Please be patient as the page loads
|
SIM2_HUMAN
|
||||||
| SwissProt Accessions | Q14190, O60766, Q15470, Q15471, Q15472, Q15473, Q16532 | Gene names | SIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
SIM1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.988909 (rank : 3) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P81133, Q5TDP7 | Gene names | SIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1. | |||||
|
SIM1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.988506 (rank : 4) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q61045, O70284, P70183 | Gene names | Sim1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1 (mSIM1). | |||||
|
SIM2_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | Q14190, O60766, Q15470, Q15471, Q15472, Q15473, Q16532 | Gene names | SIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2. | |||||
|
SIM2_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.995700 (rank : 2) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q61079, O35391, Q61046, Q61904 | Gene names | Sim2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2 (SIM transcription factor) (mSIM). | |||||
|
NPAS3_HUMAN
|
||||||
| θ value | 2.0926e-81 (rank : 5) | NC score | 0.946602 (rank : 7) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8IXF0, Q86US6, Q86US7, Q8IXF2, Q9BY81, Q9H323, Q9Y4L8 | Gene names | NPAS3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
NPAS3_MOUSE
|
||||||
| θ value | 2.0926e-81 (rank : 6) | NC score | 0.945507 (rank : 8) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9QZQ0, Q9EQP4 | Gene names | Npas3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
NPAS1_MOUSE
|
||||||
| θ value | 5.8972e-76 (rank : 7) | NC score | 0.954256 (rank : 5) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P97459 | Gene names | Npas1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1). | |||||
|
HIF1A_MOUSE
|
||||||
| θ value | 1.89707e-74 (rank : 8) | NC score | 0.936152 (rank : 9) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q61221, O08741, O08993, Q61664, Q61665, Q8C681, Q8CC19, Q8CCB6, Q8R385, Q9CYA8 | Gene names | Hif1a | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein). | |||||
|
HIF1A_HUMAN
|
||||||
| θ value | 9.41505e-74 (rank : 9) | NC score | 0.933296 (rank : 10) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q16665, Q96PT9, Q9UPB1 | Gene names | HIF1A | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein) (Member of PAS protein 1) (MOP1). | |||||
|
NPAS1_HUMAN
|
||||||
| θ value | 2.09745e-73 (rank : 10) | NC score | 0.953012 (rank : 6) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q99742, Q99632, Q9BY83 | Gene names | NPAS1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1) (Member of PAS protein 5) (MOP5). | |||||
|
EPAS1_HUMAN
|
||||||
| θ value | 9.76566e-63 (rank : 11) | NC score | 0.924438 (rank : 11) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q99814, Q86VA2, Q99630 | Gene names | EPAS1, HIF2A | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Member of PAS protein 2) (MOP2) (Hypoxia-inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (HLF). | |||||
|
EPAS1_MOUSE
|
||||||
| θ value | 4.10295e-61 (rank : 12) | NC score | 0.923527 (rank : 12) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P97481, O08787, O55046 | Gene names | Epas1, Hif2a | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Hypoxia- inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (MHLF) (HIF-related factor) (HRF). | |||||
|
AHR_MOUSE
|
||||||
| θ value | 1.28732e-30 (rank : 13) | NC score | 0.841801 (rank : 14) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P30561, Q8QZX6, Q8R4S3, Q99P79, Q9QVY1 | Gene names | Ahr | |||
|
Domain Architecture |
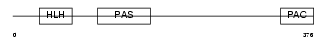
|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
AHR_HUMAN
|
||||||
| θ value | 3.17079e-29 (rank : 14) | NC score | 0.842738 (rank : 13) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P35869, Q13728, Q13803, Q13804 | Gene names | AHR | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
BMAL1_MOUSE
|
||||||
| θ value | 4.14116e-29 (rank : 15) | NC score | 0.791811 (rank : 16) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9WTL8, O88295, Q921S4, Q9R0U2, Q9WTL9 | Gene names | Arntl, Bmal1 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Arnt3). | |||||
|
BMAL1_HUMAN
|
||||||
| θ value | 3.50572e-28 (rank : 16) | NC score | 0.793198 (rank : 15) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O00327, O00313, O00314, O00315, O00316, O00317, Q99631, Q99649 | Gene names | ARNTL, BMAL1, MOP3 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Member of PAS protein 3) (Basic-helix-loop- helix-PAS orphan MOP3) (bHLH-PAS protein JAP3). | |||||
|
ARNT2_HUMAN
|
||||||
| θ value | 2.77775e-25 (rank : 17) | NC score | 0.748487 (rank : 19) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9HBZ2, O15024 | Gene names | ARNT2, KIAA0307 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
ARNT2_MOUSE
|
||||||
| θ value | 2.77775e-25 (rank : 18) | NC score | 0.747171 (rank : 20) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q61324 | Gene names | Arnt2 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
CLOCK_HUMAN
|
||||||
| θ value | 1.6852e-22 (rank : 19) | NC score | 0.760858 (rank : 17) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O15516, O14516, Q9UIT8 | Gene names | CLOCK, KIAA0334 | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (hCLOCK). | |||||
|
CLOCK_MOUSE
|
||||||
| θ value | 2.20094e-22 (rank : 20) | NC score | 0.757698 (rank : 18) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O08785 | Gene names | Clock | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (mCLOCK). | |||||
|
NPAS4_HUMAN
|
||||||
| θ value | 2.20094e-22 (rank : 21) | NC score | 0.704444 (rank : 25) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8IUM7, Q8N8S5, Q8N9Q9 | Gene names | NPAS4, NXF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF). | |||||
|
ARNT_HUMAN
|
||||||
| θ value | 3.17815e-21 (rank : 22) | NC score | 0.736844 (rank : 21) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P27540 | Gene names | ARNT | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
ARNT_MOUSE
|
||||||
| θ value | 3.17815e-21 (rank : 23) | NC score | 0.724561 (rank : 24) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P53762, Q60661 | Gene names | Arnt | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
NPAS4_MOUSE
|
||||||
| θ value | 1.2077e-20 (rank : 24) | NC score | 0.688795 (rank : 26) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8BGD7, Q3V3U3 | Gene names | Npas4, Nxf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF) (Limbic-enhanced PAS protein) (LE-PAS). | |||||
|
NPAS2_MOUSE
|
||||||
