Please be patient as the page loads
|
HEY2_MOUSE
|
||||||
| SwissProt Accessions | Q9QUS4, Q3TZ99, Q8CD44 | Gene names | Hey2, Chf1, Herp, Herp1, Hesr2, Hrt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES-related repressor protein 2). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
HEY2_MOUSE
|
||||||
| θ value | 5.26011e-125 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | Q9QUS4, Q3TZ99, Q8CD44 | Gene names | Hey2, Chf1, Herp, Herp1, Hesr2, Hrt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES-related repressor protein 2). | |||||
|
HEY2_HUMAN
|
||||||
| θ value | 4.61877e-113 (rank : 2) | NC score | 0.996757 (rank : 2) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9UBP5 | Gene names | HEY2, CHF1, GRL, HERP, HERP1, HRT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Cardiovascular helix-loop-helix factor 1) (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES- related repressor protein 2) (Protein gridlock homolog). | |||||
|
HEY1_HUMAN
|
||||||
| θ value | 1.0458e-56 (rank : 3) | NC score | 0.961396 (rank : 3) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9Y5J3, Q5TZS3, Q9NYP4 | Gene names | HEY1, CHF2, HESR1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1) (Cardiovascular helix-loop-helix factor 2) (HES-related repressor protein 2) (HERP2). | |||||
|
HEY1_MOUSE
|
||||||
| θ value | 6.56568e-51 (rank : 4) | NC score | 0.960095 (rank : 4) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9WV93 | Gene names | Hey1, Hesr1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1). | |||||
|
HES1_HUMAN
|
||||||
| θ value | 5.60996e-18 (rank : 5) | NC score | 0.787111 (rank : 7) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q14469 | Gene names | HES1, HL, HRY | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor HES-1 (Hairy and enhancer of split 1) (Hairy- like) (HHL) (Hairy homolog). | |||||
|
HES1_MOUSE
|
||||||
| θ value | 5.60996e-18 (rank : 6) | NC score | 0.787638 (rank : 6) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P35428 | Gene names | Hes1, Hes-1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-1 (Hairy and enhancer of split 1). | |||||
|
HES4_HUMAN
|
||||||
| θ value | 8.10077e-17 (rank : 7) | NC score | 0.813024 (rank : 5) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9HCC6 | Gene names | HES4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-4 (Hairy and enhancer of split 4) (bHLH factor Hes4). | |||||
|
HES2_HUMAN
|
||||||
| θ value | 4.30538e-10 (rank : 8) | NC score | 0.720168 (rank : 10) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9Y543, Q96EN4, Q9Y542 | Gene names | HES2 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor HES-2 (Hairy and enhancer of split 2). | |||||
|
HES2_MOUSE
|
||||||
| θ value | 5.62301e-10 (rank : 9) | NC score | 0.724373 (rank : 9) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O54792 | Gene names | Hes2 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor HES-2 (Hairy and enhancer of split 2). | |||||
|
HES6_HUMAN
|
||||||
| θ value | 9.59137e-10 (rank : 10) | NC score | 0.684061 (rank : 11) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q96HZ4, Q8N2J2, Q96T93, Q9P2S3 | Gene names | HES6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription cofactor HES-6 (Hairy and enhancer of split 6) (C- HAIRY1). | |||||
|
HES6_MOUSE
|
||||||
| θ value | 4.76016e-09 (rank : 11) | NC score | 0.670130 (rank : 12) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9JHE6, Q3U2D7 | Gene names | Hes6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription cofactor HES-6 (Hairy and enhancer of split 6). | |||||
|
HES5_MOUSE
|
||||||
| θ value | 5.26297e-08 (rank : 12) | NC score | 0.744202 (rank : 8) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P70120, Q8BVI1 | Gene names | Hes5, Hes-5 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-5 (Hairy and enhancer of split 5). | |||||
|
BHLH3_MOUSE
|
||||||
| θ value | 6.87365e-08 (rank : 13) | NC score | 0.561823 (rank : 16) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q99PV5 | Gene names | Bhlhb3, Dec2 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (mDEC2). | |||||
|
BHLH3_HUMAN
|
||||||
| θ value | 3.41135e-07 (rank : 14) | NC score | 0.558761 (rank : 17) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9C0J9 | Gene names | BHLHB3, DEC2, SHARP1 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (hDEC2) (Enhancer-of-split and hairy-related protein 1) (SHARP-1). | |||||
|
BHLH2_HUMAN
|
||||||
| θ value | 5.81887e-07 (rank : 15) | NC score | 0.535193 (rank : 18) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O14503, Q96TD3 | Gene names | BHLHB2, DEC1, SHARP2, STRA13 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 2 (bHLHB2) (Differentially expressed in chondrocytes protein 1) (DEC1) (Enhancer-of-split and hairy-related protein 2) (SHARP-2) (Stimulated with retinoic acid 13). | |||||
|
BHLH2_MOUSE
|
||||||
| θ value | 2.88788e-06 (rank : 16) | NC score | 0.525737 (rank : 19) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O35185, P97289 | Gene names | Bhlhb2, Clast5, Stra13 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 2 (bHLHB2) (Stimulated with retinoic acid 13) (E47 interaction protein 1) (eipl). | |||||
|
HES7_MOUSE
|
||||||
| θ value | 4.92598e-06 (rank : 17) | NC score | 0.584886 (rank : 14) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8BKT2, Q99JA6 | Gene names | Hes7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor HES-7 (Hairy and enhancer of split 7) (bHLH factor Hes7). | |||||
|
HES7_HUMAN
|
||||||
| θ value | 1.09739e-05 (rank : 18) | NC score | 0.585908 (rank : 13) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9BYE0 | Gene names | HES7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor HES-7 (Hairy and enhancer of split 7) (bHLH factor Hes7). | |||||
|
BMAL1_HUMAN
|
||||||
| θ value | 9.29e-05 (rank : 19) | NC score | 0.279833 (rank : 28) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O00327, O00313, O00314, O00315, O00316, O00317, Q99631, Q99649 | Gene names | ARNTL, BMAL1, MOP3 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Member of PAS protein 3) (Basic-helix-loop- helix-PAS orphan MOP3) (bHLH-PAS protein JAP3). | |||||
|
BMAL1_MOUSE
|
||||||
| θ value | 9.29e-05 (rank : 20) | NC score | 0.283651 (rank : 26) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9WTL8, O88295, Q921S4, Q9R0U2, Q9WTL9 | Gene names | Arntl, Bmal1 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Arnt3). | |||||
|
HES3_MOUSE
|
||||||
| θ value | 0.00102713 (rank : 21) | NC score | 0.568544 (rank : 15) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q61657, O09083, O09150 | Gene names | Hes3, Hes-3 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor HES-3 (Hairy and enhancer of split 3). | |||||
|
MLX_HUMAN
|
||||||
| θ value | 0.00134147 (rank : 22) | NC score | 0.315946 (rank : 21) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9UH92, Q53XM6, Q96FL2, Q9H2V0, Q9H2V1, Q9H2V2, Q9NXN3 | Gene names | MLX, TCFL4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
MLX_MOUSE
|
||||||
| θ value | 0.00134147 (rank : 23) | NC score | 0.320280 (rank : 20) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O08609, Q9EQJ7, Q9EQJ8 | Gene names | Mlx, Tcfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
ARNT2_HUMAN
|
||||||
| θ value | 0.00298849 (rank : 24) | NC score | 0.228539 (rank : 36) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9HBZ2, O15024 | Gene names | ARNT2, KIAA0307 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
ARNT2_MOUSE
|
||||||
| θ value | 0.00298849 (rank : 25) | NC score | 0.229817 (rank : 35) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q61324 | Gene names | Arnt2 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
USF1_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 26) | NC score | 0.312070 (rank : 22) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P22415 | Gene names | USF1, USF | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF1_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 27) | NC score | 0.311969 (rank : 23) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q61069 | Gene names | Usf1, Usf | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF2_MOUSE
|
||||||
| θ value | 0.00509761 (rank : 28) | NC score | 0.301537 (rank : 24) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q64705, Q3UIL2, Q8VE59 | Gene names | Usf2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (Major late transcription factor 2). | |||||
|
USF2_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 29) | NC score | 0.298020 (rank : 25) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q15853, O00671, O00709, Q05750, Q07952, Q15851, Q15852 | Gene names | USF2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (FOS- interacting protein) (FIP) (Major late transcription factor 2). | |||||
|
ARNT_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 30) | NC score | 0.225281 (rank : 37) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P27540 | Gene names | ARNT | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
ARNT_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 31) | NC score | 0.221920 (rank : 38) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P53762, Q60661 | Gene names | Arnt | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
SRBP2_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 32) | NC score | 0.242855 (rank : 33) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q12772, Q9UH04 | Gene names | SREBF2, SREBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 2 (SREBP-2) (Sterol regulatory element-binding transcription factor 2). | |||||
|
TFE3_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 33) | NC score | 0.281243 (rank : 27) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P19532, Q92757, Q92758, Q99964 | Gene names | TFE3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E3. | |||||
|
SRBP1_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 34) | NC score | 0.241524 (rank : 34) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P36956, Q16062 | Gene names | SREBF1, SREBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
TFEB_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 35) | NC score | 0.256768 (rank : 32) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P19484, Q709B3, Q7Z6P9, Q9BRJ5, Q9UJD8 | Gene names | TFEB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
TFEB_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 36) | NC score | 0.260350 (rank : 31) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9R210 | Gene names | Tfeb, Tcfeb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
MITF_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 37) | NC score | 0.269763 (rank : 29) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O75030, Q14841, Q9P2V0, Q9P2V1, Q9P2V2, Q9P2Y8 | Gene names | MITF | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
MITF_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 38) | NC score | 0.267196 (rank : 30) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q08874, O08885, O88203, Q08843, Q60781, Q60782, Q9JIJ0, Q9JIJ1, Q9JIJ2, Q9JIJ3, Q9JIJ4, Q9JIJ5, Q9JIJ6, Q9JKX9 | Gene names | Mitf, Bw, Mi, Vit | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
SRBP1_MOUSE
|
||||||
| θ value | 0.125558 (rank : 39) | NC score | 0.219028 (rank : 39) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9WTN3 | Gene names | Srebf1, Srebp1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
TAL1_HUMAN
|
||||||
| θ value | 0.125558 (rank : 40) | NC score | 0.115563 (rank : 46) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P17542 | Gene names | TAL1, SCL, TCL5 | |||
|
Domain Architecture |
|
|||||
| Description | T-cell acute lymphocytic leukemia-1 protein (TAL-1 protein) (Stem cell protein) (T-cell leukemia/lymphoma-5 protein). | |||||
|
CLOCK_MOUSE
|
||||||
| θ value | 0.365318 (rank : 41) | NC score | 0.165378 (rank : 40) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O08785 | Gene names | Clock | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (mCLOCK). | |||||
|
LYL1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 42) | NC score | 0.097462 (rank : 57) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P12980, O76102 | Gene names | LYL1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein lyl-1 (Lymphoblastic leukemia-derived sequence 1). | |||||
|
CLOCK_HUMAN
|
||||||
| θ value | 0.62314 (rank : 43) | NC score | 0.164215 (rank : 41) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O15516, O14516, Q9UIT8 | Gene names | CLOCK, KIAA0334 | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (hCLOCK). | |||||
|
LYL1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 44) | NC score | 0.095328 (rank : 58) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P27792 | Gene names | Lyl1, Lyl-1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein lyl-1 (Lymphoblastic leukemia-derived sequence 1). | |||||
|
HNF1A_MOUSE
|
||||||
| θ value | 0.813845 (rank : 45) | NC score | 0.024203 (rank : 110) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P22361 | Gene names | Tcf1, Hnf-1, Hnf-1a, Hnf1a | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 1-alpha (HNF-1A) (Liver-specific transcription factor LF-B1) (LFB1). | |||||
|
TAL1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 46) | NC score | 0.108774 (rank : 51) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P22091 | Gene names | Tal1, Scl, Tal-1 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell acute lymphocytic leukemia-1 protein homolog (TAL-1 protein) (Stem cell protein). | |||||
|
ALAT1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 47) | NC score | 0.021485 (rank : 112) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P24298, P78398, Q93076 | Gene names | GPT, AAT1, GPT1 | |||
|
Domain Architecture |

