Please be patient as the page loads
|
MBD1_HUMAN
|
||||||
| SwissProt Accessions | Q9UIS9, O15248, O95241, Q7Z7B5, Q8N4W4, Q9UNZ6, Q9UNZ7, Q9UNZ8, Q9UNZ9 | Gene names | MBD1, PCM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 1 (Methyl-CpG-binding protein MBD1) (Protein containing methyl-CpG-binding domain 1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
MBD1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | Q9UIS9, O15248, O95241, Q7Z7B5, Q8N4W4, Q9UNZ6, Q9UNZ7, Q9UNZ8, Q9UNZ9 | Gene names | MBD1, PCM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 1 (Methyl-CpG-binding protein MBD1) (Protein containing methyl-CpG-binding domain 1). | |||||
|
MBD1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.980165 (rank : 2) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9Z2E2, Q792D6, Q8CCL9, Q9DC19 | Gene names | Mbd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 1 (Methyl-CpG-binding protein MBD1). | |||||
|
MBD2_MOUSE
|
||||||
| θ value | 4.74913e-17 (rank : 3) | NC score | 0.536210 (rank : 3) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Z2E1, Q811D9, Q9Z2D9 | Gene names | Mbd2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 2 (Methyl-CpG-binding protein MBD2). | |||||
|
MBD2_HUMAN
|
||||||
| θ value | 1.38178e-16 (rank : 4) | NC score | 0.532367 (rank : 4) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UBB5, O95242, Q9UIS8 | Gene names | MBD2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 2 (Methyl-CpG-binding protein MBD2) (Demethylase) (DMTase). | |||||
|
MBD3_HUMAN
|
||||||
| θ value | 4.16044e-13 (rank : 5) | NC score | 0.507762 (rank : 6) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O95983, Q6PIL9, Q6PJZ9, Q86XF4 | Gene names | MBD3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 3 (Methyl-CpG-binding protein MBD3). | |||||
|
MBD3_MOUSE
|
||||||
| θ value | 4.16044e-13 (rank : 6) | NC score | 0.508180 (rank : 5) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Z2D8, Q792D3, Q8CFJ1 | Gene names | Mbd3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 3 (Methyl-CpG-binding protein MBD3). | |||||
|
CXCC1_MOUSE
|
||||||
| θ value | 7.84624e-12 (rank : 7) | NC score | 0.436752 (rank : 7) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9CWW7 | Gene names | Cxxc1, Cgbp, Pccx1 | |||
|
Domain Architecture |

|
|||||
| Description | CpG-binding protein (PHD finger and CXXC domain-containing protein 1) (CXXC-type zinc finger protein 1). | |||||
|
HRX_HUMAN
|
||||||
| θ value | 7.84624e-12 (rank : 8) | NC score | 0.280838 (rank : 19) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
CXCC1_HUMAN
|
||||||
| θ value | 1.02475e-11 (rank : 9) | NC score | 0.432665 (rank : 8) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9P0U4, Q8N2W4, Q96BC8, Q9P2V7 | Gene names | CXXC1, CGBP, PCCX1, PHF18 | |||
|
Domain Architecture |
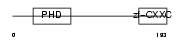
|
|||||
| Description | CpG-binding protein (PHD finger and CXXC domain-containing protein 1) (CXXC-type zinc finger protein 1). | |||||
|
MLL4_HUMAN
|
||||||
| θ value | 2.79066e-09 (rank : 10) | NC score | 0.275080 (rank : 20) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9UMN6, O15022, O95836, Q96GP2, Q96IP3, Q9UK25, Q9Y668, Q9Y669 | Gene names | MLL4, HRX2, KIAA0304, MLL2, TRX2 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 4 (Trithorax homolog 2). | |||||
|
DNMT1_HUMAN
|
||||||
| θ value | 1.38499e-08 (rank : 11) | NC score | 0.343524 (rank : 14) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P26358, Q9UHG5, Q9ULA2, Q9UMZ6 | Gene names | DNMT1, AIM, DNMT | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 1 (EC 2.1.1.37) (Dnmt1) (DNA methyltransferase HsaI) (DNA MTase HsaI) (MCMT) (M.HsaI). | |||||
|
JHD1B_HUMAN
|
||||||
| θ value | 1.99992e-07 (rank : 12) | NC score | 0.313742 (rank : 16) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8NHM5, Q8NCI2, Q96HC7, Q96SL0, Q96T03, Q9NS96, Q9UF75 | Gene names | FBXL10, CXXC2, FBL10, JHDM1B, PCCX2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1B (EC 1.14.11.-) (F-box/LRR-repeat protein 10) (F-box and leucine-rich repeat protein 10) (F-box protein FBL10) (Protein JEMMA) (Jumonji domain-containing EMSY-interactor methyltransferase motif protein) (CXXC-type zinc finger protein 2) (Protein-containing CXXC domain 2). | |||||
|
JHD1B_MOUSE
|
||||||
| θ value | 1.99992e-07 (rank : 13) | NC score | 0.314277 (rank : 15) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6P1G2, Q3V396, Q6PFD0, Q6ZPE8, Q9CSF7, Q9QZN6 | Gene names | Fbxl10, Fbl10, Jhdm1b, Kiaa3014 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1B (EC 1.14.11.-) (F-box/LRR-repeat protein 10) (F-box and leucine-rich repeat protein 10) (F-box protein FBL10). | |||||
|
DNMT1_MOUSE
|
||||||
| θ value | 1.29631e-06 (rank : 14) | NC score | 0.367349 (rank : 11) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P13864, P97413, Q80ZU3, Q9CSC6, Q9QXX6 | Gene names | Dnmt1, Dnmt, Met1, Uim | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 1 (EC 2.1.1.37) (Dnmt1) (DNA methyltransferase MmuI) (DNA MTase MmuI) (MCMT) (M.MmuI) (Met-1). | |||||
|
FXL19_HUMAN
|
||||||
| θ value | 1.29631e-06 (rank : 15) | NC score | 0.370865 (rank : 9) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6PCT2, Q8N789, Q9NT14 | Gene names | FBXL19, FBL19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 19 (F-box and leucine-rich repeat protein 19). | |||||
|
FXL19_MOUSE
|
||||||
| θ value | 1.29631e-06 (rank : 16) | NC score | 0.366222 (rank : 12) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q6PB97, Q7TSH0, Q8BIB4 | Gene names | Fbxl19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 19 (F-box and leucine-rich repeat protein 19). | |||||
|
JHD1A_HUMAN
|
||||||
| θ value | 1.09739e-05 (rank : 17) | NC score | 0.301308 (rank : 18) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Y2K7, Q49A21, Q4G0M3, Q69YY8, Q9BVH5, Q9H7H5, Q9UK66 | Gene names | FBXL11, FBL7, JHDM1A, KIAA1004 | |||
|
Domain Architecture |