| θ value | 2.27762e-19 (rank : 25) | NC score | 0.735514 (rank : 22) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P97460 | Gene names | Npas2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2). | |||||
|
NPAS2_HUMAN
|
||||||
| θ value | 5.60996e-18 (rank : 26) | NC score | 0.727511 (rank : 23) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q99743, Q99629 | Gene names | NPAS2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2) (Member of PAS protein 4) (MOP4). | |||||
|
NCOA1_MOUSE
|
||||||
| θ value | 5.25075e-16 (rank : 27) | NC score | 0.633834 (rank : 27) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P70365, P70366, Q61202, Q8CBI9 | Gene names | Ncoa1, Src1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (Nuclear receptor coactivator protein 1) (mNRC-1). | |||||
|
NCOA1_HUMAN
|
||||||
| θ value | 6.85773e-16 (rank : 28) | NC score | 0.630315 (rank : 28) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q15788, O00150, O43792, O43793, Q13071, Q13420, Q6GVI5, Q7KYV3 | Gene names | NCOA1, SRC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (RIP160) (Protein Hin-2) (NY-REN-52 antigen). | |||||
|
NCOA2_HUMAN
|
||||||
| θ value | 2.28291e-11 (rank : 29) | NC score | 0.556651 (rank : 29) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q15596 | Gene names | NCOA2, TIF2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 2 (NCoA-2) (Transcriptional intermediary factor 2). | |||||
|
NCOA2_MOUSE
|
||||||
| θ value | 2.52405e-10 (rank : 30) | NC score | 0.554927 (rank : 30) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q61026, O09001, P97759 | Gene names | Ncoa2, Grip1, Tif2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 2 (NCoA-2) (Transcriptional intermediary factor 2) (Glucocorticoid receptor-interacting protein 1) (GRIP-1). | |||||
|
NCOA3_HUMAN
|
||||||
| θ value | 1.38499e-08 (rank : 31) | NC score | 0.553234 (rank : 31) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9Y6Q9, Q5JYD9, Q5JYE0, Q9BR49, Q9UPC9, Q9UPG4, Q9UPG7 | Gene names | NCOA3, AIB1, RAC3, TRAM1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP-interacting protein) (pCIP). | |||||
|
NCOA3_MOUSE
|
||||||
| θ value | 1.53129e-07 (rank : 32) | NC score | 0.517247 (rank : 32) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O09000, Q9CRD5 | Gene names | Ncoa3, Aib1, Pcip, Rac3, Tram1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein homolog) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP- interacting protein) (p/CIP) (pCIP). | |||||
|
BSN_HUMAN
|
||||||
| θ value | 0.365318 (rank : 33) | NC score | 0.031121 (rank : 58) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
ZN645_HUMAN
|
||||||
| θ value | 0.62314 (rank : 34) | NC score | 0.030760 (rank : 59) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 265 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8N7E2, Q6DJY9 | Gene names | ZNF645 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 645. | |||||
|
PER2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 35) | NC score | 0.216490 (rank : 34) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O15055 | Gene names | PER2, KIAA0347 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2. | |||||
|
PER2_MOUSE
|
||||||
| θ value | 1.06291 (rank : 36) | NC score | 0.196936 (rank : 38) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O54943, O54954 | Gene names | Per2 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2 (mPER2). | |||||
|
RTN4_HUMAN
|
||||||
| θ value | 1.06291 (rank : 37) | NC score | 0.013308 (rank : 65) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NQC3, O94962, Q9BXG5, Q9H212, Q9H3I3, Q9UQ42, Q9Y293, Q9Y2Y7, Q9Y5U6 | Gene names | RTN4, KIAA0886, NOGO | |||
|
Domain Architecture |

|
|||||
| Description | Reticulon-4 (Neurite outgrowth inhibitor) (Nogo protein) (Foocen) (Neuroendocrine-specific protein) (NSP) (Neuroendocrine-specific protein C homolog) (RTN-x) (Reticulon-5). | |||||
|
EXOC3_MOUSE
|
||||||
| θ value | 1.38821 (rank : 38) | NC score | 0.024503 (rank : 61) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6KAR6 | Gene names | Exoc3, Sec6l1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Exocyst complex component 3 (Exocyst complex component Sec6). | |||||
|
HXB3_HUMAN
|
||||||
| θ value | 1.38821 (rank : 39) | NC score | 0.005041 (rank : 82) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 559 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P14651, O95615, P17484 | Gene names | HOXB3, HOX2G | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-B3 (Hox-2G) (Hox-2.7). | |||||
|
US6NL_HUMAN
|
||||||
| θ value | 1.38821 (rank : 40) | NC score | 0.028054 (rank : 60) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q92738, Q15400, Q7L0K9 | Gene names | USP6NL, KIAA0019 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USP6 N-terminal-like protein (Related to the N-terminus of tre) (RN- tre). | |||||
|
HEY2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 41) | NC score | 0.114995 (rank : 39) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9UBP5 | Gene names | HEY2, CHF1, GRL, HERP, HERP1, HRT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Cardiovascular helix-loop-helix factor 1) (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES- related repressor protein 2) (Protein gridlock homolog). | |||||
|
HEY2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 42) | NC score | 0.113097 (rank : 40) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9QUS4, Q3TZ99, Q8CD44 | Gene names | Hey2, Chf1, Herp, Herp1, Hesr2, Hrt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES-related repressor protein 2). | |||||
|
MAGC1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 43) | NC score | 0.031317 (rank : 57) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 431 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O60732, O75451, Q8TCV4 | Gene names | MAGEC1 | |||
|
Domain Architecture |

|
|||||
| Description | Melanoma-associated antigen C1 (MAGE-C1 antigen) (Cancer-testis antigen CT7). | |||||
|
PER3_HUMAN
|
||||||
| θ value | 1.81305 (rank : 44) | NC score | 0.202410 (rank : 36) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P56645, Q5H8X4, Q969K6, Q96S77, Q96S78, Q9C0J3, Q9NSP9, Q9UGU8 | Gene names | PER3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (hPER3). | |||||
|
WASIP_HUMAN
|
||||||
| θ value | 1.81305 (rank : 45) | NC score | 0.014081 (rank : 64) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O43516, Q15220, Q6MZU9, Q9BU37, Q9UNP1 | Gene names | WASPIP, WIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein) (PRPL-2 protein). | |||||
|
DYN1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 46) | NC score | 0.008335 (rank : 73) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P39053, Q61358, Q61359, Q61360, Q8JZZ4, Q9CSY7, Q9QXX1 | Gene names | Dnm1, Dnm | |||