|
|||||
| Description | Alanine aminotransferase 1 (EC 2.6.1.2) (ALT1) (Glutamic--pyruvic transaminase 1) (GPT 1) (Glutamic--alanine transaminase 1). | |||||
|
BHLH8_HUMAN
|
||||||
| θ value | 1.38821 (rank : 48) | NC score | 0.083304 (rank : 67) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q7RTS1 | Gene names | BHLHB8, MIST1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Class B basic helix-loop-helix protein 8 (bHLHB8) (Muscle, intestine and stomach expression 1) (MIST-1). | |||||
|
BHLH8_MOUSE
|
||||||
| θ value | 1.38821 (rank : 49) | NC score | 0.082197 (rank : 68) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9QYC3, Q9QYE4 | Gene names | Bhlhb8, Mist1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Class B basic helix-loop-helix protein 8 (bHLHB8) (Muscle, intestine and stomach expression 1) (MIST-1). | |||||
|
TAL2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 50) | NC score | 0.089189 (rank : 63) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q62282 | Gene names | Tal2 | |||
|
Domain Architecture |
|
|||||
| Description | T-cell acute lymphocytic leukemia-2 protein homolog (TAL-2 protein). | |||||
|
APLP1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 51) | NC score | 0.023408 (rank : 111) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q03157, Q8VC38 | Gene names | Aplp1 | |||
|
Domain Architecture |
|
|||||
| Description | Amyloid-like protein 1 precursor (APLP) (APLP-1) [Contains: C30]. | |||||
|
ID1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 52) | NC score | 0.075512 (rank : 71) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P20067, Q61101, Q9D897 | Gene names | Id1, Id, Id-1, Idb1 | |||
|
Domain Architecture |
|
|||||
| Description | DNA-binding protein inhibitor ID-1 (Inhibitor of DNA binding 1). | |||||
|
SIM2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 53) | NC score | 0.113097 (rank : 47) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q14190, O60766, Q15470, Q15471, Q15472, Q15473, Q16532 | Gene names | SIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2. | |||||
|
SIM2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 54) | NC score | 0.111543 (rank : 50) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q61079, O35391, Q61046, Q61904 | Gene names | Sim2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2 (SIM transcription factor) (mSIM). | |||||
|
ELK3_MOUSE
|
||||||
| θ value | 2.36792 (rank : 55) | NC score | 0.015843 (rank : 113) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P41971, P97747, Q62346 | Gene names | Elk3, Erp, Net | |||
|
Domain Architecture |
|
|||||
| Description | ETS domain-containing protein Elk-3 (ETS-related protein NET) (ETS- related protein ERP). | |||||
|
ID4_HUMAN
|
||||||
| θ value | 2.36792 (rank : 56) | NC score | 0.071522 (rank : 73) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P47928, Q13005 | Gene names | ID4 | |||
|
Domain Architecture |
|
|||||
| Description | DNA-binding protein inhibitor ID-4 (Inhibitor of DNA binding 4). | |||||
|
ID4_MOUSE
|
||||||
| θ value | 2.36792 (rank : 57) | NC score | 0.071177 (rank : 74) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P41139 | Gene names | Id4, Id-4, Idb4 | |||
|
Domain Architecture |
|
|||||
| Description | DNA-binding protein inhibitor ID-4 (Inhibitor of DNA binding 4). | |||||
|
NCOA1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 58) | NC score | 0.112560 (rank : 49) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q15788, O00150, O43792, O43793, Q13071, Q13420, Q6GVI5, Q7KYV3 | Gene names | NCOA1, SRC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (RIP160) (Protein Hin-2) (NY-REN-52 antigen). | |||||
|
NCOA1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 59) | NC score | 0.113081 (rank : 48) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P70365, P70366, Q61202, Q8CBI9 | Gene names | Ncoa1, Src1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (Nuclear receptor coactivator protein 1) (mNRC-1). | |||||
|
NDF4_HUMAN
|
||||||
| θ value | 3.0926 (rank : 60) | NC score | 0.054198 (rank : 102) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9HD90 | Gene names | NEUROD4 | |||
|
Domain Architecture |
|
|||||
| Description | Neurogenic differentiation factor 4 (NeuroD4). | |||||
|
NDF4_MOUSE
|
||||||
| θ value | 3.0926 (rank : 61) | NC score | 0.052024 (rank : 105) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O09105 | Gene names | Neurod4, Ath3, Atoh3 | |||
|
Domain Architecture |
|
|||||
| Description | Neurogenic differentiation factor 4 (NeuroD4) (Atonal protein homolog 3) (Helix-loop-helix protein mATH-3) (MATH3). | |||||
|
ALS2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 62) | NC score | 0.011995 (rank : 116) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q920R0, Q8JZR1, Q9CXJ3 | Gene names | Als2 | |||
|
Domain Architecture |

