Please be patient as the page loads
|
K1718_MOUSE
|
||||||
| SwissProt Accessions | Q3UWM4, Q3UWN8, Q6ZPJ5, Q8C969, Q8C9E0, Q91VX8 | Gene names | Kiaa1718 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1718. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
K1718_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.991569 (rank : 2) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q6ZMT4, Q6MZL8, Q9C0E5 | Gene names | KIAA1718 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1718. | |||||
|
K1718_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 98 | |
| SwissProt Accessions | Q3UWM4, Q3UWN8, Q6ZPJ5, Q8C969, Q8C9E0, Q91VX8 | Gene names | Kiaa1718 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1718. | |||||
|
PHF8_MOUSE
|
||||||
| θ value | 2.48782e-175 (rank : 3) | NC score | 0.971984 (rank : 3) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q80TJ7, Q8BLX8, Q8BLY0, Q8BZ61, Q8CG26 | Gene names | Phf8, Kiaa1111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 8. | |||||
|
PHF8_HUMAN
|
||||||
| θ value | 4.6918e-174 (rank : 4) | NC score | 0.969453 (rank : 4) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9UPP1, Q5H9U5, Q5VUJ4, Q7Z6D4, Q9HAH2 | Gene names | PHF8, KIAA1111, ZNF422 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 8. | |||||
|
PHF2_HUMAN
|
||||||
| θ value | 2.04462e-161 (rank : 5) | NC score | 0.950555 (rank : 5) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | O75151, Q8N3K2, Q9Y6N4 | Gene names | PHF2, KIAA0662 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
PHF2_MOUSE
|
||||||
| θ value | 1.01473e-160 (rank : 6) | NC score | 0.948714 (rank : 6) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9WTU0, Q80WA8 | Gene names | Phf2 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
JHD1B_HUMAN
|
||||||
| θ value | 2.47765e-74 (rank : 7) | NC score | 0.766997 (rank : 9) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8NHM5, Q8NCI2, Q96HC7, Q96SL0, Q96T03, Q9NS96, Q9UF75 | Gene names | FBXL10, CXXC2, FBL10, JHDM1B, PCCX2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1B (EC 1.14.11.-) (F-box/LRR-repeat protein 10) (F-box and leucine-rich repeat protein 10) (F-box protein FBL10) (Protein JEMMA) (Jumonji domain-containing EMSY-interactor methyltransferase motif protein) (CXXC-type zinc finger protein 2) (Protein-containing CXXC domain 2). | |||||
|
JHD1B_MOUSE
|
||||||
| θ value | 4.22625e-74 (rank : 8) | NC score | 0.766520 (rank : 10) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6P1G2, Q3V396, Q6PFD0, Q6ZPE8, Q9CSF7, Q9QZN6 | Gene names | Fbxl10, Fbl10, Jhdm1b, Kiaa3014 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1B (EC 1.14.11.-) (F-box/LRR-repeat protein 10) (F-box and leucine-rich repeat protein 10) (F-box protein FBL10). | |||||
|
JHD1A_HUMAN
|
||||||
| θ value | 1.11475e-66 (rank : 9) | NC score | 0.775470 (rank : 8) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9Y2K7, Q49A21, Q4G0M3, Q69YY8, Q9BVH5, Q9H7H5, Q9UK66 | Gene names | FBXL11, FBL7, JHDM1A, KIAA1004 | |||
|
Domain Architecture |

|
|||||
| Description | JmjC domain-containing histone demethylation protein 1A (EC 1.14.11.-) (F-box/LRR-repeat protein 11) (F-box and leucine-rich repeat protein 11) (F-box protein FBL7) (F-box protein Lilina). | |||||
|
JHD1A_MOUSE
|
||||||
| θ value | 3.24342e-66 (rank : 10) | NC score | 0.776460 (rank : 7) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P59997, Q3U1M5, Q3UR56, Q3V3Q1, Q69ZT4 | Gene names | Fbxl11, Jhdm1a, Kiaa1004 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1A (EC 1.14.11.-) (F-box/LRR-repeat protein 11) (F-box and leucine-rich repeat protein 11). | |||||
|
DIDO1_HUMAN
|
||||||
| θ value | 1.33837e-11 (rank : 11) | NC score | 0.297724 (rank : 13) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
BPTF_HUMAN
|
||||||
| θ value | 5.08577e-11 (rank : 12) | NC score | 0.270670 (rank : 19) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 786 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q12830, Q6NX67, Q7Z7D6, Q9UIG2 | Gene names | FALZ, BPTF, FAC1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleosome remodeling factor subunit BPTF (Bromodomain and PHD finger- containing transcription factor) (Fetal Alzheimer antigen) (Fetal Alz- 50 clone 1 protein). | |||||
|
CXCC1_HUMAN
|
||||||
| θ value | 1.47974e-10 (rank : 13) | NC score | 0.499871 (rank : 12) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9P0U4, Q8N2W4, Q96BC8, Q9P2V7 | Gene names | CXXC1, CGBP, PCCX1, PHF18 | |||
|
Domain Architecture |
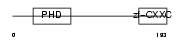
|
|||||
| Description | CpG-binding protein (PHD finger and CXXC domain-containing protein 1) (CXXC-type zinc finger protein 1). | |||||
|
CXCC1_MOUSE
|
||||||
| θ value | 3.29651e-10 (rank : 14) | NC score | 0.500364 (rank : 11) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9CWW7 | Gene names | Cxxc1, Cgbp, Pccx1 | |||
|
Domain Architecture |

|
|||||
| Description | CpG-binding protein (PHD finger and CXXC domain-containing protein 1) (CXXC-type zinc finger protein 1). | |||||
|
CN130_MOUSE
|
||||||
| θ value | 0.000158464 (rank : 15) | NC score | 0.297516 (rank : 14) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BU04, Q80UT9 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C14orf130 homolog. | |||||
|
CN130_HUMAN
|
||||||
| θ value | 0.000270298 (rank : 16) | NC score | 0.295518 (rank : 15) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8N806, Q86U21, Q86UA9, Q96BY0, Q9NVV6 | Gene names | C14orf130 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C14orf130. | |||||
|
JAD1A_HUMAN
|
||||||
| θ value | 0.000270298 (rank : 17) | NC score | 0.170570 (rank : 29) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P29375 | Gene names | JARID1A, RBBP2, RBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1A (Retinoblastoma-binding protein 2) (RBBP-2). | |||||
|
PHF3_HUMAN
|
||||||
| θ value | 0.000270298 (rank : 18) | NC score | 0.240412 (rank : 22) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q92576, Q5T1T6, Q9NQ16, Q9UI45 | Gene names | PHF3, KIAA0244 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 3. | |||||
|
PTDSR_MOUSE
|
||||||
| θ value | 0.00102713 (rank : 19) | NC score | 0.276814 (rank : 18) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9ERI5, Q80TX1 | Gene names | Ptdsr, Kiaa0585 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein PTDSR (Phosphatidylserine receptor) (Apoptotic cell clearance receptor PtdSerR). | |||||
|
PHF20_MOUSE
|
||||||
| θ value | 0.00298849 (rank : 20) | NC score | 0.184268 (rank : 28) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8BLG0, Q8BMA2, Q8BYR4, Q921N1 | Gene names | Phf20, Hca58 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58 homolog). | |||||
|
TAF3_HUMAN
|
||||||
| θ value | 0.00298849 (rank : 21) | NC score | 0.195474 (rank : 26) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 568 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q5VWG9, Q6GMS5, Q6P6B5, Q86VY6, Q9BQS9, Q9UFI8 | Gene names | TAF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
PHF20_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 22) | NC score | 0.191159 (rank : 27) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9BVI0, Q5JWY9, Q9BWV4, Q9BXA3, Q9BZW3, Q9H421, Q9H4J6, Q9NZ22 | Gene names | PHF20, C20orf104, GLEA2, HCA58, NZF, TZP | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58) (Glioma-expressed antigen 2) (Transcription factor TZP) (Novel zinc finger protein). | |||||
|
PTDSR_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 23) | NC score | 0.269445 (rank : 20) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6NYC1, Q86VY0, Q8IUM5, Q9Y4E2 | Gene names | PTDSR, KIAA0585 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein PTDSR (Phosphatidylserine receptor). | |||||
|
JAD1D_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 24) | NC score | 0.144833 (rank : 35) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9BY66, Q92509, Q92809, Q9HCU1 | Gene names | SMCY, HY, HYA, JARID1D, KIAA0234 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1D (SmcY protein) (Histocompatibility Y antigen) (H-Y). | |||||
|
TAF3_MOUSE
|
||||||
| θ value | 0.00665767 (rank : 25) | NC score | 0.201189 (rank : 25) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 463 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q5HZG4, Q3U490, Q3UWX2, Q8BIU8, Q99JH4 | Gene names | Taf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
JAD1C_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 26) | NC score | 0.133610 (rank : 39) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P41229 | Gene names | SMCX, DXS1272E, JARID1C, XE169 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1C (Protein SmcX) (Protein Xe169). | |||||
|
JAD1C_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 27) | NC score | 0.134399 (rank : 36) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P41230, O54995, Q9CVI4, Q9D0C3, Q9QVR8, Q9R039 | Gene names | Smcx, Jarid1c, Xe169 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1C (SmcX protein) (Xe169 protein). | |||||
|
TCF19_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 28) | NC score | 0.246768 (rank : 21) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9Y242, Q13176, Q15967, Q5SQ89, Q5STD6, Q5STF5, Q9BUM2, Q9UBH7 | Gene names | TCF19, SC1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 19 (Transcription factor SC1). | |||||
|
ING3_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 29) | NC score | 0.145921 (rank : 33) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9NXR8, O60394, Q6GMT3, Q7Z762, Q969G0, Q96DT4, Q9HC99, Q9P081 | Gene names | ING3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 3 (p47ING3 protein). | |||||
|
MTF2_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 30) | NC score | 0.146668 (rank : 32) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9Y483, Q9UES9, Q9UP40 | Gene names | MTF2 | |||
|
Domain Architecture |
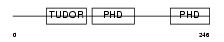
|
|||||
| Description | Metal-response element-binding transcription factor 2 (Metal-response element DNA-binding protein M96) (Metal-regulatory transcription factor 2) (PCL2). | |||||
|
MTF2_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 31) | NC score | 0.145761 (rank : 34) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q02395, Q569Z8, Q6PG89, Q8BGP9, Q8BSJ7 | Gene names | Mtf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metal-response element-binding transcription factor 2 (Zinc-regulated factor 1) (ZiRF1) (Metal-response element DNA-binding protein M96) (Metal-regulatory transcription factor 2) (PCL2). | |||||
|
PHF13_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 32) | NC score | 0.219299 (rank : 24) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q86YI8, Q8N551, Q9UJP2 | Gene names | PHF13 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 13. | |||||
|
PHF13_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 33) | NC score | 0.219939 (rank : 23) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8K2W6 | Gene names | Phf13 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 13. | |||||
|
ING1_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 34) | NC score | 0.159038 (rank : 30) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9UK53, O00532, O43658, Q9H007, Q9HD98, Q9HD99, Q9NS83, Q9P0U6, Q9UBC6, Q9UIJ1, Q9UIJ2, Q9UIJ3, Q9UIJ4, Q9UK52 | Gene names | ING1 | |||
|
Domain Architecture |

