Please be patient as the page loads
|
K1718_HUMAN
|
||||||
| SwissProt Accessions | Q6ZMT4, Q6MZL8, Q9C0E5 | Gene names | KIAA1718 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1718. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
K1718_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 114 | |
| SwissProt Accessions | Q6ZMT4, Q6MZL8, Q9C0E5 | Gene names | KIAA1718 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1718. | |||||
|
K1718_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.991569 (rank : 2) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q3UWM4, Q3UWN8, Q6ZPJ5, Q8C969, Q8C9E0, Q91VX8 | Gene names | Kiaa1718 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1718. | |||||
|
PHF8_MOUSE
|
||||||
| θ value | 2.48782e-175 (rank : 3) | NC score | 0.969096 (rank : 3) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q80TJ7, Q8BLX8, Q8BLY0, Q8BZ61, Q8CG26 | Gene names | Phf8, Kiaa1111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 8. | |||||
|
PHF8_HUMAN
|
||||||
| θ value | 2.32852e-173 (rank : 4) | NC score | 0.966548 (rank : 4) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9UPP1, Q5H9U5, Q5VUJ4, Q7Z6D4, Q9HAH2 | Gene names | PHF8, KIAA1111, ZNF422 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 8. | |||||
|
PHF2_HUMAN
|
||||||
| θ value | 2.85304e-163 (rank : 5) | NC score | 0.947579 (rank : 5) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O75151, Q8N3K2, Q9Y6N4 | Gene names | PHF2, KIAA0662 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
PHF2_MOUSE
|
||||||
| θ value | 4.11978e-162 (rank : 6) | NC score | 0.945595 (rank : 6) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9WTU0, Q80WA8 | Gene names | Phf2 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
JHD1B_HUMAN
|
||||||
| θ value | 1.60597e-73 (rank : 7) | NC score | 0.763773 (rank : 9) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8NHM5, Q8NCI2, Q96HC7, Q96SL0, Q96T03, Q9NS96, Q9UF75 | Gene names | FBXL10, CXXC2, FBL10, JHDM1B, PCCX2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1B (EC 1.14.11.-) (F-box/LRR-repeat protein 10) (F-box and leucine-rich repeat protein 10) (F-box protein FBL10) (Protein JEMMA) (Jumonji domain-containing EMSY-interactor methyltransferase motif protein) (CXXC-type zinc finger protein 2) (Protein-containing CXXC domain 2). | |||||
|
JHD1B_MOUSE
|
||||||
| θ value | 2.73936e-73 (rank : 8) | NC score | 0.763494 (rank : 10) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6P1G2, Q3V396, Q6PFD0, Q6ZPE8, Q9CSF7, Q9QZN6 | Gene names | Fbxl10, Fbl10, Jhdm1b, Kiaa3014 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1B (EC 1.14.11.-) (F-box/LRR-repeat protein 10) (F-box and leucine-rich repeat protein 10) (F-box protein FBL10). | |||||
|
JHD1A_HUMAN
|
||||||
| θ value | 1.90147e-66 (rank : 9) | NC score | 0.772707 (rank : 8) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9Y2K7, Q49A21, Q4G0M3, Q69YY8, Q9BVH5, Q9H7H5, Q9UK66 | Gene names | FBXL11, FBL7, JHDM1A, KIAA1004 | |||
|
Domain Architecture |

|
|||||
| Description | JmjC domain-containing histone demethylation protein 1A (EC 1.14.11.-) (F-box/LRR-repeat protein 11) (F-box and leucine-rich repeat protein 11) (F-box protein FBL7) (F-box protein Lilina). | |||||
|
JHD1A_MOUSE
|
||||||
| θ value | 4.23606e-66 (rank : 10) | NC score | 0.773515 (rank : 7) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P59997, Q3U1M5, Q3UR56, Q3V3Q1, Q69ZT4 | Gene names | Fbxl11, Jhdm1a, Kiaa1004 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1A (EC 1.14.11.-) (F-box/LRR-repeat protein 11) (F-box and leucine-rich repeat protein 11). | |||||
|
DIDO1_HUMAN
|
||||||
| θ value | 2.43908e-13 (rank : 11) | NC score | 0.297907 (rank : 13) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
BPTF_HUMAN
|
||||||
| θ value | 5.08577e-11 (rank : 12) | NC score | 0.273164 (rank : 19) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 786 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q12830, Q6NX67, Q7Z7D6, Q9UIG2 | Gene names | FALZ, BPTF, FAC1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleosome remodeling factor subunit BPTF (Bromodomain and PHD finger- containing transcription factor) (Fetal Alzheimer antigen) (Fetal Alz- 50 clone 1 protein). | |||||
|
CXCC1_HUMAN
|
||||||
| θ value | 4.30538e-10 (rank : 13) | NC score | 0.495115 (rank : 12) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9P0U4, Q8N2W4, Q96BC8, Q9P2V7 | Gene names | CXXC1, CGBP, PCCX1, PHF18 | |||
|
Domain Architecture |
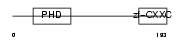
|
|||||
| Description | CpG-binding protein (PHD finger and CXXC domain-containing protein 1) (CXXC-type zinc finger protein 1). | |||||
|
CXCC1_MOUSE
|
||||||
| θ value | 9.59137e-10 (rank : 14) | NC score | 0.495770 (rank : 11) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9CWW7 | Gene names | Cxxc1, Cgbp, Pccx1 | |||
|
Domain Architecture |

|
|||||
| Description | CpG-binding protein (PHD finger and CXXC domain-containing protein 1) (CXXC-type zinc finger protein 1). | |||||
|
CN130_MOUSE
|
||||||
| θ value | 0.00020696 (rank : 15) | NC score | 0.295896 (rank : 14) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8BU04, Q80UT9 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C14orf130 homolog. | |||||
|
JAD1A_HUMAN
|
||||||
| θ value | 0.00020696 (rank : 16) | NC score | 0.172414 (rank : 29) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P29375 | Gene names | JARID1A, RBBP2, RBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1A (Retinoblastoma-binding protein 2) (RBBP-2). | |||||
|
CN130_HUMAN
|
||||||
| θ value | 0.00035302 (rank : 17) | NC score | 0.293970 (rank : 15) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8N806, Q86U21, Q86UA9, Q96BY0, Q9NVV6 | Gene names | C14orf130 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C14orf130. | |||||
|
PHF3_HUMAN
|
||||||
| θ value | 0.000602161 (rank : 18) | NC score | 0.239056 (rank : 22) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q92576, Q5T1T6, Q9NQ16, Q9UI45 | Gene names | PHF3, KIAA0244 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 3. | |||||
|
PTDSR_MOUSE
|
||||||
| θ value | 0.00175202 (rank : 19) | NC score | 0.275465 (rank : 18) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9ERI5, Q80TX1 | Gene names | Ptdsr, Kiaa0585 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein PTDSR (Phosphatidylserine receptor) (Apoptotic cell clearance receptor PtdSerR). | |||||
|
PTDSR_HUMAN
|
||||||
| θ value | 0.00298849 (rank : 20) | NC score | 0.268190 (rank : 20) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6NYC1, Q86VY0, Q8IUM5, Q9Y4E2 | Gene names | PTDSR, KIAA0585 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein PTDSR (Phosphatidylserine receptor). | |||||
|
TAF3_HUMAN
|
||||||
| θ value | 0.00298849 (rank : 21) | NC score | 0.193980 (rank : 26) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 568 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q5VWG9, Q6GMS5, Q6P6B5, Q86VY6, Q9BQS9, Q9UFI8 | Gene names | TAF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
JAD1D_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 22) | NC score | 0.146006 (rank : 35) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9BY66, Q92509, Q92809, Q9HCU1 | Gene names | SMCY, HY, HYA, JARID1D, KIAA0234 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1D (SmcY protein) (Histocompatibility Y antigen) (H-Y). | |||||
|
SFRS4_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 23) | NC score | 0.031168 (rank : 79) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q08170, Q5VXP1, Q9BUA4, Q9UEB5 | Gene names | SFRS4, SRP75 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 4 (Pre-mRNA-splicing factor SRP75) (SRP001LB). | |||||
|
TAF3_MOUSE
|
||||||
| θ value | 0.00665767 (rank : 24) | NC score | 0.201609 (rank : 25) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 463 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q5HZG4, Q3U490, Q3UWX2, Q8BIU8, Q99JH4 | Gene names | Taf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
PKP4_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 25) | NC score | 0.030796 (rank : 80) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q68FH0, Q640N0, Q68G56, Q8BK47, Q8BVH1, Q9CRE3 | Gene names | Pkp4, Armrp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plakophilin-4 (Armadillo-related protein). | |||||
|
JAD1C_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 26) | NC score | 0.134528 (rank : 37) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P41229 | Gene names | SMCX, DXS1272E, JARID1C, XE169 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1C (Protein SmcX) (Protein Xe169). | |||||
|
JAD1C_MOUSE
|
||||||
| θ value | 0.0148317 (rank : 27) | NC score | 0.135272 (rank : 36) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P41230, O54995, Q9CVI4, Q9D0C3, Q9QVR8, Q9R039 | Gene names | Smcx, Jarid1c, Xe169 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1C (SmcX protein) (Xe169 protein). | |||||
|
DSPP_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 28) | NC score | 0.036070 (rank : 78) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
PHF20_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 29) | NC score | 0.181324 (rank : 27) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9BVI0, Q5JWY9, Q9BWV4, Q9BXA3, Q9BZW3, Q9H421, Q9H4J6, Q9NZ22 | Gene names | PHF20, C20orf104, GLEA2, HCA58, NZF, TZP | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58) (Glioma-expressed antigen 2) (Transcription factor TZP) (Novel zinc finger protein). | |||||
|
PHF20_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 30) | NC score | 0.174246 (rank : 28) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8BLG0, Q8BMA2, Q8BYR4, Q921N1 | Gene names | Phf20, Hca58 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58 homolog). | |||||
|
BRWD1_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 31) | NC score | 0.024142 (rank : 83) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 448 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NSI6, O43721, Q5R2V0, Q5R2V1, Q8TCV3, Q96QG9, Q96QH0, Q9NUK1 | Gene names | BRWD1, C21orf107, WDR9 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain and WD repeat domain-containing protein 1 (WD repeat protein 9). | |||||
|
GOGA5_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 32) | NC score | 0.014805 (rank : 100) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 1434 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9QYE6, O88317, Q3TGE7, Q3U6S5, Q3UUF9 | Gene names | Golga5, Retii | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 5 (Golgin-84) (Sumiko protein) (Ret-II protein). | |||||
|
MTF2_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 33) | NC score | 0.148187 (rank : 32) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9Y483, Q9UES9, Q9UP40 | Gene names | MTF2 | |||
|
Domain Architecture |
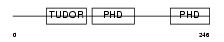
|
|||||
| Description | Metal-response element-binding transcription factor 2 (Metal-response element DNA-binding protein M96) (Metal-regulatory transcription factor 2) (PCL2). | |||||
|
MTF2_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 34) | NC score | 0.147281 (rank : 33) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q02395, Q569Z8, Q6PG89, Q8BGP9, Q8BSJ7 | Gene names | Mtf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metal-response element-binding transcription factor 2 (Zinc-regulated factor 1) (ZiRF1) (Metal-response element DNA-binding protein M96) (Metal-regulatory transcription factor 2) (PCL2). | |||||
|
TCF19_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 35) | NC score | 0.244325 (rank : 21) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9Y242, Q13176, Q15967, Q5SQ89, Q5STD6, Q5STF5, Q9BUM2, Q9UBH7 | Gene names | TCF19, SC1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 19 (Transcription factor SC1). | |||||
|
ING3_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 36) | NC score | 0.146231 (rank : 34) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9NXR8, O60394, Q6GMT3, Q7Z762, Q969G0, Q96DT4, Q9HC99, Q9P081 | Gene names | ING3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 3 (p47ING3 protein). | |||||
|
PHF13_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 37) | NC score | 0.216418 (rank : 24) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q86YI8, Q8N551, Q9UJP2 | Gene names | PHF13 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 13. | |||||
|
PHF13_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 38) | NC score | 0.217087 (rank : 23) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8K2W6 | Gene names | Phf13 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 13. | |||||
|
JAD1D_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 39) | NC score | 0.123206 (rank : 40) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q62240, Q9QVR9, Q9R040 | Gene names | Smcy, Hya, Jarid1d | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1D (SmcY protein) (Histocompatibility Y antigen) (H-Y). | |||||
|
ING1_HUMAN
|
||||||
| θ value | 0.125558 (rank : 40) | NC score | 0.158661 (rank : 30) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9UK53, O00532, O43658, Q9H007, Q9HD98, Q9HD99, Q9NS83, Q9P0U6, Q9UBC6, Q9UIJ1, Q9UIJ2, Q9UIJ3, Q9UIJ4, Q9UK52 | Gene names | ING1 | |||
|
Domain Architecture |