|
|||||
| Description | Alsin (Amyotrophic lateral sclerosis protein 2 homolog). | |||||
|
ELK3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 63) | NC score | 0.014468 (rank : 114) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P41970, Q6FG57, Q6GU29, Q9UD17 | Gene names | ELK3, NET, SAP2 | |||
|
Domain Architecture |
|
|||||
| Description | ETS domain-containing protein Elk-3 (ETS-related protein NET) (ETS- related protein ERP) (SRF accessory protein 2) (SAP-2). | |||||
|
NPAS3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 64) | NC score | 0.100058 (rank : 56) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8IXF0, Q86US6, Q86US7, Q8IXF2, Q9BY81, Q9H323, Q9Y4L8 | Gene names | NPAS3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
NPAS3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 65) | NC score | 0.101216 (rank : 55) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9QZQ0, Q9EQP4 | Gene names | Npas3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
SIM1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 66) | NC score | 0.107287 (rank : 53) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P81133, Q5TDP7 | Gene names | SIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1. | |||||
|
SIM1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 67) | NC score | 0.107336 (rank : 52) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q61045, O70284, P70183 | Gene names | Sim1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1 (mSIM1). | |||||
|
TAL2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 68) | NC score | 0.074721 (rank : 72) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q16559 | Gene names | TAL2 | |||
|
Domain Architecture |
|
|||||
| Description | T-cell acute lymphocytic leukemia-2 protein (TAL-2 protein). | |||||
|
WDR22_MOUSE
|
||||||
| θ value | 4.03905 (rank : 69) | NC score | 0.012542 (rank : 115) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80T85, Q80VT3, Q80ZW6, Q8BIP8, Q8BVK5, Q8K0Q6 | Gene names | Wdr22, Kiaa1824 | |||
|
Domain Architecture |
|
|||||
| Description | WD repeat protein 22. | |||||
|
ID1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 70) | NC score | 0.064752 (rank : 79) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P41134, O00651, O00652, Q16371, Q16377, Q5TE66, Q5TE67, Q969Z7, Q9H0Z5, Q9H109 | Gene names | ID1, ID | |||
|
Domain Architecture |
|
|||||
| Description | DNA-binding protein inhibitor ID-1 (Inhibitor of DNA binding 1). | |||||
|
TWST2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 71) | NC score | 0.053393 (rank : 104) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8WVJ9 | Gene names | TWIST2, DERMO1 | |||
|
Domain Architecture |
|
|||||
| Description | Twist-related protein 2 (Dermis expressed protein 1) (Dermo-1). | |||||
|
TWST2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 72) | NC score | 0.053462 (rank : 103) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9D030 | Gene names | Twist2, Dermo1 | |||
|
Domain Architecture |
|
|||||
| Description | Twist-related protein 2 (Dermis expressed protein 1) (Dermo-1). | |||||
|
MAX_HUMAN
|
||||||
| θ value | 6.88961 (rank : 73) | NC score | 0.149410 (rank : 43) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P61244, P25912, P52163 | Gene names | MAX | |||
|
Domain Architecture |

|
|||||
| Description | Protein max (Myc-associated factor X). | |||||
|
MAX_MOUSE
|
||||||
| θ value | 6.88961 (rank : 74) | NC score | 0.153696 (rank : 42) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P28574 | Gene names | Max, Myn | |||
|
Domain Architecture |

|
|||||
| Description | Protein max (Myc-associated factor X) (Protein myn) (Myc-binding novel HLH/LZ protein). | |||||
|
NGN3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 75) | NC score | 0.048495 (rank : 109) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P70661 | Gene names | Neurog3, Ath4b, Atoh5, Ngn3 | |||
|
Domain Architecture |
|
|||||
| Description | Neurogenin 3 (Atonal protein homolog 5) (Helix-loop-helix protein mATH-4B) (MATH4B). | |||||
|
TFAP4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 76) | NC score | 0.106661 (rank : 54) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q01664, O60409 | Gene names | TFAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor AP-4 (Activating enhancer-binding protein 4). | |||||
|
TWST1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 77) | NC score | 0.056372 (rank : 95) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q15672, Q92487, Q99804 | Gene names | TWIST1, TWIST | |||
|
Domain Architecture |

|
|||||
| Description | Twist-related protein 1 (H-twist). | |||||
|
TWST1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 78) | NC score | 0.056168 (rank : 96) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P26687 | Gene names | Twist1, Twist | |||
|
Domain Architecture |

|
|||||
| Description | Twist-related protein 1 (M-twist). | |||||
|
GPTC2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 79) | NC score | 0.009024 (rank : 117) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7TQC7, Q8BNJ9, Q8BPM1, Q8CDH9 | Gene names | Gpatc2 | |||
|
Domain Architecture |

|
|||||
| Description | G patch domain-containing protein 2. | |||||
|
HAND1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 80) | NC score | 0.059244 (rank : 88) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q64279, Q61099 | Gene names | Hand1, Ehand, Hxt, Thing1 | |||
|
Domain Architecture |
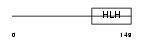
|
|||||
| Description | Heart- and neural crest derivatives-expressed protein 1 (Extraembryonic tissues, heart, autonomic nervous system and neural crest derivatives-expressed protein 1) (eHAND) (Helix-loop-helix transcription factor expressed in extraembryonic mesoderm and trophoblast) (Thing-1) (Th1). | |||||
|
AHR_HUMAN
|
||||||
| θ value | θ > 10 (rank : 81) | NC score | 0.084388 (rank : 64) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P35869, Q13728, Q13803, Q13804 | Gene names | AHR | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
AHR_MOUSE
|
||||||
| θ value | θ > 10 (rank : 82) | NC score | 0.080744 (rank : 70) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P30561, Q8QZX6, Q8R4S3, Q99P79, Q9QVY1 | Gene names | Ahr | |||
|
Domain Architecture |
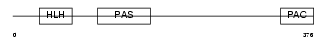
|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
EPAS1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 83) | NC score | 0.089845 (rank : 62) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q99814, Q86VA2, Q99630 | Gene names | EPAS1, HIF2A | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Member of PAS protein 2) (MOP2) (Hypoxia-inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (HLF). | |||||
|
EPAS1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 84) | NC score | 0.090280 (rank : 60) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P97481, O08787, O55046 | Gene names | Epas1, Hif2a | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Hypoxia- inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (MHLF) (HIF-related factor) (HRF). | |||||
|
HAND1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 85) | NC score | 0.055075 (rank : 100) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O96004 | Gene names | HAND1, EHAND | |||
|
Domain Architecture |
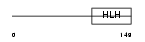
|
|||||
| Description | Heart- and neural crest derivatives-expressed protein 1 (Extraembryonic tissues, heart, autonomic nervous system and neural crest derivatives-expressed protein 1) (eHAND). | |||||
|
HEN1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 86) | NC score | 0.059760 (rank : 87) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q02575 | Gene names | NHLH1, HEN1 | |||
|
Domain Architecture |
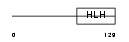
|
|||||
| Description | Helix-loop-helix protein 1 (HEN1) (Nescient helix loop helix 1) (NSCL- 1). | |||||
|
HEN1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 87) | NC score | 0.062678 (rank : 82) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q02576 | Gene names | Nhlh1, Hen1 | |||
|
Domain Architecture |
|
|||||
| Description | Helix-loop-helix protein 1 (HEN1) (Nescient helix loop helix 1) (NSCL- 1). | |||||
|
HEN2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 88) | NC score | 0.061083 (rank : 84) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q02577 | Gene names | NHLH2, HEN2 | |||
|
Domain Architecture |
|
|||||
| Description | Helix-loop-helix protein 2 (HEN2) (Nescient helix loop helix 2) (NSCL- 2). | |||||
|
HEN2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 89) | NC score | 0.061063 (rank : 85) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q64221 | Gene names | Nhlh2, Hen2 | |||
|
Domain Architecture |
|
|||||
| Description | Helix-loop-helix protein 2 (HEN2) (Nescient helix loop helix 2) (NSCL- 2). | |||||
|
HIF1A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 90) | NC score | 0.091866 (rank : 59) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q16665, Q96PT9, Q9UPB1 | Gene names | HIF1A | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein) (Member of PAS protein 1) (MOP1). | |||||
|
HIF1A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 91) | NC score | 0.090184 (rank : 61) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q61221, O08741, O08993, Q61664, Q61665, Q8C681, Q8CC19, Q8CCB6, Q8R385, Q9CYA8 | Gene names | Hif1a | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein). | |||||
|
MAD1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 92) | NC score | 0.056863 (rank : 94) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q05195 | Gene names | MXD1, MAD | |||
|
Domain Architecture |
|
|||||
| Description | MAD protein (MAX dimerizer) (MAX dimerization protein 1). | |||||
|
MAD1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 93) | NC score | 0.070155 (rank : 76) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P50538, Q60798, Q61825 | Gene names | Mxd1, Mad, Mad1 | |||
|
Domain Architecture |

