Please be patient as the page loads
|
JPH3_MOUSE
|
||||||
| SwissProt Accessions | Q9ET77, Q9EQZ2 | Gene names | Jph3, Jp3 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-3 (Junctophilin type 3) (JP-3). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
JPH3_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.996278 (rank : 2) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q8WXH2, Q8N471, Q9HDC3, Q9HDC4 | Gene names | JPH3, JP3 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-3 (Junctophilin type 3) (JP-3). | |||||
|
JPH3_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | Q9ET77, Q9EQZ2 | Gene names | Jph3, Jp3 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-3 (Junctophilin type 3) (JP-3). | |||||
|
JPH1_MOUSE
|
||||||
| θ value | 4.56552e-153 (rank : 3) | NC score | 0.967876 (rank : 4) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9ET80, Q9EQZ3 | Gene names | Jph1, Jp1 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-1 (Junctophilin type 1) (JP-1). | |||||
|
JPH1_HUMAN
|
||||||
| θ value | 1.51984e-148 (rank : 4) | NC score | 0.972786 (rank : 3) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9HDC5 | Gene names | JPH1, JP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junctophilin-1 (Junctophilin type 1) (JP-1). | |||||
|
JPH2_MOUSE
|
||||||
| θ value | 3.5201e-129 (rank : 5) | NC score | 0.923380 (rank : 7) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9ET78, Q9ET79 | Gene names | Jph2, Jp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junctophilin-2 (Junctophilin type 2) (JP-2). | |||||
|
JPH2_HUMAN
|
||||||
| θ value | 4.30305e-127 (rank : 6) | NC score | 0.867476 (rank : 8) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 639 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9BR39, O95913, Q5JY74, Q9UJN4 | Gene names | JPH2, JP2 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-2 (Junctophilin type 2) (JP-2). | |||||
|
JPH4_MOUSE
|
||||||
| θ value | 1.06727e-101 (rank : 7) | NC score | 0.952992 (rank : 6) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q80WT0, Q8BMI1 | Gene names | Jph4, Jphl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junctophilin-4 (Junctophilin-like 1 protein). | |||||
|
JPH4_HUMAN
|
||||||
| θ value | 9.98939e-100 (rank : 8) | NC score | 0.953199 (rank : 5) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q96JJ6, Q8ND44, Q96DQ0 | Gene names | JPH4, JPHL1, KIAA1831 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junctophilin-4 (Junctophilin-like 1 protein). | |||||
|
MORN3_MOUSE
|
||||||
| θ value | 3.76295e-14 (rank : 9) | NC score | 0.687373 (rank : 9) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8C5T4, Q8VE21, Q9D5H6 | Gene names | Morn3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORN repeat-containing protein 3. | |||||
|
MORN3_HUMAN
|
||||||
| θ value | 6.41864e-14 (rank : 10) | NC score | 0.680040 (rank : 10) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6PF18, Q86YQ9 | Gene names | MORN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORN repeat-containing protein 3. | |||||
|
TSGA2_HUMAN
|
||||||
| θ value | 1.42992e-13 (rank : 11) | NC score | 0.669307 (rank : 11) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8WYR4 | Gene names | TSGA2, TSA2 | |||
|
Domain Architecture |

|
|||||
| Description | Testis-specific gene A2 protein (Male meiotic metaphase chromosome- associated acidic protein) (Meichroacidin). | |||||
|
TSGA2_MOUSE
|
||||||
| θ value | 4.16044e-13 (rank : 12) | NC score | 0.661623 (rank : 12) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8VIG3, Q9DAL5 | Gene names | Tsga2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-specific gene A2 protein (Male meiotic metaphase chromosome- associated acidic protein) (Meichroacidin). | |||||
|
MORN1_HUMAN
|
||||||
| θ value | 4.59992e-12 (rank : 13) | NC score | 0.636807 (rank : 13) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q5T089, Q8WW30, Q9H852 | Gene names | MORN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORN repeat-containing protein 1. | |||||
|
ALS2_HUMAN
|
||||||
| θ value | 3.89403e-11 (rank : 14) | NC score | 0.422158 (rank : 17) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q96Q42, Q8N1E0, Q96PC4, Q96Q41, Q9H973, Q9HCK9 | Gene names | ALS2, KIAA1563 | |||
|
Domain Architecture |

|
|||||
| Description | Alsin (Amyotrophic lateral sclerosis protein 2). | |||||
|
ALS2_MOUSE
|
||||||
| θ value | 3.89403e-11 (rank : 15) | NC score | 0.422280 (rank : 16) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q920R0, Q8JZR1, Q9CXJ3 | Gene names | Als2 | |||
|
Domain Architecture |