|
Domain Architecture |

|
|||||
| Description | Dynamin-1 (EC 3.6.5.5). | |||||
|
GON4L_MOUSE
|
||||||
| θ value | 2.36792 (rank : 47) | NC score | 0.012214 (rank : 66) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9DB00, Q80TB4, Q91YI9 | Gene names | Gon4l, Gon4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
HEY1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 48) | NC score | 0.104269 (rank : 42) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9Y5J3, Q5TZS3, Q9NYP4 | Gene names | HEY1, CHF2, HESR1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1) (Cardiovascular helix-loop-helix factor 2) (HES-related repressor protein 2) (HERP2). | |||||
|
HEY1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 49) | NC score | 0.107983 (rank : 41) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9WV93 | Gene names | Hey1, Hesr1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1). | |||||
|
HXB3_MOUSE
|
||||||
| θ value | 2.36792 (rank : 50) | NC score | 0.004251 (rank : 83) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 493 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P09026, P10285, Q4PJ06, Q61680 | Gene names | Hoxb3, Hox-2.7, Hoxb-3 | |||
|
Domain Architecture |
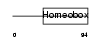
|
|||||
| Description | Homeobox protein Hox-B3 (Hox-2.7) (Homeobox protein MH-23). | |||||
|
NFH_MOUSE
|
||||||
| θ value | 2.36792 (rank : 51) | NC score | 0.006285 (rank : 79) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
CSKI2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 52) | NC score | 0.006111 (rank : 80) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 747 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8WXE0, Q7LG69, Q9ULT1 | Gene names | CASKIN2, KIAA1139 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-2. | |||||
|
DIDO1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 53) | NC score | 0.009467 (rank : 72) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
NCOA6_HUMAN
|
||||||
| θ value | 4.03905 (rank : 54) | NC score | 0.032161 (rank : 56) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 1202 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q14686, Q9NTZ9, Q9UH74, Q9UK86 | Gene names | NCOA6, AIB3, KIAA0181, RAP250, TRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (PRIP) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC RAP250) (Thyroid hormone receptor-binding protein). | |||||
|
SMOX_HUMAN
|
||||||
| θ value | 4.03905 (rank : 55) | NC score | 0.008335 (rank : 74) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NWM0, Q5TE26, Q5TE27, Q6UY28, Q8IX00, Q96LC3, Q96LC4, Q96QT3, Q9BW38, Q9H6H1, Q9NP51, Q9NPY1, Q9NPY2 | Gene names | SMOX, C20orf16, SMO | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Spermine oxidase (EC 1.5.3.-) (Polyamine oxidase 1) (PAO-1) (PAOh1). | |||||
|
TAF4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 56) | NC score | 0.010939 (rank : 68) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 687 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O00268, Q5TBP6, Q99721, Q9BR40, Q9BX42 | Gene names | TAF4, TAF2C, TAF2C1, TAF4A, TAFII130, TAFII135 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 4 (TBP-associated factor 4) (Transcription initiation factor TFIID 135 kDa subunit) (TAF(II)135) (TAFII-135) (TAFII135) (TAFII-130) (TAFII130). | |||||
|
ZEP1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 57) | NC score | 0.001185 (rank : 87) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 1036 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P15822, Q14122 | Gene names | HIVEP1, ZNF40 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 40 (Human immunodeficiency virus type I enhancer- binding protein 1) (HIV-EP1) (Major histocompatibility complex-binding protein 1) (MBP-1) (Positive regulatory domain II-binding factor 1) (PRDII-BF1). | |||||
|
BIN1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 58) | NC score | 0.008155 (rank : 75) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O00499, O00297, O00545, O43867, O60552, O60553, O60554, O60555, O75514, O75515, O75516, O75517, O75518, Q92944, Q99688 | Gene names | BIN1, AMPHL | |||
|
Domain Architecture |

|
|||||
| Description | Myc box-dependent-interacting protein 1 (Bridging integrator 1) (Amphiphysin-like protein) (Amphiphysin II) (Box-dependent myc- interacting protein 1). | |||||
|
PER3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 59) | NC score | 0.226327 (rank : 33) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O70361 | Gene names | Per3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (mPER3). | |||||
|
TNC6B_HUMAN
|
||||||
| θ value | 5.27518 (rank : 60) | NC score | 0.017178 (rank : 62) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UPQ9, Q5TH52, Q8TBX2 | Gene names | TNRC6B, KIAA1093 | |||
|
Domain Architecture |

|
|||||
| Description | Trinucleotide repeat-containing 6B protein. | |||||
|
BCL9_HUMAN
|
||||||
| θ value | 6.88961 (rank : 61) | NC score | 0.015845 (rank : 63) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 1008 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O00512 | Gene names | BCL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9) (Legless homolog). | |||||
|
ETV4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 62) | NC score | 0.003184 (rank : 84) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P28322 | Gene names | Etv4, Pea-3, Pea3 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 4 (Polyomavirus enhancer activator 3) (Protein PEA3). | |||||
|
HES5_MOUSE
|
||||||
| θ value | 6.88961 (rank : 63) | NC score | 0.073616 (rank : 43) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P70120, Q8BVI1 | Gene names | Hes5, Hes-5 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-5 (Hairy and enhancer of split 5). | |||||
|
MAST4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 64) | NC score | -0.002313 (rank : 90) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 1804 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O15021, Q6ZN07, Q96LY3 | Gene names | MAST4, KIAA0303 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase 4 (EC 2.7.11.1). | |||||
|
SYNJ1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 65) | NC score | 0.010559 (rank : 69) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O43426, O43425, O94984 | Gene names | SYNJ1, KIAA0910 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptojanin-1 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 1). | |||||
|
SYNJ1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 66) | NC score | 0.009791 (rank : 71) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8CHC4 | Gene names | Synj1, Kiaa0910 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptojanin-1 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 1). | |||||