|
|||||
| Description | MAD protein (MAX dimerizer) (MAX dimerization protein 1). | |||||
|
MAD3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 94) | NC score | 0.050924 (rank : 108) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9BW11, Q53HK1, Q7Z4Y0, Q8NDJ7, Q96ME3 | Gene names | MXD3, MAD3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Max-interacting transcriptional repressor MAD3 (Max-associated protein 3) (MAX dimerization protein 3) (Myx). | |||||
|
MAD4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 95) | NC score | 0.083646 (rank : 66) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q14582, Q5TZX4 | Gene names | MXD4, MAD4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-interacting transcriptional repressor MAD4 (Max-associated protein 4) (MAX dimerization protein 4). | |||||
|
MXI1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 96) | NC score | 0.056909 (rank : 93) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P50539, Q15887 | Gene names | MXI1 | |||
|
Domain Architecture |
|
|||||
| Description | MAX-interacting protein 1 (Protein MXI1). | |||||
|
MXI1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 97) | NC score | 0.056952 (rank : 92) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P50540 | Gene names | Mxi1 | |||
|
Domain Architecture |
|
|||||
| Description | MAX-interacting protein 1 (Protein MXI1). | |||||
|
MYCL1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 98) | NC score | 0.062685 (rank : 81) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P12524, Q14897, Q5QPL0, Q5QPL1, Q9NUE9 | Gene names | MYCL1, LMYC, MYCL | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
MYCL1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 99) | NC score | 0.066365 (rank : 78) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P10166, Q5FWI7 | Gene names | Mycl1, Lmyc1, Mycl | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
MYCN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.055631 (rank : 98) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P04198, Q6LDT9 | Gene names | MYCN, NMYC | |||
|
Domain Architecture |

|
|||||
| Description | N-myc proto-oncogene protein. | |||||
|
MYCN_MOUSE
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.055286 (rank : 99) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P03966, Q61978 | Gene names | Mycn, Nmyc, Nmyc1 | |||
|
Domain Architecture |

|
|||||
| Description | N-myc proto-oncogene protein. | |||||
|
MYCS_MOUSE
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.062798 (rank : 80) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9Z304 | Gene names | Mycs | |||
|
Domain Architecture |
|
|||||
| Description | Protein S-myc. | |||||
|
MYC_HUMAN
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.060798 (rank : 86) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P01106, P01107, Q14026 | Gene names | MYC | |||
|
Domain Architecture |

