Please be patient as the page loads
|
MLH3_HUMAN
|
||||||
| SwissProt Accessions | Q9UHC1, P49751, Q56DK9, Q9P292, Q9UHC0 | Gene names | MLH3 | |||
|
Domain Architecture |

|
|||||
| Description | DNA mismatch repair protein Mlh3 (MutL protein homolog 3). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
MLH3_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9UHC1, P49751, Q56DK9, Q9P292, Q9UHC0 | Gene names | MLH3 | |||
|
Domain Architecture |

|
|||||
| Description | DNA mismatch repair protein Mlh3 (MutL protein homolog 3). | |||||
|
MLH1_HUMAN
|
||||||
| θ value | 3.07116e-24 (rank : 2) | NC score | 0.651832 (rank : 2) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P40692 | Gene names | MLH1, COCA2 | |||
|
Domain Architecture |

|
|||||
| Description | DNA mismatch repair protein Mlh1 (MutL protein homolog 1). | |||||
|
MLH1_MOUSE
|
||||||
| θ value | 1.29031e-22 (rank : 3) | NC score | 0.644750 (rank : 3) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9JK91, Q62454 | Gene names | Mlh1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA mismatch repair protein Mlh1 (MutL protein homolog 1). | |||||
|
PMS1_HUMAN
|
||||||
| θ value | 6.40375e-22 (rank : 4) | NC score | 0.589997 (rank : 4) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P54277 | Gene names | PMS1, PMSL1 | |||
|
Domain Architecture |

|
|||||
| Description | PMS1 protein homolog 1 (DNA mismatch repair protein PMS1). | |||||
|
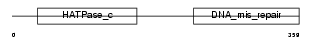
PMS2_HUMAN
|
||||||
| θ value | 6.85773e-16 (rank : 5) | NC score | 0.568280 (rank : 5) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P54278 | Gene names | PMS2, PMSL2 | |||
|
Domain Architecture |
|
|||||
| Description | PMS1 protein homolog 2 (DNA mismatch repair protein PMS2). | |||||
|
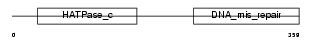
PMS2_MOUSE
|
||||||
| θ value | 2.88119e-14 (rank : 6) | NC score | 0.567305 (rank : 6) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P54279 | Gene names | Pms2 | |||
|
Domain Architecture |
|
|||||
| Description | PMS1 protein homolog 2 (DNA mismatch repair protein PMS2). | |||||
|
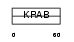
ZN514_HUMAN
|
||||||
| θ value | 0.365318 (rank : 7) | NC score | 0.003607 (rank : 42) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 738 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96K75 | Gene names | ZNF514 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger protein 514. | |||||
|
SETX_HUMAN
|
||||||
| θ value | 0.47712 (rank : 8) | NC score | 0.046369 (rank : 8) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q7Z333, O75120, Q3KQX4, Q68DW5, Q6AZD7, Q7Z3J6, Q8WX33, Q9H9D1, Q9NVP9 | Gene names | SETX, ALS4, KIAA0625, SCAR1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable helicase senataxin (EC 3.6.1.-) (SEN1 homolog). | |||||
|
TPR_HUMAN
|
||||||
| θ value | 0.47712 (rank : 9) | NC score | 0.018430 (rank : 23) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
DMP1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 10) | NC score | 0.046520 (rank : 7) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q13316, O43265 | Gene names | DMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1). | |||||
|
PREP_MOUSE
|
||||||
| θ value | 1.06291 (rank : 11) | NC score | 0.042912 (rank : 9) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8K411, Q3THC4, Q3TJI1, Q3TPP0, Q3URQ8, Q4KUG2, Q4KUG3, Q6ZPX6, Q922N1, Q9CV63 | Gene names | Pitrm1, Kiaa1104, Ntup1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Presequence protease, mitochondrial precursor (EC 3.4.24.-) (Pitrilysin metalloproteinase 1). | |||||
|
WWP1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 12) | NC score | 0.024995 (rank : 14) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9H0M0, O00307, Q96BP4 | Gene names | WWP1 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP1 (EC 6.3.2.-) (WW domain- containing protein 1) (Atropin-1-interacting protein 5) (AIP5). | |||||
|
WWP1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 13) | NC score | 0.024248 (rank : 16) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BZZ3, Q8BIV9, Q8VDP8 | Gene names | Wwp1 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP1 (EC 6.3.2.-) (WW domain- containing protein 1). | |||||
|
CSPG2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 14) | NC score | 0.014640 (rank : 28) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P13611, P20754, Q13010, Q13189, Q15123, Q9UCL9, Q9UNW5 | Gene names | CSPG2 | |||
|
Domain Architecture |

