Please be patient as the page loads
|
CXCC1_MOUSE
|
||||||
| SwissProt Accessions | Q9CWW7 | Gene names | Cxxc1, Cgbp, Pccx1 | |||
|
Domain Architecture |

|
|||||
| Description | CpG-binding protein (PHD finger and CXXC domain-containing protein 1) (CXXC-type zinc finger protein 1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
CXCC1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.992935 (rank : 2) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 111 | |
| SwissProt Accessions | Q9P0U4, Q8N2W4, Q96BC8, Q9P2V7 | Gene names | CXXC1, CGBP, PCCX1, PHF18 | |||
|
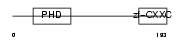
Domain Architecture |
|
|||||
| Description | CpG-binding protein (PHD finger and CXXC domain-containing protein 1) (CXXC-type zinc finger protein 1). | |||||
|
CXCC1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 124 | |
| SwissProt Accessions | Q9CWW7 | Gene names | Cxxc1, Cgbp, Pccx1 | |||
|
Domain Architecture |

|
|||||
| Description | CpG-binding protein (PHD finger and CXXC domain-containing protein 1) (CXXC-type zinc finger protein 1). | |||||
|
MBD1_MOUSE
|
||||||
| θ value | 2.69671e-12 (rank : 3) | NC score | 0.431709 (rank : 14) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Z2E2, Q792D6, Q8CCL9, Q9DC19 | Gene names | Mbd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 1 (Methyl-CpG-binding protein MBD1). | |||||
|
MBD1_HUMAN
|
||||||
| θ value | 7.84624e-12 (rank : 4) | NC score | 0.436752 (rank : 13) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9UIS9, O15248, O95241, Q7Z7B5, Q8N4W4, Q9UNZ6, Q9UNZ7, Q9UNZ8, Q9UNZ9 | Gene names | MBD1, PCM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 1 (Methyl-CpG-binding protein MBD1) (Protein containing methyl-CpG-binding domain 1). | |||||
|
PHF2_HUMAN
|
||||||
| θ value | 1.9326e-10 (rank : 5) | NC score | 0.497552 (rank : 5) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O75151, Q8N3K2, Q9Y6N4 | Gene names | PHF2, KIAA0662 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
PHF2_MOUSE
|
||||||
| θ value | 1.9326e-10 (rank : 6) | NC score | 0.498104 (rank : 4) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9WTU0, Q80WA8 | Gene names | Phf2 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
DIDO1_HUMAN
|
||||||
| θ value | 2.52405e-10 (rank : 7) | NC score | 0.320208 (rank : 29) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
K1718_MOUSE
|
||||||
| θ value | 3.29651e-10 (rank : 8) | NC score | 0.500364 (rank : 3) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q3UWM4, Q3UWN8, Q6ZPJ5, Q8C969, Q8C9E0, Q91VX8 | Gene names | Kiaa1718 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1718. | |||||
|
K1718_HUMAN
|
||||||
| θ value | 9.59137e-10 (rank : 9) | NC score | 0.495770 (rank : 6) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q6ZMT4, Q6MZL8, Q9C0E5 | Gene names | KIAA1718 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1718. | |||||
|
PHF8_HUMAN
|
||||||
| θ value | 1.25267e-09 (rank : 10) | NC score | 0.490834 (rank : 8) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9UPP1, Q5H9U5, Q5VUJ4, Q7Z6D4, Q9HAH2 | Gene names | PHF8, KIAA1111, ZNF422 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 8. | |||||
|
PHF8_MOUSE
|
||||||
| θ value | 2.79066e-09 (rank : 11) | NC score | 0.490890 (rank : 7) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q80TJ7, Q8BLX8, Q8BLY0, Q8BZ61, Q8CG26 | Gene names | Phf8, Kiaa1111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 8. | |||||
|
BPTF_HUMAN
|
||||||
| θ value | 4.76016e-09 (rank : 12) | NC score | 0.295613 (rank : 34) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 786 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q12830, Q6NX67, Q7Z7D6, Q9UIG2 | Gene names | FALZ, BPTF, FAC1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleosome remodeling factor subunit BPTF (Bromodomain and PHD finger- containing transcription factor) (Fetal Alzheimer antigen) (Fetal Alz- 50 clone 1 protein). | |||||
|
HRX_HUMAN
|
||||||
| θ value | 4.0297e-08 (rank : 13) | NC score | 0.280842 (rank : 38) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
PHF3_HUMAN
|
||||||
| θ value | 5.26297e-08 (rank : 14) | NC score | 0.330522 (rank : 26) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q92576, Q5T1T6, Q9NQ16, Q9UI45 | Gene names | PHF3, KIAA0244 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 3. | |||||
|
FXL19_HUMAN
|
||||||
| θ value | 2.61198e-07 (rank : 15) | NC score | 0.353533 (rank : 19) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q6PCT2, Q8N789, Q9NT14 | Gene names | FBXL19, FBL19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 19 (F-box and leucine-rich repeat protein 19). | |||||
|
FXL19_MOUSE
|
||||||
| θ value | 2.61198e-07 (rank : 16) | NC score | 0.348118 (rank : 20) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q6PB97, Q7TSH0, Q8BIB4 | Gene names | Fbxl19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 19 (F-box and leucine-rich repeat protein 19). | |||||
|
MLL4_HUMAN
|
||||||
| θ value | 2.61198e-07 (rank : 17) | NC score | 0.280906 (rank : 37) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9UMN6, O15022, O95836, Q96GP2, Q96IP3, Q9UK25, Q9Y668, Q9Y669 | Gene names | MLL4, HRX2, KIAA0304, MLL2, TRX2 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 4 (Trithorax homolog 2). | |||||
|
JHD1B_HUMAN
|
||||||
| θ value | 4.45536e-07 (rank : 18) | NC score | 0.446326 (rank : 12) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8NHM5, Q8NCI2, Q96HC7, Q96SL0, Q96T03, Q9NS96, Q9UF75 | Gene names | FBXL10, CXXC2, FBL10, JHDM1B, PCCX2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1B (EC 1.14.11.-) (F-box/LRR-repeat protein 10) (F-box and leucine-rich repeat protein 10) (F-box protein FBL10) (Protein JEMMA) (Jumonji domain-containing EMSY-interactor methyltransferase motif protein) (CXXC-type zinc finger protein 2) (Protein-containing CXXC domain 2). | |||||
|
JHD1B_MOUSE
|
||||||
| θ value | 4.45536e-07 (rank : 19) | NC score | 0.447000 (rank : 11) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q6P1G2, Q3V396, Q6PFD0, Q6ZPE8, Q9CSF7, Q9QZN6 | Gene names | Fbxl10, Fbl10, Jhdm1b, Kiaa3014 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1B (EC 1.14.11.-) (F-box/LRR-repeat protein 10) (F-box and leucine-rich repeat protein 10) (F-box protein FBL10). | |||||
|
DNMT1_HUMAN
|
||||||
| θ value | 5.81887e-07 (rank : 20) | NC score | 0.309588 (rank : 33) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P26358, Q9UHG5, Q9ULA2, Q9UMZ6 | Gene names | DNMT1, AIM, DNMT | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 1 (EC 2.1.1.37) (Dnmt1) (DNA methyltransferase HsaI) (DNA MTase HsaI) (MCMT) (M.HsaI). | |||||
|
DNMT1_MOUSE
|
||||||
| θ value | 9.92553e-07 (rank : 21) | NC score | 0.325879 (rank : 28) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P13864, P97413, Q80ZU3, Q9CSC6, Q9QXX6 | Gene names | Dnmt1, Dnmt, Met1, Uim | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 1 (EC 2.1.1.37) (Dnmt1) (DNA methyltransferase MmuI) (DNA MTase MmuI) (MCMT) (M.MmuI) (Met-1). | |||||
|
JHD1A_HUMAN
|
||||||
| θ value | 1.29631e-06 (rank : 22) | NC score | 0.447451 (rank : 9) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9Y2K7, Q49A21, Q4G0M3, Q69YY8, Q9BVH5, Q9H7H5, Q9UK66 | Gene names | FBXL11, FBL7, JHDM1A, KIAA1004 | |||
|
Domain Architecture |

|
|||||
| Description | JmjC domain-containing histone demethylation protein 1A (EC 1.14.11.-) (F-box/LRR-repeat protein 11) (F-box and leucine-rich repeat protein 11) (F-box protein FBL7) (F-box protein Lilina). | |||||
|
JHD1A_MOUSE
|
||||||
| θ value | 1.29631e-06 (rank : 23) | NC score | 0.447433 (rank : 10) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P59997, Q3U1M5, Q3UR56, Q3V3Q1, Q69ZT4 | Gene names | Fbxl11, Jhdm1a, Kiaa1004 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1A (EC 1.14.11.-) (F-box/LRR-repeat protein 11) (F-box and leucine-rich repeat protein 11). | |||||
|
TAF3_MOUSE
|
||||||
| θ value | 1.69304e-06 (rank : 24) | NC score | 0.347500 (rank : 22) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 463 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q5HZG4, Q3U490, Q3UWX2, Q8BIU8, Q99JH4 | Gene names | Taf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
TAF3_HUMAN
|
||||||
| θ value | 2.88788e-06 (rank : 25) | NC score | 0.348086 (rank : 21) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 568 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q5VWG9, Q6GMS5, Q6P6B5, Q86VY6, Q9BQS9, Q9UFI8 | Gene names | TAF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
ING2_MOUSE
|
||||||
| θ value | 8.40245e-06 (rank : 26) | NC score | 0.361926 (rank : 17) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9ESK4, Q80VI5, Q8BGU8 | Gene names | Ing2, Ing1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 2 (p33ING2) (Inhibitor of growth 1-like protein). | |||||
|
ING1_HUMAN
|
||||||
| θ value | 1.09739e-05 (rank : 27) | NC score | 0.377760 (rank : 15) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9UK53, O00532, O43658, Q9H007, Q9HD98, Q9HD99, Q9NS83, Q9P0U6, Q9UBC6, Q9UIJ1, Q9UIJ2, Q9UIJ3, Q9UIJ4, Q9UK52 | Gene names | ING1 | |||
|
Domain Architecture |

|
|||||
| Description | Inhibitor of growth protein 1. | |||||
|
ING1_MOUSE
|
||||||
| θ value | 1.43324e-05 (rank : 28) | NC score | 0.377124 (rank : 16) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9QXV3, Q9QUP8, Q9QXV4, Q9QZX3 | Gene names | Ing1 | |||
|
Domain Architecture |