|
|||||
| Description | Myc proto-oncogene protein (c-Myc) (Transcription factor p64). | |||||
|
MYC_MOUSE
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.056069 (rank : 97) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P01108, P70247, Q3UM70, Q61422 | Gene names | Myc | |||
|
Domain Architecture |
|
|||||
| Description | Myc proto-oncogene protein (c-Myc) (Transcription factor p64). | |||||
|
NCOA2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.058354 (rank : 90) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q15596 | Gene names | NCOA2, TIF2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 2 (NCoA-2) (Transcriptional intermediary factor 2). | |||||
|
NCOA2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 106) | NC score | 0.058610 (rank : 89) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q61026, O09001, P97759 | Gene names | Ncoa2, Grip1, Tif2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 2 (NCoA-2) (Transcriptional intermediary factor 2) (Glucocorticoid receptor-interacting protein 1) (GRIP-1). | |||||
|
NCOA3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 107) | NC score | 0.070662 (rank : 75) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Y6Q9, Q5JYD9, Q5JYE0, Q9BR49, Q9UPC9, Q9UPG4, Q9UPG7 | Gene names | NCOA3, AIB1, RAC3, TRAM1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP-interacting protein) (pCIP). | |||||
|
NCOA3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 108) | NC score | 0.062359 (rank : 83) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O09000, Q9CRD5 | Gene names | Ncoa3, Aib1, Pcip, Rac3, Tram1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein homolog) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP- interacting protein) (p/CIP) (pCIP). | |||||
|
NPAS1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 109) | NC score | 0.081527 (rank : 69) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q99742, Q99632, Q9BY83 | Gene names | NPAS1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1) (Member of PAS protein 5) (MOP5). | |||||
|
NPAS1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 110) | NC score | 0.083912 (rank : 65) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P97459 | Gene names | Npas1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1). | |||||
|
NPAS2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 111) | NC score | 0.130818 (rank : 44) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q99743, Q99629 | Gene names | NPAS2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2) (Member of PAS protein 4) (MOP4). | |||||
|
NPAS2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 112) | NC score | 0.129758 (rank : 45) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P97460 | Gene names | Npas2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2). | |||||
|
NPAS4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 113) | NC score | 0.057197 (rank : 91) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8IUM7, Q8N8S5, Q8N9Q9 | Gene names | NPAS4, NXF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF). | |||||
|
NPAS4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 114) | NC score | 0.054624 (rank : 101) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8BGD7, Q3V3U3 | Gene names | Npas4, Nxf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF) (Limbic-enhanced PAS protein) (LE-PAS). | |||||
|
PER2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 115) | NC score | 0.050982 (rank : 107) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O15055 | Gene names | PER2, KIAA0347 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2. | |||||
|
PER3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 116) | NC score | 0.051488 (rank : 106) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O70361 | Gene names | Per3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (mPER3). | |||||
|
TCFL5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 117) | NC score | 0.068575 (rank : 77) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9UL49, O94771, Q9BYW0 | Gene names | TCFL5, CHA, E2BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor-like 5 protein (Cha transcription factor) (HPV-16 E2-binding protein 1) (E2BP-1). | |||||
|
HEY2_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 5.26011e-125 (rank : 1) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | Q9QUS4, Q3TZ99, Q8CD44 | Gene names | Hey2, Chf1, Herp, Herp1, Hesr2, Hrt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES-related repressor protein 2). | |||||
|
HEY2_HUMAN
|
||||||
| NC score | 0.996757 (rank : 2) | θ value | 4.61877e-113 (rank : 2) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9UBP5 | Gene names | HEY2, CHF1, GRL, HERP, HERP1, HRT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Cardiovascular helix-loop-helix factor 1) (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES- related repressor protein 2) (Protein gridlock homolog). | |||||
|
HEY1_HUMAN
|
||||||
| NC score | 0.961396 (rank : 3) | θ value | 1.0458e-56 (rank : 3) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9Y5J3, Q5TZS3, Q9NYP4 | Gene names | HEY1, CHF2, HESR1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1) (Cardiovascular helix-loop-helix factor 2) (HES-related repressor protein 2) (HERP2). | |||||
|
HEY1_MOUSE
|
||||||
| NC score | 0.960095 (rank : 4) | θ value | 6.56568e-51 (rank : 4) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9WV93 | Gene names | Hey1, Hesr1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1). | |||||
|
HES4_HUMAN
|
||||||
| NC score | 0.813024 (rank : 5) | θ value | 8.10077e-17 (rank : 7) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9HCC6 | Gene names | HES4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-4 (Hairy and enhancer of split 4) (bHLH factor Hes4). | |||||
|
HES1_MOUSE
|
||||||
| NC score | 0.787638 (rank : 6) | θ value | 5.60996e-18 (rank : 6) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P35428 | Gene names | Hes1, Hes-1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-1 (Hairy and enhancer of split 1). | |||||
|
HES1_HUMAN
|
||||||
| NC score | 0.787111 (rank : 7) | θ value | 5.60996e-18 (rank : 5) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q14469 | Gene names | HES1, HL, HRY | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-1 (Hairy and enhancer of split 1) (Hairy- like) (HHL) (Hairy homolog). | |||||
|
HES5_MOUSE
|
||||||
| NC score | 0.744202 (rank : 8) | θ value | 5.26297e-08 (rank : 12) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P70120, Q8BVI1 | Gene names | Hes5, Hes-5 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-5 (Hairy and enhancer of split 5). | |||||
|
HES2_MOUSE
|
||||||
| NC score | 0.724373 (rank : 9) | θ value | 5.62301e-10 (rank : 9) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O54792 | Gene names | Hes2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-2 (Hairy and enhancer of split 2). | |||||
|
HES2_HUMAN
|
||||||
| NC score | 0.720168 (rank : 10) | θ value | 4.30538e-10 (rank : 8) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9Y543, Q96EN4, Q9Y542 | Gene names | HES2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-2 (Hairy and enhancer of split 2). | |||||
|
HES6_HUMAN
|
||||||
| NC score | 0.684061 (rank : 11) | θ value | 9.59137e-10 (rank : 10) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q96HZ4, Q8N2J2, Q96T93, Q9P2S3 | Gene names | HES6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription cofactor HES-6 (Hairy and enhancer of split 6) (C- HAIRY1). | |||||
|
HES6_MOUSE
|
||||||
| NC score | 0.670130 (rank : 12) | θ value | 4.76016e-09 (rank : 11) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9JHE6, Q3U2D7 | Gene names | Hes6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription cofactor HES-6 (Hairy and enhancer of split 6). | |||||
|
HES7_HUMAN
|
||||||
| NC score | 0.585908 (rank : 13) | θ value | 1.09739e-05 (rank : 18) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9BYE0 | Gene names | HES7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor HES-7 (Hairy and enhancer of split 7) (bHLH factor Hes7). | |||||
|
HES7_MOUSE
|
||||||
| NC score | 0.584886 (rank : 14) | θ value | 4.92598e-06 (rank : 17) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8BKT2, Q99JA6 | Gene names | Hes7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor HES-7 (Hairy and enhancer of split 7) (bHLH factor Hes7). | |||||
|
HES3_MOUSE
|
||||||
| NC score | 0.568544 (rank : 15) | θ value | 0.00102713 (rank : 21) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q61657, O09083, O09150 | Gene names | Hes3, Hes-3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-3 (Hairy and enhancer of split 3). | |||||
|
BHLH3_MOUSE
|
||||||
| NC score | 0.561823 (rank : 16) | θ value | 6.87365e-08 (rank : 13) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q99PV5 | Gene names | Bhlhb3, Dec2 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (mDEC2). | |||||
|
BHLH3_HUMAN
|
||||||
| NC score | 0.558761 (rank : 17) | θ value | 3.41135e-07 (rank : 14) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9C0J9 | Gene names | BHLHB3, DEC2, SHARP1 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (hDEC2) (Enhancer-of-split and hairy-related protein 1) (SHARP-1). | |||||
|
BHLH2_HUMAN
|
||||||
| NC score | 0.535193 (rank : 18) | θ value | 5.81887e-07 (rank : 15) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O14503, Q96TD3 | Gene names | BHLHB2, DEC1, SHARP2, STRA13 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 2 (bHLHB2) (Differentially expressed in chondrocytes protein 1) (DEC1) (Enhancer-of-split and hairy-related protein 2) (SHARP-2) (Stimulated with retinoic acid 13). | |||||
|
BHLH2_MOUSE
|
||||||
| NC score | 0.525737 (rank : 19) | θ value | 2.88788e-06 (rank : 16) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O35185, P97289 | Gene names | Bhlhb2, Clast5, Stra13 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 2 (bHLHB2) (Stimulated with retinoic acid 13) (E47 interaction protein 1) (eipl). | |||||
|
MLX_MOUSE
|
||||||
| NC score | 0.320280 (rank : 20) | θ value | 0.00134147 (rank : 23) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O08609, Q9EQJ7, Q9EQJ8 | Gene names | Mlx, Tcfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
MLX_HUMAN
|
||||||
| NC score | 0.315946 (rank : 21) | θ value | 0.00134147 (rank : 22) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9UH92, Q53XM6, Q96FL2, Q9H2V0, Q9H2V1, Q9H2V2, Q9NXN3 | Gene names | MLX, TCFL4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
USF1_HUMAN
|
||||||
| NC score | 0.312070 (rank : 22) | θ value | 0.00390308 (rank : 26) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P22415 | Gene names | USF1, USF | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF1_MOUSE
|
||||||
| NC score | 0.311969 (rank : 23) | θ value | 0.00390308 (rank : 27) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q61069 | Gene names | Usf1, Usf | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF2_MOUSE
|
||||||
| NC score | 0.301537 (rank : 24) | θ value | 0.00509761 (rank : 28) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q64705, Q3UIL2, Q8VE59 | Gene names | Usf2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (Major late transcription factor 2). | |||||
|
USF2_HUMAN
|
||||||
| NC score | 0.298020 (rank : 25) | θ value | 0.00869519 (rank : 29) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q15853, O00671, O00709, Q05750, Q07952, Q15851, Q15852 | Gene names | USF2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (FOS- interacting protein) (FIP) (Major late transcription factor 2). | |||||
|
BMAL1_MOUSE
|
||||||
| NC score | 0.283651 (rank : 26) | θ value | 9.29e-05 (rank : 20) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9WTL8, O88295, Q921S4, Q9R0U2, Q9WTL9 | Gene names | Arntl, Bmal1 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Arnt3). | |||||
|
TFE3_HUMAN
|
||||||
| NC score | 0.281243 (rank : 27) | θ value | 0.0193708 (rank : 33) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P19532, Q92757, Q92758, Q99964 | Gene names | TFE3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E3. | |||||
|
BMAL1_HUMAN
|
||||||
| NC score | 0.279833 (rank : 28) | θ value | 9.29e-05 (rank : 19) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O00327, O00313, O00314, O00315, O00316, O00317, Q99631, Q99649 | Gene names | ARNTL, BMAL1, MOP3 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Member of PAS protein 3) (Basic-helix-loop- helix-PAS orphan MOP3) (bHLH-PAS protein JAP3). | |||||
|
MITF_HUMAN
|
||||||
| NC score | 0.269763 (rank : 29) | θ value | 0.0563607 (rank : 37) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O75030, Q14841, Q9P2V0, Q9P2V1, Q9P2V2, Q9P2Y8 | Gene names | MITF | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
MITF_MOUSE
|
||||||
| NC score | 0.267196 (rank : 30) | θ value | 0.0563607 (rank : 38) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q08874, O08885, O88203, Q08843, Q60781, Q60782, Q9JIJ0, Q9JIJ1, Q9JIJ2, Q9JIJ3, Q9JIJ4, Q9JIJ5, Q9JIJ6, Q9JKX9 | Gene names | Mitf, Bw, Mi, Vit | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
TFEB_MOUSE
|
||||||
| NC score | 0.260350 (rank : 31) | θ value | 0.0431538 (rank : 36) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9R210 | Gene names | Tfeb, Tcfeb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
TFEB_HUMAN
|
||||||
| NC score | 0.256768 (rank : 32) | θ value | 0.0431538 (rank : 35) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P19484, Q709B3, Q7Z6P9, Q9BRJ5, Q9UJD8 | Gene names | TFEB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
SRBP2_HUMAN
|
||||||
| NC score | 0.242855 (rank : 33) | θ value | 0.0193708 (rank : 32) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q12772, Q9UH04 | Gene names | SREBF2, SREBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 2 (SREBP-2) (Sterol regulatory element-binding transcription factor 2). | |||||
|
SRBP1_HUMAN
|
||||||
| NC score | 0.241524 (rank : 34) | θ value | 0.0252991 (rank : 34) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P36956, Q16062 | Gene names | SREBF1, SREBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
ARNT2_MOUSE
|
||||||
| NC score | 0.229817 (rank : 35) | θ value | 0.00298849 (rank : 25) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q61324 | Gene names | Arnt2 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
ARNT2_HUMAN
|
||||||
| NC score | 0.228539 (rank : 36) | θ value | 0.00298849 (rank : 24) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9HBZ2, O15024 | Gene names | ARNT2, KIAA0307 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
ARNT_HUMAN
|
||||||
| NC score | 0.225281 (rank : 37) | θ value | 0.0113563 (rank : 30) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P27540 | Gene names | ARNT | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
ARNT_MOUSE
|
||||||
| NC score | 0.221920 (rank : 38) | θ value | 0.0113563 (rank : 31) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P53762, Q60661 | Gene names | Arnt | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
SRBP1_MOUSE
|
||||||
| NC score | 0.219028 (rank : 39) | θ value | 0.125558 (rank : 39) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9WTN3 | Gene names | Srebf1, Srebp1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
CLOCK_MOUSE
|
||||||
| NC score | 0.165378 (rank : 40) | θ value | 0.365318 (rank : 41) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O08785 | Gene names | Clock | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (mCLOCK). | |||||
|
CLOCK_HUMAN
|
||||||
| NC score | 0.164215 (rank : 41) | θ value | 0.62314 (rank : 43) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O15516, O14516, Q9UIT8 | Gene names | CLOCK, KIAA0334 | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (hCLOCK). | |||||
|
MAX_MOUSE
|
||||||
| NC score | 0.153696 (rank : 42) | θ value | 6.88961 (rank : 74) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P28574 | Gene names | Max, Myn | |||
|
Domain Architecture |