|
|||||
| Description | Inhibitor of growth protein 1. | |||||
|
ING1_MOUSE
|
||||||
| θ value | 0.125558 (rank : 41) | NC score | 0.156406 (rank : 31) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9QXV3, Q9QUP8, Q9QXV4, Q9QZX3 | Gene names | Ing1 | |||
|
Domain Architecture |

|
|||||
| Description | Inhibitor of growth protein 1. | |||||
|
PHF1_HUMAN
|
||||||
| θ value | 0.125558 (rank : 42) | NC score | 0.113646 (rank : 45) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O43189, O60929, Q96KM7 | Gene names | PHF1 | |||
|
Domain Architecture |
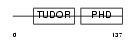
|
|||||
| Description | PHD finger protein 1 (Protein PHF1). | |||||
|
PHF1_MOUSE
|
||||||
| θ value | 0.125558 (rank : 43) | NC score | 0.113478 (rank : 46) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9Z1B8, O54808 | Gene names | Phf1, Tctex-3, Tctex3 | |||
|
Domain Architecture |
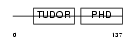
|
|||||
| Description | PHD finger protein 1 (Protein PHF1) (T-complex testis-expressed 3) (Polycomblike 1) (mPCl1). | |||||
|
ING3_MOUSE
|
||||||
| θ value | 0.163984 (rank : 44) | NC score | 0.134206 (rank : 38) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8VEK6, Q99JS6, Q9ERB2 | Gene names | Ing3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 3 (p47ING3 protein). | |||||
|
MPP8_HUMAN
|
||||||
| θ value | 0.47712 (rank : 45) | NC score | 0.017737 (rank : 93) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 948 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q99549, Q86TK3, Q9BTP1 | Gene names | MPHOSPH8, MPP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | M-phase phosphoprotein 8. | |||||
|
NOP14_MOUSE
|
||||||
| θ value | 0.813845 (rank : 46) | NC score | 0.021875 (rank : 88) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 324 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8R3N1 | Gene names | Nop14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable nucleolar complex protein 14. | |||||
|
ZF106_HUMAN
|
||||||
| θ value | 0.813845 (rank : 47) | NC score | 0.022618 (rank : 85) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 470 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9H2Y7, Q6NSD9, Q6PEK1, Q86T43, Q86T45, Q86T50, Q86T58, Q86TA9, Q96M37, Q9H7B8 | Gene names | ZFP106, ZNF474 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 106 homolog (Zfp-106) (Zinc finger protein 474). | |||||
|
ZRAB2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 48) | NC score | 0.054409 (rank : 71) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O95218, Q9UP63 | Gene names | ZRANB2, ZIS, ZNF265 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger Ran-binding domain-containing protein 2 Zinc finger protein 265 (Zinc finger, splicing). | |||||
|
DDX46_MOUSE
|
||||||
| θ value | 1.06291 (rank : 49) | NC score | 0.009936 (rank : 128) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 358 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q569Z5, Q6ZQ42, Q8R0R6 | Gene names | Ddx46, Kiaa0801 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX46 (EC 3.6.1.-) (DEAD box protein 46). | |||||
|
HPS5_MOUSE
|
||||||
| θ value | 1.06291 (rank : 50) | NC score | 0.020993 (rank : 89) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P59438 | Gene names | Hps5, Ru2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hermansky-Pudlak syndrome 5 protein homolog (Ruby-eye protein 2) (Ru2). | |||||
|
MLL4_HUMAN
|
||||||
| θ value | 1.06291 (rank : 51) | NC score | 0.129289 (rank : 39) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9UMN6, O15022, O95836, Q96GP2, Q96IP3, Q9UK25, Q9Y668, Q9Y669 | Gene names | MLL4, HRX2, KIAA0304, MLL2, TRX2 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 4 (Trithorax homolog 2). | |||||
|
MYH9_HUMAN
|
||||||
| θ value | 1.06291 (rank : 52) | NC score | 0.005485 (rank : 134) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P35579, O60805 | Gene names | MYH9 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
LMO7_HUMAN
|
||||||
| θ value | 1.38821 (rank : 53) | NC score | 0.024029 (rank : 84) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8WWI1, O15462, O95346, Q9UKC1, Q9UQM5, Q9Y6A7 | Gene names | LMO7, FBX20, FBXO20, KIAA0858 | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain only protein 7 (LOMP) (F-box only protein 20). | |||||
|
RAD50_MOUSE
|
||||||
| θ value | 1.38821 (rank : 54) | NC score | 0.010010 (rank : 127) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 1366 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P70388, Q6PEB0, Q8C2T7, Q9CU59 | Gene names | Rad50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (mRad50). | |||||
|
ING4_HUMAN
|
||||||
| θ value | 1.81305 (rank : 55) | NC score | 0.099962 (rank : 49) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UNL4, Q96E15, Q9H3J0 | Gene names | ING4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 4 (p29ING4). | |||||
|
ING4_MOUSE
|
||||||
| θ value | 1.81305 (rank : 56) | NC score | 0.099049 (rank : 52) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8C0D7, Q8C1S7, Q8K3Q5, Q8K3Q6, Q8K3Q7, Q9D7F9 | Gene names | Ing4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 4 (p29ING4). | |||||
|
MYST3_HUMAN
|
||||||
| θ value | 1.81305 (rank : 57) | NC score | 0.050043 (rank : 77) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q92794 | Gene names | MYST3, MOZ, RUNXBP2, ZNF220 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Runt-related transcription factor-binding protein 2) (Monocytic leukemia zinc finger protein) (Zinc finger protein 220). | |||||
|
UTRO_HUMAN
|
||||||
| θ value | 1.81305 (rank : 58) | NC score | 0.011372 (rank : 116) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P46939 | Gene names | UTRN, DMDL | |||
|
Domain Architecture |

|
|||||
| Description | Utrophin (Dystrophin-related protein 1) (DRP1) (DRP). | |||||
|
WDR22_MOUSE
|
||||||
| θ value | 1.81305 (rank : 59) | NC score | 0.013101 (rank : 109) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80T85, Q80VT3, Q80ZW6, Q8BIP8, Q8BVK5, Q8K0Q6 | Gene names | Wdr22, Kiaa1824 | |||
|
Domain Architecture |
|
|||||
| Description | WD repeat protein 22. | |||||
|
BORIS_HUMAN
|
||||||
| θ value | 2.36792 (rank : 60) | NC score | -0.001354 (rank : 141) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 743 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8NI51, Q5JUG4, Q9BZ30, Q9NQJ3 | Gene names | CTCFL, BORIS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcriptional repressor CTCFL (CCCTC-binding factor) (Brother of the regulator of imprinted sites) (Zinc finger protein CTCF-T) (CTCF paralog). | |||||
|
CEP55_MOUSE
|
||||||
| θ value | 2.36792 (rank : 61) | NC score | 0.012308 (rank : 112) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 693 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BT07, Q8C2J0, Q8K2I8, Q8R2Y4, Q9DBZ8 | Gene names | Cep55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome protein 55. | |||||
|
K1383_HUMAN
|
||||||
| θ value | 2.36792 (rank : 62) | NC score | 0.028502 (rank : 81) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9P2G4, Q58EZ9, Q5VV83 | Gene names | KIAA1383 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1383. | |||||
|
MLL2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 63) | NC score | 0.061473 (rank : 67) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
PININ_HUMAN
|
||||||
| θ value | 2.36792 (rank : 64) | NC score | 0.014997 (rank : 98) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 753 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9H307, O60899, Q53EM7, Q6P5X4, Q7KYL1, Q99738, Q9UHZ9, Q9UQR9 | Gene names | PNN, DRS, MEMA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pinin (140 kDa nuclear and cell adhesion-related phosphoprotein) (Domain-rich serine protein) (DRS-protein) (DRSP) (Melanoma metastasis clone A protein) (Desmosome-associated protein) (SR-like protein) (Nuclear protein SDK3). | |||||
|
RXINP_MOUSE
|
||||||
| θ value | 2.36792 (rank : 65) | NC score | 0.017941 (rank : 92) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5U5Q9, Q60811 | Gene names | Rxrip110, Rip110 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoid X receptor-interacting protein 110. | |||||
|
SMG7_MOUSE
|
||||||
| θ value | 2.36792 (rank : 66) | NC score | 0.019557 (rank : 91) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q5RJH6, Q63ZW5, Q6ZQF3 | Gene names | Smg7, Est1c, Kiaa0250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein SMG7 (SMG-7 homolog) (EST1-like protein C). | |||||
|
CA103_MOUSE
|
||||||
| θ value | 3.0926 (rank : 67) | NC score | 0.016125 (rank : 96) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CDD9, Q8C893 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C1orf103 homolog. | |||||
|
DPOLZ_HUMAN
|
||||||
| θ value | 3.0926 (rank : 68) | NC score | 0.012848 (rank : 110) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O60673, O43214 | Gene names | REV3L, POLZ, REV3 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase zeta catalytic subunit (EC 2.7.7.7) (hREV3). | |||||
|
EYA4_HUMAN
|
||||||
| θ value | 3.0926 (rank : 69) | NC score | 0.016475 (rank : 94) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O95677, O95464, O95679, Q8IW39, Q9NTR7 | Gene names | EYA4 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 4 (EC 3.1.3.48). | |||||
|
EYA4_MOUSE
|
||||||
| θ value | 3.0926 (rank : 70) | NC score | 0.016418 (rank : 95) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Z191 | Gene names | Eya4 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 4 (EC 3.1.3.48). | |||||
|
OSBL8_HUMAN
|
||||||
| θ value | 3.0926 (rank : 71) | NC score | 0.011529 (rank : 115) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9BZF1, Q8WXP8, Q9P277 | Gene names | OSBPL8, KIAA1451, ORP8, OSBP10 | |||
|
Domain Architecture |

|
|||||
| Description | Oxysterol-binding protein-related protein 8 (OSBP-related protein 8) (ORP-8). | |||||
|
A4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 72) | NC score | 0.012690 (rank : 111) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P05067, P09000, P78438, Q13764, Q13778, Q13793, Q16011, Q16014, Q16019, Q16020, Q9BT38, Q9UCA9, Q9UCB6, Q9UCC8, Q9UCD1, Q9UQ58 | Gene names | APP, A4, AD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 protein precursor (APP) (ABPP) (Alzheimer disease amyloid protein) (Cerebral vascular amyloid peptide) (CVAP) (Protease nexin-II) (PN-II) (APPI) (PreA4) [Contains: Soluble APP-alpha (S-APP- alpha); Soluble APP-beta (S-APP-beta); C99; Beta-amyloid protein 42 (Beta-APP42); Beta-amyloid protein 40 (Beta-APP40); C83; P3(42); P3(40); Gamma-CTF(59) (Gamma-secretase C-terminal fragment 59) (Amyloid intracellular domain 59) (AID(59)); Gamma-CTF(57) (Gamma- secretase C-terminal fragment 57) (Amyloid intracellular domain 57) (AID(57)); Gamma-CTF(50) (Gamma-secretase C-terminal fragment 50) (Amyloid intracellular domain 50) (AID(50)); C31]. | |||||
|
APLP1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 73) | NC score | 0.014860 (rank : 99) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q03157, Q8VC38 | Gene names | Aplp1 | |||
|
Domain Architecture |
|
|||||
| Description | Amyloid-like protein 1 precursor (APLP) (APLP-1) [Contains: C30]. | |||||
|
CASC5_HUMAN
|
||||||
| θ value | 4.03905 (rank : 74) | NC score | 0.013669 (rank : 105) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 376 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8NG31, Q8NHE1, Q8WXA6, Q9HCK2, Q9NR92 | Gene names | CASC5, KIAA1570 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein CASC5 (Cancer susceptibility candidate gene 5 protein) (ALL1- fused gene from chromosome 15q14) (AF15q14) (D40/AF15q14 protein). | |||||
|
CJ118_HUMAN
|
||||||
| θ value | 4.03905 (rank : 75) | NC score | 0.010560 (rank : 122) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 937 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q7Z3E2, Q2M2V6, Q6NS91, Q7RTP1, Q8N117, Q8N3G3, Q8N6C2, Q9NWA3 | Gene names | C10orf118 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf118 (CTCL tumor antigen HD-CL-01/L14-2). | |||||
|
DDX46_HUMAN
|
||||||
| θ value | 4.03905 (rank : 76) | NC score | 0.007897 (rank : 130) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7L014, O94894, Q96EI0, Q9Y658 | Gene names | DDX46, KIAA0801 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX46 (EC 3.6.1.-) (DEAD box protein 46) (PRP5 homolog). | |||||
|
ETFD_HUMAN
|
||||||
| θ value | 4.03905 (rank : 77) | NC score | 0.027774 (rank : 82) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q16134, Q7Z347 | Gene names | ETFDH | |||
|
Domain Architecture |