|
|||||
| Description | Inhibitor of growth protein 1. | |||||
|
ING3_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 35) | NC score | 0.133909 (rank : 37) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8VEK6, Q99JS6, Q9ERB2 | Gene names | Ing3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 3 (p47ING3 protein). | |||||
|
ING1_MOUSE
|
||||||
| θ value | 0.125558 (rank : 36) | NC score | 0.156384 (rank : 31) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9QXV3, Q9QUP8, Q9QXV4, Q9QZX3 | Gene names | Ing1 | |||
|
Domain Architecture |

|
|||||
| Description | Inhibitor of growth protein 1. | |||||
|
JAD1D_MOUSE
|
||||||
| θ value | 0.125558 (rank : 37) | NC score | 0.121987 (rank : 40) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q62240, Q9QVR9, Q9R040 | Gene names | Smcy, Hya, Jarid1d | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1D (SmcY protein) (Histocompatibility Y antigen) (H-Y). | |||||
|
UBP34_HUMAN
|
||||||
| θ value | 0.125558 (rank : 38) | NC score | 0.028321 (rank : 79) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q70CQ2, O60316, O94834, Q3B777, Q6P6C9, Q7L8P6, Q8N3T9, Q8TBW2, Q9UGA1 | Gene names | USP34, KIAA0570, KIAA0729 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 34 (EC 3.1.2.15) (Ubiquitin thioesterase 34) (Ubiquitin-specific-processing protease 34) (Deubiquitinating enzyme 34). | |||||
|
PHF1_HUMAN
|
||||||
| θ value | 0.21417 (rank : 39) | NC score | 0.112001 (rank : 46) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O43189, O60929, Q96KM7 | Gene names | PHF1 | |||
|
Domain Architecture |
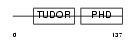
|
|||||
| Description | PHD finger protein 1 (Protein PHF1). | |||||
|
PHF1_MOUSE
|
||||||
| θ value | 0.21417 (rank : 40) | NC score | 0.111720 (rank : 47) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9Z1B8, O54808 | Gene names | Phf1, Tctex-3, Tctex3 | |||
|
Domain Architecture |
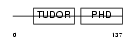
|
|||||
| Description | PHD finger protein 1 (Protein PHF1) (T-complex testis-expressed 3) (Polycomblike 1) (mPCl1). | |||||
|
SRPK2_MOUSE
|
||||||
| θ value | 0.365318 (rank : 41) | NC score | 0.008797 (rank : 108) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 765 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O54781, Q8VCD9 | Gene names | Srpk2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SRPK2 (EC 2.7.11.1) (Serine/arginine- rich protein-specific kinase 2) (SR-protein-specific kinase 2) (SFRS protein kinase 2). | |||||
|
ANR12_HUMAN
|
||||||
| θ value | 0.47712 (rank : 42) | NC score | 0.015967 (rank : 92) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1081 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q6UB98, O94951, Q658K1, Q6QMF7, Q9H231, Q9H784 | Gene names | ANKRD12, ANCO2, KIAA0874 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 12 (Ankyrin repeat-containing cofactor 2) (GAC-1 protein). | |||||
|
IF2P_HUMAN
|
||||||
| θ value | 0.47712 (rank : 43) | NC score | 0.022907 (rank : 88) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 889 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O60841, O95805, Q9UF81, Q9UMN7 | Gene names | EIF5B, IF2, KIAA0741 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 5B (eIF-5B) (Translation initiation factor IF-2). | |||||
|
MLL4_HUMAN
|
||||||
| θ value | 0.62314 (rank : 44) | NC score | 0.133846 (rank : 38) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9UMN6, O15022, O95836, Q96GP2, Q96IP3, Q9UK25, Q9Y668, Q9Y669 | Gene names | MLL4, HRX2, KIAA0304, MLL2, TRX2 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 4 (Trithorax homolog 2). | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | 0.62314 (rank : 45) | NC score | 0.027228 (rank : 81) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
UBP34_MOUSE
|
||||||
| θ value | 0.62314 (rank : 46) | NC score | 0.025965 (rank : 82) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6ZQ93, Q3UPN0, Q6P563, Q7TMJ6, Q8CCH0 | Gene names | Usp34, Kiaa0570 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 34 (EC 3.1.2.15) (Ubiquitin thioesterase 34) (Ubiquitin-specific-processing protease 34) (Deubiquitinating enzyme 34). | |||||
|
UTRO_HUMAN
|
||||||
| θ value | 0.62314 (rank : 47) | NC score | 0.009466 (rank : 106) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P46939 | Gene names | UTRN, DMDL | |||
|
Domain Architecture |

|
|||||
| Description | Utrophin (Dystrophin-related protein 1) (DRP1) (DRP). | |||||
|
MLL2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 48) | NC score | 0.063445 (rank : 66) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
HBP1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 49) | NC score | 0.016432 (rank : 91) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O60381, Q8TBM1, Q8TE93, Q96AJ2 | Gene names | HBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HMG box-containing protein 1 (HMG box transcription factor 1) (High mobility group box transcription factor 1). | |||||
|
KIF9_HUMAN
|
||||||
| θ value | 1.06291 (rank : 50) | NC score | 0.008119 (rank : 109) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9HAQ2, Q86Z28, Q9H8A4 | Gene names | KIF9 | |||
|
Domain Architecture |
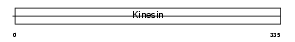
|
|||||
| Description | Kinesin-like protein KIF9. | |||||
|
DSPP_MOUSE
|
||||||
| θ value | 1.38821 (rank : 51) | NC score | 0.019468 (rank : 89) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 52) | NC score | 0.025167 (rank : 83) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
WDR48_HUMAN
|
||||||
| θ value | 1.38821 (rank : 53) | NC score | 0.010781 (rank : 104) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8TAF3, Q63HJ2, Q658Y1, Q8N3Z1, Q9NSK8, Q9P279 | Gene names | WDR48, KIAA1449 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 48 (WD repeat endosomal protein) (p80). | |||||
|
WDR48_MOUSE
|
||||||
| θ value | 1.38821 (rank : 54) | NC score | 0.010909 (rank : 102) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BH57, Q80TD4, Q80XI0, Q8BRM0, Q8CBK0, Q922Z9, Q9CRR1, Q9CSL0 | Gene names | Wdr48, Kiaa1449 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 48. | |||||
|
ZRAB2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 55) | NC score | 0.056046 (rank : 70) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O95218, Q9UP63 | Gene names | ZRANB2, ZIS, ZNF265 | |||
|
Domain Architecture |
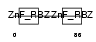
|
|||||
| Description | Zinc finger Ran-binding domain-containing protein 2 Zinc finger protein 265 (Zinc finger, splicing). | |||||
|
ETFD_HUMAN
|
||||||
| θ value | 1.81305 (rank : 56) | NC score | 0.033249 (rank : 75) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q16134, Q7Z347 | Gene names | ETFDH | |||
|
Domain Architecture |

