Please be patient as the page loads
|
US6NL_MOUSE
|
||||||
| SwissProt Accessions | Q80XC3, Q6ZQK8 | Gene names | Usp6nl, Kiaa0019 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USP6 N-terminal-like protein. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
US6NL_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.958673 (rank : 2) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q92738, Q15400, Q7L0K9 | Gene names | USP6NL, KIAA0019 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USP6 N-terminal-like protein (Related to the N-terminus of tre) (RN- tre). | |||||
|
US6NL_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 157 | |
| SwissProt Accessions | Q80XC3, Q6ZQK8 | Gene names | Usp6nl, Kiaa0019 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USP6 N-terminal-like protein. | |||||
|
UBP6_HUMAN
|
||||||
| θ value | 2.09745e-73 (rank : 3) | NC score | 0.520774 (rank : 17) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P35125, Q15634, Q86WP6, Q8IWT4 | Gene names | USP6, TRE2 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 6 (EC 3.1.2.15) (Ubiquitin thioesterase 6) (Ubiquitin-specific-processing protease 6) (Deubiquitinating enzyme 6) (Proto-oncogene TRE-2). | |||||
|
TBCD3_HUMAN
|
||||||
| θ value | 3.03575e-64 (rank : 4) | NC score | 0.870449 (rank : 3) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8IZP1, Q9H0B9, Q9UDD4 | Gene names | TBC1D3, PRC17 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 3 (Rab GTPase-activating protein PRC17) (Prostate cancer gene 17 protein) (TRE17 alpha protein). | |||||
|
TBC10_MOUSE
|
||||||
| θ value | 9.8567e-31 (rank : 5) | NC score | 0.805776 (rank : 5) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P58802 | Gene names | Tbc1d10a, Epi64, Tbc1d10 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 10A (EBP50-PDX interactor of 64 kDa) (EPI64 protein). | |||||
|
TBC10_HUMAN
|
||||||
| θ value | 8.3442e-30 (rank : 6) | NC score | 0.809232 (rank : 4) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9BXI6, O76053, Q543A2 | Gene names | TBC1D10A, EPI64, TBC1D10 | |||
|
Domain Architecture |
|
|||||
| Description | TBC1 domain family member 10A (EBP50-PDX interactor of 64 kDa) (EPI64 protein). | |||||
|
TBCD1_HUMAN
|
||||||
| θ value | 1.33217e-27 (rank : 7) | NC score | 0.699563 (rank : 9) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q86TI0, Q96K82, Q9UPP4 | Gene names | TBC1D1, KIAA1108 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 1. | |||||
|
TBCD1_MOUSE
|
||||||
| θ value | 1.33217e-27 (rank : 8) | NC score | 0.693071 (rank : 10) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q60949, Q80TJ9, Q923F8 | Gene names | Tbc1d1, Kiaa1108, Tbc1 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 1. | |||||
|
TBCD4_MOUSE
|
||||||
| θ value | 7.30988e-26 (rank : 9) | NC score | 0.668202 (rank : 11) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8BYJ6, Q5DU23, Q66JU2, Q6P2M2, Q8BMH6, Q8BXM2 | Gene names | Tbc1d4, Kiaa0603 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 4 (Akt substrate of 160 kDa) (AS160). | |||||
|
TBCD4_HUMAN
|
||||||
| θ value | 1.37858e-24 (rank : 10) | NC score | 0.658094 (rank : 13) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O60343, Q5W0B9 | Gene names | TBC1D4, KIAA0603 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 4 (Akt substrate of 160 kDa) (AS160). | |||||
|
TBC12_HUMAN
|
||||||
| θ value | 1.80048e-24 (rank : 11) | NC score | 0.736634 (rank : 6) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O60347, Q5VYA6, Q8WX26, Q8WX59, Q9UG83 | Gene names | TBC1D12, KIAA0608 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 12. | |||||
|
TBC14_HUMAN
|
||||||
| θ value | 5.79196e-23 (rank : 12) | NC score | 0.707853 (rank : 7) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9P2M4, Q8IW15 | Gene names | TBC1D14, KIAA1322 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 14. | |||||
|
TBC14_MOUSE
|
||||||
| θ value | 6.40375e-22 (rank : 13) | NC score | 0.703514 (rank : 8) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8CGA2, Q8CHA5 | Gene names | Tbc1d14, D5Ertd110e, Kiaa1322 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 14. | |||||
|
TBCD2_HUMAN
|
||||||
| θ value | 7.82807e-20 (rank : 14) | NC score | 0.642058 (rank : 15) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9BYX2 | Gene names | TBC1D2, PARIS1 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 2 (Prostate antigen recognized and identified by SEREX) (PARIS-1). | |||||
|
TBCD8_HUMAN
|
||||||
| θ value | 1.33526e-19 (rank : 15) | NC score | 0.654078 (rank : 14) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O95759, Q9UQ32 | Gene names | TBC1D8, VRP | |||
|
Domain Architecture |
|
|||||
| Description | TBC1 domain family member 8 (Vascular Rab-GAP/TBC-containing protein) (AD 3). | |||||
|
TBCD8_MOUSE
|
||||||
| θ value | 6.62687e-19 (rank : 16) | NC score | 0.658989 (rank : 12) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9Z1A9 | Gene names | Tbc1d8, Hblp1, Vrp | |||
|
Domain Architecture |
|
|||||
| Description | TBC1 domain family member 8 (Vascular Rab-GAP/TBC-containing protein) (BUB2-like protein 1). | |||||
|
TBC16_HUMAN
|
||||||
| θ value | 1.68911e-14 (rank : 17) | NC score | 0.607441 (rank : 16) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8TBP0 | Gene names | TBC1D16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 16. | |||||
|
TBC17_MOUSE
|
||||||
| θ value | 6.64225e-11 (rank : 18) | NC score | 0.474905 (rank : 19) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BYH7 | Gene names | Tbc1d17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 17. | |||||
|
TBC15_HUMAN
|
||||||
| θ value | 1.47974e-10 (rank : 19) | NC score | 0.471316 (rank : 20) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8TC07 | Gene names | TBC1D15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 15. | |||||
|
TBC15_MOUSE
|
||||||
| θ value | 1.9326e-10 (rank : 20) | NC score | 0.475562 (rank : 18) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9CXF4, Q3UI41 | Gene names | Tbc1d15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 15. | |||||
|
TBC17_HUMAN
|
||||||
| θ value | 1.25267e-09 (rank : 21) | NC score | 0.463924 (rank : 21) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9HA65 | Gene names | TBC1D17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 17. | |||||
|
TB22A_HUMAN
|
||||||
| θ value | 3.41135e-07 (rank : 22) | NC score | 0.361663 (rank : 24) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8WUA7, Q5TE47, Q6ZUH2, Q92680, Q9BVD6, Q9UGG0, Q9UGT2, Q9UGU6, Q9UH25, Q9Y4W5 | Gene names | TBC1D22A, C22orf4 | |||
|
Domain Architecture |
|
|||||
| Description | TBC1 domain family member 22A. | |||||
|
TB22A_MOUSE
|
||||||
| θ value | 7.59969e-07 (rank : 23) | NC score | 0.364189 (rank : 23) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8R5A6, Q3U268, Q3U3T0, Q8CA49 | Gene names | Tbc1d22a, D15Ertd781e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 22A. | |||||
|
TB22B_HUMAN
|
||||||
| θ value | 0.000158464 (rank : 24) | NC score | 0.366128 (rank : 22) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NU19, Q5VUK9, Q7Z6P7, Q9BPV6, Q9BUT5, Q9NXB6 | Gene names | TBC1D22B, C6orf197 | |||
|
Domain Architecture |
|
|||||
| Description | TBC1 domain family member 22B. | |||||
|
TARA_HUMAN
|
||||||
| θ value | 0.000602161 (rank : 25) | NC score | 0.059624 (rank : 49) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9H2D6, O94797, Q2PZW8, Q2Q3Z9, Q2Q400, Q5R3M6, Q96DW1, Q9BT77, Q9BTL7, Q9BY98, Q9Y3L4 | Gene names | TRIOBP, KIAA1662, TARA | |||
|
Domain Architecture |

