Please be patient as the page loads
|
ALS2_HUMAN
|
||||||
| SwissProt Accessions | Q96Q42, Q8N1E0, Q96PC4, Q96Q41, Q9H973, Q9HCK9 | Gene names | ALS2, KIAA1563 | |||
|
Domain Architecture |

|
|||||
| Description | Alsin (Amyotrophic lateral sclerosis protein 2). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
ALS2_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 121 | |
| SwissProt Accessions | Q96Q42, Q8N1E0, Q96PC4, Q96Q41, Q9H973, Q9HCK9 | Gene names | ALS2, KIAA1563 | |||
|
Domain Architecture |

|
|||||
| Description | Alsin (Amyotrophic lateral sclerosis protein 2). | |||||
|
ALS2_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.993643 (rank : 2) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 90 | |
| SwissProt Accessions | Q920R0, Q8JZR1, Q9CXJ3 | Gene names | Als2 | |||
|
Domain Architecture |

|
|||||
| Description | Alsin (Amyotrophic lateral sclerosis protein 2 homolog). | |||||
|
HERC3_HUMAN
|
||||||
| θ value | 1.63225e-17 (rank : 3) | NC score | 0.415846 (rank : 25) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q15034 | Gene names | HERC3, KIAA0032 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT domain and RCC1-like domain protein 3. | |||||
|
RCBT1_MOUSE
|
||||||
| θ value | 1.058e-16 (rank : 4) | NC score | 0.476897 (rank : 20) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q6NXM2, Q8BTZ6, Q8BZV0 | Gene names | Rcbtb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RCC1 and BTB domain-containing protein 1 (Regulator of chromosome condensation and BTB domain-containing protein 1). | |||||
|
TSGA2_HUMAN
|
||||||
| θ value | 6.85773e-16 (rank : 5) | NC score | 0.539903 (rank : 6) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8WYR4 | Gene names | TSGA2, TSA2 | |||
|
Domain Architecture |

|
|||||
| Description | Testis-specific gene A2 protein (Male meiotic metaphase chromosome- associated acidic protein) (Meichroacidin). | |||||
|
HERC2_HUMAN
|
||||||
| θ value | 8.95645e-16 (rank : 6) | NC score | 0.416483 (rank : 24) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O95714, Q86SV7, Q86SV8, Q86SV9, Q86YY3, Q86YY4, Q86YY5, Q86YY6, Q86YY7, Q86YY8, Q86YY9, Q86YZ0, Q86YZ1 | Gene names | HERC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT domain and RCC1-like domain-containing protein 2. | |||||
|
HERC2_MOUSE
|
||||||
| θ value | 8.95645e-16 (rank : 7) | NC score | 0.414944 (rank : 26) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q4U2R1, O88473, Q3TRJ8, Q3TS47, Q3TST2, Q3UFQ6, Q3URH7, Q5DU32, Q7TPR5, Q80VV7, Q9QYT1, Q9Z168, Q9Z171 | Gene names | Herc2, jdf2, Kiaa0393, rjs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT domain and RCC1-like domain-containing protein 2. | |||||
|
RCBT1_HUMAN
|
||||||
| θ value | 1.16975e-15 (rank : 8) | NC score | 0.474526 (rank : 21) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8NDN9, Q969U9 | Gene names | RCBTB1, CLLD7, E4.5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RCC1 and BTB domain-containing protein 1 (Regulator of chromosome condensation and BTB domain-containing protein 1) (Chronic lymphocytic leukemia deletion region gene 7 protein) (CLL deletion region gene 7 protein). | |||||
|
MORN1_HUMAN
|
||||||
| θ value | 3.40345e-15 (rank : 9) | NC score | 0.522228 (rank : 12) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q5T089, Q8WW30, Q9H852 | Gene names | MORN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORN repeat-containing protein 1. | |||||
|
MORN3_MOUSE
|
||||||
| θ value | 1.09485e-13 (rank : 10) | NC score | 0.522224 (rank : 13) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8C5T4, Q8VE21, Q9D5H6 | Gene names | Morn3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORN repeat-containing protein 3. | |||||
|
TSGA2_MOUSE
|
||||||
| θ value | 1.09485e-13 (rank : 11) | NC score | 0.522938 (rank : 10) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8VIG3, Q9DAL5 | Gene names | Tsga2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-specific gene A2 protein (Male meiotic metaphase chromosome- associated acidic protein) (Meichroacidin). | |||||
|
RCBT2_MOUSE
|
||||||
| θ value | 1.42992e-13 (rank : 12) | NC score | 0.483813 (rank : 18) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q99LJ7, Q3TUA3, Q8BMG2 | Gene names | Rcbtb2, Chc1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RCC1 and BTB domain-containing protein 2 (Regulator of chromosome condensation and BTB domain-containing protein 2) (Chromosome condensation 1-like). | |||||
|
RPGR_MOUSE
|
||||||
| θ value | 1.86753e-13 (rank : 13) | NC score | 0.530670 (rank : 8) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9R0X5, O88408, Q9CU92 | Gene names | Rpgr | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator (mRpgr). | |||||
|
MORN3_HUMAN
|
||||||
| θ value | 3.18553e-13 (rank : 14) | NC score | 0.518648 (rank : 14) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6PF18, Q86YQ9 | Gene names | MORN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORN repeat-containing protein 3. | |||||
|
RPGR_HUMAN
|
||||||
| θ value | 5.43371e-13 (rank : 15) | NC score | 0.526639 (rank : 9) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q92834, O00702, O00737, Q8N5T6, Q93039, Q9HD29, Q9UMR1 | Gene names | RPGR, RP3, XLRP3 | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator. | |||||
|
HERC5_HUMAN
|
||||||
| θ value | 9.26847e-13 (rank : 16) | NC score | 0.404522 (rank : 32) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9UII4 | Gene names | HERC5, CEB1, CEBP1 | |||
|
Domain Architecture |

|
|||||
| Description | HECT domain and RCC1-like domain-containing protein 5 (Cyclin-E- binding protein 1). | |||||
|
RCC2_HUMAN
|
||||||
| θ value | 1.2105e-12 (rank : 17) | NC score | 0.567947 (rank : 3) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9P258, Q8IVL9, Q9BSN6, Q9NPV8 | Gene names | RCC2, KIAA1470, TD60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein RCC2 (Telophase disk protein of 60 kDa) (RCC1-like protein TD- 60). | |||||
|
RCBT2_HUMAN
|
||||||
| θ value | 1.58096e-12 (rank : 18) | NC score | 0.483673 (rank : 19) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O95199 | Gene names | RCBTB2, CHC1L, RLG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RCC1 and BTB domain-containing protein 2 (Regulator of chromosome condensation and BTB domain-containing protein 2) (Chromosome condensation 1-like) (CHC1-L) (RCC1-like G exchanging factor). | |||||
|
RCC2_MOUSE
|
||||||
| θ value | 1.58096e-12 (rank : 19) | NC score | 0.567581 (rank : 4) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8BK67, Q6ZPQ0 | Gene names | Rcc2, Kiaa1470 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein RCC2. | |||||
|
NEK8_HUMAN
|
||||||
| θ value | 4.59992e-12 (rank : 20) | NC score | 0.131797 (rank : 55) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 872 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q86SG6, Q2M1S6, Q8NDH1 | Gene names | NEK8, JCK, NEK12A | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Nek8 (EC 2.7.11.1) (Never in mitosis A-related kinase 8) (NimA-related protein kinase 8) (NIMA-related kinase 12a). | |||||
|
JPH2_HUMAN
|
||||||
| θ value | 2.98157e-11 (rank : 21) | NC score | 0.382160 (rank : 33) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 639 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9BR39, O95913, Q5JY74, Q9UJN4 | Gene names | JPH2, JP2 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-2 (Junctophilin type 2) (JP-2). | |||||
|
JPH2_MOUSE
|
||||||
| θ value | 2.98157e-11 (rank : 22) | NC score | 0.407916 (rank : 30) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9ET78, Q9ET79 | Gene names | Jph2, Jp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junctophilin-2 (Junctophilin type 2) (JP-2). | |||||
|
JPH3_HUMAN
|
||||||
| θ value | 3.89403e-11 (rank : 23) | NC score | 0.422168 (rank : 22) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8WXH2, Q8N471, Q9HDC3, Q9HDC4 | Gene names | JPH3, JP3 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-3 (Junctophilin type 3) (JP-3). | |||||
|
JPH3_MOUSE
|
||||||
| θ value | 3.89403e-11 (rank : 24) | NC score | 0.422158 (rank : 23) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9ET77, Q9EQZ2 | Gene names | Jph3, Jp3 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-3 (Junctophilin type 3) (JP-3). | |||||
|
JPH1_MOUSE
|
||||||
| θ value | 1.9326e-10 (rank : 25) | NC score | 0.413604 (rank : 27) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9ET80, Q9EQZ3 | Gene names | Jph1, Jp1 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-1 (Junctophilin type 1) (JP-1). | |||||
|
NEK8_MOUSE
|
||||||
| θ value | 4.30538e-10 (rank : 26) | NC score | 0.130741 (rank : 57) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 872 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q91ZR4, Q3U498, Q9D685 | Gene names | Nek8, Jck | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Nek8 (EC 2.7.11.1) (Never in mitosis A-related kinase 8) (NimA-related protein kinase 8). | |||||
|
JPH1_HUMAN
|
||||||
| θ value | 1.25267e-09 (rank : 27) | NC score | 0.411031 (rank : 28) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9HDC5 | Gene names | JPH1, JP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junctophilin-1 (Junctophilin type 1) (JP-1). | |||||
|
JPH4_HUMAN
|
||||||
| θ value | 2.79066e-09 (rank : 28) | NC score | 0.409207 (rank : 29) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96JJ6, Q8ND44, Q96DQ0 | Gene names | JPH4, JPHL1, KIAA1831 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junctophilin-4 (Junctophilin-like 1 protein). | |||||
|
JPH4_MOUSE
|
||||||
| θ value | 2.79066e-09 (rank : 29) | NC score | 0.406507 (rank : 31) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q80WT0, Q8BMI1 | Gene names | Jph4, Jphl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junctophilin-4 (Junctophilin-like 1 protein). | |||||
|
SRGEF_MOUSE
|
||||||
| θ value | 2.79066e-09 (rank : 30) | NC score | 0.542221 (rank : 5) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q80YD6, Q9QXB7 | Gene names | Sergef, Delgef, Gnefr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Secretion-regulating guanine nucleotide exchange factor (Guanine nucleotide exchange factor-related protein) (Deafness locus-associated putative guanine nucleotide exchange factor) (DelGEF). | |||||
|
SRGEF_HUMAN
|
||||||
| θ value | 1.80886e-08 (rank : 31) | NC score | 0.536960 (rank : 7) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9UGK8, Q9UGK9 | Gene names | SERGEF, DELGEF, GNEFR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Secretion-regulating guanine nucleotide exchange factor (Guanine nucleotide exchange factor-related protein) (Deafness locus-associated putative guanine nucleotide exchange factor) (DelGEF). | |||||
|
RCC1_MOUSE
|
||||||
| θ value | 6.87365e-08 (rank : 32) | NC score | 0.522577 (rank : 11) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8VE37, Q3UDB6 | Gene names | Rcc1, Chc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of chromosome condensation (Chromosome condensation protein 1). | |||||
|
MYCB2_HUMAN
|
||||||
| θ value | 1.17247e-07 (rank : 33) | NC score | 0.331789 (rank : 36) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 390 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O75592, Q5JSX8, Q5VZN6, Q6PIB6, Q9UQ11, Q9Y6E4 | Gene names | MYCBP2, KIAA0916, PAM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ubiquitin ligase protein MYCBP2 (EC 6.3.2.-) (Myc-binding protein 2) (Protein associated with Myc) (Pam/highwire/rpm-1 protein). | |||||
|
NEK9_HUMAN
|
||||||
| θ value | 2.61198e-07 (rank : 34) | NC score | 0.130259 (rank : 58) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 900 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8TD19, Q52LK6, Q8NCN0, Q8TCY4, Q9UPI4, Q9Y6S4, Q9Y6S5, Q9Y6S6 | Gene names | NEK9, KIAA1995, NEK8, NERCC | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Nek9 (EC 2.7.11.1) (NimA-related protein kinase 9) (Never in mitosis A-related kinase 9) (Nercc1 kinase) (NIMA-related kinase 8) (Nek8). | |||||
|
NEK9_MOUSE
|
||||||
| θ value | 5.81887e-07 (rank : 35) | NC score | 0.130951 (rank : 56) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 890 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8K1R7, Q8R3P1 | Gene names | Nek9, Nercc | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Nek9 (EC 2.7.11.1) (NimA-related protein kinase 9) (Never in mitosis A-related kinase 9). | |||||
|
RCC1_HUMAN
|
||||||
| θ value | 7.59969e-07 (rank : 36) | NC score | 0.518430 (rank : 15) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P18754 | Gene names | RCC1, CHC1 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of chromosome condensation (Chromosome condensation protein 1) (Cell cycle regulatory protein). | |||||
|
WBS16_MOUSE
|
||||||
| θ value | 7.59969e-07 (rank : 37) | NC score | 0.506162 (rank : 17) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9CYF5 | Gene names | Wbscr16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Williams-Beuren syndrome chromosome region 16 protein homolog. | |||||
|
WBS16_HUMAN
|
||||||
| θ value | 9.92553e-07 (rank : 38) | NC score | 0.508561 (rank : 16) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q96I51, Q548B1, Q9H0G7 | Gene names | WBSCR16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Williams-Beuren syndrome chromosome region 16 protein (RCC1-like G exchanging factor-like protein). | |||||
|
MYCB2_MOUSE
|
||||||
| θ value | 1.29631e-06 (rank : 39) | NC score | 0.325887 (rank : 37) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q7TPH6, Q6PCM8 | Gene names | Mycbp2, Pam, Phr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ubiquitin ligase protein MYCBP2 (EC 6.3.2.-) (Myc-binding protein 2) (Protein associated with Myc) (Pam/highwire/rpm-1 protein). | |||||
|
PKHF2_HUMAN
|
||||||
| θ value | 1.87187e-05 (rank : 40) | NC score | 0.198113 (rank : 43) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9H8W4 | Gene names | PLEKHF2, ZFYVE18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family F member 2 (PH domain- containing family F member 2) (PH and FYVE domain-containing protein 2) (Phafin-2) (Zinc finger FYVE domain-containing protein 18). | |||||
|
PKHF2_MOUSE
|
||||||
| θ value | 2.44474e-05 (rank : 41) | NC score | 0.196619 (rank : 44) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q91WB4 | Gene names | Plekhf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family F member 2 (PH domain- containing family F member 2). | |||||
|
FARP1_HUMAN
|
||||||
| θ value | 7.1131e-05 (rank : 42) | NC score | 0.120430 (rank : 60) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9Y4F1, Q6IQ29 | Gene names | FARP1, CDEP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FERM, RhoGEF and pleckstrin domain-containing protein 1 (Chondrocyte- derived ezrin-like protein). | |||||
|
ANKY1_HUMAN
|
||||||
| θ value | 0.000158464 (rank : 43) | NC score | 0.211279 (rank : 41) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9P2S6, Q8IYX5, Q8NDK5, Q9NX10 | Gene names | ANKMY1, TSAL1, ZMYND13 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin repeat and MYND domain-containing protein 1 (Testis-specific ankyrin-like protein 1) (Zinc-finger MYND domain-containing protein 13). | |||||
|
ARHG6_HUMAN
|
||||||
| θ value | 0.000270298 (rank : 44) | NC score | 0.106530 (rank : 64) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 376 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q15052, Q15396, Q5JQ66, Q86XH0 | Gene names | ARHGEF6, COOL2, KIAA0006, PIXA | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 6 (Rac/Cdc42 guanine nucleotide exchange factor 6) (PAK-interacting exchange factor alpha) (Alpha-Pix) (COOL-2). | |||||
|
FGD2_MOUSE
|
||||||
| θ value | 0.000270298 (rank : 45) | NC score | 0.138470 (rank : 51) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BY35, O88841, Q7TSE3, Q8VDH4 | Gene names | Fgd2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 2. | |||||
|
FARP2_MOUSE
|
||||||
| θ value | 0.00035302 (rank : 46) | NC score | 0.112810 (rank : 63) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q91VS8, Q3TAP2, Q69ZZ0 | Gene names | Farp2, Kiaa0793 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FERM, RhoGEF and pleckstrin domain-containing protein 2 (FERM domain including RhoGEF) (FIR). | |||||
|
PKHF1_MOUSE
|
||||||
| θ value | 0.000786445 (rank : 47) | NC score | 0.186109 (rank : 45) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q3TB82, Q99M16 | Gene names | Plekhf1, Lapf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family F member 1 (PH domain- containing family F member 1) (Lysosome-associated apoptosis-inducing protein containing PH and FYVE domains). | |||||
|
FGD6_HUMAN
|
||||||
| θ value | 0.00102713 (rank : 48) | NC score | 0.142912 (rank : 48) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 237 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6ZV73, Q6ZR53, Q7Z2Z7, Q96D44, Q9NUR8, Q9P2I5 | Gene names | FGD6, KIAA1362, ZFYVE24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 6 (Zinc finger FYVE domain-containing protein 24). | |||||
|
RIN3_MOUSE
|
||||||
| θ value | 0.00102713 (rank : 49) | NC score | 0.140151 (rank : 50) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P59729 | Gene names | Rin3 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 3 (Ras interaction/interference protein 3). | |||||
|
ARHG6_MOUSE
|
||||||
| θ value | 0.00228821 (rank : 50) | NC score | 0.098573 (rank : 68) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8K4I3, Q8C9V4 | Gene names | Arhgef6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 6 (Rac/Cdc42 guanine nucleotide exchange factor 6). | |||||
|
FGD6_MOUSE
|
||||||
| θ value | 0.00228821 (rank : 51) | NC score | 0.142519 (rank : 49) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q69ZL1, Q8C8W5, Q8K3B0, Q9D3Y7 | Gene names | Fgd6, Kiaa1362 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 6. | |||||
|
PKHF1_HUMAN
|
||||||
| θ value | 0.00228821 (rank : 52) | NC score | 0.179832 (rank : 46) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q96S99, Q96K11, Q9BUB9 | Gene names | PLEKHF1, APPD, LAPF, ZFYVE15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family F member 1 (PH domain- containing family F member 1) (PH and FYVE domain-containing protein 1) (Phafin-1) (Lysosome-associated apoptosis-inducing protein containing PH and FYVE domains) (Apoptosis-inducing protein) (Zinc finger FYVE domain-containing protein 15). | |||||
|
RIN3_HUMAN
|
||||||
| θ value | 0.00228821 (rank : 53) | NC score | 0.138401 (rank : 52) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8TB24, Q8NF30, Q8TEE8, Q8WYP4, Q9H6A5, Q9HAG1 | Gene names | RIN3 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 3 (Ras interaction/interference protein 3). | |||||
|
RABX5_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 54) | NC score | 0.145957 (rank : 47) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9JM13, Q3UIW0 | Gene names | Rabgef1, Rabex5 | |||
|
Domain Architecture |