|
|||||
| Description | Versican core protein precursor (Large fibroblast proteoglycan) (Chondroitin sulfate proteoglycan core protein 2) (PG-M) (Glial hyaluronate-binding protein) (GHAP). | |||||
|
UBP31_HUMAN
|
||||||
| θ value | 1.38821 (rank : 15) | NC score | 0.023635 (rank : 18) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 293 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q70CQ4, Q6AW97, Q6ZTC0, Q6ZTN2, Q9ULL7 | Gene names | USP31, KIAA1203 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 31 (EC 3.1.2.15) (Ubiquitin thioesterase 31) (Ubiquitin-specific-processing protease 31) (Deubiquitinating enzyme 31). | |||||
|
CT2NL_HUMAN
|
||||||
| θ value | 1.81305 (rank : 16) | NC score | 0.024870 (rank : 15) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 1059 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9P2B4, Q96B40 | Gene names | CTTNBP2NL, KIAA1433 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CTTNBP2 N-terminal-like protein. | |||||
|
DIAP3_MOUSE
|
||||||
| θ value | 1.81305 (rank : 17) | NC score | 0.021270 (rank : 20) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Z207 | Gene names | Diaph3, Diap3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 3 (Diaphanous-related formin-3) (DRF3) (mDIA2) (p134mDIA2). | |||||
|
CIR_MOUSE
|
||||||
| θ value | 2.36792 (rank : 18) | NC score | 0.034583 (rank : 10) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9DA19, Q3V2N9, Q4KL44, Q52KL9, Q5FW66 | Gene names | Cir | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CBF1-interacting corepressor. | |||||
|
OPTN_MOUSE
|
||||||
| θ value | 3.0926 (rank : 19) | NC score | 0.014477 (rank : 29) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 1120 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8K3K8 | Gene names | Optn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Optineurin. | |||||
|
PDZK4_HUMAN
|
||||||
| θ value | 3.0926 (rank : 20) | NC score | 0.019216 (rank : 22) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q76G19, Q8NB75, Q9BUH9, Q9P284 | Gene names | PDZK4, KIAA1444, PDZRN4L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 4 (PDZ domain-containing RING finger protein 4-like protein). | |||||
|
PML_MOUSE
|
||||||
| θ value | 3.0926 (rank : 21) | NC score | 0.023060 (rank : 19) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q60953 | Gene names | Pml | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable transcription factor PML. | |||||
|
CT2NL_MOUSE
|
||||||
| θ value | 4.03905 (rank : 22) | NC score | 0.025501 (rank : 13) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 902 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q99LJ0, Q6ZPR0, Q8BSV1 | Gene names | Cttnbp2nl, Kiaa1433 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CTTNBP2 N-terminal-like protein. | |||||
|
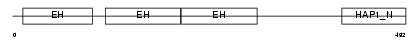
EP15_HUMAN
|
||||||
| θ value | 4.03905 (rank : 23) | NC score | 0.015498 (rank : 25) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 1021 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P42566 | Gene names | EPS15, AF1P | |||
|
Domain Architecture |
|
|||||
| Description | Epidermal growth factor receptor substrate 15 (Protein Eps15) (AF-1p protein). | |||||
|
MYH3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 24) | NC score | 0.008172 (rank : 40) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 1660 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P11055, Q15492 | Gene names | MYH3 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, fast skeletal muscle, embryonic (Muscle embryonic myosin heavy chain) (SMHCE). | |||||
|
PALB2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 25) | NC score | 0.026015 (rank : 12) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q86YC2, Q8N7Y6, Q8ND31, Q9H6W1 | Gene names | PALB2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Partner and localizer of BRCA2. | |||||
|
ZN318_MOUSE
|
||||||
| θ value | 4.03905 (rank : 26) | NC score | 0.023811 (rank : 17) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 642 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q99PP2, Q3TZL5, Q9JJ01 | Gene names | Znf318, Tzf, Zfp318 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 318 (Testicular zinc finger protein). | |||||
|
BRCA2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 27) | NC score | 0.020352 (rank : 21) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P51587, O00183, O15008, Q13879 | Gene names | BRCA2, FANCD1 | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer type 2 susceptibility protein (Fanconi anemia group D1 protein). | |||||
|
FGD3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 28) | NC score | 0.009609 (rank : 38) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5JSP0, Q7Z7D9, Q8N5G1 | Gene names | FGD3, ZFYVE5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 3 (Zinc finger FYVE domain-containing protein 5). | |||||
|
STAM2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 29) | NC score | 0.011492 (rank : 32) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O88811, Q8C8Y4 | Gene names | Stam2, Hbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal transducing adapter molecule 2 (STAM-2) (Hrs-binding protein). | |||||
|
DPOLZ_HUMAN
|
||||||
| θ value | 6.88961 (rank : 30) | NC score | 0.014739 (rank : 26) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O60673, O43214 | Gene names | REV3L, POLZ, REV3 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase zeta catalytic subunit (EC 2.7.7.7) (hREV3). | |||||
|
K0408_HUMAN
|
||||||
| θ value | 6.88961 (rank : 31) | NC score | 0.027307 (rank : 11) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6ZU52, O43158, Q5TF20, Q7L2M2 | Gene names | KIAA0408 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0408. | |||||
|
TRHY_HUMAN
|
||||||
| θ value | 6.88961 (rank : 32) | NC score | 0.010819 (rank : 34) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 1385 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q07283 | Gene names | THH, THL, TRHY | |||
|
Domain Architecture |