|
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
ANK2_HUMAN
|
||||||
| θ value | 0.00228821 (rank : 26) | NC score | 0.029511 (rank : 81) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
RPB1_HUMAN
|
||||||
| θ value | 0.00228821 (rank : 27) | NC score | 0.098576 (rank : 33) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P24928 | Gene names | POLR2A | |||
|
Domain Architecture |

|
|||||
| Description | DNA-directed RNA polymerase II largest subunit (EC 2.7.7.6) (RPB1). | |||||
|
RPB1_MOUSE
|
||||||
| θ value | 0.00228821 (rank : 28) | NC score | 0.099401 (rank : 32) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 363 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P08775 | Gene names | Polr2a, Rpii215, Rpo2-1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-directed RNA polymerase II largest subunit (EC 2.7.7.6) (RPB1). | |||||
|
SSXT_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 29) | NC score | 0.065779 (rank : 43) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q62280 | Gene names | Ss18, Ssxt, Syt | |||
|
Domain Architecture |
|
|||||
| Description | SSXT protein (SYT protein) (Synovial sarcoma-associated Ss18-alpha). | |||||
|
DCC_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 30) | NC score | 0.017723 (rank : 124) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P70211 | Gene names | Dcc | |||
|
Domain Architecture |

|
|||||
| Description | Netrin receptor DCC precursor (Tumor suppressor protein DCC). | |||||
|
SSXT_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 31) | NC score | 0.065337 (rank : 45) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q15532, Q16404, Q9BXC6 | Gene names | SS18, SSXT, SYT | |||
|
Domain Architecture |
|
|||||
| Description | SSXT protein (Synovial sarcoma, translocated to X chromosome) (SYT protein). | |||||
|
ATN1_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 32) | NC score | 0.053728 (rank : 54) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
E41L1_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 33) | NC score | 0.031109 (rank : 76) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Z2H5 | Gene names | Epb41l1, Epb4, Epb4.1l1 | |||
|
Domain Architecture |

|
|||||
| Description | Band 4.1-like protein 1 (Neuronal protein 4.1) (4.1N). | |||||
|
SMCA2_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 34) | NC score | 0.018352 (rank : 123) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 503 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P51531 | Gene names | SMARCA2, BRM, SNF2A, SNF2L2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L2 (EC 3.6.1.-) (ATP- dependent helicase SMARCA2) (SNF2-alpha) (SWI/SNF-related matrix- associated actin-dependent regulator of chromatin subfamily A member 2) (hBRM). | |||||
|
E41L1_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 35) | NC score | 0.030271 (rank : 78) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9H4G0, O15046, Q4VXN4 | Gene names | EPB41L1, KIAA0338 | |||
|
Domain Architecture |