|
|||||
| Description | Rab5 GDP/GTP exchange factor (Rabex-5). | |||||
|
RIN2_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 55) | NC score | 0.136578 (rank : 53) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8WYP3, Q00425, Q5TFT8, Q9BQL3, Q9H071 | Gene names | RIN2, RASSF4 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 2 (Ras interaction/interference protein 2) (Ras inhibitor JC265) (Ras association domain family 4). | |||||
|
RIN2_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 56) | NC score | 0.135705 (rank : 54) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9D684, Q99K06 | Gene names | Rin2 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 2 (Ras interaction/interference protein 2). | |||||
|
MORN2_HUMAN
|
||||||
| θ value | 0.00665767 (rank : 57) | NC score | 0.368749 (rank : 34) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q502X0, Q6UL00 | Gene names | MORN2, MOPT | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORN repeat-containing protein 2 (MORN motif protein in testis). | |||||
|
VAV2_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 58) | NC score | 0.068052 (rank : 84) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 453 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P52735 | Gene names | VAV2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-2. | |||||
|
MORN2_MOUSE
|
||||||
| θ value | 0.0148317 (rank : 59) | NC score | 0.351045 (rank : 35) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6UL01 | Gene names | Morn2, Mopt | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORN repeat-containing protein 2 (MORN motif protein in testis). | |||||
|
RABX5_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 60) | NC score | 0.124671 (rank : 59) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UJ41 | Gene names | RABGEF1, RABEX5 | |||
|
Domain Architecture |

|
|||||
| Description | Rab5 GDP/GTP exchange factor (Rabex-5) (Rabaptin-5-associated exchange factor for Rab5). | |||||
|
SETD7_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 61) | NC score | 0.228384 (rank : 40) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8WTS6, Q9C0E6 | Gene names | SETD7, KIAA1717, SET7 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET7 (EC 2.1.1.43) (Histone H3-K4 methyltransferase) (H3-K4-HMTase) (SET domain-containing protein 7) (Set9) (SET7/9). | |||||
|
SETD7_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 62) | NC score | 0.228638 (rank : 39) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8VHL1 | Gene names | Setd7, Set7 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET7 (EC 2.1.1.43) (Histone H3-K4 methyltransferase) (H3-K4-HMTase) (SET domain-containing protein 7). | |||||
|
FARP2_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 63) | NC score | 0.094352 (rank : 72) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O94887, Q53QM5, Q8WU27, Q9UFE7 | Gene names | FARP2, KIAA0793 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FERM, RhoGEF and pleckstrin domain-containing protein 2 (FERM domain including RhoGEF) (FIR). | |||||
|
VAV2_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 64) | NC score | 0.064141 (rank : 88) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 456 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q60992 | Gene names | Vav2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-2. | |||||
|
FGD4_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 65) | NC score | 0.118563 (rank : 62) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q91ZT5, Q3UEB6, Q8BW60, Q8BZI7, Q91ZT3, Q91ZT4 | Gene names | Fgd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 4 (Actin filament- binding protein frabin) (FGD1-related F-actin-binding protein). | |||||
|
ARHGA_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 66) | NC score | 0.095502 (rank : 70) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O15013, O14665, Q68D55, Q8IWD9, Q8IY77 | Gene names | ARHGEF10, KIAA0294 | |||
|
Domain Architecture |
|
|||||
| Description | Rho guanine nucleotide exchange factor 10. | |||||
|
FBX24_HUMAN
|
||||||
| θ value | 0.163984 (rank : 67) | NC score | 0.234567 (rank : 38) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O75426, Q9H0G1 | Gene names | FBXO24, FBX24 | |||
|
Domain Architecture |
|
|||||
| Description | F-box only protein 24. | |||||
|
OGT1_HUMAN
|
||||||
| θ value | 0.163984 (rank : 68) | NC score | 0.044619 (rank : 124) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O15294, Q7Z3K0, Q8WWM8, Q96CC1, Q9UG57 | Gene names | OGT | |||
|
Domain Architecture |
|
|||||
| Description | UDP-N-acetylglucosamine--peptide N-acetylglucosaminyltransferase 110 kDa subunit (EC 2.4.1.-) (O-GlcNAc transferase p110 subunit). | |||||
|
E41L3_HUMAN
|
||||||
| θ value | 0.21417 (rank : 69) | NC score | 0.039458 (rank : 125) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y2J2, O95713, Q9BRP5 | Gene names | EPB41L3, DAL1, KIAA0987 | |||
|
Domain Architecture |

|
|||||
| Description | Band 4.1-like protein 3 (4.1B) (Differentially expressed in adenocarcinoma of the lung protein 1) (DAL-1). | |||||
|
FBX24_MOUSE
|
||||||
| θ value | 0.21417 (rank : 70) | NC score | 0.210135 (rank : 42) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9D417 | Gene names | Fbxo24, Fbx24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 24. | |||||
|
FGD2_HUMAN
|
||||||
| θ value | 0.279714 (rank : 71) | NC score | 0.120354 (rank : 61) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q7Z6J4, Q5T8I1, Q6P6A8, Q6ZNL5, Q8IZ32, Q8N868, Q9H7M2 | Gene names | FGD2, ZFYVE4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 2 (Zinc finger FYVE domain-containing protein 4). | |||||
|
PARC_MOUSE
|
||||||
| θ value | 0.279714 (rank : 72) | NC score | 0.053439 (rank : 110) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q80TT8, Q8BKL9, Q8CGC0, Q8R363 | Gene names | Parc, Kiaa0708 | |||
|
Domain Architecture |

|
|||||
| Description | p53-associated parkin-like cytoplasmic protein. | |||||
|
PANK4_HUMAN
|
||||||
| θ value | 0.365318 (rank : 73) | NC score | 0.037713 (rank : 126) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NVE7, Q53EU3, Q5TA84, Q7RTX3, Q9H3X5 | Gene names | PANK4 | |||
|
Domain Architecture |

|
|||||
| Description | Pantothenate kinase 4 (EC 2.7.1.33) (Pantothenic acid kinase 4) (hPanK4). | |||||
|
CP007_MOUSE
|
||||||
| θ value | 0.47712 (rank : 74) | NC score | 0.073727 (rank : 78) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8C190, Q6PFD1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C16orf7 homolog (5-day ovary-specific transcript 1 protein). | |||||
|
FIP1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 75) | NC score | 0.034889 (rank : 127) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9D824, Q8BWX7, Q99LH0, Q9DBB2 | Gene names | Fip1l1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pre-mRNA 3'-end-processing factor FIP1 (FIP1-like 1). | |||||
|
HECD1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 76) | NC score | 0.057122 (rank : 97) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 371 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9ULT8, Q6P445, Q86VJ1, Q96F34, Q9UFZ7 | Gene names | HECTD1, KIAA1131 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin-protein ligase HECTD1 (HECT domain-containing protein 1) (E3 ligase for inhibin receptor) (EULIR). | |||||
|
MYO10_HUMAN
|
||||||
| θ value | 0.62314 (rank : 77) | NC score | 0.009953 (rank : 157) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 920 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9HD67, Q9NYM7, Q9P110, Q9P111, Q9UHF6 | Gene names | MYO10 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-10 (Myosin X). | |||||
|
PRG4_HUMAN
|
||||||
| θ value | 0.62314 (rank : 78) | NC score | 0.019913 (rank : 143) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
SNX1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 79) | NC score | 0.030186 (rank : 129) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q13596, O60750, O60751 | Gene names | SNX1 | |||
|
Domain Architecture |
|
|||||
| Description | Sorting nexin-1. | |||||
|
TBCD4_MOUSE
|
||||||
| θ value | 0.813845 (rank : 80) | NC score | 0.016429 (rank : 147) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BYJ6, Q5DU23, Q66JU2, Q6P2M2, Q8BMH6, Q8BXM2 | Gene names | Tbc1d4, Kiaa0603 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 4 (Akt substrate of 160 kDa) (AS160). | |||||
|
BAG4_MOUSE
|
||||||
| θ value | 1.38821 (rank : 81) | NC score | 0.025302 (rank : 136) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CI61, Q3TRL9, Q91VT5, Q9CWG2 | Gene names | Bag4, Sodd | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 4 (BCL2-associated athanogene 4) (BAG-4) (Silencer of death domains). | |||||
|
FGD4_HUMAN
|
||||||
| θ value | 1.38821 (rank : 82) | NC score | 0.104029 (rank : 66) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q96M96, Q6ULS2, Q8TCP6 | Gene names | FGD4, FRABP, ZFYVE6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 4 (Actin filament- binding protein frabin) (FGD1-related F-actin-binding protein) (Zinc finger FYVE domain-containing protein 6). | |||||
|
POPD2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 83) | NC score | 0.027210 (rank : 133) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9HBU9, Q86UE7 | Gene names | POPDC2, POP2 | |||
|
Domain Architecture |

|
|||||
| Description | Popeye domain-containing protein 2 (Popeye protein 2). | |||||
|
ARHGA_MOUSE
|
||||||
| θ value | 1.81305 (rank : 84) | NC score | 0.081292 (rank : 77) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8C033, Q5DU38, Q80VH8, Q8BW76, Q922S7 | Gene names | Arhgef10, Kiaa0294 | |||
|
Domain Architecture |
|
|||||
| Description | Rho guanine nucleotide exchange factor 10. | |||||
|
FGD1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 85) | NC score | 0.103744 (rank : 67) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 367 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P98174, Q8N4D9 | Gene names | FGD1, ZFYVE3 | |||
|
Domain Architecture |

