958 (rank : 49) | θ value | θ > 10 (rank : 67) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UII4 | Gene names | HERC5, CEB1, CEBP1 | |||
|
Domain Architecture |

|
|||||
| Description | HECT domain and RCC1-like domain-containing protein 5 (Cyclin-E- binding protein 1). | |||||
|
HERC3_HUMAN
|
||||||
| NC score | 0.152482 (rank : 50) | θ value | θ > 10 (rank : 66) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q15034 | Gene names | HERC3, KIAA0032 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT domain and RCC1-like domain protein 3. | |||||
|
HECD3_MOUSE
|
||||||
| NC score | 0.132098 (rank : 51) | θ value | θ > 10 (rank : 63) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q3U487, Q3TN76, Q641P3, Q8BQ74, Q8R1L6 | Gene names | Hectd3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase HECTD3 (HECT domain-containing protein 3). | |||||
|
EDD1_MOUSE
|
||||||
| NC score | 0.130963 (rank : 52) | θ value | θ > 10 (rank : 58) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q80TP3, Q698K9, Q6PEQ8, Q6PFQ9, Q80VL4, Q810V6, Q9CXE9 | Gene names | Edd1, Edd, Kiaa0896 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase EDD1 (EC 6.3.2.-) (Hyperplastic discs protein homolog). | |||||
|
EDD1_HUMAN
|
||||||
| NC score | 0.130800 (rank : 53) | θ value | θ > 10 (rank : 57) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O95071, O94970, Q9NPL3 | Gene names | EDD1, EDD, HYD, KIAA0896 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase EDD1 (EC 6.3.2.-) (Hyperplastic discs protein homolog) (hHYD) (Progestin-induced protein). | |||||
|
HECD3_HUMAN
|
||||||
| NC score | 0.129792 (rank : 54) | θ value | θ > 10 (rank : 62) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q5T447, Q5T448, Q9H783 | Gene names | HECTD3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase HECTD3 (HECT domain-containing protein 3). | |||||
|
HERC2_MOUSE
|
||||||
| NC score | 0.123484 (rank : 55) | θ value | θ > 10 (rank : 65) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q4U2R1, O88473, Q3TRJ8, Q3TS47, Q3TST2, Q3UFQ6, Q3URH7, Q5DU32, Q7TPR5, Q80VV7, Q9QYT1, Q9Z168, Q9Z171 | Gene names | Herc2, jdf2, Kiaa0393, rjs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT domain and RCC1-like domain-containing protein 2. | |||||
|
HERC2_HUMAN
|
||||||
| NC score | 0.122090 (rank : 56) | θ value | θ > 10 (rank : 64) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O95714, Q86SV7, Q86SV8, Q86SV9, Q86YY3, Q86YY4, Q86YY5, Q86YY6, Q86YY7, Q86YY8, Q86YY9, Q86YZ0, Q86YZ1 | Gene names | HERC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT domain and RCC1-like domain-containing protein 2. | |||||
|
HECD1_HUMAN
|
||||||
| NC score | 0.115877 (rank : 57) | θ value | θ > 10 (rank : 59) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 371 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9ULT8, Q6P445, Q86VJ1, Q96F34, Q9UFZ7 | Gene names | HECTD1, KIAA1131 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin-protein ligase HECTD1 (HECT domain-containing protein 1) (E3 ligase for inhibin receptor) (EULIR). | |||||
|
DGCR8_HUMAN
|
||||||
| NC score | 0.111621 (rank : 58) | θ value | 0.47712 (rank : 38) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8WYQ5, Q6DCB2, Q6MZE9, Q6Y2L0, Q96G39, Q96GP8, Q9H6L8, Q9H6T7, Q9NRW2 | Gene names | DGCR8, C22orf12, DGCRK6 | |||
|
Domain Architecture |