|
|||||
| Description | Band 4.1-like protein 1 (Neuronal protein 4.1) (4.1N). | |||||
|
MBD5_HUMAN
|
||||||
| θ value | 0.125558 (rank : 36) | NC score | 0.054558 (rank : 52) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9P267, Q9NUV6 | Gene names | MBD5, KIAA1461 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 5 (Methyl-CpG-binding protein MBD5). | |||||
|
TBCD7_MOUSE
|
||||||
| θ value | 0.125558 (rank : 37) | NC score | 0.147914 (rank : 29) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9D0K0, Q3U0V0 | Gene names | Tbc1d7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 7. | |||||
|
NELFB_HUMAN
|
||||||
| θ value | 0.163984 (rank : 38) | NC score | 0.060839 (rank : 47) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8WX92, Q96EW5, Q9H9R4, Q9ULN8, Q9Y3W0 | Gene names | NELFB, COBRA1, KIAA1182 | |||
|
Domain Architecture |

|
|||||
| Description | Negative elongation factor B (NELF-B) (Cofactor of BRCA1). | |||||
|
NELFB_MOUSE
|
||||||
| θ value | 0.163984 (rank : 39) | NC score | 0.061032 (rank : 46) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8C4Y3, Q99J41, Q9JJA5 | Gene names | Nelfb | |||
|
Domain Architecture |

|
|||||
| Description | Negative elongation factor B (NELF-B). | |||||
|
RB15B_MOUSE
|
||||||
| θ value | 0.163984 (rank : 40) | NC score | 0.045725 (rank : 59) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 284 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q6PHZ5, Q8C6G2 | Gene names | Rbm15b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative RNA-binding protein 15B (RNA-binding motif protein 15B). | |||||
|
TR150_HUMAN
|
||||||
| θ value | 0.163984 (rank : 41) | NC score | 0.060093 (rank : 48) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9Y2W1, Q5VTK6 | Gene names | THRAP3, TRAP150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid hormone receptor-associated protein 3 (Thyroid hormone receptor-associated protein complex 150 kDa component) (Trap150). | |||||
|
TR150_MOUSE
|
||||||
| θ value | 0.163984 (rank : 42) | NC score | 0.054471 (rank : 53) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 468 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q569Z6 | Gene names | Thrap3, Trap150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid hormone receptor-associated protein 3 (Thyroid hormone receptor-associated protein complex 150 kDa component) (Trap150). | |||||
|
NFIB_MOUSE
|
||||||
| θ value | 0.279714 (rank : 43) | NC score | 0.022425 (rank : 110) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P97863, P70252, P70253, P70254, Q9R1G4 | Gene names | Nfib | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor 1 B-type (Nuclear factor 1/B) (NF1-B) (NFI-B) (NF-I/B) (CCAAT-box-binding transcription factor) (CTF) (TGGCA-binding protein). | |||||
|
PTN21_HUMAN
|
||||||
| θ value | 0.279714 (rank : 44) | NC score | 0.021612 (rank : 114) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q16825 | Gene names | PTPN21, PTPD1 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 21 (EC 3.1.3.48) (Protein-tyrosine phosphatase D1). | |||||
|
RB15B_HUMAN
|
||||||
| θ value | 0.279714 (rank : 45) | NC score | 0.038920 (rank : 65) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8NDT2, Q6QE19, Q9BV96 | Gene names | RBM15B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative RNA-binding protein 15B (RNA-binding motif protein 15B). | |||||
|
SPR1A_MOUSE
|
||||||
| θ value | 0.365318 (rank : 46) | NC score | 0.029754 (rank : 79) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q62266 | Gene names | Sprr1a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cornifin A (Small proline-rich protein 1A) (SPR1A) (SPR1 A). | |||||
|
DAZP2_HUMAN
|
||||||
| θ value | 0.47712 (rank : 47) | NC score | 0.065875 (rank : 42) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q15038 | Gene names | DAZAP2, KIAA0058 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DAZ-associated protein 2 (Deleted in azoospermia-associated protein 2). | |||||
|
DAZP2_MOUSE
|
||||||
| θ value | 0.47712 (rank : 48) | NC score | 0.065766 (rank : 44) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9DCP9, O88675, Q3UVI4 | Gene names | Dazap2, Prtb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DAZ-associated protein 2 (Deleted in azoospermia-associated protein 2) (Proline-rich protein expressed in brain). | |||||
|
NUCKS_HUMAN
|
||||||
| θ value | 0.47712 (rank : 49) | NC score | 0.088169 (rank : 35) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9H1E3, Q9H1D6, Q9H723 | Gene names | NUCKS1, NUCKS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (P1). | |||||
|
CNOT3_MOUSE
|
||||||
| θ value | 0.62314 (rank : 50) | NC score | 0.038400 (rank : 66) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8K0V4 | Gene names | Cnot3, Not3 | |||
|
Domain Architecture |