|
|||||
| Description | JmjC domain-containing histone demethylation protein 1A (EC 1.14.11.-) (F-box/LRR-repeat protein 11) (F-box and leucine-rich repeat protein 11) (F-box protein FBL7) (F-box protein Lilina). | |||||
|
JHD1A_MOUSE
|
||||||
| θ value | 1.09739e-05 (rank : 18) | NC score | 0.302045 (rank : 17) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P59997, Q3U1M5, Q3UR56, Q3V3Q1, Q69ZT4 | Gene names | Fbxl11, Jhdm1a, Kiaa1004 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1A (EC 1.14.11.-) (F-box/LRR-repeat protein 11) (F-box and leucine-rich repeat protein 11). | |||||
|
MECP2_HUMAN
|
||||||
| θ value | 4.1701e-05 (rank : 19) | NC score | 0.368785 (rank : 10) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P51608, O15233 | Gene names | MECP2 | |||
|
Domain Architecture |
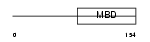
|
|||||
| Description | Methyl-CpG-binding protein 2 (MeCP-2 protein) (MeCP2). | |||||
|
MECP2_MOUSE
|
||||||
| θ value | 4.1701e-05 (rank : 20) | NC score | 0.363611 (rank : 13) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Z2D6 | Gene names | Mecp2 | |||
|
Domain Architecture |
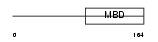
|
|||||
| Description | Methyl-CpG-binding protein 2 (MeCP-2 protein) (MeCP2). | |||||
|
CXXC6_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 21) | NC score | 0.214275 (rank : 24) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8NFU7, Q5VUP7, Q7Z6B6, Q8TCR1, Q9C0I7 | Gene names | CXXC6, KIAA1676, LCX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CXXC-type zinc finger protein 6 (Leukemia-associated protein with a CXXC domain). | |||||
|
MBD4_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 22) | NC score | 0.247904 (rank : 22) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O95243, Q7Z4T3, Q96F09 | Gene names | MBD4, MED1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 4 (EC 3.2.2.-) (Methyl-CpG-binding protein MBD4) (Methyl-CpG-binding endonuclease 1) (Mismatch-specific DNA N-glycosylase). | |||||
|
MBD4_MOUSE
|
||||||
| θ value | 0.0148317 (rank : 23) | NC score | 0.252838 (rank : 21) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Z2D7, Q792D2, Q8R3R3 | Gene names | Mbd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 4 (EC 3.2.2.-) (Methyl-CpG-binding protein MBD4) (Mismatch-specific DNA N-glycosylase). | |||||
|
LAMB2_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 24) | NC score | 0.049025 (rank : 57) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q61292, Q62182 | Gene names | Lamb2, Lams | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-2 chain precursor (S-laminin) (S-LAM). | |||||
|
KRA59_HUMAN
|
||||||
| θ value | 0.163984 (rank : 25) | NC score | 0.060926 (rank : 44) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P26371, Q14564 | Gene names | KRTAP5-9, KAP5.9, KRN1, KRTAP5.9, UHSK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-9 (Keratin-associated protein 5.9) (Ultrahigh sulfur keratin-associated protein 5.9) (Keratin, cuticle, ultrahigh sulfur 1) (Keratin, ultra high-sulfur matrix protein A) (UHS keratin A) (UHS KerA). | |||||
|
ATS18_MOUSE
|
||||||
| θ value | 0.279714 (rank : 26) | NC score | 0.013823 (rank : 84) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q4VC17, Q8BZD1 | Gene names | Adamts18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAMTS-18 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase with thrombospondin motifs 18) (ADAM-TS 18) (ADAM-TS18). | |||||
|
DMRTD_HUMAN
|
||||||
| θ value | 0.47712 (rank : 27) | NC score | 0.040852 (rank : 59) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8IXT2, Q8N6Q2, Q96M39, Q96SD4 | Gene names | DMRTC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Doublesex- and mab-3-related transcription factor C2. | |||||
|
PHLPP_HUMAN
|
||||||
| θ value | 0.47712 (rank : 28) | NC score | 0.016774 (rank : 83) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 604 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O60346, Q641Q7, Q6P4C4, Q6PJI6, Q86TN6, Q96FK2, Q9NUY1 | Gene names | PHLPP, KIAA0606, PLEKHE1, SCOP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH domain leucine-rich repeat-containing protein phosphatase (EC 3.1.3.16) (PH domain leucine-rich repeat protein phosphatase) (Pleckstrin homology domain-containing family E protein 1) (Suprachiasmatic nucleus circadian oscillatory protein) (hSCOP). | |||||
|
ZN672_HUMAN
|
||||||
| θ value | 0.47712 (rank : 29) | NC score | 0.003816 (rank : 98) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q499Z4, Q96H65, Q96IM3, Q9H6G5 | Gene names | ZNF672 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 672. | |||||
|
MKRN1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 30) | NC score | 0.022689 (rank : 76) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9QXP6 | Gene names | Mkrn1 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-1. | |||||
|
MLL3_HUMAN
|
||||||
| θ value | 0.813845 (rank : 31) | NC score | 0.076486 (rank : 34) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
MAGE1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 32) | NC score | 0.019466 (rank : 79) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
ITB6_MOUSE
|
||||||
| θ value | 1.38821 (rank : 33) | NC score | 0.028396 (rank : 66) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9Z0T9 | Gene names | Itgb6 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-6 precursor. | |||||
|
MLL3_MOUSE
|
||||||
| θ value | 1.38821 (rank : 34) | NC score | 0.079210 (rank : 33) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
BAZ2A_HUMAN
|
||||||
| θ value | 1.81305 (rank : 35) | NC score | 0.054220 (rank : 53) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 780 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9UIF9, O00536, O15030, Q96H26 | Gene names | BAZ2A, KIAA0314, TIP5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5) (hWALp3). | |||||
|
BAZ2A_MOUSE
|
||||||
| θ value | 1.81305 (rank : 36) | NC score | 0.053472 (rank : 54) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q91YE5 | Gene names | Baz2a, Tip5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5). | |||||
|
MEGF8_MOUSE
|
||||||
| θ value | 1.81305 (rank : 37) | NC score | 0.024979 (rank : 73) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 520 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P60882, Q80TR3, Q80V41, Q8BMN9, Q8JZW7, Q8K0J3 | Gene names | Megf8, Egfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 8 (EGF-like domain- containing protein 4) (Multiple EGF-like domain protein 4). | |||||
|
ZN628_HUMAN
|
||||||
| θ value | 1.81305 (rank : 38) | NC score | 0.002185 (rank : 101) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 814 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q5EBL2, Q86X34 | Gene names | ZNF628 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 628. | |||||
|
CHD5_HUMAN
|
||||||
| θ value | 2.36792 (rank : 39) | NC score | 0.020546 (rank : 77) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8TDI0, O75032, Q5TG89, Q9UFR9 | Gene names | CHD5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain helicase-DNA-binding protein 5 (EC 3.6.1.-) (ATP- dependent helicase CHD5) (CHD-5). | |||||
|
LAMB2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 40) | NC score | 0.040402 (rank : 60) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 692 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P55268, Q16321 | Gene names | LAMB2, LAMS | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-2 chain precursor (S-laminin) (Laminin B1s chain). | |||||
|
SRCH_HUMAN
|
||||||
| θ value | 2.36792 (rank : 41) | NC score | 0.050147 (rank : 56) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P23327, Q504Y6 | Gene names | HRC, HCP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcoplasmic reticulum histidine-rich calcium-binding protein precursor. | |||||
|
ZN507_HUMAN
|
||||||
| θ value | 2.36792 (rank : 42) | NC score | 0.005437 (rank : 94) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 739 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8TCN5 | Gene names | ZNF507 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 507. | |||||
|
GA2L2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 43) | NC score | 0.029301 (rank : 65) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5SSG4 | Gene names | Gas2l2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GAS2-like protein 2 (Growth arrest-specific 2-like 2). | |||||
|
MMP17_HUMAN
|
||||||
| θ value | 3.0926 (rank : 44) | NC score | 0.007833 (rank : 90) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9ULZ9, Q14850 | Gene names | MMP17, MT4MMP | |||
|
Domain Architecture |

|
|||||
| Description | Matrix metalloproteinase-17 precursor (EC 3.4.24.-) (MMP-17) (Membrane-type matrix metalloproteinase 4) (MT-MMP 4) (Membrane-type-4 matrix metalloproteinase) (MT4-MMP). | |||||
|
TPR_HUMAN
|
||||||
| θ value | 3.0926 (rank : 45) | NC score | 0.004024 (rank : 97) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
BSN_MOUSE
|
||||||
| θ value | 4.03905 (rank : 46) | NC score | 0.027454 (rank : 68) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
KRA51_HUMAN
|
||||||
| θ value | 4.03905 (rank : 47) | NC score | 0.038760 (rank : 61) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 288 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6L8H4 | Gene names | KRTAP5-1, KAP5.1, KRTAP5.1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-1 (Keratin-associated protein 5.1) (Ultrahigh sulfur keratin-associated protein 5.1). | |||||
|
KRA54_HUMAN
|
||||||
| θ value | 4.03905 (rank : 48) | NC score | 0.045108 (rank : 58) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 270 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q6L8H1 | Gene names | KRTAP5-4, KAP5.4, KRTAP5.4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-4 (Keratin-associated protein 5.4) (Ultrahigh sulfur keratin-associated protein 5.4). | |||||
|
LAMC1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 49) | NC score | 0.025583 (rank : 71) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 922 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P11047 | Gene names | LAMC1, LAMB2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-1 chain precursor (Laminin B2 chain). | |||||
|
RSPO4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 50) | NC score | 0.017528 (rank : 81) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q2I0M5, Q9UGB2 | Gene names | RSPO4, C20orf182 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | R-spondin-4 precursor (Roof plate-specific spondin-4) (hRspo4). | |||||
|
TNR25_HUMAN
|
||||||
| θ value | 4.03905 (rank : 51) | NC score | 0.025811 (rank : 70) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q93038, O00275, O00276, O00277, O00278, O00279, O00280, O14865, O14866, P78507, P78515, Q92983, Q93036, Q93037, Q99722, Q99830, Q99831, Q9BY86, Q9UME0, Q9UME1, Q9UME5 | Gene names | TNFRSF25, APO3, DDR3, DR3, TNFRSF12, WSL, WSL1 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 25 precursor (WSL-1 protein) (Apoptosis-mediating receptor DR3) (Apoptosis-mediating receptor TRAMP) (Death domain receptor 3) (WSL protein) (Apoptosis- inducing receptor AIR) (Apo-3) (Lymphocyte-associated receptor of death) (LARD). | |||||
|
AMOT_HUMAN
|
||||||
| θ value | 5.27518 (rank : 52) | NC score | 0.013014 (rank : 86) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 712 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q4VCS5, Q504X5, Q9HD27, Q9UPT1 | Gene names | AMOT, KIAA1071 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin. | |||||
|
DAB1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 53) | NC score | 0.022989 (rank : 75) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O75553, Q9NYA8 | Gene names | DAB1 | |||
|
Domain Architecture |
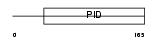
|
|||||
| Description | Disabled homolog 1. | |||||
|
DFRC_MOUSE
|
||||||
| θ value | 5.27518 (rank : 54) | NC score | 0.025170 (rank : 72) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P17533, Q9QVW7 | Gene names | Defcr-rs1 | |||
|
Domain Architecture |