|
|||||
| Description | Inhibitor of growth protein 1. | |||||
|
ING2_HUMAN
|
||||||
| θ value | 1.87187e-05 (rank : 29) | NC score | 0.358415 (rank : 18) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9H160, O95698 | Gene names | ING2, ING1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 2 (p33ING2) (Inhibitor of growth 1-like protein) (ING1Lp) (p32). | |||||
|
ING3_HUMAN
|
||||||
| θ value | 0.00020696 (rank : 30) | NC score | 0.335821 (rank : 23) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9NXR8, O60394, Q6GMT3, Q7Z762, Q969G0, Q96DT4, Q9HC99, Q9P081 | Gene names | ING3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 3 (p47ING3 protein). | |||||
|
ING4_HUMAN
|
||||||
| θ value | 0.000270298 (rank : 31) | NC score | 0.334227 (rank : 24) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9UNL4, Q96E15, Q9H3J0 | Gene names | ING4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 4 (p29ING4). | |||||
|
ING4_MOUSE
|
||||||
| θ value | 0.000270298 (rank : 32) | NC score | 0.332521 (rank : 25) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8C0D7, Q8C1S7, Q8K3Q5, Q8K3Q6, Q8K3Q7, Q9D7F9 | Gene names | Ing4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 4 (p29ING4). | |||||
|
ING3_MOUSE
|
||||||
| θ value | 0.000602161 (rank : 33) | NC score | 0.326789 (rank : 27) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8VEK6, Q99JS6, Q9ERB2 | Gene names | Ing3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 3 (p47ING3 protein). | |||||
|
ING5_HUMAN
|
||||||
| θ value | 0.000602161 (rank : 34) | NC score | 0.314433 (rank : 31) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8WYH8, Q9BS30 | Gene names | ING5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 5 (p28ING5). | |||||
|
ING5_MOUSE
|
||||||
| θ value | 0.000602161 (rank : 35) | NC score | 0.312977 (rank : 32) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9D8Y8, Q3UL57, Q6P292, Q9CV64, Q9D9V8 | Gene names | Ing5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 5. | |||||
|
PHF13_HUMAN
|
||||||
| θ value | 0.00102713 (rank : 36) | NC score | 0.289500 (rank : 35) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q86YI8, Q8N551, Q9UJP2 | Gene names | PHF13 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 13. | |||||
|
PHF13_MOUSE
|
||||||
| θ value | 0.00175202 (rank : 37) | NC score | 0.287501 (rank : 36) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8K2W6 | Gene names | Phf13 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 13. | |||||
|
BAZ1A_HUMAN
|
||||||
| θ value | 0.00228821 (rank : 38) | NC score | 0.187522 (rank : 46) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 693 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9NRL2, Q9NZ15, Q9P065, Q9UIG1, Q9Y3V3 | Gene names | BAZ1A, ACF1, WCRF180 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1A (ATP-utilizing chromatin assembly and remodeling factor 1) (hACF1) (ATP-dependent chromatin remodelling protein) (Williams syndrome transcription factor-related chromatin remodeling factor 180) (WCRF180) (hWALp1) (CHRAC subunit ACF1). | |||||
|
JAD1A_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 39) | NC score | 0.214745 (rank : 41) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | P29375 | Gene names | JARID1A, RBBP2, RBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1A (Retinoblastoma-binding protein 2) (RBBP-2). | |||||
|
BAZ2A_MOUSE
|
||||||
| θ value | 0.0148317 (rank : 40) | NC score | 0.153533 (rank : 55) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q91YE5 | Gene names | Baz2a, Tip5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5). | |||||
|
CXXC6_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 41) | NC score | 0.192502 (rank : 45) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8NFU7, Q5VUP7, Q7Z6B6, Q8TCR1, Q9C0I7 | Gene names | CXXC6, KIAA1676, LCX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CXXC-type zinc finger protein 6 (Leukemia-associated protein with a CXXC domain). | |||||
|
PHF20_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 42) | NC score | 0.198527 (rank : 44) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9BVI0, Q5JWY9, Q9BWV4, Q9BXA3, Q9BZW3, Q9H421, Q9H4J6, Q9NZ22 | Gene names | PHF20, C20orf104, GLEA2, HCA58, NZF, TZP | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58) (Glioma-expressed antigen 2) (Transcription factor TZP) (Novel zinc finger protein). | |||||
|
PHF20_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 43) | NC score | 0.198716 (rank : 43) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8BLG0, Q8BMA2, Q8BYR4, Q921N1 | Gene names | Phf20, Hca58 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58 homolog). | |||||
|
C8AP2_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 44) | NC score | 0.085144 (rank : 87) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 548 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9UKL3, Q6PH76, Q7LCQ7, Q86YD9, Q9NUQ4, Q9NZV9, Q9P2N1, Q9Y563 | Gene names | CASP8AP2, FLASH, KIAA1315, RIP25 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
CN130_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 45) | NC score | 0.220311 (rank : 40) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8N806, Q86U21, Q86UA9, Q96BY0, Q9NVV6 | Gene names | C14orf130 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C14orf130. | |||||
|
JAD1D_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 46) | NC score | 0.161112 (rank : 49) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q62240, Q9QVR9, Q9R040 | Gene names | Smcy, Hya, Jarid1d | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1D (SmcY protein) (Histocompatibility Y antigen) (H-Y). | |||||
|
BAZ2A_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 47) | NC score | 0.150675 (rank : 56) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 780 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q9UIF9, O00536, O15030, Q96H26 | Gene names | BAZ2A, KIAA0314, TIP5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5) (hWALp3). | |||||
|
CN130_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 48) | NC score | 0.220478 (rank : 39) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8BU04, Q80UT9 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C14orf130 homolog. | |||||
|
DPF1_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 49) | NC score | 0.143955 (rank : 59) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q92782 | Gene names | DPF1, NEUD4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein neuro-d4 (D4, zinc and double PHD fingers family 1). | |||||
|
UHRF1_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 50) | NC score | 0.140038 (rank : 63) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 272 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q96T88, Q8J022, Q9H6S6, Q9P115, Q9P1U7 | Gene names | UHRF1, ICBP90, NP95, RNF106 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 1 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 1) (Inverted CCAAT box-binding protein of 90 kDa) (Transcription factor ICBP90) (Nuclear zinc finger protein Np95) (Nuclear protein 95) (HuNp95) (RING finger protein 106). | |||||
|
BAZ1B_MOUSE
|
||||||
| θ value | 0.125558 (rank : 51) | NC score | 0.164534 (rank : 48) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9Z277, Q9CU68 | Gene names | Baz1b, Wbscr9 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein homolog) (WBRS9). | |||||
|
INGX_HUMAN
|
||||||
| θ value | 0.125558 (rank : 52) | NC score | 0.317975 (rank : 30) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9P0U5 | Gene names | INGX, ING2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein, X-linked (Inhibitor of growth protein 2) (ING1-like tumor suppressor protein). | |||||
|
JAD1D_HUMAN
|
||||||
| θ value | 0.125558 (rank : 53) | NC score | 0.173940 (rank : 47) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q9BY66, Q92509, Q92809, Q9HCU1 | Gene names | SMCY, HY, HYA, JARID1D, KIAA0234 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1D (SmcY protein) (Histocompatibility Y antigen) (H-Y). | |||||
|
PHF1_HUMAN
|
||||||
| θ value | 0.125558 (rank : 54) | NC score | 0.147804 (rank : 57) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O43189, O60929, Q96KM7 | Gene names | PHF1 | |||
|
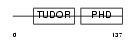
Domain Architecture |
|
|||||
| Description | PHD finger protein 1 (Protein PHF1). | |||||
|
PHF1_MOUSE
|
||||||
| θ value | 0.125558 (rank : 55) | NC score | 0.147706 (rank : 58) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9Z1B8, O54808 | Gene names | Phf1, Tctex-3, Tctex3 | |||
|
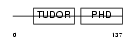
Domain Architecture |
|
|||||
| Description | PHD finger protein 1 (Protein PHF1) (T-complex testis-expressed 3) (Polycomblike 1) (mPCl1). | |||||
|
BAZ1B_HUMAN
|
||||||
| θ value | 0.21417 (rank : 56) | NC score | 0.159312 (rank : 52) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 701 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q9UIG0, O95039, O95247, O95277 | Gene names | BAZ1B, WBSC10, WBSCR9, WSTF | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein) (Williams syndrome transcription factor) (hWALP2). | |||||
|
MLL2_HUMAN
|
||||||
| θ value | 0.279714 (rank : 57) | NC score | 0.140097 (rank : 62) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
RSF1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 58) | NC score | 0.139819 (rank : 64) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 539 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q96T23, Q86X86, Q9H3L8, Q9NVZ8, Q9NYU0 | Gene names | RSF1, HBXAP, XAP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Remodeling and spacing factor 1 (Rsf-1) (Hepatitis B virus X- associated protein) (HBV pX-associated protein 8) (p325 subunit of RSF chromatin remodelling complex). | |||||
|
BAZ2B_HUMAN
|
||||||
| θ value | 0.365318 (rank : 59) | NC score | 0.122993 (rank : 69) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 803 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q9UIF8, Q96EA1, Q96SQ8, Q9P252, Q9Y4N8 | Gene names | BAZ2B, KIAA1476 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2B (hWALp4). | |||||
|
JAD1C_HUMAN
|
||||||
| θ value | 0.365318 (rank : 60) | NC score | 0.160871 (rank : 50) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P41229 | Gene names | SMCX, DXS1272E, JARID1C, XE169 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1C (Protein SmcX) (Protein Xe169). | |||||
|
JAD1C_MOUSE
|
||||||
| θ value | 0.365318 (rank : 61) | NC score | 0.160529 (rank : 51) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P41230, O54995, Q9CVI4, Q9D0C3, Q9QVR8, Q9R039 | Gene names | Smcx, Jarid1c, Xe169 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1C (SmcX protein) (Xe169 protein). | |||||
|
RIMS2_HUMAN
|
||||||
| θ value | 0.365318 (rank : 62) | NC score | 0.048124 (rank : 137) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UQ26, O43413, Q86XL9, Q8IWV9, Q8IWW1 | Gene names | RIMS2, KIAA0751, RIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 2 (Rab3-interacting molecule 2) (RIM 2). | |||||
|
RIMS2_MOUSE
|
||||||
| θ value | 0.365318 (rank : 63) | NC score | 0.049744 (rank : 136) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 376 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9EQZ7, Q8C433, Q8CCK2 | Gene names | Rims2, Rab3ip2, Rim2 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 2 (Rab3-interacting molecule 2) (RIM 2) (Rab3-interacting protein 2). | |||||
|
TCF19_HUMAN
|
||||||
| θ value | 0.365318 (rank : 64) | NC score | 0.199045 (rank : 42) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9Y242, Q13176, Q15967, Q5SQ89, Q5STD6, Q5STF5, Q9BUM2, Q9UBH7 | Gene names | TCF19, SC1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 19 (Transcription factor SC1). | |||||
|
TNIK_HUMAN
|
||||||
| θ value | 0.47712 (rank : 65) | NC score | 0.011563 (rank : 168) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1570 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9UKE5, O60298, Q8WUY7, Q9UKD8, Q9UKD9, Q9UKE0, Q9UKE1, Q9UKE2, Q9UKE3, Q9UKE4 | Gene names | TNIK, KIAA0551 | |||
|
Domain Architecture |

|
|||||
| Description | TRAF2 and NCK-interacting protein kinase (EC 2.7.11.1). | |||||
|
DPF1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 66) | NC score | 0.128651 (rank : 65) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9QX66, Q9QX65, Q9QYA4 | Gene names | Dpf1, Neud4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein neuro-d4 (D4, zinc and double PHD fingers family 1). | |||||
|
KI20A_HUMAN
|
||||||
| θ value | 1.06291 (rank : 67) | NC score | 0.020037 (rank : 157) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O95235 | Gene names | KIF20A, RAB6KIFL | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin family member 20A (Rabkinesin-6) (Rab6-interacting kinesin- like protein) (GG10_2). | |||||
|
MTF2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 68) | NC score | 0.157394 (rank : 53) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9Y483, Q9UES9, Q9UP40 | Gene names | MTF2 | |||
|
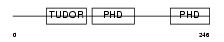
Domain Architecture |
|
|||||
| Description | Metal-response element-binding transcription factor 2 (Metal-response element DNA-binding protein M96) (Metal-regulatory transcription factor 2) (PCL2). | |||||
|
MTF2_MOUSE
|
||||||
| θ value | 1.06291 (rank : 69) | NC score | 0.156445 (rank : 54) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q02395, Q569Z8, Q6PG89, Q8BGP9, Q8BSJ7 | Gene names | Mtf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metal-response element-binding transcription factor 2 (Zinc-regulated factor 1) (ZiRF1) (Metal-response element DNA-binding protein M96) (Metal-regulatory transcription factor 2) (PCL2). | |||||
|
MYST4_HUMAN
|
||||||
| θ value | 1.06291 (rank : 70) | NC score | 0.116846 (rank : 74) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q8WYB5, O15087, Q86Y05, Q8WU81, Q9UKW2, Q9UKW3, Q9UKX0 | Gene names | MYST4, KIAA0383, MORF, MOZ2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Histone acetyltransferase MOZ2) (Monocytic leukemia zinc finger protein- related factor) (Histone acetyltransferase MORF). | |||||
|
RIMS1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 71) | NC score | 0.035769 (rank : 139) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 437 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q86UR5, O15048, Q8TDY9, Q8TDZ5, Q9HBA1, Q9HBA2, Q9HBA3, Q9HBA4, Q9HBA5, Q9HBA6 | Gene names | RIMS1, KIAA0340, RIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 1 (Rab3-interacting molecule 1) (RIM 1). | |||||
|
EVPL_HUMAN
|
||||||
| θ value | 1.38821 (rank : 72) | NC score | 0.022550 (rank : 154) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 895 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q92817 | Gene names | EVPL | |||
|
Domain Architecture |

|
|||||
| Description | Envoplakin (210 kDa paraneoplastic pemphigus antigen) (p210) (210 kDa cornified envelope precursor protein). | |||||
|
MYST4_MOUSE
|
||||||
| θ value | 1.38821 (rank : 73) | NC score | 0.113656 (rank : 76) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q8BRB7, Q7TNW5, Q8BG35, Q8C441, Q9JKX5 | Gene names | Myst4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Querkopf protein). | |||||
|
ZN598_MOUSE
|
||||||
| θ value | 1.38821 (rank : 74) | NC score | 0.027293 (rank : 148) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80YR4, Q6KAT0, Q8R3S1 | Gene names | Znf598, Zfp598 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 598. | |||||
|
K1045_HUMAN
|
||||||
| θ value | 1.81305 (rank : 75) | NC score | 0.035728 (rank : 140) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UPV7, Q58FE9, Q5T662 | Gene names | KIAA1045 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1045. | |||||
|
RBBP6_MOUSE
|
||||||
| θ value | 1.81305 (rank : 76) | NC score | 0.054590 (rank : 120) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
SMC1A_HUMAN
|
||||||
| θ value | 1.81305 (rank : 77) | NC score | 0.027516 (rank : 147) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1346 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q14683, O14995, Q16351 | Gene names | SMC1L1, DXS423E, KIAA0178, SB1.8, SMC1 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Sb1.8). | |||||
|
CTTB2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 78) | NC score | 0.010166 (rank : 169) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1061 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8WZ74, O43389, Q7LG11, Q9C0A5 | Gene names | CTTNBP2, C7orf8, CORTBP2, KIAA1758 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cortactin-binding protein 2 (CortBP2). | |||||
|
NC6IP_MOUSE
|
||||||
| θ value | 2.36792 (rank : 79) | NC score | 0.024522 (rank : 152) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q923W1, Q6DI60, Q6PEA7, Q8R0W9 | Gene names | Ncoa6ip, Pimt | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative RNA methyltransferase NCOA6IP (EC 2.1.1.-) (Nuclear receptor coactivator 6-interacting protein) (PRIP-interacting protein with methyltransferase motif) (PIPMT) (PIMT). | |||||
|
PKCB1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 80) | NC score | 0.119539 (rank : 72) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 488 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q9ULU4, Q13517, Q4JJ94, Q4JJ95, Q5TH09, Q6MZM1, Q8WXC5, Q9H1F3, Q9H1F4, Q9H1F5, Q9H1L8, Q9H1L9, Q9H2G5, Q9NYN3, Q9UIX6 | Gene names | PRKCBP1, KIAA1125, RACK7, ZMYND8 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C-binding protein 1 (Rack7) (Cutaneous T-cell lymphoma- associated antigen se14-3) (CTCL tumor antigen se14-3) (Zinc finger MYND domain-containing protein 8). | |||||
|
SMC1A_MOUSE
|
||||||
| θ value | 2.36792 (rank : 81) | NC score | 0.025917 (rank : 151) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1384 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9CU62, Q3V480, Q9CUX9, Q9D959, Q9WTU1 | Gene names | Smc1l1, Sb1.8, Smc1, Smcb | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Chromosome segregation protein SmcB) (Sb1.8). | |||||
|
MLL3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 82) | NC score | 0.118424 (rank : 73) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
MYST3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 83) | NC score | 0.107873 (rank : 78) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q92794 | Gene names | MYST3, MOZ, RUNXBP2, ZNF220 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Runt-related transcription factor-binding protein 2) (Monocytic leukemia zinc finger protein) (Zinc finger protein 220). | |||||
|
REQU_HUMAN
|
||||||
| θ value | 3.0926 (rank : 84) | NC score | 0.125535 (rank : 67) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 523 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q92785 | Gene names | DPF2, REQ, UBID4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein ubi-d4 (Requiem) (Apoptosis response zinc finger protein) (D4, zinc and double PHD fingers family 2). | |||||
|
REQU_MOUSE
|
||||||
| θ value | 3.0926 (rank : 85) | NC score | 0.122248 (rank : 70) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q61103, Q3UNP5, Q60663, Q9QYA3 | Gene names | Dpf2, Req, Ubid4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein ubi-d4 (Requiem) (Apoptosis response zinc finger protein) (D4, zinc and double PHD fingers family 2). | |||||
|
SYBU_MOUSE
|
||||||
| θ value | 3.0926 (rank : 86) | NC score | 0.032577 (rank : 141) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BHS8, Q3TQN7, Q401F3, Q401F4, Q571C1, Q6P1J2, Q80XH0, Q8BHS7, Q8BI27 | Gene names | Sybu, Golsyn, Kiaa1472 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntabulin (Syntaxin-1-binding protein) (Golgi-localized syntaphilin- related protein) (m-Golsyn). | |||||
|
TRHY_HUMAN
|
||||||
| θ value | 3.0926 (rank : 87) | NC score | 0.019982 (rank : 158) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1385 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q07283 | Gene names | THH, THL, TRHY | |||
|
Domain Architecture |