|
|||||
| Description | CCR4-NOT transcription complex subunit 3 (CCR4-associated factor 3). | |||||
|
FILA_HUMAN
|
||||||
| θ value | 0.62314 (rank : 51) | NC score | 0.037480 (rank : 67) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 685 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P20930, Q01720, Q5T583 | Gene names | FLG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filaggrin. | |||||
|
TB182_HUMAN
|
||||||
| θ value | 0.62314 (rank : 52) | NC score | 0.040578 (rank : 64) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9C0C2 | Gene names | TNKS1BP1, KIAA1741, TAB182 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 182 kDa tankyrase 1-binding protein. | |||||
|
YLPM1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 53) | NC score | 0.045791 (rank : 58) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P49750, P49752, Q96I64, Q9P1V7 | Gene names | YLPM1, C14orf170, ZAP3 | |||
|
Domain Architecture |

|
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3) (ZAP113). | |||||
|
DC1L2_MOUSE
|
||||||
| θ value | 0.813845 (rank : 54) | NC score | 0.024262 (rank : 104) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6PDL0 | Gene names | Dync1li2, Dncli2, Dnclic2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoplasmic dynein 1 light intermediate chain 2 (Dynein light intermediate chain 2, cytosolic). | |||||
|
LTB1L_HUMAN
|
||||||
| θ value | 0.813845 (rank : 55) | NC score | 0.005684 (rank : 155) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 553 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q14766 | Gene names | LTBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein, isoform 1L precursor (LTBP-1) (Transforming growth factor beta-1-binding protein 1) (TGF-beta1-BP-1). | |||||
|
NUCKS_MOUSE
|
||||||
| θ value | 0.813845 (rank : 56) | NC score | 0.074625 (rank : 39) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q80XU3, Q8BVD8 | Gene names | Nucks1, Nucks | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (JC7). | |||||
|
SRRM1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 57) | NC score | 0.080786 (rank : 37) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 58) | NC score | 0.074252 (rank : 41) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 0.813845 (rank : 59) | NC score | 0.074440 (rank : 40) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
BSN_MOUSE
|
||||||
| θ value | 1.06291 (rank : 60) | NC score | 0.041659 (rank : 63) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
IASPP_MOUSE
|
||||||
| θ value | 1.06291 (rank : 61) | NC score | 0.026138 (rank : 97) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 1024 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q5I1X5 | Gene names | Ppp1r13l, Nkip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RelA-associated inhibitor (Inhibitor of ASPP protein) (Protein iASPP) (PPP1R13B-like protein) (NFkB-interacting protein 1). | |||||
|
MAGE1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 62) | NC score | 0.032480 (rank : 74) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
RFX1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 63) | NC score | 0.018689 (rank : 121) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P48377 | Gene names | Rfx1 | |||
|
Domain Architecture |
|
|||||
| Description | DNA-binding protein RFX1. | |||||
|
SNIP1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 64) | NC score | 0.048955 (rank : 56) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8TAD8, Q96SP9, Q9H9T7 | Gene names | SNIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Smad nuclear-interacting protein 1. | |||||
|
THAP7_MOUSE
|
||||||
| θ value | 1.06291 (rank : 65) | NC score | 0.025717 (rank : 98) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8VCZ3 | Gene names | Thap7 | |||
|
Domain Architecture |
|
|||||
| Description | THAP domain-containing protein 7. | |||||
|
ZBTB5_HUMAN
|
||||||
| θ value | 1.06291 (rank : 66) | NC score | 0.000792 (rank : 162) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 811 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O15062 | Gene names | ZBTB5, KIAA0354 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger and BTB domain-containing protein 5. | |||||
|
HTSF1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 67) | NC score | 0.026929 (rank : 92) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8BGC0, Q1WWK0, Q9CT41, Q9DAU3 | Gene names | Htatsf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIV Tat-specific factor 1 homolog. | |||||
|
IWS1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 68) | NC score | 0.029676 (rank : 80) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
K0280_HUMAN
|
||||||
| θ value | 1.38821 (rank : 69) | NC score | 0.042111 (rank : 61) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q92567, Q86UG2 | Gene names | KIAA0280 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0280. | |||||
|
K0280_MOUSE
|
||||||
| θ value | 1.38821 (rank : 70) | NC score | 0.042111 (rank : 62) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BGZ2, Q3UVC2, Q80U50, Q8BGN7 | Gene names | Kiaa0280 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0280. | |||||
|
K1802_MOUSE
|
||||||
| θ value | 1.38821 (rank : 71) | NC score | 0.025312 (rank : 100) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
NIPBL_HUMAN
|
||||||
| θ value | 1.38821 (rank : 72) | NC score | 0.035035 (rank : 70) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
SMAD9_HUMAN
|
||||||
| θ value | 1.38821 (rank : 73) | NC score | 0.010962 (rank : 140) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O15198, O14989 | Gene names | SMAD9, MADH6, MADH9 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 9 (SMAD 9) (Mothers against DPP homolog 9) (Smad9) (Madh6). | |||||
|
SOX30_MOUSE
|
||||||
| θ value | 1.38821 (rank : 74) | NC score | 0.033649 (rank : 73) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8CGW4 | Gene names | Sox30 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-30. | |||||
|
STAT6_HUMAN
|
||||||
| θ value | 1.38821 (rank : 75) | NC score | 0.021791 (rank : 113) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P42226 | Gene names | STAT6 | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 6 (IL-4 Stat). | |||||
|
WASIP_HUMAN
|
||||||
| θ value | 1.38821 (rank : 76) | NC score | 0.046386 (rank : 57) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O43516, Q15220, Q6MZU9, Q9BU37, Q9UNP1 | Gene names | WASPIP, WIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein) (PRPL-2 protein). | |||||
|
XE7_HUMAN
|
||||||
| θ value | 1.38821 (rank : 77) | NC score | 0.018691 (rank : 120) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 738 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q02040, Q02832 | Gene names | CXYorf3, DXYS155E, XE7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-lymphocyte antigen precursor (B-lymphocyte surface antigen) (721P) (Protein XE7). | |||||
|
CREB5_HUMAN
|
||||||
| θ value | 1.81305 (rank : 78) | NC score | 0.018585 (rank : 122) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q02930, Q05886, Q06246 | Gene names | CREB5, CREBPA | |||
|
Domain Architecture |