|
|||||
| Description | Electron transfer flavoprotein-ubiquinone oxidoreductase, mitochondrial precursor (EC 1.5.5.1) (ETF-QO) (ETF-ubiquinone oxidoreductase) (ETF dehydrogenase) (Electron-transferring- flavoprotein dehydrogenase). | |||||
|
EXOSX_HUMAN
|
||||||
| θ value | 4.03905 (rank : 78) | NC score | 0.013626 (rank : 106) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q01780, Q15158 | Gene names | EXOSC10, PMSCL, PMSCL2 | |||
|
Domain Architecture |

|
|||||
| Description | Exosome component 10 (Polymyositis/scleroderma autoantigen 2) (Autoantigen PM/Scl 2) (Polymyositis/scleroderma autoantigen 100 kDa) (PM/Scl-100) (P100 polymyositis-scleroderma overlap syndrome- associated autoantigen). | |||||
|
FANCM_MOUSE
|
||||||
| θ value | 4.03905 (rank : 79) | NC score | 0.012302 (rank : 113) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BGE5, Q69ZF5, Q8BKB7, Q8BUA8 | Gene names | Fancm, Kiaa1596 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fanconi anemia group M protein homolog (EC 3.6.1.-) (ATP-dependent RNA helicase FANCM) (Protein FACM). | |||||
|
GP158_HUMAN
|
||||||
| θ value | 4.03905 (rank : 80) | NC score | 0.013146 (rank : 108) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5T848, Q6QR81, Q9ULT3 | Gene names | GPR158, KIAA1136 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 158 precursor. | |||||
|
ING2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 81) | NC score | 0.119166 (rank : 42) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9H160, O95698 | Gene names | ING2, ING1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 2 (p33ING2) (Inhibitor of growth 1-like protein) (ING1Lp) (p32). | |||||
|
ING2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 82) | NC score | 0.119471 (rank : 41) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9ESK4, Q80VI5, Q8BGU8 | Gene names | Ing2, Ing1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 2 (p33ING2) (Inhibitor of growth 1-like protein). | |||||
|
LAP2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 83) | NC score | 0.010558 (rank : 123) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 499 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96RT1, Q86W38, Q9NR18, Q9NW48, Q9ULJ5 | Gene names | ERBB2IP, ERBIN, KIAA1225, LAP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LAP2 (Erbb2-interacting protein) (Erbin) (Densin-180-like protein). | |||||
|
NEK11_HUMAN
|
||||||
| θ value | 4.03905 (rank : 84) | NC score | -0.003086 (rank : 144) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 1043 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NG66, Q5JPC0, Q8NG65, Q8TBY1, Q9H5F4 | Gene names | NEK11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase Nek11 (EC 2.7.11.1) (NimA-related protein kinase 11) (Never in mitosis A-related kinase 11). | |||||
|
PKP4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 85) | NC score | 0.019856 (rank : 90) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99569 | Gene names | PKP4 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-4 (p0071). | |||||
|
RAI14_MOUSE
|
||||||
| θ value | 4.03905 (rank : 86) | NC score | 0.005231 (rank : 136) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 1505 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9EP71, Q3URT3, Q6ZPT6 | Gene names | Rai14, Kiaa1334, Norpeg | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankycorbin (Ankyrin repeat and coiled-coil structure-containing protein) (Retinoic acid-induced protein 14) (Novel retinal pigment epithelial cell protein) (p125). | |||||
|
DC12_HUMAN
|
||||||
| θ value | 5.27518 (rank : 87) | NC score | 0.013352 (rank : 107) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96FZ2, Q96G34, Q9NRP3 | Gene names | C3orf37 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UPF0361 protein DC12. | |||||
|
EDEM1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 88) | NC score | 0.014039 (rank : 102) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q92611 | Gene names | EDEM1, EDEM, KIAA0212 | |||
|
Domain Architecture |

|
|||||
| Description | ER degradation-enhancing alpha-mannosidase-like 1. | |||||
|
EDEM1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 89) | NC score | 0.011723 (rank : 114) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q925U4 | Gene names | Edem1, Edem | |||
|
Domain Architecture |

|
|||||
| Description | ER degradation-enhancing alpha-mannosidase-like 1. | |||||
|
HTSF1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 90) | NC score | 0.013772 (rank : 104) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8BGC0, Q1WWK0, Q9CT41, Q9DAU3 | Gene names | Htatsf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIV Tat-specific factor 1 homolog. | |||||
|
LAP2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 91) | NC score | 0.009059 (rank : 129) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 482 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q80TH2, Q8BQ14, Q8CE41, Q8K171, Q99JU3, Q9JI47 | Gene names | Erbb2ip, Erbin, Kiaa1225, Lap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LAP2 (Erbb2-interacting protein) (Erbin) (Densin-180-like protein). | |||||
|
PDYN_HUMAN
|
||||||
| θ value | 5.27518 (rank : 92) | NC score | 0.013773 (rank : 103) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P01213 | Gene names | PDYN | |||
|
Domain Architecture |
|
|||||
| Description | Beta-neoendorphin-dynorphin precursor (Proenkephalin B) (Preprodynorphin) [Contains: Beta-neoendorphin; Dynorphin; Leu- enkephalin; Rimorphin; Leumorphin]. | |||||
|
R3HD1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 93) | NC score | 0.011057 (rank : 118) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15032, Q8IW32 | Gene names | R3HDM1, KIAA0029, R3HDM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | R3H domain-containing protein 1. | |||||
|
ANR12_HUMAN
|
||||||
| θ value | 6.88961 (rank : 94) | NC score | 0.014647 (rank : 101) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 1081 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q6UB98, O94951, Q658K1, Q6QMF7, Q9H231, Q9H784 | Gene names | ANKRD12, ANCO2, KIAA0874 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 12 (Ankyrin repeat-containing cofactor 2) (GAC-1 protein). | |||||
|
BNC1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 95) | NC score | 0.005648 (rank : 132) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q01954, Q15840 | Gene names | BNC1, BNC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein basonuclin-1. | |||||
|
CJ118_MOUSE
|
||||||
| θ value | 6.88961 (rank : 96) | NC score | 0.011344 (rank : 117) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 1132 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8C9S4, Q6P3D6, Q8CJC0 | Gene names | Otg1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf118 homolog (Oocyte-testis gene 1 protein). | |||||
|
CSF1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 97) | NC score | 0.010039 (rank : 126) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P09603, Q13130, Q14086, Q14806, Q9UQR8 | Gene names | CSF1 | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage colony-stimulating factor 1 precursor (CSF-1) (MCSF) (M- CSF) (Lanimostim). | |||||
|
MINT_MOUSE
|
||||||
| θ value | 6.88961 (rank : 98) | NC score | 0.021903 (rank : 87) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
PLAL1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 99) | NC score | -0.001920 (rank : 142) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 728 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UM63 | Gene names | PLAGL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein PLAGL1 (Pleiomorphic adenoma-like protein 1). | |||||
|
ZDHC8_MOUSE
|
||||||
| θ value | 6.88961 (rank : 100) | NC score | 0.003996 (rank : 137) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5Y5T5, Q7TNF7 | Gene names | Zdhhc8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable palmitoyltransferase ZDHHC8 (EC 2.3.1.-) (Zinc finger DHHC domain-containing protein 8) (DHHC-8). | |||||
|
A4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 101) | NC score | 0.010721 (rank : 121) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P12023, P97487, P97942, Q99K32 | Gene names | App | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 protein precursor (APP) (ABPP) (Alzheimer disease amyloid protein homolog) (Amyloidogenic glycoprotein) (AG) [Contains: Soluble APP-alpha (S-APP-alpha); Soluble APP-beta (S-APP-beta); C99 (APP-C99); Beta-amyloid protein 42 (Beta-APP42); Beta-amyloid protein 40 (Beta-APP40); C83; P3(42); P3(40); Gamma-CTF(59) (Gamma-secretase C-terminal fragment 59) (Amyloid intracellular domain 59) (AID(59)) (APP-C59); Gamma-CTF(57) (Gamma-secretase C-terminal fragment 57) (Amyloid intracellular domain 57) (AID(57)) (APP-C57); Gamma-CTF(50) (Gamma-secretase C-terminal fragment 50) (Amyloid intracellular domain 50) (AID(50)); C31]. | |||||
|
AFF1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 102) | NC score | 0.010757 (rank : 120) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P51825 | Gene names | AFF1, AF4, FEL, MLLT2 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4) (Protein FEL). | |||||
|
ANKS6_HUMAN
|
||||||
| θ value | 8.99809 (rank : 103) | NC score | 0.002667 (rank : 139) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 411 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q68DC2, Q5VSL0, Q5VSL2, Q5VSL3, Q5VSL4, Q68DB8, Q6P2R2, Q8N9L6, Q96D62 | Gene names | ANKS6, ANKRD14, SAMD6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat and SAM domain-containing protein 6 (Sterile alpha motif domain-containing protein 6) (Ankyrin repeat domain-containing protein 14). | |||||
|
CCD21_MOUSE
|
||||||
| θ value | 8.99809 (rank : 104) | NC score | 0.010096 (rank : 125) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8BMK0, Q8BUF1, Q8K0E6 | Gene names | Ccdc21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 21. | |||||
|
CF146_MOUSE
|
||||||
| θ value | 8.99809 (rank : 105) | NC score | 0.010296 (rank : 124) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9D9W6 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf146 homolog. | |||||
|
CYTSA_MOUSE
|
||||||
| θ value | 8.99809 (rank : 106) | NC score | 0.005553 (rank : 133) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 1104 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q2KN98, Q8CHF9 | Gene names | Cytsa, Kiaa0376 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytospin-A. | |||||
|
ETFD_MOUSE
|
||||||
| θ value | 8.99809 (rank : 107) | NC score | 0.021930 (rank : 86) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q921G7, Q3U7K2, Q8BK82, Q9DCT9 | Gene names | Etfdh | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Electron transfer flavoprotein-ubiquinone oxidoreductase, mitochondrial precursor (EC 1.5.5.1) (ETF-QO) (ETF-ubiquinone oxidoreductase) (ETF dehydrogenase) (Electron-transferring- flavoprotein dehydrogenase). | |||||
|
FA21C_HUMAN
|
||||||
| θ value | 8.99809 (rank : 108) | NC score | 0.010763 (rank : 119) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y4E1, Q5SQU4, Q5SQU5, Q7L521, Q9UG79 | Gene names | FAM21C, KIAA0592 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM21C. | |||||
|
FBX46_HUMAN
|
||||||
| θ value | 8.99809 (rank : 109) | NC score | 0.006803 (rank : 131) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6PJ61 | Gene names | FBXO46, FBX46, FBXO34L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 46 (F-box only protein 34-like). | |||||
|
KI18A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 110) | NC score | 0.002262 (rank : 140) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q91WD7, Q3TP77, Q8BLL1 | Gene names | Kif18a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 18A. | |||||
|
PP2CG_MOUSE
|
||||||
| θ value | 8.99809 (rank : 111) | NC score | 0.005433 (rank : 135) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q61074 | Gene names | Ppm1g, Fin13, Ppm1c | |||
|
Domain Architecture |