|
|||||
| Description | Defensin-related sequence cryptdin peptide precursor (Cryptdin-related protein 1C) (CRS1C). | |||||
|
GCR_MOUSE
|
||||||
| θ value | 5.27518 (rank : 55) | NC score | 0.006342 (rank : 93) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P06537, Q61628, Q61629 | Gene names | Nr3c1, Grl, Grl1 | |||
|
Domain Architecture |

|
|||||
| Description | Glucocorticoid receptor (GR). | |||||
|
ADAM2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 56) | NC score | 0.010827 (rank : 87) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q60718, Q60814, Q9D4G3, Q9QWJ0 | Gene names | Adam2, Ftnb | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 2 precursor (A disintegrin and metalloproteinase domain 2) (Fertilin subunit beta) (PH-30) (PH30) (PH30-beta). | |||||
|
COBA1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 57) | NC score | 0.004031 (rank : 96) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 487 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61245, Q64047 | Gene names | Col11a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XI) chain precursor. | |||||
|
FXR2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 58) | NC score | 0.010326 (rank : 88) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9WVR4, Q9WVR5 | Gene names | Fxr2, Fxr2h | |||
|
Domain Architecture |

|
|||||
| Description | Fragile X mental retardation syndrome-related protein 2. | |||||
|
KRA58_HUMAN
|
||||||
| θ value | 6.88961 (rank : 59) | NC score | 0.019507 (rank : 78) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 324 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O75690, Q6L8G7, Q6UTX6 | Gene names | KRTAP5-8, KAP5.8, KRTAP5.8, UHSKB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-8 (Keratin-associated protein 5.8) (Ultrahigh sulfur keratin-associated protein 5.8) (Keratin, ultra high-sulfur matrix protein B) (UHS keratin B) (UHS KerB). | |||||
|
LAMB1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 60) | NC score | 0.030817 (rank : 62) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 1018 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P07942 | Gene names | LAMB1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-1 chain precursor (Laminin B1 chain). | |||||
|
LAMC3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 61) | NC score | 0.027735 (rank : 67) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 588 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9Y6N6 | Gene names | LAMC3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-3 chain precursor (Laminin 12 gamma 3 subunit). | |||||
|
LAMC3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 62) | NC score | 0.027433 (rank : 69) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9R0B6, Q9WTW6 | Gene names | Lamc3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-3 chain precursor (Laminin 12 gamma 3 subunit). | |||||
|
NET2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 63) | NC score | 0.030564 (rank : 63) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O00634 | Gene names | NTN2L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin-2-like protein precursor. | |||||
|
RORG_HUMAN
|
||||||
| θ value | 6.88961 (rank : 64) | NC score | 0.005221 (rank : 95) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P51449, Q8N5V7, Q8NCY8 | Gene names | RORC, NR1F3, RORG, RZRG | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor ROR-gamma (Nuclear receptor RZR-gamma). | |||||
|
WHRN_MOUSE
|
||||||
| θ value | 6.88961 (rank : 65) | NC score | 0.007231 (rank : 91) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80VW5, Q80TC2, Q80VW4 | Gene names | Whrn, Kiaa1526 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Whirlin. | |||||
|
ZN335_HUMAN
|
||||||
| θ value | 6.88961 (rank : 66) | NC score | 0.002466 (rank : 99) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 973 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9H4Z2, Q9H684 | Gene names | ZNF335 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 335. | |||||
|
CSPG5_MOUSE
|
||||||
| θ value | 8.99809 (rank : 67) | NC score | 0.013016 (rank : 85) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q71M36, Q71M37, Q7TNT8, Q8BPJ5, Q9QY32 | Gene names | Cspg5, Caleb, Ngc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate proteoglycan 5 precursor (Neuroglycan C) (Acidic leucine-rich EGF-like domain-containing brain protein). | |||||
|
ITB1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 68) | NC score | 0.019062 (rank : 80) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P09055 | Gene names | Itgb1 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-1 precursor (Fibronectin receptor subunit beta) (Integrin VLA-4 subunit beta) (CD29 antigen). | |||||
|
JAG2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 69) | NC score | 0.010214 (rank : 89) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9QYE5, O55139, O70219 | Gene names | Jag2 | |||
|
Domain Architecture |