|
|||||
| Description | cAMP response element-binding protein 5 (CRE-BPa). | |||||
|
MBB1A_HUMAN
|
||||||
| θ value | 1.81305 (rank : 79) | NC score | 0.034807 (rank : 71) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 665 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9BQG0, Q86VM3, Q9BW49, Q9P0V5, Q9UF99 | Gene names | MYBBP1A, P160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myb-binding protein 1A. | |||||
|
SPR1B_MOUSE
|
||||||
| θ value | 1.81305 (rank : 80) | NC score | 0.026564 (rank : 94) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q62267 | Gene names | Sprr1b | |||
|
Domain Architecture |

|
|||||
| Description | Cornifin B (Small proline-rich protein 1B) (SPR1B) (SPR1 B). | |||||
|
SRRM1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 81) | NC score | 0.087942 (rank : 36) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
ADA19_HUMAN
|
||||||
| θ value | 2.36792 (rank : 82) | NC score | 0.012170 (rank : 138) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9H013, Q9BZL5, Q9UHP2 | Gene names | ADAM19, MLTNB | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 19 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 19) (Meltrin beta) (Metalloprotease and disintegrin dentritic antigen marker) (MADDAM). | |||||
|
CCD39_HUMAN
|
||||||
| θ value | 2.36792 (rank : 83) | NC score | 0.005504 (rank : 156) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 868 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UFE4 | Gene names | CCDC39 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 39. | |||||
|
CCNL2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 84) | NC score | 0.029470 (rank : 82) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9JJA7, Q5XK66, Q60995, Q8C136, Q8CIJ8, Q99L73, Q9QXH5 | Gene names | Ccnl2, Ania6b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-L2 (Cyclin Ania-6b) (Paneth cell-enhanced expression protein) (PCEE). | |||||
|
MARK2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 85) | NC score | -0.000990 (rank : 164) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 1056 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q7KZI7, Q15449, Q15524, Q5XGA3, Q68A18, Q96HB3, Q96RG0 | Gene names | MARK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MARK2 (EC 2.7.11.1) (MAP/microtubule affinity-regulating kinase 2) (ELKL motif kinase) (EMK1) (PAR1 homolog). | |||||
|
PDZ11_MOUSE
|
||||||
| θ value | 2.36792 (rank : 86) | NC score | 0.013976 (rank : 133) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9CZG9 | Gene names | Pdzd11, Pdzk11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 11. | |||||
|
PTN21_MOUSE
|
||||||
| θ value | 2.36792 (rank : 87) | NC score | 0.016858 (rank : 127) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q62136 | Gene names | Ptpn21 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 21 (EC 3.1.3.48) (Protein-tyrosine phosphatase PTP-RL10). | |||||
|
SYCP2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 88) | NC score | 0.024744 (rank : 103) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BX26, O75763, Q5JX11, Q9NTX8, Q9UG27 | Gene names | SYCP2, SCP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptonemal complex protein 2 (SCP-2) (Synaptonemal complex lateral element protein) (hsSCP2). | |||||
|
TBCD7_HUMAN
|
||||||
| θ value | 2.36792 (rank : 89) | NC score | 0.124408 (rank : 30) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9P0N9 | Gene names | TBC1D7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 7. | |||||
|
TSSC4_MOUSE
|
||||||
| θ value | 2.36792 (rank : 90) | NC score | 0.026404 (rank : 95) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9JHE7 | Gene names | Tssc4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein TSSC4. | |||||
|
ADAM8_MOUSE
|
||||||
| θ value | 3.0926 (rank : 91) | NC score | 0.007541 (rank : 152) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 326 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q05910 | Gene names | Adam8, Ms2 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 8 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 8) (Cell surface antigen MS2) (Macrophage cysteine-rich glycoprotein) (CD156 antigen). | |||||
|
DEDD2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 92) | NC score | 0.035291 (rank : 69) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8WXF8, Q8NBR2, Q8NES1, Q8TAA8, Q96D35 | Gene names | DEDD2, FLAME3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA-binding death effector domain-containing protein 2 (FADD-like anti-apoptotic molecule 3) (DED-containing protein FLAME-3). | |||||
|
DTX2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 93) | NC score | 0.013822 (rank : 134) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q86UW9, Q96H69, Q9H890, Q9P200 | Gene names | DTX2, KIAA1528 | |||
|
Domain Architecture |
|
|||||
| Description | Protein deltex-2 (Deltex-2) (Deltex2) (hDTX2). | |||||
|
FANCJ_HUMAN
|
||||||
| θ value | 3.0926 (rank : 94) | NC score | 0.019596 (rank : 117) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BX63, Q8NCI5 | Gene names | BRIP1, BACH1, FANCJ | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fanconi anemia group J protein (EC 3.6.1.-) (ATP-dependent RNA helicase BRIP1) (Protein FACJ) (BRCA1-interacting protein C-terminal helicase 1) (BRCA1-interacting protein 1) (BRCA1-associated C-terminal helicase 1). | |||||
|
FAT2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 95) | NC score | 0.002274 (rank : 158) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9NYQ8, O75091, Q9NSR7 | Gene names | FAT2, CDHF8, KIAA0811, MEGF1 | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin Fat 2 precursor (hFat2) (Multiple epidermal growth factor-like domains 1). | |||||
|
GGN_MOUSE
|
||||||
| θ value | 3.0926 (rank : 96) | NC score | 0.025687 (rank : 99) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q80WJ1, Q5EBP4, Q80WI9, Q80WJ0 | Gene names | Ggn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin. | |||||
|
MEF2D_HUMAN
|
||||||
| θ value | 3.0926 (rank : 97) | NC score | 0.017145 (rank : 126) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q14814, Q14815 | Gene names | MEF2D | |||
|
Domain Architecture |
|
|||||
| Description | Myocyte-specific enhancer factor 2D. | |||||
|
MEF2D_MOUSE
|
||||||
| θ value | 3.0926 (rank : 98) | NC score | 0.016733 (rank : 128) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q63943, Q63944 | Gene names | Mef2d | |||
|
Domain Architecture |
|
|||||
| Description | Myocyte-specific enhancer factor 2D. | |||||
|
MK15_HUMAN
|
||||||
| θ value | 3.0926 (rank : 99) | NC score | -0.001270 (rank : 165) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 1038 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8TD08, Q2TCF9, Q8N362 | Gene names | MAPK15, ERK8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase 15 (EC 2.7.11.24) (Extracellular signal-regulated kinase 8). | |||||
|
NKTR_HUMAN
|
||||||
| θ value | 3.0926 (rank : 100) | NC score | 0.020269 (rank : 116) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 658 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P30414 | Gene names | NKTR | |||
|
Domain Architecture |