|
|||||
| Description | DGCR8 protein (DiGeorge syndrome critical region 8). | |||||
|
DGCR8_MOUSE
|
||||||
| NC score | 0.110836 (rank : 59) | θ value | 0.47712 (rank : 39) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9EQM6 | Gene names | Dgcr8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DGCR8 protein (DiGeorge syndrome critical region 8 homolog) (Gy1). | |||||
|
PRP40_HUMAN
|
||||||
| NC score | 0.098124 (rank : 60) | θ value | θ > 10 (rank : 72) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 442 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O75400, O43856, O75404, Q8TBQ1, Q9H782, Q9NWU9, Q9P0Q2, Q9Y5A8 | Gene names | PRPF40A, FLAF1, FNBP3, HYPA | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Huntingtin yeast partner A) (Huntingtin-interacting protein HYPA/FBP11) (Fas ligand-associated factor 1) (NY-REN-6 antigen). | |||||
|
PRP40_MOUSE
|
||||||
| NC score | 0.091606 (rank : 61) | θ value | θ > 10 (rank : 73) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9R1C7, Q61049, Q8BQ76, Q8BRW4, Q8C2U1 | Gene names | Prpf40a, Fbp11, Fnbp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Formin-binding protein 11) (FBP-11). | |||||
|
DRP2_HUMAN
|
||||||
| NC score | 0.078995 (rank : 62) | θ value | θ > 10 (rank : 56) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q13474 | Gene names | DRP2 | |||
|
Domain Architecture |

|
|||||
| Description | Dystrophin-related protein 2. | |||||
|
SETD2_HUMAN
|
||||||
| NC score | 0.075424 (rank : 63) | θ value | θ > 10 (rank : 76) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9BYW2, O75397, O75405, Q17RW8, Q5BKS9, Q5QGN2, Q69YI5, Q6IN64, Q6ZN53, Q6ZS25, Q8N3R0, Q8TCN0, Q9C0D1, Q9H696, Q9NZW9 | Gene names | SETD2, HIF1, HYPB, KIAA1732, SET2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36-specific (EC 2.1.1.43) (SET domain-containing protein 2) (hSET2) (Huntingtin- interacting protein HYPB) (Huntingtin-interacting protein 1) (HIF-1) (p231HBP). | |||||
|
RHG12_MOUSE
|
||||||
| NC score | 0.068133 (rank : 64) | θ value | θ > 10 (rank : 75) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8C0D4, Q8BVP8 | Gene names | Arhgap12 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 12. | |||||
|
WWC3_HUMAN
|
||||||
| NC score | 0.059776 (rank : 65) | θ value | θ > 10 (rank : 83) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9ULE0, Q659C1, Q9BTQ1 | Gene names | WWC3, KIAA1280 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WWC family member 3. | |||||
|
RHG12_HUMAN
|
||||||
| NC score | 0.057968 (rank : 66) | θ value | θ > 10 (rank : 74) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8IWW6, Q86UB3, Q8IWW7, Q8N3L1, Q9NT76 | Gene names | ARHGAP12 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 12. | |||||
|
APBB2_HUMAN
|
||||||
| NC score | 0.057252 (rank : 67) | θ value | θ > 10 (rank : 55) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q92870, Q8IUI6 | Gene names | APBB2, FE65L, FE65L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 2 (Fe65-like protein). | |||||
|
WBP4_MOUSE
|
||||||
| NC score | 0.056232 (rank : 68) | θ value | θ > 10 (rank : 82) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q61048, Q8K1Z9, Q9CS45 | Gene names | Wbp4, Fbp21, Fnbp21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 4 (WBP-4) (Formin-binding protein 21). | |||||
|
BTBDC_HUMAN
|
||||||
| NC score | 0.049062 (rank : 69) | θ value | 1.38821 (rank : 42) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8IY92, Q96JP1 | Gene names | BTBD12, KIAA1784 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein 12. | |||||
|
DMD_HUMAN
|
||||||
| NC score | 0.045075 (rank : 70) | θ value | 0.813845 (rank : 40) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P11532, Q02295, Q14169, Q14170, Q5JYU0, Q7KZ48 | Gene names | DMD | |||
|
Domain Architecture |

|
|||||
| Description | Dystrophin. | |||||
|
DMD_MOUSE
|
||||||
| NC score | 0.042302 (rank : 71) | θ value | 0.813845 (rank : 41) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1111 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P11531, O35653, Q60703 | Gene names | Dmd | |||
|
Domain Architecture |