|
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 1 (Faciogenital dysplasia 1 protein) (Zinc finger FYVE domain-containing protein 3) (Rho/Rac guanine nucleotide exchange factor FGD1) (Rho/Rac GEF). | |||||
|
FGD3_MOUSE
|
||||||
| θ value | 1.81305 (rank : 86) | NC score | 0.104345 (rank : 65) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O88842, Q8BQ72 | Gene names | Fgd3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 3. | |||||
|
TR11B_MOUSE
|
||||||
| θ value | 1.81305 (rank : 87) | NC score | 0.027261 (rank : 132) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O08712, O70202 | Gene names | Tnfrsf11b, Ocif, Opg | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 11B precursor (Osteoprotegerin) (Osteoclastogenesis inhibitory factor). | |||||
|
CADH2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 88) | NC score | 0.002212 (rank : 165) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P19022, Q14923 | Gene names | CDH2, CDHN, NCAD | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin-2 precursor (Neural-cadherin) (N-cadherin) (CDw325 antigen). | |||||
|
CHMP3_MOUSE
|
||||||
| θ value | 2.36792 (rank : 89) | NC score | 0.026929 (rank : 134) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9CQ10, Q9D2Z2, Q9D7A5 | Gene names | Vps24, Chmp3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Charged multivesicular body protein 3 (Chromatin-modifying protein 3) (Vacuolar protein sorting-associated protein 24). | |||||
|
LPIN2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 90) | NC score | 0.022886 (rank : 138) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q92539 | Gene names | LPIN2, KIAA0249 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipin-2. | |||||
|
LPIN2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 91) | NC score | 0.022136 (rank : 139) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99PI5, Q8C357, Q8C7I8, Q8CC85, Q8CHR7 | Gene names | Lpin2 | |||
|
Domain Architecture |
|
|||||
| Description | Lipin-2. | |||||
|
PKHA1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 92) | NC score | 0.028894 (rank : 131) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9HB21, Q9BVK0 | Gene names | PLEKHA1, TAPP1 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 1 (Tandem PH domain-containing protein 1) (TAPP-1). | |||||
|
RBBP6_MOUSE
|
||||||
| θ value | 2.36792 (rank : 93) | NC score | 0.013738 (rank : 150) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
CENA1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 94) | NC score | 0.025662 (rank : 135) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O75689 | Gene names | CENTA1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centaurin-alpha 1 (Putative MAPK-activating protein PM25). | |||||
|
CRSP2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 95) | NC score | 0.020427 (rank : 141) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O60244, Q9UNB3 | Gene names | CRSP2, ARC150, DRIP150, EXLM1, TRAP170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CRSP complex subunit 2 (Cofactor required for Sp1 transcriptional activation subunit 2) (Transcriptional coactivator CRSP150) (Vitamin D3 receptor-interacting protein complex 150 kDa component) (DRIP150) (Thyroid hormone receptor-associated protein complex 170 kDa component) (Trap170) (Activator-recruited cofactor 150 kDa component) (ARC150). | |||||
|
MARK2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 96) | NC score | 0.011721 (rank : 153) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 1018 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q05512, Q6PDR4, Q8BR95 | Gene names | Mark2, Emk | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase MARK2 (EC 2.7.11.1) (MAP/microtubule affinity-regulating kinase 2) (ELKL Motif Kinase) (EMK1). | |||||
|
PRG4_MOUSE
|
||||||
| θ value | 3.0926 (rank : 97) | NC score | 0.014696 (rank : 148) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9JM99, Q3UEL1, Q3V198 | Gene names | Prg4, Msf, Szp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
BTBDG_HUMAN
|
||||||
| θ value | 4.03905 (rank : 98) | NC score | 0.020413 (rank : 142) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q32M84, Q4VXL1, Q96LN0 | Gene names | BTBD16, C10orf87 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein 16. | |||||
|
PANK4_MOUSE
|
||||||
| θ value | 4.03905 (rank : 99) | NC score | 0.024925 (rank : 137) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80YV4, Q7M751, Q8BQE9 | Gene names | Pank4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pantothenate kinase 4 (EC 2.7.1.33) (Pantothenic acid kinase 4) (mPanK4). | |||||
|
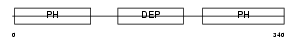
PLEK_HUMAN
|
||||||
| θ value | 4.03905 (rank : 100) | NC score | 0.033237 (rank : 128) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P08567, Q53SU8, Q6FGM8, Q8WV81 | Gene names | PLEK, P47 | |||
|
Domain Architecture |
|
|||||
| Description | Pleckstrin (Platelet p47 protein). | |||||
|
ZN292_HUMAN
|
||||||
| θ value | 4.03905 (rank : 101) | NC score | 0.005619 (rank : 164) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 918 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O60281, Q9H8G3, Q9H8J4 | Gene names | ZNF292, KIAA0530 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 292. | |||||
|
ATR_HUMAN
|
||||||
| θ value | 5.27518 (rank : 102) | NC score | 0.013993 (rank : 149) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13535, Q7KYL3, Q93051, Q9BXK4 | Gene names | ATR, FRP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase ATR (EC 2.7.11.1) (Ataxia telangiectasia and Rad3-related protein) (FRAP-related protein 1). | |||||
|
CEND1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 103) | NC score | 0.018598 (rank : 144) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 284 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8WZ64, Q4W5D2, Q7Z2L5, Q96L70, Q96P49, Q9Y4E4 | Gene names | CENTD1, ARAP2, KIAA0580 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centaurin-delta 1 (Cnt-d1) (Arf-GAP, Rho-GAP, ankyrin repeat and pleckstrin homology domain-containing protein 2) (PARX protein). | |||||
|
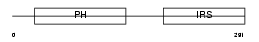
IRS2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 104) | NC score | 0.012486 (rank : 152) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y4H2, Q96RR2, Q9BZG0, Q9Y6I5 | Gene names | IRS2 | |||
|
Domain Architecture |
|
|||||
| Description | Insulin receptor substrate 2 (IRS-2). | |||||
|
MARK2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 105) | NC score | 0.010263 (rank : 156) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 1056 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q7KZI7, Q15449, Q15524, Q5XGA3, Q68A18, Q96HB3, Q96RG0 | Gene names | MARK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MARK2 (EC 2.7.11.1) (MAP/microtubule affinity-regulating kinase 2) (ELKL motif kinase) (EMK1) (PAR1 homolog). | |||||
|
PKCB1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 106) | NC score | 0.013634 (rank : 151) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 488 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9ULU4, Q13517, Q4JJ94, Q4JJ95, Q5TH09, Q6MZM1, Q8WXC5, Q9H1F3, Q9H1F4, Q9H1F5, Q9H1L8, Q9H1L9, Q9H2G5, Q9NYN3, Q9UIX6 | Gene names | PRKCBP1, KIAA1125, RACK7, ZMYND8 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C-binding protein 1 (Rack7) (Cutaneous T-cell lymphoma- associated antigen se14-3) (CTCL tumor antigen se14-3) (Zinc finger MYND domain-containing protein 8). | |||||
|
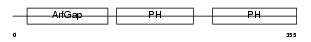
CENA2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 107) | NC score | 0.017947 (rank : 145) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NPF8, Q8N4Q6, Q96SD5 | Gene names | CENTA2 | |||
|
Domain Architecture |
|
|||||
| Description | Centaurin-alpha 2. | |||||
|
CHMP3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 108) | NC score | 0.021313 (rank : 140) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y3E7, Q53S71, Q53SU5, Q9NZ51 | Gene names | VPS24, CHMP3, NEDF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Charged multivesicular body protein 3 (Chromatin-modifying protein 3) (Vacuolar protein sorting-associated protein 24) (hVps24) (Neuroendocrine differentiation factor). | |||||
|
E41L2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 109) | NC score | 0.029830 (rank : 130) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O43491 | Gene names | EPB41L2 | |||
|
Domain Architecture |

|
|||||
| Description | Band 4.1-like protein 2 (Generally expressed protein 4.1) (4.1G). | |||||
|
GCC2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 110) | NC score | -0.000493 (rank : 167) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 1558 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8IWJ2, O15045, Q8TDH3, Q9H2G8 | Gene names | GCC2, KIAA0336 | |||
|
Domain Architecture |

|
|||||
| Description | GRIP and coiled-coil domain-containing protein 2 (Golgi coiled coil protein GCC185) (CTCL tumor antigen se1-1) (CLL-associated antigen KW- 11) (NY-REN-53 antigen). | |||||
|
JIP4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 111) | NC score | 0.010904 (rank : 155) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 585 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O60271, O60905, Q3KQU8, Q3MKM7, Q86WC7, Q86WC8, Q8IZX7, Q96II0, Q9H811 | Gene names | SPAG9, HSS, KIAA0516, MAPK8IP4, SYD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 4 (JNK-interacting protein 4) (JIP-4) (JNK-associated leucine-zipper protein) (JLP) (Sperm-associated antigen 9) (Mitogen-activated protein kinase 8- interacting protein 4) (Human lung cancer protein 6) (HLC-6) (Proliferation-inducing protein 6) (Sperm-specific protein) (Sperm surface protein) (Protein highly expressed in testis) (PHET) (Sunday driver 1). | |||||
|
PK3CD_MOUSE
|
||||||
| θ value | 6.88961 (rank : 112) | NC score | 0.009709 (rank : 159) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O35904 | Gene names | Pik3cd | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit delta isoform (EC 2.7.1.153) (PI3-kinase p110 subunit delta) (PtdIns-3- kinase p110) (PI3K) (p110delta). | |||||
|
RBP24_HUMAN
|
||||||
| θ value | 6.88961 (rank : 113) | NC score | -0.000478 (rank : 166) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q4ZG46 | Gene names | RANBP2L4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 2-like 4 (RanBP2L4). | |||||
|
SL9A7_HUMAN
|
||||||
| θ value | 6.88961 (rank : 114) | NC score | 0.006343 (rank : 162) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96T83, O75827, Q5JXP9 | Gene names | SLC9A7, NHE7 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/hydrogen exchanger 7 (Na(+)/H(+) exchanger 7) (NHE-7) (Solute carrier family 9 member 7). | |||||
|
ZN236_HUMAN
|
||||||
| θ value | 6.88961 (rank : 115) | NC score | -0.004647 (rank : 168) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 844 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UL36, Q9UL37 | Gene names | ZNF236 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 236. | |||||
|
CEND1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 116) | NC score | 0.016430 (rank : 146) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8BZ05, Q80TX2, Q8C3T2, Q8VEL6 | Gene names | Centd1, Kiaa0580 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centaurin-delta 1 (Cnt-d1). | |||||
|
EMIL2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 117) | NC score | 0.007894 (rank : 160) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 573 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9BXX0, Q8NBH3, Q96JQ4 | Gene names | EMILIN2 | |||
|
Domain Architecture |

|
|||||
| Description | EMILIN-2 precursor (Elastin microfibril interface-located protein 2) (Elastin microfibril interfacer 2) (Protein FOAP-10). | |||||
|
MYCN_MOUSE
|
||||||
| θ value | 8.99809 (rank : 118) | NC score | 0.006039 (rank : 163) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P03966, Q61978 | Gene names | Mycn, Nmyc, Nmyc1 | |||
|
Domain Architecture |

|
|||||
| Description | N-myc proto-oncogene protein. | |||||
|
PCLO_HUMAN
|
||||||
| θ value | 8.99809 (rank : 119) | NC score | 0.009925 (rank : 158) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
RFIP5_MOUSE
|
||||||
| θ value | 8.99809 (rank : 120) | NC score | 0.007074 (rank : 161) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8R361 | Gene names | Rab11fip5, D6Ertd32e, Rip11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab11 family-interacting protein 5 (Rab11-FIP5) (Rab11-interacting protein Rip11). | |||||
|
VP13A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 121) | NC score | 0.011096 (rank : 154) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96RL7, Q5JSX9, Q5JSY0, Q5VYR5, Q702P4, Q709D0, Q86YF8, Q96S61, Q9H995, Q9Y2J1 | Gene names | VPS13A, CHAC, KIAA0986 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 13A (Chorein) (Chorea- acanthocytosis protein). | |||||
|
ARHG4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 122) | NC score | 0.056535 (rank : 100) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9NR80, Q9HDC6, Q9UPP0 | Gene names | ARHGEF4, KIAA1112 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 4 (APC-stimulated guanine nucleotide exchange factor) (Asef). | |||||
|
ARHG4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 123) | NC score | 0.058331 (rank : 94) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q7TNR9, Q80TJ6 | Gene names | Arhgef4, Kiaa1112 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 4 (APC-stimulated guanine nucleotide exchange factor) (Asef). | |||||
|
ARHG7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 124) | NC score | 0.057945 (rank : 96) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q14155, Q6P9G3, Q6PII2, Q86W63, Q8N3M1 | Gene names | ARHGEF7, COOL1, KIAA0142, P85SPR, PAK3BP, PIXB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 7 (PAK-interacting exchange factor beta) (Beta-Pix) (COOL-1) (p85). | |||||
|
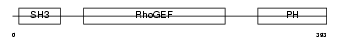
ARHG7_MOUSE
|
||||||
| θ value | θ > 10 (rank : 125) | NC score | 0.053319 (rank : 111) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 396 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9ES28, O08757, Q9ES27 | Gene names | Arhgef7, Pak3bp | |||
|
Domain Architecture |
|
|||||
| Description | Rho guanine nucleotide exchange factor 7 (PAK-interacting exchange factor beta) (Beta-Pix) (p85SPR). | |||||
|
ARHG9_HUMAN
|
||||||
| θ value | θ > 10 (rank : 126) | NC score | 0.050639 (rank : 120) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O43307, Q5JSL6 | Gene names | ARHGEF9, KIAA0424 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 9 (Rac/Cdc42 guanine nucleotide exchange factor 9) (Collybistin) (PEM-2 homolog). | |||||
|
ARHG9_MOUSE
|
||||||
| θ value | θ > 10 (rank : 127) | NC score | 0.050272 (rank : 122) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q3UTH8, Q3TQ60, Q80U06, Q8CAF9 | Gene names | Arhgef9, Kiaa0424 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 9 (Rac/Cdc42 guanine nucleotide exchange factor 9) (Collybistin). | |||||
|
CUL7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 128) | NC score | 0.050968 (rank : 118) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14999, Q5T654 | Gene names | CUL7, KIAA0076 | |||
|
Domain Architecture |