|
|||||
| Description | Protein phosphatase 2C isoform gamma (EC 3.1.3.16) (PP2C-gamma) (Protein phosphatase magnesium-dependent 1 gamma) (Protein phosphatase 1C) (Fibroblast growth factor-inducible protein 13) (FIN13). | |||||
|
RHG06_HUMAN
|
||||||
| θ value | 8.99809 (rank : 112) | NC score | 0.003540 (rank : 138) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O43182, O43437, Q9P1B3, Q9UK81, Q9UK82 | Gene names | ARHGAP6, RHOGAP6 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 6 (Rho-type GTPase-activating protein RhoGAPX-1). | |||||
|
SMG7_HUMAN
|
||||||
| θ value | 8.99809 (rank : 113) | NC score | 0.015202 (rank : 97) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q92540, Q5T1Q0, Q6PIE0, Q7Z7H9, Q8IXC1, Q8IXC2 | Gene names | SMG7, C1orf16, EST1C, KIAA0250 | |||
|
Domain Architecture |

|
|||||
| Description | Protein SMG7 (SMG-7 homolog) (EST1-like protein C). | |||||
|
TCF8_HUMAN
|
||||||
| θ value | 8.99809 (rank : 114) | NC score | -0.001934 (rank : 143) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 1057 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P37275, Q12924, Q13800 | Gene names | TCF8, AREB6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 8 (NIL-2-A zinc finger protein) (Negative regulator of IL2). | |||||
|
BAZ1B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 115) | NC score | 0.052566 (rank : 75) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 701 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9UIG0, O95039, O95247, O95277 | Gene names | BAZ1B, WBSC10, WBSCR9, WSTF | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein) (Williams syndrome transcription factor) (hWALP2). | |||||
|
BAZ1B_MOUSE
|
||||||
| θ value | θ > 10 (rank : 116) | NC score | 0.050446 (rank : 76) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9Z277, Q9CU68 | Gene names | Baz1b, Wbscr9 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein homolog) (WBRS9). | |||||
|
CENPC_MOUSE
|
||||||
| θ value | θ > 10 (rank : 117) | NC score | 0.053204 (rank : 74) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P49452 | Gene names | Cenpc1, Cenpc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein C 1 (CENP-C) (Centromere autoantigen C). | |||||
|
CXXC6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 118) | NC score | 0.053214 (rank : 73) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8NFU7, Q5VUP7, Q7Z6B6, Q8TCR1, Q9C0I7 | Gene names | CXXC6, KIAA1676, LCX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CXXC-type zinc finger protein 6 (Leukemia-associated protein with a CXXC domain). | |||||
|
DNMT1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 119) | NC score | 0.109064 (rank : 48) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P26358, Q9UHG5, Q9ULA2, Q9UMZ6 | Gene names | DNMT1, AIM, DNMT | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 1 (EC 2.1.1.37) (Dnmt1) (DNA methyltransferase HsaI) (DNA MTase HsaI) (MCMT) (M.HsaI). | |||||
|
DNMT1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 120) | NC score | 0.114921 (rank : 44) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P13864, P97413, Q80ZU3, Q9CSC6, Q9QXX6 | Gene names | Dnmt1, Dnmt, Met1, Uim | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 1 (EC 2.1.1.37) (Dnmt1) (DNA methyltransferase MmuI) (DNA MTase MmuI) (MCMT) (M.MmuI) (Met-1). | |||||
|
FBX37_HUMAN
|
||||||
| θ value | θ > 10 (rank : 121) | NC score | 0.059111 (rank : 68) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H469, Q49AL7, Q5JWA5 | Gene names | FBXL15, FBXO37 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 37 (F-box/LRR-repeat protein 15). | |||||
|
FBXL2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 122) | NC score | 0.080910 (rank : 61) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UKC9, Q9NVQ8, Q9UK27, Q9UKA5, Q9Y3Y9 | Gene names | FBXL2, FBL2, FBL3 | |||
|
Domain Architecture |
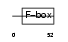
|
|||||
| Description | F-box/LRR-repeat protein 2 (F-box and leucine-rich repeat protein 2) (F-box protein FBL2/FBL3). | |||||
|
FBXL2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 123) | NC score | 0.083801 (rank : 60) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BH16, Q8BXM4 | Gene names | Fbxl2, Fbl2 | |||
|
Domain Architecture |
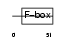
|
|||||
| Description | F-box/LRR-repeat protein 2 (F-box and leucine-rich repeat protein 2). | |||||
|
FBXL7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 124) | NC score | 0.099679 (rank : 50) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UJT9, O94926 | Gene names | FBXL7, FBL6, FBL7, KIAA0840 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 7 (F-box and leucine-rich repeat protein 7) (F-box protein FBL6/FBL7). | |||||
|
FXL12_HUMAN
|
||||||
| θ value | θ > 10 (rank : 125) | NC score | 0.062786 (rank : 66) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NXK8, Q9H5K4 | Gene names | FBXL12, FBL12 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 12 (F-box and leucine-rich repeat protein 12) (F-box protein FBL12). | |||||
|
FXL12_MOUSE
|
||||||
| θ value | θ > 10 (rank : 126) | NC score | 0.069462 (rank : 65) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9EPX5, Q3UVH7, Q8CDX0, Q9CY04, Q9QZN5 | Gene names | Fbxl12, Fbl12 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 12 (F-box and leucine-rich repeat protein 12) (F-box protein FBL12). | |||||
|
FXL13_HUMAN
|
||||||
| θ value | θ > 10 (rank : 127) | NC score | 0.078006 (rank : 62) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NEE6, Q6UVW7, Q6UVW8, Q75MN5, Q86UJ5, Q8N7Y4, Q8TCL2, Q8WUF9, Q8WUG0 | Gene names | FBXL13, FBL13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 13 (F-box and leucine-rich repeat protein 13). | |||||
|
FXL13_MOUSE
|
||||||
| θ value | θ > 10 (rank : 128) | NC score | 0.088377 (rank : 58) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8CDU4, Q8CDE9 | Gene names | Fbxl13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 13 (F-box and leucine-rich repeat protein 13). | |||||
|
FXL14_HUMAN
|
||||||
| θ value | θ > 10 (rank : 129) | NC score | 0.077572 (rank : 64) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N1E6 | Gene names | FBXL14, FBL14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 14 (F-box and leucine-rich repeat protein 14). | |||||
|
FXL14_MOUSE
|
||||||
| θ value | θ > 10 (rank : 130) | NC score | 0.077792 (rank : 63) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BID8, Q3U2Q9, Q8R5H7, Q8VDT7, Q922N5 | Gene names | Fbxl14, Fbl14, Ppa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 14 (F-box and leucine-rich repeat protein 14). | |||||
|
FXL16_HUMAN
|
||||||
| θ value | θ > 10 (rank : 131) | NC score | 0.089816 (rank : 55) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N461, Q2MHR2, Q96S14, Q9UJI0 | Gene names | FBXL16, C16orf22, FBL16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 16 (F-box and leucine-rich repeat protein 16). | |||||
|
FXL19_HUMAN
|
||||||
| θ value | θ > 10 (rank : 132) | NC score | 0.285503 (rank : 16) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6PCT2, Q8N789, Q9NT14 | Gene names | FBXL19, FBL19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 19 (F-box and leucine-rich repeat protein 19). | |||||
|
FXL19_MOUSE
|
||||||
| θ value | θ > 10 (rank : 133) | NC score | 0.280534 (rank : 17) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q6PB97, Q7TSH0, Q8BIB4 | Gene names | Fbxl19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 19 (F-box and leucine-rich repeat protein 19). | |||||
|
FXL20_HUMAN
|
||||||
| θ value | θ > 10 (rank : 134) | NC score | 0.090381 (rank : 54) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96IG2 | Gene names | FBXL20, FBL2 | |||
|
Domain Architecture |
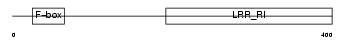
|
|||||
| Description | F-box/LRR-repeat protein 20 (F-box and leucine-rich repeat protein 20) (F-box/LRR-repeat protein 2-like). | |||||
|
FXL20_MOUSE
|
||||||
| θ value | θ > 10 (rank : 135) | NC score | 0.089740 (rank : 56) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9CZV8, Q8BZ95 | Gene names | Fbxl20, Fbl2 | |||
|
Domain Architecture |
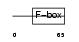
|
|||||
| Description | F-box/LRR-repeat protein 20 (F-box and leucine-rich repeat protein 20) (F-box/LRR-repeat protein 2-like). | |||||
|
HRX_HUMAN
|
||||||
| θ value | θ > 10 (rank : 136) | NC score | 0.098830 (rank : 53) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
ING5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 137) | NC score | 0.056632 (rank : 69) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8WYH8, Q9BS30 | Gene names | ING5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 5 (p28ING5). | |||||
|
ING5_MOUSE
|
||||||
| θ value | θ > 10 (rank : 138) | NC score | 0.055706 (rank : 70) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9D8Y8, Q3UL57, Q6P292, Q9CV64, Q9D9V8 | Gene names | Ing5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 5. | |||||
|
INGX_HUMAN
|
||||||
| θ value | θ > 10 (rank : 139) | NC score | 0.054249 (rank : 72) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9P0U5 | Gene names | INGX, ING2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein, X-linked (Inhibitor of growth protein 2) (ING1-like tumor suppressor protein). | |||||
|
LRC29_HUMAN
|
||||||
| θ value | θ > 10 (rank : 140) | NC score | 0.084186 (rank : 59) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8WV35, Q9UKA0 | Gene names | LRRC29, FBL9, FBXL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 29 (F-box/LRR-repeat protein 9) (F-box and leucine-rich repeat protein 9) (F-box protein FBL9). | |||||
|
MBD1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 141) | NC score | 0.118306 (rank : 43) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UIS9, O15248, O95241, Q7Z7B5, Q8N4W4, Q9UNZ6, Q9UNZ7, Q9UNZ8, Q9UNZ9 | Gene names | MBD1, PCM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 1 (Methyl-CpG-binding protein MBD1) (Protein containing methyl-CpG-binding domain 1). | |||||
|
MBD1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 142) | NC score | 0.112765 (rank : 47) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Z2E2, Q792D6, Q8CCL9, Q9DC19 | Gene names | Mbd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 1 (Methyl-CpG-binding protein MBD1). | |||||
|
SKP2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 143) | NC score | 0.099270 (rank : 51) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13309, Q8TDZ0, Q8TDZ1, Q9BV69 | Gene names | SKP2, FBXL1 | |||
|
Domain Architecture |

|
|||||
| Description | S-phase kinase-associated protein 2 (F-box protein Skp2) (Cyclin A/CDK2-associated protein p45) (p45skp2) (F-box/LRR-repeat protein 1). | |||||
|
SKP2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 144) | NC score | 0.088526 (rank : 57) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Z0Z3, Q8BSG7, Q8C8Y9, Q8CB22, Q99J53 | Gene names | Skp2 | |||
|
Domain Architecture |

|
|||||
| Description | S-phase kinase-associated protein 2 (F-box protein Skp2) (Cyclin A/CDK2-associated protein p45) (F-box/WD-40 protein 1) (FWD1). | |||||
|
K1718_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 114 | |
| SwissProt Accessions | Q6ZMT4, Q6MZL8, Q9C0E5 | Gene names | KIAA1718 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1718. | |||||
|
K1718_MOUSE
|
||||||
| NC score | 0.991569 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q3UWM4, Q3UWN8, Q6ZPJ5, Q8C969, Q8C9E0, Q91VX8 | Gene names | Kiaa1718 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1718. | |||||
|
PHF8_MOUSE
|
||||||
| NC score | 0.969096 (rank : 3) | θ value | 2.48782e-175 (rank : 3) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q80TJ7, Q8BLX8, Q8BLY0, Q8BZ61, Q8CG26 | Gene names | Phf8, Kiaa1111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 8. | |||||
|
PHF8_HUMAN
|
||||||
| NC score | 0.966548 (rank : 4) | θ value | 2.32852e-173 (rank : 4) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9UPP1, Q5H9U5, Q5VUJ4, Q7Z6D4, Q9HAH2 | Gene names | PHF8, KIAA1111, ZNF422 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 8. | |||||
|
PHF2_HUMAN
|
||||||
| NC score | 0.947579 (rank : 5) | θ value | 2.85304e-163 (rank : 5) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O75151, Q8N3K2, Q9Y6N4 | Gene names | PHF2, KIAA0662 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
PHF2_MOUSE
|
||||||
| NC score | 0.945595 (rank : 6) | θ value | 4.11978e-162 (rank : 6) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9WTU0, Q80WA8 | Gene names | Phf2 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
JHD1A_MOUSE
|
||||||
| NC score | 0.773515 (rank : 7) | θ value | 4.23606e-66 (rank : 10) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P59997, Q3U1M5, Q3UR56, Q3V3Q1, Q69ZT4 | Gene names | Fbxl11, Jhdm1a, Kiaa1004 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1A (EC 1.14.11.-) (F-box/LRR-repeat protein 11) (F-box and leucine-rich repeat protein 11). | |||||
|
JHD1A_HUMAN
|
||||||
| NC score | 0.772707 (rank : 8) | θ value | 1.90147e-66 (rank : 9) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9Y2K7, Q49A21, Q4G0M3, Q69YY8, Q9BVH5, Q9H7H5, Q9UK66 | Gene names | FBXL11, FBL7, JHDM1A, KIAA1004 | |||
|
Domain Architecture |