|
|||||
| Description | Trichohyalin. | |||||
|
ASPP2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 33) | NC score | 0.008720 (rank : 39) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 1156 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q13625, Q12892, Q86X75, Q96KQ3 | Gene names | TP53BP2, ASPP2, BBP | |||
|
Domain Architecture |

|
|||||
| Description | Apoptosis-stimulating of p53 protein 2 (Tumor suppressor p53-binding protein 2) (p53-binding protein 2) (p53BP2) (53BP2) (Bcl2-binding protein) (Bbp) (NY-REN-51 antigen). | |||||
|
DCP1A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 34) | NC score | 0.014671 (rank : 27) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q91YD3, Q6NZE3 | Gene names | Dcp1a, Mitc1, Smif | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | mRNA decapping enzyme 1A (EC 3.-.-.-) (Transcription factor SMIF) (MAD homolog 4-interacting transcription coactivator 1) (Smad4-interacting transcriptional co-activator). | |||||
|
DNMT1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 35) | NC score | 0.013391 (rank : 30) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P13864, P97413, Q80ZU3, Q9CSC6, Q9QXX6 | Gene names | Dnmt1, Dnmt, Met1, Uim | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 1 (EC 2.1.1.37) (Dnmt1) (DNA methyltransferase MmuI) (DNA MTase MmuI) (MCMT) (M.MmuI) (Met-1). | |||||
|
IWS1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 36) | NC score | 0.010993 (rank : 33) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
LRP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 37) | NC score | 0.004106 (rank : 41) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 617 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q07954, Q2PP12, Q8IVG8 | Gene names | LRP1, A2MR | |||
|
Domain Architecture |

|
|||||
| Description | Low-density lipoprotein receptor-related protein 1 precursor (LRP) (Alpha-2-macroglobulin receptor) (A2MR) (Apolipoprotein E receptor) (APOER) (CD91 antigen). | |||||
|
MMAA_HUMAN
|
||||||
| θ value | 8.99809 (rank : 38) | NC score | 0.017924 (rank : 24) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8IVH4 | Gene names | MMAA | |||
|
Domain Architecture |