|
|||||
| Description | Electron transfer flavoprotein-ubiquinone oxidoreductase, mitochondrial precursor (EC 1.5.5.1) (ETF-QO) (ETF-ubiquinone oxidoreductase) (ETF dehydrogenase) (Electron-transferring- flavoprotein dehydrogenase). | |||||
|
TXLNB_MOUSE
|
||||||
| θ value | 1.81305 (rank : 57) | NC score | 0.008107 (rank : 110) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 640 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8VBT1, Q3UVB8, Q8BUK2 | Gene names | Txlnb, Mdp77 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-taxilin (Muscle-derived protein 77). | |||||
|
ARI1B_HUMAN
|
||||||
| θ value | 2.36792 (rank : 58) | NC score | 0.028413 (rank : 78) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 567 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8NFD5, Q5JRD1, Q5VYC4, Q8IZY8, Q8TEV0, Q8TF02, Q99491, Q9ULI5 | Gene names | ARID1B, DAN15, KIAA1235, OSA2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-rich interactive domain-containing protein 1B (ARID domain- containing protein 1B) (Osa homolog 2) (hOsa2) (p250R) (BRG1-binding protein hELD/OSA1) (BRG1-associated factor 250b) (BAF250B). | |||||
|
AIOL_MOUSE
|
||||||
| θ value | 3.0926 (rank : 59) | NC score | -0.001625 (rank : 124) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 759 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O08900 | Gene names | Ikzf3, Zfpn1a3, Znfn1a3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein Aiolos (IKAROS family zinc finger protein 3). | |||||
|
DJC12_HUMAN
|
||||||
| θ value | 3.0926 (rank : 60) | NC score | 0.012576 (rank : 100) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UKB3, Q9UKB2 | Gene names | DNAJC12, JDP1 | |||
|
Domain Architecture |
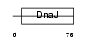
|
|||||
| Description | DnaJ homolog subfamily C member 12 (J domain-containing protein 1). | |||||
|
ING4_HUMAN
|
||||||
| θ value | 3.0926 (rank : 61) | NC score | 0.099462 (rank : 52) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UNL4, Q96E15, Q9H3J0 | Gene names | ING4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 4 (p29ING4). | |||||
|
ING4_MOUSE
|
||||||
| θ value | 3.0926 (rank : 62) | NC score | 0.098438 (rank : 53) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8C0D7, Q8C1S7, Q8K3Q5, Q8K3Q6, Q8K3Q7, Q9D7F9 | Gene names | Ing4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 4 (p29ING4). | |||||
|
IWS1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 63) | NC score | 0.024499 (rank : 85) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
MAML1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 64) | NC score | 0.030947 (rank : 76) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6T264, Q505D8, Q5SUC2, Q6PDK3, Q6ZQG5, Q8BIU5, Q8R3T0 | Gene names | Maml1, Kiaa0200 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 1 (Mam-1). | |||||
|
SFR12_MOUSE
|
||||||
| θ value | 3.0926 (rank : 65) | NC score | 0.029298 (rank : 77) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8BZX4, Q8BLM8, Q8BWK7, Q8BYK0, Q8BYZ4 | Gene names | Sfrs12, Srrp86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86). | |||||
|
TAOK3_MOUSE
|
||||||
| θ value | 3.0926 (rank : 66) | NC score | -0.002112 (rank : 125) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1458 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BYC6, Q3V3B3 | Gene names | Taok3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase TAO3 (EC 2.7.11.1) (Thousand and one amino acid protein 3). | |||||
|
ETFD_MOUSE
|
||||||
| θ value | 4.03905 (rank : 67) | NC score | 0.027401 (rank : 80) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q921G7, Q3U7K2, Q8BK82, Q9DCT9 | Gene names | Etfdh | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Electron transfer flavoprotein-ubiquinone oxidoreductase, mitochondrial precursor (EC 1.5.5.1) (ETF-QO) (ETF-ubiquinone oxidoreductase) (ETF dehydrogenase) (Electron-transferring- flavoprotein dehydrogenase). | |||||
|
ING2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 68) | NC score | 0.118422 (rank : 43) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9H160, O95698 | Gene names | ING2, ING1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 2 (p33ING2) (Inhibitor of growth 1-like protein) (ING1Lp) (p32). | |||||
|
ING2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 69) | NC score | 0.118457 (rank : 42) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9ESK4, Q80VI5, Q8BGU8 | Gene names | Ing2, Ing1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 2 (p33ING2) (Inhibitor of growth 1-like protein). | |||||
|
JPH3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 70) | NC score | 0.007556 (rank : 114) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9ET77, Q9EQZ2 | Gene names | Jph3, Jp3 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-3 (Junctophilin type 3) (JP-3). | |||||
|
MINT_MOUSE
|
||||||
| θ value | 4.03905 (rank : 71) | NC score | 0.023895 (rank : 86) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
PIAS1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 72) | NC score | 0.007497 (rank : 115) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O88907, Q8C6H5 | Gene names | Pias1, Ddxbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein inhibitor of activated STAT protein 1 (DEAD/H box-binding protein 1). | |||||
|
RTEL1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 73) | NC score | 0.013657 (rank : 98) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NZ71, Q5JTV4, Q9BW37, Q9H402, Q9H4X6, Q9NX25, Q9NZ73, Q9UPR4, Q9Y4R6 | Gene names | RTEL1, C20orf41, KIAA1088, NHL | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of telomere elongation helicase 1 (EC 3.6.1.-) (Helicase- like protein NHL). | |||||
|
CBP_HUMAN
|
||||||
| θ value | 5.27518 (rank : 74) | NC score | 0.015406 (rank : 93) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
EYA4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 75) | NC score | 0.014096 (rank : 97) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O95677, O95464, O95679, Q8IW39, Q9NTR7 | Gene names | EYA4 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 4 (EC 3.1.3.48). | |||||
|
EYA4_MOUSE
|
||||||
| θ value | 5.27518 (rank : 76) | NC score | 0.014504 (rank : 96) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Z191 | Gene names | Eya4 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 4 (EC 3.1.3.48). | |||||
|
ITIH4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 77) | NC score | 0.007495 (rank : 116) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q14624, Q15135, Q9P190, Q9UQ54 | Gene names | ITIH4, IHRP, ITIHL1, PK120 | |||
|
Domain Architecture |

|
|||||
| Description | Inter-alpha-trypsin inhibitor heavy chain H4 precursor (ITI heavy chain H4) (Inter-alpha-inhibitor heavy chain 4) (Inter-alpha-trypsin inhibitor family heavy chain-related protein) (IHRP) (Plasma kallikrein sensitive glycoprotein 120) (PK-120) (GP120) [Contains: 70 kDa inter-alpha-trypsin inhibitor heavy chain H4; 35 kDa inter-alpha- trypsin inhibitor heavy chain H4]. | |||||
|
K1C23_HUMAN
|
||||||
| θ value | 5.27518 (rank : 78) | NC score | 0.001522 (rank : 120) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9C075, Q9NUR6 | Gene names | KRT23 | |||
|
Domain Architecture |
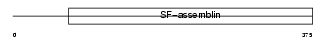
|
|||||
| Description | Keratin, type I cytoskeletal 23 (Cytokeratin-23) (CK-23) (Keratin-23) (K23). | |||||
|
LAMP1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 79) | NC score | 0.010904 (rank : 103) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P11279 | Gene names | LAMP1 | |||
|
Domain Architecture |
|
|||||
| Description | Lysosome-associated membrane glycoprotein 1 precursor (LAMP-1) (CD107a antigen). | |||||
|
LAP2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 80) | NC score | 0.010243 (rank : 105) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 499 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q96RT1, Q86W38, Q9NR18, Q9NW48, Q9ULJ5 | Gene names | ERBB2IP, ERBIN, KIAA1225, LAP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LAP2 (Erbb2-interacting protein) (Erbin) (Densin-180-like protein). | |||||
|
RRP5_HUMAN
|
||||||
| θ value | 5.27518 (rank : 81) | NC score | 0.023121 (rank : 87) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q14690, Q86VQ8 | Gene names | PDCD11, KIAA0185 | |||
|
Domain Architecture |

|
|||||
| Description | RRP5 protein homolog (Programmed cell death protein 11). | |||||
|
SFRS4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 82) | NC score | 0.024707 (rank : 84) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q08170, Q5VXP1, Q9BUA4, Q9UEB5 | Gene names | SFRS4, SRP75 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 4 (Pre-mRNA-splicing factor SRP75) (SRP001LB). | |||||
|
BORIS_HUMAN
|
||||||
| θ value | 6.88961 (rank : 83) | NC score | -0.001589 (rank : 123) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 743 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NI51, Q5JUG4, Q9BZ30, Q9NQJ3 | Gene names | CTCFL, BORIS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcriptional repressor CTCFL (CCCTC-binding factor) (Brother of the regulator of imprinted sites) (Zinc finger protein CTCF-T) (CTCF paralog). | |||||
|
HRX_HUMAN
|
||||||
| θ value | 6.88961 (rank : 84) | NC score | 0.104446 (rank : 49) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
LAP2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 85) | NC score | 0.009067 (rank : 107) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 482 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q80TH2, Q8BQ14, Q8CE41, Q8K171, Q99JU3, Q9JI47 | Gene names | Erbb2ip, Erbin, Kiaa1225, Lap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LAP2 (Erbb2-interacting protein) (Erbin) (Densin-180-like protein). | |||||
|
LRCH2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 86) | NC score | 0.003840 (rank : 117) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q5VUJ6, Q9HA88, Q9P233 | Gene names | LRCH2, KIAA1495 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 2. | |||||
|
MAML1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 87) | NC score | 0.014719 (rank : 95) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q92585, Q9NZ12 | Gene names | MAML1, KIAA0200 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 1 (Mam-1). | |||||
|
PER1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 88) | NC score | 0.008018 (rank : 111) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O15534 | Gene names | PER1, KIAA0482, PER, RIGUI | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (hPER). | |||||
|
SEPP1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 89) | NC score | 0.011774 (rank : 101) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P49908, Q6PD59, Q6PI43, Q6PI87, Q6PJF9 | Gene names | SEPP1, SELP | |||
|
Domain Architecture |