|
|||||
| Description | Cullin-7 (CUL-7). | |||||
|
ECT2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 129) | NC score | 0.054302 (rank : 109) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9H8V3, Q9NSV8, Q9NVW9 | Gene names | ECT2 | |||
|
Domain Architecture |

|
|||||
| Description | ECT2 protein (Epithelial cell-transforming sequence 2 oncogene). | |||||
|
ECT2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 130) | NC score | 0.055112 (rank : 105) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q07139 | Gene names | Ect2 | |||
|
Domain Architecture |

|
|||||
| Description | ECT2 protein (Epithelial cell-transforming sequence 2 oncogene). | |||||
|
EDD1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 131) | NC score | 0.051556 (rank : 115) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O95071, O94970, Q9NPL3 | Gene names | EDD1, EDD, HYD, KIAA0896 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase EDD1 (EC 6.3.2.-) (Hyperplastic discs protein homolog) (hHYD) (Progestin-induced protein). | |||||
|
EDD1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 132) | NC score | 0.051321 (rank : 116) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q80TP3, Q698K9, Q6PEQ8, Q6PFQ9, Q80VL4, Q810V6, Q9CXE9 | Gene names | Edd1, Edd, Kiaa0896 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase EDD1 (EC 6.3.2.-) (Hyperplastic discs protein homolog). | |||||
|
FGD1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 133) | NC score | 0.097507 (rank : 69) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P52734 | Gene names | Fgd1 | |||
|
Domain Architecture |

|
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 1 (Faciogenital dysplasia 1 protein homolog) (Zinc finger FYVE domain-containing protein 3) (Rho/Rac guanine nucleotide exchange factor FGD1) (Rho/Rac GEF). | |||||
|
FGD3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 134) | NC score | 0.094819 (rank : 71) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q5JSP0, Q7Z7D9, Q8N5G1 | Gene names | FGD3, ZFYVE5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 3 (Zinc finger FYVE domain-containing protein 5). | |||||
|
FGD5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 135) | NC score | 0.093914 (rank : 73) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6ZNL6, Q6MZY1, Q7Z303, Q8IYP3, Q8N861, Q8N8G4 | Gene names | FGD5, ZFYVE23 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 5 (Zinc finger FYVE domain-containing protein 23). | |||||
|
FGD5_MOUSE
|
||||||
| θ value | θ > 10 (rank : 136) | NC score | 0.092668 (rank : 74) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q80UZ0, Q8BHM5 | Gene names | Fgd5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 5. | |||||
|
HECD2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 137) | NC score | 0.071825 (rank : 82) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5U5R9, Q5VZ97, Q5VZ99, Q8TCP5 | Gene names | HECTD2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase HECTD2 (HECT domain-containing protein 2). | |||||
|
HECD2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 138) | NC score | 0.070882 (rank : 83) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CDU6, Q8CBQ9 | Gene names | Hectd2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase HECTD2 (HECT domain-containing protein 2). | |||||
|
HECD3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 139) | NC score | 0.084093 (rank : 76) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q5T447, Q5T448, Q9H783 | Gene names | HECTD3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase HECTD3 (HECT domain-containing protein 3). | |||||
|
HECD3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 140) | NC score | 0.084134 (rank : 75) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q3U487, Q3TN76, Q641P3, Q8BQ74, Q8R1L6 | Gene names | Hectd3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase HECTD3 (HECT domain-containing protein 3). | |||||
|
HUWE1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 141) | NC score | 0.056883 (rank : 99) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q7Z6Z7, O15029, Q4G2Z2, Q5H961, Q6P4D0, Q8NG67, Q9BUI0, Q9HCJ4, Q9NSL6, Q9P0A9 | Gene names | HUWE1, KIAA0312, KIAA1578, UREB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (Mcl-1 ubiquitin ligase E3) (Mule) (ARF-binding protein 1) (ARF-BP1). | |||||
|
HUWE1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 142) | NC score | 0.058135 (rank : 95) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q7TMY8, Q4G2Z1, Q5BMM7, Q6NS61, Q8BNJ7, Q8CFH2, Q8VD14, Q921M5, Q9R0P2 | Gene names | Huwe1, Kiaa0312, Ureb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (E3Histone). | |||||
|
ITCH_HUMAN
|
||||||
| θ value | θ > 10 (rank : 143) | NC score | 0.054736 (rank : 106) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96J02, O43584, Q5TEL0, Q96F66, Q9BY75, Q9H451, Q9H4U5 | Gene names | ITCH | |||
|
Domain Architecture |

|
|||||
| Description | Itchy homolog E3 ubiquitin protein ligase (EC 6.3.2.-) (Itch) (Atrophin-1-interacting protein 4) (AIP4) (NFE2-associated polypeptide 1) (NAPP1). | |||||
|
ITCH_MOUSE
|
||||||
| θ value | θ > 10 (rank : 144) | NC score | 0.055163 (rank : 104) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8C863, O54971 | Gene names | Itch | |||
|
Domain Architecture |

|
|||||
| Description | Itchy E3 ubiquitin protein ligase (EC 6.3.2.-). | |||||
|
KR202_HUMAN
|
||||||
| θ value | θ > 10 (rank : 145) | NC score | 0.057065 (rank : 98) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 4 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q3LI61 | Gene names | KRTAP20-2, KAP20.2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 20-2. | |||||
|
LORI_HUMAN
|
||||||
| θ value | θ > 10 (rank : 146) | NC score | 0.073470 (rank : 79) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P23490 | Gene names | LOR, LRN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Loricrin. | |||||
|
LORI_MOUSE
|
||||||
| θ value | θ > 10 (rank : 147) | NC score | 0.067303 (rank : 85) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P18165, Q8R0I9 | Gene names | Lor | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Loricrin. | |||||
|
NED4L_HUMAN
|
||||||
| θ value | θ > 10 (rank : 148) | NC score | 0.050560 (rank : 121) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96PU5, O43165, Q7Z5F1, Q7Z5F2, Q7Z5N3, Q8N5A7, Q8WUU9, Q9BW58, Q9H2W4, Q9NT88 | Gene names | NEDD4L, KIAA0439, NEDL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin-protein ligase NEDD4-like protein (EC 6.3.2.-) (Nedd4-2) (NEDD4.2). | |||||
|
NED4L_MOUSE
|
||||||
| θ value | θ > 10 (rank : 149) | NC score | 0.050715 (rank : 119) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8CFI0, Q8BRT9, Q8BS42, Q99PK2 | Gene names | Nedd4l, Kiaa0439, Nedd4b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin-protein ligase NEDD4-like protein (EC 6.3.2.-) (Nedd4-2) (NEDD4.2). | |||||
|
NEDD4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 150) | NC score | 0.050030 (rank : 123) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P46934 | Gene names | NEDD4, KIAA0093 | |||
|
Domain Architecture |

|
|||||
| Description | E3 ubiquitin-protein ligase NEDD4 (EC 6.3.2.-). | |||||
|
NEDD4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 151) | NC score | 0.051152 (rank : 117) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P46935, O08758, Q3UZI2, Q8BGB3 | Gene names | Nedd4, Kiaa0093, Nedd-4, Nedd4a | |||
|
Domain Architecture |

|
|||||
| Description | E3 ubiquitin-protein ligase NEDD4 (EC 6.3.2.-) (Neural precursor cell expressed developmentally down-regulated protein 4). | |||||
|
PKHG1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 152) | NC score | 0.059327 (rank : 93) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9ULL1, Q5T1F2 | Gene names | PLEKHG1, KIAA1209 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family G member 1. | |||||
|
PREX1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 153) | NC score | 0.066502 (rank : 86) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8TCU6, Q5JS95, Q5JS96, Q69YL2, Q7Z2L9, Q9BQH0, Q9BX55, Q9H4Q6, Q9P2D2, Q9UGQ4 | Gene names | PREX1, KIAA1415 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 3,4,5-trisphosphate-dependent Rac exchanger 1 protein (P-Rex1 protein). | |||||
|
RIN1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 154) | NC score | 0.059529 (rank : 92) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q13671, O15010, Q00427, Q96CC8 | Gene names | RIN1 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 1 (Ras interaction/interference protein 1) (Ras inhibitor JC99). | |||||
|
RIN1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 155) | NC score | 0.060753 (rank : 91) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q921Q7 | Gene names | Rin1 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 1 (Ras interaction/interference protein 1). | |||||
|
SMUF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 156) | NC score | 0.055775 (rank : 102) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9HCE7, O75853, Q9UJT8 | Gene names | SMURF1, KIAA1625 | |||
|
Domain Architecture |

|
|||||
| Description | Smad ubiquitination regulatory factor 1 (EC 6.3.2.-) (Ubiquitin-- protein ligase SMURF1) (Smad-specific E3 ubiquitin ligase 1) (hSMURF1). | |||||
|
SMUF1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 157) | NC score | 0.055501 (rank : 103) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9CUN6, Q3U412, Q8K300 | Gene names | Smurf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Smad ubiquitination regulatory factor 1 (EC 6.3.2.-) (Ubiquitin-- protein ligase SMURF1) (Smad-specific E3 ubiquitin ligase 1). | |||||
|
SMUF2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 158) | NC score | 0.055981 (rank : 101) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9HAU4, Q9H260 | Gene names | SMURF2 | |||
|
Domain Architecture |

|
|||||
| Description | Smad ubiquitination regulatory factor 2 (EC 6.3.2.-) (Ubiquitin-- protein ligase SMURF2) (Smad-specific E3 ubiquitin ligase 2) (hSMURF2). | |||||
|
TRIPC_HUMAN
|
||||||
| θ value | θ > 10 (rank : 159) | NC score | 0.065559 (rank : 87) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 319 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q14669, Q15644 | Gene names | TRIP12, KIAA0045 | |||
|
Domain Architecture |

|
|||||
| Description | Thyroid receptor-interacting protein 12 (TRIP12). | |||||
|
UBE3A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 160) | NC score | 0.073399 (rank : 80) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q05086, P78355, Q93066, Q9UEP4, Q9UEP5, Q9UEP6, Q9UEP7, Q9UEP8, Q9UEP9 | Gene names | UBE3A, E6AP | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase E3A (EC 6.3.2.-) (E6AP ubiquitin-protein ligase) (Oncogenic protein-associated protein E6-AP) (Human papillomavirus E6-associated protein) (NY-REN-54 antigen). | |||||
|
UBE3A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 161) | NC score | 0.072923 (rank : 81) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O08759, P97482 | Gene names | Ube3a | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase E3A (EC 6.3.2.-) (Oncogenic protein- associated protein E6-AP). | |||||
|
UBE3C_HUMAN
|
||||||
| θ value | θ > 10 (rank : 162) | NC score | 0.061455 (rank : 89) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q15386, Q8TC15, Q96CR4, Q9UDU3 | Gene names | UBE3C, KIAA0010 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase E3C (EC 6.3.2.-). | |||||
|
UBE3C_MOUSE
|
||||||
| θ value | θ > 10 (rank : 163) | NC score | 0.061269 (rank : 90) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80U95, Q8BQZ6, Q8C7W6, Q8CDJ1, Q8VDL5 | Gene names | Ube3c, Kiaa0010 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase E3C (EC 6.3.2.-). | |||||
|
WWP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 164) | NC score | 0.052370 (rank : 114) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9H0M0, O00307, Q96BP4 | Gene names | WWP1 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP1 (EC 6.3.2.-) (WW domain- containing protein 1) (Atropin-1-interacting protein 5) (AIP5). | |||||
|
WWP1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 165) | NC score | 0.052745 (rank : 113) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BZZ3, Q8BIV9, Q8VDP8 | Gene names | Wwp1 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP1 (EC 6.3.2.-) (WW domain- containing protein 1). | |||||
|
WWP2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 166) | NC score | 0.054646 (rank : 108) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O00308, Q96CZ2, Q9BWN6 | Gene names | WWP2 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP2 (EC 6.3.2.-) (WW domain- containing protein 2) (Atrophin-1-interacting protein 2) (AIP2). | |||||
|
WWP2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 167) | NC score | 0.054736 (rank : 107) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9DBH0, Q8BTG4, Q923F6 | Gene names | Wwp2 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP2 (EC 6.3.2.-) (WW domain- containing protein 2). | |||||
|
ZFY28_HUMAN
|
||||||
| θ value | θ > 10 (rank : 168) | NC score | 0.052982 (rank : 112) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9HCC9 | Gene names | ZFYVE28, KIAA1643 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger FYVE domain-containing protein 28. | |||||
|
ALS2_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 121 | |
| SwissProt Accessions | Q96Q42, Q8N1E0, Q96PC4, Q96Q41, Q9H973, Q9HCK9 | Gene names | ALS2, KIAA1563 | |||
|
Domain Architecture |

|
|||||
| Description | Alsin (Amyotrophic lateral sclerosis protein 2). | |||||
|
ALS2_MOUSE
|
||||||
| NC score | 0.993643 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 90 | |
| SwissProt Accessions | Q920R0, Q8JZR1, Q9CXJ3 | Gene names | Als2 | |||
|
Domain Architecture |

|
|||||
| Description | Alsin (Amyotrophic lateral sclerosis protein 2 homolog). | |||||
|
RCC2_HUMAN
|
||||||
| NC score | 0.567947 (rank : 3) | θ value | 1.2105e-12 (rank : 17) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9P258, Q8IVL9, Q9BSN6, Q9NPV8 | Gene names | RCC2, KIAA1470, TD60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein RCC2 (Telophase disk protein of 60 kDa) (RCC1-like protein TD- 60). | |||||
|
RCC2_MOUSE
|
||||||
| NC score | 0.567581 (rank : 4) | θ value | 1.58096e-12 (rank : 19) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8BK67, Q6ZPQ0 | Gene names | Rcc2, Kiaa1470 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein RCC2. | |||||
|
SRGEF_MOUSE
|
||||||
| NC score | 0.542221 (rank : 5) | θ value | 2.79066e-09 (rank : 30) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q80YD6, Q9QXB7 | Gene names | Sergef, Delgef, Gnefr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Secretion-regulating guanine nucleotide exchange factor (Guanine nucleotide exchange factor-related protein) (Deafness locus-associated putative guanine nucleotide exchange factor) (DelGEF). | |||||
|
TSGA2_HUMAN
|
||||||
| NC score | 0.539903 (rank : 6) | θ value | 6.85773e-16 (rank : 5) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8WYR4 | Gene names | TSGA2, TSA2 | |||
|
Domain Architecture |