|
TXLNB_HUMAN
|
||||||
| θ value | 6.88961 (rank : 67) | NC score | 0.001516 (rank : 86) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 912 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8N3L3, Q76L25, Q86T52, Q8N3S2 | Gene names | TXLNB, C6orf198, MDP77 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-taxilin (Muscle-derived protein 77) (hMDP77). | |||||
|
WNK2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 68) | NC score | -0.000749 (rank : 89) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 1407 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9Y3S1, Q8IY36, Q9C0A3, Q9H3P4 | Gene names | WNK2, KIAA1760, PRKWNK2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK2 (EC 2.7.11.1) (Protein kinase with no lysine 2) (Protein kinase, lysine-deficient 2). | |||||
|
ANK2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 69) | NC score | 0.002065 (rank : 85) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
DUT_HUMAN
|
||||||
| θ value | 8.99809 (rank : 70) | NC score | 0.011279 (rank : 67) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P33316, O14785, Q16708, Q16860, Q96Q81 | Gene names | DUT | |||
|
Domain Architecture |

|
|||||
| Description | Deoxyuridine 5'-triphosphate nucleotidohydrolase, mitochondrial precursor (EC 3.6.1.23) (dUTPase) (dUTP pyrophosphatase). | |||||
|
GPTC3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 71) | NC score | 0.006805 (rank : 77) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BIY1 | Gene names | Gpatc3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 3. | |||||
|
ILDR1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 72) | NC score | 0.006837 (rank : 76) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q86SU0, Q6ZP61, Q7Z578 | Gene names | ILDR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Immunoglobulin-like domain-containing receptor 1 precursor. | |||||
|
NCOR2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 73) | NC score | 0.005441 (rank : 81) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Y618, O00613, O15416, Q13354, Q9Y5U0 | Gene names | NCOR2, CTG26 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC) (CTG repeat protein 26) (SMAP270). | |||||
|
NDC1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 74) | NC score | 0.006751 (rank : 78) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8VCB1 | Gene names | Tmem48, Ndc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoporin NDC1 (Transmembrane protein 48). | |||||
|
TMPSD_MOUSE
|
||||||
| θ value | 8.99809 (rank : 75) | NC score | 0.000470 (rank : 88) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 334 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5U405, Q8CFE0, Q91VQ8 | Gene names | Tmprss13, Msp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protease, serine 13 (EC 3.4.21.-) (Mosaic serine protease) (Membrane-type mosaic serine protease). | |||||
|
US6NL_MOUSE
|
||||||
| θ value | 8.99809 (rank : 76) | NC score | 0.009817 (rank : 70) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q80XC3, Q6ZQK8 | Gene names | Usp6nl, Kiaa0019 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USP6 N-terminal-like protein. | |||||
|
MITF_HUMAN
|
||||||
| θ value | θ > 10 (rank : 77) | NC score | 0.069012 (rank : 47) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O75030, Q14841, Q9P2V0, Q9P2V1, Q9P2V2, Q9P2Y8 | Gene names | MITF | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
MITF_MOUSE
|
||||||
| θ value | θ > 10 (rank : 78) | NC score | 0.070716 (rank : 44) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q08874, O08885, O88203, Q08843, Q60781, Q60782, Q9JIJ0, Q9JIJ1, Q9JIJ2, Q9JIJ3, Q9JIJ4, Q9JIJ5, Q9JIJ6, Q9JKX9 | Gene names | Mitf, Bw, Mi, Vit | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
MLX_HUMAN
|
||||||
| θ value | θ > 10 (rank : 79) | NC score | 0.064821 (rank : 53) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UH92, Q53XM6, Q96FL2, Q9H2V0, Q9H2V1, Q9H2V2, Q9NXN3 | Gene names | MLX, TCFL4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
MLX_MOUSE
|
||||||
| θ value | θ > 10 (rank : 80) | NC score | 0.064860 (rank : 52) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O08609, Q9EQJ7, Q9EQJ8 | Gene names | Mlx, Tcfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
PER1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 81) | NC score | 0.202517 (rank : 35) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O15534 | Gene names | PER1, KIAA0482, PER, RIGUI | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (hPER). | |||||
|
PER1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 82) | NC score | 0.199271 (rank : 37) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O35973 | Gene names | Per1, Per, Rigui | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (mPER) (M-Rigui). | |||||
|
TCFL5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 83) | NC score | 0.070143 (rank : 45) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9UL49, O94771, Q9BYW0 | Gene names | TCFL5, CHA, E2BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor-like 5 protein (Cha transcription factor) (HPV-16 E2-binding protein 1) (E2BP-1). | |||||
|
TFE3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 84) | NC score | 0.069028 (rank : 46) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P19532, Q92757, Q92758, Q99964 | Gene names | TFE3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E3. | |||||
|
TFEB_HUMAN
|
||||||
| θ value | θ > 10 (rank : 85) | NC score | 0.065360 (rank : 50) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P19484, Q709B3, Q7Z6P9, Q9BRJ5, Q9UJD8 | Gene names | TFEB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
TFEB_MOUSE
|
||||||
| θ value | θ > 10 (rank : 86) | NC score | 0.065537 (rank : 48) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9R210 | Gene names | Tfeb, Tcfeb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
USF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 87) | NC score | 0.065360 (rank : 49) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P22415 | Gene names | USF1, USF | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 88) | NC score | 0.065330 (rank : 51) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q61069 | Gene names | Usf1, Usf | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 89) | NC score | 0.063002 (rank : 54) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q15853, O00671, O00709, Q05750, Q07952, Q15851, Q15852 | Gene names | USF2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (FOS- interacting protein) (FIP) (Major late transcription factor 2). | |||||