|
|||||
| Description | Methylmalonic aciduria type A protein, mitochondrial precursor. | |||||
|
PPRB_HUMAN
|
||||||
| θ value | 8.99809 (rank : 39) | NC score | 0.013309 (rank : 31) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15648, O43810, O75447, Q9HD39 | Gene names | PPARBP, ARC205, DRIP205, TRAP220, TRIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor-binding protein (PBP) (PPAR-binding protein) (Thyroid hormone receptor-associated protein complex 220 kDa component) (Trap220) (Thyroid receptor-interacting protein 2) (TRIP-2) (p53 regulatory protein RB18A) (Vitamin D receptor-interacting protein complex component DRIP205) (Activator- recruited cofactor 205 kDa component) (ARC205). | |||||
|
SYCP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 40) | NC score | 0.010069 (rank : 37) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 1313 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q15431, O14963 | Gene names | SYCP1, SCP1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptonemal complex protein 1 (SCP-1). | |||||
|
TCPA1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 41) | NC score | 0.010722 (rank : 36) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P11984 | Gene names | Cct1, Ccta, Tcp1 | |||
|
Domain Architecture |
|
|||||
| Description | T-complex protein 1 subunit alpha A (TCP-1-alpha) (CCT-alpha) (Tailless complex polypeptide 1A) (TCP-1-A). | |||||
|
TCPA2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 42) | NC score | 0.010745 (rank : 35) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P11983 | Gene names | Cct1, Ccta, Tcp1 | |||
|
Domain Architecture |

|
|||||
| Description | T-complex protein 1 subunit alpha B (TCP-1-alpha) (CCT-alpha) (Tailless complex polypeptide 1B) (TCP-1-B). | |||||
|
MLH3_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9UHC1, P49751, Q56DK9, Q9P292, Q9UHC0 | Gene names | MLH3 | |||
|
Domain Architecture |

|
|||||
| Description | DNA mismatch repair protein Mlh3 (MutL protein homolog 3). | |||||
|
MLH1_HUMAN
|
||||||
| NC score | 0.651832 (rank : 2) | θ value | 3.07116e-24 (rank : 2) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P40692 | Gene names | MLH1, COCA2 | |||
|
Domain Architecture |

|
|||||
| Description | DNA mismatch repair protein Mlh1 (MutL protein homolog 1). | |||||
|
MLH1_MOUSE
|
||||||
| NC score | 0.644750 (rank : 3) | θ value | 1.29031e-22 (rank : 3) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9JK91, Q62454 | Gene names | Mlh1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA mismatch repair protein Mlh1 (MutL protein homolog 1). | |||||
|
PMS1_HUMAN
|
||||||
| NC score | 0.589997 (rank : 4) | θ value | 6.40375e-22 (rank : 4) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P54277 | Gene names | PMS1, PMSL1 | |||
|
Domain Architecture |

|
|||||
| Description | PMS1 protein homolog 1 (DNA mismatch repair protein PMS1). | |||||
|
PMS2_HUMAN
|
||||||
| NC score | 0.568280 (rank : 5) | θ value | 6.85773e-16 (rank : 5) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P54278 | Gene names | PMS2, PMSL2 | |||
|
Domain Architecture |

|
|||||
| Description | PMS1 protein homolog 2 (DNA mismatch repair protein PMS2). | |||||
|
PMS2_MOUSE
|
||||||
| NC score | 0.567305 (rank : 6) | θ value | 2.88119e-14 (rank : 6) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P54279 | Gene names | Pms2 | |||
|
Domain Architecture |