|
|||||
| Description | Protein max (Myc-associated factor X) (Protein myn) (Myc-binding novel HLH/LZ protein). | |||||
|
MAX_HUMAN
|
||||||
| NC score | 0.149410 (rank : 43) | θ value | 6.88961 (rank : 73) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P61244, P25912, P52163 | Gene names | MAX | |||
|
Domain Architecture |

|
|||||
| Description | Protein max (Myc-associated factor X). | |||||
|
NPAS2_HUMAN
|
||||||
| NC score | 0.130818 (rank : 44) | θ value | θ > 10 (rank : 111) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q99743, Q99629 | Gene names | NPAS2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2) (Member of PAS protein 4) (MOP4). | |||||
|
NPAS2_MOUSE
|
||||||
| NC score | 0.129758 (rank : 45) | θ value | θ > 10 (rank : 112) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P97460 | Gene names | Npas2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2). | |||||
|
TAL1_HUMAN
|
||||||
| NC score | 0.115563 (rank : 46) | θ value | 0.125558 (rank : 40) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P17542 | Gene names | TAL1, SCL, TCL5 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell acute lymphocytic leukemia-1 protein (TAL-1 protein) (Stem cell protein) (T-cell leukemia/lymphoma-5 protein). | |||||
|
SIM2_HUMAN
|
||||||
| NC score | 0.113097 (rank : 47) | θ value | 1.81305 (rank : 53) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q14190, O60766, Q15470, Q15471, Q15472, Q15473, Q16532 | Gene names | SIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2. | |||||
|
NCOA1_MOUSE
|
||||||
| NC score | 0.113081 (rank : 48) | θ value | 2.36792 (rank : 59) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P70365, P70366, Q61202, Q8CBI9 | Gene names | Ncoa1, Src1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (Nuclear receptor coactivator protein 1) (mNRC-1). | |||||
|
NCOA1_HUMAN
|
||||||
| NC score | 0.112560 (rank : 49) | θ value | 2.36792 (rank : 58) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q15788, O00150, O43792, O43793, Q13071, Q13420, Q6GVI5, Q7KYV3 | Gene names | NCOA1, SRC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (RIP160) (Protein Hin-2) (NY-REN-52 antigen). | |||||
|
SIM2_MOUSE
|
||||||
| NC score | 0.111543 (rank : 50) | θ value | 1.81305 (rank : 54) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q61079, O35391, Q61046, Q61904 | Gene names | Sim2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2 (SIM transcription factor) (mSIM). | |||||
|
TAL1_MOUSE
|
||||||
| NC score | 0.108774 (rank : 51) | θ value | 1.06291 (rank : 46) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P22091 | Gene names | Tal1, Scl, Tal-1 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell acute lymphocytic leukemia-1 protein homolog (TAL-1 protein) (Stem cell protein). | |||||
|
SIM1_MOUSE
|
||||||
| NC score | 0.107336 (rank : 52) | θ value | 4.03905 (rank : 67) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q61045, O70284, P70183 | Gene names | Sim1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1 (mSIM1). | |||||
|
SIM1_HUMAN
|
||||||
| NC score | 0.107287 (rank : 53) | θ value | 4.03905 (rank : 66) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P81133, Q5TDP7 | Gene names | SIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1. | |||||
|
TFAP4_HUMAN
|
||||||
| NC score | 0.106661 (rank : 54) | θ value | 6.88961 (rank : 76) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q01664, O60409 | Gene names | TFAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor AP-4 (Activating enhancer-binding protein 4). | |||||
|
NPAS3_MOUSE
|
||||||
| NC score | 0.101216 (rank : 55) | θ value | 4.03905 (rank : 65) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9QZQ0, Q9EQP4 | Gene names | Npas3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
NPAS3_HUMAN
|
||||||
| NC score | 0.100058 (rank : 56) | θ value | 4.03905 (rank : 64) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8IXF0, Q86US6, Q86US7, Q8IXF2, Q9BY81, Q9H323, Q9Y4L8 | Gene names | NPAS3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
LYL1_HUMAN
|
||||||
| NC score | 0.097462 (rank : 57) | θ value | 0.47712 (rank : 42) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P12980, O76102 | Gene names | LYL1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein lyl-1 (Lymphoblastic leukemia-derived sequence 1). | |||||
|
LYL1_MOUSE
|
||||||
| NC score | 0.095328 (rank : 58) | θ value | 0.62314 (rank : 44) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P27792 | Gene names | Lyl1, Lyl-1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein lyl-1 (Lymphoblastic leukemia-derived sequence 1). | |||||
|
HIF1A_HUMAN
|
||||||
| NC score | 0.091866 (rank : 59) | θ value | θ > 10 (rank : 90) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q16665, Q96PT9, Q9UPB1 | Gene names | HIF1A | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein) (Member of PAS protein 1) (MOP1). | |||||
|
EPAS1_MOUSE
|
||||||
| NC score | 0.090280 (rank : 60) | θ value | θ > 10 (rank : 84) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P97481, O08787, O55046 | Gene names | Epas1, Hif2a | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Hypoxia- inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (MHLF) (HIF-related factor) (HRF). | |||||
|
HIF1A_MOUSE
|
||||||
| NC score | 0.090184 (rank : 61) | θ value | θ > 10 (rank : 91) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q61221, O08741, O08993, Q61664, Q61665, Q8C681, Q8CC19, Q8CCB6, Q8R385, Q9CYA8 | Gene names | Hif1a | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein). | |||||
|
EPAS1_HUMAN
|
||||||
| NC score | 0.089845 (rank : 62) | θ value | θ > 10 (rank : 83) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q99814, Q86VA2, Q99630 | Gene names | EPAS1, HIF2A | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Member of PAS protein 2) (MOP2) (Hypoxia-inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (HLF). | |||||
|
TAL2_MOUSE
|
||||||
| NC score | 0.089189 (rank : 63) | θ value | 1.38821 (rank : 50) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q62282 | Gene names | Tal2 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell acute lymphocytic leukemia-2 protein homolog (TAL-2 protein). | |||||
|
AHR_HUMAN
|
||||||
| NC score | 0.084388 (rank : 64) | θ value | θ > 10 (rank : 81) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P35869, Q13728, Q13803, Q13804 | Gene names | AHR | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
NPAS1_MOUSE
|
||||||
| NC score | 0.083912 (rank : 65) | θ value | θ > 10 (rank : 110) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P97459 | Gene names | Npas1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1). | |||||
|
MAD4_HUMAN
|
||||||
| NC score | 0.083646 (rank : 66) | θ value | θ > 10 (rank : 95) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q14582, Q5TZX4 | Gene names | MXD4, MAD4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-interacting transcriptional repressor MAD4 (Max-associated protein 4) (MAX dimerization protein 4). | |||||
|
BHLH8_HUMAN
|
||||||
| NC score | 0.083304 (rank : 67) | θ value | 1.38821 (rank : 48) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q7RTS1 | Gene names | BHLHB8, MIST1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Class B basic helix-loop-helix protein 8 (bHLHB8) (Muscle, intestine and stomach expression 1) (MIST-1). | |||||
|
BHLH8_MOUSE
|
||||||
| NC score | 0.082197 (rank : 68) | θ value | 1.38821 (rank : 49) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9QYC3, Q9QYE4 | Gene names | Bhlhb8, Mist1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Class B basic helix-loop-helix protein 8 (bHLHB8) (Muscle, intestine and stomach expression 1) (MIST-1). | |||||
|
NPAS1_HUMAN
|
||||||
| NC score | 0.081527 (rank : 69) | θ value | θ > 10 (rank : 109) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q99742, Q99632, Q9BY83 | Gene names | NPAS1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1) (Member of PAS protein 5) (MOP5). | |||||
|
AHR_MOUSE
|
||||||
| NC score | 0.080744 (rank : 70) | θ value | θ > 10 (rank : 82) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P30561, Q8QZX6, Q8R4S3, Q99P79, Q9QVY1 | Gene names | Ahr | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
ID1_MOUSE
|
||||||
| NC score | 0.075512 (rank : 71) | θ value | 1.81305 (rank : 52) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P20067, Q61101, Q9D897 | Gene names | Id1, Id, Id-1, Idb1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-binding protein inhibitor ID-1 (Inhibitor of DNA binding 1). | |||||
|
TAL2_HUMAN
|
||||||
| NC score | 0.074721 (rank : 72) | θ value | 4.03905 (rank : 68) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q16559 | Gene names | TAL2 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell acute lymphocytic leukemia-2 protein (TAL-2 protein). | |||||
|
ID4_HUMAN
|
||||||
| NC score | 0.071522 (rank : 73) | θ value | 2.36792 (rank : 56) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P47928, Q13005 | Gene names | ID4 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-binding protein inhibitor ID-4 (Inhibitor of DNA binding 4). | |||||
|
ID4_MOUSE
|
||||||
| NC score | 0.071177 (rank : 74) | θ value | 2.36792 (rank : 57) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P41139 | Gene names | Id4, Id-4, Idb4 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-binding protein inhibitor ID-4 (Inhibitor of DNA binding 4). | |||||
|
NCOA3_HUMAN
|
||||||
| NC score | 0.070662 (rank : 75) | θ value | θ > 10 (rank : 107) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Y6Q9, Q5JYD9, Q5JYE0, Q9BR49, Q9UPC9, Q9UPG4, Q9UPG7 | Gene names | NCOA3, AIB1, RAC3, TRAM1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP-interacting protein) (pCIP). | |||||
|
MAD1_MOUSE
|
||||||
| NC score | 0.070155 (rank : 76) | θ value | θ > 10 (rank : 93) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P50538, Q60798, Q61825 | Gene names | Mxd1, Mad, Mad1 | |||
|
Domain Architecture |