|
|||||
| Description | Jagged-2 precursor (Jagged2). | |||||
|
KCD12_MOUSE
|
||||||
| θ value | 8.99809 (rank : 70) | NC score | 0.007120 (rank : 92) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6WVG3 | Gene names | Kctd12, Pfet1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD12 (Pfetin) (Predominantly fetal expressed T1 domain). | |||||
|
LAMA5_HUMAN
|
||||||
| θ value | 8.99809 (rank : 71) | NC score | 0.023643 (rank : 74) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 976 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O15230, Q8WZA7, Q9H1P1 | Gene names | LAMA5, KIAA0533, KIAA1907 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-5 chain precursor. | |||||
|
NET2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 72) | NC score | 0.029935 (rank : 64) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9R1A3, Q9QY49, Q9WVA6 | Gene names | Ntn2l, Ntn3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin-2-like protein precursor (Netrin-3). | |||||
|
NTAL_HUMAN
|
||||||
| θ value | 8.99809 (rank : 73) | NC score | 0.017364 (rank : 82) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9GZY6, Q9BXX8, Q9NZY9 | Gene names | LAT2, LAB, NTAL, WBS15, WBSCR15, WBSCR5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Linker for activation of T-cells family member 2 (Non-T-cell activation linker) (Linker for activation of B-cells) (Membrane- associated adapter molecule) (Williams-Beuren syndrome chromosome region 15 protein). | |||||
|
OBSCN_HUMAN
|
||||||
| θ value | 8.99809 (rank : 74) | NC score | 0.000646 (rank : 104) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 1931 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q5VST9, Q2A664, Q5T7G8, Q5T7G9, Q5VSU2, Q86YC7, Q8NHN0, Q8NHN1, Q8NHN2, Q8NHN4, Q8NHN5, Q8NHN6, Q8NHN7, Q8NHN8, Q8NHN9, Q96AA2, Q9HCD3, Q9HCL6 | Gene names | OBSCN, KIAA1556, KIAA1639 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Obscurin (Obscurin-myosin light chain kinase) (Obscurin-MLCK) (Obscurin-RhoGEF). | |||||
|
TRI10_MOUSE
|
||||||
| θ value | 8.99809 (rank : 75) | NC score | 0.002216 (rank : 100) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9WUH5, Q80WA9, Q9CY03 | Gene names | Trim10, Herf1, Rnf9 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 10 (RING finger protein 9) (Hematopoietic RING finger 1). | |||||
|
ZN689_HUMAN
|
||||||
| θ value | 8.99809 (rank : 76) | NC score | 0.000820 (rank : 102) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 736 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96CS4, Q658J5 | Gene names | ZNF689 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 689. | |||||
|
ZN689_MOUSE
|
||||||
| θ value | 8.99809 (rank : 77) | NC score | 0.000812 (rank : 103) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 736 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BKK5, Q8C1C6 | Gene names | Znf689, Zfp689 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 689. | |||||
|
FBXL2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 78) | NC score | 0.054343 (rank : 52) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UKC9, Q9NVQ8, Q9UK27, Q9UKA5, Q9Y3Y9 | Gene names | FBXL2, FBL2, FBL3 | |||
|
Domain Architecture |
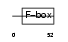
|
|||||
| Description | F-box/LRR-repeat protein 2 (F-box and leucine-rich repeat protein 2) (F-box protein FBL2/FBL3). | |||||
|
FBXL2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 79) | NC score | 0.056765 (rank : 51) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BH16, Q8BXM4 | Gene names | Fbxl2, Fbl2 | |||
|
Domain Architecture |
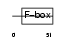
|
|||||
| Description | F-box/LRR-repeat protein 2 (F-box and leucine-rich repeat protein 2). | |||||
|
FBXL7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 80) | NC score | 0.067971 (rank : 36) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UJT9, O94926 | Gene names | FBXL7, FBL6, FBL7, KIAA0840 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 7 (F-box and leucine-rich repeat protein 7) (F-box protein FBL6/FBL7). | |||||
|
FXL13_HUMAN
|
||||||
| θ value | θ > 10 (rank : 81) | NC score | 0.051236 (rank : 55) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8NEE6, Q6UVW7, Q6UVW8, Q75MN5, Q86UJ5, Q8N7Y4, Q8TCL2, Q8WUF9, Q8WUG0 | Gene names | FBXL13, FBL13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 13 (F-box and leucine-rich repeat protein 13). | |||||
|
FXL13_MOUSE
|
||||||
| θ value | θ > 10 (rank : 82) | NC score | 0.059300 (rank : 47) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CDU4, Q8CDE9 | Gene names | Fbxl13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 13 (F-box and leucine-rich repeat protein 13). | |||||
|
FXL14_HUMAN
|
||||||
| θ value | θ > 10 (rank : 83) | NC score | 0.057410 (rank : 50) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8N1E6 | Gene names | FBXL14, FBL14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 14 (F-box and leucine-rich repeat protein 14). | |||||
|
FXL14_MOUSE
|
||||||
| θ value | θ > 10 (rank : 84) | NC score | 0.057573 (rank : 49) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BID8, Q3U2Q9, Q8R5H7, Q8VDT7, Q922N5 | Gene names | Fbxl14, Fbl14, Ppa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 14 (F-box and leucine-rich repeat protein 14). | |||||
|
FXL16_HUMAN
|
||||||
| θ value | θ > 10 (rank : 85) | NC score | 0.060087 (rank : 45) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8N461, Q2MHR2, Q96S14, Q9UJI0 | Gene names | FBXL16, C16orf22, FBL16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 16 (F-box and leucine-rich repeat protein 16). | |||||
|
FXL20_HUMAN
|
||||||
| θ value | θ > 10 (rank : 86) | NC score | 0.061168 (rank : 42) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96IG2 | Gene names | FBXL20, FBL2 | |||
|
Domain Architecture |
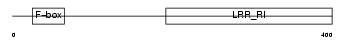
|
|||||
| Description | F-box/LRR-repeat protein 20 (F-box and leucine-rich repeat protein 20) (F-box/LRR-repeat protein 2-like). | |||||
|
FXL20_MOUSE
|
||||||
| θ value | θ > 10 (rank : 87) | NC score | 0.061040 (rank : 43) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9CZV8, Q8BZ95 | Gene names | Fbxl20, Fbl2 | |||
|
Domain Architecture |
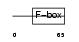
|
|||||
| Description | F-box/LRR-repeat protein 20 (F-box and leucine-rich repeat protein 20) (F-box/LRR-repeat protein 2-like). | |||||
|
K1718_HUMAN
|
||||||
| θ value | θ > 10 (rank : 88) | NC score | 0.118306 (rank : 31) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6ZMT4, Q6MZL8, Q9C0E5 | Gene names | KIAA1718 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1718. | |||||
|
K1718_MOUSE
|
||||||
| θ value | θ > 10 (rank : 89) | NC score | 0.121163 (rank : 27) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q3UWM4, Q3UWN8, Q6ZPJ5, Q8C969, Q8C9E0, Q91VX8 | Gene names | Kiaa1718 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1718. | |||||
|
LRC29_HUMAN
|
||||||
| θ value | θ > 10 (rank : 90) | NC score | 0.062579 (rank : 40) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8WV35, Q9UKA0 | Gene names | LRRC29, FBL9, FBXL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 29 (F-box/LRR-repeat protein 9) (F-box and leucine-rich repeat protein 9) (F-box protein FBL9). | |||||
|
MB3L1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 91) | NC score | 0.229250 (rank : 23) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8WWY6 | Gene names | MBD3L1, MBD3L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 3-like 1 (MBD3-like 1). | |||||
|
MB3L1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 92) | NC score | 0.205041 (rank : 25) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D9H3, Q6P8W5, Q9D2S3 | Gene names | Mbd3l1, Mbd3l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 3-like 1 (MBD3-like 1). | |||||
|
MB3L2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 93) | NC score | 0.190529 (rank : 26) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8NHZ7 | Gene names | MBD3L2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 3-like 2 (MBD3-like 2). | |||||
|
MLL2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 94) | NC score | 0.063432 (rank : 38) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
PHF11_HUMAN
|
||||||
| θ value | θ > 10 (rank : 95) | NC score | 0.066987 (rank : 37) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UIL8, Q9Y5A2 | Gene names | PHF11, BCAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 11 (BRCA1 C-terminus-associated protein) (NY-REN-34 antigen). | |||||
|
PHF2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 96) | NC score | 0.118295 (rank : 32) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O75151, Q8N3K2, Q9Y6N4 | Gene names | PHF2, KIAA0662 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
PHF2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 97) | NC score | 0.119181 (rank : 30) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9WTU0, Q80WA8 | Gene names | Phf2 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
PHF8_HUMAN
|
||||||
| θ value | θ > 10 (rank : 98) | NC score | 0.121157 (rank : 28) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UPP1, Q5H9U5, Q5VUJ4, Q7Z6D4, Q9HAH2 | Gene names | PHF8, KIAA1111, ZNF422 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 8. | |||||
|
PHF8_MOUSE
|
||||||
| θ value | θ > 10 (rank : 99) | NC score | 0.120033 (rank : 29) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q80TJ7, Q8BLX8, Q8BLY0, Q8BZ61, Q8CG26 | Gene names | Phf8, Kiaa1111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 8. | |||||
|
SET1A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.072534 (rank : 35) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O15047, Q6PIF3, Q8TAJ6 | Gene names | SETD1A, KIAA0339, SET1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET1 (EC 2.1.1.43) (Set1/Ash2 histone methyltransferase complex subunit SET1) (SET-domain-containing protein 1A). | |||||
|
SETD8_HUMAN
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.062971 (rank : 39) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NQR1, Q86W83, Q8TD09 | Gene names | SETD8, PRSET7, SET07, SET8 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H4 lysine-20 specific (EC 2.1.1.43) (Histone H4-K20 methyltransferase) (H4-K20-HMTase) (SET domain-containing lysine methyltransferase 8) (SET domain-containing protein 8) (PR/SET domain-containing protein 07) (PR/SET07) (PR-Set7). | |||||
|
SETD8_MOUSE
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.059184 (rank : 48) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q2YDW7, Q8C0J9 | Gene names | SETD8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H4 lysine-20 specific (EC 2.1.1.43) (Histone H4-K20 methyltransferase) (H4-K20-HMTase) (SET domain-containing lysine methyltransferase 8) (SET domain-containing protein 8). | |||||
|
SKP2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.062267 (rank : 41) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q13309, Q8TDZ0, Q8TDZ1, Q9BV69 | Gene names | SKP2, FBXL1 | |||
|
Domain Architecture |

|
|||||
| Description | S-phase kinase-associated protein 2 (F-box protein Skp2) (Cyclin A/CDK2-associated protein p45) (p45skp2) (F-box/LRR-repeat protein 1). | |||||
|
SKP2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.060052 (rank : 46) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Z0Z3, Q8BSG7, Q8C8Y9, Q8CB22, Q99J53 | Gene names | Skp2 | |||
|
Domain Architecture |