|
|||||
| Description | Alsin (Amyotrophic lateral sclerosis protein 2 homolog). | |||||
|
MORN2_HUMAN
|
||||||
| θ value | 0.00102713 (rank : 16) | NC score | 0.520168 (rank : 14) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q502X0, Q6UL00 | Gene names | MORN2, MOPT | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORN repeat-containing protein 2 (MORN motif protein in testis). | |||||
|
MORN2_MOUSE
|
||||||
| θ value | 0.00175202 (rank : 17) | NC score | 0.502608 (rank : 15) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6UL01 | Gene names | Morn2, Mopt | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORN repeat-containing protein 2 (MORN motif protein in testis). | |||||
|
DHX9_MOUSE
|
||||||
| θ value | 0.00509761 (rank : 18) | NC score | 0.074510 (rank : 32) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O70133, O35931, O54703, Q5FWY1, Q6R5F7, Q9CSA2 | Gene names | Dhx9, Ddx9 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent RNA helicase A (EC 3.6.1.-) (Nuclear DNA helicase II) (NDH II) (DEAH box protein 9) (mHEL-5). | |||||
|
ANKY1_HUMAN
|
||||||
| θ value | 0.00665767 (rank : 19) | NC score | 0.226944 (rank : 22) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9P2S6, Q8IYX5, Q8NDK5, Q9NX10 | Gene names | ANKMY1, TSAL1, ZMYND13 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin repeat and MYND domain-containing protein 1 (Testis-specific ankyrin-like protein 1) (Zinc-finger MYND domain-containing protein 13). | |||||
|
AFF3_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 20) | NC score | 0.047002 (rank : 36) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P51826 | Gene names | AFF3, LAF4 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 3 (Protein LAF-4) (Lymphoid nuclear protein related to AF4). | |||||
|
LORI_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 21) | NC score | 0.267874 (rank : 19) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P18165, Q8R0I9 | Gene names | Lor | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Loricrin. | |||||
|
ROA1_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 22) | NC score | 0.075614 (rank : 31) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P09651, Q6PJZ7 | Gene names | HNRPA1 | |||
|
Domain Architecture |
|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein A1 (Helix-destabilizing protein) (Single-strand RNA-binding protein) (hnRNP core protein A1). | |||||
|
K1C10_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 23) | NC score | 0.030278 (rank : 46) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 526 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P13645, Q14664, Q8N175 | Gene names | KRT10 | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type I cytoskeletal 10 (Cytokeratin-10) (CK-10) (Keratin-10) (K10). | |||||
|
ARSD_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 24) | NC score | 0.043648 (rank : 37) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P51689, Q9UHJ8 | Gene names | ARSD | |||
|
Domain Architecture |
|
|||||
| Description | Arylsulfatase D precursor (EC 3.1.6.-) (ASD). | |||||
|
SETD7_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 25) | NC score | 0.247114 (rank : 20) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8WTS6, Q9C0E6 | Gene names | SETD7, KIAA1717, SET7 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET7 (EC 2.1.1.43) (Histone H3-K4 methyltransferase) (H3-K4-HMTase) (SET domain-containing protein 7) (Set9) (SET7/9). | |||||
|
SETD7_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 26) | NC score | 0.245565 (rank : 21) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8VHL1 | Gene names | Setd7, Set7 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET7 (EC 2.1.1.43) (Histone H3-K4 methyltransferase) (H3-K4-HMTase) (SET domain-containing protein 7). | |||||
|
FA98A_HUMAN
|
||||||
| θ value | 0.125558 (rank : 27) | NC score | 0.140748 (rank : 25) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8NCA5 | Gene names | FAM98A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM98A. | |||||
|
FA98A_MOUSE
|
||||||
| θ value | 0.125558 (rank : 28) | NC score | 0.133289 (rank : 26) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q3TJZ6, Q99KJ2 | Gene names | Fam98a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM98A. | |||||
|
DHX9_HUMAN
|
||||||
| θ value | 0.163984 (rank : 29) | NC score | 0.053353 (rank : 34) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q08211, Q5VY62, Q6PD69, Q99556 | Gene names | DHX9, DDX9, NDH2 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent RNA helicase A (EC 3.6.1.-) (Nuclear DNA helicase II) (NDH II) (DEAH box protein 9). | |||||
|
ROA3_MOUSE
|
||||||
| θ value | 0.163984 (rank : 30) | NC score | 0.037855 (rank : 38) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BG05, Q8BHF8 | Gene names | Hnrpa3, Hnrnpa3 | |||
|
Domain Architecture |
|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein A3 (hnRNP A3). | |||||
|
TMC2_MOUSE
|
||||||
| θ value | 0.21417 (rank : 31) | NC score | 0.033938 (rank : 44) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8R4P4 | Gene names | Tmc2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane cochlear-expressed protein 2. | |||||
|
K1C10_MOUSE
|
||||||
| θ value | 0.279714 (rank : 32) | NC score | 0.023713 (rank : 52) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 466 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P02535, P08731, Q8BUX3, Q8BV09, Q8BVU3, Q9CXH6, Q9JKB4 | Gene names | Krt10, Krt1-10 | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type I cytoskeletal 10 (Cytokeratin-10) (CK-10) (Keratin-10) (K10) (56 kDa cytokeratin) (Keratin, type I cytoskeletal 59 kDa). | |||||
|
MRC1_MOUSE
|
||||||
| θ value | 0.279714 (rank : 33) | NC score | 0.023346 (rank : 54) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q61830, Q8C502 | Gene names | Mrc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Macrophage mannose receptor 1 precursor (MMR) (CD206 antigen). | |||||
|
HORN_MOUSE
|
||||||
| θ value | 0.365318 (rank : 34) | NC score | 0.087968 (rank : 28) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8VHD8 | Gene names | Hrnr | |||
|
Domain Architecture |