|
|||||
| Description | Selenoprotein P precursor (SeP). | |||||
|
TPR_HUMAN
|
||||||
| θ value | 6.88961 (rank : 90) | NC score | 0.001959 (rank : 118) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
ATX7_HUMAN
|
||||||
| θ value | 8.99809 (rank : 91) | NC score | 0.017872 (rank : 90) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O15265, O75328, O75329, Q9Y6P8 | Gene names | ATXN7, SCA7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-7 (Spinocerebellar ataxia type 7 protein). | |||||
|
CIR_MOUSE
|
||||||
| θ value | 8.99809 (rank : 92) | NC score | 0.012626 (rank : 99) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9DA19, Q3V2N9, Q4KL44, Q52KL9, Q5FW66 | Gene names | Cir | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CBF1-interacting corepressor. | |||||
|
KIF5C_HUMAN
|
||||||
| θ value | 8.99809 (rank : 93) | NC score | 0.001384 (rank : 121) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 974 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O60282, O95079 | Gene names | KIF5C, KIAA0531, NKHC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin heavy chain isoform 5C (Kinesin heavy chain neuron-specific 2). | |||||
|
NCOR2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 94) | NC score | 0.015095 (rank : 94) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Y618, O00613, O15416, Q13354, Q9Y5U0 | Gene names | NCOR2, CTG26 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC) (CTG repeat protein 26) (SMAP270). | |||||
|
PER1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 95) | NC score | 0.007846 (rank : 112) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O35973 | Gene names | Per1, Per, Rigui | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (mPER) (M-Rigui). | |||||
|
RLF_HUMAN
|
||||||
| θ value | 8.99809 (rank : 96) | NC score | -0.001277 (rank : 122) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 775 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q13129, Q9NU60 | Gene names | RLF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein Rlf (Rearranged L-myc fusion gene protein) (Zn-15- related protein). | |||||
|
RNF8_HUMAN
|
||||||
| θ value | 8.99809 (rank : 97) | NC score | 0.001627 (rank : 119) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 622 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O76064 | Gene names | RNF8, KIAA0646 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin ligase protein RNF8 (EC 6.3.2.-) (RING finger protein 8). | |||||
|
ZCH14_HUMAN
|
||||||
| θ value | 8.99809 (rank : 98) | NC score | 0.007749 (rank : 113) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8WYQ9, O60324, Q9UFP0 | Gene names | ZCCHC14, KIAA0579 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 14 (BDG-29). | |||||
|
CENPC_MOUSE
|
||||||
| θ value | θ > 10 (rank : 99) | NC score | 0.052681 (rank : 74) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P49452 | Gene names | Cenpc1, Cenpc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein C 1 (CENP-C) (Centromere autoantigen C). | |||||
|
CXXC6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.055629 (rank : 71) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8NFU7, Q5VUP7, Q7Z6B6, Q8TCR1, Q9C0I7 | Gene names | CXXC6, KIAA1676, LCX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CXXC-type zinc finger protein 6 (Leukemia-associated protein with a CXXC domain). | |||||
|
DNMT1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.110658 (rank : 48) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P26358, Q9UHG5, Q9ULA2, Q9UMZ6 | Gene names | DNMT1, AIM, DNMT | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 1 (EC 2.1.1.37) (Dnmt1) (DNA methyltransferase HsaI) (DNA MTase HsaI) (MCMT) (M.HsaI). | |||||
|
DNMT1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.117135 (rank : 44) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P13864, P97413, Q80ZU3, Q9CSC6, Q9QXX6 | Gene names | Dnmt1, Dnmt, Met1, Uim | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 1 (EC 2.1.1.37) (Dnmt1) (DNA methyltransferase MmuI) (DNA MTase MmuI) (MCMT) (M.MmuI) (Met-1). | |||||
|
FBX37_HUMAN
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.059510 (rank : 68) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H469, Q49AL7, Q5JWA5 | Gene names | FBXL15, FBXO37 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 37 (F-box/LRR-repeat protein 15). | |||||
|
FBXL2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.081431 (rank : 61) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UKC9, Q9NVQ8, Q9UK27, Q9UKA5, Q9Y3Y9 | Gene names | FBXL2, FBL2, FBL3 | |||
|
Domain Architecture |
|
|||||
| Description | F-box/LRR-repeat protein 2 (F-box and leucine-rich repeat protein 2) (F-box protein FBL2/FBL3). | |||||
|
FBXL2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.084337 (rank : 60) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BH16, Q8BXM4 | Gene names | Fbxl2, Fbl2 | |||
|
Domain Architecture |
|
|||||
| Description | F-box/LRR-repeat protein 2 (F-box and leucine-rich repeat protein 2). | |||||
|
FBXL7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 106) | NC score | 0.100211 (rank : 50) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UJT9, O94926 | Gene names | FBXL7, FBL6, FBL7, KIAA0840 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 7 (F-box and leucine-rich repeat protein 7) (F-box protein FBL6/FBL7). | |||||
|
FXL12_HUMAN
|
||||||
| θ value | θ > 10 (rank : 107) | NC score | 0.062967 (rank : 67) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NXK8, Q9H5K4 | Gene names | FBXL12, FBL12 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 12 (F-box and leucine-rich repeat protein 12) (F-box protein FBL12). | |||||
|
FXL12_MOUSE
|
||||||
| θ value | θ > 10 (rank : 108) | NC score | 0.069699 (rank : 65) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9EPX5, Q3UVH7, Q8CDX0, Q9CY04, Q9QZN5 | Gene names | Fbxl12, Fbl12 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 12 (F-box and leucine-rich repeat protein 12) (F-box protein FBL12). | |||||
|
FXL13_HUMAN
|
||||||
| θ value | θ > 10 (rank : 109) | NC score | 0.078493 (rank : 62) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NEE6, Q6UVW7, Q6UVW8, Q75MN5, Q86UJ5, Q8N7Y4, Q8TCL2, Q8WUF9, Q8WUG0 | Gene names | FBXL13, FBL13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 13 (F-box and leucine-rich repeat protein 13). | |||||
|
FXL13_MOUSE
|
||||||
| θ value | θ > 10 (rank : 110) | NC score | 0.088922 (rank : 58) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8CDU4, Q8CDE9 | Gene names | Fbxl13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 13 (F-box and leucine-rich repeat protein 13). | |||||
|
FXL14_HUMAN
|
||||||
| θ value | θ > 10 (rank : 111) | NC score | 0.078060 (rank : 64) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N1E6 | Gene names | FBXL14, FBL14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 14 (F-box and leucine-rich repeat protein 14). | |||||
|
FXL14_MOUSE
|
||||||
| θ value | θ > 10 (rank : 112) | NC score | 0.078281 (rank : 63) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BID8, Q3U2Q9, Q8R5H7, Q8VDT7, Q922N5 | Gene names | Fbxl14, Fbl14, Ppa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 14 (F-box and leucine-rich repeat protein 14). | |||||
|
FXL16_HUMAN
|
||||||
| θ value | θ > 10 (rank : 113) | NC score | 0.090377 (rank : 55) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N461, Q2MHR2, Q96S14, Q9UJI0 | Gene names | FBXL16, C16orf22, FBL16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 16 (F-box and leucine-rich repeat protein 16). | |||||
|
FXL19_HUMAN
|
||||||
| θ value | θ > 10 (rank : 114) | NC score | 0.288271 (rank : 16) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6PCT2, Q8N789, Q9NT14 | Gene names | FBXL19, FBL19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 19 (F-box and leucine-rich repeat protein 19). | |||||
|
FXL19_MOUSE
|
||||||
| θ value | θ > 10 (rank : 115) | NC score | 0.283444 (rank : 17) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6PB97, Q7TSH0, Q8BIB4 | Gene names | Fbxl19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 19 (F-box and leucine-rich repeat protein 19). | |||||
|
FXL20_HUMAN
|
||||||
| θ value | θ > 10 (rank : 116) | NC score | 0.090953 (rank : 54) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96IG2 | Gene names | FBXL20, FBL2 | |||
|
Domain Architecture |
|
|||||
| Description | F-box/LRR-repeat protein 20 (F-box and leucine-rich repeat protein 20) (F-box/LRR-repeat protein 2-like). | |||||
|
FXL20_MOUSE
|
||||||
| θ value | θ > 10 (rank : 117) | NC score | 0.090308 (rank : 56) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9CZV8, Q8BZ95 | Gene names | Fbxl20, Fbl2 | |||
|
Domain Architecture |
|
|||||
| Description | F-box/LRR-repeat protein 20 (F-box and leucine-rich repeat protein 20) (F-box/LRR-repeat protein 2-like). | |||||
|
ING5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 118) | NC score | 0.056069 (rank : 69) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8WYH8, Q9BS30 | Gene names | ING5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 5 (p28ING5). | |||||
|
ING5_MOUSE
|
||||||
| θ value | θ > 10 (rank : 119) | NC score | 0.055226 (rank : 72) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9D8Y8, Q3UL57, Q6P292, Q9CV64, Q9D9V8 | Gene names | Ing5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 5. | |||||
|
INGX_HUMAN
|
||||||
| θ value | θ > 10 (rank : 120) | NC score | 0.055078 (rank : 73) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9P0U5 | Gene names | INGX, ING2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein, X-linked (Inhibitor of growth protein 2) (ING1-like tumor suppressor protein). | |||||
|
LRC29_HUMAN
|
||||||
| θ value | θ > 10 (rank : 121) | NC score | 0.084675 (rank : 59) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8WV35, Q9UKA0 | Gene names | LRRC29, FBL9, FBXL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 29 (F-box/LRR-repeat protein 9) (F-box and leucine-rich repeat protein 9) (F-box protein FBL9). | |||||
|
MBD1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 122) | NC score | 0.121163 (rank : 41) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UIS9, O15248, O95241, Q7Z7B5, Q8N4W4, Q9UNZ6, Q9UNZ7, Q9UNZ8, Q9UNZ9 | Gene names | MBD1, PCM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 1 (Methyl-CpG-binding protein MBD1) (Protein containing methyl-CpG-binding domain 1). | |||||
|
MBD1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 123) | NC score | 0.115364 (rank : 45) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Z2E2, Q792D6, Q8CCL9, Q9DC19 | Gene names | Mbd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 1 (Methyl-CpG-binding protein MBD1). | |||||
|
SKP2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 124) | NC score | 0.099883 (rank : 51) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13309, Q8TDZ0, Q8TDZ1, Q9BV69 | Gene names | SKP2, FBXL1 | |||
|
Domain Architecture |

|
|||||
| Description | S-phase kinase-associated protein 2 (F-box protein Skp2) (Cyclin A/CDK2-associated protein p45) (p45skp2) (F-box/LRR-repeat protein 1). | |||||
|
SKP2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 125) | NC score | 0.089214 (rank : 57) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Z0Z3, Q8BSG7, Q8C8Y9, Q8CB22, Q99J53 | Gene names | Skp2 | |||
|
Domain Architecture |