|
|||||
| Description | Trichohyalin. | |||||
|
UHRF1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 88) | NC score | 0.121286 (rank : 71) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8VDF2, Q8C6F1, Q8VIA1, Q9Z1H6 | Gene names | Uhrf1, Np95 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 1 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 1) (Nuclear zinc finger protein Np95) (Nuclear protein 95). | |||||
|
CCD46_HUMAN
|
||||||
| θ value | 4.03905 (rank : 89) | NC score | 0.017634 (rank : 160) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1430 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8N8E3, Q6PIB5, Q8NCR4, Q8NFR4 | Gene names | CCDC46 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 46. | |||||
|
HNRLL_MOUSE
|
||||||
| θ value | 4.03905 (rank : 90) | NC score | 0.012153 (rank : 165) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q921F4, Q8BIP6, Q99J40 | Gene names | Hnrpll | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heterogeneous nuclear ribonucleoprotein L-like. | |||||
|
IF3A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 91) | NC score | 0.028543 (rank : 146) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 875 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q14152, O00653 | Gene names | EIF3S10, KIAA0139 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 10 (eIF-3 theta) (eIF3 p167) (eIF3 p180) (eIF3 p185) (eIF3a). | |||||
|
LAMA1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 92) | NC score | 0.005084 (rank : 176) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 804 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P25391 | Gene names | LAMA1, LAMA | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-1 chain precursor (Laminin A chain). | |||||
|
MLL3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 93) | NC score | 0.124156 (rank : 68) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
MOR2A_MOUSE
|
||||||
| θ value | 4.03905 (rank : 94) | NC score | 0.029703 (rank : 143) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q69ZX6, Q5QNQ7, Q6P547 | Gene names | Morc2a, Kiaa0852, Zcwcc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORC family CW-type zinc finger protein 2A (Zinc finger CW-type coiled-coil domain protein 1). | |||||
|
PLCG1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 95) | NC score | 0.009642 (rank : 170) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 314 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q62077, Q6P1G1 | Gene names | Plcg1, Plcg-1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 1-phosphatidylinositol-4,5-bisphosphate phosphodiesterase gamma 1 (EC 3.1.4.11) (Phosphoinositide phospholipase C) (PLC-gamma-1) (Phospholipase C-gamma-1) (PLC-II) (PLC-148). | |||||
|
RUFY2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 96) | NC score | 0.017300 (rank : 163) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1208 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8WXA3, Q96P51, Q9P1Z1 | Gene names | RUFY2, KIAA1537, RABIP4R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and FYVE domain-containing protein 2 (Rab4-interacting protein related). | |||||
|
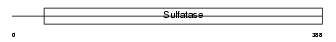
SULF2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 97) | NC score | 0.020487 (rank : 156) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CFG0, Q8BM68, Q8BUZ4, Q8BX28, Q8BZ51, Q8C169, Q9D8E2 | Gene names | Sulf2 | |||
|
Domain Architecture |
|
|||||
| Description | Extracellular sulfatase Sulf-2 precursor (EC 3.1.6.-) (MSulf-2). | |||||
|
TNIP3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 98) | NC score | 0.026752 (rank : 150) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 511 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q96KP6, Q96PQ3, Q9H780 | Gene names | TNIP3, ABIN3, LIND | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TNFAIP3-interacting protein 3 (Listeria-induced gene protein) (A20- binding inhibitor of NF-kappa-B activation 3) (ABIN-3). | |||||
|
AN30A_HUMAN
|
||||||
| θ value | 5.27518 (rank : 99) | NC score | 0.014967 (rank : 164) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1119 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9BXX3, Q5W025 | Gene names | ANKRD30A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 30A (NY-BR-1 antigen). | |||||
|
E2AK4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 100) | NC score | 0.005500 (rank : 175) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1012 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9P2K8, Q69YL7, Q6DC97, Q96GN6, Q9H5K1, Q9NSQ3, Q9NSZ5, Q9UJ56 | Gene names | EIF2AK4, GCN2, KIAA1338 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 2-alpha kinase 4 (EC 2.7.11.1) (GCN2-like protein). | |||||
|
IF3A_MOUSE
|
||||||
| θ value | 5.27518 (rank : 101) | NC score | 0.029649 (rank : 144) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 871 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P23116, Q60697, Q62162 | Gene names | Eif3s10, Csma, Eif3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 10 (eIF-3 theta) (eIF3 p167) (eIF3 p180) (eIF3 p185) (eIF3a) (p162 protein) (Centrosomin). | |||||
|
MINK1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 102) | NC score | 0.006659 (rank : 172) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8N4C8, Q9P1X1, Q9P2R8 | Gene names | MINK1, MAP4K6, MINK | |||
|
Domain Architecture |

|
|||||
| Description | Misshapen-like kinase 1 (EC 2.7.11.1) (Mitogen-activated protein kinase kinase kinase kinase 6) (MAPK/ERK kinase kinase kinase 6) (MEK kinase kinase 6) (MEKKK 6) (Misshapen/NIK-related kinase) (GCK family kinase MiNK). | |||||
|
MINK1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 103) | NC score | 0.006952 (rank : 171) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1451 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9JM52, Q921M6, Q9JM92 | Gene names | Mink1, Map4k6, Mink | |||
|
Domain Architecture |

|
|||||
| Description | Misshapen-like kinase 1 (EC 2.7.11.1) (Mitogen-activated protein kinase kinase kinase kinase 6) (MAPK/ERK kinase kinase kinase 6) (MEK kinase kinase 6) (MEKKK 6) (Misshapen/NIK-related kinase) (GCK family kinase MiNK). | |||||
|
NHS_HUMAN
|
||||||
| θ value | 5.27518 (rank : 104) | NC score | 0.032403 (rank : 142) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6T4R5, Q5J7Q0, Q5J7Q1, Q68DR5 | Gene names | NHS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nance-Horan syndrome protein (Congenital cataracts and dental anomalies protein). | |||||
|
NSD1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 105) | NC score | 0.105984 (rank : 79) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q96L73, Q96PD8, Q96RN7 | Gene names | NSD1, ARA267 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein) (Androgen receptor-associated coregulator 267). | |||||
|
NSD1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 106) | NC score | 0.113586 (rank : 77) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | O88491, Q8C480, Q9CT70 | Gene names | Nsd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein). | |||||
|
PF21A_MOUSE
|
||||||
| θ value | 5.27518 (rank : 107) | NC score | 0.127107 (rank : 66) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q6ZPK0, Q6XVG0, Q80Z33, Q8CAZ4, Q8VEC8 | Gene names | Phf21a, Bhc80, Kiaa1696, Pftf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (mBHC80) (BHC80a). | |||||
|
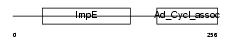
TTC28_HUMAN
|
||||||
| θ value | 5.27518 (rank : 108) | NC score | 0.011745 (rank : 166) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96AY4, O95928, O95929, Q9NTE4, Q9UG31, Q9UGG5, Q9UPV8, Q9Y3S5 | Gene names | TTC28, KIAA1043 | |||
|
Domain Architecture |
|
|||||
| Description | Tetratricopeptide repeat protein 28 (TPR repeat protein 28). | |||||
|
E2AK4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 109) | NC score | 0.004740 (rank : 177) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 984 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9QZ05, Q6GQT4, Q6ZPT5, Q8C5S0, Q8CIF5, Q9CT30, Q9CUV9, Q9ESB6, Q9ESB7, Q9ESB8 | Gene names | Eif2ak4, Gcn2, Kiaa1338 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 2-alpha kinase 4 (EC 2.7.11.1) (GCN2-like protein) (mGCN2). | |||||
|
JADE3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 110) | NC score | 0.068119 (rank : 98) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q92613, Q6IE79 | Gene names | PHF16, JADE3, KIAA0215 | |||
|
Domain Architecture |

|
|||||
| Description | Protein Jade-3 (PHD finger protein 16). | |||||
|
JADE3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 111) | NC score | 0.066477 (rank : 99) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q6IE82, Q6ZQG2 | Gene names | Phf16, Jade3, Kiaa0215 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-3 (PHD finger protein 16). | |||||
|
LRRC6_HUMAN
|
||||||
| θ value | 6.88961 (rank : 112) | NC score | 0.017377 (rank : 162) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q86X45, Q13648, Q4G183 | Gene names | LRRC6, LRTP, TSLRP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 6 (Leucine-rich testis-specific protein) (Testis-specific leucine-rich repeat protein). | |||||
|
M4K4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 113) | NC score | 0.006317 (rank : 174) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1589 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O95819, O75172, Q9NST7 | Gene names | MAP4K4, HGK, KIAA0687, NIK | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase kinase 4 (EC 2.7.11.1) (MAPK/ERK kinase kinase kinase 4) (MEK kinase kinase 4) (MEKKK 4) (HPK/GCK-like kinase HGK) (Nck-interacting kinase). | |||||
|
M4K4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 114) | NC score | 0.006573 (rank : 173) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1560 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P97820 | Gene names | Map4k4, Nik | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase kinase 4 (EC 2.7.11.1) (MAPK/ERK kinase kinase kinase 4) (MEK kinase kinase 4) (MEKKK 4) (HPK/GCK-like kinase HGK) (Nck-interacting kinase). | |||||
|
ZCHC7_HUMAN
|
||||||
| θ value | 6.88961 (rank : 115) | NC score | 0.029313 (rank : 145) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8N3Z6, Q5T0Q8, Q5T0Q9, Q5T0R0, Q8N2M1, Q8N4J2, Q8TBK8, Q9H648, Q9P0F0 | Gene names | ZCCHC7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 7. | |||||
|
BRD2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 116) | NC score | 0.042658 (rank : 138) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P25440, O00699, O00700, Q15310, Q5STC9, Q969U4 | Gene names | BRD2, KIAA9001, RING3 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 2 (Protein RING3) (O27.1.1). | |||||
|
CCD46_MOUSE
|
||||||
| θ value | 8.99809 (rank : 117) | NC score | 0.020838 (rank : 155) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q5PR68, Q80ZN6, Q99JS4, Q9D9T1 | Gene names | Ccdc46 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled coil domain-containing protein 46. | |||||
|
CNTRB_HUMAN
|
||||||
| θ value | 8.99809 (rank : 118) | NC score | 0.017598 (rank : 161) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8N137, Q331K3, Q69YV7, Q8NCB8, Q8WXV3, Q96CQ7, Q9C060 | Gene names | CNTROB, LIP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrobin (LYST-interacting protein 8). | |||||
|
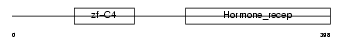
COT1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 119) | NC score | 0.004285 (rank : 178) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q60632, Q61438 | Gene names | Nr2f1, Erbal3, Tfcoup1 | |||
|
Domain Architecture |
|
|||||
| Description | COUP transcription factor 1 (COUP-TF1) (COUP-TF I). | |||||
|
CYLN2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 120) | NC score | 0.011742 (rank : 167) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1004 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9UDT6, O14527, O43611 | Gene names | CYLN2, KIAA0291, WBSCR4, WSCR4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoplasmic linker protein 2 (Cytoplasmic linker protein 115) (CLIP- 115) (Williams-Beuren syndrome chromosome region 4 protein). | |||||
|
FA47B_HUMAN
|
||||||
| θ value | 8.99809 (rank : 121) | NC score | 0.019789 (rank : 159) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 431 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8NA70, Q5JQN5, Q6PIG3 | Gene names | FAM47B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM47B. | |||||
|
MRCKG_HUMAN
|
||||||
| θ value | 8.99809 (rank : 122) | NC score | 0.001724 (rank : 179) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1519 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q6DT37, O00565 | Gene names | CDC42BPG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK gamma (EC 2.7.11.1) (CDC42- binding protein kinase gamma) (Myotonic dystrophy kinase-related CDC42-binding kinase gamma) (Myotonic dystrophy protein kinase-like alpha) (MRCK gamma) (DMPK-like gamma). | |||||
|
PYGO1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 123) | NC score | 0.026940 (rank : 149) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y3Y4 | Gene names | PYGO1 | |||
|
Domain Architecture |

|
|||||
| Description | Pygopus homolog 1. | |||||
|
SYBU_HUMAN
|
||||||
| θ value | 8.99809 (rank : 124) | NC score | 0.024258 (rank : 153) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9NX95, Q5R1T1, Q5R1T2, Q5R1T3, Q5Y2M6, Q8ND49, Q8TCR6, Q9P256 | Gene names | SYBU, GOLSYN, KIAA1472 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntabulin (Syntaxin-1-binding protein) (Golgi-localized syntaphilin- related protein). | |||||
|
AIRE_HUMAN
|
||||||
| θ value | θ > 10 (rank : 125) | NC score | 0.096325 (rank : 82) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O43918, O43922, O43932, O75745 | Gene names | AIRE, APECED | |||
|
Domain Architecture |

|
|||||
| Description | Autoimmune regulator (Autoimmune polyendocrinopathy candidiasis ectodermal dystrophy protein) (APECED protein). | |||||
|
AIRE_MOUSE
|
||||||
| θ value | θ > 10 (rank : 126) | NC score | 0.095112 (rank : 83) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9Z0E3, Q9JLW0, Q9JLW1, Q9JLW2, Q9JLW3, Q9JLW4, Q9JLW5, Q9JLW6, Q9JLW7, Q9JLW8, Q9JLW9, Q9JLX0 | Gene names | Aire | |||
|
Domain Architecture |

|
|||||
| Description | Autoimmune regulator (Autoimmune polyendocrinopathy candidiasis ectodermal dystrophy protein homolog) (APECED protein homolog). | |||||
|
BRD1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 127) | NC score | 0.057388 (rank : 109) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O95696 | Gene names | BRD1, BRL, BRPF2 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 1 (BR140-like protein). | |||||
|
BRPF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 128) | NC score | 0.059419 (rank : 107) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P55201, Q9UHI0 | Gene names | BRPF1, BR140 | |||
|
Domain Architecture |

|
|||||
| Description | Peregrin (Bromodomain and PHD finger-containing protein 1) (BR140 protein). | |||||
|
CECR2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 129) | NC score | 0.056728 (rank : 112) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9BXF3, Q658Z4, Q96P58, Q9C0C3 | Gene names | CECR2, KIAA1740 | |||
|
Domain Architecture |