|
|||||
| Description | Hornerin. | |||||
|
MYH6_HUMAN
|
||||||
| θ value | 0.47712 (rank : 35) | NC score | 0.010655 (rank : 70) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1426 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P13533, Q13943, Q14906, Q14907 | Gene names | MYH6, MYHCA | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
SNUT1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 36) | NC score | 0.034929 (rank : 41) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O43290, Q53GB5 | Gene names | SART1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U4/U6.U5 tri-snRNP-associated protein 1 (U4/U6.U5 tri-snRNP-associated 110 kDa protein) (Squamous cell carcinoma antigen recognized by T cells 1) (SART-1) (hSART-1) (hSnu66). | |||||
|
K1C9_HUMAN
|
||||||
| θ value | 0.62314 (rank : 37) | NC score | 0.031843 (rank : 45) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 526 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P35527, O00109, Q14665 | Gene names | KRT9 | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type I cytoskeletal 9 (Cytokeratin-9) (CK-9) (Keratin-9) (K9). | |||||
|
PCLO_HUMAN
|
||||||
| θ value | 0.813845 (rank : 38) | NC score | 0.024370 (rank : 51) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
HORN_HUMAN
|
||||||
| θ value | 1.06291 (rank : 39) | NC score | 0.101153 (rank : 27) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q86YZ3 | Gene names | HRNR, S100A18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hornerin. | |||||
|
CAC1A_HUMAN
|
||||||
| θ value | 1.38821 (rank : 40) | NC score | 0.007702 (rank : 78) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O00555, P78510, P78511, Q16290, Q92690, Q99790, Q99791, Q99792, Q99793 | Gene names | CACNA1A, CACH4, CACN3, CACNL1A4 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent P/Q-type calcium channel subunit alpha-1A (Voltage- gated calcium channel subunit alpha Cav2.1) (Calcium channel, L type, alpha-1 polypeptide isoform 4) (Brain calcium channel I) (BI). | |||||
|
CUTL1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 41) | NC score | 0.014978 (rank : 58) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P53564, O08994, P70301, Q571L6, Q91ZD2 | Gene names | Cutl1, Cux, Cux1, Kiaa4047 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP) (Homeobox protein Cux). | |||||
|
LORI_HUMAN
|
||||||
| θ value | 1.38821 (rank : 42) | NC score | 0.280605 (rank : 18) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P23490 | Gene names | LOR, LRN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Loricrin. | |||||
|
ROA2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 43) | NC score | 0.035885 (rank : 40) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O88569 | Gene names | Hnrpa2b1 | |||
|
Domain Architecture |
|
|||||
| Description | Heterogeneous nuclear ribonucleoproteins A2/B1 (hnRNP A2 / hnRNP B1). | |||||
|
ARSE_HUMAN
|
||||||
| θ value | 1.81305 (rank : 44) | NC score | 0.034893 (rank : 42) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P51690, Q53FT2, Q53FU8 | Gene names | ARSE | |||
|
Domain Architecture |
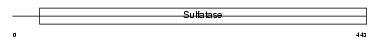
|
|||||
| Description | Arylsulfatase E precursor (EC 3.1.6.-) (ASE). | |||||
|
CUTL1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 45) | NC score | 0.012732 (rank : 64) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1574 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P39880, Q6NYH4, Q9UEV5 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP). | |||||
|
RMP_HUMAN
|
||||||
| θ value | 1.81305 (rank : 46) | NC score | 0.050171 (rank : 35) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O94763 | Gene names | RMP, C19orf2 | |||
|
Domain Architecture |
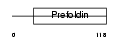
|
|||||
| Description | RNA polymerase II subunit 5-mediating protein (RPB5-mediating protein). | |||||
|
CT012_MOUSE
|
||||||
| θ value | 2.36792 (rank : 47) | NC score | 0.013793 (rank : 62) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 229 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8C008, Q8C060 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C20orf12 homolog. | |||||
|
FA98B_MOUSE
|
||||||
| θ value | 2.36792 (rank : 48) | NC score | 0.080867 (rank : 29) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q80VD1, Q9CS28, Q9CS37 | Gene names | Fam98b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM98B. | |||||
|
FREM1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 49) | NC score | 0.014199 (rank : 61) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5H8C1, Q5VV00, Q5VV01, Q6MZI4, Q8NEG9, Q96LI3 | Gene names | FREM1, C9orf154 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FRAS1-related extracellular matrix protein 1 precursor (Protein QBRICK). | |||||
|
SRMS_HUMAN
|
||||||
| θ value | 2.36792 (rank : 50) | NC score | -0.001336 (rank : 83) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H3Y6 | Gene names | SRMS, C20orf148 | |||
|
Domain Architecture |
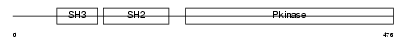
|
|||||
| Description | Tyrosine-protein kinase Srms (EC 2.7.10.2). | |||||
|
LSM4_MOUSE
|
||||||
| θ value | 3.0926 (rank : 51) | NC score | 0.021097 (rank : 55) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9QXA5 | Gene names | Lsm4 | |||
|
Domain Architecture |
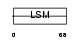
|
|||||
| Description | U6 snRNA-associated Sm-like protein LSm4. | |||||
|
MYH6_MOUSE
|
||||||
| θ value | 3.0926 (rank : 52) | NC score | 0.008299 (rank : 75) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1501 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q02566, Q64258, Q64738 | Gene names | Myh6, Myhca | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
PDE9A_HUMAN
|
||||||
| θ value | 3.0926 (rank : 53) | NC score | 0.007104 (rank : 81) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O76083, O75490, O75491, O95225, Q86SF7, Q86SI6, Q86SJ3, Q86WN3, Q86WN4, Q86WN5, Q86WN6, Q86WN7, Q86WN8, Q86WN9, Q86WP0 | Gene names | PDE9A | |||
|
Domain Architecture |

|
|||||
| Description | High-affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (EC 3.1.4.35). | |||||
|
RBP56_HUMAN
|
||||||
| θ value | 3.0926 (rank : 54) | NC score | 0.077277 (rank : 30) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q92804, Q92751 | Gene names | TAF15, RBP56, TAF2N | |||
|
Domain Architecture |

|
|||||
| Description | TATA-binding protein-associated factor 2N (RNA-binding protein 56) (TAFII68) (TAF(II)68). | |||||
|
TACC2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 55) | NC score | 0.015272 (rank : 57) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 856 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O95359, Q9NZ41, Q9NZR5 | Gene names | TACC2 | |||
|
Domain Architecture |

|
|||||
| Description | Transforming acidic coiled-coil-containing protein 2 (Anti Zuai-1) (AZU-1). | |||||
|
ARSF_HUMAN
|
||||||
| θ value | 4.03905 (rank : 56) | NC score | 0.027704 (rank : 47) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P54793, Q8TCC5 | Gene names | ARSF | |||
|
Domain Architecture |
|
|||||
| Description | Arylsulfatase F precursor (EC 3.1.6.-) (ASF). | |||||
|
HEMGN_HUMAN
|
||||||
| θ value | 4.03905 (rank : 57) | NC score | 0.023533 (rank : 53) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9BXL5, Q6XAR3, Q86XY5, Q9NPC0 | Gene names | HEMGN, EDAG, NDR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hemogen (Hemopoietic gene protein) (Negative differentiation regulator protein) (Erythroid differentiation-associated gene protein) (EDAG-1). | |||||
|
ILF3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 58) | NC score | 0.021087 (rank : 56) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Z1X4, Q80VD5, Q812A1, Q8BP80, Q8C2H8, Q8K588 | Gene names | Ilf3 | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin enhancer-binding factor 3. | |||||
|
K1718_MOUSE
|
||||||
| θ value | 4.03905 (rank : 59) | NC score | 0.007556 (rank : 79) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q3UWM4, Q3UWN8, Q6ZPJ5, Q8C969, Q8C9E0, Q91VX8 | Gene names | Kiaa1718 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1718. | |||||
|
MYH7_HUMAN
|
||||||
| θ value | 4.03905 (rank : 60) | NC score | 0.007749 (rank : 77) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P12883, Q14836, Q14837, Q14904, Q16579, Q92679, Q9H1D5, Q9UMM8 | Gene names | MYH7, MYHCB | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle beta isoform (MyHC-beta). | |||||
|
ROA3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 61) | NC score | 0.027478 (rank : 48) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P51991, Q6URK5 | Gene names | HNRPA3 | |||
|
Domain Architecture |
|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein A3 (hnRNP A3). | |||||
|
SNUT1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 62) | NC score | 0.027394 (rank : 49) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Z315, Q8K155, Q9R1I9, Q9R270 | Gene names | Sart1, Haf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U4/U6.U5 tri-snRNP-associated protein 1 (Squamous cell carcinoma antigen recognized by T-cells 1) (SART-1) (mSART-1) (Hypoxia- associated factor). | |||||
|
TCF7_MOUSE
|
||||||
| θ value | 4.03905 (rank : 63) | NC score | 0.011092 (rank : 68) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q00417 | Gene names | Tcf7, Tcf-1, Tcf1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 7 (T-cell-specific transcription factor 1) (TCF- 1) (T-cell factor 1). | |||||
|
ESCO2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 64) | NC score | 0.010866 (rank : 69) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CIB9, Q3UMT6, Q6IQX5, Q8BNG9, Q8BQF8, Q9CRI8 | Gene names | Esco2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyltransferase ESCO2 (EC 2.3.1.-) (Establishment of cohesion 1 homolog 2) (ECO1 homolog 2). | |||||
|
FILA_HUMAN
|
||||||
| θ value | 5.27518 (rank : 65) | NC score | 0.034206 (rank : 43) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 685 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P20930, Q01720, Q5T583 | Gene names | FLG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filaggrin. | |||||
|
PACN3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 66) | NC score | 0.009236 (rank : 73) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q99JB8, Q9EQP9 | Gene names | Pacsin3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C and casein kinase II substrate protein 3. | |||||
|
RBM19_HUMAN
|
||||||
| θ value | 5.27518 (rank : 67) | NC score | 0.011973 (rank : 66) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y4C8, Q9BPY6, Q9UFN5 | Gene names | RBM19, KIAA0682 | |||
|
Domain Architecture |