|
|||||
| Description | S-phase kinase-associated protein 2 (F-box protein Skp2) (Cyclin A/CDK2-associated protein p45) (F-box/WD-40 protein 1) (FWD1). | |||||
|
K1718_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 98 | |
| SwissProt Accessions | Q3UWM4, Q3UWN8, Q6ZPJ5, Q8C969, Q8C9E0, Q91VX8 | Gene names | Kiaa1718 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1718. | |||||
|
K1718_HUMAN
|
||||||
| NC score | 0.991569 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q6ZMT4, Q6MZL8, Q9C0E5 | Gene names | KIAA1718 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1718. | |||||
|
PHF8_MOUSE
|
||||||
| NC score | 0.971984 (rank : 3) | θ value | 2.48782e-175 (rank : 3) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q80TJ7, Q8BLX8, Q8BLY0, Q8BZ61, Q8CG26 | Gene names | Phf8, Kiaa1111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 8. | |||||
|
PHF8_HUMAN
|
||||||
| NC score | 0.969453 (rank : 4) | θ value | 4.6918e-174 (rank : 4) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9UPP1, Q5H9U5, Q5VUJ4, Q7Z6D4, Q9HAH2 | Gene names | PHF8, KIAA1111, ZNF422 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 8. | |||||
|
PHF2_HUMAN
|
||||||
| NC score | 0.950555 (rank : 5) | θ value | 2.04462e-161 (rank : 5) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | O75151, Q8N3K2, Q9Y6N4 | Gene names | PHF2, KIAA0662 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
PHF2_MOUSE
|
||||||
| NC score | 0.948714 (rank : 6) | θ value | 1.01473e-160 (rank : 6) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9WTU0, Q80WA8 | Gene names | Phf2 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
JHD1A_MOUSE
|
||||||
| NC score | 0.776460 (rank : 7) | θ value | 3.24342e-66 (rank : 10) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P59997, Q3U1M5, Q3UR56, Q3V3Q1, Q69ZT4 | Gene names | Fbxl11, Jhdm1a, Kiaa1004 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1A (EC 1.14.11.-) (F-box/LRR-repeat protein 11) (F-box and leucine-rich repeat protein 11). | |||||
|
JHD1A_HUMAN
|
||||||
| NC score | 0.775470 (rank : 8) | θ value | 1.11475e-66 (rank : 9) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9Y2K7, Q49A21, Q4G0M3, Q69YY8, Q9BVH5, Q9H7H5, Q9UK66 | Gene names | FBXL11, FBL7, JHDM1A, KIAA1004 | |||
|
Domain Architecture |

|
|||||
| Description | JmjC domain-containing histone demethylation protein 1A (EC 1.14.11.-) (F-box/LRR-repeat protein 11) (F-box and leucine-rich repeat protein 11) (F-box protein FBL7) (F-box protein Lilina). | |||||
|
JHD1B_HUMAN
|
||||||
| NC score | 0.766997 (rank : 9) | θ value | 2.47765e-74 (rank : 7) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8NHM5, Q8NCI2, Q96HC7, Q96SL0, Q96T03, Q9NS96, Q9UF75 | Gene names | FBXL10, CXXC2, FBL10, JHDM1B, PCCX2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1B (EC 1.14.11.-) (F-box/LRR-repeat protein 10) (F-box and leucine-rich repeat protein 10) (F-box protein FBL10) (Protein JEMMA) (Jumonji domain-containing EMSY-interactor methyltransferase motif protein) (CXXC-type zinc finger protein 2) (Protein-containing CXXC domain 2). | |||||
|
JHD1B_MOUSE
|
||||||
| NC score | 0.766520 (rank : 10) | θ value | 4.22625e-74 (rank : 8) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6P1G2, Q3V396, Q6PFD0, Q6ZPE8, Q9CSF7, Q9QZN6 | Gene names | Fbxl10, Fbl10, Jhdm1b, Kiaa3014 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1B (EC 1.14.11.-) (F-box/LRR-repeat protein 10) (F-box and leucine-rich repeat protein 10) (F-box protein FBL10). | |||||
|
CXCC1_MOUSE
|
||||||
| NC score | 0.500364 (rank : 11) | θ value | 3.29651e-10 (rank : 14) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9CWW7 | Gene names | Cxxc1, Cgbp, Pccx1 | |||
|
Domain Architecture |

|
|||||
| Description | CpG-binding protein (PHD finger and CXXC domain-containing protein 1) (CXXC-type zinc finger protein 1). | |||||
|
CXCC1_HUMAN
|
||||||
| NC score | 0.499871 (rank : 12) | θ value | 1.47974e-10 (rank : 13) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9P0U4, Q8N2W4, Q96BC8, Q9P2V7 | Gene names | CXXC1, CGBP, PCCX1, PHF18 | |||
|
Domain Architecture |

|
|||||
| Description | CpG-binding protein (PHD finger and CXXC domain-containing protein 1) (CXXC-type zinc finger protein 1). | |||||
|
DIDO1_HUMAN
|
||||||
| NC score | 0.297724 (rank : 13) | θ value | 1.33837e-11 (rank : 11) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
CN130_MOUSE
|
||||||
| NC score | 0.297516 (rank : 14) | θ value | 0.000158464 (rank : 15) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BU04, Q80UT9 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C14orf130 homolog. | |||||
|
CN130_HUMAN
|
||||||
| NC score | 0.295518 (rank : 15) | θ value | 0.000270298 (rank : 16) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8N806, Q86U21, Q86UA9, Q96BY0, Q9NVV6 | Gene names | C14orf130 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C14orf130. | |||||
|
FXL19_HUMAN
|
||||||
| NC score | 0.288271 (rank : 16) | θ value | θ > 10 (rank : 114) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6PCT2, Q8N789, Q9NT14 | Gene names | FBXL19, FBL19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 19 (F-box and leucine-rich repeat protein 19). | |||||
|
FXL19_MOUSE
|
||||||
| NC score | 0.283444 (rank : 17) | θ value | θ > 10 (rank : 115) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6PB97, Q7TSH0, Q8BIB4 | Gene names | Fbxl19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 19 (F-box and leucine-rich repeat protein 19). | |||||
|
PTDSR_MOUSE
|
||||||
| NC score | 0.276814 (rank : 18) | θ value | 0.00102713 (rank : 19) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9ERI5, Q80TX1 | Gene names | Ptdsr, Kiaa0585 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein PTDSR (Phosphatidylserine receptor) (Apoptotic cell clearance receptor PtdSerR). | |||||
|
BPTF_HUMAN
|
||||||
| NC score | 0.270670 (rank : 19) | θ value | 5.08577e-11 (rank : 12) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 786 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q12830, Q6NX67, Q7Z7D6, Q9UIG2 | Gene names | FALZ, BPTF, FAC1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleosome remodeling factor subunit BPTF (Bromodomain and PHD finger- containing transcription factor) (Fetal Alzheimer antigen) (Fetal Alz- 50 clone 1 protein). | |||||
|
PTDSR_HUMAN
|
||||||
| NC score | 0.269445 (rank : 20) | θ value | 0.00390308 (rank : 23) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6NYC1, Q86VY0, Q8IUM5, Q9Y4E2 | Gene names | PTDSR, KIAA0585 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein PTDSR (Phosphatidylserine receptor). | |||||
|
TCF19_HUMAN
|
||||||
| NC score | 0.246768 (rank : 21) | θ value | 0.0193708 (rank : 28) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9Y242, Q13176, Q15967, Q5SQ89, Q5STD6, Q5STF5, Q9BUM2, Q9UBH7 | Gene names | TCF19, SC1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 19 (Transcription factor SC1). | |||||
|
PHF3_HUMAN
|
||||||
| NC score | 0.240412 (rank : 22) | θ value | 0.000270298 (rank : 18) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q92576, Q5T1T6, Q9NQ16, Q9UI45 | Gene names | PHF3, KIAA0244 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 3. | |||||
|
PHF13_MOUSE
|
||||||
| NC score | 0.219939 (rank : 23) | θ value | 0.0736092 (rank : 33) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8K2W6 | Gene names | Phf13 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 13. | |||||
|
PHF13_HUMAN
|
||||||
| NC score | 0.219299 (rank : 24) | θ value | 0.0736092 (rank : 32) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q86YI8, Q8N551, Q9UJP2 | Gene names | PHF13 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 13. | |||||
|
TAF3_MOUSE
|
||||||
| NC score | 0.201189 (rank : 25) | θ value | 0.00665767 (rank : 25) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 463 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q5HZG4, Q3U490, Q3UWX2, Q8BIU8, Q99JH4 | Gene names | Taf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
TAF3_HUMAN
|
||||||
| NC score | 0.195474 (rank : 26) | θ value | 0.00298849 (rank : 21) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 568 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q5VWG9, Q6GMS5, Q6P6B5, Q86VY6, Q9BQS9, Q9UFI8 | Gene names | TAF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
PHF20_HUMAN
|
||||||
| NC score | 0.191159 (rank : 27) | θ value | 0.00390308 (rank : 22) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9BVI0, Q5JWY9, Q9BWV4, Q9BXA3, Q9BZW3, Q9H421, Q9H4J6, Q9NZ22 | Gene names | PHF20, C20orf104, GLEA2, HCA58, NZF, TZP | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58) (Glioma-expressed antigen 2) (Transcription factor TZP) (Novel zinc finger protein). | |||||
|
PHF20_MOUSE
|
||||||
| NC score | 0.184268 (rank : 28) | θ value | 0.00298849 (rank : 20) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8BLG0, Q8BMA2, Q8BYR4, Q921N1 | Gene names | Phf20, Hca58 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58 homolog). | |||||
|
JAD1A_HUMAN
|
||||||
| NC score | 0.170570 (rank : 29) | θ value | 0.000270298 (rank : 17) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P29375 | Gene names | JARID1A, RBBP2, RBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1A (Retinoblastoma-binding protein 2) (RBBP-2). | |||||
|
ING1_HUMAN
|
||||||
| NC score | 0.159038 (rank : 30) | θ value | 0.0961366 (rank : 34) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9UK53, O00532, O43658, Q9H007, Q9HD98, Q9HD99, Q9NS83, Q9P0U6, Q9UBC6, Q9UIJ1, Q9UIJ2, Q9UIJ3, Q9UIJ4, Q9UK52 | Gene names | ING1 | |||
|
Domain Architecture |

|
|||||
| Description | Inhibitor of growth protein 1. | |||||
|
ING1_MOUSE
|
||||||
| NC score | 0.156384 (rank : 31) | θ value | 0.125558 (rank : 36) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9QXV3, Q9QUP8, Q9QXV4, Q9QZX3 | Gene names | Ing1 | |||
|
Domain Architecture |

|
|||||
| Description | Inhibitor of growth protein 1. | |||||
|
MTF2_HUMAN
|
||||||
| NC score | 0.146668 (rank : 32) | θ value | 0.0563607 (rank : 30) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9Y483, Q9UES9, Q9UP40 | Gene names | MTF2 | |||
|
Domain Architecture |