|
|||||
| Description | NK-tumor recognition protein (Natural-killer cells cyclophilin-related protein) (NK-TR protein). | |||||
|
RGS3_MOUSE
|
||||||
| θ value | 3.0926 (rank : 101) | NC score | 0.026150 (rank : 96) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
SATB1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 102) | NC score | 0.022966 (rank : 107) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q01826 | Gene names | SATB1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-binding protein SATB1 (Special AT-rich sequence-binding protein 1). | |||||
|
SATB1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 103) | NC score | 0.022084 (rank : 112) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q60611 | Gene names | Satb1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-binding protein SATB1 (Special AT-rich sequence-binding protein 1). | |||||
|
SFPQ_HUMAN
|
||||||
| θ value | 3.0926 (rank : 104) | NC score | 0.026986 (rank : 91) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 986 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P23246, P30808 | Gene names | SFPQ, PSF | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, proline- and glutamine-rich (Polypyrimidine tract- binding protein-associated-splicing factor) (PTB-associated-splicing factor) (PSF) (DNA-binding p52/p100 complex, 100 kDa subunit) (100 kDa DNA-pairing protein) (hPOMp100). | |||||
|
SFPQ_MOUSE
|
||||||
| θ value | 3.0926 (rank : 105) | NC score | 0.022884 (rank : 108) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8VIJ6, Q9ERW2 | Gene names | Sfpq | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, proline- and glutamine-rich (Polypyrimidine tract- binding protein-associated-splicing factor) (PTB-associated-splicing factor) (PSF) (DNA-binding p52/p100 complex, 100 kDa subunit). | |||||
|
TAB3_MOUSE
|
||||||
| θ value | 3.0926 (rank : 106) | NC score | 0.028036 (rank : 87) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q571K4 | Gene names | Map3k7ip3, Kiaa4135, Tab3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase 7-interacting protein 3 (TAK1-binding protein 3). | |||||
|
XYLT1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 107) | NC score | 0.013337 (rank : 136) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q86Y38, Q9H1B6 | Gene names | XYLT1, XT1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Xylosyltransferase 1 (EC 2.4.2.26) (Xylosyltransferase I) (XylT-I) (XT-I) (Peptide O-xylosyltransferase 1). | |||||
|
CCNL1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 108) | NC score | 0.031319 (rank : 75) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q52KE7, Q8BQ75, Q8R5H9, Q922K0, Q9CSZ3, Q9WV44 | Gene names | Ccnl1, Ania6a, Ccn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-L1 (Cyclin-L) (Cyclin Ania-6a). | |||||
|
CCNL2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 109) | NC score | 0.030840 (rank : 77) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q96S94, Q5T2N5, Q5T2N6, Q6IQ12, Q7Z4Z8, Q8N3C9, Q8N3D5, Q8NHE3, Q8TEL0, Q96B00 | Gene names | CCNL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-L2 (Paneth cell-enhanced expression protein). | |||||
|
CF055_MOUSE
|
||||||
| θ value | 4.03905 (rank : 110) | NC score | 0.019359 (rank : 118) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9CR26 | Gene names | ||||
|
Domain Architecture |
|
|||||
| Description | Protein C6orf55 homolog. | |||||
|
EWS_MOUSE
|
||||||
| θ value | 4.03905 (rank : 111) | NC score | 0.028206 (rank : 85) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q61545 | Gene names | Ewsr1, Ews, Ewsh | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein EWS. | |||||
|
FYB_HUMAN
|
||||||
| θ value | 4.03905 (rank : 112) | NC score | 0.026763 (rank : 93) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 379 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O15117, O00359 | Gene names | FYB, SLAP130 | |||
|
Domain Architecture |