|
|||||
| Description | Cat eye syndrome critical region protein 2. | |||||
|
CHD3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 130) | NC score | 0.063002 (rank : 105) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q12873, Q9Y4I0 | Gene names | CHD3 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain helicase-DNA-binding protein 3 (EC 3.6.1.-) (ATP- dependent helicase CHD3) (CHD-3) (Mi-2 autoantigen 240 kDa protein) (Mi2-alpha) (Zinc-finger helicase) (hZFH). | |||||
|
CHD4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 131) | NC score | 0.063282 (rank : 104) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 365 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q14839, Q8IXZ5 | Gene names | CHD4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain helicase-DNA-binding protein 4 (EC 3.6.1.-) (ATP- dependent helicase CHD4) (CHD-4) (Mi-2 autoantigen 218 kDa protein) (Mi2-beta). | |||||
|
CHD4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 132) | NC score | 0.064076 (rank : 102) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q6PDQ2 | Gene names | Chd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain helicase-DNA-binding protein 4 (CHD-4). | |||||
|
CHD5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 133) | NC score | 0.065849 (rank : 101) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q8TDI0, O75032, Q5TG89, Q9UFR9 | Gene names | CHD5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain helicase-DNA-binding protein 5 (EC 3.6.1.-) (ATP- dependent helicase CHD5) (CHD-5). | |||||
|
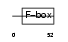
FBXL2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 134) | NC score | 0.050707 (rank : 134) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UKC9, Q9NVQ8, Q9UK27, Q9UKA5, Q9Y3Y9 | Gene names | FBXL2, FBL2, FBL3 | |||
|
Domain Architecture |
|
|||||
| Description | F-box/LRR-repeat protein 2 (F-box and leucine-rich repeat protein 2) (F-box protein FBL2/FBL3). | |||||
|
FBXL2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 135) | NC score | 0.053194 (rank : 128) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BH16, Q8BXM4 | Gene names | Fbxl2, Fbl2 | |||
|
Domain Architecture |
|
|||||
| Description | F-box/LRR-repeat protein 2 (F-box and leucine-rich repeat protein 2). | |||||
|
FBXL7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 136) | NC score | 0.063910 (rank : 103) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UJT9, O94926 | Gene names | FBXL7, FBL6, FBL7, KIAA0840 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 7 (F-box and leucine-rich repeat protein 7) (F-box protein FBL6/FBL7). | |||||
|
FXL13_MOUSE
|
||||||
| θ value | θ > 10 (rank : 137) | NC score | 0.055679 (rank : 118) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CDU4, Q8CDE9 | Gene names | Fbxl13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 13 (F-box and leucine-rich repeat protein 13). | |||||
|
FXL14_HUMAN
|
||||||
| θ value | θ > 10 (rank : 138) | NC score | 0.053993 (rank : 124) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8N1E6 | Gene names | FBXL14, FBL14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 14 (F-box and leucine-rich repeat protein 14). | |||||
|
FXL14_MOUSE
|
||||||
| θ value | θ > 10 (rank : 139) | NC score | 0.054146 (rank : 122) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BID8, Q3U2Q9, Q8R5H7, Q8VDT7, Q922N5 | Gene names | Fbxl14, Fbl14, Ppa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 14 (F-box and leucine-rich repeat protein 14). | |||||
|
FXL16_HUMAN
|
||||||
| θ value | θ > 10 (rank : 140) | NC score | 0.056175 (rank : 115) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8N461, Q2MHR2, Q96S14, Q9UJI0 | Gene names | FBXL16, C16orf22, FBL16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 16 (F-box and leucine-rich repeat protein 16). | |||||
|
FXL20_HUMAN
|
||||||
| θ value | θ > 10 (rank : 141) | NC score | 0.057226 (rank : 110) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96IG2 | Gene names | FBXL20, FBL2 | |||
|
Domain Architecture |
|
|||||
| Description | F-box/LRR-repeat protein 20 (F-box and leucine-rich repeat protein 20) (F-box/LRR-repeat protein 2-like). | |||||
|
FXL20_MOUSE
|
||||||
| θ value | θ > 10 (rank : 142) | NC score | 0.057115 (rank : 111) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9CZV8, Q8BZ95 | Gene names | Fbxl20, Fbl2 | |||
|
Domain Architecture |
|
|||||
| Description | F-box/LRR-repeat protein 20 (F-box and leucine-rich repeat protein 20) (F-box/LRR-repeat protein 2-like). | |||||
|
JADE1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 143) | NC score | 0.054438 (rank : 121) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q6IE81, Q4W5D5, Q6ZSL7, Q8NC41, Q96JL8, Q96SQ1, Q9H692 | Gene names | PHF17, JADE1, KIAA1807 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-1 (PHD finger protein 17). | |||||
|
JADE1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 144) | NC score | 0.054017 (rank : 123) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q6ZPI0, Q6IE84, Q8C7S6, Q8CFM2, Q8R362 | Gene names | Phf17, Jade1, Kiaa1807 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-1 (PHD finger protein 17). | |||||
|
JADE2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 145) | NC score | 0.050686 (rank : 135) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9NQC1, Q6IE80, Q8TEK0, Q92513, Q96GQ6 | Gene names | PHF15, JADE2, KIAA0239 | |||
|
Domain Architecture |

|
|||||
| Description | Protein Jade-2 (PHD finger protein 15). | |||||
|
JADE2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 146) | NC score | 0.056068 (rank : 116) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q6ZQF7, Q3UHD5, Q6IE83 | Gene names | Phf15, Jade2, Kiaa0239 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-2 (PHD finger protein 15). | |||||
|
LRC29_HUMAN
|
||||||
| θ value | θ > 10 (rank : 147) | NC score | 0.059475 (rank : 106) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8WV35, Q9UKA0 | Gene names | LRRC29, FBL9, FBXL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 29 (F-box/LRR-repeat protein 9) (F-box and leucine-rich repeat protein 9) (F-box protein FBL9). | |||||
|
MBD2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 148) | NC score | 0.051343 (rank : 131) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UBB5, O95242, Q9UIS8 | Gene names | MBD2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 2 (Methyl-CpG-binding protein MBD2) (Demethylase) (DMTase). | |||||
|
MBD2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 149) | NC score | 0.051991 (rank : 130) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Z2E1, Q811D9, Q9Z2D9 | Gene names | Mbd2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 2 (Methyl-CpG-binding protein MBD2). | |||||
|
MYST3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 150) | NC score | 0.100792 (rank : 80) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 688 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q8BZ21, Q8BW52, Q8BYH1, Q8C1F3 | Gene names | Myst3, Moz | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Monocytic leukemia zinc finger protein) (Monocytic leukemia zinc finger homolog). | |||||
|
PF21A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 151) | NC score | 0.115054 (rank : 75) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q96BD5, Q6AWA2, Q9C0G7, Q9H8V9, Q9HAK6, Q9NZE9 | Gene names | PHF21A, BHC80, KIAA1696 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (BHC80a). | |||||
|
PF21B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 152) | NC score | 0.092390 (rank : 86) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q96EK2, Q5TFL2, Q6ICC0 | Gene names | PHF21B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
PF21B_MOUSE
|
||||||
| θ value | θ > 10 (rank : 153) | NC score | 0.098246 (rank : 81) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q8C966 | Gene names | Phf21b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
PHF10_HUMAN
|
||||||
| θ value | θ > 10 (rank : 154) | NC score | 0.143347 (rank : 60) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q8WUB8, Q2YDA3, Q9BXD2, Q9NV26 | Gene names | PHF10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 10 (XAP135). | |||||
|
PHF10_MOUSE
|
||||||
| θ value | θ > 10 (rank : 155) | NC score | 0.140227 (rank : 61) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9D8M7, Q99LV5 | Gene names | Phf10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 10. | |||||
|
PHF11_HUMAN
|
||||||
| θ value | θ > 10 (rank : 156) | NC score | 0.053664 (rank : 126) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UIL8, Q9Y5A2 | Gene names | PHF11, BCAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 11 (BRCA1 C-terminus-associated protein) (NY-REN-34 antigen). | |||||
|
PHF12_HUMAN
|
||||||
| θ value | θ > 10 (rank : 157) | NC score | 0.079323 (rank : 89) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q96QT6, Q6ZML2, Q9BV34, Q9H7U9, Q9P205 | Gene names | PHF12, KIAA1523 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 12 (PHD factor 1) (Pf1). | |||||
|
PHF12_MOUSE
|
||||||
| θ value | θ > 10 (rank : 158) | NC score | 0.077065 (rank : 91) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q5SPL2, Q5SPL3, Q6ZPN8, Q80W50 | Gene names | Phf12, Kiaa1523 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 12 (PHD factor 1) (Pf1). | |||||
|
PHF14_HUMAN
|
||||||
| θ value | θ > 10 (rank : 159) | NC score | 0.071740 (rank : 95) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | O94880 | Gene names | PHF14, KIAA0783 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 14. | |||||
|
PHF14_MOUSE
|
||||||
| θ value | θ > 10 (rank : 160) | NC score | 0.072378 (rank : 94) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9D4H9, Q810Z6, Q8C284, Q8CGF8, Q9D1Q8 | Gene names | Phf14 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 14. | |||||
|
PHF22_HUMAN
|
||||||
| θ value | θ > 10 (rank : 161) | NC score | 0.050743 (rank : 133) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q96CB8, Q9HD71 | Gene names | INTS12, PHF22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integrator complex subunit 12 (PHD finger protein 22). | |||||
|
PHF22_MOUSE
|
||||||
| θ value | θ > 10 (rank : 162) | NC score | 0.056304 (rank : 114) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9D168, Q3U2Q5, Q921U2 | Gene names | Ints12, Phf22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integrator complex subunit 12 (PHD finger protein 22). | |||||
|
PTDSR_HUMAN
|
||||||
| θ value | θ > 10 (rank : 163) | NC score | 0.078618 (rank : 90) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q6NYC1, Q86VY0, Q8IUM5, Q9Y4E2 | Gene names | PTDSR, KIAA0585 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein PTDSR (Phosphatidylserine receptor). | |||||
|
PTDSR_MOUSE
|
||||||
| θ value | θ > 10 (rank : 164) | NC score | 0.080886 (rank : 88) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9ERI5, Q80TX1 | Gene names | Ptdsr, Kiaa0585 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein PTDSR (Phosphatidylserine receptor) (Apoptotic cell clearance receptor PtdSerR). | |||||
|
SET1A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 165) | NC score | 0.071045 (rank : 96) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O15047, Q6PIF3, Q8TAJ6 | Gene names | SETD1A, KIAA0339, SET1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET1 (EC 2.1.1.43) (Set1/Ash2 histone methyltransferase complex subunit SET1) (SET-domain-containing protein 1A). | |||||
|
SETD8_HUMAN
|
||||||
| θ value | θ > 10 (rank : 166) | NC score | 0.055921 (rank : 117) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9NQR1, Q86W83, Q8TD09 | Gene names | SETD8, PRSET7, SET07, SET8 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H4 lysine-20 specific (EC 2.1.1.43) (Histone H4-K20 methyltransferase) (H4-K20-HMTase) (SET domain-containing lysine methyltransferase 8) (SET domain-containing protein 8) (PR/SET domain-containing protein 07) (PR/SET07) (PR-Set7). | |||||
|
SETD8_MOUSE
|
||||||
| θ value | θ > 10 (rank : 167) | NC score | 0.053229 (rank : 127) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q2YDW7, Q8C0J9 | Gene names | SETD8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H4 lysine-20 specific (EC 2.1.1.43) (Histone H4-K20 methyltransferase) (H4-K20-HMTase) (SET domain-containing lysine methyltransferase 8) (SET domain-containing protein 8). | |||||
|
SKP2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 168) | NC score | 0.058290 (rank : 108) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q13309, Q8TDZ0, Q8TDZ1, Q9BV69 | Gene names | SKP2, FBXL1 | |||
|
Domain Architecture |

|
|||||
| Description | S-phase kinase-associated protein 2 (F-box protein Skp2) (Cyclin A/CDK2-associated protein p45) (p45skp2) (F-box/LRR-repeat protein 1). | |||||
|
SKP2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 169) | NC score | 0.056307 (rank : 113) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Z0Z3, Q8BSG7, Q8C8Y9, Q8CB22, Q99J53 | Gene names | Skp2 | |||
|
Domain Architecture |

|
|||||
| Description | S-phase kinase-associated protein 2 (F-box protein Skp2) (Cyclin A/CDK2-associated protein p45) (F-box/WD-40 protein 1) (FWD1). | |||||
|
TIF1A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 170) | NC score | 0.069344 (rank : 97) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O15164, O95854 | Gene names | TRIM24, RNF82, TIF1, TIF1A | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-alpha (TIF1-alpha) (Tripartite motif-containing protein 24) (RING finger protein 82). | |||||
|
TIF1A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 171) | NC score | 0.072897 (rank : 93) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q64127, Q64126 | Gene names | Trim24, Tif1, Tif1a | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-alpha (TIF1-alpha) (Tripartite motif-containing protein 24). | |||||
|
TIF1G_HUMAN
|
||||||
| θ value | θ > 10 (rank : 172) | NC score | 0.050823 (rank : 132) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 410 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9UPN9, O95855, Q9C017, Q9UJ79 | Gene names | TRIM33, KIAA1113, RFG7, TIF1G | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-gamma (TIF1-gamma) (Tripartite motif-containing protein 33) (RET-fused gene 7 protein) (Rfg7 protein). | |||||
|
TIF1G_MOUSE
|
||||||
| θ value | θ > 10 (rank : 173) | NC score | 0.053783 (rank : 125) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q99PP7, Q6SI71, Q6ZPX5 | Gene names | Trim33, Kiaa1113 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription intermediary factor 1-gamma (TIF1-gamma) (Tripartite motif-containing protein 33). | |||||
|
TRI66_HUMAN
|
||||||
| θ value | θ > 10 (rank : 174) | NC score | 0.065915 (rank : 100) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | O15016, Q9BQQ4 | Gene names | TRIM66, KIAA0298 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
TRI66_MOUSE
|
||||||
| θ value | θ > 10 (rank : 175) | NC score | 0.072902 (rank : 92) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q924W6 | Gene names | Trim66 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
UHRF2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 176) | NC score | 0.093642 (rank : 84) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q96PU4, Q5VYR1, Q5VYR3, Q659C8, Q8TAG7 | Gene names | UHRF2, NIRF, RNF107 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 2 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 2) (Np95/ICBP90-like RING finger protein) (Np95-like RING finger protein) (Nuclear zinc finger protein Np97) (RING finger protein 107). | |||||
|
UHRF2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 177) | NC score | 0.092447 (rank : 85) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q7TMI3, Q8BG56, Q8BJP6, Q8BY30, Q8K1I5 | Gene names | Uhrf2, Nirf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 2 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 2) (Np95-like ring finger protein) (Nuclear zinc finger protein Np97) (NIRF). | |||||
|
ZMY11_HUMAN
|
||||||
| θ value | θ > 10 (rank : 178) | NC score | 0.055512 (rank : 119) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q15326 | Gene names | ZMYND11, BS69 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger MYND domain-containing protein 11 (Adenovirus 5 E1A- binding protein) (BS69 protein). | |||||
|
ZMY11_MOUSE
|
||||||
| θ value | θ > 10 (rank : 179) | NC score | 0.052493 (rank : 129) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8R5C8, Q8C155 | Gene names | Zmynd11 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger MYND domain-containing protein 11. | |||||
|
CXCC1_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 124 | |
| SwissProt Accessions | Q9CWW7 | Gene names | Cxxc1, Cgbp, Pccx1 | |||
|
Domain Architecture |

|
|||||
| Description | CpG-binding protein (PHD finger and CXXC domain-containing protein 1) (CXXC-type zinc finger protein 1). | |||||
|
CXCC1_HUMAN
|
||||||
| NC score | 0.992935 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 111 | |
| SwissProt Accessions | Q9P0U4, Q8N2W4, Q96BC8, Q9P2V7 | Gene names | CXXC1, CGBP, PCCX1, PHF18 | |||
|
Domain Architecture |