|
|||||
| Description | Metal-response element-binding transcription factor 2 (Metal-response element DNA-binding protein M96) (Metal-regulatory transcription factor 2) (PCL2). | |||||
|
ING3_HUMAN
|
||||||
| NC score | 0.145921 (rank : 33) | θ value | 0.0431538 (rank : 29) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9NXR8, O60394, Q6GMT3, Q7Z762, Q969G0, Q96DT4, Q9HC99, Q9P081 | Gene names | ING3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 3 (p47ING3 protein). | |||||
|
MTF2_MOUSE
|
||||||
| NC score | 0.145761 (rank : 34) | θ value | 0.0563607 (rank : 31) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q02395, Q569Z8, Q6PG89, Q8BGP9, Q8BSJ7 | Gene names | Mtf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metal-response element-binding transcription factor 2 (Zinc-regulated factor 1) (ZiRF1) (Metal-response element DNA-binding protein M96) (Metal-regulatory transcription factor 2) (PCL2). | |||||
|
JAD1D_HUMAN
|
||||||
| NC score | 0.144833 (rank : 35) | θ value | 0.00509761 (rank : 24) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9BY66, Q92509, Q92809, Q9HCU1 | Gene names | SMCY, HY, HYA, JARID1D, KIAA0234 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1D (SmcY protein) (Histocompatibility Y antigen) (H-Y). | |||||
|
JAD1C_MOUSE
|
||||||
| NC score | 0.134399 (rank : 36) | θ value | 0.0193708 (rank : 27) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P41230, O54995, Q9CVI4, Q9D0C3, Q9QVR8, Q9R039 | Gene names | Smcx, Jarid1c, Xe169 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1C (SmcX protein) (Xe169 protein). | |||||
|
ING3_MOUSE
|
||||||
| NC score | 0.133909 (rank : 37) | θ value | 0.0961366 (rank : 35) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8VEK6, Q99JS6, Q9ERB2 | Gene names | Ing3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 3 (p47ING3 protein). | |||||
|
MLL4_HUMAN
|
||||||
| NC score | 0.133846 (rank : 38) | θ value | 0.62314 (rank : 44) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9UMN6, O15022, O95836, Q96GP2, Q96IP3, Q9UK25, Q9Y668, Q9Y669 | Gene names | MLL4, HRX2, KIAA0304, MLL2, TRX2 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 4 (Trithorax homolog 2). | |||||
|
JAD1C_HUMAN
|
||||||
| NC score | 0.133610 (rank : 39) | θ value | 0.0193708 (rank : 26) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P41229 | Gene names | SMCX, DXS1272E, JARID1C, XE169 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1C (Protein SmcX) (Protein Xe169). | |||||
|
JAD1D_MOUSE
|
||||||
| NC score | 0.121987 (rank : 40) | θ value | 0.125558 (rank : 37) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q62240, Q9QVR9, Q9R040 | Gene names | Smcy, Hya, Jarid1d | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1D (SmcY protein) (Histocompatibility Y antigen) (H-Y). | |||||
|
MBD1_HUMAN
|
||||||
| NC score | 0.121163 (rank : 41) | θ value | θ > 10 (rank : 122) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UIS9, O15248, O95241, Q7Z7B5, Q8N4W4, Q9UNZ6, Q9UNZ7, Q9UNZ8, Q9UNZ9 | Gene names | MBD1, PCM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 1 (Methyl-CpG-binding protein MBD1) (Protein containing methyl-CpG-binding domain 1). | |||||
|
ING2_MOUSE
|
||||||
| NC score | 0.118457 (rank : 42) | θ value | 4.03905 (rank : 69) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9ESK4, Q80VI5, Q8BGU8 | Gene names | Ing2, Ing1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 2 (p33ING2) (Inhibitor of growth 1-like protein). | |||||
|
ING2_HUMAN
|
||||||
| NC score | 0.118422 (rank : 43) | θ value | 4.03905 (rank : 68) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9H160, O95698 | Gene names | ING2, ING1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 2 (p33ING2) (Inhibitor of growth 1-like protein) (ING1Lp) (p32). | |||||
|
DNMT1_MOUSE
|
||||||
| NC score | 0.117135 (rank : 44) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P13864, P97413, Q80ZU3, Q9CSC6, Q9QXX6 | Gene names | Dnmt1, Dnmt, Met1, Uim | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 1 (EC 2.1.1.37) (Dnmt1) (DNA methyltransferase MmuI) (DNA MTase MmuI) (MCMT) (M.MmuI) (Met-1). | |||||
|
MBD1_MOUSE
|
||||||
| NC score | 0.115364 (rank : 45) | θ value | θ > 10 (rank : 123) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Z2E2, Q792D6, Q8CCL9, Q9DC19 | Gene names | Mbd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 1 (Methyl-CpG-binding protein MBD1). | |||||
|
PHF1_HUMAN
|
||||||
| NC score | 0.112001 (rank : 46) | θ value | 0.21417 (rank : 39) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O43189, O60929, Q96KM7 | Gene names | PHF1 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 1 (Protein PHF1). | |||||
|
PHF1_MOUSE
|
||||||
| NC score | 0.111720 (rank : 47) | θ value | 0.21417 (rank : 40) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9Z1B8, O54808 | Gene names | Phf1, Tctex-3, Tctex3 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 1 (Protein PHF1) (T-complex testis-expressed 3) (Polycomblike 1) (mPCl1). | |||||
|
DNMT1_HUMAN
|
||||||
| NC score | 0.110658 (rank : 48) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P26358, Q9UHG5, Q9ULA2, Q9UMZ6 | Gene names | DNMT1, AIM, DNMT | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 1 (EC 2.1.1.37) (Dnmt1) (DNA methyltransferase HsaI) (DNA MTase HsaI) (MCMT) (M.HsaI). | |||||
|
HRX_HUMAN
|
||||||
| NC score | 0.104446 (rank : 49) | θ value | 6.88961 (rank : 84) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
FBXL7_HUMAN
|
||||||
| NC score | 0.100211 (rank : 50) | θ value | θ > 10 (rank : 106) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UJT9, O94926 | Gene names | FBXL7, FBL6, FBL7, KIAA0840 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 7 (F-box and leucine-rich repeat protein 7) (F-box protein FBL6/FBL7). | |||||
|
SKP2_HUMAN
|
||||||
| NC score | 0.099883 (rank : 51) | θ value | θ > 10 (rank : 124) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13309, Q8TDZ0, Q8TDZ1, Q9BV69 | Gene names | SKP2, FBXL1 | |||
|
Domain Architecture |

|
|||||
| Description | S-phase kinase-associated protein 2 (F-box protein Skp2) (Cyclin A/CDK2-associated protein p45) (p45skp2) (F-box/LRR-repeat protein 1). | |||||
|
ING4_HUMAN
|
||||||
| NC score | 0.099462 (rank : 52) | θ value | 3.0926 (rank : 61) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UNL4, Q96E15, Q9H3J0 | Gene names | ING4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 4 (p29ING4). | |||||
|
ING4_MOUSE
|
||||||
| NC score | 0.098438 (rank : 53) | θ value | 3.0926 (rank : 62) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8C0D7, Q8C1S7, Q8K3Q5, Q8K3Q6, Q8K3Q7, Q9D7F9 | Gene names | Ing4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 4 (p29ING4). | |||||
|
FXL20_HUMAN
|
||||||
| NC score | 0.090953 (rank : 54) | θ value | θ > 10 (rank : 116) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96IG2 | Gene names | FBXL20, FBL2 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 20 (F-box and leucine-rich repeat protein 20) (F-box/LRR-repeat protein 2-like). | |||||
|
FXL16_HUMAN
|
||||||
| NC score | 0.090377 (rank : 55) | θ value | θ > 10 (rank : 113) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N461, Q2MHR2, Q96S14, Q9UJI0 | Gene names | FBXL16, C16orf22, FBL16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 16 (F-box and leucine-rich repeat protein 16). | |||||
|
FXL20_MOUSE
|
||||||
| NC score | 0.090308 (rank : 56) | θ value | θ > 10 (rank : 117) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9CZV8, Q8BZ95 | Gene names | Fbxl20, Fbl2 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 20 (F-box and leucine-rich repeat protein 20) (F-box/LRR-repeat protein 2-like). | |||||
|
SKP2_MOUSE
|
||||||
| NC score | 0.089214 (rank : 57) | θ value | θ > 10 (rank : 125) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Z0Z3, Q8BSG7, Q8C8Y9, Q8CB22, Q99J53 | Gene names | Skp2 | |||
|
Domain Architecture |