|
|||||
| Description | Testis-specific gene A2 protein (Male meiotic metaphase chromosome- associated acidic protein) (Meichroacidin). | |||||
|
SRGEF_HUMAN
|
||||||
| NC score | 0.536960 (rank : 7) | θ value | 1.80886e-08 (rank : 31) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9UGK8, Q9UGK9 | Gene names | SERGEF, DELGEF, GNEFR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Secretion-regulating guanine nucleotide exchange factor (Guanine nucleotide exchange factor-related protein) (Deafness locus-associated putative guanine nucleotide exchange factor) (DelGEF). | |||||
|
RPGR_MOUSE
|
||||||
| NC score | 0.530670 (rank : 8) | θ value | 1.86753e-13 (rank : 13) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9R0X5, O88408, Q9CU92 | Gene names | Rpgr | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator (mRpgr). | |||||
|
RPGR_HUMAN
|
||||||
| NC score | 0.526639 (rank : 9) | θ value | 5.43371e-13 (rank : 15) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q92834, O00702, O00737, Q8N5T6, Q93039, Q9HD29, Q9UMR1 | Gene names | RPGR, RP3, XLRP3 | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator. | |||||
|
TSGA2_MOUSE
|
||||||
| NC score | 0.522938 (rank : 10) | θ value | 1.09485e-13 (rank : 11) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8VIG3, Q9DAL5 | Gene names | Tsga2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-specific gene A2 protein (Male meiotic metaphase chromosome- associated acidic protein) (Meichroacidin). | |||||
|
RCC1_MOUSE
|
||||||
| NC score | 0.522577 (rank : 11) | θ value | 6.87365e-08 (rank : 32) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8VE37, Q3UDB6 | Gene names | Rcc1, Chc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of chromosome condensation (Chromosome condensation protein 1). | |||||
|
MORN1_HUMAN
|
||||||
| NC score | 0.522228 (rank : 12) | θ value | 3.40345e-15 (rank : 9) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q5T089, Q8WW30, Q9H852 | Gene names | MORN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORN repeat-containing protein 1. | |||||
|
MORN3_MOUSE
|
||||||
| NC score | 0.522224 (rank : 13) | θ value | 1.09485e-13 (rank : 10) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8C5T4, Q8VE21, Q9D5H6 | Gene names | Morn3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORN repeat-containing protein 3. | |||||
|
MORN3_HUMAN
|
||||||
| NC score | 0.518648 (rank : 14) | θ value | 3.18553e-13 (rank : 14) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6PF18, Q86YQ9 | Gene names | MORN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORN repeat-containing protein 3. | |||||
|
RCC1_HUMAN
|
||||||
| NC score | 0.518430 (rank : 15) | θ value | 7.59969e-07 (rank : 36) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P18754 | Gene names | RCC1, CHC1 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of chromosome condensation (Chromosome condensation protein 1) (Cell cycle regulatory protein). | |||||
|
WBS16_HUMAN
|
||||||
| NC score | 0.508561 (rank : 16) | θ value | 9.92553e-07 (rank : 38) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q96I51, Q548B1, Q9H0G7 | Gene names | WBSCR16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Williams-Beuren syndrome chromosome region 16 protein (RCC1-like G exchanging factor-like protein). | |||||
|
WBS16_MOUSE
|
||||||
| NC score | 0.506162 (rank : 17) | θ value | 7.59969e-07 (rank : 37) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9CYF5 | Gene names | Wbscr16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Williams-Beuren syndrome chromosome region 16 protein homolog. | |||||
|
RCBT2_MOUSE
|
||||||
| NC score | 0.483813 (rank : 18) | θ value | 1.42992e-13 (rank : 12) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q99LJ7, Q3TUA3, Q8BMG2 | Gene names | Rcbtb2, Chc1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RCC1 and BTB domain-containing protein 2 (Regulator of chromosome condensation and BTB domain-containing protein 2) (Chromosome condensation 1-like). | |||||
|
RCBT2_HUMAN
|
||||||
| NC score | 0.483673 (rank : 19) | θ value | 1.58096e-12 (rank : 18) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O95199 | Gene names | RCBTB2, CHC1L, RLG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RCC1 and BTB domain-containing protein 2 (Regulator of chromosome condensation and BTB domain-containing protein 2) (Chromosome condensation 1-like) (CHC1-L) (RCC1-like G exchanging factor). | |||||
|
RCBT1_MOUSE
|
||||||
| NC score | 0.476897 (rank : 20) | θ value | 1.058e-16 (rank : 4) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q6NXM2, Q8BTZ6, Q8BZV0 | Gene names | Rcbtb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RCC1 and BTB domain-containing protein 1 (Regulator of chromosome condensation and BTB domain-containing protein 1). | |||||
|
RCBT1_HUMAN
|
||||||
| NC score | 0.474526 (rank : 21) | θ value | 1.16975e-15 (rank : 8) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8NDN9, Q969U9 | Gene names | RCBTB1, CLLD7, E4.5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RCC1 and BTB domain-containing protein 1 (Regulator of chromosome condensation and BTB domain-containing protein 1) (Chronic lymphocytic leukemia deletion region gene 7 protein) (CLL deletion region gene 7 protein). | |||||
|
JPH3_HUMAN
|
||||||
| NC score | 0.422168 (rank : 22) | θ value | 3.89403e-11 (rank : 23) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8WXH2, Q8N471, Q9HDC3, Q9HDC4 | Gene names | JPH3, JP3 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-3 (Junctophilin type 3) (JP-3). | |||||
|
JPH3_MOUSE
|
||||||
| NC score | 0.422158 (rank : 23) | θ value | 3.89403e-11 (rank : 24) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9ET77, Q9EQZ2 | Gene names | Jph3, Jp3 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-3 (Junctophilin type 3) (JP-3). | |||||
|
HERC2_HUMAN
|
||||||
| NC score | 0.416483 (rank : 24) | θ value | 8.95645e-16 (rank : 6) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O95714, Q86SV7, Q86SV8, Q86SV9, Q86YY3, Q86YY4, Q86YY5, Q86YY6, Q86YY7, Q86YY8, Q86YY9, Q86YZ0, Q86YZ1 | Gene names | HERC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT domain and RCC1-like domain-containing protein 2. | |||||
|
HERC3_HUMAN
|
||||||
| NC score | 0.415846 (rank : 25) | θ value | 1.63225e-17 (rank : 3) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q15034 | Gene names | HERC3, KIAA0032 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT domain and RCC1-like domain protein 3. | |||||
|
HERC2_MOUSE
|
||||||
| NC score | 0.414944 (rank : 26) | θ value | 8.95645e-16 (rank : 7) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q4U2R1, O88473, Q3TRJ8, Q3TS47, Q3TST2, Q3UFQ6, Q3URH7, Q5DU32, Q7TPR5, Q80VV7, Q9QYT1, Q9Z168, Q9Z171 | Gene names | Herc2, jdf2, Kiaa0393, rjs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT domain and RCC1-like domain-containing protein 2. | |||||
|
JPH1_MOUSE
|
||||||
| NC score | 0.413604 (rank : 27) | θ value | 1.9326e-10 (rank : 25) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9ET80, Q9EQZ3 | Gene names | Jph1, Jp1 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-1 (Junctophilin type 1) (JP-1). | |||||
|
JPH1_HUMAN
|
||||||
| NC score | 0.411031 (rank : 28) | θ value | 1.25267e-09 (rank : 27) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9HDC5 | Gene names | JPH1, JP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junctophilin-1 (Junctophilin type 1) (JP-1). | |||||
|
JPH4_HUMAN
|
||||||
| NC score | 0.409207 (rank : 29) | θ value | 2.79066e-09 (rank : 28) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96JJ6, Q8ND44, Q96DQ0 | Gene names | JPH4, JPHL1, KIAA1831 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junctophilin-4 (Junctophilin-like 1 protein). | |||||
|
JPH2_MOUSE
|
||||||
| NC score | 0.407916 (rank : 30) | θ value | 2.98157e-11 (rank : 22) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9ET78, Q9ET79 | Gene names | Jph2, Jp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junctophilin-2 (Junctophilin type 2) (JP-2). | |||||
|
JPH4_MOUSE
|
||||||
| NC score | 0.406507 (rank : 31) | θ value | 2.79066e-09 (rank : 29) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q80WT0, Q8BMI1 | Gene names | Jph4, Jphl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junctophilin-4 (Junctophilin-like 1 protein). | |||||
|
HERC5_HUMAN
|
||||||
| NC score | 0.404522 (rank : 32) | θ value | 9.26847e-13 (rank : 16) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9UII4 | Gene names | HERC5, CEB1, CEBP1 | |||
|
Domain Architecture |

|
|||||
| Description | HECT domain and RCC1-like domain-containing protein 5 (Cyclin-E- binding protein 1). | |||||
|
JPH2_HUMAN
|
||||||
| NC score | 0.382160 (rank : 33) | θ value | 2.98157e-11 (rank : 21) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 639 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9BR39, O95913, Q5JY74, Q9UJN4 | Gene names | JPH2, JP2 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-2 (Junctophilin type 2) (JP-2). | |||||
|
MORN2_HUMAN
|
||||||
| NC score | 0.368749 (rank : 34) | θ value | 0.00665767 (rank : 57) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q502X0, Q6UL00 | Gene names | MORN2, MOPT | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORN repeat-containing protein 2 (MORN motif protein in testis). | |||||
|
MORN2_MOUSE
|
||||||
| NC score | 0.351045 (rank : 35) | θ value | 0.0148317 (rank : 59) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6UL01 | Gene names | Morn2, Mopt | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORN repeat-containing protein 2 (MORN motif protein in testis). | |||||
|
MYCB2_HUMAN
|
||||||
| NC score | 0.331789 (rank : 36) | θ value | 1.17247e-07 (rank : 33) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 390 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O75592, Q5JSX8, Q5VZN6, Q6PIB6, Q9UQ11, Q9Y6E4 | Gene names | MYCBP2, KIAA0916, PAM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ubiquitin ligase protein MYCBP2 (EC 6.3.2.-) (Myc-binding protein 2) (Protein associated with Myc) (Pam/highwire/rpm-1 protein). | |||||
|
MYCB2_MOUSE
|
||||||
| NC score | 0.325887 (rank : 37) | θ value | 1.29631e-06 (rank : 39) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q7TPH6, Q6PCM8 | Gene names | Mycbp2, Pam, Phr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ubiquitin ligase protein MYCBP2 (EC 6.3.2.-) (Myc-binding protein 2) (Protein associated with Myc) (Pam/highwire/rpm-1 protein). | |||||
|
FBX24_HUMAN
|
||||||
| NC score | 0.234567 (rank : 38) | θ value | 0.163984 (rank : 67) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O75426, Q9H0G1 | Gene names | FBXO24, FBX24 | |||
|
Domain Architecture |

|
|||||
| Description | F-box only protein 24. | |||||
|
SETD7_MOUSE
|
||||||
| NC score | 0.228638 (rank : 39) | θ value | 0.0252991 (rank : 62) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8VHL1 | Gene names | Setd7, Set7 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET7 (EC 2.1.1.43) (Histone H3-K4 methyltransferase) (H3-K4-HMTase) (SET domain-containing protein 7). | |||||
|
SETD7_HUMAN
|
||||||
| NC score | 0.228384 (rank : 40) | θ value | 0.0252991 (rank : 61) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8WTS6, Q9C0E6 | Gene names | SETD7, KIAA1717, SET7 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET7 (EC 2.1.1.43) (Histone H3-K4 methyltransferase) (H3-K4-HMTase) (SET domain-containing protein 7) (Set9) (SET7/9). | |||||
|
ANKY1_HUMAN
|
||||||
| NC score | 0.211279 (rank : 41) | θ value | 0.000158464 (rank : 43) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9P2S6, Q8IYX5, Q8NDK5, Q9NX10 | Gene names | ANKMY1, TSAL1, ZMYND13 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin repeat and MYND domain-containing protein 1 (Testis-specific ankyrin-like protein 1) (Zinc-finger MYND domain-containing protein 13). | |||||
|
FBX24_MOUSE
|
||||||
| NC score | 0.210135 (rank : 42) | θ value | 0.21417 (rank : 70) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9D417 | Gene names | Fbxo24, Fbx24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 24. | |||||
|
PKHF2_HUMAN
|
||||||
| NC score | 0.198113 (rank : 43) | θ value | 1.87187e-05 (rank : 40) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9H8W4 | Gene names | PLEKHF2, ZFYVE18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family F member 2 (PH domain- containing family F member 2) (PH and FYVE domain-containing protein 2) (Phafin-2) (Zinc finger FYVE domain-containing protein 18). | |||||
|
PKHF2_MOUSE
|
||||||
| NC score | 0.196619 (rank : 44) | θ value | 2.44474e-05 (rank : 41) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q91WB4 | Gene names | Plekhf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family F member 2 (PH domain- containing family F member 2). | |||||
|
PKHF1_MOUSE
|
||||||
| NC score | 0.186109 (rank : 45) | θ value | 0.000786445 (rank : 47) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q3TB82, Q99M16 | Gene names | Plekhf1, Lapf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family F member 1 (PH domain- containing family F member 1) (Lysosome-associated apoptosis-inducing protein containing PH and FYVE domains). | |||||
|
PKHF1_HUMAN
|
||||||
| NC score | 0.179832 (rank : 46) | θ value | 0.00228821 (rank : 52) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q96S99, Q96K11, Q9BUB9 | Gene names | PLEKHF1, APPD, LAPF, ZFYVE15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family F member 1 (PH domain- containing family F member 1) (PH and FYVE domain-containing protein 1) (Phafin-1) (Lysosome-associated apoptosis-inducing protein containing PH and FYVE domains) (Apoptosis-inducing protein) (Zinc finger FYVE domain-containing protein 15). | |||||
|
RABX5_MOUSE
|
||||||
| NC score | 0.145957 (rank : 47) | θ value | 0.00390308 (rank : 54) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9JM13, Q3UIW0 | Gene names | Rabgef1, Rabex5 | |||
|
Domain Architecture |