|
|||||
| Description | Probable RNA-binding protein 19 (RNA-binding motif protein 19). | |||||
|
ROA2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 68) | NC score | 0.025888 (rank : 50) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P22626, P22627 | Gene names | HNRPA2B1 | |||
|
Domain Architecture |
|
|||||
| Description | Heterogeneous nuclear ribonucleoproteins A2/B1 (hnRNP A2 / hnRNP B1). | |||||
|
SPEG_MOUSE
|
||||||
| θ value | 5.27518 (rank : 69) | NC score | 0.008263 (rank : 76) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
IMPG1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 70) | NC score | 0.010427 (rank : 71) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q17R60, O43686, O95094, Q9BWZ1 | Gene names | IMPG1, IPM150, SPACR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interphotoreceptor matrix proteoglycan 1 precursor (Interphotoreceptor matrix proteoglycan of 150 kDa) (IPM-150) (Sialoprotein associated with cones and rods). | |||||
|
MFA3L_MOUSE
|
||||||
| θ value | 6.88961 (rank : 71) | NC score | 0.011457 (rank : 67) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D3X9, Q80TV6, Q8BJA9 | Gene names | Mfap3l, Kiaa0626 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microfibrillar-associated protein 3-like precursor. | |||||
|
MRC1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 72) | NC score | 0.014787 (rank : 59) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P22897 | Gene names | MRC1 | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage mannose receptor 1 precursor (MMR) (CD206 antigen). | |||||
|
PCLO_MOUSE
|
||||||
| θ value | 6.88961 (rank : 73) | NC score | 0.013777 (rank : 63) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
XPC_MOUSE
|
||||||
| θ value | 6.88961 (rank : 74) | NC score | 0.010363 (rank : 72) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P51612, P54732, Q3TKI2, Q920M1, Q9DBW7 | Gene names | Xpc | |||
|
Domain Architecture |

|
|||||
| Description | DNA-repair protein complementing XP-C cells homolog (Xeroderma pigmentosum group C-complementing protein homolog) (p125). | |||||
|
BAT2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 75) | NC score | 0.037355 (rank : 39) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
CD2L5_MOUSE
|
||||||
| θ value | 8.99809 (rank : 76) | NC score | -0.000663 (rank : 82) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1011 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q69ZA1, Q80V11, Q8BZG1, Q8K0A4 | Gene names | Cdc2l5, Kiaa1791 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle 2-like protein kinase 5 (EC 2.7.11.22) (CDC2- related protein kinase 5). | |||||
|
CS034_HUMAN
|
||||||
| θ value | 8.99809 (rank : 77) | NC score | 0.014562 (rank : 60) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NCQ2 | Gene names | C19orf34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C19orf34. | |||||
|
RRBP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 78) | NC score | 0.012627 (rank : 65) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
UB2J1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 79) | NC score | 0.009021 (rank : 74) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9JJZ4, Q9DC92 | Gene names | Ube2j1, Ncube1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-conjugating enzyme E2 J1 (EC 6.3.2.19) (Non-canonical ubiquitin-conjugating enzyme 1) (NCUBE1). | |||||
|
ZN541_HUMAN
|
||||||
| θ value | 8.99809 (rank : 80) | NC score | 0.007488 (rank : 80) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H0D2, Q8NDK8 | Gene names | ZNF541 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 541. | |||||
|
KR193_HUMAN
|
||||||
| θ value | θ > 10 (rank : 81) | NC score | 0.150311 (rank : 24) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 5 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7Z4W3 | Gene names | KRTAP19-3, KAP19.3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 19-3 (Glycine/tyrosine-rich protein) (GTHRP). | |||||
|
KR197_HUMAN
|
||||||
| θ value | θ > 10 (rank : 82) | NC score | 0.064384 (rank : 33) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q3SYF9 | Gene names | KRTAP19-7, KAP19.7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 19-7. | |||||
|
KR202_HUMAN
|
||||||
| θ value | θ > 10 (rank : 83) | NC score | 0.184769 (rank : 23) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 4 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q3LI61 | Gene names | KRTAP20-2, KAP20.2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 20-2. | |||||
|
JPH3_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | Q9ET77, Q9EQZ2 | Gene names | Jph3, Jp3 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-3 (Junctophilin type 3) (JP-3). | |||||
|
JPH3_HUMAN
|
||||||
| NC score | 0.996278 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q8WXH2, Q8N471, Q9HDC3, Q9HDC4 | Gene names | JPH3, JP3 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-3 (Junctophilin type 3) (JP-3). | |||||
|
JPH1_HUMAN
|
||||||
| NC score | 0.972786 (rank : 3) | θ value | 1.51984e-148 (rank : 4) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9HDC5 | Gene names | JPH1, JP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junctophilin-1 (Junctophilin type 1) (JP-1). | |||||
|
JPH1_MOUSE
|
||||||
| NC score | 0.967876 (rank : 4) | θ value | 4.56552e-153 (rank : 3) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9ET80, Q9EQZ3 | Gene names | Jph1, Jp1 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-1 (Junctophilin type 1) (JP-1). | |||||
|
JPH4_HUMAN
|
||||||
| NC score | 0.953199 (rank : 5) | θ value | 9.98939e-100 (rank : 8) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q96JJ6, Q8ND44, Q96DQ0 | Gene names | JPH4, JPHL1, KIAA1831 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junctophilin-4 (Junctophilin-like 1 protein). | |||||
|
JPH4_MOUSE
|
||||||
| NC score | 0.952992 (rank : 6) | θ value | 1.06727e-101 (rank : 7) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q80WT0, Q8BMI1 | Gene names | Jph4, Jphl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junctophilin-4 (Junctophilin-like 1 protein). | |||||
|
JPH2_MOUSE
|
||||||
| NC score | 0.923380 (rank : 7) | θ value | 3.5201e-129 (rank : 5) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9ET78, Q9ET79 | Gene names | Jph2, Jp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junctophilin-2 (Junctophilin type 2) (JP-2). | |||||
|
JPH2_HUMAN
|
||||||
| NC score | 0.867476 (rank : 8) | θ value | 4.30305e-127 (rank : 6) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 639 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9BR39, O95913, Q5JY74, Q9UJN4 | Gene names | JPH2, JP2 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-2 (Junctophilin type 2) (JP-2). | |||||
|
MORN3_MOUSE
|
||||||
| NC score | 0.687373 (rank : 9) | θ value | 3.76295e-14 (rank : 9) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8C5T4, Q8VE21, Q9D5H6 | Gene names | Morn3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORN repeat-containing protein 3. | |||||
|
MORN3_HUMAN
|
||||||
| NC score | 0.680040 (rank : 10) | θ value | 6.41864e-14 (rank : 10) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6PF18, Q86YQ9 | Gene names | MORN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORN repeat-containing protein 3. | |||||
|
TSGA2_HUMAN
|
||||||
| NC score | 0.669307 (rank : 11) | θ value | 1.42992e-13 (rank : 11) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8WYR4 | Gene names | TSGA2, TSA2 | |||
|
Domain Architecture |