|
|||||
| Description | Dystrophin. | |||||
|
USF2_MOUSE
|
||||||
| NC score | 0.029898 (rank : 72) | θ value | 3.0926 (rank : 45) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q64705, Q3UIL2, Q8VE59 | Gene names | Usf2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (Major late transcription factor 2). | |||||
|
USF2_HUMAN
|
||||||
| NC score | 0.028958 (rank : 73) | θ value | 4.03905 (rank : 47) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15853, O00671, O00709, Q05750, Q07952, Q15851, Q15852 | Gene names | USF2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (FOS- interacting protein) (FIP) (Major late transcription factor 2). | |||||
|
CIZ1_HUMAN
|
||||||
| NC score | 0.016654 (rank : 74) | θ value | 5.27518 (rank : 49) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9ULV3, Q9NYM8, Q9UHK4, Q9Y3F9, Q9Y3G0 | Gene names | CIZ1, LSFR1, NP94 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cip1-interacting zinc finger protein (Nuclear protein NP94). | |||||
|
GABR2_HUMAN
|
||||||
| NC score | 0.014648 (rank : 75) | θ value | 4.03905 (rank : 46) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O75899, O75974, O75975, Q5VXZ2, Q8WX04, Q9P1R2, Q9UNR1, Q9UNS9 | Gene names | GABBR2, GPR51, GPRC3B | |||
|
Domain Architecture |

|
|||||
| Description | Gamma-aminobutyric acid type B receptor, subunit 2 precursor (GABA-B receptor 2) (GABA-B-R2) (Gb2) (GABABR2) (G-protein coupled receptor 51) (HG20). | |||||
|
K0157_MOUSE
|
||||||
| NC score | 0.011495 (rank : 76) | θ value | 8.99809 (rank : 53) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q3TCJ1, Q3TB08, Q8K0R4 | Gene names | Kiaa0157 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0157. | |||||
|
MYH7_HUMAN
|
||||||
| NC score | 0.008653 (rank : 77) | θ value | 3.0926 (rank : 44) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P12883, Q14836, Q14837, Q14904, Q16579, Q92679, Q9H1D5, Q9UMM8 | Gene names | MYH7, MYHCB | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle beta isoform (MyHC-beta). | |||||
|
ANKY1_HUMAN
|
||||||
| NC score | 0.008405 (rank : 78) | θ value | 8.99809 (rank : 51) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9P2S6, Q8IYX5, Q8NDK5, Q9NX10 | Gene names | ANKMY1, TSAL1, ZMYND13 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin repeat and MYND domain-containing protein 1 (Testis-specific ankyrin-like protein 1) (Zinc-finger MYND domain-containing protein 13). | |||||
|
TRPM6_MOUSE
|
||||||
| NC score | 0.006862 (rank : 79) | θ value | 8.99809 (rank : 54) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CIR4 | Gene names | Trpm6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transient receptor potential cation channel subfamily M member 6 (EC 2.7.11.1) (Channel kinase 2) (Melastatin-related TRP cation channel 6). | |||||
|
BRCA1_HUMAN
|
||||||
| NC score | 0.006082 (rank : 80) | θ value | 8.99809 (rank : 52) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P38398 | Gene names | BRCA1, RNF53 | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer type 1 susceptibility protein (RING finger protein 53). | |||||
|
ADA29_HUMAN
|
||||||
| NC score | 0.004386 (rank : 81) | θ value | 5.27518 (rank : 48) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UKF5, Q9UHP1, Q9UKF3, Q9UKF4 | Gene names | ADAM29 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 29 precursor (A disintegrin and metalloproteinase domain 29). | |||||
|
TLK2_MOUSE
|
||||||
| NC score | 0.003410 (rank : 82) | θ value | 1.81305 (rank : 43) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 1088 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O55047, Q9D5Y5 | Gene names | Tlk2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase tousled-like 2 (EC 2.7.11.1) (Tousled- like kinase 2). | |||||
|
INSR_MOUSE
|
||||||
| NC score | -0.001131 (rank : 83) | θ value | 5.27518 (rank : 50) | |||
| Query Neighborhood Hits | 54 | Target Neighborhood Hits | 968 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P15208 | Gene names | Insr | |||
|
Domain Architecture |

|
|||||
| Description | Insulin receptor precursor (EC 2.7.10.1) (IR) (CD220 antigen) [Contains: Insulin receptor subunit alpha; Insulin receptor subunit beta]. | |||||