|
|||||
| Description | Rab5 GDP/GTP exchange factor (Rabex-5). | |||||
|
FGD6_HUMAN
|
||||||
| NC score | 0.142912 (rank : 48) | θ value | 0.00102713 (rank : 48) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 237 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6ZV73, Q6ZR53, Q7Z2Z7, Q96D44, Q9NUR8, Q9P2I5 | Gene names | FGD6, KIAA1362, ZFYVE24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 6 (Zinc finger FYVE domain-containing protein 24). | |||||
|
FGD6_MOUSE
|
||||||
| NC score | 0.142519 (rank : 49) | θ value | 0.00228821 (rank : 51) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q69ZL1, Q8C8W5, Q8K3B0, Q9D3Y7 | Gene names | Fgd6, Kiaa1362 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 6. | |||||
|
RIN3_MOUSE
|
||||||
| NC score | 0.140151 (rank : 50) | θ value | 0.00102713 (rank : 49) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P59729 | Gene names | Rin3 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 3 (Ras interaction/interference protein 3). | |||||
|
FGD2_MOUSE
|
||||||
| NC score | 0.138470 (rank : 51) | θ value | 0.000270298 (rank : 45) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BY35, O88841, Q7TSE3, Q8VDH4 | Gene names | Fgd2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 2. | |||||
|
RIN3_HUMAN
|
||||||
| NC score | 0.138401 (rank : 52) | θ value | 0.00228821 (rank : 53) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8TB24, Q8NF30, Q8TEE8, Q8WYP4, Q9H6A5, Q9HAG1 | Gene names | RIN3 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 3 (Ras interaction/interference protein 3). | |||||
|
RIN2_HUMAN
|
||||||
| NC score | 0.136578 (rank : 53) | θ value | 0.00390308 (rank : 55) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8WYP3, Q00425, Q5TFT8, Q9BQL3, Q9H071 | Gene names | RIN2, RASSF4 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 2 (Ras interaction/interference protein 2) (Ras inhibitor JC265) (Ras association domain family 4). | |||||
|
RIN2_MOUSE
|
||||||
| NC score | 0.135705 (rank : 54) | θ value | 0.00390308 (rank : 56) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9D684, Q99K06 | Gene names | Rin2 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 2 (Ras interaction/interference protein 2). | |||||
|
NEK8_HUMAN
|
||||||
| NC score | 0.131797 (rank : 55) | θ value | 4.59992e-12 (rank : 20) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 872 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q86SG6, Q2M1S6, Q8NDH1 | Gene names | NEK8, JCK, NEK12A | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Nek8 (EC 2.7.11.1) (Never in mitosis A-related kinase 8) (NimA-related protein kinase 8) (NIMA-related kinase 12a). | |||||
|
NEK9_MOUSE
|
||||||
| NC score | 0.130951 (rank : 56) | θ value | 5.81887e-07 (rank : 35) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 890 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8K1R7, Q8R3P1 | Gene names | Nek9, Nercc | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Nek9 (EC 2.7.11.1) (NimA-related protein kinase 9) (Never in mitosis A-related kinase 9). | |||||
|
NEK8_MOUSE
|
||||||
| NC score | 0.130741 (rank : 57) | θ value | 4.30538e-10 (rank : 26) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 872 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q91ZR4, Q3U498, Q9D685 | Gene names | Nek8, Jck | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Nek8 (EC 2.7.11.1) (Never in mitosis A-related kinase 8) (NimA-related protein kinase 8). | |||||
|
NEK9_HUMAN
|
||||||
| NC score | 0.130259 (rank : 58) | θ value | 2.61198e-07 (rank : 34) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 900 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8TD19, Q52LK6, Q8NCN0, Q8TCY4, Q9UPI4, Q9Y6S4, Q9Y6S5, Q9Y6S6 | Gene names | NEK9, KIAA1995, NEK8, NERCC | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Nek9 (EC 2.7.11.1) (NimA-related protein kinase 9) (Never in mitosis A-related kinase 9) (Nercc1 kinase) (NIMA-related kinase 8) (Nek8). | |||||
|
RABX5_HUMAN
|
||||||
| NC score | 0.124671 (rank : 59) | θ value | 0.0193708 (rank : 60) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UJ41 | Gene names | RABGEF1, RABEX5 | |||
|
Domain Architecture |

|
|||||
| Description | Rab5 GDP/GTP exchange factor (Rabex-5) (Rabaptin-5-associated exchange factor for Rab5). | |||||
|
FARP1_HUMAN
|
||||||
| NC score | 0.120430 (rank : 60) | θ value | 7.1131e-05 (rank : 42) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9Y4F1, Q6IQ29 | Gene names | FARP1, CDEP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FERM, RhoGEF and pleckstrin domain-containing protein 1 (Chondrocyte- derived ezrin-like protein). | |||||
|
FGD2_HUMAN
|
||||||
| NC score | 0.120354 (rank : 61) | θ value | 0.279714 (rank : 71) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q7Z6J4, Q5T8I1, Q6P6A8, Q6ZNL5, Q8IZ32, Q8N868, Q9H7M2 | Gene names | FGD2, ZFYVE4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 2 (Zinc finger FYVE domain-containing protein 4). | |||||
|
FGD4_MOUSE
|
||||||
| NC score | 0.118563 (rank : 62) | θ value | 0.0563607 (rank : 65) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q91ZT5, Q3UEB6, Q8BW60, Q8BZI7, Q91ZT3, Q91ZT4 | Gene names | Fgd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 4 (Actin filament- binding protein frabin) (FGD1-related F-actin-binding protein). | |||||
|
FARP2_MOUSE
|
||||||
| NC score | 0.112810 (rank : 63) | θ value | 0.00035302 (rank : 46) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q91VS8, Q3TAP2, Q69ZZ0 | Gene names | Farp2, Kiaa0793 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FERM, RhoGEF and pleckstrin domain-containing protein 2 (FERM domain including RhoGEF) (FIR). | |||||
|
ARHG6_HUMAN
|
||||||
| NC score | 0.106530 (rank : 64) | θ value | 0.000270298 (rank : 44) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 376 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q15052, Q15396, Q5JQ66, Q86XH0 | Gene names | ARHGEF6, COOL2, KIAA0006, PIXA | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 6 (Rac/Cdc42 guanine nucleotide exchange factor 6) (PAK-interacting exchange factor alpha) (Alpha-Pix) (COOL-2). | |||||
|
FGD3_MOUSE
|
||||||
| NC score | 0.104345 (rank : 65) | θ value | 1.81305 (rank : 86) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O88842, Q8BQ72 | Gene names | Fgd3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 3. | |||||
|
FGD4_HUMAN
|
||||||
| NC score | 0.104029 (rank : 66) | θ value | 1.38821 (rank : 82) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q96M96, Q6ULS2, Q8TCP6 | Gene names | FGD4, FRABP, ZFYVE6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 4 (Actin filament- binding protein frabin) (FGD1-related F-actin-binding protein) (Zinc finger FYVE domain-containing protein 6). | |||||
|
FGD1_HUMAN
|
||||||
| NC score | 0.103744 (rank : 67) | θ value | 1.81305 (rank : 85) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 367 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P98174, Q8N4D9 | Gene names | FGD1, ZFYVE3 | |||
|
Domain Architecture |

|
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 1 (Faciogenital dysplasia 1 protein) (Zinc finger FYVE domain-containing protein 3) (Rho/Rac guanine nucleotide exchange factor FGD1) (Rho/Rac GEF). | |||||
|
ARHG6_MOUSE
|
||||||
| NC score | 0.098573 (rank : 68) | θ value | 0.00228821 (rank : 50) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8K4I3, Q8C9V4 | Gene names | Arhgef6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 6 (Rac/Cdc42 guanine nucleotide exchange factor 6). | |||||
|
FGD1_MOUSE
|
||||||
| NC score | 0.097507 (rank : 69) | θ value | θ > 10 (rank : 133) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P52734 | Gene names | Fgd1 | |||
|
Domain Architecture |

|
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 1 (Faciogenital dysplasia 1 protein homolog) (Zinc finger FYVE domain-containing protein 3) (Rho/Rac guanine nucleotide exchange factor FGD1) (Rho/Rac GEF). | |||||
|
ARHGA_HUMAN
|
||||||
| NC score | 0.095502 (rank : 70) | θ value | 0.0736092 (rank : 66) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O15013, O14665, Q68D55, Q8IWD9, Q8IY77 | Gene names | ARHGEF10, KIAA0294 | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 10. | |||||
|
FGD3_HUMAN
|
||||||
| NC score | 0.094819 (rank : 71) | θ value | θ > 10 (rank : 134) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q5JSP0, Q7Z7D9, Q8N5G1 | Gene names | FGD3, ZFYVE5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 3 (Zinc finger FYVE domain-containing protein 5). | |||||
|
FARP2_HUMAN
|
||||||
| NC score | 0.094352 (rank : 72) | θ value | 0.0330416 (rank : 63) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O94887, Q53QM5, Q8WU27, Q9UFE7 | Gene names | FARP2, KIAA0793 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FERM, RhoGEF and pleckstrin domain-containing protein 2 (FERM domain including RhoGEF) (FIR). | |||||
|
FGD5_HUMAN
|
||||||
| NC score | 0.093914 (rank : 73) | θ value | θ > 10 (rank : 135) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6ZNL6, Q6MZY1, Q7Z303, Q8IYP3, Q8N861, Q8N8G4 | Gene names | FGD5, ZFYVE23 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 5 (Zinc finger FYVE domain-containing protein 23). | |||||
|
FGD5_MOUSE
|
||||||
| NC score | 0.092668 (rank : 74) | θ value | θ > 10 (rank : 136) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q80UZ0, Q8BHM5 | Gene names | Fgd5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 5. | |||||
|
HECD3_MOUSE
|
||||||
| NC score | 0.084134 (rank : 75) | θ value | θ > 10 (rank : 140) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q3U487, Q3TN76, Q641P3, Q8BQ74, Q8R1L6 | Gene names | Hectd3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase HECTD3 (HECT domain-containing protein 3). | |||||
|
HECD3_HUMAN
|
||||||
| NC score | 0.084093 (rank : 76) | θ value | θ > 10 (rank : 139) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q5T447, Q5T448, Q9H783 | Gene names | HECTD3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase HECTD3 (HECT domain-containing protein 3). | |||||
|
ARHGA_MOUSE
|
||||||
| NC score | 0.081292 (rank : 77) | θ value | 1.81305 (rank : 84) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8C033, Q5DU38, Q80VH8, Q8BW76, Q922S7 | Gene names | Arhgef10, Kiaa0294 | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 10. | |||||
|
CP007_MOUSE
|
||||||
| NC score | 0.073727 (rank : 78) | θ value | 0.47712 (rank : 74) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8C190, Q6PFD1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C16orf7 homolog (5-day ovary-specific transcript 1 protein). | |||||
|
LORI_HUMAN
|
||||||
| NC score | 0.073470 (rank : 79) | θ value | θ > 10 (rank : 146) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P23490 | Gene names | LOR, LRN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Loricrin. | |||||
|
UBE3A_HUMAN
|
||||||
| NC score | 0.073399 (rank : 80) | θ value | θ > 10 (rank : 160) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q05086, P78355, Q93066, Q9UEP4, Q9UEP5, Q9UEP6, Q9UEP7, Q9UEP8, Q9UEP9 | Gene names | UBE3A, E6AP | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase E3A (EC 6.3.2.-) (E6AP ubiquitin-protein ligase) (Oncogenic protein-associated protein E6-AP) (Human papillomavirus E6-associated protein) (NY-REN-54 antigen). | |||||
|
UBE3A_MOUSE
|
||||||
| NC score | 0.072923 (rank : 81) | θ value | θ > 10 (rank : 161) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O08759, P97482 | Gene names | Ube3a | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase E3A (EC 6.3.2.-) (Oncogenic protein- associated protein E6-AP). | |||||
|
HECD2_HUMAN
|
||||||
| NC score | 0.071825 (rank : 82) | θ value | θ > 10 (rank : 137) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5U5R9, Q5VZ97, Q5VZ99, Q8TCP5 | Gene names | HECTD2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase HECTD2 (HECT domain-containing protein 2). | |||||
|
HECD2_MOUSE
|
||||||
| NC score | 0.070882 (rank : 83) | θ value | θ > 10 (rank : 138) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CDU6, Q8CBQ9 | Gene names | Hectd2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase HECTD2 (HECT domain-containing protein 2). | |||||
|
VAV2_HUMAN
|
||||||
| NC score | 0.068052 (rank : 84) | θ value | 0.00869519 (rank : 58) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 453 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P52735 | Gene names | VAV2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-2. | |||||
|
LORI_MOUSE
|
||||||
| NC score | 0.067303 (rank : 85) | θ value | θ > 10 (rank : 147) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P18165, Q8R0I9 | Gene names | Lor | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Loricrin. | |||||
|
PREX1_HUMAN
|
||||||
| NC score | 0.066502 (rank : 86) | θ value | θ > 10 (rank : 153) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8TCU6, Q5JS95, Q5JS96, Q69YL2, Q7Z2L9, Q9BQH0, Q9BX55, Q9H4Q6, Q9P2D2, Q9UGQ4 | Gene names | PREX1, KIAA1415 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 3,4,5-trisphosphate-dependent Rac exchanger 1 protein (P-Rex1 protein). | |||||
|
TRIPC_HUMAN
|
||||||
| NC score | 0.065559 (rank : 87) | θ value | θ > 10 (rank : 159) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 319 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q14669, Q15644 | Gene names | TRIP12, KIAA0045 | |||
|
Domain Architecture |