|
|||||
| Description | PMS1 protein homolog 2 (DNA mismatch repair protein PMS2). | |||||
|
DMP1_HUMAN
|
||||||
| NC score | 0.046520 (rank : 7) | θ value | 0.813845 (rank : 10) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q13316, O43265 | Gene names | DMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1). | |||||
|
SETX_HUMAN
|
||||||
| NC score | 0.046369 (rank : 8) | θ value | 0.47712 (rank : 8) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q7Z333, O75120, Q3KQX4, Q68DW5, Q6AZD7, Q7Z3J6, Q8WX33, Q9H9D1, Q9NVP9 | Gene names | SETX, ALS4, KIAA0625, SCAR1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable helicase senataxin (EC 3.6.1.-) (SEN1 homolog). | |||||
|
PREP_MOUSE
|
||||||
| NC score | 0.042912 (rank : 9) | θ value | 1.06291 (rank : 11) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8K411, Q3THC4, Q3TJI1, Q3TPP0, Q3URQ8, Q4KUG2, Q4KUG3, Q6ZPX6, Q922N1, Q9CV63 | Gene names | Pitrm1, Kiaa1104, Ntup1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Presequence protease, mitochondrial precursor (EC 3.4.24.-) (Pitrilysin metalloproteinase 1). | |||||
|
CIR_MOUSE
|
||||||
| NC score | 0.034583 (rank : 10) | θ value | 2.36792 (rank : 18) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9DA19, Q3V2N9, Q4KL44, Q52KL9, Q5FW66 | Gene names | Cir | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CBF1-interacting corepressor. | |||||
|
K0408_HUMAN
|
||||||
| NC score | 0.027307 (rank : 11) | θ value | 6.88961 (rank : 31) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6ZU52, O43158, Q5TF20, Q7L2M2 | Gene names | KIAA0408 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0408. | |||||
|
PALB2_HUMAN
|
||||||
| NC score | 0.026015 (rank : 12) | θ value | 4.03905 (rank : 25) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q86YC2, Q8N7Y6, Q8ND31, Q9H6W1 | Gene names | PALB2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Partner and localizer of BRCA2. | |||||
|
CT2NL_MOUSE
|
||||||
| NC score | 0.025501 (rank : 13) | θ value | 4.03905 (rank : 22) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 902 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q99LJ0, Q6ZPR0, Q8BSV1 | Gene names | Cttnbp2nl, Kiaa1433 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CTTNBP2 N-terminal-like protein. | |||||
|
WWP1_HUMAN
|
||||||
| NC score | 0.024995 (rank : 14) | θ value | 1.06291 (rank : 12) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9H0M0, O00307, Q96BP4 | Gene names | WWP1 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP1 (EC 6.3.2.-) (WW domain- containing protein 1) (Atropin-1-interacting protein 5) (AIP5). | |||||
|
CT2NL_HUMAN
|
||||||
| NC score | 0.024870 (rank : 15) | θ value | 1.81305 (rank : 16) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 1059 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9P2B4, Q96B40 | Gene names | CTTNBP2NL, KIAA1433 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CTTNBP2 N-terminal-like protein. | |||||
|
WWP1_MOUSE
|
||||||
| NC score | 0.024248 (rank : 16) | θ value | 1.06291 (rank : 13) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BZZ3, Q8BIV9, Q8VDP8 | Gene names | Wwp1 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP1 (EC 6.3.2.-) (WW domain- containing protein 1). | |||||
|
ZN318_MOUSE
|
||||||
| NC score | 0.023811 (rank : 17) | θ value | 4.03905 (rank : 26) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 642 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q99PP2, Q3TZL5, Q9JJ01 | Gene names | Znf318, Tzf, Zfp318 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 318 (Testicular zinc finger protein). | |||||
|
UBP31_HUMAN
|
||||||
| NC score | 0.023635 (rank : 18) | θ value | 1.38821 (rank : 15) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 293 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q70CQ4, Q6AW97, Q6ZTC0, Q6ZTN2, Q9ULL7 | Gene names | USP31, KIAA1203 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 31 (EC 3.1.2.15) (Ubiquitin thioesterase 31) (Ubiquitin-specific-processing protease 31) (Deubiquitinating enzyme 31). | |||||
|
PML_MOUSE
|
||||||
| NC score | 0.023060 (rank : 19) | θ value | 3.0926 (rank : 21) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q60953 | Gene names | Pml | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable transcription factor PML. | |||||
|
DIAP3_MOUSE
|
||||||
| NC score | 0.021270 (rank : 20) | θ value | 1.81305 (rank : 17) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Z207 | Gene names | Diaph3, Diap3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 3 (Diaphanous-related formin-3) (DRF3) (mDIA2) (p134mDIA2). | |||||
|
BRCA2_HUMAN
|
||||||
| NC score | 0.020352 (rank : 21) | θ value | 5.27518 (rank : 27) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P51587, O00183, O15008, Q13879 | Gene names | BRCA2, FANCD1 | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer type 2 susceptibility protein (Fanconi anemia group D1 protein). | |||||
|
PDZK4_HUMAN
|
||||||
| NC score | 0.019216 (rank : 22) | θ value | 3.0926 (rank : 20) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q76G19, Q8NB75, Q9BUH9, Q9P284 | Gene names | PDZK4, KIAA1444, PDZRN4L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 4 (PDZ domain-containing RING finger protein 4-like protein). | |||||
|
TPR_HUMAN
|
||||||
| NC score | 0.018430 (rank : 23) | θ value | 0.47712 (rank : 9) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
MMAA_HUMAN
|
||||||
| NC score | 0.017924 (rank : 24) | θ value | 8.99809 (rank : 38) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8IVH4 | Gene names | MMAA | |||
|
Domain Architecture |