|
|||||
| Description | FYN-binding protein (FYN-T-binding protein) (FYB-120/130) (p120/p130) (SLP-76-associated phosphoprotein) (SLAP-130). | |||||
|
K1210_HUMAN
|
||||||
| θ value | 4.03905 (rank : 113) | NC score | 0.027481 (rank : 90) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9ULL0, Q5JPN4 | Gene names | KIAA1210 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1210. | |||||
|
PHC3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 114) | NC score | 0.017501 (rank : 125) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8CHP6, Q3TLV5 | Gene names | Phc3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyhomeotic-like protein 3. | |||||
|
RAI1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 115) | NC score | 0.022831 (rank : 109) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q61818, Q6ZPH7, Q8CEV1 | Gene names | Rai1, Kiaa1820 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 1. | |||||
|
RTN4_MOUSE
|
||||||
| θ value | 4.03905 (rank : 116) | NC score | 0.028423 (rank : 84) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q99P72, Q5DTK9, Q7TNB7, Q80W95, Q8BGK7, Q8BGM9, Q8K290, Q8K3G8, Q9CTE3 | Gene names | Rtn4, Kiaa0886, Nogo | |||
|
Domain Architecture |

|
|||||
| Description | Reticulon-4 (Neurite outgrowth inhibitor) (Nogo protein). | |||||
|
SURF6_MOUSE
|
||||||
| θ value | 4.03905 (rank : 117) | NC score | 0.015507 (rank : 131) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 195 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P70279, Q8BPI4, Q8BRL7, Q8R0Q7 | Gene names | Surf6, Surf-6 | |||
|
Domain Architecture |
|
|||||
| Description | Surfeit locus protein 6. | |||||
|
SYN1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 118) | NC score | 0.021563 (rank : 115) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 363 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O88935, Q62279, Q8QZT8 | Gene names | Syn1, Syn-1 | |||
|
Domain Architecture |
|
|||||
| Description | Synapsin-1 (Synapsin I). | |||||
|
TBC19_HUMAN
|
||||||
| θ value | 4.03905 (rank : 119) | NC score | 0.055206 (rank : 50) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8N5T2 | Gene names | TBC1D19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 19. | |||||
|
ADA19_MOUSE
|
||||||
| θ value | 5.27518 (rank : 120) | NC score | 0.007698 (rank : 151) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O35674 | Gene names | Adam19, Mltnb | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 19 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 19) (Meltrin beta). | |||||
|
CSKI1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 121) | NC score | 0.012457 (rank : 137) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 1031 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q6P9K8, Q6ZPU2, Q8BWU2, Q8BX99, Q9CXH0 | Gene names | Caskin1, Kiaa1306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-1 (CASK-interacting protein 1). | |||||
|
PRR8_HUMAN
|
||||||
| θ value | 5.27518 (rank : 122) | NC score | 0.036639 (rank : 68) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9NSV0 | Gene names | PRR8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 8. | |||||
|
SFR11_HUMAN
|
||||||
| θ value | 5.27518 (rank : 123) | NC score | 0.043111 (rank : 60) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q05519 | Gene names | SFRS11 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor arginine/serine-rich 11 (Arginine-rich 54 kDa nuclear protein) (p54). | |||||
|
USH1C_MOUSE
|
||||||
| θ value | 5.27518 (rank : 124) | NC score | 0.008733 (rank : 149) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 557 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9ES64, Q91XD1, Q9CVG7, Q9ES65 | Gene names | Ush1c | |||
|
Domain Architecture |