|
|||||
| Description | MAD protein (MAX dimerizer) (MAX dimerization protein 1). | |||||
|
TCFL5_HUMAN
|
||||||
| NC score | 0.068575 (rank : 77) | θ value | θ > 10 (rank : 117) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9UL49, O94771, Q9BYW0 | Gene names | TCFL5, CHA, E2BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor-like 5 protein (Cha transcription factor) (HPV-16 E2-binding protein 1) (E2BP-1). | |||||
|
MYCL1_MOUSE
|
||||||
| NC score | 0.066365 (rank : 78) | θ value | θ > 10 (rank : 99) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P10166, Q5FWI7 | Gene names | Mycl1, Lmyc1, Mycl | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
ID1_HUMAN
|
||||||
| NC score | 0.064752 (rank : 79) | θ value | 5.27518 (rank : 70) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P41134, O00651, O00652, Q16371, Q16377, Q5TE66, Q5TE67, Q969Z7, Q9H0Z5, Q9H109 | Gene names | ID1, ID | |||
|
Domain Architecture |

|
|||||
| Description | DNA-binding protein inhibitor ID-1 (Inhibitor of DNA binding 1). | |||||
|
MYCS_MOUSE
|
||||||
| NC score | 0.062798 (rank : 80) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9Z304 | Gene names | Mycs | |||
|
Domain Architecture |

|
|||||
| Description | Protein S-myc. | |||||
|
MYCL1_HUMAN
|
||||||
| NC score | 0.062685 (rank : 81) | θ value | θ > 10 (rank : 98) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P12524, Q14897, Q5QPL0, Q5QPL1, Q9NUE9 | Gene names | MYCL1, LMYC, MYCL | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
HEN1_MOUSE
|
||||||
| NC score | 0.062678 (rank : 82) | θ value | θ > 10 (rank : 87) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q02576 | Gene names | Nhlh1, Hen1 | |||
|
Domain Architecture |

|
|||||
| Description | Helix-loop-helix protein 1 (HEN1) (Nescient helix loop helix 1) (NSCL- 1). | |||||
|
NCOA3_MOUSE
|
||||||
| NC score | 0.062359 (rank : 83) | θ value | θ > 10 (rank : 108) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O09000, Q9CRD5 | Gene names | Ncoa3, Aib1, Pcip, Rac3, Tram1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein homolog) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP- interacting protein) (p/CIP) (pCIP). | |||||
|
HEN2_HUMAN
|
||||||
| NC score | 0.061083 (rank : 84) | θ value | θ > 10 (rank : 88) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q02577 | Gene names | NHLH2, HEN2 | |||
|
Domain Architecture |

|
|||||
| Description | Helix-loop-helix protein 2 (HEN2) (Nescient helix loop helix 2) (NSCL- 2). | |||||
|
HEN2_MOUSE
|
||||||
| NC score | 0.061063 (rank : 85) | θ value | θ > 10 (rank : 89) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q64221 | Gene names | Nhlh2, Hen2 | |||
|
Domain Architecture |

|
|||||
| Description | Helix-loop-helix protein 2 (HEN2) (Nescient helix loop helix 2) (NSCL- 2). | |||||
|
MYC_HUMAN
|
||||||
| NC score | 0.060798 (rank : 86) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P01106, P01107, Q14026 | Gene names | MYC | |||
|
Domain Architecture |

|
|||||
| Description | Myc proto-oncogene protein (c-Myc) (Transcription factor p64). | |||||
|
HEN1_HUMAN
|
||||||
| NC score | 0.059760 (rank : 87) | θ value | θ > 10 (rank : 86) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q02575 | Gene names | NHLH1, HEN1 | |||
|
Domain Architecture |

|
|||||
| Description | Helix-loop-helix protein 1 (HEN1) (Nescient helix loop helix 1) (NSCL- 1). | |||||
|
HAND1_MOUSE
|
||||||
| NC score | 0.059244 (rank : 88) | θ value | 8.99809 (rank : 80) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q64279, Q61099 | Gene names | Hand1, Ehand, Hxt, Thing1 | |||
|
Domain Architecture |

|
|||||
| Description | Heart- and neural crest derivatives-expressed protein 1 (Extraembryonic tissues, heart, autonomic nervous system and neural crest derivatives-expressed protein 1) (eHAND) (Helix-loop-helix transcription factor expressed in extraembryonic mesoderm and trophoblast) (Thing-1) (Th1). | |||||
|
NCOA2_MOUSE
|
||||||
| NC score | 0.058610 (rank : 89) | θ value | θ > 10 (rank : 106) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q61026, O09001, P97759 | Gene names | Ncoa2, Grip1, Tif2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 2 (NCoA-2) (Transcriptional intermediary factor 2) (Glucocorticoid receptor-interacting protein 1) (GRIP-1). | |||||
|
NCOA2_HUMAN
|
||||||
| NC score | 0.058354 (rank : 90) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q15596 | Gene names | NCOA2, TIF2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 2 (NCoA-2) (Transcriptional intermediary factor 2). | |||||
|
NPAS4_HUMAN
|
||||||
| NC score | 0.057197 (rank : 91) | θ value | θ > 10 (rank : 113) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8IUM7, Q8N8S5, Q8N9Q9 | Gene names | NPAS4, NXF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF). | |||||
|
MXI1_MOUSE
|
||||||
| NC score | 0.056952 (rank : 92) | θ value | θ > 10 (rank : 97) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P50540 | Gene names | Mxi1 | |||
|
Domain Architecture |