|
|||||
| Description | Methylmalonic aciduria type A protein, mitochondrial precursor. | |||||
|
EP15_HUMAN
|
||||||
| NC score | 0.015498 (rank : 25) | θ value | 4.03905 (rank : 23) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 1021 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P42566 | Gene names | EPS15, AF1P | |||
|
Domain Architecture |

|
|||||
| Description | Epidermal growth factor receptor substrate 15 (Protein Eps15) (AF-1p protein). | |||||
|
DPOLZ_HUMAN
|
||||||
| NC score | 0.014739 (rank : 26) | θ value | 6.88961 (rank : 30) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O60673, O43214 | Gene names | REV3L, POLZ, REV3 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase zeta catalytic subunit (EC 2.7.7.7) (hREV3). | |||||
|
DCP1A_MOUSE
|
||||||
| NC score | 0.014671 (rank : 27) | θ value | 8.99809 (rank : 34) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q91YD3, Q6NZE3 | Gene names | Dcp1a, Mitc1, Smif | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | mRNA decapping enzyme 1A (EC 3.-.-.-) (Transcription factor SMIF) (MAD homolog 4-interacting transcription coactivator 1) (Smad4-interacting transcriptional co-activator). | |||||
|
CSPG2_HUMAN
|
||||||
| NC score | 0.014640 (rank : 28) | θ value | 1.38821 (rank : 14) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P13611, P20754, Q13010, Q13189, Q15123, Q9UCL9, Q9UNW5 | Gene names | CSPG2 | |||
|
Domain Architecture |

|
|||||
| Description | Versican core protein precursor (Large fibroblast proteoglycan) (Chondroitin sulfate proteoglycan core protein 2) (PG-M) (Glial hyaluronate-binding protein) (GHAP). | |||||
|
OPTN_MOUSE
|
||||||
| NC score | 0.014477 (rank : 29) | θ value | 3.0926 (rank : 19) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 1120 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8K3K8 | Gene names | Optn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Optineurin. | |||||
|
DNMT1_MOUSE
|
||||||
| NC score | 0.013391 (rank : 30) | θ value | 8.99809 (rank : 35) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P13864, P97413, Q80ZU3, Q9CSC6, Q9QXX6 | Gene names | Dnmt1, Dnmt, Met1, Uim | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 1 (EC 2.1.1.37) (Dnmt1) (DNA methyltransferase MmuI) (DNA MTase MmuI) (MCMT) (M.MmuI) (Met-1). | |||||
|
PPRB_HUMAN
|
||||||
| NC score | 0.013309 (rank : 31) | θ value | 8.99809 (rank : 39) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15648, O43810, O75447, Q9HD39 | Gene names | PPARBP, ARC205, DRIP205, TRAP220, TRIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor-binding protein (PBP) (PPAR-binding protein) (Thyroid hormone receptor-associated protein complex 220 kDa component) (Trap220) (Thyroid receptor-interacting protein 2) (TRIP-2) (p53 regulatory protein RB18A) (Vitamin D receptor-interacting protein complex component DRIP205) (Activator- recruited cofactor 205 kDa component) (ARC205). | |||||
|
STAM2_MOUSE
|
||||||
| NC score | 0.011492 (rank : 32) | θ value | 5.27518 (rank : 29) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O88811, Q8C8Y4 | Gene names | Stam2, Hbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal transducing adapter molecule 2 (STAM-2) (Hrs-binding protein). | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.010993 (rank : 33) | θ value | 8.99809 (rank : 36) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
TRHY_HUMAN
|
||||||
| NC score | 0.010819 (rank : 34) | θ value | 6.88961 (rank : 32) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 1385 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q07283 | Gene names | THH, THL, TRHY | |||
|
Domain Architecture |