|
|||||
| Description | S-phase kinase-associated protein 2 (F-box protein Skp2) (Cyclin A/CDK2-associated protein p45) (F-box/WD-40 protein 1) (FWD1). | |||||
|
MBD1_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | Q9UIS9, O15248, O95241, Q7Z7B5, Q8N4W4, Q9UNZ6, Q9UNZ7, Q9UNZ8, Q9UNZ9 | Gene names | MBD1, PCM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 1 (Methyl-CpG-binding protein MBD1) (Protein containing methyl-CpG-binding domain 1). | |||||
|
MBD1_MOUSE
|
||||||
| NC score | 0.980165 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9Z2E2, Q792D6, Q8CCL9, Q9DC19 | Gene names | Mbd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 1 (Methyl-CpG-binding protein MBD1). | |||||
|
MBD2_MOUSE
|
||||||
| NC score | 0.536210 (rank : 3) | θ value | 4.74913e-17 (rank : 3) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Z2E1, Q811D9, Q9Z2D9 | Gene names | Mbd2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 2 (Methyl-CpG-binding protein MBD2). | |||||
|
MBD2_HUMAN
|
||||||
| NC score | 0.532367 (rank : 4) | θ value | 1.38178e-16 (rank : 4) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UBB5, O95242, Q9UIS8 | Gene names | MBD2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 2 (Methyl-CpG-binding protein MBD2) (Demethylase) (DMTase). | |||||
|
MBD3_MOUSE
|
||||||
| NC score | 0.508180 (rank : 5) | θ value | 4.16044e-13 (rank : 6) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Z2D8, Q792D3, Q8CFJ1 | Gene names | Mbd3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 3 (Methyl-CpG-binding protein MBD3). | |||||
|
MBD3_HUMAN
|
||||||
| NC score | 0.507762 (rank : 6) | θ value | 4.16044e-13 (rank : 5) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O95983, Q6PIL9, Q6PJZ9, Q86XF4 | Gene names | MBD3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 3 (Methyl-CpG-binding protein MBD3). | |||||
|
CXCC1_MOUSE
|
||||||
| NC score | 0.436752 (rank : 7) | θ value | 7.84624e-12 (rank : 7) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9CWW7 | Gene names | Cxxc1, Cgbp, Pccx1 | |||
|
Domain Architecture |

|
|||||
| Description | CpG-binding protein (PHD finger and CXXC domain-containing protein 1) (CXXC-type zinc finger protein 1). | |||||
|
CXCC1_HUMAN
|
||||||
| NC score | 0.432665 (rank : 8) | θ value | 1.02475e-11 (rank : 9) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9P0U4, Q8N2W4, Q96BC8, Q9P2V7 | Gene names | CXXC1, CGBP, PCCX1, PHF18 | |||
|
Domain Architecture |

|
|||||
| Description | CpG-binding protein (PHD finger and CXXC domain-containing protein 1) (CXXC-type zinc finger protein 1). | |||||
|
FXL19_HUMAN
|
||||||
| NC score | 0.370865 (rank : 9) | θ value | 1.29631e-06 (rank : 15) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6PCT2, Q8N789, Q9NT14 | Gene names | FBXL19, FBL19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 19 (F-box and leucine-rich repeat protein 19). | |||||
|
MECP2_HUMAN
|
||||||
| NC score | 0.368785 (rank : 10) | θ value | 4.1701e-05 (rank : 19) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P51608, O15233 | Gene names | MECP2 | |||
|
Domain Architecture |

|
|||||
| Description | Methyl-CpG-binding protein 2 (MeCP-2 protein) (MeCP2). | |||||
|
DNMT1_MOUSE
|
||||||
| NC score | 0.367349 (rank : 11) | θ value | 1.29631e-06 (rank : 14) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P13864, P97413, Q80ZU3, Q9CSC6, Q9QXX6 | Gene names | Dnmt1, Dnmt, Met1, Uim | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 1 (EC 2.1.1.37) (Dnmt1) (DNA methyltransferase MmuI) (DNA MTase MmuI) (MCMT) (M.MmuI) (Met-1). | |||||
|
FXL19_MOUSE
|
||||||
| NC score | 0.366222 (rank : 12) | θ value | 1.29631e-06 (rank : 16) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q6PB97, Q7TSH0, Q8BIB4 | Gene names | Fbxl19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 19 (F-box and leucine-rich repeat protein 19). | |||||
|
MECP2_MOUSE
|
||||||
| NC score | 0.363611 (rank : 13) | θ value | 4.1701e-05 (rank : 20) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Z2D6 | Gene names | Mecp2 | |||
|
Domain Architecture |

|
|||||
| Description | Methyl-CpG-binding protein 2 (MeCP-2 protein) (MeCP2). | |||||
|
DNMT1_HUMAN
|
||||||
| NC score | 0.343524 (rank : 14) | θ value | 1.38499e-08 (rank : 11) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P26358, Q9UHG5, Q9ULA2, Q9UMZ6 | Gene names | DNMT1, AIM, DNMT | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 1 (EC 2.1.1.37) (Dnmt1) (DNA methyltransferase HsaI) (DNA MTase HsaI) (MCMT) (M.HsaI). | |||||
|
JHD1B_MOUSE
|
||||||
| NC score | 0.314277 (rank : 15) | θ value | 1.99992e-07 (rank : 13) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6P1G2, Q3V396, Q6PFD0, Q6ZPE8, Q9CSF7, Q9QZN6 | Gene names | Fbxl10, Fbl10, Jhdm1b, Kiaa3014 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1B (EC 1.14.11.-) (F-box/LRR-repeat protein 10) (F-box and leucine-rich repeat protein 10) (F-box protein FBL10). | |||||
|
JHD1B_HUMAN
|
||||||
| NC score | 0.313742 (rank : 16) | θ value | 1.99992e-07 (rank : 12) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8NHM5, Q8NCI2, Q96HC7, Q96SL0, Q96T03, Q9NS96, Q9UF75 | Gene names | FBXL10, CXXC2, FBL10, JHDM1B, PCCX2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1B (EC 1.14.11.-) (F-box/LRR-repeat protein 10) (F-box and leucine-rich repeat protein 10) (F-box protein FBL10) (Protein JEMMA) (Jumonji domain-containing EMSY-interactor methyltransferase motif protein) (CXXC-type zinc finger protein 2) (Protein-containing CXXC domain 2). | |||||
|
JHD1A_MOUSE
|
||||||
| NC score | 0.302045 (rank : 17) | θ value | 1.09739e-05 (rank : 18) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P59997, Q3U1M5, Q3UR56, Q3V3Q1, Q69ZT4 | Gene names | Fbxl11, Jhdm1a, Kiaa1004 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | JmjC domain-containing histone demethylation protein 1A (EC 1.14.11.-) (F-box/LRR-repeat protein 11) (F-box and leucine-rich repeat protein 11). | |||||
|
JHD1A_HUMAN
|
||||||
| NC score | 0.301308 (rank : 18) | θ value | 1.09739e-05 (rank : 17) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Y2K7, Q49A21, Q4G0M3, Q69YY8, Q9BVH5, Q9H7H5, Q9UK66 | Gene names | FBXL11, FBL7, JHDM1A, KIAA1004 | |||
|
Domain Architecture |