|
|||||
| Description | Testis-specific gene A2 protein (Male meiotic metaphase chromosome- associated acidic protein) (Meichroacidin). | |||||
|
TSGA2_MOUSE
|
||||||
| NC score | 0.661623 (rank : 12) | θ value | 4.16044e-13 (rank : 12) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8VIG3, Q9DAL5 | Gene names | Tsga2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-specific gene A2 protein (Male meiotic metaphase chromosome- associated acidic protein) (Meichroacidin). | |||||
|
MORN1_HUMAN
|
||||||
| NC score | 0.636807 (rank : 13) | θ value | 4.59992e-12 (rank : 13) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q5T089, Q8WW30, Q9H852 | Gene names | MORN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORN repeat-containing protein 1. | |||||
|
MORN2_HUMAN
|
||||||
| NC score | 0.520168 (rank : 14) | θ value | 0.00102713 (rank : 16) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q502X0, Q6UL00 | Gene names | MORN2, MOPT | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORN repeat-containing protein 2 (MORN motif protein in testis). | |||||
|
MORN2_MOUSE
|
||||||
| NC score | 0.502608 (rank : 15) | θ value | 0.00175202 (rank : 17) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6UL01 | Gene names | Morn2, Mopt | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORN repeat-containing protein 2 (MORN motif protein in testis). | |||||
|
ALS2_MOUSE
|
||||||
| NC score | 0.422280 (rank : 16) | θ value | 3.89403e-11 (rank : 15) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q920R0, Q8JZR1, Q9CXJ3 | Gene names | Als2 | |||
|
Domain Architecture |

|
|||||
| Description | Alsin (Amyotrophic lateral sclerosis protein 2 homolog). | |||||
|
ALS2_HUMAN
|
||||||
| NC score | 0.422158 (rank : 17) | θ value | 3.89403e-11 (rank : 14) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q96Q42, Q8N1E0, Q96PC4, Q96Q41, Q9H973, Q9HCK9 | Gene names | ALS2, KIAA1563 | |||
|
Domain Architecture |

|
|||||
| Description | Alsin (Amyotrophic lateral sclerosis protein 2). | |||||
|
LORI_HUMAN
|
||||||
| NC score | 0.280605 (rank : 18) | θ value | 1.38821 (rank : 42) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P23490 | Gene names | LOR, LRN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Loricrin. | |||||
|
LORI_MOUSE
|
||||||
| NC score | 0.267874 (rank : 19) | θ value | 0.0252991 (rank : 21) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P18165, Q8R0I9 | Gene names | Lor | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Loricrin. | |||||
|
SETD7_HUMAN
|
||||||
| NC score | 0.247114 (rank : 20) | θ value | 0.0961366 (rank : 25) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8WTS6, Q9C0E6 | Gene names | SETD7, KIAA1717, SET7 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET7 (EC 2.1.1.43) (Histone H3-K4 methyltransferase) (H3-K4-HMTase) (SET domain-containing protein 7) (Set9) (SET7/9). | |||||
|
SETD7_MOUSE
|
||||||
| NC score | 0.245565 (rank : 21) | θ value | 0.0961366 (rank : 26) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8VHL1 | Gene names | Setd7, Set7 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET7 (EC 2.1.1.43) (Histone H3-K4 methyltransferase) (H3-K4-HMTase) (SET domain-containing protein 7). | |||||
|
ANKY1_HUMAN
|
||||||
| NC score | 0.226944 (rank : 22) | θ value | 0.00665767 (rank : 19) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9P2S6, Q8IYX5, Q8NDK5, Q9NX10 | Gene names | ANKMY1, TSAL1, ZMYND13 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin repeat and MYND domain-containing protein 1 (Testis-specific ankyrin-like protein 1) (Zinc-finger MYND domain-containing protein 13). | |||||
|
KR202_HUMAN
|
||||||
| NC score | 0.184769 (rank : 23) | θ value | θ > 10 (rank : 83) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 4 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q3LI61 | Gene names | KRTAP20-2, KAP20.2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 20-2. | |||||
|
KR193_HUMAN
|
||||||
| NC score | 0.150311 (rank : 24) | θ value | θ > 10 (rank : 81) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 5 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7Z4W3 | Gene names | KRTAP19-3, KAP19.3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 19-3 (Glycine/tyrosine-rich protein) (GTHRP). | |||||
|
FA98A_HUMAN
|
||||||
| NC score | 0.140748 (rank : 25) | θ value | 0.125558 (rank : 27) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8NCA5 | Gene names | FAM98A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM98A. | |||||
|
FA98A_MOUSE
|
||||||
| NC score | 0.133289 (rank : 26) | θ value | 0.125558 (rank : 28) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q3TJZ6, Q99KJ2 | Gene names | Fam98a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM98A. | |||||
|
HORN_HUMAN
|
||||||
| NC score | 0.101153 (rank : 27) | θ value | 1.06291 (rank : 39) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q86YZ3 | Gene names | HRNR, S100A18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hornerin. | |||||
|
HORN_MOUSE
|
||||||
| NC score | 0.087968 (rank : 28) | θ value | 0.365318 (rank : 34) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8VHD8 | Gene names | Hrnr | |||
|
Domain Architecture |