|
USF2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 90) | NC score | 0.059265 (rank : 55) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q64705, Q3UIL2, Q8VE59 | Gene names | Usf2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (Major late transcription factor 2). | |||||
|
SIM2_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | Q14190, O60766, Q15470, Q15471, Q15472, Q15473, Q16532 | Gene names | SIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2. | |||||
|
SIM2_MOUSE
|
||||||
| NC score | 0.995700 (rank : 2) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q61079, O35391, Q61046, Q61904 | Gene names | Sim2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2 (SIM transcription factor) (mSIM). | |||||
|
SIM1_HUMAN
|
||||||
| NC score | 0.988909 (rank : 3) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P81133, Q5TDP7 | Gene names | SIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1. | |||||
|
SIM1_MOUSE
|
||||||
| NC score | 0.988506 (rank : 4) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q61045, O70284, P70183 | Gene names | Sim1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1 (mSIM1). | |||||
|
NPAS1_MOUSE
|
||||||
| NC score | 0.954256 (rank : 5) | θ value | 5.8972e-76 (rank : 7) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P97459 | Gene names | Npas1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1). | |||||
|
NPAS1_HUMAN
|
||||||
| NC score | 0.953012 (rank : 6) | θ value | 2.09745e-73 (rank : 10) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q99742, Q99632, Q9BY83 | Gene names | NPAS1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1) (Member of PAS protein 5) (MOP5). | |||||
|
NPAS3_HUMAN
|
||||||
| NC score | 0.946602 (rank : 7) | θ value | 2.0926e-81 (rank : 5) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8IXF0, Q86US6, Q86US7, Q8IXF2, Q9BY81, Q9H323, Q9Y4L8 | Gene names | NPAS3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
NPAS3_MOUSE
|
||||||
| NC score | 0.945507 (rank : 8) | θ value | 2.0926e-81 (rank : 6) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9QZQ0, Q9EQP4 | Gene names | Npas3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
HIF1A_MOUSE
|
||||||
| NC score | 0.936152 (rank : 9) | θ value | 1.89707e-74 (rank : 8) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q61221, O08741, O08993, Q61664, Q61665, Q8C681, Q8CC19, Q8CCB6, Q8R385, Q9CYA8 | Gene names | Hif1a | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein). | |||||
|
HIF1A_HUMAN
|
||||||
| NC score | 0.933296 (rank : 10) | θ value | 9.41505e-74 (rank : 9) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q16665, Q96PT9, Q9UPB1 | Gene names | HIF1A | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein) (Member of PAS protein 1) (MOP1). | |||||
|
EPAS1_HUMAN
|
||||||
| NC score | 0.924438 (rank : 11) | θ value | 9.76566e-63 (rank : 11) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q99814, Q86VA2, Q99630 | Gene names | EPAS1, HIF2A | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Member of PAS protein 2) (MOP2) (Hypoxia-inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (HLF). | |||||
|
EPAS1_MOUSE
|
||||||
| NC score | 0.923527 (rank : 12) | θ value | 4.10295e-61 (rank : 12) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P97481, O08787, O55046 | Gene names | Epas1, Hif2a | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Hypoxia- inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (MHLF) (HIF-related factor) (HRF). | |||||
|
AHR_HUMAN
|
||||||
| NC score | 0.842738 (rank : 13) | θ value | 3.17079e-29 (rank : 14) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P35869, Q13728, Q13803, Q13804 | Gene names | AHR | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
AHR_MOUSE
|
||||||
| NC score | 0.841801 (rank : 14) | θ value | 1.28732e-30 (rank : 13) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P30561, Q8QZX6, Q8R4S3, Q99P79, Q9QVY1 | Gene names | Ahr | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
BMAL1_HUMAN
|
||||||
| NC score | 0.793198 (rank : 15) | θ value | 3.50572e-28 (rank : 16) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O00327, O00313, O00314, O00315, O00316, O00317, Q99631, Q99649 | Gene names | ARNTL, BMAL1, MOP3 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Member of PAS protein 3) (Basic-helix-loop- helix-PAS orphan MOP3) (bHLH-PAS protein JAP3). | |||||
|
BMAL1_MOUSE
|
||||||
| NC score | 0.791811 (rank : 16) | θ value | 4.14116e-29 (rank : 15) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9WTL8, O88295, Q921S4, Q9R0U2, Q9WTL9 | Gene names | Arntl, Bmal1 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Arnt3). | |||||
|
CLOCK_HUMAN
|
||||||
| NC score | 0.760858 (rank : 17) | θ value | 1.6852e-22 (rank : 19) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O15516, O14516, Q9UIT8 | Gene names | CLOCK, KIAA0334 | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (hCLOCK). | |||||
|
CLOCK_MOUSE
|
||||||
| NC score | 0.757698 (rank : 18) | θ value | 2.20094e-22 (rank : 20) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O08785 | Gene names | Clock | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (mCLOCK). | |||||
|
ARNT2_HUMAN
|
||||||
| NC score | 0.748487 (rank : 19) | θ value | 2.77775e-25 (rank : 17) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9HBZ2, O15024 | Gene names | ARNT2, KIAA0307 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
ARNT2_MOUSE
|
||||||
| NC score | 0.747171 (rank : 20) | θ value | 2.77775e-25 (rank : 18) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q61324 | Gene names | Arnt2 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
ARNT_HUMAN
|
||||||
| NC score | 0.736844 (rank : 21) | θ value | 3.17815e-21 (rank : 22) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P27540 | Gene names | ARNT | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
NPAS2_MOUSE
|
||||||
| NC score | 0.735514 (rank : 22) | θ value | 2.27762e-19 (rank : 25) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P97460 | Gene names | Npas2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2). | |||||
|
NPAS2_HUMAN
|
||||||
| NC score | 0.727511 (rank : 23) | θ value | 5.60996e-18 (rank : 26) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q99743, Q99629 | Gene names | NPAS2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2) (Member of PAS protein 4) (MOP4). | |||||
|
ARNT_MOUSE
|
||||||
| NC score | 0.724561 (rank : 24) | θ value | 3.17815e-21 (rank : 23) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P53762, Q60661 | Gene names | Arnt | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
NPAS4_HUMAN
|
||||||