|
|||||
| Description | CpG-binding protein (PHD finger and CXXC domain-containing protein 1) (CXXC-type zinc finger protein 1). | |||||
|
K1718_MOUSE
|
||||||
| NC score | 0.500364 (rank : 3) | θ value | 3.29651e-10 (rank : 8) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q3UWM4, Q3UWN8, Q6ZPJ5, Q8C969, Q8C9E0, Q91VX8 | Gene names | Kiaa1718 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1718. | |||||
|
PHF2_MOUSE
|
||||||
| NC score | 0.498104 (rank : 4) | θ value | 1.9326e-10 (rank : 6) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9WTU0, Q80WA8 | Gene names | Phf2 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
PHF2_HUMAN
|
||||||
| NC score | 0.497552 (rank : 5) | θ value | 1.9326e-10 (rank : 5) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O75151, Q8N3K2, Q9Y6N4 | Gene names | PHF2, KIAA0662 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
K1718_HUMAN
|
||||||
| NC score | 0.495770 (rank : 6) | θ value | 9.59137e-10 (rank : 9) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q6ZMT4, Q6MZL8, Q9C0E5 | Gene names | KIAA1718 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1718. | |||||
|
PHF8_MOUSE
|
||||||
| NC score | 0.490890 (rank : 7) | θ value | 2.79066e-09 (rank : 11) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q80TJ7, Q8BLX8, Q8BLY0, Q8BZ61, Q8CG26 | Gene names | Phf8, Kiaa1111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 8. | |||||
|
PHF8_HUMAN
|
||||||
| NC score | 0.490834 (rank : 8) | θ value | 1.25267e-09 (rank : 10) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9UPP1, Q5H9U5, Q5VUJ4, Q7Z6D4, Q9HAH2 | Gene names | PHF8, KIAA1111, ZNF422 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 8. | |||||
|
JHD1A_HUMAN
|
||||||
| NC score | 0.447451 (rank : 9) | θ value | 1.29631e-06 (rank : 22) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9Y2K7, Q49A21, Q4G0M3, Q69YY8, Q9BVH5, Q9H7H5, Q9UK66 | Gene names | FBXL11, FBL7, JHDM1A, KIAA1004 | |||
|
Domain Architecture |

|
|||||
| Description | JmjC domain-containing histone demethylation protein 1A (EC 1.14.11.-) (F-box/LRR-repeat protein 11) (F-box and leucine-rich repeat protein 11) (F-box protein FBL7) (F-box protein Lilina). | |||||
|
JHD1A_MOUSE
|
||||||
| NC score | 0.447433 (rank : 10) | θ value | 1.29631e-06 (rank : 23) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P59997, Q3U1M5, Q3UR56, Q3V3Q1, Q69ZT4 | Gene names | Fbxl11, Jhdm1a, Kiaa1004 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1A (EC 1.14.11.-) (F-box/LRR-repeat protein 11) (F-box and leucine-rich repeat protein 11). | |||||
|
JHD1B_MOUSE
|
||||||
| NC score | 0.447000 (rank : 11) | θ value | 4.45536e-07 (rank : 19) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q6P1G2, Q3V396, Q6PFD0, Q6ZPE8, Q9CSF7, Q9QZN6 | Gene names | Fbxl10, Fbl10, Jhdm1b, Kiaa3014 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1B (EC 1.14.11.-) (F-box/LRR-repeat protein 10) (F-box and leucine-rich repeat protein 10) (F-box protein FBL10). | |||||
|
JHD1B_HUMAN
|
||||||
| NC score | 0.446326 (rank : 12) | θ value | 4.45536e-07 (rank : 18) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8NHM5, Q8NCI2, Q96HC7, Q96SL0, Q96T03, Q9NS96, Q9UF75 | Gene names | FBXL10, CXXC2, FBL10, JHDM1B, PCCX2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1B (EC 1.14.11.-) (F-box/LRR-repeat protein 10) (F-box and leucine-rich repeat protein 10) (F-box protein FBL10) (Protein JEMMA) (Jumonji domain-containing EMSY-interactor methyltransferase motif protein) (CXXC-type zinc finger protein 2) (Protein-containing CXXC domain 2). | |||||
|
MBD1_HUMAN
|
||||||
| NC score | 0.436752 (rank : 13) | θ value | 7.84624e-12 (rank : 4) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9UIS9, O15248, O95241, Q7Z7B5, Q8N4W4, Q9UNZ6, Q9UNZ7, Q9UNZ8, Q9UNZ9 | Gene names | MBD1, PCM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 1 (Methyl-CpG-binding protein MBD1) (Protein containing methyl-CpG-binding domain 1). | |||||
|
MBD1_MOUSE
|
||||||
| NC score | 0.431709 (rank : 14) | θ value | 2.69671e-12 (rank : 3) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Z2E2, Q792D6, Q8CCL9, Q9DC19 | Gene names | Mbd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 1 (Methyl-CpG-binding protein MBD1). | |||||
|
ING1_HUMAN
|
||||||
| NC score | 0.377760 (rank : 15) | θ value | 1.09739e-05 (rank : 27) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9UK53, O00532, O43658, Q9H007, Q9HD98, Q9HD99, Q9NS83, Q9P0U6, Q9UBC6, Q9UIJ1, Q9UIJ2, Q9UIJ3, Q9UIJ4, Q9UK52 | Gene names | ING1 | |||
|
Domain Architecture |

|
|||||
| Description | Inhibitor of growth protein 1. | |||||
|
ING1_MOUSE
|
||||||
| NC score | 0.377124 (rank : 16) | θ value | 1.43324e-05 (rank : 28) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9QXV3, Q9QUP8, Q9QXV4, Q9QZX3 | Gene names | Ing1 | |||
|
Domain Architecture |

|
|||||
| Description | Inhibitor of growth protein 1. | |||||
|
ING2_MOUSE
|
||||||
| NC score | 0.361926 (rank : 17) | θ value | 8.40245e-06 (rank : 26) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9ESK4, Q80VI5, Q8BGU8 | Gene names | Ing2, Ing1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 2 (p33ING2) (Inhibitor of growth 1-like protein). | |||||
|
ING2_HUMAN
|
||||||
| NC score | 0.358415 (rank : 18) | θ value | 1.87187e-05 (rank : 29) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9H160, O95698 | Gene names | ING2, ING1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 2 (p33ING2) (Inhibitor of growth 1-like protein) (ING1Lp) (p32). | |||||
|
FXL19_HUMAN
|
||||||
| NC score | 0.353533 (rank : 19) | θ value | 2.61198e-07 (rank : 15) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q6PCT2, Q8N789, Q9NT14 | Gene names | FBXL19, FBL19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 19 (F-box and leucine-rich repeat protein 19). | |||||
|
FXL19_MOUSE
|
||||||
| NC score | 0.348118 (rank : 20) | θ value | 2.61198e-07 (rank : 16) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q6PB97, Q7TSH0, Q8BIB4 | Gene names | Fbxl19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 19 (F-box and leucine-rich repeat protein 19). | |||||
|
TAF3_HUMAN
|
||||||
| NC score | 0.348086 (rank : 21) | θ value | 2.88788e-06 (rank : 25) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 568 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q5VWG9, Q6GMS5, Q6P6B5, Q86VY6, Q9BQS9, Q9UFI8 | Gene names | TAF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
TAF3_MOUSE
|
||||||
| NC score | 0.347500 (rank : 22) | θ value | 1.69304e-06 (rank : 24) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 463 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q5HZG4, Q3U490, Q3UWX2, Q8BIU8, Q99JH4 | Gene names | Taf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
ING3_HUMAN
|
||||||
| NC score | 0.335821 (rank : 23) | θ value | 0.00020696 (rank : 30) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9NXR8, O60394, Q6GMT3, Q7Z762, Q969G0, Q96DT4, Q9HC99, Q9P081 | Gene names | ING3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 3 (p47ING3 protein). | |||||
|
ING4_HUMAN
|
||||||
| NC score | 0.334227 (rank : 24) | θ value | 0.000270298 (rank : 31) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9UNL4, Q96E15, Q9H3J0 | Gene names | ING4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 4 (p29ING4). | |||||
|
ING4_MOUSE
|
||||||
| NC score | 0.332521 (rank : 25) | θ value | 0.000270298 (rank : 32) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8C0D7, Q8C1S7, Q8K3Q5, Q8K3Q6, Q8K3Q7, Q9D7F9 | Gene names | Ing4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 4 (p29ING4). | |||||
|
PHF3_HUMAN
|
||||||
| NC score | 0.330522 (rank : 26) | θ value | 5.26297e-08 (rank : 14) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q92576, Q5T1T6, Q9NQ16, Q9UI45 | Gene names | PHF3, KIAA0244 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 3. | |||||
|
ING3_MOUSE
|
||||||
| NC score | 0.326789 (rank : 27) | θ value | 0.000602161 (rank : 33) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8VEK6, Q99JS6, Q9ERB2 | Gene names | Ing3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 3 (p47ING3 protein). | |||||
|
DNMT1_MOUSE
|
||||||
| NC score | 0.325879 (rank : 28) | θ value | 9.92553e-07 (rank : 21) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P13864, P97413, Q80ZU3, Q9CSC6, Q9QXX6 | Gene names | Dnmt1, Dnmt, Met1, Uim | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 1 (EC 2.1.1.37) (Dnmt1) (DNA methyltransferase MmuI) (DNA MTase MmuI) (MCMT) (M.MmuI) (Met-1). | |||||
|
DIDO1_HUMAN
|
||||||
| NC score | 0.320208 (rank : 29) | θ value | 2.52405e-10 (rank : 7) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
INGX_HUMAN
|
||||||
| NC score | 0.317975 (rank : 30) | θ value | 0.125558 (rank : 52) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9P0U5 | Gene names | INGX, ING2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein, X-linked (Inhibitor of growth protein 2) (ING1-like tumor suppressor protein). | |||||
|
ING5_HUMAN
|
||||||
| NC score | 0.314433 (rank : 31) | θ value | 0.000602161 (rank : 34) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8WYH8, Q9BS30 | Gene names | ING5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 5 (p28ING5). | |||||
|
ING5_MOUSE
|
||||||
| NC score | 0.312977 (rank : 32) | θ value | 0.000602161 (rank : 35) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9D8Y8, Q3UL57, Q6P292, Q9CV64, Q9D9V8 | Gene names | Ing5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inhibitor of growth protein 5. | |||||
|
DNMT1_HUMAN
|
||||||
| NC score | 0.309588 (rank : 33) | θ value | 5.81887e-07 (rank : 20) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P26358, Q9UHG5, Q9ULA2, Q9UMZ6 | Gene names | DNMT1, AIM, DNMT | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 1 (EC 2.1.1.37) (Dnmt1) (DNA methyltransferase HsaI) (DNA MTase HsaI) (MCMT) (M.HsaI). | |||||
|
BPTF_HUMAN
|
||||||
| NC score | 0.295613 (rank : 34) | θ value | 4.76016e-09 (rank : 12) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 786 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q12830, Q6NX67, Q7Z7D6, Q9UIG2 | Gene names | FALZ, BPTF, FAC1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleosome remodeling factor subunit BPTF (Bromodomain and PHD finger- containing transcription factor) (Fetal Alzheimer antigen) (Fetal Alz- 50 clone 1 protein). | |||||
|
PHF13_HUMAN
|
||||||
| NC score | 0.289500 (rank : 35) | θ value | 0.00102713 (rank : 36) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q86YI8, Q8N551, Q9UJP2 | Gene names | PHF13 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 13. | |||||
|
PHF13_MOUSE
|
||||||
| NC score | 0.287501 (rank : 36) | θ value | 0.00175202 (rank : 37) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8K2W6 | Gene names | Phf13 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 13. | |||||
|
MLL4_HUMAN
|
||||||
| NC score | 0.280906 (rank : 37) | θ value | 2.61198e-07 (rank : 17) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9UMN6, O15022, O95836, Q96GP2, Q96IP3, Q9UK25, Q9Y668, Q9Y669 | Gene names | MLL4, HRX2, KIAA0304, MLL2, TRX2 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 4 (Trithorax homolog 2). | |||||
|
HRX_HUMAN
|
||||||
| NC score | 0.280842 (rank : 38) | θ value | 4.0297e-08 (rank : 13) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
CN130_MOUSE
|
||||||
| NC score | 0.220478 (rank : 39) | θ value | 0.0961366 (rank : 48) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8BU04, Q80UT9 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C14orf130 homolog. | |||||
|
CN130_HUMAN
|
||||||
| NC score | 0.220311 (rank : 40) | θ value | 0.0431538 (rank : 45) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8N806, Q86U21, Q86UA9, Q96BY0, Q9NVV6 | Gene names | C14orf130 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C14orf130. | |||||
|
JAD1A_HUMAN
|
||||||
| NC score | 0.214745 (rank : 41) | θ value | 0.00509761 (rank : 39) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | P29375 | Gene names | JARID1A, RBBP2, RBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1A (Retinoblastoma-binding protein 2) (RBBP-2). | |||||
|
TCF19_HUMAN
|
||||||
| NC score | 0.199045 (rank : 42) | θ value | 0.365318 (rank : 64) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9Y242, Q13176, Q15967, Q5SQ89, Q5STD6, Q5STF5, Q9BUM2, Q9UBH7 | Gene names | TCF19, SC1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 19 (Transcription factor SC1). | |||||
|
PHF20_MOUSE
|
||||||
| NC score | 0.198716 (rank : 43) | θ value | 0.0252991 (rank : 43) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8BLG0, Q8BMA2, Q8BYR4, Q921N1 | Gene names | Phf20, Hca58 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58 homolog). | |||||
|
PHF20_HUMAN
|
||||||
| NC score | 0.198527 (rank : 44) | θ value | 0.0252991 (rank : 42) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9BVI0, Q5JWY9, Q9BWV4, Q9BXA3, Q9BZW3, Q9H421, Q9H4J6, Q9NZ22 | Gene names | PHF20, C20orf104, GLEA2, HCA58, NZF, TZP | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58) (Glioma-expressed antigen 2) (Transcription factor TZP) (Novel zinc finger protein). | |||||
|
CXXC6_HUMAN
|
||||||
| NC score | 0.192502 (rank : 45) | θ value | 0.0252991 (rank : 41) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8NFU7, Q5VUP7, Q7Z6B6, Q8TCR1, Q9C0I7 | Gene names | CXXC6, KIAA1676, LCX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CXXC-type zinc finger protein 6 (Leukemia-associated protein with a CXXC domain). | |||||
|
BAZ1A_HUMAN
|
||||||
| NC score | 0.187522 (rank : 46) | θ value | 0.00228821 (rank : 38) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 693 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9NRL2, Q9NZ15, Q9P065, Q9UIG1, Q9Y3V3 | Gene names | BAZ1A, ACF1, WCRF180 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1A (ATP-utilizing chromatin assembly and remodeling factor 1) (hACF1) (ATP-dependent chromatin remodelling protein) (Williams syndrome transcription factor-related chromatin remodeling factor 180) (WCRF180) (hWALp1) (CHRAC subunit ACF1). | |||||
|
JAD1D_HUMAN
|
||||||
| NC score | 0.173940 (rank : 47) | θ value | 0.125558 (rank : 53) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q9BY66, Q92509, Q92809, Q9HCU1 | Gene names | SMCY, HY, HYA, JARID1D, KIAA0234 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1D (SmcY protein) (Histocompatibility Y antigen) (H-Y). | |||||
|
BAZ1B_MOUSE
|
||||||
| NC score | 0.164534 (rank : 48) | θ value | 0.125558 (rank : 51) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9Z277, Q9CU68 | Gene names | Baz1b, Wbscr9 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein homolog) (WBRS9). | |||||
|
JAD1D_MOUSE
|
||||||
| NC score | 0.161112 (rank : 49) | θ value | 0.0736092 (rank : 46) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q62240, Q9QVR9, Q9R040 | Gene names | Smcy, Hya, Jarid1d | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1D (SmcY protein) (Histocompatibility Y antigen) (H-Y). | |||||
|
JAD1C_HUMAN
|
||||||
| NC score | 0.160871 (rank : 50) | θ value | 0.365318 (rank : 60) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P41229 | Gene names | SMCX, DXS1272E, JARID1C, XE169 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1C (Protein SmcX) (Protein Xe169). | |||||
|
JAD1C_MOUSE
|
||||||
| NC score | 0.160529 (rank : 51) | θ value | 0.365318 (rank : 61) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P41230, O54995, Q9CVI4, Q9D0C3, Q9QVR8, Q9R039 | Gene names | Smcx, Jarid1c, Xe169 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1C (SmcX protein) (Xe169 protein). | |||||
|
BAZ1B_HUMAN
|
||||||
| NC score | 0.159312 (rank : 52) | θ value | 0.21417 (rank : 56) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 701 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q9UIG0, O95039, O95247, O95277 | Gene names | BAZ1B, WBSC10, WBSCR9, WSTF | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein) (Williams syndrome transcription factor) (hWALP2). | |||||
|
MTF2_HUMAN
|
||||||
| NC score | 0.157394 (rank : 53) | θ value | 1.06291 (rank : 68) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9Y483, Q9UES9, Q9UP40 | Gene names | MTF2 | |||
|
Domain Architecture |