|
|||||
| Description | JmjC domain-containing histone demethylation protein 1A (EC 1.14.11.-) (F-box/LRR-repeat protein 11) (F-box and leucine-rich repeat protein 11) (F-box protein FBL7) (F-box protein Lilina). | |||||
|
HRX_HUMAN
|
||||||
| NC score | 0.280838 (rank : 19) | θ value | 7.84624e-12 (rank : 8) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
MLL4_HUMAN
|
||||||
| NC score | 0.275080 (rank : 20) | θ value | 2.79066e-09 (rank : 10) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9UMN6, O15022, O95836, Q96GP2, Q96IP3, Q9UK25, Q9Y668, Q9Y669 | Gene names | MLL4, HRX2, KIAA0304, MLL2, TRX2 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 4 (Trithorax homolog 2). | |||||
|
MBD4_MOUSE
|
||||||
| NC score | 0.252838 (rank : 21) | θ value | 0.0148317 (rank : 23) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Z2D7, Q792D2, Q8R3R3 | Gene names | Mbd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 4 (EC 3.2.2.-) (Methyl-CpG-binding protein MBD4) (Mismatch-specific DNA N-glycosylase). | |||||
|
MBD4_HUMAN
|
||||||
| NC score | 0.247904 (rank : 22) | θ value | 0.0148317 (rank : 22) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O95243, Q7Z4T3, Q96F09 | Gene names | MBD4, MED1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 4 (EC 3.2.2.-) (Methyl-CpG-binding protein MBD4) (Methyl-CpG-binding endonuclease 1) (Mismatch-specific DNA N-glycosylase). | |||||
|
MB3L1_HUMAN
|
||||||
| NC score | 0.229250 (rank : 23) | θ value | θ > 10 (rank : 91) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8WWY6 | Gene names | MBD3L1, MBD3L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 3-like 1 (MBD3-like 1). | |||||
|
CXXC6_HUMAN
|
||||||
| NC score | 0.214275 (rank : 24) | θ value | 0.00509761 (rank : 21) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8NFU7, Q5VUP7, Q7Z6B6, Q8TCR1, Q9C0I7 | Gene names | CXXC6, KIAA1676, LCX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CXXC-type zinc finger protein 6 (Leukemia-associated protein with a CXXC domain). | |||||
|
MB3L1_MOUSE
|
||||||
| NC score | 0.205041 (rank : 25) | θ value | θ > 10 (rank : 92) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D9H3, Q6P8W5, Q9D2S3 | Gene names | Mbd3l1, Mbd3l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 3-like 1 (MBD3-like 1). | |||||
|
MB3L2_HUMAN
|
||||||
| NC score | 0.190529 (rank : 26) | θ value | θ > 10 (rank : 93) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8NHZ7 | Gene names | MBD3L2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 3-like 2 (MBD3-like 2). | |||||
|
K1718_MOUSE
|
||||||
| NC score | 0.121163 (rank : 27) | θ value | θ > 10 (rank : 89) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q3UWM4, Q3UWN8, Q6ZPJ5, Q8C969, Q8C9E0, Q91VX8 | Gene names | Kiaa1718 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1718. | |||||
|
PHF8_HUMAN
|
||||||
| NC score | 0.121157 (rank : 28) | θ value | θ > 10 (rank : 98) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UPP1, Q5H9U5, Q5VUJ4, Q7Z6D4, Q9HAH2 | Gene names | PHF8, KIAA1111, ZNF422 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 8. | |||||
|
PHF8_MOUSE
|
||||||
| NC score | 0.120033 (rank : 29) | θ value | θ > 10 (rank : 99) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q80TJ7, Q8BLX8, Q8BLY0, Q8BZ61, Q8CG26 | Gene names | Phf8, Kiaa1111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 8. | |||||
|
PHF2_MOUSE
|
||||||
| NC score | 0.119181 (rank : 30) | θ value | θ > 10 (rank : 97) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9WTU0, Q80WA8 | Gene names | Phf2 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
K1718_HUMAN
|
||||||
| NC score | 0.118306 (rank : 31) | θ value | θ > 10 (rank : 88) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6ZMT4, Q6MZL8, Q9C0E5 | Gene names | KIAA1718 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1718. | |||||
|
PHF2_HUMAN
|
||||||
| NC score | 0.118295 (rank : 32) | θ value | θ > 10 (rank : 96) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O75151, Q8N3K2, Q9Y6N4 | Gene names | PHF2, KIAA0662 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
MLL3_MOUSE
|
||||||
| NC score | 0.079210 (rank : 33) | θ value | 1.38821 (rank : 34) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
MLL3_HUMAN
|
||||||
| NC score | 0.076486 (rank : 34) | θ value | 0.813845 (rank : 31) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
SET1A_HUMAN
|
||||||
| NC score | 0.072534 (rank : 35) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O15047, Q6PIF3, Q8TAJ6 | Gene names | SETD1A, KIAA0339, SET1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET1 (EC 2.1.1.43) (Set1/Ash2 histone methyltransferase complex subunit SET1) (SET-domain-containing protein 1A). | |||||
|
FBXL7_HUMAN
|
||||||
| NC score | 0.067971 (rank : 36) | θ value | θ > 10 (rank : 80) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UJT9, O94926 | Gene names | FBXL7, FBL6, FBL7, KIAA0840 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 7 (F-box and leucine-rich repeat protein 7) (F-box protein FBL6/FBL7). | |||||
|
PHF11_HUMAN
|
||||||
| NC score | 0.066987 (rank : 37) | θ value | θ > 10 (rank : 95) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UIL8, Q9Y5A2 | Gene names | PHF11, BCAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 11 (BRCA1 C-terminus-associated protein) (NY-REN-34 antigen). | |||||
|
MLL2_HUMAN
|
||||||
| NC score | 0.063432 (rank : 38) | θ value | θ > 10 (rank : 94) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
SETD8_HUMAN
|
||||||
| NC score | 0.062971 (rank : 39) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NQR1, Q86W83, Q8TD09 | Gene names | SETD8, PRSET7, SET07, SET8 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H4 lysine-20 specific (EC 2.1.1.43) (Histone H4-K20 methyltransferase) (H4-K20-HMTase) (SET domain-containing lysine methyltransferase 8) (SET domain-containing protein 8) (PR/SET domain-containing protein 07) (PR/SET07) (PR-Set7). | |||||
|
LRC29_HUMAN
|
||||||
| NC score | 0.062579 (rank : 40) | θ value | θ > 10 (rank : 90) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8WV35, Q9UKA0 | Gene names | LRRC29, FBL9, FBXL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 29 (F-box/LRR-repeat protein 9) (F-box and leucine-rich repeat protein 9) (F-box protein FBL9). | |||||
|
SKP2_HUMAN
|
||||||
| NC score | 0.062267 (rank : 41) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q13309, Q8TDZ0, Q8TDZ1, Q9BV69 | Gene names | SKP2, FBXL1 | |||
|
Domain Architecture |

|
|||||
| Description | S-phase kinase-associated protein 2 (F-box protein Skp2) (Cyclin A/CDK2-associated protein p45) (p45skp2) (F-box/LRR-repeat protein 1). | |||||
|
FXL20_HUMAN
|
||||||
| NC score | 0.061168 (rank : 42) | θ value | θ > 10 (rank : 86) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96IG2 | Gene names | FBXL20, FBL2 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 20 (F-box and leucine-rich repeat protein 20) (F-box/LRR-repeat protein 2-like). | |||||
|
FXL20_MOUSE
|
||||||
| NC score | 0.061040 (rank : 43) | θ value | θ > 10 (rank : 87) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9CZV8, Q8BZ95 | Gene names | Fbxl20, Fbl2 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 20 (F-box and leucine-rich repeat protein 20) (F-box/LRR-repeat protein 2-like). | |||||
|
KRA59_HUMAN
|
||||||
| NC score | 0.060926 (rank : 44) | θ value | 0.163984 (rank : 25) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P26371, Q14564 | Gene names | KRTAP5-9, KAP5.9, KRN1, KRTAP5.9, UHSK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-9 (Keratin-associated protein 5.9) (Ultrahigh sulfur keratin-associated protein 5.9) (Keratin, cuticle, ultrahigh sulfur 1) (Keratin, ultra high-sulfur matrix protein A) (UHS keratin A) (UHS KerA). | |||||
|
FXL16_HUMAN
|
||||||
| NC score | 0.060087 (rank : 45) | θ value | θ > 10 (rank : 85) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8N461, Q2MHR2, Q96S14, Q9UJI0 | Gene names | FBXL16, C16orf22, FBL16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 16 (F-box and leucine-rich repeat protein 16). | |||||
|
SKP2_MOUSE
|
||||||
| NC score | 0.060052 (rank : 46) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Z0Z3, Q8BSG7, Q8C8Y9, Q8CB22, Q99J53 | Gene names | Skp2 | |||
|
Domain Architecture |