|
|||||
| Description | Thyroid receptor-interacting protein 12 (TRIP12). | |||||
|
VAV2_MOUSE
|
||||||
| NC score | 0.064141 (rank : 88) | θ value | 0.0431538 (rank : 64) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 456 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q60992 | Gene names | Vav2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein vav-2. | |||||
|
UBE3C_HUMAN
|
||||||
| NC score | 0.061455 (rank : 89) | θ value | θ > 10 (rank : 162) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q15386, Q8TC15, Q96CR4, Q9UDU3 | Gene names | UBE3C, KIAA0010 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase E3C (EC 6.3.2.-). | |||||
|
UBE3C_MOUSE
|
||||||
| NC score | 0.061269 (rank : 90) | θ value | θ > 10 (rank : 163) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80U95, Q8BQZ6, Q8C7W6, Q8CDJ1, Q8VDL5 | Gene names | Ube3c, Kiaa0010 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase E3C (EC 6.3.2.-). | |||||
|
RIN1_MOUSE
|
||||||
| NC score | 0.060753 (rank : 91) | θ value | θ > 10 (rank : 155) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q921Q7 | Gene names | Rin1 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 1 (Ras interaction/interference protein 1). | |||||
|
RIN1_HUMAN
|
||||||
| NC score | 0.059529 (rank : 92) | θ value | θ > 10 (rank : 154) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q13671, O15010, Q00427, Q96CC8 | Gene names | RIN1 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 1 (Ras interaction/interference protein 1) (Ras inhibitor JC99). | |||||
|
PKHG1_HUMAN
|
||||||
| NC score | 0.059327 (rank : 93) | θ value | θ > 10 (rank : 152) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9ULL1, Q5T1F2 | Gene names | PLEKHG1, KIAA1209 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family G member 1. | |||||
|
ARHG4_MOUSE
|
||||||
| NC score | 0.058331 (rank : 94) | θ value | θ > 10 (rank : 123) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q7TNR9, Q80TJ6 | Gene names | Arhgef4, Kiaa1112 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 4 (APC-stimulated guanine nucleotide exchange factor) (Asef). | |||||
|
HUWE1_MOUSE
|
||||||
| NC score | 0.058135 (rank : 95) | θ value | θ > 10 (rank : 142) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q7TMY8, Q4G2Z1, Q5BMM7, Q6NS61, Q8BNJ7, Q8CFH2, Q8VD14, Q921M5, Q9R0P2 | Gene names | Huwe1, Kiaa0312, Ureb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (E3Histone). | |||||
|
ARHG7_HUMAN
|
||||||
| NC score | 0.057945 (rank : 96) | θ value | θ > 10 (rank : 124) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q14155, Q6P9G3, Q6PII2, Q86W63, Q8N3M1 | Gene names | ARHGEF7, COOL1, KIAA0142, P85SPR, PAK3BP, PIXB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 7 (PAK-interacting exchange factor beta) (Beta-Pix) (COOL-1) (p85). | |||||
|
HECD1_HUMAN
|
||||||
| NC score | 0.057122 (rank : 97) | θ value | 0.62314 (rank : 76) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 371 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9ULT8, Q6P445, Q86VJ1, Q96F34, Q9UFZ7 | Gene names | HECTD1, KIAA1131 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin-protein ligase HECTD1 (HECT domain-containing protein 1) (E3 ligase for inhibin receptor) (EULIR). | |||||
|
KR202_HUMAN
|
||||||
| NC score | 0.057065 (rank : 98) | θ value | θ > 10 (rank : 145) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 4 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q3LI61 | Gene names | KRTAP20-2, KAP20.2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 20-2. | |||||
|
HUWE1_HUMAN
|
||||||
| NC score | 0.056883 (rank : 99) | θ value | θ > 10 (rank : 141) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q7Z6Z7, O15029, Q4G2Z2, Q5H961, Q6P4D0, Q8NG67, Q9BUI0, Q9HCJ4, Q9NSL6, Q9P0A9 | Gene names | HUWE1, KIAA0312, KIAA1578, UREB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (Mcl-1 ubiquitin ligase E3) (Mule) (ARF-binding protein 1) (ARF-BP1). | |||||
|
ARHG4_HUMAN
|
||||||
| NC score | 0.056535 (rank : 100) | θ value | θ > 10 (rank : 122) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9NR80, Q9HDC6, Q9UPP0 | Gene names | ARHGEF4, KIAA1112 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 4 (APC-stimulated guanine nucleotide exchange factor) (Asef). | |||||
|
SMUF2_HUMAN
|
||||||
| NC score | 0.055981 (rank : 101) | θ value | θ > 10 (rank : 158) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9HAU4, Q9H260 | Gene names | SMURF2 | |||
|
Domain Architecture |

|
|||||
| Description | Smad ubiquitination regulatory factor 2 (EC 6.3.2.-) (Ubiquitin-- protein ligase SMURF2) (Smad-specific E3 ubiquitin ligase 2) (hSMURF2). | |||||
|
SMUF1_HUMAN
|
||||||
| NC score | 0.055775 (rank : 102) | θ value | θ > 10 (rank : 156) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9HCE7, O75853, Q9UJT8 | Gene names | SMURF1, KIAA1625 | |||
|
Domain Architecture |

|
|||||
| Description | Smad ubiquitination regulatory factor 1 (EC 6.3.2.-) (Ubiquitin-- protein ligase SMURF1) (Smad-specific E3 ubiquitin ligase 1) (hSMURF1). | |||||
|
SMUF1_MOUSE
|
||||||
| NC score | 0.055501 (rank : 103) | θ value | θ > 10 (rank : 157) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9CUN6, Q3U412, Q8K300 | Gene names | Smurf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Smad ubiquitination regulatory factor 1 (EC 6.3.2.-) (Ubiquitin-- protein ligase SMURF1) (Smad-specific E3 ubiquitin ligase 1). | |||||
|
ITCH_MOUSE
|
||||||
| NC score | 0.055163 (rank : 104) | θ value | θ > 10 (rank : 144) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8C863, O54971 | Gene names | Itch | |||
|
Domain Architecture |

|
|||||
| Description | Itchy E3 ubiquitin protein ligase (EC 6.3.2.-). | |||||
|
ECT2_MOUSE
|
||||||
| NC score | 0.055112 (rank : 105) | θ value | θ > 10 (rank : 130) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q07139 | Gene names | Ect2 | |||
|
Domain Architecture |

|
|||||
| Description | ECT2 protein (Epithelial cell-transforming sequence 2 oncogene). | |||||
|
ITCH_HUMAN
|
||||||
| NC score | 0.054736 (rank : 106) | θ value | θ > 10 (rank : 143) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96J02, O43584, Q5TEL0, Q96F66, Q9BY75, Q9H451, Q9H4U5 | Gene names | ITCH | |||
|
Domain Architecture |

|
|||||
| Description | Itchy homolog E3 ubiquitin protein ligase (EC 6.3.2.-) (Itch) (Atrophin-1-interacting protein 4) (AIP4) (NFE2-associated polypeptide 1) (NAPP1). | |||||
|
WWP2_MOUSE
|
||||||
| NC score | 0.054736 (rank : 107) | θ value | θ > 10 (rank : 167) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9DBH0, Q8BTG4, Q923F6 | Gene names | Wwp2 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP2 (EC 6.3.2.-) (WW domain- containing protein 2). | |||||
|
WWP2_HUMAN
|
||||||
| NC score | 0.054646 (rank : 108) | θ value | θ > 10 (rank : 166) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O00308, Q96CZ2, Q9BWN6 | Gene names | WWP2 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP2 (EC 6.3.2.-) (WW domain- containing protein 2) (Atrophin-1-interacting protein 2) (AIP2). | |||||
|
ECT2_HUMAN
|
||||||
| NC score | 0.054302 (rank : 109) | θ value | θ > 10 (rank : 129) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9H8V3, Q9NSV8, Q9NVW9 | Gene names | ECT2 | |||
|
Domain Architecture |

|
|||||
| Description | ECT2 protein (Epithelial cell-transforming sequence 2 oncogene). | |||||
|
PARC_MOUSE
|
||||||
| NC score | 0.053439 (rank : 110) | θ value | 0.279714 (rank : 72) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q80TT8, Q8BKL9, Q8CGC0, Q8R363 | Gene names | Parc, Kiaa0708 | |||
|
Domain Architecture |

|
|||||
| Description | p53-associated parkin-like cytoplasmic protein. | |||||
|
ARHG7_MOUSE
|
||||||
| NC score | 0.053319 (rank : 111) | θ value | θ > 10 (rank : 125) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 396 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9ES28, O08757, Q9ES27 | Gene names | Arhgef7, Pak3bp | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 7 (PAK-interacting exchange factor beta) (Beta-Pix) (p85SPR). | |||||
|
ZFY28_HUMAN
|
||||||
| NC score | 0.052982 (rank : 112) | θ value | θ > 10 (rank : 168) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9HCC9 | Gene names | ZFYVE28, KIAA1643 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger FYVE domain-containing protein 28. | |||||
|
WWP1_MOUSE
|
||||||
| NC score | 0.052745 (rank : 113) | θ value | θ > 10 (rank : 165) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BZZ3, Q8BIV9, Q8VDP8 | Gene names | Wwp1 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP1 (EC 6.3.2.-) (WW domain- containing protein 1). | |||||
|
WWP1_HUMAN
|
||||||
| NC score | 0.052370 (rank : 114) | θ value | θ > 10 (rank : 164) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9H0M0, O00307, Q96BP4 | Gene names | WWP1 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-like E3 ubiquitin-protein ligase WWP1 (EC 6.3.2.-) (WW domain- containing protein 1) (Atropin-1-interacting protein 5) (AIP5). | |||||
|
EDD1_HUMAN
|
||||||
| NC score | 0.051556 (rank : 115) | θ value | θ > 10 (rank : 131) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O95071, O94970, Q9NPL3 | Gene names | EDD1, EDD, HYD, KIAA0896 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase EDD1 (EC 6.3.2.-) (Hyperplastic discs protein homolog) (hHYD) (Progestin-induced protein). | |||||
|
EDD1_MOUSE
|
||||||
| NC score | 0.051321 (rank : 116) | θ value | θ > 10 (rank : 132) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q80TP3, Q698K9, Q6PEQ8, Q6PFQ9, Q80VL4, Q810V6, Q9CXE9 | Gene names | Edd1, Edd, Kiaa0896 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase EDD1 (EC 6.3.2.-) (Hyperplastic discs protein homolog). | |||||
|
NEDD4_MOUSE
|
||||||
| NC score | 0.051152 (rank : 117) | θ value | θ > 10 (rank : 151) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P46935, O08758, Q3UZI2, Q8BGB3 | Gene names | Nedd4, Kiaa0093, Nedd-4, Nedd4a | |||
|
Domain Architecture |

|
|||||
| Description | E3 ubiquitin-protein ligase NEDD4 (EC 6.3.2.-) (Neural precursor cell expressed developmentally down-regulated protein 4). | |||||
|
CUL7_HUMAN
|
||||||
| NC score | 0.050968 (rank : 118) | θ value | θ > 10 (rank : 128) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14999, Q5T654 | Gene names | CUL7, KIAA0076 | |||
|
Domain Architecture |

|
|||||
| Description | Cullin-7 (CUL-7). | |||||
|
NED4L_MOUSE
|
||||||
| NC score | 0.050715 (rank : 119) | θ value | θ > 10 (rank : 149) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8CFI0, Q8BRT9, Q8BS42, Q99PK2 | Gene names | Nedd4l, Kiaa0439, Nedd4b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin-protein ligase NEDD4-like protein (EC 6.3.2.-) (Nedd4-2) (NEDD4.2). | |||||
|
ARHG9_HUMAN
|
||||||
| NC score | 0.050639 (rank : 120) | θ value | θ > 10 (rank : 126) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O43307, Q5JSL6 | Gene names | ARHGEF9, KIAA0424 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 9 (Rac/Cdc42 guanine nucleotide exchange factor 9) (Collybistin) (PEM-2 homolog). | |||||
|
NED4L_HUMAN
|
||||||
| NC score | 0.050560 (rank : 121) | θ value | θ > 10 (rank : 148) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96PU5, O43165, Q7Z5F1, Q7Z5F2, Q7Z5N3, Q8N5A7, Q8WUU9, Q9BW58, Q9H2W4, Q9NT88 | Gene names | NEDD4L, KIAA0439, NEDL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin-protein ligase NEDD4-like protein (EC 6.3.2.-) (Nedd4-2) (NEDD4.2). | |||||
|
ARHG9_MOUSE
|
||||||
| NC score | 0.050272 (rank : 122) | θ value | θ > 10 (rank : 127) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q3UTH8, Q3TQ60, Q80U06, Q8CAF9 | Gene names | Arhgef9, Kiaa0424 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 9 (Rac/Cdc42 guanine nucleotide exchange factor 9) (Collybistin). | |||||
|
NEDD4_HUMAN
|
||||||
| NC score | 0.050030 (rank : 123) | θ value | θ > 10 (rank : 150) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P46934 | Gene names | NEDD4, KIAA0093 | |||
|
Domain Architecture |

|
|||||
| Description | E3 ubiquitin-protein ligase NEDD4 (EC 6.3.2.-). | |||||
|
OGT1_HUMAN
|
||||||
| NC score | 0.044619 (rank : 124) | θ value | 0.163984 (rank : 68) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O15294, Q7Z3K0, Q8WWM8, Q96CC1, Q9UG57 | Gene names | OGT | |||
|
Domain Architecture |

|
|||||
| Description | UDP-N-acetylglucosamine--peptide N-acetylglucosaminyltransferase 110 kDa subunit (EC 2.4.1.-) (O-GlcNAc transferase p110 subunit). | |||||
|
E41L3_HUMAN
|
||||||
| NC score | 0.039458 (rank : 125) | θ value | 0.21417 (rank : 69) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y2J2, O95713, Q9BRP5 | Gene names | EPB41L3, DAL1, KIAA0987 | |||
|
Domain Architecture |

|
|||||
| Description | Band 4.1-like protein 3 (4.1B) (Differentially expressed in adenocarcinoma of the lung protein 1) (DAL-1). | |||||
|
PANK4_HUMAN
|
||||||
| NC score | 0.037713 (rank : 126) | θ value | 0.365318 (rank : 73) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NVE7, Q53EU3, Q5TA84, Q7RTX3, Q9H3X5 | Gene names | PANK4 | |||
|
Domain Architecture |

|
|||||
| Description | Pantothenate kinase 4 (EC 2.7.1.33) (Pantothenic acid kinase 4) (hPanK4). | |||||
|
FIP1_MOUSE
|
||||||
| NC score | 0.034889 (rank : 127) | θ value | 0.47712 (rank : 75) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9D824, Q8BWX7, Q99LH0, Q9DBB2 | Gene names | Fip1l1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pre-mRNA 3'-end-processing factor FIP1 (FIP1-like 1). | |||||
|
PLEK_HUMAN
|
||||||
| NC score | 0.033237 (rank : 128) | θ value | 4.03905 (rank : 100) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P08567, Q53SU8, Q6FGM8, Q8WV81 | Gene names | PLEK, P47 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin (Platelet p47 protein). | |||||
|
SNX1_HUMAN
|
||||||
| NC score | 0.030186 (rank : 129) | θ value | 0.62314 (rank : 79) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q13596, O60750, O60751 | Gene names | SNX1 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-1. | |||||
|
E41L2_HUMAN
|
||||||
| NC score | 0.029830 (rank : 130) | θ value | 6.88961 (rank : 109) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O43491 | Gene names | EPB41L2 | |||
|
Domain Architecture |