|
|||||
| Description | Hornerin. | |||||
|
FA98B_MOUSE
|
||||||
| NC score | 0.080867 (rank : 29) | θ value | 2.36792 (rank : 48) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q80VD1, Q9CS28, Q9CS37 | Gene names | Fam98b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM98B. | |||||
|
RBP56_HUMAN
|
||||||
| NC score | 0.077277 (rank : 30) | θ value | 3.0926 (rank : 54) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q92804, Q92751 | Gene names | TAF15, RBP56, TAF2N | |||
|
Domain Architecture |

|
|||||
| Description | TATA-binding protein-associated factor 2N (RNA-binding protein 56) (TAFII68) (TAF(II)68). | |||||
|
ROA1_HUMAN
|
||||||
| NC score | 0.075614 (rank : 31) | θ value | 0.0431538 (rank : 22) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P09651, Q6PJZ7 | Gene names | HNRPA1 | |||
|
Domain Architecture |

|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein A1 (Helix-destabilizing protein) (Single-strand RNA-binding protein) (hnRNP core protein A1). | |||||
|
DHX9_MOUSE
|
||||||
| NC score | 0.074510 (rank : 32) | θ value | 0.00509761 (rank : 18) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O70133, O35931, O54703, Q5FWY1, Q6R5F7, Q9CSA2 | Gene names | Dhx9, Ddx9 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent RNA helicase A (EC 3.6.1.-) (Nuclear DNA helicase II) (NDH II) (DEAH box protein 9) (mHEL-5). | |||||
|
KR197_HUMAN
|
||||||
| NC score | 0.064384 (rank : 33) | θ value | θ > 10 (rank : 82) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q3SYF9 | Gene names | KRTAP19-7, KAP19.7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 19-7. | |||||
|
DHX9_HUMAN
|
||||||
| NC score | 0.053353 (rank : 34) | θ value | 0.163984 (rank : 29) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q08211, Q5VY62, Q6PD69, Q99556 | Gene names | DHX9, DDX9, NDH2 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent RNA helicase A (EC 3.6.1.-) (Nuclear DNA helicase II) (NDH II) (DEAH box protein 9). | |||||
|
RMP_HUMAN
|
||||||
| NC score | 0.050171 (rank : 35) | θ value | 1.81305 (rank : 46) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O94763 | Gene names | RMP, C19orf2 | |||
|
Domain Architecture |

|
|||||
| Description | RNA polymerase II subunit 5-mediating protein (RPB5-mediating protein). | |||||
|
AFF3_HUMAN
|
||||||
| NC score | 0.047002 (rank : 36) | θ value | 0.0148317 (rank : 20) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P51826 | Gene names | AFF3, LAF4 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 3 (Protein LAF-4) (Lymphoid nuclear protein related to AF4). | |||||
|
ARSD_HUMAN
|
||||||
| NC score | 0.043648 (rank : 37) | θ value | 0.0961366 (rank : 24) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P51689, Q9UHJ8 | Gene names | ARSD | |||
|
Domain Architecture |

|
|||||
| Description | Arylsulfatase D precursor (EC 3.1.6.-) (ASD). | |||||
|
ROA3_MOUSE
|
||||||
| NC score | 0.037855 (rank : 38) | θ value | 0.163984 (rank : 30) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BG05, Q8BHF8 | Gene names | Hnrpa3, Hnrnpa3 | |||
|
Domain Architecture |

|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein A3 (hnRNP A3). | |||||
|
BAT2_HUMAN
|
||||||
| NC score | 0.037355 (rank : 39) | θ value | 8.99809 (rank : 75) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
ROA2_MOUSE
|
||||||
| NC score | 0.035885 (rank : 40) | θ value | 1.38821 (rank : 43) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O88569 | Gene names | Hnrpa2b1 | |||
|
Domain Architecture |

|
|||||
| Description | Heterogeneous nuclear ribonucleoproteins A2/B1 (hnRNP A2 / hnRNP B1). | |||||
|
SNUT1_HUMAN
|
||||||
| NC score | 0.034929 (rank : 41) | θ value | 0.47712 (rank : 36) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O43290, Q53GB5 | Gene names | SART1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U4/U6.U5 tri-snRNP-associated protein 1 (U4/U6.U5 tri-snRNP-associated 110 kDa protein) (Squamous cell carcinoma antigen recognized by T cells 1) (SART-1) (hSART-1) (hSnu66). | |||||
|
ARSE_HUMAN
|
||||||
| NC score | 0.034893 (rank : 42) | θ value | 1.81305 (rank : 44) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P51690, Q53FT2, Q53FU8 | Gene names | ARSE | |||
|
Domain Architecture |