|
|||||
| Description | JmjC domain-containing histone demethylation protein 1A (EC 1.14.11.-) (F-box/LRR-repeat protein 11) (F-box and leucine-rich repeat protein 11) (F-box protein FBL7) (F-box protein Lilina). | |||||
|
JHD1B_HUMAN
|
||||||
| NC score | 0.763773 (rank : 9) | θ value | 1.60597e-73 (rank : 7) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8NHM5, Q8NCI2, Q96HC7, Q96SL0, Q96T03, Q9NS96, Q9UF75 | Gene names | FBXL10, CXXC2, FBL10, JHDM1B, PCCX2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1B (EC 1.14.11.-) (F-box/LRR-repeat protein 10) (F-box and leucine-rich repeat protein 10) (F-box protein FBL10) (Protein JEMMA) (Jumonji domain-containing EMSY-interactor methyltransferase motif protein) (CXXC-type zinc finger protein 2) (Protein-containing CXXC domain 2). | |||||
|
JHD1B_MOUSE
|
||||||
| NC score | 0.763494 (rank : 10) | θ value | 2.73936e-73 (rank : 8) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6P1G2, Q3V396, Q6PFD0, Q6ZPE8, Q9CSF7, Q9QZN6 | Gene names | Fbxl10, Fbl10, Jhdm1b, Kiaa3014 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1B (EC 1.14.11.-) (F-box/LRR-repeat protein 10) (F-box and leucine-rich repeat protein 10) (F-box protein FBL10). | |||||
|
CXCC1_MOUSE
|
||||||
| NC score | 0.495770 (rank : 11) | θ value | 9.59137e-10 (rank : 14) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9CWW7 | Gene names | Cxxc1, Cgbp, Pccx1 | |||
|
Domain Architecture |

|
|||||
| Description | CpG-binding protein (PHD finger and CXXC domain-containing protein 1) (CXXC-type zinc finger protein 1). | |||||
|
CXCC1_HUMAN
|
||||||
| NC score | 0.495115 (rank : 12) | θ value | 4.30538e-10 (rank : 13) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9P0U4, Q8N2W4, Q96BC8, Q9P2V7 | Gene names | CXXC1, CGBP, PCCX1, PHF18 | |||
|
Domain Architecture |

|
|||||
| Description | CpG-binding protein (PHD finger and CXXC domain-containing protein 1) (CXXC-type zinc finger protein 1). | |||||
|
DIDO1_HUMAN
|
||||||
| NC score | 0.297907 (rank : 13) | θ value | 2.43908e-13 (rank : 11) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
CN130_MOUSE
|
||||||
| NC score | 0.295896 (rank : 14) | θ value | 0.00020696 (rank : 15) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8BU04, Q80UT9 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C14orf130 homolog. | |||||
|
CN130_HUMAN
|
||||||
| NC score | 0.293970 (rank : 15) | θ value | 0.00035302 (rank : 17) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8N806, Q86U21, Q86UA9, Q96BY0, Q9NVV6 | Gene names | C14orf130 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C14orf130. | |||||
|
FXL19_HUMAN
|
||||||
| NC score | 0.285503 (rank : 16) | θ value | θ > 10 (rank : 132) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6PCT2, Q8N789, Q9NT14 | Gene names | FBXL19, FBL19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 19 (F-box and leucine-rich repeat protein 19). | |||||
|
FXL19_MOUSE
|
||||||
| NC score | 0.280534 (rank : 17) | θ value | θ > 10 (rank : 133) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q6PB97, Q7TSH0, Q8BIB4 | Gene names | Fbxl19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 19 (F-box and leucine-rich repeat protein 19). | |||||
|
PTDSR_MOUSE
|
||||||
| NC score | 0.275465 (rank : 18) | θ value | 0.00175202 (rank : 19) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9ERI5, Q80TX1 | Gene names | Ptdsr, Kiaa0585 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein PTDSR (Phosphatidylserine receptor) (Apoptotic cell clearance receptor PtdSerR). | |||||
|
BPTF_HUMAN
|
||||||
| NC score | 0.273164 (rank : 19) | θ value | 5.08577e-11 (rank : 12) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 786 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q12830, Q6NX67, Q7Z7D6, Q9UIG2 | Gene names | FALZ, BPTF, FAC1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleosome remodeling factor subunit BPTF (Bromodomain and PHD finger- containing transcription factor) (Fetal Alzheimer antigen) (Fetal Alz- 50 clone 1 protein). | |||||
|
PTDSR_HUMAN
|
||||||
| NC score | 0.268190 (rank : 20) | θ value | 0.00298849 (rank : 20) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6NYC1, Q86VY0, Q8IUM5, Q9Y4E2 | Gene names | PTDSR, KIAA0585 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein PTDSR (Phosphatidylserine receptor). | |||||
|
TCF19_HUMAN
|
||||||
| NC score | 0.244325 (rank : 21) | θ value | 0.0431538 (rank : 35) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9Y242, Q13176, Q15967, Q5SQ89, Q5STD6, Q5STF5, Q9BUM2, Q9UBH7 | Gene names | TCF19, SC1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 19 (Transcription factor SC1). | |||||
|
PHF3_HUMAN
|
||||||
| NC score | 0.239056 (rank : 22) | θ value | 0.000602161 (rank : 18) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q92576, Q5T1T6, Q9NQ16, Q9UI45 | Gene names | PHF3, KIAA0244 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 3. | |||||
|
PHF13_MOUSE
|
||||||
| NC score | 0.217087 (rank : 23) | θ value | 0.0736092 (rank : 38) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8K2W6 | Gene names | Phf13 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 13. | |||||
|
PHF13_HUMAN
|
||||||
| NC score | 0.216418 (rank : 24) | θ value | 0.0736092 (rank : 37) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q86YI8, Q8N551, Q9UJP2 | Gene names | PHF13 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 13. | |||||
|
TAF3_MOUSE
|
||||||
| NC score | 0.201609 (rank : 25) | θ value | 0.00665767 (rank : 24) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 463 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q5HZG4, Q3U490, Q3UWX2, Q8BIU8, Q99JH4 | Gene names | Taf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
TAF3_HUMAN
|
||||||
| NC score | 0.193980 (rank : 26) | θ value | 0.00298849 (rank : 21) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 568 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q5VWG9, Q6GMS5, Q6P6B5, Q86VY6, Q9BQS9, Q9UFI8 | Gene names | TAF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
PHF20_HUMAN
|
||||||
| NC score | 0.181324 (rank : 27) | θ value | 0.0193708 (rank : 29) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9BVI0, Q5JWY9, Q9BWV4, Q9BXA3, Q9BZW3, Q9H421, Q9H4J6, Q9NZ22 | Gene names | PHF20, C20orf104, GLEA2, HCA58, NZF, TZP | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58) (Glioma-expressed antigen 2) (Transcription factor TZP) (Novel zinc finger protein). | |||||
|
PHF20_MOUSE
|
||||||
| NC score | 0.174246 (rank : 28) | θ value | 0.0193708 (rank : 30) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8BLG0, Q8BMA2, Q8BYR4, Q921N1 | Gene names | Phf20, Hca58 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58 homolog). | |||||
|
JAD1A_HUMAN
|
||||||
| NC score | 0.172414 (rank : 29) | θ value | 0.00020696 (rank : 16) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P29375 | Gene names | JARID1A, RBBP2, RBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1A (Retinoblastoma-binding protein 2) (RBBP-2). | |||||
|
ING1_HUMAN
|
||||||
| NC score | 0.158661 (rank : 30) | θ value | 0.125558 (rank : 40) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9UK53, O00532, O43658, Q9H007, Q9HD98, Q9HD99, Q9NS83, Q9P0U6, Q9UBC6, Q9UIJ1, Q9UIJ2, Q9UIJ3, Q9UIJ4, Q9UK52 | Gene names | ING1 | |||
|
Domain Architecture |

|
|||||
| Description | Inhibitor of growth protein 1. | |||||
|
ING1_MOUSE
|
||||||
| NC score | 0.156406 (rank : 31) | θ value | 0.125558 (rank : 41) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9QXV3, Q9QUP8, Q9QXV4, Q9QZX3 | Gene names | Ing1 | |||
|
Domain Architecture |

|
|||||
| Description | Inhibitor of growth protein 1. | |||||
|
MTF2_HUMAN
|
||||||
| NC score | 0.148187 (rank : 32) | θ value | 0.0431538 (rank : 33) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9Y483, Q9UES9, Q9UP40 | Gene names | MTF2 | |||
|
Domain Architecture |