|
|||||
| Description | Band 4.1-like protein 2 (Generally expressed protein 4.1) (4.1G). | |||||
|
PKHA1_HUMAN
|
||||||
| NC score | 0.028894 (rank : 131) | θ value | 2.36792 (rank : 92) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9HB21, Q9BVK0 | Gene names | PLEKHA1, TAPP1 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 1 (Tandem PH domain-containing protein 1) (TAPP-1). | |||||
|
TR11B_MOUSE
|
||||||
| NC score | 0.027261 (rank : 132) | θ value | 1.81305 (rank : 87) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O08712, O70202 | Gene names | Tnfrsf11b, Ocif, Opg | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 11B precursor (Osteoprotegerin) (Osteoclastogenesis inhibitory factor). | |||||
|
POPD2_HUMAN
|
||||||
| NC score | 0.027210 (rank : 133) | θ value | 1.38821 (rank : 83) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9HBU9, Q86UE7 | Gene names | POPDC2, POP2 | |||
|
Domain Architecture |

|
|||||
| Description | Popeye domain-containing protein 2 (Popeye protein 2). | |||||
|
CHMP3_MOUSE
|
||||||
| NC score | 0.026929 (rank : 134) | θ value | 2.36792 (rank : 89) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9CQ10, Q9D2Z2, Q9D7A5 | Gene names | Vps24, Chmp3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Charged multivesicular body protein 3 (Chromatin-modifying protein 3) (Vacuolar protein sorting-associated protein 24). | |||||
|
CENA1_HUMAN
|
||||||
| NC score | 0.025662 (rank : 135) | θ value | 3.0926 (rank : 94) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O75689 | Gene names | CENTA1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centaurin-alpha 1 (Putative MAPK-activating protein PM25). | |||||
|
BAG4_MOUSE
|
||||||
| NC score | 0.025302 (rank : 136) | θ value | 1.38821 (rank : 81) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CI61, Q3TRL9, Q91VT5, Q9CWG2 | Gene names | Bag4, Sodd | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 4 (BCL2-associated athanogene 4) (BAG-4) (Silencer of death domains). | |||||
|
PANK4_MOUSE
|
||||||
| NC score | 0.024925 (rank : 137) | θ value | 4.03905 (rank : 99) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80YV4, Q7M751, Q8BQE9 | Gene names | Pank4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pantothenate kinase 4 (EC 2.7.1.33) (Pantothenic acid kinase 4) (mPanK4). | |||||
|
LPIN2_HUMAN
|
||||||
| NC score | 0.022886 (rank : 138) | θ value | 2.36792 (rank : 90) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q92539 | Gene names | LPIN2, KIAA0249 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipin-2. | |||||
|
LPIN2_MOUSE
|
||||||
| NC score | 0.022136 (rank : 139) | θ value | 2.36792 (rank : 91) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99PI5, Q8C357, Q8C7I8, Q8CC85, Q8CHR7 | Gene names | Lpin2 | |||
|
Domain Architecture |

|
|||||
| Description | Lipin-2. | |||||
|
CHMP3_HUMAN
|
||||||
| NC score | 0.021313 (rank : 140) | θ value | 6.88961 (rank : 108) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y3E7, Q53S71, Q53SU5, Q9NZ51 | Gene names | VPS24, CHMP3, NEDF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Charged multivesicular body protein 3 (Chromatin-modifying protein 3) (Vacuolar protein sorting-associated protein 24) (hVps24) (Neuroendocrine differentiation factor). | |||||
|
CRSP2_HUMAN
|
||||||
| NC score | 0.020427 (rank : 141) | θ value | 3.0926 (rank : 95) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O60244, Q9UNB3 | Gene names | CRSP2, ARC150, DRIP150, EXLM1, TRAP170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CRSP complex subunit 2 (Cofactor required for Sp1 transcriptional activation subunit 2) (Transcriptional coactivator CRSP150) (Vitamin D3 receptor-interacting protein complex 150 kDa component) (DRIP150) (Thyroid hormone receptor-associated protein complex 170 kDa component) (Trap170) (Activator-recruited cofactor 150 kDa component) (ARC150). | |||||
|
BTBDG_HUMAN
|
||||||
| NC score | 0.020413 (rank : 142) | θ value | 4.03905 (rank : 98) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q32M84, Q4VXL1, Q96LN0 | Gene names | BTBD16, C10orf87 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein 16. | |||||
|
PRG4_HUMAN
|
||||||
| NC score | 0.019913 (rank : 143) | θ value | 0.62314 (rank : 78) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
CEND1_HUMAN
|
||||||
| NC score | 0.018598 (rank : 144) | θ value | 5.27518 (rank : 103) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 284 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8WZ64, Q4W5D2, Q7Z2L5, Q96L70, Q96P49, Q9Y4E4 | Gene names | CENTD1, ARAP2, KIAA0580 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centaurin-delta 1 (Cnt-d1) (Arf-GAP, Rho-GAP, ankyrin repeat and pleckstrin homology domain-containing protein 2) (PARX protein). | |||||
|
CENA2_HUMAN
|
||||||
| NC score | 0.017947 (rank : 145) | θ value | 6.88961 (rank : 107) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NPF8, Q8N4Q6, Q96SD5 | Gene names | CENTA2 | |||
|
Domain Architecture |

|
|||||
| Description | Centaurin-alpha 2. | |||||
|
CEND1_MOUSE
|
||||||
| NC score | 0.016430 (rank : 146) | θ value | 8.99809 (rank : 116) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8BZ05, Q80TX2, Q8C3T2, Q8VEL6 | Gene names | Centd1, Kiaa0580 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centaurin-delta 1 (Cnt-d1). | |||||
|
TBCD4_MOUSE
|
||||||
| NC score | 0.016429 (rank : 147) | θ value | 0.813845 (rank : 80) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BYJ6, Q5DU23, Q66JU2, Q6P2M2, Q8BMH6, Q8BXM2 | Gene names | Tbc1d4, Kiaa0603 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 4 (Akt substrate of 160 kDa) (AS160). | |||||
|
PRG4_MOUSE
|
||||||
| NC score | 0.014696 (rank : 148) | θ value | 3.0926 (rank : 97) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9JM99, Q3UEL1, Q3V198 | Gene names | Prg4, Msf, Szp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
ATR_HUMAN
|
||||||
| NC score | 0.013993 (rank : 149) | θ value | 5.27518 (rank : 102) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13535, Q7KYL3, Q93051, Q9BXK4 | Gene names | ATR, FRP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase ATR (EC 2.7.11.1) (Ataxia telangiectasia and Rad3-related protein) (FRAP-related protein 1). | |||||
|
RBBP6_MOUSE
|
||||||
| NC score | 0.013738 (rank : 150) | θ value | 2.36792 (rank : 93) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
PKCB1_HUMAN
|
||||||
| NC score | 0.013634 (rank : 151) | θ value | 5.27518 (rank : 106) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 488 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9ULU4, Q13517, Q4JJ94, Q4JJ95, Q5TH09, Q6MZM1, Q8WXC5, Q9H1F3, Q9H1F4, Q9H1F5, Q9H1L8, Q9H1L9, Q9H2G5, Q9NYN3, Q9UIX6 | Gene names | PRKCBP1, KIAA1125, RACK7, ZMYND8 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C-binding protein 1 (Rack7) (Cutaneous T-cell lymphoma- associated antigen se14-3) (CTCL tumor antigen se14-3) (Zinc finger MYND domain-containing protein 8). | |||||
|
IRS2_HUMAN
|
||||||
| NC score | 0.012486 (rank : 152) | θ value | 5.27518 (rank : 104) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y4H2, Q96RR2, Q9BZG0, Q9Y6I5 | Gene names | IRS2 | |||
|
Domain Architecture |

|
|||||
| Description | Insulin receptor substrate 2 (IRS-2). | |||||
|
MARK2_MOUSE
|
||||||
| NC score | 0.011721 (rank : 153) | θ value | 3.0926 (rank : 96) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 1018 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q05512, Q6PDR4, Q8BR95 | Gene names | Mark2, Emk | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase MARK2 (EC 2.7.11.1) (MAP/microtubule affinity-regulating kinase 2) (ELKL Motif Kinase) (EMK1). | |||||
|
VP13A_HUMAN
|
||||||
| NC score | 0.011096 (rank : 154) | θ value | 8.99809 (rank : 121) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96RL7, Q5JSX9, Q5JSY0, Q5VYR5, Q702P4, Q709D0, Q86YF8, Q96S61, Q9H995, Q9Y2J1 | Gene names | VPS13A, CHAC, KIAA0986 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 13A (Chorein) (Chorea- acanthocytosis protein). | |||||
|
JIP4_HUMAN
|
||||||
| NC score | 0.010904 (rank : 155) | θ value | 6.88961 (rank : 111) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 585 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O60271, O60905, Q3KQU8, Q3MKM7, Q86WC7, Q86WC8, Q8IZX7, Q96II0, Q9H811 | Gene names | SPAG9, HSS, KIAA0516, MAPK8IP4, SYD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 4 (JNK-interacting protein 4) (JIP-4) (JNK-associated leucine-zipper protein) (JLP) (Sperm-associated antigen 9) (Mitogen-activated protein kinase 8- interacting protein 4) (Human lung cancer protein 6) (HLC-6) (Proliferation-inducing protein 6) (Sperm-specific protein) (Sperm surface protein) (Protein highly expressed in testis) (PHET) (Sunday driver 1). | |||||
|
MARK2_HUMAN
|
||||||
| NC score | 0.010263 (rank : 156) | θ value | 5.27518 (rank : 105) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 1056 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q7KZI7, Q15449, Q15524, Q5XGA3, Q68A18, Q96HB3, Q96RG0 | Gene names | MARK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MARK2 (EC 2.7.11.1) (MAP/microtubule affinity-regulating kinase 2) (ELKL motif kinase) (EMK1) (PAR1 homolog). | |||||
|
MYO10_HUMAN
|
||||||
| NC score | 0.009953 (rank : 157) | θ value | 0.62314 (rank : 77) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 920 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9HD67, Q9NYM7, Q9P110, Q9P111, Q9UHF6 | Gene names | MYO10 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-10 (Myosin X). | |||||
|
PCLO_HUMAN
|
||||||
| NC score | 0.009925 (rank : 158) | θ value | 8.99809 (rank : 119) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
PK3CD_MOUSE
|
||||||
| NC score | 0.009709 (rank : 159) | θ value | 6.88961 (rank : 112) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O35904 | Gene names | Pik3cd | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit delta isoform (EC 2.7.1.153) (PI3-kinase p110 subunit delta) (PtdIns-3- kinase p110) (PI3K) (p110delta). | |||||
|
EMIL2_HUMAN
|
||||||
| NC score | 0.007894 (rank : 160) | θ value | 8.99809 (rank : 117) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 573 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9BXX0, Q8NBH3, Q96JQ4 | Gene names | EMILIN2 | |||
|
Domain Architecture |

|
|||||
| Description | EMILIN-2 precursor (Elastin microfibril interface-located protein 2) (Elastin microfibril interfacer 2) (Protein FOAP-10). | |||||
|
RFIP5_MOUSE
|
||||||
| NC score | 0.007074 (rank : 161) | θ value | 8.99809 (rank : 120) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8R361 | Gene names | Rab11fip5, D6Ertd32e, Rip11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab11 family-interacting protein 5 (Rab11-FIP5) (Rab11-interacting protein Rip11). | |||||
|
SL9A7_HUMAN
|
||||||
| NC score | 0.006343 (rank : 162) | θ value | 6.88961 (rank : 114) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96T83, O75827, Q5JXP9 | Gene names | SLC9A7, NHE7 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/hydrogen exchanger 7 (Na(+)/H(+) exchanger 7) (NHE-7) (Solute carrier family 9 member 7). | |||||
|
MYCN_MOUSE
|
||||||
| NC score | 0.006039 (rank : 163) | θ value | 8.99809 (rank : 118) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P03966, Q61978 | Gene names | Mycn, Nmyc, Nmyc1 | |||
|
Domain Architecture |

|
|||||
| Description | N-myc proto-oncogene protein. | |||||
|
ZN292_HUMAN
|
||||||
| NC score | 0.005619 (rank : 164) | θ value | 4.03905 (rank : 101) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 918 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O60281, Q9H8G3, Q9H8J4 | Gene names | ZNF292, KIAA0530 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 292. | |||||
|
CADH2_HUMAN
|
||||||
| NC score | 0.002212 (rank : 165) | θ value | 2.36792 (rank : 88) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P19022, Q14923 | Gene names | CDH2, CDHN, NCAD | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin-2 precursor (Neural-cadherin) (N-cadherin) (CDw325 antigen). | |||||
|
RBP24_HUMAN
|
||||||
| NC score | -0.000478 (rank : 166) | θ value | 6.88961 (rank : 113) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q4ZG46 | Gene names | RANBP2L4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 2-like 4 (RanBP2L4). | |||||
|
GCC2_HUMAN
|
||||||
| NC score | -0.000493 (rank : 167) | θ value | 6.88961 (rank : 110) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 1558 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8IWJ2, O15045, Q8TDH3, Q9H2G8 | Gene names | GCC2, KIAA0336 | |||
|
Domain Architecture |

|
|||||
| Description | GRIP and coiled-coil domain-containing protein 2 (Golgi coiled coil protein GCC185) (CTCL tumor antigen se1-1) (CLL-associated antigen KW- 11) (NY-REN-53 antigen). | |||||
|
ZN236_HUMAN
|
||||||
| NC score | -0.004647 (rank : 168) | θ value | 6.88961 (rank : 115) | |||
| Query Neighborhood Hits | 121 | Target Neighborhood Hits | 844 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UL36, Q9UL37 | Gene names | ZNF236 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 236. | |||||