| NC score | 0.704444 (rank : 25) | θ value | 2.20094e-22 (rank : 21) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8IUM7, Q8N8S5, Q8N9Q9 | Gene names | NPAS4, NXF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF). | |||||
|
NPAS4_MOUSE
|
||||||
| NC score | 0.688795 (rank : 26) | θ value | 1.2077e-20 (rank : 24) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8BGD7, Q3V3U3 | Gene names | Npas4, Nxf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF) (Limbic-enhanced PAS protein) (LE-PAS). | |||||
|
NCOA1_MOUSE
|
||||||
| NC score | 0.633834 (rank : 27) | θ value | 5.25075e-16 (rank : 27) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P70365, P70366, Q61202, Q8CBI9 | Gene names | Ncoa1, Src1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (Nuclear receptor coactivator protein 1) (mNRC-1). | |||||
|
NCOA1_HUMAN
|
||||||
| NC score | 0.630315 (rank : 28) | θ value | 6.85773e-16 (rank : 28) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q15788, O00150, O43792, O43793, Q13071, Q13420, Q6GVI5, Q7KYV3 | Gene names | NCOA1, SRC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (RIP160) (Protein Hin-2) (NY-REN-52 antigen). | |||||
|
NCOA2_HUMAN
|
||||||
| NC score | 0.556651 (rank : 29) | θ value | 2.28291e-11 (rank : 29) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q15596 | Gene names | NCOA2, TIF2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 2 (NCoA-2) (Transcriptional intermediary factor 2). | |||||
|
NCOA2_MOUSE
|
||||||
| NC score | 0.554927 (rank : 30) | θ value | 2.52405e-10 (rank : 30) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q61026, O09001, P97759 | Gene names | Ncoa2, Grip1, Tif2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 2 (NCoA-2) (Transcriptional intermediary factor 2) (Glucocorticoid receptor-interacting protein 1) (GRIP-1). | |||||
|
NCOA3_HUMAN
|
||||||
| NC score | 0.553234 (rank : 31) | θ value | 1.38499e-08 (rank : 31) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9Y6Q9, Q5JYD9, Q5JYE0, Q9BR49, Q9UPC9, Q9UPG4, Q9UPG7 | Gene names | NCOA3, AIB1, RAC3, TRAM1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP-interacting protein) (pCIP). | |||||
|
NCOA3_MOUSE
|
||||||
| NC score | 0.517247 (rank : 32) | θ value | 1.53129e-07 (rank : 32) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O09000, Q9CRD5 | Gene names | Ncoa3, Aib1, Pcip, Rac3, Tram1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein homolog) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP- interacting protein) (p/CIP) (pCIP). | |||||
|
PER3_MOUSE
|
||||||
| NC score | 0.226327 (rank : 33) | θ value | 5.27518 (rank : 59) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O70361 | Gene names | Per3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (mPER3). | |||||
|
PER2_HUMAN
|
||||||
| NC score | 0.216490 (rank : 34) | θ value | 0.813845 (rank : 35) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O15055 | Gene names | PER2, KIAA0347 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2. | |||||
|
PER1_HUMAN
|
||||||
| NC score | 0.202517 (rank : 35) | θ value | θ > 10 (rank : 81) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O15534 | Gene names | PER1, KIAA0482, PER, RIGUI | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (hPER). | |||||
|
PER3_HUMAN
|
||||||
| NC score | 0.202410 (rank : 36) | θ value | 1.81305 (rank : 44) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P56645, Q5H8X4, Q969K6, Q96S77, Q96S78, Q9C0J3, Q9NSP9, Q9UGU8 | Gene names | PER3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (hPER3). | |||||
|
PER1_MOUSE
|
||||||
| NC score | 0.199271 (rank : 37) | θ value | θ > 10 (rank : 82) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O35973 | Gene names | Per1, Per, Rigui | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (mPER) (M-Rigui). | |||||
|
PER2_MOUSE
|
||||||
| NC score | 0.196936 (rank : 38) | θ value | 1.06291 (rank : 36) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O54943, O54954 | Gene names | Per2 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2 (mPER2). | |||||
|
HEY2_HUMAN
|
||||||
| NC score | 0.114995 (rank : 39) | θ value | 1.81305 (rank : 41) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9UBP5 | Gene names | HEY2, CHF1, GRL, HERP, HERP1, HRT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Cardiovascular helix-loop-helix factor 1) (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES- related repressor protein 2) (Protein gridlock homolog). | |||||
|
HEY2_MOUSE
|
||||||
| NC score | 0.113097 (rank : 40) | θ value | 1.81305 (rank : 42) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9QUS4, Q3TZ99, Q8CD44 | Gene names | Hey2, Chf1, Herp, Herp1, Hesr2, Hrt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES-related repressor protein 2). | |||||
|
HEY1_MOUSE
|
||||||
| NC score | 0.107983 (rank : 41) | θ value | 2.36792 (rank : 49) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9WV93 | Gene names | Hey1, Hesr1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1). | |||||
|
HEY1_HUMAN
|
||||||
| NC score | 0.104269 (rank : 42) | θ value | 2.36792 (rank : 48) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9Y5J3, Q5TZS3, Q9NYP4 | Gene names | HEY1, CHF2, HESR1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1) (Cardiovascular helix-loop-helix factor 2) (HES-related repressor protein 2) (HERP2). | |||||
|
HES5_MOUSE
|
||||||
| NC score | 0.073616 (rank : 43) | θ value | 6.88961 (rank : 63) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P70120, Q8BVI1 | Gene names | Hes5, Hes-5 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-5 (Hairy and enhancer of split 5). | |||||
|
MITF_MOUSE
|
||||||
| NC score | 0.070716 (rank : 44) | θ value | θ > 10 (rank : 78) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q08874, O08885, O88203, Q08843, Q60781, Q60782, Q9JIJ0, Q9JIJ1, Q9JIJ2, Q9JIJ3, Q9JIJ4, Q9JIJ5, Q9JIJ6, Q9JKX9 | Gene names | Mitf, Bw, Mi, Vit | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
TCFL5_HUMAN
|
||||||
| NC score | 0.070143 (rank : 45) | θ value | θ > 10 (rank : 83) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9UL49, O94771, Q9BYW0 | Gene names | TCFL5, CHA, E2BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor-like 5 protein (Cha transcription factor) (HPV-16 E2-binding protein 1) (E2BP-1). | |||||
|
TFE3_HUMAN
|
||||||
| NC score | 0.069028 (rank : 46) | θ value | θ > 10 (rank : 84) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P19532, Q92757, Q92758, Q99964 | Gene names | TFE3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E3. | |||||
|
MITF_HUMAN
|
||||||
| NC score | 0.069012 (rank : 47) | θ value | θ > 10 (rank : 77) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O75030, Q14841, Q9P2V0, Q9P2V1, Q9P2V2, Q9P2Y8 | Gene names | MITF | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