|
|||||
| Description | Metal-response element-binding transcription factor 2 (Metal-response element DNA-binding protein M96) (Metal-regulatory transcription factor 2) (PCL2). | |||||
|
MTF2_MOUSE
|
||||||
| NC score | 0.147281 (rank : 33) | θ value | 0.0431538 (rank : 34) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q02395, Q569Z8, Q6PG89, Q8BGP9, Q8BSJ7 | Gene names | Mtf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metal-response element-binding transcription factor 2 (Zinc-regulated factor 1) (ZiRF1) (Metal-response element DNA-binding protein M96) (Metal-regulatory transcription factor 2) (PCL2). | |||||
|
ING3_HUMAN
|
||||||
| NC score | 0.146231 (rank : 34) | θ value | 0.0736092 (rank : 36) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9NXR8, O60394, Q6GMT3, Q7Z762, Q969G0, Q96DT4, Q9HC99, Q9P081 | Gene names | ING3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 3 (p47ING3 protein). | |||||
|
JAD1D_HUMAN
|
||||||
| NC score | 0.146006 (rank : 35) | θ value | 0.00390308 (rank : 22) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9BY66, Q92509, Q92809, Q9HCU1 | Gene names | SMCY, HY, HYA, JARID1D, KIAA0234 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1D (SmcY protein) (Histocompatibility Y antigen) (H-Y). | |||||
|
JAD1C_MOUSE
|
||||||
| NC score | 0.135272 (rank : 36) | θ value | 0.0148317 (rank : 27) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P41230, O54995, Q9CVI4, Q9D0C3, Q9QVR8, Q9R039 | Gene names | Smcx, Jarid1c, Xe169 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1C (SmcX protein) (Xe169 protein). | |||||
|
JAD1C_HUMAN
|
||||||
| NC score | 0.134528 (rank : 37) | θ value | 0.0148317 (rank : 26) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P41229 | Gene names | SMCX, DXS1272E, JARID1C, XE169 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1C (Protein SmcX) (Protein Xe169). | |||||
|
ING3_MOUSE
|
||||||
| NC score | 0.134206 (rank : 38) | θ value | 0.163984 (rank : 44) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8VEK6, Q99JS6, Q9ERB2 | Gene names | Ing3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 3 (p47ING3 protein). | |||||
|
MLL4_HUMAN
|
||||||
| NC score | 0.129289 (rank : 39) | θ value | 1.06291 (rank : 51) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9UMN6, O15022, O95836, Q96GP2, Q96IP3, Q9UK25, Q9Y668, Q9Y669 | Gene names | MLL4, HRX2, KIAA0304, MLL2, TRX2 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 4 (Trithorax homolog 2). | |||||
|
JAD1D_MOUSE
|
||||||
| NC score | 0.123206 (rank : 40) | θ value | 0.0961366 (rank : 39) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q62240, Q9QVR9, Q9R040 | Gene names | Smcy, Hya, Jarid1d | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1D (SmcY protein) (Histocompatibility Y antigen) (H-Y). | |||||
|
ING2_MOUSE
|
||||||
| NC score | 0.119471 (rank : 41) | θ value | 4.03905 (rank : 82) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9ESK4, Q80VI5, Q8BGU8 | Gene names | Ing2, Ing1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 2 (p33ING2) (Inhibitor of growth 1-like protein). | |||||
|
ING2_HUMAN
|
||||||
| NC score | 0.119166 (rank : 42) | θ value | 4.03905 (rank : 81) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9H160, O95698 | Gene names | ING2, ING1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 2 (p33ING2) (Inhibitor of growth 1-like protein) (ING1Lp) (p32). | |||||
|
MBD1_HUMAN
|
||||||
| NC score | 0.118306 (rank : 43) | θ value | θ > 10 (rank : 141) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UIS9, O15248, O95241, Q7Z7B5, Q8N4W4, Q9UNZ6, Q9UNZ7, Q9UNZ8, Q9UNZ9 | Gene names | MBD1, PCM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 1 (Methyl-CpG-binding protein MBD1) (Protein containing methyl-CpG-binding domain 1). | |||||
|
DNMT1_MOUSE
|
||||||
| NC score | 0.114921 (rank : 44) | θ value | θ > 10 (rank : 120) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P13864, P97413, Q80ZU3, Q9CSC6, Q9QXX6 | Gene names | Dnmt1, Dnmt, Met1, Uim | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 1 (EC 2.1.1.37) (Dnmt1) (DNA methyltransferase MmuI) (DNA MTase MmuI) (MCMT) (M.MmuI) (Met-1). | |||||
|
PHF1_HUMAN
|
||||||
| NC score | 0.113646 (rank : 45) | θ value | 0.125558 (rank : 42) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O43189, O60929, Q96KM7 | Gene names | PHF1 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 1 (Protein PHF1). | |||||
|
PHF1_MOUSE
|
||||||
| NC score | 0.113478 (rank : 46) | θ value | 0.125558 (rank : 43) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9Z1B8, O54808 | Gene names | Phf1, Tctex-3, Tctex3 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 1 (Protein PHF1) (T-complex testis-expressed 3) (Polycomblike 1) (mPCl1). | |||||
|
MBD1_MOUSE
|
||||||
| NC score | 0.112765 (rank : 47) | θ value | θ > 10 (rank : 142) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Z2E2, Q792D6, Q8CCL9, Q9DC19 | Gene names | Mbd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 1 (Methyl-CpG-binding protein MBD1). | |||||
|
DNMT1_HUMAN
|
||||||
| NC score | 0.109064 (rank : 48) | θ value | θ > 10 (rank : 119) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P26358, Q9UHG5, Q9ULA2, Q9UMZ6 | Gene names | DNMT1, AIM, DNMT | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 1 (EC 2.1.1.37) (Dnmt1) (DNA methyltransferase HsaI) (DNA MTase HsaI) (MCMT) (M.HsaI). | |||||
|
ING4_HUMAN
|
||||||
| NC score | 0.099962 (rank : 49) | θ value | 1.81305 (rank : 55) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UNL4, Q96E15, Q9H3J0 | Gene names | ING4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 4 (p29ING4). | |||||
|
FBXL7_HUMAN
|
||||||
| NC score | 0.099679 (rank : 50) | θ value | θ > 10 (rank : 124) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UJT9, O94926 | Gene names | FBXL7, FBL6, FBL7, KIAA0840 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 7 (F-box and leucine-rich repeat protein 7) (F-box protein FBL6/FBL7). | |||||
|
SKP2_HUMAN
|
||||||
| NC score | 0.099270 (rank : 51) | θ value | θ > 10 (rank : 143) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13309, Q8TDZ0, Q8TDZ1, Q9BV69 | Gene names | SKP2, FBXL1 | |||
|
Domain Architecture |

|
|||||
| Description | S-phase kinase-associated protein 2 (F-box protein Skp2) (Cyclin A/CDK2-associated protein p45) (p45skp2) (F-box/LRR-repeat protein 1). | |||||
|
ING4_MOUSE
|
||||||
| NC score | 0.099049 (rank : 52) | θ value | 1.81305 (rank : 56) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8C0D7, Q8C1S7, Q8K3Q5, Q8K3Q6, Q8K3Q7, Q9D7F9 | Gene names | Ing4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 4 (p29ING4). | |||||
|
HRX_HUMAN
|
||||||
| NC score | 0.098830 (rank : 53) | θ value | θ > 10 (rank : 136) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
FXL20_HUMAN
|
||||||
| NC score | 0.090381 (rank : 54) | θ value | θ > 10 (rank : 134) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96IG2 | Gene names | FBXL20, FBL2 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 20 (F-box and leucine-rich repeat protein 20) (F-box/LRR-repeat protein 2-like). | |||||
|
FXL16_HUMAN
|
||||||
| NC score | 0.089816 (rank : 55) | θ value | θ > 10 (rank : 131) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N461, Q2MHR2, Q96S14, Q9UJI0 | Gene names | FBXL16, C16orf22, FBL16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 16 (F-box and leucine-rich repeat protein 16). | |||||
|
FXL20_MOUSE
|
||||||
| NC score | 0.089740 (rank : 56) | θ value | θ > 10 (rank : 135) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9CZV8, Q8BZ95 | Gene names | Fbxl20, Fbl2 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 20 (F-box and leucine-rich repeat protein 20) (F-box/LRR-repeat protein 2-like). | |||||
|
SKP2_MOUSE
|
||||||
| NC score | 0.088526 (rank : 57) | θ value | θ > 10 (rank : 144) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Z0Z3, Q8BSG7, Q8C8Y9, Q8CB22, Q99J53 | Gene names | Skp2 | |||
|
Domain Architecture |

|
|||||
| Description | S-phase kinase-associated protein 2 (F-box protein Skp2) (Cyclin A/CDK2-associated protein p45) (F-box/WD-40 protein 1) (FWD1). | |||||
|
FXL13_MOUSE
|
||||||
| NC score | 0.088377 (rank : 58) | θ value | θ > 10 (rank : 128) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8CDU4, Q8CDE9 | Gene names | Fbxl13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 13 (F-box and leucine-rich repeat protein 13). | |||||
|
LRC29_HUMAN
|
||||||
| NC score | 0.084186 (rank : 59) | θ value | θ > 10 (rank : 140) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8WV35, Q9UKA0 | Gene names | LRRC29, FBL9, FBXL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 29 (F-box/LRR-repeat protein 9) (F-box and leucine-rich repeat protein 9) (F-box protein FBL9). | |||||
|
FBXL2_MOUSE
|
||||||
| NC score | 0.083801 (rank : 60) | θ value | θ > 10 (rank : 123) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BH16, Q8BXM4 | Gene names | Fbxl2, Fbl2 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 2 (F-box and leucine-rich repeat protein 2). | |||||
|
FBXL2_HUMAN
|
||||||
| NC score | 0.080910 (rank : 61) | θ value | θ > 10 (rank : 122) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UKC9, Q9NVQ8, Q9UK27, Q9UKA5, Q9Y3Y9 | Gene names | FBXL2, FBL2, FBL3 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 2 (F-box and leucine-rich repeat protein 2) (F-box protein FBL2/FBL3). | |||||
|
FXL13_HUMAN
|
||||||
| NC score | 0.078006 (rank : 62) | θ value | θ > 10 (rank : 127) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NEE6, Q6UVW7, Q6UVW8, Q75MN5, Q86UJ5, Q8N7Y4, Q8TCL2, Q8WUF9, Q8WUG0 | Gene names | FBXL13, FBL13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 13 (F-box and leucine-rich repeat protein 13). | |||||
|
FXL14_MOUSE
|
||||||
| NC score | 0.077792 (rank : 63) | θ value | θ > 10 (rank : 130) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BID8, Q3U2Q9, Q8R5H7, Q8VDT7, Q922N5 | Gene names | Fbxl14, Fbl14, Ppa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 14 (F-box and leucine-rich repeat protein 14). | |||||
|
FXL14_HUMAN
|
||||||
| NC score | 0.077572 (rank : 64) | θ value | θ > 10 (rank : 129) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N1E6 | Gene names | FBXL14, FBL14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 14 (F-box and leucine-rich repeat protein 14). | |||||
|
FXL12_MOUSE
|
||||||
| NC score | 0.069462 (rank : 65) | θ value | θ > 10 (rank : 126) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9EPX5, Q3UVH7, Q8CDX0, Q9CY04, Q9QZN5 | Gene names | Fbxl12, Fbl12 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 12 (F-box and leucine-rich repeat protein 12) (F-box protein FBL12). | |||||
|
FXL12_HUMAN
|
||||||
| NC score | 0.062786 (rank : 66) | θ value | θ > 10 (rank : 125) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NXK8, Q9H5K4 | Gene names | FBXL12, FBL12 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 12 (F-box and leucine-rich repeat protein 12) (F-box protein FBL12). | |||||
|
MLL2_HUMAN
|
||||||
| NC score | 0.061473 (rank : 67) | θ value | 2.36792 (rank : 63) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
FBX37_HUMAN
|
||||||
| NC score | 0.059111 (rank : 68) | θ value | θ > 10 (rank : 121) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H469, Q49AL7, Q5JWA5 | Gene names | FBXL15, FBXO37 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 37 (F-box/LRR-repeat protein 15). | |||||
|
ING5_HUMAN
|
||||||
| NC score | 0.056632 (rank : 69) | θ value | θ > 10 (rank : 137) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8WYH8, Q9BS30 | Gene names | ING5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 5 (p28ING5). | |||||
|
ING5_MOUSE
|
||||||
| NC score | 0.055706 (rank : 70) | θ value | θ > 10 (rank : 138) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9D8Y8, Q3UL57, Q6P292, Q9CV64, Q9D9V8 | Gene names | Ing5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 5. | |||||
|
ZRAB2_HUMAN
|
||||||
| NC score | 0.054409 (rank : 71) | θ value | 0.813845 (rank : 48) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O95218, Q9UP63 | Gene names | ZRANB2, ZIS, ZNF265 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger Ran-binding domain-containing protein 2 Zinc finger protein 265 (Zinc finger, splicing). | |||||
|
INGX_HUMAN
|
||||||
| NC score | 0.054249 (rank : 72) | θ value | θ > 10 (rank : 139) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9P0U5 | Gene names | INGX, ING2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein, X-linked (Inhibitor of growth protein 2) (ING1-like tumor suppressor protein). | |||||
|
CXXC6_HUMAN
|
||||||
| NC score | 0.053214 (rank : 73) | θ value | θ > 10 (rank : 118) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8NFU7, Q5VUP7, Q7Z6B6, Q8TCR1, Q9C0I7 | Gene names | CXXC6, KIAA1676, LCX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CXXC-type zinc finger protein 6 (Leukemia-associated protein with a CXXC domain). | |||||
|
CENPC_MOUSE
|
||||||
| NC score | 0.053204 (rank : 74) | θ value | θ > 10 (rank : 117) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P49452 | Gene names | Cenpc1, Cenpc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein C 1 (CENP-C) (Centromere autoantigen C). | |||||
|
BAZ1B_HUMAN
|
||||||
| NC score | 0.052566 (rank : 75) | θ value | θ > 10 (rank : 115) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 701 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9UIG0, O95039, O95247, O95277 | Gene names | BAZ1B, WBSC10, WBSCR9, WSTF | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein) (Williams syndrome transcription factor) (hWALP2). | |||||
|
BAZ1B_MOUSE
|
||||||
| NC score | 0.050446 (rank : 76) | θ value | θ > 10 (rank : 116) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9Z277, Q9CU68 | Gene names | Baz1b, Wbscr9 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein homolog) (WBRS9). | |||||
|
MYST3_HUMAN
|
||||||
| NC score | 0.050043 (rank : 77) | θ value | 1.81305 (rank : 57) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q92794 | Gene names | MYST3, MOZ, RUNXBP2, ZNF220 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Runt-related transcription factor-binding protein 2) (Monocytic leukemia zinc finger protein) (Zinc finger protein 220). | |||||
|
DSPP_HUMAN
|
||||||
| NC score | 0.036070 (rank : 78) | θ value | 0.0193708 (rank : 28) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
SFRS4_HUMAN
|
||||||
| NC score | 0.031168 (rank : 79) | θ value | 0.00509761 (rank : 23) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q08170, Q5VXP1, Q9BUA4, Q9UEB5 | Gene names | SFRS4, SRP75 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 4 (Pre-mRNA-splicing factor SRP75) (SRP001LB). | |||||
|
PKP4_MOUSE
|
||||||
| NC score | 0.030796 (rank : 80) | θ value | 0.0113563 (rank : 25) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q68FH0, Q640N0, Q68G56, Q8BK47, Q8BVH1, Q9CRE3 | Gene names | Pkp4, Armrp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plakophilin-4 (Armadillo-related protein). | |||||
|
K1383_HUMAN
|
||||||
| NC score | 0.028502 (rank : 81) | θ value | 2.36792 (rank : 62) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9P2G4, Q58EZ9, Q5VV83 | Gene names | KIAA1383 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1383. | |||||
|
ETFD_HUMAN
|
||||||
| NC score | 0.027774 (rank : 82) | θ value | 4.03905 (rank : 77) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q16134, Q7Z347 | Gene names | ETFDH | |||
|
Domain Architecture |