|
|||||
| Description | S-phase kinase-associated protein 2 (F-box protein Skp2) (Cyclin A/CDK2-associated protein p45) (F-box/WD-40 protein 1) (FWD1). | |||||
|
FXL13_MOUSE
|
||||||
| NC score | 0.088922 (rank : 58) | θ value | θ > 10 (rank : 110) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8CDU4, Q8CDE9 | Gene names | Fbxl13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 13 (F-box and leucine-rich repeat protein 13). | |||||
|
LRC29_HUMAN
|
||||||
| NC score | 0.084675 (rank : 59) | θ value | θ > 10 (rank : 121) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8WV35, Q9UKA0 | Gene names | LRRC29, FBL9, FBXL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 29 (F-box/LRR-repeat protein 9) (F-box and leucine-rich repeat protein 9) (F-box protein FBL9). | |||||
|
FBXL2_MOUSE
|
||||||
| NC score | 0.084337 (rank : 60) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BH16, Q8BXM4 | Gene names | Fbxl2, Fbl2 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 2 (F-box and leucine-rich repeat protein 2). | |||||
|
FBXL2_HUMAN
|
||||||
| NC score | 0.081431 (rank : 61) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UKC9, Q9NVQ8, Q9UK27, Q9UKA5, Q9Y3Y9 | Gene names | FBXL2, FBL2, FBL3 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 2 (F-box and leucine-rich repeat protein 2) (F-box protein FBL2/FBL3). | |||||
|
FXL13_HUMAN
|
||||||
| NC score | 0.078493 (rank : 62) | θ value | θ > 10 (rank : 109) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NEE6, Q6UVW7, Q6UVW8, Q75MN5, Q86UJ5, Q8N7Y4, Q8TCL2, Q8WUF9, Q8WUG0 | Gene names | FBXL13, FBL13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 13 (F-box and leucine-rich repeat protein 13). | |||||
|
FXL14_MOUSE
|
||||||
| NC score | 0.078281 (rank : 63) | θ value | θ > 10 (rank : 112) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BID8, Q3U2Q9, Q8R5H7, Q8VDT7, Q922N5 | Gene names | Fbxl14, Fbl14, Ppa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 14 (F-box and leucine-rich repeat protein 14). | |||||
|
FXL14_HUMAN
|
||||||
| NC score | 0.078060 (rank : 64) | θ value | θ > 10 (rank : 111) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N1E6 | Gene names | FBXL14, FBL14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 14 (F-box and leucine-rich repeat protein 14). | |||||
|
FXL12_MOUSE
|
||||||
| NC score | 0.069699 (rank : 65) | θ value | θ > 10 (rank : 108) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9EPX5, Q3UVH7, Q8CDX0, Q9CY04, Q9QZN5 | Gene names | Fbxl12, Fbl12 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 12 (F-box and leucine-rich repeat protein 12) (F-box protein FBL12). | |||||
|
MLL2_HUMAN
|
||||||
| NC score | 0.063445 (rank : 66) | θ value | 0.813845 (rank : 48) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
FXL12_HUMAN
|
||||||
| NC score | 0.062967 (rank : 67) | θ value | θ > 10 (rank : 107) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NXK8, Q9H5K4 | Gene names | FBXL12, FBL12 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 12 (F-box and leucine-rich repeat protein 12) (F-box protein FBL12). | |||||
|
FBX37_HUMAN
|
||||||
| NC score | 0.059510 (rank : 68) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H469, Q49AL7, Q5JWA5 | Gene names | FBXL15, FBXO37 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 37 (F-box/LRR-repeat protein 15). | |||||
|
ING5_HUMAN
|
||||||
| NC score | 0.056069 (rank : 69) | θ value | θ > 10 (rank : 118) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8WYH8, Q9BS30 | Gene names | ING5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 5 (p28ING5). | |||||
|
ZRAB2_HUMAN
|
||||||
| NC score | 0.056046 (rank : 70) | θ value | 1.38821 (rank : 55) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O95218, Q9UP63 | Gene names | ZRANB2, ZIS, ZNF265 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger Ran-binding domain-containing protein 2 Zinc finger protein 265 (Zinc finger, splicing). | |||||
|
CXXC6_HUMAN
|
||||||
| NC score | 0.055629 (rank : 71) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8NFU7, Q5VUP7, Q7Z6B6, Q8TCR1, Q9C0I7 | Gene names | CXXC6, KIAA1676, LCX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CXXC-type zinc finger protein 6 (Leukemia-associated protein with a CXXC domain). | |||||
|
ING5_MOUSE
|
||||||
| NC score | 0.055226 (rank : 72) | θ value | θ > 10 (rank : 119) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9D8Y8, Q3UL57, Q6P292, Q9CV64, Q9D9V8 | Gene names | Ing5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 5. | |||||
|
INGX_HUMAN
|
||||||
| NC score | 0.055078 (rank : 73) | θ value | θ > 10 (rank : 120) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9P0U5 | Gene names | INGX, ING2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein, X-linked (Inhibitor of growth protein 2) (ING1-like tumor suppressor protein). | |||||
|
CENPC_MOUSE
|
||||||
| NC score | 0.052681 (rank : 74) | θ value | θ > 10 (rank : 99) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P49452 | Gene names | Cenpc1, Cenpc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein C 1 (CENP-C) (Centromere autoantigen C). | |||||
|
ETFD_HUMAN
|
||||||
| NC score | 0.033249 (rank : 75) | θ value | 1.81305 (rank : 56) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q16134, Q7Z347 | Gene names | ETFDH | |||
|
Domain Architecture |

|
|||||
| Description | Electron transfer flavoprotein-ubiquinone oxidoreductase, mitochondrial precursor (EC 1.5.5.1) (ETF-QO) (ETF-ubiquinone oxidoreductase) (ETF dehydrogenase) (Electron-transferring- flavoprotein dehydrogenase). | |||||
|
MAML1_MOUSE
|
||||||
| NC score | 0.030947 (rank : 76) | θ value | 3.0926 (rank : 64) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6T264, Q505D8, Q5SUC2, Q6PDK3, Q6ZQG5, Q8BIU5, Q8R3T0 | Gene names | Maml1, Kiaa0200 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 1 (Mam-1). | |||||
|
SFR12_MOUSE
|
||||||
| NC score | 0.029298 (rank : 77) | θ value | 3.0926 (rank : 65) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8BZX4, Q8BLM8, Q8BWK7, Q8BYK0, Q8BYZ4 | Gene names | Sfrs12, Srrp86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86). | |||||
|
ARI1B_HUMAN
|
||||||
| NC score | 0.028413 (rank : 78) | θ value | 2.36792 (rank : 58) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 567 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8NFD5, Q5JRD1, Q5VYC4, Q8IZY8, Q8TEV0, Q8TF02, Q99491, Q9ULI5 | Gene names | ARID1B, DAN15, KIAA1235, OSA2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-rich interactive domain-containing protein 1B (ARID domain- containing protein 1B) (Osa homolog 2) (hOsa2) (p250R) (BRG1-binding protein hELD/OSA1) (BRG1-associated factor 250b) (BAF250B). | |||||
|
UBP34_HUMAN
|
||||||
| NC score | 0.028321 (rank : 79) | θ value | 0.125558 (rank : 38) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q70CQ2, O60316, O94834, Q3B777, Q6P6C9, Q7L8P6, Q8N3T9, Q8TBW2, Q9UGA1 | Gene names | USP34, KIAA0570, KIAA0729 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 34 (EC 3.1.2.15) (Ubiquitin thioesterase 34) (Ubiquitin-specific-processing protease 34) (Deubiquitinating enzyme 34). | |||||
|
ETFD_MOUSE
|
||||||
| NC score | 0.027401 (rank : 80) | θ value | 4.03905 (rank : 67) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q921G7, Q3U7K2, Q8BK82, Q9DCT9 | Gene names | Etfdh | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Electron transfer flavoprotein-ubiquinone oxidoreductase, mitochondrial precursor (EC 1.5.5.1) (ETF-QO) (ETF-ubiquinone oxidoreductase) (ETF dehydrogenase) (Electron-transferring- flavoprotein dehydrogenase). | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.027228 (rank : 81) | θ value | 0.62314 (rank : 45) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
UBP34_MOUSE
|
||||||
| NC score | 0.025965 (rank : 82) | θ value | 0.62314 (rank : 46) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6ZQ93, Q3UPN0, Q6P563, Q7TMJ6, Q8CCH0 | Gene names | Usp34, Kiaa0570 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 34 (EC 3.1.2.15) (Ubiquitin thioesterase 34) (Ubiquitin-specific-processing protease 34) (Deubiquitinating enzyme 34). | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.025167 (rank : 83) | θ value | 1.38821 (rank : 52) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
SFRS4_HUMAN
|
||||||
| NC score | 0.024707 (rank : 84) | θ value | 5.27518 (rank : 82) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q08170, Q5VXP1, Q9BUA4, Q9UEB5 | Gene names | SFRS4, SRP75 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 4 (Pre-mRNA-splicing factor SRP75) (SRP001LB). | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.024499 (rank : 85) | θ value | 3.0926 (rank : 63) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
MINT_MOUSE
|
||||||
| NC score | 0.023895 (rank : 86) | θ value | 4.03905 (rank : 71) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
RRP5_HUMAN
|
||||||
| NC score | 0.023121 (rank : 87) | θ value | 5.27518 (rank : 81) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q14690, Q86VQ8 | Gene names | PDCD11, KIAA0185 | |||
|
Domain Architecture |

|
|||||
| Description | RRP5 protein homolog (Programmed cell death protein 11). | |||||
|
IF2P_HUMAN
|
||||||
| NC score | 0.022907 (rank : 88) | θ value | 0.47712 (rank : 43) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 889 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O60841, O95805, Q9UF81, Q9UMN7 | Gene names | EIF5B, IF2, KIAA0741 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 5B (eIF-5B) (Translation initiation factor IF-2). | |||||
|
DSPP_MOUSE
|
||||||
| NC score | 0.019468 (rank : 89) | θ value | 1.38821 (rank : 51) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
ATX7_HUMAN
|
||||||
| NC score | 0.017872 (rank : 90) | θ value | 8.99809 (rank : 91) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O15265, O75328, O75329, Q9Y6P8 | Gene names | ATXN7, SCA7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-7 (Spinocerebellar ataxia type 7 protein). | |||||
|
HBP1_HUMAN
|
||||||
| NC score | 0.016432 (rank : 91) | θ value | 1.06291 (rank : 49) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O60381, Q8TBM1, Q8TE93, Q96AJ2 | Gene names | HBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HMG box-containing protein 1 (HMG box transcription factor 1) (High mobility group box transcription factor 1). | |||||
|
ANR12_HUMAN
|
||||||
| NC score | 0.015967 (rank : 92) | θ value | 0.47712 (rank : 42) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1081 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q6UB98, O94951, Q658K1, Q6QMF7, Q9H231, Q9H784 | Gene names | ANKRD12, ANCO2, KIAA0874 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 12 (Ankyrin repeat-containing cofactor 2) (GAC-1 protein). | |||||
|
CBP_HUMAN
|
||||||
| NC score | 0.015406 (rank : 93) | θ value | 5.27518 (rank : 74) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
NCOR2_HUMAN
|
||||||
| NC score | 0.015095 (rank : 94) | θ value | 8.99809 (rank : 94) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Y618, O00613, O15416, Q13354, Q9Y5U0 | Gene names | NCOR2, CTG26 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC) (CTG repeat protein 26) (SMAP270). | |||||
|
MAML1_HUMAN
|
||||||
| NC score | 0.014719 (rank : 95) | θ value | 6.88961 (rank : 87) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q92585, Q9NZ12 | Gene names | MAML1, KIAA0200 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 1 (Mam-1). | |||||
|
EYA4_MOUSE
|
||||||
| NC score | 0.014504 (rank : 96) | θ value | 5.27518 (rank : 76) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Z191 | Gene names | Eya4 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 4 (EC 3.1.3.48). | |||||
|
EYA4_HUMAN
|
||||||
| NC score | 0.014096 (rank : 97) | θ value | 5.27518 (rank : 75) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O95677, O95464, O95679, Q8IW39, Q9NTR7 | Gene names | EYA4 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 4 (EC 3.1.3.48). | |||||
|
RTEL1_HUMAN
|
||||||
| NC score | 0.013657 (rank : 98) | θ value | 4.03905 (rank : 73) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NZ71, Q5JTV4, Q9BW37, Q9H402, Q9H4X6, Q9NX25, Q9NZ73, Q9UPR4, Q9Y4R6 | Gene names | RTEL1, C20orf41, KIAA1088, NHL | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of telomere elongation helicase 1 (EC 3.6.1.-) (Helicase- like protein NHL). | |||||
|
CIR_MOUSE
|
||||||
| NC score | 0.012626 (rank : 99) | θ value | 8.99809 (rank : 92) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9DA19, Q3V2N9, Q4KL44, Q52KL9, Q5FW66 | Gene names | Cir | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CBF1-interacting corepressor. | |||||
|
DJC12_HUMAN
|
||||||
| NC score | 0.012576 (rank : 100) | θ value | 3.0926 (rank : 60) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UKB3, Q9UKB2 | Gene names | DNAJC12, JDP1 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 12 (J domain-containing protein 1). | |||||
|
SEPP1_HUMAN
|
||||||
| NC score | 0.011774 (rank : 101) | θ value | 6.88961 (rank : 89) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P49908, Q6PD59, Q6PI43, Q6PI87, Q6PJF9 | Gene names | SEPP1, SELP | |||
|
Domain Architecture |