|
|||||
| Description | Arylsulfatase E precursor (EC 3.1.6.-) (ASE). | |||||
|
FILA_HUMAN
|
||||||
| NC score | 0.034206 (rank : 43) | θ value | 5.27518 (rank : 65) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 685 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P20930, Q01720, Q5T583 | Gene names | FLG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filaggrin. | |||||
|
TMC2_MOUSE
|
||||||
| NC score | 0.033938 (rank : 44) | θ value | 0.21417 (rank : 31) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8R4P4 | Gene names | Tmc2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane cochlear-expressed protein 2. | |||||
|
K1C9_HUMAN
|
||||||
| NC score | 0.031843 (rank : 45) | θ value | 0.62314 (rank : 37) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 526 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P35527, O00109, Q14665 | Gene names | KRT9 | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type I cytoskeletal 9 (Cytokeratin-9) (CK-9) (Keratin-9) (K9). | |||||
|
K1C10_HUMAN
|
||||||
| NC score | 0.030278 (rank : 46) | θ value | 0.0736092 (rank : 23) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 526 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P13645, Q14664, Q8N175 | Gene names | KRT10 | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type I cytoskeletal 10 (Cytokeratin-10) (CK-10) (Keratin-10) (K10). | |||||
|
ARSF_HUMAN
|
||||||
| NC score | 0.027704 (rank : 47) | θ value | 4.03905 (rank : 56) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P54793, Q8TCC5 | Gene names | ARSF | |||
|
Domain Architecture |

|
|||||
| Description | Arylsulfatase F precursor (EC 3.1.6.-) (ASF). | |||||
|
ROA3_HUMAN
|
||||||
| NC score | 0.027478 (rank : 48) | θ value | 4.03905 (rank : 61) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P51991, Q6URK5 | Gene names | HNRPA3 | |||
|
Domain Architecture |

|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein A3 (hnRNP A3). | |||||
|
SNUT1_MOUSE
|
||||||
| NC score | 0.027394 (rank : 49) | θ value | 4.03905 (rank : 62) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Z315, Q8K155, Q9R1I9, Q9R270 | Gene names | Sart1, Haf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U4/U6.U5 tri-snRNP-associated protein 1 (Squamous cell carcinoma antigen recognized by T-cells 1) (SART-1) (mSART-1) (Hypoxia- associated factor). | |||||
|
ROA2_HUMAN
|
||||||
| NC score | 0.025888 (rank : 50) | θ value | 5.27518 (rank : 68) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P22626, P22627 | Gene names | HNRPA2B1 | |||
|
Domain Architecture |

|
|||||
| Description | Heterogeneous nuclear ribonucleoproteins A2/B1 (hnRNP A2 / hnRNP B1). | |||||
|
PCLO_HUMAN
|
||||||
| NC score | 0.024370 (rank : 51) | θ value | 0.813845 (rank : 38) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
K1C10_MOUSE
|
||||||
| NC score | 0.023713 (rank : 52) | θ value | 0.279714 (rank : 32) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 466 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P02535, P08731, Q8BUX3, Q8BV09, Q8BVU3, Q9CXH6, Q9JKB4 | Gene names | Krt10, Krt1-10 | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type I cytoskeletal 10 (Cytokeratin-10) (CK-10) (Keratin-10) (K10) (56 kDa cytokeratin) (Keratin, type I cytoskeletal 59 kDa). | |||||
|
HEMGN_HUMAN
|
||||||
| NC score | 0.023533 (rank : 53) | θ value | 4.03905 (rank : 57) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9BXL5, Q6XAR3, Q86XY5, Q9NPC0 | Gene names | HEMGN, EDAG, NDR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hemogen (Hemopoietic gene protein) (Negative differentiation regulator protein) (Erythroid differentiation-associated gene protein) (EDAG-1). | |||||
|
MRC1_MOUSE
|
||||||
| NC score | 0.023346 (rank : 54) | θ value | 0.279714 (rank : 33) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q61830, Q8C502 | Gene names | Mrc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Macrophage mannose receptor 1 precursor (MMR) (CD206 antigen). | |||||
|
LSM4_MOUSE
|
||||||
| NC score | 0.021097 (rank : 55) | θ value | 3.0926 (rank : 51) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9QXA5 | Gene names | Lsm4 | |||
|
Domain Architecture |

|
|||||
| Description | U6 snRNA-associated Sm-like protein LSm4. | |||||
|
ILF3_MOUSE
|
||||||
| NC score | 0.021087 (rank : 56) | θ value | 4.03905 (rank : 58) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Z1X4, Q80VD5, Q812A1, Q8BP80, Q8C2H8, Q8K588 | Gene names | Ilf3 | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin enhancer-binding factor 3. | |||||
|
TACC2_HUMAN
|
||||||
| NC score | 0.015272 (rank : 57) | θ value | 3.0926 (rank : 55) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 856 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O95359, Q9NZ41, Q9NZR5 | Gene names | TACC2 | |||
|
Domain Architecture |

|
|||||
| Description | Transforming acidic coiled-coil-containing protein 2 (Anti Zuai-1) (AZU-1). | |||||
|
CUTL1_MOUSE
|
||||||
| NC score | 0.014978 (rank : 58) | θ value | 1.38821 (rank : 41) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P53564, O08994, P70301, Q571L6, Q91ZD2 | Gene names | Cutl1, Cux, Cux1, Kiaa4047 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP) (Homeobox protein Cux). | |||||
|
MRC1_HUMAN
|
||||||
| NC score | 0.014787 (rank : 59) | θ value | 6.88961 (rank : 72) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P22897 | Gene names | MRC1 | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage mannose receptor 1 precursor (MMR) (CD206 antigen). | |||||
|
CS034_HUMAN
|
||||||
| NC score | 0.014562 (rank : 60) | θ value | 8.99809 (rank : 77) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NCQ2 | Gene names | C19orf34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C19orf34. | |||||
|
FREM1_HUMAN
|
||||||
| NC score | 0.014199 (rank : 61) | θ value | 2.36792 (rank : 49) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5H8C1, Q5VV00, Q5VV01, Q6MZI4, Q8NEG9, Q96LI3 | Gene names | FREM1, C9orf154 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FRAS1-related extracellular matrix protein 1 precursor (Protein QBRICK). | |||||
|
CT012_MOUSE
|
||||||
| NC score | 0.013793 (rank : 62) | θ value | 2.36792 (rank : 47) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 229 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8C008, Q8C060 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C20orf12 homolog. | |||||
|
PCLO_MOUSE
|
||||||
| NC score | 0.013777 (rank : 63) | θ value | 6.88961 (rank : 73) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
CUTL1_HUMAN
|
||||||
| NC score | 0.012732 (rank : 64) | θ value | 1.81305 (rank : 45) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1574 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P39880, Q6NYH4, Q9UEV5 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP). | |||||
|
RRBP1_MOUSE
|
||||||
| NC score | 0.012627 (rank : 65) | θ value | 8.99809 (rank : 78) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
RBM19_HUMAN
|
||||||
| NC score | 0.011973 (rank : 66) | θ value | 5.27518 (rank : 67) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y4C8, Q9BPY6, Q9UFN5 | Gene names | RBM19, KIAA0682 | |||
|
Domain Architecture |