|
|||||
| Description | Electron transfer flavoprotein-ubiquinone oxidoreductase, mitochondrial precursor (EC 1.5.5.1) (ETF-QO) (ETF-ubiquinone oxidoreductase) (ETF dehydrogenase) (Electron-transferring- flavoprotein dehydrogenase). | |||||
|
BRWD1_HUMAN
|
||||||
| NC score | 0.024142 (rank : 83) | θ value | 0.0252991 (rank : 31) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 448 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NSI6, O43721, Q5R2V0, Q5R2V1, Q8TCV3, Q96QG9, Q96QH0, Q9NUK1 | Gene names | BRWD1, C21orf107, WDR9 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain and WD repeat domain-containing protein 1 (WD repeat protein 9). | |||||
|
LMO7_HUMAN
|
||||||
| NC score | 0.024029 (rank : 84) | θ value | 1.38821 (rank : 53) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8WWI1, O15462, O95346, Q9UKC1, Q9UQM5, Q9Y6A7 | Gene names | LMO7, FBX20, FBXO20, KIAA0858 | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain only protein 7 (LOMP) (F-box only protein 20). | |||||
|
ZF106_HUMAN
|
||||||
| NC score | 0.022618 (rank : 85) | θ value | 0.813845 (rank : 47) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 470 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9H2Y7, Q6NSD9, Q6PEK1, Q86T43, Q86T45, Q86T50, Q86T58, Q86TA9, Q96M37, Q9H7B8 | Gene names | ZFP106, ZNF474 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 106 homolog (Zfp-106) (Zinc finger protein 474). | |||||
|
ETFD_MOUSE
|
||||||
| NC score | 0.021930 (rank : 86) | θ value | 8.99809 (rank : 107) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q921G7, Q3U7K2, Q8BK82, Q9DCT9 | Gene names | Etfdh | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Electron transfer flavoprotein-ubiquinone oxidoreductase, mitochondrial precursor (EC 1.5.5.1) (ETF-QO) (ETF-ubiquinone oxidoreductase) (ETF dehydrogenase) (Electron-transferring- flavoprotein dehydrogenase). | |||||
|
MINT_MOUSE
|
||||||
| NC score | 0.021903 (rank : 87) | θ value | 6.88961 (rank : 98) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
NOP14_MOUSE
|
||||||
| NC score | 0.021875 (rank : 88) | θ value | 0.813845 (rank : 46) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 324 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8R3N1 | Gene names | Nop14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable nucleolar complex protein 14. | |||||
|
HPS5_MOUSE
|
||||||
| NC score | 0.020993 (rank : 89) | θ value | 1.06291 (rank : 50) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P59438 | Gene names | Hps5, Ru2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hermansky-Pudlak syndrome 5 protein homolog (Ruby-eye protein 2) (Ru2). | |||||
|
PKP4_HUMAN
|
||||||
| NC score | 0.019856 (rank : 90) | θ value | 4.03905 (rank : 85) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99569 | Gene names | PKP4 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-4 (p0071). | |||||
|
SMG7_MOUSE
|
||||||
| NC score | 0.019557 (rank : 91) | θ value | 2.36792 (rank : 66) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q5RJH6, Q63ZW5, Q6ZQF3 | Gene names | Smg7, Est1c, Kiaa0250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein SMG7 (SMG-7 homolog) (EST1-like protein C). | |||||
|
RXINP_MOUSE
|
||||||
| NC score | 0.017941 (rank : 92) | θ value | 2.36792 (rank : 65) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5U5Q9, Q60811 | Gene names | Rxrip110, Rip110 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoid X receptor-interacting protein 110. | |||||
|
MPP8_HUMAN
|
||||||
| NC score | 0.017737 (rank : 93) | θ value | 0.47712 (rank : 45) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 948 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q99549, Q86TK3, Q9BTP1 | Gene names | MPHOSPH8, MPP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | M-phase phosphoprotein 8. | |||||
|
EYA4_HUMAN
|
||||||
| NC score | 0.016475 (rank : 94) | θ value | 3.0926 (rank : 69) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O95677, O95464, O95679, Q8IW39, Q9NTR7 | Gene names | EYA4 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 4 (EC 3.1.3.48). | |||||
|
EYA4_MOUSE
|
||||||
| NC score | 0.016418 (rank : 95) | θ value | 3.0926 (rank : 70) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Z191 | Gene names | Eya4 | |||
|
Domain Architecture |

|
|||||
| Description | Eyes absent homolog 4 (EC 3.1.3.48). | |||||
|
CA103_MOUSE
|
||||||
| NC score | 0.016125 (rank : 96) | θ value | 3.0926 (rank : 67) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CDD9, Q8C893 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C1orf103 homolog. | |||||
|
SMG7_HUMAN
|
||||||
| NC score | 0.015202 (rank : 97) | θ value | 8.99809 (rank : 113) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q92540, Q5T1Q0, Q6PIE0, Q7Z7H9, Q8IXC1, Q8IXC2 | Gene names | SMG7, C1orf16, EST1C, KIAA0250 | |||
|
Domain Architecture |

|
|||||
| Description | Protein SMG7 (SMG-7 homolog) (EST1-like protein C). | |||||
|
PININ_HUMAN
|
||||||
| NC score | 0.014997 (rank : 98) | θ value | 2.36792 (rank : 64) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 753 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9H307, O60899, Q53EM7, Q6P5X4, Q7KYL1, Q99738, Q9UHZ9, Q9UQR9 | Gene names | PNN, DRS, MEMA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pinin (140 kDa nuclear and cell adhesion-related phosphoprotein) (Domain-rich serine protein) (DRS-protein) (DRSP) (Melanoma metastasis clone A protein) (Desmosome-associated protein) (SR-like protein) (Nuclear protein SDK3). | |||||
|
APLP1_MOUSE
|
||||||
| NC score | 0.014860 (rank : 99) | θ value | 4.03905 (rank : 73) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q03157, Q8VC38 | Gene names | Aplp1 | |||
|
Domain Architecture |

|
|||||
| Description | Amyloid-like protein 1 precursor (APLP) (APLP-1) [Contains: C30]. | |||||
|
GOGA5_MOUSE
|
||||||
| NC score | 0.014805 (rank : 100) | θ value | 0.0431538 (rank : 32) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 1434 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9QYE6, O88317, Q3TGE7, Q3U6S5, Q3UUF9 | Gene names | Golga5, Retii | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 5 (Golgin-84) (Sumiko protein) (Ret-II protein). | |||||
|
ANR12_HUMAN
|
||||||
| NC score | 0.014647 (rank : 101) | θ value | 6.88961 (rank : 94) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 1081 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q6UB98, O94951, Q658K1, Q6QMF7, Q9H231, Q9H784 | Gene names | ANKRD12, ANCO2, KIAA0874 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 12 (Ankyrin repeat-containing cofactor 2) (GAC-1 protein). | |||||
|
EDEM1_HUMAN
|
||||||
| NC score | 0.014039 (rank : 102) | θ value | 5.27518 (rank : 88) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q92611 | Gene names | EDEM1, EDEM, KIAA0212 | |||
|
Domain Architecture |

|
|||||
| Description | ER degradation-enhancing alpha-mannosidase-like 1. | |||||
|
PDYN_HUMAN
|
||||||
| NC score | 0.013773 (rank : 103) | θ value | 5.27518 (rank : 92) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P01213 | Gene names | PDYN | |||
|
Domain Architecture |

|
|||||
| Description | Beta-neoendorphin-dynorphin precursor (Proenkephalin B) (Preprodynorphin) [Contains: Beta-neoendorphin; Dynorphin; Leu- enkephalin; Rimorphin; Leumorphin]. | |||||
|
HTSF1_MOUSE
|
||||||
| NC score | 0.013772 (rank : 104) | θ value | 5.27518 (rank : 90) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8BGC0, Q1WWK0, Q9CT41, Q9DAU3 | Gene names | Htatsf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIV Tat-specific factor 1 homolog. | |||||
|
CASC5_HUMAN
|
||||||
| NC score | 0.013669 (rank : 105) | θ value | 4.03905 (rank : 74) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 376 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8NG31, Q8NHE1, Q8WXA6, Q9HCK2, Q9NR92 | Gene names | CASC5, KIAA1570 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein CASC5 (Cancer susceptibility candidate gene 5 protein) (ALL1- fused gene from chromosome 15q14) (AF15q14) (D40/AF15q14 protein). | |||||
|
EXOSX_HUMAN
|
||||||
| NC score | 0.013626 (rank : 106) | θ value | 4.03905 (rank : 78) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q01780, Q15158 | Gene names | EXOSC10, PMSCL, PMSCL2 | |||
|
Domain Architecture |

|
|||||
| Description | Exosome component 10 (Polymyositis/scleroderma autoantigen 2) (Autoantigen PM/Scl 2) (Polymyositis/scleroderma autoantigen 100 kDa) (PM/Scl-100) (P100 polymyositis-scleroderma overlap syndrome- associated autoantigen). | |||||
|
DC12_HUMAN
|
||||||
| NC score | 0.013352 (rank : 107) | θ value | 5.27518 (rank : 87) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96FZ2, Q96G34, Q9NRP3 | Gene names | C3orf37 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UPF0361 protein DC12. | |||||
|
GP158_HUMAN
|
||||||
| NC score | 0.013146 (rank : 108) | θ value | 4.03905 (rank : 80) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5T848, Q6QR81, Q9ULT3 | Gene names | GPR158, KIAA1136 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 158 precursor. | |||||
|
WDR22_MOUSE
|
||||||
| NC score | 0.013101 (rank : 109) | θ value | 1.81305 (rank : 59) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80T85, Q80VT3, Q80ZW6, Q8BIP8, Q8BVK5, Q8K0Q6 | Gene names | Wdr22, Kiaa1824 | |||
|
Domain Architecture |