TFEB_MOUSE
|
||||||
| NC score | 0.065537 (rank : 48) | θ value | θ > 10 (rank : 86) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9R210 | Gene names | Tfeb, Tcfeb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
USF1_HUMAN
|
||||||
| NC score | 0.065360 (rank : 49) | θ value | θ > 10 (rank : 87) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P22415 | Gene names | USF1, USF | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
TFEB_HUMAN
|
||||||
| NC score | 0.065360 (rank : 50) | θ value | θ > 10 (rank : 85) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P19484, Q709B3, Q7Z6P9, Q9BRJ5, Q9UJD8 | Gene names | TFEB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
USF1_MOUSE
|
||||||
| NC score | 0.065330 (rank : 51) | θ value | θ > 10 (rank : 88) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q61069 | Gene names | Usf1, Usf | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
MLX_MOUSE
|
||||||
| NC score | 0.064860 (rank : 52) | θ value | θ > 10 (rank : 80) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O08609, Q9EQJ7, Q9EQJ8 | Gene names | Mlx, Tcfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
MLX_HUMAN
|
||||||
| NC score | 0.064821 (rank : 53) | θ value | θ > 10 (rank : 79) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UH92, Q53XM6, Q96FL2, Q9H2V0, Q9H2V1, Q9H2V2, Q9NXN3 | Gene names | MLX, TCFL4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
USF2_HUMAN
|
||||||
| NC score | 0.063002 (rank : 54) | θ value | θ > 10 (rank : 89) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q15853, O00671, O00709, Q05750, Q07952, Q15851, Q15852 | Gene names | USF2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (FOS- interacting protein) (FIP) (Major late transcription factor 2). | |||||
|
USF2_MOUSE
|
||||||
| NC score | 0.059265 (rank : 55) | θ value | θ > 10 (rank : 90) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q64705, Q3UIL2, Q8VE59 | Gene names | Usf2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (Major late transcription factor 2). | |||||
|
NCOA6_HUMAN
|
||||||
| NC score | 0.032161 (rank : 56) | θ value | 4.03905 (rank : 54) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 1202 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q14686, Q9NTZ9, Q9UH74, Q9UK86 | Gene names | NCOA6, AIB3, KIAA0181, RAP250, TRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (PRIP) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC RAP250) (Thyroid hormone receptor-binding protein). | |||||
|
MAGC1_HUMAN
|
||||||
| NC score | 0.031317 (rank : 57) | θ value | 1.81305 (rank : 43) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 431 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O60732, O75451, Q8TCV4 | Gene names | MAGEC1 | |||
|
Domain Architecture |

|
|||||
| Description | Melanoma-associated antigen C1 (MAGE-C1 antigen) (Cancer-testis antigen CT7). | |||||
|
BSN_HUMAN
|
||||||
| NC score | 0.031121 (rank : 58) | θ value | 0.365318 (rank : 33) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
ZN645_HUMAN
|
||||||
| NC score | 0.030760 (rank : 59) | θ value | 0.62314 (rank : 34) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 265 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8N7E2, Q6DJY9 | Gene names | ZNF645 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 645. | |||||
|
US6NL_HUMAN
|
||||||
| NC score | 0.028054 (rank : 60) | θ value | 1.38821 (rank : 40) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q92738, Q15400, Q7L0K9 | Gene names | USP6NL, KIAA0019 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USP6 N-terminal-like protein (Related to the N-terminus of tre) (RN- tre). | |||||
|
EXOC3_MOUSE
|
||||||
| NC score | 0.024503 (rank : 61) | θ value | 1.38821 (rank : 38) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6KAR6 | Gene names | Exoc3, Sec6l1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Exocyst complex component 3 (Exocyst complex component Sec6). | |||||
|
TNC6B_HUMAN
|
||||||
| NC score | 0.017178 (rank : 62) | θ value | 5.27518 (rank : 60) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UPQ9, Q5TH52, Q8TBX2 | Gene names | TNRC6B, KIAA1093 | |||
|
Domain Architecture |

|
|||||
| Description | Trinucleotide repeat-containing 6B protein. | |||||
|
BCL9_HUMAN
|
||||||
| NC score | 0.015845 (rank : 63) | θ value | 6.88961 (rank : 61) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 1008 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O00512 | Gene names | BCL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9) (Legless homolog). | |||||
|
WASIP_HUMAN
|
||||||
| NC score | 0.014081 (rank : 64) | θ value | 1.81305 (rank : 45) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O43516, Q15220, Q6MZU9, Q9BU37, Q9UNP1 | Gene names | WASPIP, WIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein) (PRPL-2 protein). | |||||
|
RTN4_HUMAN
|
||||||
| NC score | 0.013308 (rank : 65) | θ value | 1.06291 (rank : 37) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NQC3, O94962, Q9BXG5, Q9H212, Q9H3I3, Q9UQ42, Q9Y293, Q9Y2Y7, Q9Y5U6 | Gene names | RTN4, KIAA0886, NOGO | |||
|
Domain Architecture |

|
|||||
| Description | Reticulon-4 (Neurite outgrowth inhibitor) (Nogo protein) (Foocen) (Neuroendocrine-specific protein) (NSP) (Neuroendocrine-specific protein C homolog) (RTN-x) (Reticulon-5). | |||||
|
GON4L_MOUSE
|
||||||
| NC score | 0.012214 (rank : 66) | θ value | 2.36792 (rank : 47) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9DB00, Q80TB4, Q91YI9 | Gene names | Gon4l, Gon4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
DUT_HUMAN
|
||||||
| NC score | 0.011279 (rank : 67) | θ value | 8.99809 (rank : 70) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P33316, O14785, Q16708, Q16860, Q96Q81 | Gene names | DUT | |||
|
Domain Architecture |

|
|||||
| Description | Deoxyuridine 5'-triphosphate nucleotidohydrolase, mitochondrial precursor (EC 3.6.1.23) (dUTPase) (dUTP pyrophosphatase). | |||||
|
TAF4_HUMAN
|
||||||
| NC score | 0.010939 (rank : 68) | θ value | 4.03905 (rank : 56) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 687 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O00268, Q5TBP6, Q99721, Q9BR40, Q9BX42 | Gene names | TAF4, TAF2C, TAF2C1, TAF4A, TAFII130, TAFII135 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 4 (TBP-associated factor 4) (Transcription initiation factor TFIID 135 kDa subunit) (TAF(II)135) (TAFII-135) (TAFII135) (TAFII-130) (TAFII130). | |||||
|
SYNJ1_HUMAN
|
||||||
| NC score | 0.010559 (rank : 69) | θ value | 6.88961 (rank : 65) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O43426, O43425, O94984 | Gene names | SYNJ1, KIAA0910 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptojanin-1 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 1). | |||||