|
|||||
| Description | MAX-interacting protein 1 (Protein MXI1). | |||||
|
MXI1_HUMAN
|
||||||
| NC score | 0.056909 (rank : 93) | θ value | θ > 10 (rank : 96) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P50539, Q15887 | Gene names | MXI1 | |||
|
Domain Architecture |

|
|||||
| Description | MAX-interacting protein 1 (Protein MXI1). | |||||
|
MAD1_HUMAN
|
||||||
| NC score | 0.056863 (rank : 94) | θ value | θ > 10 (rank : 92) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q05195 | Gene names | MXD1, MAD | |||
|
Domain Architecture |

|
|||||
| Description | MAD protein (MAX dimerizer) (MAX dimerization protein 1). | |||||
|
TWST1_HUMAN
|
||||||
| NC score | 0.056372 (rank : 95) | θ value | 6.88961 (rank : 77) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q15672, Q92487, Q99804 | Gene names | TWIST1, TWIST | |||
|
Domain Architecture |

|
|||||
| Description | Twist-related protein 1 (H-twist). | |||||
|
TWST1_MOUSE
|
||||||
| NC score | 0.056168 (rank : 96) | θ value | 6.88961 (rank : 78) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P26687 | Gene names | Twist1, Twist | |||
|
Domain Architecture |

|
|||||
| Description | Twist-related protein 1 (M-twist). | |||||
|
MYC_MOUSE
|
||||||
| NC score | 0.056069 (rank : 97) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P01108, P70247, Q3UM70, Q61422 | Gene names | Myc | |||
|
Domain Architecture |

|
|||||
| Description | Myc proto-oncogene protein (c-Myc) (Transcription factor p64). | |||||
|
MYCN_HUMAN
|
||||||
| NC score | 0.055631 (rank : 98) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P04198, Q6LDT9 | Gene names | MYCN, NMYC | |||
|
Domain Architecture |

|
|||||
| Description | N-myc proto-oncogene protein. | |||||
|
MYCN_MOUSE
|
||||||
| NC score | 0.055286 (rank : 99) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P03966, Q61978 | Gene names | Mycn, Nmyc, Nmyc1 | |||
|
Domain Architecture |

|
|||||
| Description | N-myc proto-oncogene protein. | |||||
|
HAND1_HUMAN
|
||||||
| NC score | 0.055075 (rank : 100) | θ value | θ > 10 (rank : 85) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O96004 | Gene names | HAND1, EHAND | |||
|
Domain Architecture |

|
|||||
| Description | Heart- and neural crest derivatives-expressed protein 1 (Extraembryonic tissues, heart, autonomic nervous system and neural crest derivatives-expressed protein 1) (eHAND). | |||||
|
NPAS4_MOUSE
|
||||||
| NC score | 0.054624 (rank : 101) | θ value | θ > 10 (rank : 114) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8BGD7, Q3V3U3 | Gene names | Npas4, Nxf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF) (Limbic-enhanced PAS protein) (LE-PAS). | |||||
|
NDF4_HUMAN
|
||||||
| NC score | 0.054198 (rank : 102) | θ value | 3.0926 (rank : 60) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9HD90 | Gene names | NEUROD4 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic differentiation factor 4 (NeuroD4). | |||||
|
TWST2_MOUSE
|
||||||
| NC score | 0.053462 (rank : 103) | θ value | 5.27518 (rank : 72) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9D030 | Gene names | Twist2, Dermo1 | |||
|
Domain Architecture |

|
|||||
| Description | Twist-related protein 2 (Dermis expressed protein 1) (Dermo-1). | |||||
|
TWST2_HUMAN
|
||||||
| NC score | 0.053393 (rank : 104) | θ value | 5.27518 (rank : 71) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8WVJ9 | Gene names | TWIST2, DERMO1 | |||
|
Domain Architecture |

|
|||||
| Description | Twist-related protein 2 (Dermis expressed protein 1) (Dermo-1). | |||||
|
NDF4_MOUSE
|
||||||
| NC score | 0.052024 (rank : 105) | θ value | 3.0926 (rank : 61) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O09105 | Gene names | Neurod4, Ath3, Atoh3 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic differentiation factor 4 (NeuroD4) (Atonal protein homolog 3) (Helix-loop-helix protein mATH-3) (MATH3). | |||||
|
PER3_MOUSE
|
||||||
| NC score | 0.051488 (rank : 106) | θ value | θ > 10 (rank : 116) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O70361 | Gene names | Per3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (mPER3). | |||||
|
PER2_HUMAN
|
||||||
| NC score | 0.050982 (rank : 107) | θ value | θ > 10 (rank : 115) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O15055 | Gene names | PER2, KIAA0347 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2. | |||||
|
MAD3_HUMAN
|
||||||
| NC score | 0.050924 (rank : 108) | θ value | θ > 10 (rank : 94) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9BW11, Q53HK1, Q7Z4Y0, Q8NDJ7, Q96ME3 | Gene names | MXD3, MAD3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Max-interacting transcriptional repressor MAD3 (Max-associated protein 3) (MAX dimerization protein 3) (Myx). | |||||
|
NGN3_MOUSE
|
||||||
| NC score | 0.048495 (rank : 109) | θ value | 6.88961 (rank : 75) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P70661 | Gene names | Neurog3, Ath4b, Atoh5, Ngn3 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenin 3 (Atonal protein homolog 5) (Helix-loop-helix protein mATH-4B) (MATH4B). | |||||
|
HNF1A_MOUSE
|
||||||
| NC score | 0.024203 (rank : 110) | θ value | 0.813845 (rank : 45) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P22361 | Gene names | Tcf1, Hnf-1, Hnf-1a, Hnf1a | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 1-alpha (HNF-1A) (Liver-specific transcription factor LF-B1) (LFB1). | |||||
|
APLP1_MOUSE
|
||||||
| NC score | 0.023408 (rank : 111) | θ value | 1.81305 (rank : 51) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q03157, Q8VC38 | Gene names | Aplp1 | |||
|
Domain Architecture |

|
|||||
| Description | Amyloid-like protein 1 precursor (APLP) (APLP-1) [Contains: C30]. | |||||
|
ALAT1_HUMAN
|
||||||
| NC score | 0.021485 (rank : 112) | θ value | 1.38821 (rank : 47) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P24298, P78398, Q93076 | Gene names | GPT, AAT1, GPT1 | |||
|
Domain Architecture |

|
|||||
| Description | Alanine aminotransferase 1 (EC 2.6.1.2) (ALT1) (Glutamic--pyruvic transaminase 1) (GPT 1) (Glutamic--alanine transaminase 1). | |||||
|
ELK3_MOUSE
|
||||||
| NC score | 0.015843 (rank : 113) | θ value | 2.36792 (rank : 55) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P41971, P97747, Q62346 | Gene names | Elk3, Erp, Net | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-3 (ETS-related protein NET) (ETS- related protein ERP). | |||||
|
ELK3_HUMAN
|
||||||
| NC score | 0.014468 (rank : 114) | θ value | 4.03905 (rank : 63) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P41970, Q6FG57, Q6GU29, Q9UD17 | Gene names | ELK3, NET, SAP2 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-3 (ETS-related protein NET) (ETS- related protein ERP) (SRF accessory protein 2) (SAP-2). | |||||
|
WDR22_MOUSE
|
||||||
| NC score | 0.012542 (rank : 115) | θ value | 4.03905 (rank : 69) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80T85, Q80VT3, Q80ZW6, Q8BIP8, Q8BVK5, Q8K0Q6 | Gene names | Wdr22, Kiaa1824 | |||
|
Domain Architecture |

|
|||||
| Description | WD repeat protein 22. | |||||
|
ALS2_MOUSE
|
||||||
| NC score | 0.011995 (rank : 116) | θ value | 4.03905 (rank : 62) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q920R0, Q8JZR1, Q9CXJ3 | Gene names | Als2 | |||
|
Domain Architecture |

|
|||||
| Description | Alsin (Amyotrophic lateral sclerosis protein 2 homolog). | |||||
|
GPTC2_MOUSE
|
||||||
| NC score | 0.009024 (rank : 117) | θ value | 8.99809 (rank : 79) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7TQC7, Q8BNJ9, Q8BPM1, Q8CDH9 | Gene names | Gpatc2 | |||
|
Domain Architecture |

|
|||||
| Description | G patch domain-containing protein 2. | |||||