|
|||||
| Description | WD repeat protein 22. | |||||
|
DPOLZ_HUMAN
|
||||||
| NC score | 0.012848 (rank : 110) | θ value | 3.0926 (rank : 68) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O60673, O43214 | Gene names | REV3L, POLZ, REV3 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase zeta catalytic subunit (EC 2.7.7.7) (hREV3). | |||||
|
A4_HUMAN
|
||||||
| NC score | 0.012690 (rank : 111) | θ value | 4.03905 (rank : 72) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P05067, P09000, P78438, Q13764, Q13778, Q13793, Q16011, Q16014, Q16019, Q16020, Q9BT38, Q9UCA9, Q9UCB6, Q9UCC8, Q9UCD1, Q9UQ58 | Gene names | APP, A4, AD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 protein precursor (APP) (ABPP) (Alzheimer disease amyloid protein) (Cerebral vascular amyloid peptide) (CVAP) (Protease nexin-II) (PN-II) (APPI) (PreA4) [Contains: Soluble APP-alpha (S-APP- alpha); Soluble APP-beta (S-APP-beta); C99; Beta-amyloid protein 42 (Beta-APP42); Beta-amyloid protein 40 (Beta-APP40); C83; P3(42); P3(40); Gamma-CTF(59) (Gamma-secretase C-terminal fragment 59) (Amyloid intracellular domain 59) (AID(59)); Gamma-CTF(57) (Gamma- secretase C-terminal fragment 57) (Amyloid intracellular domain 57) (AID(57)); Gamma-CTF(50) (Gamma-secretase C-terminal fragment 50) (Amyloid intracellular domain 50) (AID(50)); C31]. | |||||
|
CEP55_MOUSE
|
||||||
| NC score | 0.012308 (rank : 112) | θ value | 2.36792 (rank : 61) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 693 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BT07, Q8C2J0, Q8K2I8, Q8R2Y4, Q9DBZ8 | Gene names | Cep55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome protein 55. | |||||
|
FANCM_MOUSE
|
||||||
| NC score | 0.012302 (rank : 113) | θ value | 4.03905 (rank : 79) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BGE5, Q69ZF5, Q8BKB7, Q8BUA8 | Gene names | Fancm, Kiaa1596 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fanconi anemia group M protein homolog (EC 3.6.1.-) (ATP-dependent RNA helicase FANCM) (Protein FACM). | |||||
|
EDEM1_MOUSE
|
||||||
| NC score | 0.011723 (rank : 114) | θ value | 5.27518 (rank : 89) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q925U4 | Gene names | Edem1, Edem | |||
|
Domain Architecture |

|
|||||
| Description | ER degradation-enhancing alpha-mannosidase-like 1. | |||||
|
OSBL8_HUMAN
|
||||||
| NC score | 0.011529 (rank : 115) | θ value | 3.0926 (rank : 71) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9BZF1, Q8WXP8, Q9P277 | Gene names | OSBPL8, KIAA1451, ORP8, OSBP10 | |||
|
Domain Architecture |

|
|||||
| Description | Oxysterol-binding protein-related protein 8 (OSBP-related protein 8) (ORP-8). | |||||
|
UTRO_HUMAN
|
||||||
| NC score | 0.011372 (rank : 116) | θ value | 1.81305 (rank : 58) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P46939 | Gene names | UTRN, DMDL | |||
|
Domain Architecture |

|
|||||
| Description | Utrophin (Dystrophin-related protein 1) (DRP1) (DRP). | |||||
|
CJ118_MOUSE
|
||||||
| NC score | 0.011344 (rank : 117) | θ value | 6.88961 (rank : 96) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 1132 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8C9S4, Q6P3D6, Q8CJC0 | Gene names | Otg1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf118 homolog (Oocyte-testis gene 1 protein). | |||||
|
R3HD1_HUMAN
|
||||||
| NC score | 0.011057 (rank : 118) | θ value | 5.27518 (rank : 93) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15032, Q8IW32 | Gene names | R3HDM1, KIAA0029, R3HDM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | R3H domain-containing protein 1. | |||||
|
FA21C_HUMAN
|
||||||
| NC score | 0.010763 (rank : 119) | θ value | 8.99809 (rank : 108) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y4E1, Q5SQU4, Q5SQU5, Q7L521, Q9UG79 | Gene names | FAM21C, KIAA0592 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM21C. | |||||
|
AFF1_HUMAN
|
||||||
| NC score | 0.010757 (rank : 120) | θ value | 8.99809 (rank : 102) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P51825 | Gene names | AFF1, AF4, FEL, MLLT2 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4) (Protein FEL). | |||||
|
A4_MOUSE
|
||||||
| NC score | 0.010721 (rank : 121) | θ value | 8.99809 (rank : 101) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P12023, P97487, P97942, Q99K32 | Gene names | App | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 protein precursor (APP) (ABPP) (Alzheimer disease amyloid protein homolog) (Amyloidogenic glycoprotein) (AG) [Contains: Soluble APP-alpha (S-APP-alpha); Soluble APP-beta (S-APP-beta); C99 (APP-C99); Beta-amyloid protein 42 (Beta-APP42); Beta-amyloid protein 40 (Beta-APP40); C83; P3(42); P3(40); Gamma-CTF(59) (Gamma-secretase C-terminal fragment 59) (Amyloid intracellular domain 59) (AID(59)) (APP-C59); Gamma-CTF(57) (Gamma-secretase C-terminal fragment 57) (Amyloid intracellular domain 57) (AID(57)) (APP-C57); Gamma-CTF(50) (Gamma-secretase C-terminal fragment 50) (Amyloid intracellular domain 50) (AID(50)); C31]. | |||||
|
CJ118_HUMAN
|
||||||
| NC score | 0.010560 (rank : 122) | θ value | 4.03905 (rank : 75) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 937 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q7Z3E2, Q2M2V6, Q6NS91, Q7RTP1, Q8N117, Q8N3G3, Q8N6C2, Q9NWA3 | Gene names | C10orf118 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf118 (CTCL tumor antigen HD-CL-01/L14-2). | |||||
|
LAP2_HUMAN
|
||||||
| NC score | 0.010558 (rank : 123) | θ value | 4.03905 (rank : 83) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 499 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96RT1, Q86W38, Q9NR18, Q9NW48, Q9ULJ5 | Gene names | ERBB2IP, ERBIN, KIAA1225, LAP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LAP2 (Erbb2-interacting protein) (Erbin) (Densin-180-like protein). | |||||
|
CF146_MOUSE
|
||||||
| NC score | 0.010296 (rank : 124) | θ value | 8.99809 (rank : 105) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9D9W6 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf146 homolog. | |||||
|
CCD21_MOUSE
|
||||||
| NC score | 0.010096 (rank : 125) | θ value | 8.99809 (rank : 104) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8BMK0, Q8BUF1, Q8K0E6 | Gene names | Ccdc21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 21. | |||||
|
CSF1_HUMAN
|
||||||
| NC score | 0.010039 (rank : 126) | θ value | 6.88961 (rank : 97) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P09603, Q13130, Q14086, Q14806, Q9UQR8 | Gene names | CSF1 | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage colony-stimulating factor 1 precursor (CSF-1) (MCSF) (M- CSF) (Lanimostim). | |||||
|
RAD50_MOUSE
|
||||||
| NC score | 0.010010 (rank : 127) | θ value | 1.38821 (rank : 54) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 1366 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P70388, Q6PEB0, Q8C2T7, Q9CU59 | Gene names | Rad50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (mRad50). | |||||
|
DDX46_MOUSE
|
||||||
| NC score | 0.009936 (rank : 128) | θ value | 1.06291 (rank : 49) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 358 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q569Z5, Q6ZQ42, Q8R0R6 | Gene names | Ddx46, Kiaa0801 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX46 (EC 3.6.1.-) (DEAD box protein 46). | |||||
|
LAP2_MOUSE
|
||||||
| NC score | 0.009059 (rank : 129) | θ value | 5.27518 (rank : 91) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 482 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q80TH2, Q8BQ14, Q8CE41, Q8K171, Q99JU3, Q9JI47 | Gene names | Erbb2ip, Erbin, Kiaa1225, Lap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LAP2 (Erbb2-interacting protein) (Erbin) (Densin-180-like protein). | |||||
|
DDX46_HUMAN
|
||||||
| NC score | 0.007897 (rank : 130) | θ value | 4.03905 (rank : 76) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7L014, O94894, Q96EI0, Q9Y658 | Gene names | DDX46, KIAA0801 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX46 (EC 3.6.1.-) (DEAD box protein 46) (PRP5 homolog). | |||||
|
FBX46_HUMAN
|
||||||
| NC score | 0.006803 (rank : 131) | θ value | 8.99809 (rank : 109) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6PJ61 | Gene names | FBXO46, FBX46, FBXO34L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 46 (F-box only protein 34-like). | |||||
|
BNC1_HUMAN
|
||||||
| NC score | 0.005648 (rank : 132) | θ value | 6.88961 (rank : 95) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q01954, Q15840 | Gene names | BNC1, BNC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein basonuclin-1. | |||||
|
CYTSA_MOUSE
|
||||||
| NC score | 0.005553 (rank : 133) | θ value | 8.99809 (rank : 106) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 1104 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q2KN98, Q8CHF9 | Gene names | Cytsa, Kiaa0376 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytospin-A. | |||||
|
MYH9_HUMAN
|
||||||
| NC score | 0.005485 (rank : 134) | θ value | 1.06291 (rank : 52) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P35579, O60805 | Gene names | MYH9 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
PP2CG_MOUSE
|
||||||
| NC score | 0.005433 (rank : 135) | θ value | 8.99809 (rank : 111) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q61074 | Gene names | Ppm1g, Fin13, Ppm1c | |||
|
Domain Architecture |

|
|||||
| Description | Protein phosphatase 2C isoform gamma (EC 3.1.3.16) (PP2C-gamma) (Protein phosphatase magnesium-dependent 1 gamma) (Protein phosphatase 1C) (Fibroblast growth factor-inducible protein 13) (FIN13). | |||||
|
RAI14_MOUSE
|
||||||
| NC score | 0.005231 (rank : 136) | θ value | 4.03905 (rank : 86) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 1505 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9EP71, Q3URT3, Q6ZPT6 | Gene names | Rai14, Kiaa1334, Norpeg | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankycorbin (Ankyrin repeat and coiled-coil structure-containing protein) (Retinoic acid-induced protein 14) (Novel retinal pigment epithelial cell protein) (p125). | |||||
|
ZDHC8_MOUSE
|
||||||
| NC score | 0.003996 (rank : 137) | θ value | 6.88961 (rank : 100) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5Y5T5, Q7TNF7 | Gene names | Zdhhc8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable palmitoyltransferase ZDHHC8 (EC 2.3.1.-) (Zinc finger DHHC domain-containing protein 8) (DHHC-8). | |||||
|
RHG06_HUMAN
|
||||||
| NC score | 0.003540 (rank : 138) | θ value | 8.99809 (rank : 112) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O43182, O43437, Q9P1B3, Q9UK81, Q9UK82 | Gene names | ARHGAP6, RHOGAP6 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 6 (Rho-type GTPase-activating protein RhoGAPX-1). | |||||
|
ANKS6_HUMAN
|
||||||
| NC score | 0.002667 (rank : 139) | θ value | 8.99809 (rank : 103) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 411 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q68DC2, Q5VSL0, Q5VSL2, Q5VSL3, Q5VSL4, Q68DB8, Q6P2R2, Q8N9L6, Q96D62 | Gene names | ANKS6, ANKRD14, SAMD6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat and SAM domain-containing protein 6 (Sterile alpha motif domain-containing protein 6) (Ankyrin repeat domain-containing protein 14). | |||||
|
KI18A_MOUSE
|
||||||
| NC score | 0.002262 (rank : 140) | θ value | 8.99809 (rank : 110) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q91WD7, Q3TP77, Q8BLL1 | Gene names | Kif18a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 18A. | |||||
|
BORIS_HUMAN
|
||||||
| NC score | -0.001354 (rank : 141) | θ value | 2.36792 (rank : 60) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 743 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8NI51, Q5JUG4, Q9BZ30, Q9NQJ3 | Gene names | CTCFL, BORIS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcriptional repressor CTCFL (CCCTC-binding factor) (Brother of the regulator of imprinted sites) (Zinc finger protein CTCF-T) (CTCF paralog). | |||||
|
PLAL1_HUMAN
|
||||||
| NC score | -0.001920 (rank : 142) | θ value | 6.88961 (rank : 99) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 728 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UM63 | Gene names | PLAGL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein PLAGL1 (Pleiomorphic adenoma-like protein 1). | |||||
|
TCF8_HUMAN
|
||||||
| NC score | -0.001934 (rank : 143) | θ value | 8.99809 (rank : 114) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 1057 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P37275, Q12924, Q13800 | Gene names | TCF8, AREB6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 8 (NIL-2-A zinc finger protein) (Negative regulator of IL2). | |||||
|
NEK11_HUMAN
|
||||||
| NC score | -0.003086 (rank : 144) | θ value | 4.03905 (rank : 84) | |||
| Query Neighborhood Hits | 114 | Target Neighborhood Hits | 1043 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NG66, Q5JPC0, Q8NG65, Q8TBY1, Q9H5F4 | Gene names | NEK11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase Nek11 (EC 2.7.11.1) (NimA-related protein kinase 11) (Never in mitosis A-related kinase 11). | |||||