|
|||||
| Description | Selenoprotein P precursor (SeP). | |||||
|
WDR48_MOUSE
|
||||||
| NC score | 0.010909 (rank : 102) | θ value | 1.38821 (rank : 54) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BH57, Q80TD4, Q80XI0, Q8BRM0, Q8CBK0, Q922Z9, Q9CRR1, Q9CSL0 | Gene names | Wdr48, Kiaa1449 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 48. | |||||
|
LAMP1_HUMAN
|
||||||
| NC score | 0.010904 (rank : 103) | θ value | 5.27518 (rank : 79) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P11279 | Gene names | LAMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Lysosome-associated membrane glycoprotein 1 precursor (LAMP-1) (CD107a antigen). | |||||
|
WDR48_HUMAN
|
||||||
| NC score | 0.010781 (rank : 104) | θ value | 1.38821 (rank : 53) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8TAF3, Q63HJ2, Q658Y1, Q8N3Z1, Q9NSK8, Q9P279 | Gene names | WDR48, KIAA1449 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 48 (WD repeat endosomal protein) (p80). | |||||
|
LAP2_HUMAN
|
||||||
| NC score | 0.010243 (rank : 105) | θ value | 5.27518 (rank : 80) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 499 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q96RT1, Q86W38, Q9NR18, Q9NW48, Q9ULJ5 | Gene names | ERBB2IP, ERBIN, KIAA1225, LAP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LAP2 (Erbb2-interacting protein) (Erbin) (Densin-180-like protein). | |||||
|
UTRO_HUMAN
|
||||||
| NC score | 0.009466 (rank : 106) | θ value | 0.62314 (rank : 47) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P46939 | Gene names | UTRN, DMDL | |||
|
Domain Architecture |

|
|||||
| Description | Utrophin (Dystrophin-related protein 1) (DRP1) (DRP). | |||||
|
LAP2_MOUSE
|
||||||
| NC score | 0.009067 (rank : 107) | θ value | 6.88961 (rank : 85) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 482 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q80TH2, Q8BQ14, Q8CE41, Q8K171, Q99JU3, Q9JI47 | Gene names | Erbb2ip, Erbin, Kiaa1225, Lap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LAP2 (Erbb2-interacting protein) (Erbin) (Densin-180-like protein). | |||||
|
SRPK2_MOUSE
|
||||||
| NC score | 0.008797 (rank : 108) | θ value | 0.365318 (rank : 41) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 765 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O54781, Q8VCD9 | Gene names | Srpk2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SRPK2 (EC 2.7.11.1) (Serine/arginine- rich protein-specific kinase 2) (SR-protein-specific kinase 2) (SFRS protein kinase 2). | |||||
|
KIF9_HUMAN
|
||||||
| NC score | 0.008119 (rank : 109) | θ value | 1.06291 (rank : 50) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9HAQ2, Q86Z28, Q9H8A4 | Gene names | KIF9 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF9. | |||||
|
TXLNB_MOUSE
|
||||||
| NC score | 0.008107 (rank : 110) | θ value | 1.81305 (rank : 57) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 640 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8VBT1, Q3UVB8, Q8BUK2 | Gene names | Txlnb, Mdp77 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-taxilin (Muscle-derived protein 77). | |||||
|
PER1_HUMAN
|
||||||
| NC score | 0.008018 (rank : 111) | θ value | 6.88961 (rank : 88) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O15534 | Gene names | PER1, KIAA0482, PER, RIGUI | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (hPER). | |||||
|
PER1_MOUSE
|
||||||
| NC score | 0.007846 (rank : 112) | θ value | 8.99809 (rank : 95) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O35973 | Gene names | Per1, Per, Rigui | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (mPER) (M-Rigui). | |||||
|
ZCH14_HUMAN
|
||||||
| NC score | 0.007749 (rank : 113) | θ value | 8.99809 (rank : 98) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8WYQ9, O60324, Q9UFP0 | Gene names | ZCCHC14, KIAA0579 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 14 (BDG-29). | |||||
|
JPH3_MOUSE
|
||||||
| NC score | 0.007556 (rank : 114) | θ value | 4.03905 (rank : 70) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9ET77, Q9EQZ2 | Gene names | Jph3, Jp3 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-3 (Junctophilin type 3) (JP-3). | |||||
|
PIAS1_MOUSE
|
||||||
| NC score | 0.007497 (rank : 115) | θ value | 4.03905 (rank : 72) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O88907, Q8C6H5 | Gene names | Pias1, Ddxbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein inhibitor of activated STAT protein 1 (DEAD/H box-binding protein 1). | |||||
|
ITIH4_HUMAN
|
||||||
| NC score | 0.007495 (rank : 116) | θ value | 5.27518 (rank : 77) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q14624, Q15135, Q9P190, Q9UQ54 | Gene names | ITIH4, IHRP, ITIHL1, PK120 | |||
|
Domain Architecture |

|
|||||
| Description | Inter-alpha-trypsin inhibitor heavy chain H4 precursor (ITI heavy chain H4) (Inter-alpha-inhibitor heavy chain 4) (Inter-alpha-trypsin inhibitor family heavy chain-related protein) (IHRP) (Plasma kallikrein sensitive glycoprotein 120) (PK-120) (GP120) [Contains: 70 kDa inter-alpha-trypsin inhibitor heavy chain H4; 35 kDa inter-alpha- trypsin inhibitor heavy chain H4]. | |||||
|
LRCH2_HUMAN
|
||||||
| NC score | 0.003840 (rank : 117) | θ value | 6.88961 (rank : 86) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q5VUJ6, Q9HA88, Q9P233 | Gene names | LRCH2, KIAA1495 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 2. | |||||
|
TPR_HUMAN
|
||||||
| NC score | 0.001959 (rank : 118) | θ value | 6.88961 (rank : 90) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
RNF8_HUMAN
|
||||||
| NC score | 0.001627 (rank : 119) | θ value | 8.99809 (rank : 97) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 622 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O76064 | Gene names | RNF8, KIAA0646 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin ligase protein RNF8 (EC 6.3.2.-) (RING finger protein 8). | |||||
|
K1C23_HUMAN
|
||||||
| NC score | 0.001522 (rank : 120) | θ value | 5.27518 (rank : 78) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9C075, Q9NUR6 | Gene names | KRT23 | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type I cytoskeletal 23 (Cytokeratin-23) (CK-23) (Keratin-23) (K23). | |||||
|
KIF5C_HUMAN
|
||||||
| NC score | 0.001384 (rank : 121) | θ value | 8.99809 (rank : 93) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 974 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O60282, O95079 | Gene names | KIF5C, KIAA0531, NKHC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin heavy chain isoform 5C (Kinesin heavy chain neuron-specific 2). | |||||
|
RLF_HUMAN
|
||||||
| NC score | -0.001277 (rank : 122) | θ value | 8.99809 (rank : 96) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 775 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q13129, Q9NU60 | Gene names | RLF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein Rlf (Rearranged L-myc fusion gene protein) (Zn-15- related protein). | |||||
|
BORIS_HUMAN
|
||||||
| NC score | -0.001589 (rank : 123) | θ value | 6.88961 (rank : 83) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 743 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NI51, Q5JUG4, Q9BZ30, Q9NQJ3 | Gene names | CTCFL, BORIS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcriptional repressor CTCFL (CCCTC-binding factor) (Brother of the regulator of imprinted sites) (Zinc finger protein CTCF-T) (CTCF paralog). | |||||
|
AIOL_MOUSE
|
||||||
| NC score | -0.001625 (rank : 124) | θ value | 3.0926 (rank : 59) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 759 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O08900 | Gene names | Ikzf3, Zfpn1a3, Znfn1a3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein Aiolos (IKAROS family zinc finger protein 3). | |||||
|
TAOK3_MOUSE
|
||||||
| NC score | -0.002112 (rank : 125) | θ value | 3.0926 (rank : 66) | |||
| Query Neighborhood Hits | 98 | Target Neighborhood Hits | 1458 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BYC6, Q3V3B3 | Gene names | Taok3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase TAO3 (EC 2.7.11.1) (Thousand and one amino acid protein 3). | |||||