|
|||||
| Description | Probable RNA-binding protein 19 (RNA-binding motif protein 19). | |||||
|
MFA3L_MOUSE
|
||||||
| NC score | 0.011457 (rank : 67) | θ value | 6.88961 (rank : 71) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D3X9, Q80TV6, Q8BJA9 | Gene names | Mfap3l, Kiaa0626 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microfibrillar-associated protein 3-like precursor. | |||||
|
TCF7_MOUSE
|
||||||
| NC score | 0.011092 (rank : 68) | θ value | 4.03905 (rank : 63) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q00417 | Gene names | Tcf7, Tcf-1, Tcf1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 7 (T-cell-specific transcription factor 1) (TCF- 1) (T-cell factor 1). | |||||
|
ESCO2_MOUSE
|
||||||
| NC score | 0.010866 (rank : 69) | θ value | 5.27518 (rank : 64) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CIB9, Q3UMT6, Q6IQX5, Q8BNG9, Q8BQF8, Q9CRI8 | Gene names | Esco2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyltransferase ESCO2 (EC 2.3.1.-) (Establishment of cohesion 1 homolog 2) (ECO1 homolog 2). | |||||
|
MYH6_HUMAN
|
||||||
| NC score | 0.010655 (rank : 70) | θ value | 0.47712 (rank : 35) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1426 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P13533, Q13943, Q14906, Q14907 | Gene names | MYH6, MYHCA | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
IMPG1_HUMAN
|
||||||
| NC score | 0.010427 (rank : 71) | θ value | 6.88961 (rank : 70) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q17R60, O43686, O95094, Q9BWZ1 | Gene names | IMPG1, IPM150, SPACR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interphotoreceptor matrix proteoglycan 1 precursor (Interphotoreceptor matrix proteoglycan of 150 kDa) (IPM-150) (Sialoprotein associated with cones and rods). | |||||
|
XPC_MOUSE
|
||||||
| NC score | 0.010363 (rank : 72) | θ value | 6.88961 (rank : 74) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P51612, P54732, Q3TKI2, Q920M1, Q9DBW7 | Gene names | Xpc | |||
|
Domain Architecture |

|
|||||
| Description | DNA-repair protein complementing XP-C cells homolog (Xeroderma pigmentosum group C-complementing protein homolog) (p125). | |||||
|
PACN3_MOUSE
|
||||||
| NC score | 0.009236 (rank : 73) | θ value | 5.27518 (rank : 66) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q99JB8, Q9EQP9 | Gene names | Pacsin3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C and casein kinase II substrate protein 3. | |||||
|
UB2J1_MOUSE
|
||||||
| NC score | 0.009021 (rank : 74) | θ value | 8.99809 (rank : 79) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9JJZ4, Q9DC92 | Gene names | Ube2j1, Ncube1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-conjugating enzyme E2 J1 (EC 6.3.2.19) (Non-canonical ubiquitin-conjugating enzyme 1) (NCUBE1). | |||||
|
MYH6_MOUSE
|
||||||
| NC score | 0.008299 (rank : 75) | θ value | 3.0926 (rank : 52) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1501 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q02566, Q64258, Q64738 | Gene names | Myh6, Myhca | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
SPEG_MOUSE
|
||||||
| NC score | 0.008263 (rank : 76) | θ value | 5.27518 (rank : 69) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
MYH7_HUMAN
|
||||||
| NC score | 0.007749 (rank : 77) | θ value | 4.03905 (rank : 60) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P12883, Q14836, Q14837, Q14904, Q16579, Q92679, Q9H1D5, Q9UMM8 | Gene names | MYH7, MYHCB | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle beta isoform (MyHC-beta). | |||||
|
CAC1A_HUMAN
|
||||||
| NC score | 0.007702 (rank : 78) | θ value | 1.38821 (rank : 40) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O00555, P78510, P78511, Q16290, Q92690, Q99790, Q99791, Q99792, Q99793 | Gene names | CACNA1A, CACH4, CACN3, CACNL1A4 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent P/Q-type calcium channel subunit alpha-1A (Voltage- gated calcium channel subunit alpha Cav2.1) (Calcium channel, L type, alpha-1 polypeptide isoform 4) (Brain calcium channel I) (BI). | |||||
|
K1718_MOUSE
|
||||||
| NC score | 0.007556 (rank : 79) | θ value | 4.03905 (rank : 59) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q3UWM4, Q3UWN8, Q6ZPJ5, Q8C969, Q8C9E0, Q91VX8 | Gene names | Kiaa1718 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1718. | |||||
|
ZN541_HUMAN
|
||||||
| NC score | 0.007488 (rank : 80) | θ value | 8.99809 (rank : 80) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H0D2, Q8NDK8 | Gene names | ZNF541 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 541. | |||||
|
PDE9A_HUMAN
|
||||||
| NC score | 0.007104 (rank : 81) | θ value | 3.0926 (rank : 53) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O76083, O75490, O75491, O95225, Q86SF7, Q86SI6, Q86SJ3, Q86WN3, Q86WN4, Q86WN5, Q86WN6, Q86WN7, Q86WN8, Q86WN9, Q86WP0 | Gene names | PDE9A | |||
|
Domain Architecture |

|
|||||
| Description | High-affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A (EC 3.1.4.35). | |||||
|
CD2L5_MOUSE
|
||||||
| NC score | -0.000663 (rank : 82) | θ value | 8.99809 (rank : 76) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1011 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q69ZA1, Q80V11, Q8BZG1, Q8K0A4 | Gene names | Cdc2l5, Kiaa1791 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle 2-like protein kinase 5 (EC 2.7.11.22) (CDC2- related protein kinase 5). | |||||
|
SRMS_HUMAN
|
||||||
| NC score | -0.001336 (rank : 83) | θ value | 2.36792 (rank : 50) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H3Y6 | Gene names | SRMS, C20orf148 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein kinase Srms (EC 2.7.10.2). | |||||