|
|||||
| Description | S-phase kinase-associated protein 2 (F-box protein Skp2) (Cyclin A/CDK2-associated protein p45) (F-box/WD-40 protein 1) (FWD1). | |||||
|
FXL13_MOUSE
|
||||||
| NC score | 0.059300 (rank : 47) | θ value | θ > 10 (rank : 82) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CDU4, Q8CDE9 | Gene names | Fbxl13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 13 (F-box and leucine-rich repeat protein 13). | |||||
|
SETD8_MOUSE
|
||||||
| NC score | 0.059184 (rank : 48) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q2YDW7, Q8C0J9 | Gene names | SETD8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H4 lysine-20 specific (EC 2.1.1.43) (Histone H4-K20 methyltransferase) (H4-K20-HMTase) (SET domain-containing lysine methyltransferase 8) (SET domain-containing protein 8). | |||||
|
FXL14_MOUSE
|
||||||
| NC score | 0.057573 (rank : 49) | θ value | θ > 10 (rank : 84) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BID8, Q3U2Q9, Q8R5H7, Q8VDT7, Q922N5 | Gene names | Fbxl14, Fbl14, Ppa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 14 (F-box and leucine-rich repeat protein 14). | |||||
|
FXL14_HUMAN
|
||||||
| NC score | 0.057410 (rank : 50) | θ value | θ > 10 (rank : 83) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8N1E6 | Gene names | FBXL14, FBL14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 14 (F-box and leucine-rich repeat protein 14). | |||||
|
FBXL2_MOUSE
|
||||||
| NC score | 0.056765 (rank : 51) | θ value | θ > 10 (rank : 79) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BH16, Q8BXM4 | Gene names | Fbxl2, Fbl2 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 2 (F-box and leucine-rich repeat protein 2). | |||||
|
FBXL2_HUMAN
|
||||||
| NC score | 0.054343 (rank : 52) | θ value | θ > 10 (rank : 78) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UKC9, Q9NVQ8, Q9UK27, Q9UKA5, Q9Y3Y9 | Gene names | FBXL2, FBL2, FBL3 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/LRR-repeat protein 2 (F-box and leucine-rich repeat protein 2) (F-box protein FBL2/FBL3). | |||||
|
BAZ2A_HUMAN
|
||||||
| NC score | 0.054220 (rank : 53) | θ value | 1.81305 (rank : 35) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 780 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9UIF9, O00536, O15030, Q96H26 | Gene names | BAZ2A, KIAA0314, TIP5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5) (hWALp3). | |||||
|
BAZ2A_MOUSE
|
||||||
| NC score | 0.053472 (rank : 54) | θ value | 1.81305 (rank : 36) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q91YE5 | Gene names | Baz2a, Tip5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5). | |||||
|
FXL13_HUMAN
|
||||||
| NC score | 0.051236 (rank : 55) | θ value | θ > 10 (rank : 81) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8NEE6, Q6UVW7, Q6UVW8, Q75MN5, Q86UJ5, Q8N7Y4, Q8TCL2, Q8WUF9, Q8WUG0 | Gene names | FBXL13, FBL13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 13 (F-box and leucine-rich repeat protein 13). | |||||
|
SRCH_HUMAN
|
||||||
| NC score | 0.050147 (rank : 56) | θ value | 2.36792 (rank : 41) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P23327, Q504Y6 | Gene names | HRC, HCP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcoplasmic reticulum histidine-rich calcium-binding protein precursor. | |||||
|
LAMB2_MOUSE
|
||||||
| NC score | 0.049025 (rank : 57) | θ value | 0.0330416 (rank : 24) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q61292, Q62182 | Gene names | Lamb2, Lams | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-2 chain precursor (S-laminin) (S-LAM). | |||||
|
KRA54_HUMAN
|
||||||
| NC score | 0.045108 (rank : 58) | θ value | 4.03905 (rank : 48) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 270 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q6L8H1 | Gene names | KRTAP5-4, KAP5.4, KRTAP5.4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-4 (Keratin-associated protein 5.4) (Ultrahigh sulfur keratin-associated protein 5.4). | |||||
|
DMRTD_HUMAN
|
||||||
| NC score | 0.040852 (rank : 59) | θ value | 0.47712 (rank : 27) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8IXT2, Q8N6Q2, Q96M39, Q96SD4 | Gene names | DMRTC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Doublesex- and mab-3-related transcription factor C2. | |||||
|
LAMB2_HUMAN
|
||||||
| NC score | 0.040402 (rank : 60) | θ value | 2.36792 (rank : 40) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 692 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P55268, Q16321 | Gene names | LAMB2, LAMS | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-2 chain precursor (S-laminin) (Laminin B1s chain). | |||||
|
KRA51_HUMAN
|
||||||
| NC score | 0.038760 (rank : 61) | θ value | 4.03905 (rank : 47) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 288 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6L8H4 | Gene names | KRTAP5-1, KAP5.1, KRTAP5.1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-1 (Keratin-associated protein 5.1) (Ultrahigh sulfur keratin-associated protein 5.1). | |||||
|
LAMB1_HUMAN
|
||||||
| NC score | 0.030817 (rank : 62) | θ value | 6.88961 (rank : 60) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 1018 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P07942 | Gene names | LAMB1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-1 chain precursor (Laminin B1 chain). | |||||
|
NET2_HUMAN
|
||||||
| NC score | 0.030564 (rank : 63) | θ value | 6.88961 (rank : 63) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O00634 | Gene names | NTN2L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin-2-like protein precursor. | |||||
|
NET2_MOUSE
|
||||||
| NC score | 0.029935 (rank : 64) | θ value | 8.99809 (rank : 72) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9R1A3, Q9QY49, Q9WVA6 | Gene names | Ntn2l, Ntn3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Netrin-2-like protein precursor (Netrin-3). | |||||
|
GA2L2_MOUSE
|
||||||
| NC score | 0.029301 (rank : 65) | θ value | 3.0926 (rank : 43) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5SSG4 | Gene names | Gas2l2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GAS2-like protein 2 (Growth arrest-specific 2-like 2). | |||||
|
ITB6_MOUSE
|
||||||
| NC score | 0.028396 (rank : 66) | θ value | 1.38821 (rank : 33) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9Z0T9 | Gene names | Itgb6 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-6 precursor. | |||||
|
LAMC3_HUMAN
|
||||||
| NC score | 0.027735 (rank : 67) | θ value | 6.88961 (rank : 61) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 588 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9Y6N6 | Gene names | LAMC3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-3 chain precursor (Laminin 12 gamma 3 subunit). | |||||
|
BSN_MOUSE
|
||||||
| NC score | 0.027454 (rank : 68) | θ value | 4.03905 (rank : 46) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
LAMC3_MOUSE
|
||||||
| NC score | 0.027433 (rank : 69) | θ value | 6.88961 (rank : 62) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9R0B6, Q9WTW6 | Gene names | Lamc3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-3 chain precursor (Laminin 12 gamma 3 subunit). | |||||
|
TNR25_HUMAN
|
||||||
| NC score | 0.025811 (rank : 70) | θ value | 4.03905 (rank : 51) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q93038, O00275, O00276, O00277, O00278, O00279, O00280, O14865, O14866, P78507, P78515, Q92983, Q93036, Q93037, Q99722, Q99830, Q99831, Q9BY86, Q9UME0, Q9UME1, Q9UME5 | Gene names | TNFRSF25, APO3, DDR3, DR3, TNFRSF12, WSL, WSL1 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 25 precursor (WSL-1 protein) (Apoptosis-mediating receptor DR3) (Apoptosis-mediating receptor TRAMP) (Death domain receptor 3) (WSL protein) (Apoptosis- inducing receptor AIR) (Apo-3) (Lymphocyte-associated receptor of death) (LARD). | |||||
|
LAMC1_HUMAN
|
||||||
| NC score | 0.025583 (rank : 71) | θ value | 4.03905 (rank : 49) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 922 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P11047 | Gene names | LAMC1, LAMB2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-1 chain precursor (Laminin B2 chain). | |||||
|
DFRC_MOUSE
|
||||||
| NC score | 0.025170 (rank : 72) | θ value | 5.27518 (rank : 54) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P17533, Q9QVW7 | Gene names | Defcr-rs1 | |||
|
Domain Architecture |

|
|||||
| Description | Defensin-related sequence cryptdin peptide precursor (Cryptdin-related protein 1C) (CRS1C). | |||||
|
MEGF8_MOUSE
|
||||||
| NC score | 0.024979 (rank : 73) | θ value | 1.81305 (rank : 37) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 520 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P60882, Q80TR3, Q80V41, Q8BMN9, Q8JZW7, Q8K0J3 | Gene names | Megf8, Egfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 8 (EGF-like domain- containing protein 4) (Multiple EGF-like domain protein 4). | |||||
|
LAMA5_HUMAN
|
||||||
| NC score | 0.023643 (rank : 74) | θ value | 8.99809 (rank : 71) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 976 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O15230, Q8WZA7, Q9H1P1 | Gene names | LAMA5, KIAA0533, KIAA1907 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-5 chain precursor. | |||||
|
DAB1_HUMAN
|
||||||
| NC score | 0.022989 (rank : 75) | θ value | 5.27518 (rank : 53) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O75553, Q9NYA8 | Gene names | DAB1 | |||
|
Domain Architecture |