|
|||||
| Description | Metal-response element-binding transcription factor 2 (Metal-response element DNA-binding protein M96) (Metal-regulatory transcription factor 2) (PCL2). | |||||
|
MTF2_MOUSE
|
||||||
| NC score | 0.156445 (rank : 54) | θ value | 1.06291 (rank : 69) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q02395, Q569Z8, Q6PG89, Q8BGP9, Q8BSJ7 | Gene names | Mtf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metal-response element-binding transcription factor 2 (Zinc-regulated factor 1) (ZiRF1) (Metal-response element DNA-binding protein M96) (Metal-regulatory transcription factor 2) (PCL2). | |||||
|
BAZ2A_MOUSE
|
||||||
| NC score | 0.153533 (rank : 55) | θ value | 0.0148317 (rank : 40) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q91YE5 | Gene names | Baz2a, Tip5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5). | |||||
|
BAZ2A_HUMAN
|
||||||
| NC score | 0.150675 (rank : 56) | θ value | 0.0961366 (rank : 47) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 780 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q9UIF9, O00536, O15030, Q96H26 | Gene names | BAZ2A, KIAA0314, TIP5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5) (hWALp3). | |||||
|
PHF1_HUMAN
|
||||||
| NC score | 0.147804 (rank : 57) | θ value | 0.125558 (rank : 54) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O43189, O60929, Q96KM7 | Gene names | PHF1 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 1 (Protein PHF1). | |||||
|
PHF1_MOUSE
|
||||||
| NC score | 0.147706 (rank : 58) | θ value | 0.125558 (rank : 55) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9Z1B8, O54808 | Gene names | Phf1, Tctex-3, Tctex3 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 1 (Protein PHF1) (T-complex testis-expressed 3) (Polycomblike 1) (mPCl1). | |||||
|
DPF1_HUMAN
|
||||||
| NC score | 0.143955 (rank : 59) | θ value | 0.0961366 (rank : 49) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q92782 | Gene names | DPF1, NEUD4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein neuro-d4 (D4, zinc and double PHD fingers family 1). | |||||
|
PHF10_HUMAN
|
||||||
| NC score | 0.143347 (rank : 60) | θ value | θ > 10 (rank : 154) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q8WUB8, Q2YDA3, Q9BXD2, Q9NV26 | Gene names | PHF10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 10 (XAP135). | |||||
|
PHF10_MOUSE
|
||||||
| NC score | 0.140227 (rank : 61) | θ value | θ > 10 (rank : 155) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9D8M7, Q99LV5 | Gene names | Phf10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 10. | |||||
|
MLL2_HUMAN
|
||||||
| NC score | 0.140097 (rank : 62) | θ value | 0.279714 (rank : 57) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
UHRF1_HUMAN
|
||||||
| NC score | 0.140038 (rank : 63) | θ value | 0.0961366 (rank : 50) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 272 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q96T88, Q8J022, Q9H6S6, Q9P115, Q9P1U7 | Gene names | UHRF1, ICBP90, NP95, RNF106 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 1 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 1) (Inverted CCAAT box-binding protein of 90 kDa) (Transcription factor ICBP90) (Nuclear zinc finger protein Np95) (Nuclear protein 95) (HuNp95) (RING finger protein 106). | |||||
|
RSF1_HUMAN
|
||||||
| NC score | 0.139819 (rank : 64) | θ value | 0.279714 (rank : 58) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 539 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q96T23, Q86X86, Q9H3L8, Q9NVZ8, Q9NYU0 | Gene names | RSF1, HBXAP, XAP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Remodeling and spacing factor 1 (Rsf-1) (Hepatitis B virus X- associated protein) (HBV pX-associated protein 8) (p325 subunit of RSF chromatin remodelling complex). | |||||
|
DPF1_MOUSE
|
||||||
| NC score | 0.128651 (rank : 65) | θ value | 1.06291 (rank : 66) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9QX66, Q9QX65, Q9QYA4 | Gene names | Dpf1, Neud4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein neuro-d4 (D4, zinc and double PHD fingers family 1). | |||||
|
PF21A_MOUSE
|
||||||
| NC score | 0.127107 (rank : 66) | θ value | 5.27518 (rank : 107) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q6ZPK0, Q6XVG0, Q80Z33, Q8CAZ4, Q8VEC8 | Gene names | Phf21a, Bhc80, Kiaa1696, Pftf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (mBHC80) (BHC80a). | |||||
|
REQU_HUMAN
|
||||||
| NC score | 0.125535 (rank : 67) | θ value | 3.0926 (rank : 84) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 523 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q92785 | Gene names | DPF2, REQ, UBID4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein ubi-d4 (Requiem) (Apoptosis response zinc finger protein) (D4, zinc and double PHD fingers family 2). | |||||
|
MLL3_MOUSE
|
||||||
| NC score | 0.124156 (rank : 68) | θ value | 4.03905 (rank : 93) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
BAZ2B_HUMAN
|
||||||
| NC score | 0.122993 (rank : 69) | θ value | 0.365318 (rank : 59) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 803 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q9UIF8, Q96EA1, Q96SQ8, Q9P252, Q9Y4N8 | Gene names | BAZ2B, KIAA1476 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2B (hWALp4). | |||||
|
REQU_MOUSE
|
||||||
| NC score | 0.122248 (rank : 70) | θ value | 3.0926 (rank : 85) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q61103, Q3UNP5, Q60663, Q9QYA3 | Gene names | Dpf2, Req, Ubid4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein ubi-d4 (Requiem) (Apoptosis response zinc finger protein) (D4, zinc and double PHD fingers family 2). | |||||
|
UHRF1_MOUSE
|
||||||
| NC score | 0.121286 (rank : 71) | θ value | 3.0926 (rank : 88) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8VDF2, Q8C6F1, Q8VIA1, Q9Z1H6 | Gene names | Uhrf1, Np95 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 1 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 1) (Nuclear zinc finger protein Np95) (Nuclear protein 95). | |||||
|
PKCB1_HUMAN
|
||||||
| NC score | 0.119539 (rank : 72) | θ value | 2.36792 (rank : 80) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 488 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q9ULU4, Q13517, Q4JJ94, Q4JJ95, Q5TH09, Q6MZM1, Q8WXC5, Q9H1F3, Q9H1F4, Q9H1F5, Q9H1L8, Q9H1L9, Q9H2G5, Q9NYN3, Q9UIX6 | Gene names | PRKCBP1, KIAA1125, RACK7, ZMYND8 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C-binding protein 1 (Rack7) (Cutaneous T-cell lymphoma- associated antigen se14-3) (CTCL tumor antigen se14-3) (Zinc finger MYND domain-containing protein 8). | |||||
|
MLL3_HUMAN
|
||||||
| NC score | 0.118424 (rank : 73) | θ value | 3.0926 (rank : 82) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
MYST4_HUMAN
|
||||||
| NC score | 0.116846 (rank : 74) | θ value | 1.06291 (rank : 70) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q8WYB5, O15087, Q86Y05, Q8WU81, Q9UKW2, Q9UKW3, Q9UKX0 | Gene names | MYST4, KIAA0383, MORF, MOZ2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Histone acetyltransferase MOZ2) (Monocytic leukemia zinc finger protein- related factor) (Histone acetyltransferase MORF). | |||||
|
PF21A_HUMAN
|
||||||
| NC score | 0.115054 (rank : 75) | θ value | θ > 10 (rank : 151) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q96BD5, Q6AWA2, Q9C0G7, Q9H8V9, Q9HAK6, Q9NZE9 | Gene names | PHF21A, BHC80, KIAA1696 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (BHC80a). | |||||
|
MYST4_MOUSE
|
||||||
| NC score | 0.113656 (rank : 76) | θ value | 1.38821 (rank : 73) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q8BRB7, Q7TNW5, Q8BG35, Q8C441, Q9JKX5 | Gene names | Myst4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Querkopf protein). | |||||
|
NSD1_MOUSE
|
||||||
| NC score | 0.113586 (rank : 77) | θ value | 5.27518 (rank : 106) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | O88491, Q8C480, Q9CT70 | Gene names | Nsd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein). | |||||
|
MYST3_HUMAN
|
||||||
| NC score | 0.107873 (rank : 78) | θ value | 3.0926 (rank : 83) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q92794 | Gene names | MYST3, MOZ, RUNXBP2, ZNF220 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Runt-related transcription factor-binding protein 2) (Monocytic leukemia zinc finger protein) (Zinc finger protein 220). | |||||
|
NSD1_HUMAN
|
||||||
| NC score | 0.105984 (rank : 79) | θ value | 5.27518 (rank : 105) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q96L73, Q96PD8, Q96RN7 | Gene names | NSD1, ARA267 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein) (Androgen receptor-associated coregulator 267). | |||||
|
MYST3_MOUSE
|
||||||
| NC score | 0.100792 (rank : 80) | θ value | θ > 10 (rank : 150) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 688 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q8BZ21, Q8BW52, Q8BYH1, Q8C1F3 | Gene names | Myst3, Moz | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Monocytic leukemia zinc finger protein) (Monocytic leukemia zinc finger homolog). | |||||
|
PF21B_MOUSE
|
||||||
| NC score | 0.098246 (rank : 81) | θ value | θ > 10 (rank : 153) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q8C966 | Gene names | Phf21b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
AIRE_HUMAN
|
||||||
| NC score | 0.096325 (rank : 82) | θ value | θ > 10 (rank : 125) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O43918, O43922, O43932, O75745 | Gene names | AIRE, APECED | |||
|
Domain Architecture |

|
|||||
| Description | Autoimmune regulator (Autoimmune polyendocrinopathy candidiasis ectodermal dystrophy protein) (APECED protein). | |||||
|
AIRE_MOUSE
|
||||||
| NC score | 0.095112 (rank : 83) | θ value | θ > 10 (rank : 126) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9Z0E3, Q9JLW0, Q9JLW1, Q9JLW2, Q9JLW3, Q9JLW4, Q9JLW5, Q9JLW6, Q9JLW7, Q9JLW8, Q9JLW9, Q9JLX0 | Gene names | Aire | |||
|
Domain Architecture |

|
|||||
| Description | Autoimmune regulator (Autoimmune polyendocrinopathy candidiasis ectodermal dystrophy protein homolog) (APECED protein homolog). | |||||
|
UHRF2_HUMAN
|
||||||
| NC score | 0.093642 (rank : 84) | θ value | θ > 10 (rank : 176) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q96PU4, Q5VYR1, Q5VYR3, Q659C8, Q8TAG7 | Gene names | UHRF2, NIRF, RNF107 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 2 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 2) (Np95/ICBP90-like RING finger protein) (Np95-like RING finger protein) (Nuclear zinc finger protein Np97) (RING finger protein 107). | |||||
|
UHRF2_MOUSE
|
||||||
| NC score | 0.092447 (rank : 85) | θ value | θ > 10 (rank : 177) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q7TMI3, Q8BG56, Q8BJP6, Q8BY30, Q8K1I5 | Gene names | Uhrf2, Nirf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 2 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 2) (Np95-like ring finger protein) (Nuclear zinc finger protein Np97) (NIRF). | |||||
|
PF21B_HUMAN
|
||||||
| NC score | 0.092390 (rank : 86) | θ value | θ > 10 (rank : 152) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q96EK2, Q5TFL2, Q6ICC0 | Gene names | PHF21B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
C8AP2_HUMAN
|
||||||
| NC score | 0.085144 (rank : 87) | θ value | 0.0330416 (rank : 44) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 548 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9UKL3, Q6PH76, Q7LCQ7, Q86YD9, Q9NUQ4, Q9NZV9, Q9P2N1, Q9Y563 | Gene names | CASP8AP2, FLASH, KIAA1315, RIP25 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
PTDSR_MOUSE
|
||||||
| NC score | 0.080886 (rank : 88) | θ value | θ > 10 (rank : 164) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9ERI5, Q80TX1 | Gene names | Ptdsr, Kiaa0585 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein PTDSR (Phosphatidylserine receptor) (Apoptotic cell clearance receptor PtdSerR). | |||||
|
PHF12_HUMAN
|
||||||
| NC score | 0.079323 (rank : 89) | θ value | θ > 10 (rank : 157) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q96QT6, Q6ZML2, Q9BV34, Q9H7U9, Q9P205 | Gene names | PHF12, KIAA1523 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 12 (PHD factor 1) (Pf1). | |||||
|
PTDSR_HUMAN
|
||||||
| NC score | 0.078618 (rank : 90) | θ value | θ > 10 (rank : 163) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q6NYC1, Q86VY0, Q8IUM5, Q9Y4E2 | Gene names | PTDSR, KIAA0585 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein PTDSR (Phosphatidylserine receptor). | |||||
|
PHF12_MOUSE
|
||||||
| NC score | 0.077065 (rank : 91) | θ value | θ > 10 (rank : 158) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q5SPL2, Q5SPL3, Q6ZPN8, Q80W50 | Gene names | Phf12, Kiaa1523 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 12 (PHD factor 1) (Pf1). | |||||
|
TRI66_MOUSE
|
||||||
| NC score | 0.072902 (rank : 92) | θ value | θ > 10 (rank : 175) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q924W6 | Gene names | Trim66 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
TIF1A_MOUSE
|
||||||
| NC score | 0.072897 (rank : 93) | θ value | θ > 10 (rank : 171) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q64127, Q64126 | Gene names | Trim24, Tif1, Tif1a | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-alpha (TIF1-alpha) (Tripartite motif-containing protein 24). | |||||
|
PHF14_MOUSE
|
||||||
| NC score | 0.072378 (rank : 94) | θ value | θ > 10 (rank : 160) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9D4H9, Q810Z6, Q8C284, Q8CGF8, Q9D1Q8 | Gene names | Phf14 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 14. | |||||
|
PHF14_HUMAN
|
||||||
| NC score | 0.071740 (rank : 95) | θ value | θ > 10 (rank : 159) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | O94880 | Gene names | PHF14, KIAA0783 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 14. | |||||
|
SET1A_HUMAN
|
||||||
| NC score | 0.071045 (rank : 96) | θ value | θ > 10 (rank : 165) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O15047, Q6PIF3, Q8TAJ6 | Gene names | SETD1A, KIAA0339, SET1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET1 (EC 2.1.1.43) (Set1/Ash2 histone methyltransferase complex subunit SET1) (SET-domain-containing protein 1A). | |||||
|
TIF1A_HUMAN
|
||||||
| NC score | 0.069344 (rank : 97) | θ value | θ > 10 (rank : 170) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O15164, O95854 | Gene names | TRIM24, RNF82, TIF1, TIF1A | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-alpha (TIF1-alpha) (Tripartite motif-containing protein 24) (RING finger protein 82). | |||||
|
JADE3_HUMAN
|
||||||
| NC score | 0.068119 (rank : 98) | θ value | 6.88961 (rank : 110) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q92613, Q6IE79 | Gene names | PHF16, JADE3, KIAA0215 | |||
|
Domain Architecture |