|
US6NL_MOUSE
|
||||||
| NC score | 0.009817 (rank : 70) | θ value | 8.99809 (rank : 76) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q80XC3, Q6ZQK8 | Gene names | Usp6nl, Kiaa0019 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USP6 N-terminal-like protein. | |||||
|
SYNJ1_MOUSE
|
||||||
| NC score | 0.009791 (rank : 71) | θ value | 6.88961 (rank : 66) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8CHC4 | Gene names | Synj1, Kiaa0910 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptojanin-1 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 1). | |||||
|
DIDO1_HUMAN
|
||||||
| NC score | 0.009467 (rank : 72) | θ value | 4.03905 (rank : 53) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
DYN1_MOUSE
|
||||||
| NC score | 0.008335 (rank : 73) | θ value | 2.36792 (rank : 46) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P39053, Q61358, Q61359, Q61360, Q8JZZ4, Q9CSY7, Q9QXX1 | Gene names | Dnm1, Dnm | |||
|
Domain Architecture |

|
|||||
| Description | Dynamin-1 (EC 3.6.5.5). | |||||
|
SMOX_HUMAN
|
||||||
| NC score | 0.008335 (rank : 74) | θ value | 4.03905 (rank : 55) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NWM0, Q5TE26, Q5TE27, Q6UY28, Q8IX00, Q96LC3, Q96LC4, Q96QT3, Q9BW38, Q9H6H1, Q9NP51, Q9NPY1, Q9NPY2 | Gene names | SMOX, C20orf16, SMO | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Spermine oxidase (EC 1.5.3.-) (Polyamine oxidase 1) (PAO-1) (PAOh1). | |||||
|
BIN1_HUMAN
|
||||||
| NC score | 0.008155 (rank : 75) | θ value | 5.27518 (rank : 58) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O00499, O00297, O00545, O43867, O60552, O60553, O60554, O60555, O75514, O75515, O75516, O75517, O75518, Q92944, Q99688 | Gene names | BIN1, AMPHL | |||
|
Domain Architecture |

|
|||||
| Description | Myc box-dependent-interacting protein 1 (Bridging integrator 1) (Amphiphysin-like protein) (Amphiphysin II) (Box-dependent myc- interacting protein 1). | |||||
|
ILDR1_HUMAN
|
||||||
| NC score | 0.006837 (rank : 76) | θ value | 8.99809 (rank : 72) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q86SU0, Q6ZP61, Q7Z578 | Gene names | ILDR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Immunoglobulin-like domain-containing receptor 1 precursor. | |||||
|
GPTC3_MOUSE
|
||||||
| NC score | 0.006805 (rank : 77) | θ value | 8.99809 (rank : 71) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BIY1 | Gene names | Gpatc3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 3. | |||||
|
NDC1_MOUSE
|
||||||
| NC score | 0.006751 (rank : 78) | θ value | 8.99809 (rank : 74) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8VCB1 | Gene names | Tmem48, Ndc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoporin NDC1 (Transmembrane protein 48). | |||||
|
NFH_MOUSE
|
||||||
| NC score | 0.006285 (rank : 79) | θ value | 2.36792 (rank : 51) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
CSKI2_HUMAN
|
||||||
| NC score | 0.006111 (rank : 80) | θ value | 3.0926 (rank : 52) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 747 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8WXE0, Q7LG69, Q9ULT1 | Gene names | CASKIN2, KIAA1139 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-2. | |||||
|
NCOR2_HUMAN
|
||||||
| NC score | 0.005441 (rank : 81) | θ value | 8.99809 (rank : 73) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Y618, O00613, O15416, Q13354, Q9Y5U0 | Gene names | NCOR2, CTG26 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC) (CTG repeat protein 26) (SMAP270). | |||||
|
HXB3_HUMAN
|
||||||
| NC score | 0.005041 (rank : 82) | θ value | 1.38821 (rank : 39) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 559 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P14651, O95615, P17484 | Gene names | HOXB3, HOX2G | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-B3 (Hox-2G) (Hox-2.7). | |||||
|
HXB3_MOUSE
|
||||||
| NC score | 0.004251 (rank : 83) | θ value | 2.36792 (rank : 50) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 493 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P09026, P10285, Q4PJ06, Q61680 | Gene names | Hoxb3, Hox-2.7, Hoxb-3 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-B3 (Hox-2.7) (Homeobox protein MH-23). | |||||
|
ETV4_MOUSE
|
||||||
| NC score | 0.003184 (rank : 84) | θ value | 6.88961 (rank : 62) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P28322 | Gene names | Etv4, Pea-3, Pea3 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 4 (Polyomavirus enhancer activator 3) (Protein PEA3). | |||||
|
ANK2_HUMAN
|
||||||
| NC score | 0.002065 (rank : 85) | θ value | 8.99809 (rank : 69) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
TXLNB_HUMAN
|
||||||
| NC score | 0.001516 (rank : 86) | θ value | 6.88961 (rank : 67) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 912 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8N3L3, Q76L25, Q86T52, Q8N3S2 | Gene names | TXLNB, C6orf198, MDP77 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-taxilin (Muscle-derived protein 77) (hMDP77). | |||||
|
ZEP1_HUMAN
|
||||||
| NC score | 0.001185 (rank : 87) | θ value | 4.03905 (rank : 57) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 1036 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P15822, Q14122 | Gene names | HIVEP1, ZNF40 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 40 (Human immunodeficiency virus type I enhancer- binding protein 1) (HIV-EP1) (Major histocompatibility complex-binding protein 1) (MBP-1) (Positive regulatory domain II-binding factor 1) (PRDII-BF1). | |||||
|
TMPSD_MOUSE
|
||||||
| NC score | 0.000470 (rank : 88) | θ value | 8.99809 (rank : 75) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 334 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5U405, Q8CFE0, Q91VQ8 | Gene names | Tmprss13, Msp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protease, serine 13 (EC 3.4.21.-) (Mosaic serine protease) (Membrane-type mosaic serine protease). | |||||
|
WNK2_HUMAN
|
||||||
| NC score | -0.000749 (rank : 89) | θ value | 6.88961 (rank : 68) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 1407 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9Y3S1, Q8IY36, Q9C0A3, Q9H3P4 | Gene names | WNK2, KIAA1760, PRKWNK2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK2 (EC 2.7.11.1) (Protein kinase with no lysine 2) (Protein kinase, lysine-deficient 2). | |||||
|
MAST4_HUMAN
|
||||||
| NC score | -0.002313 (rank : 90) | θ value | 6.88961 (rank : 64) | |||
| Query Neighborhood Hits | 76 | Target Neighborhood Hits | 1804 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O15021, Q6ZN07, Q96LY3 | Gene names | MAST4, KIAA0303 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase 4 (EC 2.7.11.1). | |||||