|
|||||
| Description | Disabled homolog 1. | |||||
|
MKRN1_MOUSE
|
||||||
| NC score | 0.022689 (rank : 76) | θ value | 0.813845 (rank : 30) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9QXP6 | Gene names | Mkrn1 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-1. | |||||
|
CHD5_HUMAN
|
||||||
| NC score | 0.020546 (rank : 77) | θ value | 2.36792 (rank : 39) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8TDI0, O75032, Q5TG89, Q9UFR9 | Gene names | CHD5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain helicase-DNA-binding protein 5 (EC 3.6.1.-) (ATP- dependent helicase CHD5) (CHD-5). | |||||
|
KRA58_HUMAN
|
||||||
| NC score | 0.019507 (rank : 78) | θ value | 6.88961 (rank : 59) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 324 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O75690, Q6L8G7, Q6UTX6 | Gene names | KRTAP5-8, KAP5.8, KRTAP5.8, UHSKB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin-associated protein 5-8 (Keratin-associated protein 5.8) (Ultrahigh sulfur keratin-associated protein 5.8) (Keratin, ultra high-sulfur matrix protein B) (UHS keratin B) (UHS KerB). | |||||
|
MAGE1_HUMAN
|
||||||
| NC score | 0.019466 (rank : 79) | θ value | 1.06291 (rank : 32) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
ITB1_MOUSE
|
||||||
| NC score | 0.019062 (rank : 80) | θ value | 8.99809 (rank : 68) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P09055 | Gene names | Itgb1 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-1 precursor (Fibronectin receptor subunit beta) (Integrin VLA-4 subunit beta) (CD29 antigen). | |||||
|
RSPO4_HUMAN
|
||||||
| NC score | 0.017528 (rank : 81) | θ value | 4.03905 (rank : 50) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q2I0M5, Q9UGB2 | Gene names | RSPO4, C20orf182 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | R-spondin-4 precursor (Roof plate-specific spondin-4) (hRspo4). | |||||
|
NTAL_HUMAN
|
||||||
| NC score | 0.017364 (rank : 82) | θ value | 8.99809 (rank : 73) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9GZY6, Q9BXX8, Q9NZY9 | Gene names | LAT2, LAB, NTAL, WBS15, WBSCR15, WBSCR5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Linker for activation of T-cells family member 2 (Non-T-cell activation linker) (Linker for activation of B-cells) (Membrane- associated adapter molecule) (Williams-Beuren syndrome chromosome region 15 protein). | |||||
|
PHLPP_HUMAN
|
||||||
| NC score | 0.016774 (rank : 83) | θ value | 0.47712 (rank : 28) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 604 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O60346, Q641Q7, Q6P4C4, Q6PJI6, Q86TN6, Q96FK2, Q9NUY1 | Gene names | PHLPP, KIAA0606, PLEKHE1, SCOP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH domain leucine-rich repeat-containing protein phosphatase (EC 3.1.3.16) (PH domain leucine-rich repeat protein phosphatase) (Pleckstrin homology domain-containing family E protein 1) (Suprachiasmatic nucleus circadian oscillatory protein) (hSCOP). | |||||
|
ATS18_MOUSE
|
||||||
| NC score | 0.013823 (rank : 84) | θ value | 0.279714 (rank : 26) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q4VC17, Q8BZD1 | Gene names | Adamts18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAMTS-18 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase with thrombospondin motifs 18) (ADAM-TS 18) (ADAM-TS18). | |||||
|
CSPG5_MOUSE
|
||||||
| NC score | 0.013016 (rank : 85) | θ value | 8.99809 (rank : 67) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q71M36, Q71M37, Q7TNT8, Q8BPJ5, Q9QY32 | Gene names | Cspg5, Caleb, Ngc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate proteoglycan 5 precursor (Neuroglycan C) (Acidic leucine-rich EGF-like domain-containing brain protein). | |||||
|
AMOT_HUMAN
|
||||||
| NC score | 0.013014 (rank : 86) | θ value | 5.27518 (rank : 52) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 712 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q4VCS5, Q504X5, Q9HD27, Q9UPT1 | Gene names | AMOT, KIAA1071 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin. | |||||
|
ADAM2_MOUSE
|
||||||
| NC score | 0.010827 (rank : 87) | θ value | 6.88961 (rank : 56) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q60718, Q60814, Q9D4G3, Q9QWJ0 | Gene names | Adam2, Ftnb | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 2 precursor (A disintegrin and metalloproteinase domain 2) (Fertilin subunit beta) (PH-30) (PH30) (PH30-beta). | |||||
|
FXR2_MOUSE
|
||||||
| NC score | 0.010326 (rank : 88) | θ value | 6.88961 (rank : 58) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9WVR4, Q9WVR5 | Gene names | Fxr2, Fxr2h | |||
|
Domain Architecture |

|
|||||
| Description | Fragile X mental retardation syndrome-related protein 2. | |||||
|
JAG2_MOUSE
|
||||||
| NC score | 0.010214 (rank : 89) | θ value | 8.99809 (rank : 69) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9QYE5, O55139, O70219 | Gene names | Jag2 | |||
|
Domain Architecture |

|
|||||
| Description | Jagged-2 precursor (Jagged2). | |||||
|
MMP17_HUMAN
|
||||||
| NC score | 0.007833 (rank : 90) | θ value | 3.0926 (rank : 44) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9ULZ9, Q14850 | Gene names | MMP17, MT4MMP | |||
|
Domain Architecture |

|
|||||
| Description | Matrix metalloproteinase-17 precursor (EC 3.4.24.-) (MMP-17) (Membrane-type matrix metalloproteinase 4) (MT-MMP 4) (Membrane-type-4 matrix metalloproteinase) (MT4-MMP). | |||||
|
WHRN_MOUSE
|
||||||
| NC score | 0.007231 (rank : 91) | θ value | 6.88961 (rank : 65) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80VW5, Q80TC2, Q80VW4 | Gene names | Whrn, Kiaa1526 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Whirlin. | |||||
|
KCD12_MOUSE
|
||||||
| NC score | 0.007120 (rank : 92) | θ value | 8.99809 (rank : 70) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6WVG3 | Gene names | Kctd12, Pfet1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD12 (Pfetin) (Predominantly fetal expressed T1 domain). | |||||
|
GCR_MOUSE
|
||||||
| NC score | 0.006342 (rank : 93) | θ value | 5.27518 (rank : 55) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P06537, Q61628, Q61629 | Gene names | Nr3c1, Grl, Grl1 | |||
|
Domain Architecture |

|
|||||
| Description | Glucocorticoid receptor (GR). | |||||
|
ZN507_HUMAN
|
||||||
| NC score | 0.005437 (rank : 94) | θ value | 2.36792 (rank : 42) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 739 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8TCN5 | Gene names | ZNF507 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 507. | |||||
|
RORG_HUMAN
|
||||||
| NC score | 0.005221 (rank : 95) | θ value | 6.88961 (rank : 64) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P51449, Q8N5V7, Q8NCY8 | Gene names | RORC, NR1F3, RORG, RZRG | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor ROR-gamma (Nuclear receptor RZR-gamma). | |||||
|
COBA1_MOUSE
|
||||||
| NC score | 0.004031 (rank : 96) | θ value | 6.88961 (rank : 57) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 487 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61245, Q64047 | Gene names | Col11a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XI) chain precursor. | |||||
|
TPR_HUMAN
|
||||||
| NC score | 0.004024 (rank : 97) | θ value | 3.0926 (rank : 45) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
ZN672_HUMAN
|
||||||
| NC score | 0.003816 (rank : 98) | θ value | 0.47712 (rank : 29) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q499Z4, Q96H65, Q96IM3, Q9H6G5 | Gene names | ZNF672 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 672. | |||||
|
ZN335_HUMAN
|
||||||
| NC score | 0.002466 (rank : 99) | θ value | 6.88961 (rank : 66) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 973 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9H4Z2, Q9H684 | Gene names | ZNF335 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 335. | |||||
|
TRI10_MOUSE
|
||||||
| NC score | 0.002216 (rank : 100) | θ value | 8.99809 (rank : 75) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9WUH5, Q80WA9, Q9CY03 | Gene names | Trim10, Herf1, Rnf9 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 10 (RING finger protein 9) (Hematopoietic RING finger 1). | |||||
|
ZN628_HUMAN
|
||||||
| NC score | 0.002185 (rank : 101) | θ value | 1.81305 (rank : 38) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 814 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q5EBL2, Q86X34 | Gene names | ZNF628 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 628. | |||||
|
ZN689_HUMAN
|
||||||
| NC score | 0.000820 (rank : 102) | θ value | 8.99809 (rank : 76) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 736 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96CS4, Q658J5 | Gene names | ZNF689 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 689. | |||||
|
ZN689_MOUSE
|
||||||
| NC score | 0.000812 (rank : 103) | θ value | 8.99809 (rank : 77) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 736 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BKK5, Q8C1C6 | Gene names | Znf689, Zfp689 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 689. | |||||
|
OBSCN_HUMAN
|
||||||
| NC score | 0.000646 (rank : 104) | θ value | 8.99809 (rank : 74) | |||
| Query Neighborhood Hits | 77 | Target Neighborhood Hits | 1931 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q5VST9, Q2A664, Q5T7G8, Q5T7G9, Q5VSU2, Q86YC7, Q8NHN0, Q8NHN1, Q8NHN2, Q8NHN4, Q8NHN5, Q8NHN6, Q8NHN7, Q8NHN8, Q8NHN9, Q96AA2, Q9HCD3, Q9HCL6 | Gene names | OBSCN, KIAA1556, KIAA1639 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Obscurin (Obscurin-myosin light chain kinase) (Obscurin-MLCK) (Obscurin-RhoGEF). | |||||