|
|||||
| Description | Protein Jade-3 (PHD finger protein 16). | |||||
|
JADE3_MOUSE
|
||||||
| NC score | 0.066477 (rank : 99) | θ value | 6.88961 (rank : 111) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q6IE82, Q6ZQG2 | Gene names | Phf16, Jade3, Kiaa0215 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-3 (PHD finger protein 16). | |||||
|
TRI66_HUMAN
|
||||||
| NC score | 0.065915 (rank : 100) | θ value | θ > 10 (rank : 174) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | O15016, Q9BQQ4 | Gene names | TRIM66, KIAA0298 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
CHD5_HUMAN
|
||||||
| NC score | 0.065849 (rank : 101) | θ value | θ > 10 (rank : 133) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q8TDI0, O75032, Q5TG89, Q9UFR9 | Gene names | CHD5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain helicase-DNA-binding protein 5 (EC 3.6.1.-) (ATP- dependent helicase CHD5) (CHD-5). | |||||
|
CHD4_MOUSE
|
||||||
| NC score | 0.064076 (rank : 102) | θ value | θ > 10 (rank : 132) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q6PDQ2 | Gene names | Chd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain helicase-DNA-binding protein 4 (CHD-4). | |||||
|
FBXL7_HUMAN
|
||||||
| NC score | 0.063910 (rank : 103) | θ value | θ > 10 (rank : 136) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UJT9, O94926 | Gene names | FBXL7, FBL6, FBL7, KIAA0840 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 7 (F-box and leucine-rich repeat protein 7) (F-box protein FBL6/FBL7). | |||||
|
CHD4_HUMAN
|
||||||
| NC score | 0.063282 (rank : 104) | θ value | θ > 10 (rank : 131) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 365 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q14839, Q8IXZ5 | Gene names | CHD4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain helicase-DNA-binding protein 4 (EC 3.6.1.-) (ATP- dependent helicase CHD4) (CHD-4) (Mi-2 autoantigen 218 kDa protein) (Mi2-beta). | |||||
|
CHD3_HUMAN
|
||||||
| NC score | 0.063002 (rank : 105) | θ value | θ > 10 (rank : 130) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q12873, Q9Y4I0 | Gene names | CHD3 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain helicase-DNA-binding protein 3 (EC 3.6.1.-) (ATP- dependent helicase CHD3) (CHD-3) (Mi-2 autoantigen 240 kDa protein) (Mi2-alpha) (Zinc-finger helicase) (hZFH). | |||||
|
LRC29_HUMAN
|
||||||
| NC score | 0.059475 (rank : 106) | θ value | θ > 10 (rank : 147) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8WV35, Q9UKA0 | Gene names | LRRC29, FBL9, FBXL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 29 (F-box/LRR-repeat protein 9) (F-box and leucine-rich repeat protein 9) (F-box protein FBL9). | |||||
|
BRPF1_HUMAN
|
||||||
| NC score | 0.059419 (rank : 107) | θ value | θ > 10 (rank : 128) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P55201, Q9UHI0 | Gene names | BRPF1, BR140 | |||
|
Domain Architecture |

|
|||||
| Description | Peregrin (Bromodomain and PHD finger-containing protein 1) (BR140 protein). | |||||
|
SKP2_HUMAN
|
||||||
| NC score | 0.058290 (rank : 108) | θ value | θ > 10 (rank : 168) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q13309, Q8TDZ0, Q8TDZ1, Q9BV69 | Gene names | SKP2, FBXL1 | |||
|
Domain Architecture |

|
|||||
| Description | S-phase kinase-associated protein 2 (F-box protein Skp2) (Cyclin A/CDK2-associated protein p45) (p45skp2) (F-box/LRR-repeat protein 1). | |||||
|
BRD1_HUMAN
|
||||||
| NC score | 0.057388 (rank : 109) | θ value | θ > 10 (rank : 127) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O95696 | Gene names | BRD1, BRL, BRPF2 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 1 (BR140-like protein). | |||||
|
FXL20_HUMAN
|
||||||
| NC score | 0.057226 (rank : 110) | θ value | θ > 10 (rank : 141) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96IG2 | Gene names | FBXL20, FBL2 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 20 (F-box and leucine-rich repeat protein 20) (F-box/LRR-repeat protein 2-like). | |||||
|
FXL20_MOUSE
|
||||||
| NC score | 0.057115 (rank : 111) | θ value | θ > 10 (rank : 142) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9CZV8, Q8BZ95 | Gene names | Fbxl20, Fbl2 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 20 (F-box and leucine-rich repeat protein 20) (F-box/LRR-repeat protein 2-like). | |||||
|
CECR2_HUMAN
|
||||||
| NC score | 0.056728 (rank : 112) | θ value | θ > 10 (rank : 129) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9BXF3, Q658Z4, Q96P58, Q9C0C3 | Gene names | CECR2, KIAA1740 | |||
|
Domain Architecture |

|
|||||
| Description | Cat eye syndrome critical region protein 2. | |||||
|
SKP2_MOUSE
|
||||||
| NC score | 0.056307 (rank : 113) | θ value | θ > 10 (rank : 169) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Z0Z3, Q8BSG7, Q8C8Y9, Q8CB22, Q99J53 | Gene names | Skp2 | |||
|
Domain Architecture |

|
|||||
| Description | S-phase kinase-associated protein 2 (F-box protein Skp2) (Cyclin A/CDK2-associated protein p45) (F-box/WD-40 protein 1) (FWD1). | |||||
|
PHF22_MOUSE
|
||||||
| NC score | 0.056304 (rank : 114) | θ value | θ > 10 (rank : 162) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9D168, Q3U2Q5, Q921U2 | Gene names | Ints12, Phf22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integrator complex subunit 12 (PHD finger protein 22). | |||||
|
FXL16_HUMAN
|
||||||
| NC score | 0.056175 (rank : 115) | θ value | θ > 10 (rank : 140) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8N461, Q2MHR2, Q96S14, Q9UJI0 | Gene names | FBXL16, C16orf22, FBL16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 16 (F-box and leucine-rich repeat protein 16). | |||||
|
JADE2_MOUSE
|
||||||
| NC score | 0.056068 (rank : 116) | θ value | θ > 10 (rank : 146) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q6ZQF7, Q3UHD5, Q6IE83 | Gene names | Phf15, Jade2, Kiaa0239 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-2 (PHD finger protein 15). | |||||
|
SETD8_HUMAN
|
||||||
| NC score | 0.055921 (rank : 117) | θ value | θ > 10 (rank : 166) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9NQR1, Q86W83, Q8TD09 | Gene names | SETD8, PRSET7, SET07, SET8 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H4 lysine-20 specific (EC 2.1.1.43) (Histone H4-K20 methyltransferase) (H4-K20-HMTase) (SET domain-containing lysine methyltransferase 8) (SET domain-containing protein 8) (PR/SET domain-containing protein 07) (PR/SET07) (PR-Set7). | |||||
|
FXL13_MOUSE
|
||||||
| NC score | 0.055679 (rank : 118) | θ value | θ > 10 (rank : 137) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CDU4, Q8CDE9 | Gene names | Fbxl13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 13 (F-box and leucine-rich repeat protein 13). | |||||
|
ZMY11_HUMAN
|
||||||
| NC score | 0.055512 (rank : 119) | θ value | θ > 10 (rank : 178) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q15326 | Gene names | ZMYND11, BS69 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger MYND domain-containing protein 11 (Adenovirus 5 E1A- binding protein) (BS69 protein). | |||||
|
RBBP6_MOUSE
|
||||||
| NC score | 0.054590 (rank : 120) | θ value | 1.81305 (rank : 76) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
JADE1_HUMAN
|
||||||
| NC score | 0.054438 (rank : 121) | θ value | θ > 10 (rank : 143) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q6IE81, Q4W5D5, Q6ZSL7, Q8NC41, Q96JL8, Q96SQ1, Q9H692 | Gene names | PHF17, JADE1, KIAA1807 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-1 (PHD finger protein 17). | |||||
|
FXL14_MOUSE
|
||||||
| NC score | 0.054146 (rank : 122) | θ value | θ > 10 (rank : 139) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BID8, Q3U2Q9, Q8R5H7, Q8VDT7, Q922N5 | Gene names | Fbxl14, Fbl14, Ppa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 14 (F-box and leucine-rich repeat protein 14). | |||||
|
JADE1_MOUSE
|
||||||
| NC score | 0.054017 (rank : 123) | θ value | θ > 10 (rank : 144) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q6ZPI0, Q6IE84, Q8C7S6, Q8CFM2, Q8R362 | Gene names | Phf17, Jade1, Kiaa1807 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-1 (PHD finger protein 17). | |||||
|
FXL14_HUMAN
|
||||||
| NC score | 0.053993 (rank : 124) | θ value | θ > 10 (rank : 138) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8N1E6 | Gene names | FBXL14, FBL14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 14 (F-box and leucine-rich repeat protein 14). | |||||
|
TIF1G_MOUSE
|
||||||
| NC score | 0.053783 (rank : 125) | θ value | θ > 10 (rank : 173) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q99PP7, Q6SI71, Q6ZPX5 | Gene names | Trim33, Kiaa1113 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription intermediary factor 1-gamma (TIF1-gamma) (Tripartite motif-containing protein 33). | |||||
|
PHF11_HUMAN
|
||||||
| NC score | 0.053664 (rank : 126) | θ value | θ > 10 (rank : 156) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UIL8, Q9Y5A2 | Gene names | PHF11, BCAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 11 (BRCA1 C-terminus-associated protein) (NY-REN-34 antigen). | |||||
|
SETD8_MOUSE
|
||||||
| NC score | 0.053229 (rank : 127) | θ value | θ > 10 (rank : 167) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q2YDW7, Q8C0J9 | Gene names | SETD8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H4 lysine-20 specific (EC 2.1.1.43) (Histone H4-K20 methyltransferase) (H4-K20-HMTase) (SET domain-containing lysine methyltransferase 8) (SET domain-containing protein 8). | |||||
|
FBXL2_MOUSE
|
||||||
| NC score | 0.053194 (rank : 128) | θ value | θ > 10 (rank : 135) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BH16, Q8BXM4 | Gene names | Fbxl2, Fbl2 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 2 (F-box and leucine-rich repeat protein 2). | |||||
|
ZMY11_MOUSE
|
||||||
| NC score | 0.052493 (rank : 129) | θ value | θ > 10 (rank : 179) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8R5C8, Q8C155 | Gene names | Zmynd11 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger MYND domain-containing protein 11. | |||||
|
MBD2_MOUSE
|
||||||
| NC score | 0.051991 (rank : 130) | θ value | θ > 10 (rank : 149) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Z2E1, Q811D9, Q9Z2D9 | Gene names | Mbd2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 2 (Methyl-CpG-binding protein MBD2). | |||||
|
MBD2_HUMAN
|
||||||
| NC score | 0.051343 (rank : 131) | θ value | θ > 10 (rank : 148) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UBB5, O95242, Q9UIS8 | Gene names | MBD2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 2 (Methyl-CpG-binding protein MBD2) (Demethylase) (DMTase). | |||||
|
TIF1G_HUMAN
|
||||||
| NC score | 0.050823 (rank : 132) | θ value | θ > 10 (rank : 172) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 410 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9UPN9, O95855, Q9C017, Q9UJ79 | Gene names | TRIM33, KIAA1113, RFG7, TIF1G | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-gamma (TIF1-gamma) (Tripartite motif-containing protein 33) (RET-fused gene 7 protein) (Rfg7 protein). | |||||
|
PHF22_HUMAN
|
||||||
| NC score | 0.050743 (rank : 133) | θ value | θ > 10 (rank : 161) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q96CB8, Q9HD71 | Gene names | INTS12, PHF22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integrator complex subunit 12 (PHD finger protein 22). | |||||
|
FBXL2_HUMAN
|
||||||
| NC score | 0.050707 (rank : 134) | θ value | θ > 10 (rank : 134) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UKC9, Q9NVQ8, Q9UK27, Q9UKA5, Q9Y3Y9 | Gene names | FBXL2, FBL2, FBL3 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 2 (F-box and leucine-rich repeat protein 2) (F-box protein FBL2/FBL3). | |||||
|
JADE2_HUMAN
|
||||||
| NC score | 0.050686 (rank : 135) | θ value | θ > 10 (rank : 145) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9NQC1, Q6IE80, Q8TEK0, Q92513, Q96GQ6 | Gene names | PHF15, JADE2, KIAA0239 | |||
|
Domain Architecture |