|
|||||
| Description | Trichohyalin. | |||||
|
TCPA2_MOUSE
|
||||||
| NC score | 0.010745 (rank : 35) | θ value | 8.99809 (rank : 42) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P11983 | Gene names | Cct1, Ccta, Tcp1 | |||
|
Domain Architecture |

|
|||||
| Description | T-complex protein 1 subunit alpha B (TCP-1-alpha) (CCT-alpha) (Tailless complex polypeptide 1B) (TCP-1-B). | |||||
|
TCPA1_MOUSE
|
||||||
| NC score | 0.010722 (rank : 36) | θ value | 8.99809 (rank : 41) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P11984 | Gene names | Cct1, Ccta, Tcp1 | |||
|
Domain Architecture |

|
|||||
| Description | T-complex protein 1 subunit alpha A (TCP-1-alpha) (CCT-alpha) (Tailless complex polypeptide 1A) (TCP-1-A). | |||||
|
SYCP1_HUMAN
|
||||||
| NC score | 0.010069 (rank : 37) | θ value | 8.99809 (rank : 40) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 1313 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q15431, O14963 | Gene names | SYCP1, SCP1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptonemal complex protein 1 (SCP-1). | |||||
|
FGD3_HUMAN
|
||||||
| NC score | 0.009609 (rank : 38) | θ value | 5.27518 (rank : 28) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5JSP0, Q7Z7D9, Q8N5G1 | Gene names | FGD3, ZFYVE5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 3 (Zinc finger FYVE domain-containing protein 5). | |||||
|
ASPP2_HUMAN
|
||||||
| NC score | 0.008720 (rank : 39) | θ value | 8.99809 (rank : 33) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 1156 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q13625, Q12892, Q86X75, Q96KQ3 | Gene names | TP53BP2, ASPP2, BBP | |||
|
Domain Architecture |

|
|||||
| Description | Apoptosis-stimulating of p53 protein 2 (Tumor suppressor p53-binding protein 2) (p53-binding protein 2) (p53BP2) (53BP2) (Bcl2-binding protein) (Bbp) (NY-REN-51 antigen). | |||||
|
MYH3_HUMAN
|
||||||
| NC score | 0.008172 (rank : 40) | θ value | 4.03905 (rank : 24) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 1660 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P11055, Q15492 | Gene names | MYH3 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, fast skeletal muscle, embryonic (Muscle embryonic myosin heavy chain) (SMHCE). | |||||
|
LRP1_HUMAN
|
||||||
| NC score | 0.004106 (rank : 41) | θ value | 8.99809 (rank : 37) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 617 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q07954, Q2PP12, Q8IVG8 | Gene names | LRP1, A2MR | |||
|
Domain Architecture |

|
|||||
| Description | Low-density lipoprotein receptor-related protein 1 precursor (LRP) (Alpha-2-macroglobulin receptor) (A2MR) (Apolipoprotein E receptor) (APOER) (CD91 antigen). | |||||
|
ZN514_HUMAN
|
||||||
| NC score | 0.003607 (rank : 42) | θ value | 0.365318 (rank : 7) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 738 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96K75 | Gene names | ZNF514 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 514. | |||||