|
|||||
| Description | Protein Jade-2 (PHD finger protein 15). | |||||
|
RIMS2_MOUSE
|
||||||
| NC score | 0.049744 (rank : 136) | θ value | 0.365318 (rank : 63) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 376 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9EQZ7, Q8C433, Q8CCK2 | Gene names | Rims2, Rab3ip2, Rim2 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 2 (Rab3-interacting molecule 2) (RIM 2) (Rab3-interacting protein 2). | |||||
|
RIMS2_HUMAN
|
||||||
| NC score | 0.048124 (rank : 137) | θ value | 0.365318 (rank : 62) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UQ26, O43413, Q86XL9, Q8IWV9, Q8IWW1 | Gene names | RIMS2, KIAA0751, RIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 2 (Rab3-interacting molecule 2) (RIM 2). | |||||
|
BRD2_HUMAN
|
||||||
| NC score | 0.042658 (rank : 138) | θ value | 8.99809 (rank : 116) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P25440, O00699, O00700, Q15310, Q5STC9, Q969U4 | Gene names | BRD2, KIAA9001, RING3 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 2 (Protein RING3) (O27.1.1). | |||||
|
RIMS1_HUMAN
|
||||||
| NC score | 0.035769 (rank : 139) | θ value | 1.06291 (rank : 71) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 437 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q86UR5, O15048, Q8TDY9, Q8TDZ5, Q9HBA1, Q9HBA2, Q9HBA3, Q9HBA4, Q9HBA5, Q9HBA6 | Gene names | RIMS1, KIAA0340, RIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 1 (Rab3-interacting molecule 1) (RIM 1). | |||||
|
K1045_HUMAN
|
||||||
| NC score | 0.035728 (rank : 140) | θ value | 1.81305 (rank : 75) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UPV7, Q58FE9, Q5T662 | Gene names | KIAA1045 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1045. | |||||
|
SYBU_MOUSE
|
||||||
| NC score | 0.032577 (rank : 141) | θ value | 3.0926 (rank : 86) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BHS8, Q3TQN7, Q401F3, Q401F4, Q571C1, Q6P1J2, Q80XH0, Q8BHS7, Q8BI27 | Gene names | Sybu, Golsyn, Kiaa1472 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntabulin (Syntaxin-1-binding protein) (Golgi-localized syntaphilin- related protein) (m-Golsyn). | |||||
|
NHS_HUMAN
|
||||||
| NC score | 0.032403 (rank : 142) | θ value | 5.27518 (rank : 104) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6T4R5, Q5J7Q0, Q5J7Q1, Q68DR5 | Gene names | NHS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nance-Horan syndrome protein (Congenital cataracts and dental anomalies protein). | |||||
|
MOR2A_MOUSE
|
||||||
| NC score | 0.029703 (rank : 143) | θ value | 4.03905 (rank : 94) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q69ZX6, Q5QNQ7, Q6P547 | Gene names | Morc2a, Kiaa0852, Zcwcc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORC family CW-type zinc finger protein 2A (Zinc finger CW-type coiled-coil domain protein 1). | |||||
|
IF3A_MOUSE
|
||||||
| NC score | 0.029649 (rank : 144) | θ value | 5.27518 (rank : 101) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 871 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P23116, Q60697, Q62162 | Gene names | Eif3s10, Csma, Eif3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 10 (eIF-3 theta) (eIF3 p167) (eIF3 p180) (eIF3 p185) (eIF3a) (p162 protein) (Centrosomin). | |||||
|
ZCHC7_HUMAN
|
||||||
| NC score | 0.029313 (rank : 145) | θ value | 6.88961 (rank : 115) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8N3Z6, Q5T0Q8, Q5T0Q9, Q5T0R0, Q8N2M1, Q8N4J2, Q8TBK8, Q9H648, Q9P0F0 | Gene names | ZCCHC7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 7. | |||||
|
IF3A_HUMAN
|
||||||
| NC score | 0.028543 (rank : 146) | θ value | 4.03905 (rank : 91) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 875 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q14152, O00653 | Gene names | EIF3S10, KIAA0139 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 10 (eIF-3 theta) (eIF3 p167) (eIF3 p180) (eIF3 p185) (eIF3a). | |||||
|
SMC1A_HUMAN
|
||||||
| NC score | 0.027516 (rank : 147) | θ value | 1.81305 (rank : 77) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1346 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q14683, O14995, Q16351 | Gene names | SMC1L1, DXS423E, KIAA0178, SB1.8, SMC1 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Sb1.8). | |||||
|
ZN598_MOUSE
|
||||||
| NC score | 0.027293 (rank : 148) | θ value | 1.38821 (rank : 74) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80YR4, Q6KAT0, Q8R3S1 | Gene names | Znf598, Zfp598 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 598. | |||||
|
PYGO1_HUMAN
|
||||||
| NC score | 0.026940 (rank : 149) | θ value | 8.99809 (rank : 123) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y3Y4 | Gene names | PYGO1 | |||
|
Domain Architecture |

|
|||||
| Description | Pygopus homolog 1. | |||||
|
TNIP3_HUMAN
|
||||||
| NC score | 0.026752 (rank : 150) | θ value | 4.03905 (rank : 98) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 511 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q96KP6, Q96PQ3, Q9H780 | Gene names | TNIP3, ABIN3, LIND | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TNFAIP3-interacting protein 3 (Listeria-induced gene protein) (A20- binding inhibitor of NF-kappa-B activation 3) (ABIN-3). | |||||
|
SMC1A_MOUSE
|
||||||
| NC score | 0.025917 (rank : 151) | θ value | 2.36792 (rank : 81) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1384 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9CU62, Q3V480, Q9CUX9, Q9D959, Q9WTU1 | Gene names | Smc1l1, Sb1.8, Smc1, Smcb | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Chromosome segregation protein SmcB) (Sb1.8). | |||||
|
NC6IP_MOUSE
|
||||||
| NC score | 0.024522 (rank : 152) | θ value | 2.36792 (rank : 79) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q923W1, Q6DI60, Q6PEA7, Q8R0W9 | Gene names | Ncoa6ip, Pimt | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative RNA methyltransferase NCOA6IP (EC 2.1.1.-) (Nuclear receptor coactivator 6-interacting protein) (PRIP-interacting protein with methyltransferase motif) (PIPMT) (PIMT). | |||||
|
SYBU_HUMAN
|
||||||
| NC score | 0.024258 (rank : 153) | θ value | 8.99809 (rank : 124) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9NX95, Q5R1T1, Q5R1T2, Q5R1T3, Q5Y2M6, Q8ND49, Q8TCR6, Q9P256 | Gene names | SYBU, GOLSYN, KIAA1472 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntabulin (Syntaxin-1-binding protein) (Golgi-localized syntaphilin- related protein). | |||||
|
EVPL_HUMAN
|
||||||
| NC score | 0.022550 (rank : 154) | θ value | 1.38821 (rank : 72) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 895 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q92817 | Gene names | EVPL | |||
|
Domain Architecture |

|
|||||
| Description | Envoplakin (210 kDa paraneoplastic pemphigus antigen) (p210) (210 kDa cornified envelope precursor protein). | |||||
|
CCD46_MOUSE
|
||||||
| NC score | 0.020838 (rank : 155) | θ value | 8.99809 (rank : 117) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q5PR68, Q80ZN6, Q99JS4, Q9D9T1 | Gene names | Ccdc46 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled coil domain-containing protein 46. | |||||
|
SULF2_MOUSE
|
||||||
| NC score | 0.020487 (rank : 156) | θ value | 4.03905 (rank : 97) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CFG0, Q8BM68, Q8BUZ4, Q8BX28, Q8BZ51, Q8C169, Q9D8E2 | Gene names | Sulf2 | |||
|
Domain Architecture |

|
|||||
| Description | Extracellular sulfatase Sulf-2 precursor (EC 3.1.6.-) (MSulf-2). | |||||
|
KI20A_HUMAN
|
||||||
| NC score | 0.020037 (rank : 157) | θ value | 1.06291 (rank : 67) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O95235 | Gene names | KIF20A, RAB6KIFL | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin family member 20A (Rabkinesin-6) (Rab6-interacting kinesin- like protein) (GG10_2). | |||||
|
TRHY_HUMAN
|
||||||
| NC score | 0.019982 (rank : 158) | θ value | 3.0926 (rank : 87) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1385 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q07283 | Gene names | THH, THL, TRHY | |||
|
Domain Architecture |

|
|||||
| Description | Trichohyalin. | |||||
|
FA47B_HUMAN
|
||||||
| NC score | 0.019789 (rank : 159) | θ value | 8.99809 (rank : 121) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 431 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8NA70, Q5JQN5, Q6PIG3 | Gene names | FAM47B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM47B. | |||||
|
CCD46_HUMAN
|
||||||
| NC score | 0.017634 (rank : 160) | θ value | 4.03905 (rank : 89) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1430 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8N8E3, Q6PIB5, Q8NCR4, Q8NFR4 | Gene names | CCDC46 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 46. | |||||
|
CNTRB_HUMAN
|
||||||
| NC score | 0.017598 (rank : 161) | θ value | 8.99809 (rank : 118) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8N137, Q331K3, Q69YV7, Q8NCB8, Q8WXV3, Q96CQ7, Q9C060 | Gene names | CNTROB, LIP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrobin (LYST-interacting protein 8). | |||||
|
LRRC6_HUMAN
|
||||||
| NC score | 0.017377 (rank : 162) | θ value | 6.88961 (rank : 112) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q86X45, Q13648, Q4G183 | Gene names | LRRC6, LRTP, TSLRP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 6 (Leucine-rich testis-specific protein) (Testis-specific leucine-rich repeat protein). | |||||
|
RUFY2_HUMAN
|
||||||
| NC score | 0.017300 (rank : 163) | θ value | 4.03905 (rank : 96) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1208 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8WXA3, Q96P51, Q9P1Z1 | Gene names | RUFY2, KIAA1537, RABIP4R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and FYVE domain-containing protein 2 (Rab4-interacting protein related). | |||||
|
AN30A_HUMAN
|
||||||
| NC score | 0.014967 (rank : 164) | θ value | 5.27518 (rank : 99) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1119 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9BXX3, Q5W025 | Gene names | ANKRD30A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 30A (NY-BR-1 antigen). | |||||
|
HNRLL_MOUSE
|
||||||
| NC score | 0.012153 (rank : 165) | θ value | 4.03905 (rank : 90) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q921F4, Q8BIP6, Q99J40 | Gene names | Hnrpll | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heterogeneous nuclear ribonucleoprotein L-like. | |||||
|
TTC28_HUMAN
|
||||||
| NC score | 0.011745 (rank : 166) | θ value | 5.27518 (rank : 108) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96AY4, O95928, O95929, Q9NTE4, Q9UG31, Q9UGG5, Q9UPV8, Q9Y3S5 | Gene names | TTC28, KIAA1043 | |||
|
Domain Architecture |

|
|||||
| Description | Tetratricopeptide repeat protein 28 (TPR repeat protein 28). | |||||
|
CYLN2_HUMAN
|
||||||
| NC score | 0.011742 (rank : 167) | θ value | 8.99809 (rank : 120) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1004 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9UDT6, O14527, O43611 | Gene names | CYLN2, KIAA0291, WBSCR4, WSCR4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoplasmic linker protein 2 (Cytoplasmic linker protein 115) (CLIP- 115) (Williams-Beuren syndrome chromosome region 4 protein). | |||||
|
TNIK_HUMAN
|
||||||
| NC score | 0.011563 (rank : 168) | θ value | 0.47712 (rank : 65) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1570 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9UKE5, O60298, Q8WUY7, Q9UKD8, Q9UKD9, Q9UKE0, Q9UKE1, Q9UKE2, Q9UKE3, Q9UKE4 | Gene names | TNIK, KIAA0551 | |||
|
Domain Architecture |

|
|||||
| Description | TRAF2 and NCK-interacting protein kinase (EC 2.7.11.1). | |||||
|
CTTB2_HUMAN
|
||||||
| NC score | 0.010166 (rank : 169) | θ value | 2.36792 (rank : 78) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1061 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8WZ74, O43389, Q7LG11, Q9C0A5 | Gene names | CTTNBP2, C7orf8, CORTBP2, KIAA1758 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cortactin-binding protein 2 (CortBP2). | |||||
|
PLCG1_MOUSE
|
||||||
| NC score | 0.009642 (rank : 170) | θ value | 4.03905 (rank : 95) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 314 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q62077, Q6P1G1 | Gene names | Plcg1, Plcg-1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 1-phosphatidylinositol-4,5-bisphosphate phosphodiesterase gamma 1 (EC 3.1.4.11) (Phosphoinositide phospholipase C) (PLC-gamma-1) (Phospholipase C-gamma-1) (PLC-II) (PLC-148). | |||||
|
MINK1_MOUSE
|
||||||
| NC score | 0.006952 (rank : 171) | θ value | 5.27518 (rank : 103) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1451 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9JM52, Q921M6, Q9JM92 | Gene names | Mink1, Map4k6, Mink | |||
|
Domain Architecture |

|
|||||
| Description | Misshapen-like kinase 1 (EC 2.7.11.1) (Mitogen-activated protein kinase kinase kinase kinase 6) (MAPK/ERK kinase kinase kinase 6) (MEK kinase kinase 6) (MEKKK 6) (Misshapen/NIK-related kinase) (GCK family kinase MiNK). | |||||
|
MINK1_HUMAN
|
||||||
| NC score | 0.006659 (rank : 172) | θ value | 5.27518 (rank : 102) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8N4C8, Q9P1X1, Q9P2R8 | Gene names | MINK1, MAP4K6, MINK | |||
|
Domain Architecture |

|
|||||
| Description | Misshapen-like kinase 1 (EC 2.7.11.1) (Mitogen-activated protein kinase kinase kinase kinase 6) (MAPK/ERK kinase kinase kinase 6) (MEK kinase kinase 6) (MEKKK 6) (Misshapen/NIK-related kinase) (GCK family kinase MiNK). | |||||
|
M4K4_MOUSE
|
||||||
| NC score | 0.006573 (rank : 173) | θ value | 6.88961 (rank : 114) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1560 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P97820 | Gene names | Map4k4, Nik | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase kinase 4 (EC 2.7.11.1) (MAPK/ERK kinase kinase kinase 4) (MEK kinase kinase 4) (MEKKK 4) (HPK/GCK-like kinase HGK) (Nck-interacting kinase). | |||||
|
M4K4_HUMAN
|
||||||
| NC score | 0.006317 (rank : 174) | θ value | 6.88961 (rank : 113) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1589 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O95819, O75172, Q9NST7 | Gene names | MAP4K4, HGK, KIAA0687, NIK | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase kinase 4 (EC 2.7.11.1) (MAPK/ERK kinase kinase kinase 4) (MEK kinase kinase 4) (MEKKK 4) (HPK/GCK-like kinase HGK) (Nck-interacting kinase). | |||||
|
E2AK4_HUMAN
|
||||||
| NC score | 0.005500 (rank : 175) | θ value | 5.27518 (rank : 100) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1012 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9P2K8, Q69YL7, Q6DC97, Q96GN6, Q9H5K1, Q9NSQ3, Q9NSZ5, Q9UJ56 | Gene names | EIF2AK4, GCN2, KIAA1338 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 2-alpha kinase 4 (EC 2.7.11.1) (GCN2-like protein). | |||||
|
LAMA1_HUMAN
|
||||||
| NC score | 0.005084 (rank : 176) | θ value | 4.03905 (rank : 92) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 804 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P25391 | Gene names | LAMA1, LAMA | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-1 chain precursor (Laminin A chain). | |||||
|
E2AK4_MOUSE
|
||||||
| NC score | 0.004740 (rank : 177) | θ value | 6.88961 (rank : 109) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 984 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9QZ05, Q6GQT4, Q6ZPT5, Q8C5S0, Q8CIF5, Q9CT30, Q9CUV9, Q9ESB6, Q9ESB7, Q9ESB8 | Gene names | Eif2ak4, Gcn2, Kiaa1338 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 2-alpha kinase 4 (EC 2.7.11.1) (GCN2-like protein) (mGCN2). | |||||
|
COT1_MOUSE
|
||||||
| NC score | 0.004285 (rank : 178) | θ value | 8.99809 (rank : 119) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q60632, Q61438 | Gene names | Nr2f1, Erbal3, Tfcoup1 | |||
|
Domain Architecture |

|
|||||
| Description | COUP transcription factor 1 (COUP-TF1) (COUP-TF I). | |||||
|
MRCKG_HUMAN
|
||||||
| NC score | 0.001724 (rank : 179) | θ value | 8.99809 (rank : 122) | |||
| Query Neighborhood Hits | 124 | Target Neighborhood Hits | 1519 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q6DT37, O00565 | Gene names | CDC42BPG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK gamma (EC 2.7.11.1) (CDC42- binding protein kinase gamma) (Myotonic dystrophy kinase-related CDC42-binding kinase gamma) (Myotonic dystrophy protein kinase-like alpha) (MRCK gamma) (DMPK-like gamma). | |||||