|
|||||
| Description | Harmonin (Usher syndrome type-1C protein homolog) (PDZ domain- containing protein). | |||||
|
AFF4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 125) | NC score | 0.023553 (rank : 105) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 611 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9UHB7, Q498B2, Q59FB3, Q6P592, Q8TDR1, Q9P0E4 | Gene names | AFF4, AF5Q31, MCEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4 (ALL1-fused gene from chromosome 5q31) (Major CDK9 elongation factor-associated protein). | |||||
|
CA106_HUMAN
|
||||||
| θ value | 6.88961 (rank : 126) | NC score | 0.014150 (rank : 132) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q3KP66, Q9NV65, Q9NVI0 | Gene names | C1orf106 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C1orf106. | |||||
|
CCNL1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 127) | NC score | 0.028550 (rank : 83) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UK58, Q6NVY9, Q6UWS7, Q8NI48, Q96QT0, Q9NZF3 | Gene names | CCNL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-L1 (Cyclin-L). | |||||
|
CPSF6_HUMAN
|
||||||
| θ value | 6.88961 (rank : 128) | NC score | 0.027735 (rank : 88) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q16630, Q53ES1, Q9BSJ7, Q9BW18 | Gene names | CPSF6, CFIM68 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage and polyadenylation specificity factor 6 (Cleavage and polyadenylation specificity factor 68 kDa subunit) (CPSF 68 kDa subunit) (Pre-mRNA cleavage factor Im 68 kDa subunit) (Protein HPBRII- 4/7). | |||||
|
CPSF6_MOUSE
|
||||||
| θ value | 6.88961 (rank : 129) | NC score | 0.027511 (rank : 89) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6NVF9, Q8BX86, Q8BXI8 | Gene names | Cpsf6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage and polyadenylation specificity factor 6. | |||||
|
IF4B_MOUSE
|
||||||
| θ value | 6.88961 (rank : 130) | NC score | 0.016381 (rank : 129) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BGD9, Q3TD64 | Gene names | Eif4b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4B (eIF-4B). | |||||
|
LCP2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 131) | NC score | 0.033955 (rank : 72) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 531 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q13094 | Gene names | LCP2 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphocyte cytosolic protein 2 (SH2 domain-containing leukocyte protein of 76 kDa) (SLP-76 tyrosine phosphoprotein) (SLP76). | |||||
|
REXO1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 132) | NC score | 0.024971 (rank : 102) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8N1G1, Q9ULT2 | Gene names | REXO1, ELOABP1, KIAA1138, TCEB3BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA exonuclease 1 homolog (EC 3.1.-.-) (Elongin A-binding protein 1) (EloA-BP1) (Transcription elongation factor B polypeptide 3-binding protein 1). | |||||
|
REXO1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 133) | NC score | 0.019148 (rank : 119) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q7TT28, Q3UMP1, Q69ZR0, Q6NSQ6, Q6PI95, Q9DA29 | Gene names | Rexo1, Kiaa1138, Tceb3bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA exonuclease 1 homolog (EC 3.1.-.-) (Transcription elongation factor B polypeptide 3-binding protein 1). | |||||
|
SOX10_HUMAN
|
||||||
| θ value | 6.88961 (rank : 134) | NC score | 0.005782 (rank : 154) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P56693 | Gene names | SOX10 | |||
|
Domain Architecture |
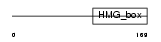
|
|||||
| Description | Transcription factor SOX-10. | |||||
|
SPG20_MOUSE
|
||||||
| θ value | 6.88961 (rank : 135) | NC score | 0.011337 (rank : 139) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8R1X6, Q6ZQ87, Q8BJD3, Q8BM37, Q8BZ63 | Gene names | Spg20, Kiaa0610 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Spartin. | |||||
|
AAKG2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 136) | NC score | 0.008254 (rank : 150) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UGJ0, Q53Y07, Q6NUI0, Q75MP4, Q9NUZ9, Q9UDN8, Q9ULX8 | Gene names | PRKAG2 | |||
|
Domain Architecture |

|
|||||
| Description | 5'-AMP-activated protein kinase subunit gamma-2 (AMPK gamma-2 chain) (AMPK gamma2) (H91620p). | |||||
|
AXN2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 137) | NC score | 0.006516 (rank : 153) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O88566, Q9QXJ6 | Gene names | Axin2 | |||
|
Domain Architecture |

|
|||||
| Description | Axin-2 (Axis inhibition protein 2) (Conductin) (Axin-like protein) (Axil). | |||||
|
CAC1A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 138) | NC score | 0.004090 (rank : 157) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P97445 | Gene names | Cacna1a, Cach4, Cacn3, Cacnl1a4, Ccha1a | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent P/Q-type calcium channel subunit alpha-1A (Voltage- gated calcium channel subunit alpha Cav2.1) (Calcium channel, L type, alpha-1 polypeptide isoform 4) (Brain calcium channel I) (BI). | |||||
|
CAD15_HUMAN
|
||||||
| θ value | 8.99809 (rank : 139) | NC score | 0.000552 (rank : 163) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P55291 | Gene names | CDH15, CDH14, CDH3 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin-15 precursor (Muscle-cadherin) (M-cadherin) (Cadherin-14). | |||||
|
CORO7_MOUSE
|
||||||
| θ value | 8.99809 (rank : 140) | NC score | 0.001800 (rank : 160) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9D2V7, Q3UDJ4, Q6P8Y8, Q8C9V7 | Gene names | Coro7 | |||
|
Domain Architecture |

|
|||||
| Description | Coronin-7 (70 kDa WD repeat tumor rejection antigen homolog). | |||||
|
HXA3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 141) | NC score | 0.001360 (rank : 161) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 558 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P02831, Q4V9Z8, Q61197 | Gene names | Hoxa3, Hox-1.5, Hoxa-3 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-A3 (Hox-1.5) (MO-10). | |||||
|
JARD2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 142) | NC score | 0.009593 (rank : 146) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q92833, Q5U5L5 | Gene names | JARID2, JMJ | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji protein (Jumonji/ARID domain-containing protein 2). | |||||
|
PDZ11_HUMAN
|
||||||
| θ value | 8.99809 (rank : 143) | NC score | 0.009616 (rank : 145) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5EBL8, Q6UWE1, Q9P0Q1 | Gene names | PDZD11, PDZK11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 11. | |||||
|
RL1D1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 144) | NC score | 0.028138 (rank : 86) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O76021, Q6PL22, Q8IWS7, Q8WUZ1, Q9HDA9, Q9Y3Z9 | Gene names | RSL1D1, CATX11, CSIG, PBK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ribosomal L1 domain-containing protein 1 (Cellular senescence- inhibited gene protein) (Protein PBK1) (CATX-11). | |||||
|
RM40_HUMAN
|
||||||
| θ value | 8.99809 (rank : 145) | NC score | 0.016119 (rank : 130) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NQ50, O95134 | Gene names | MRPL40, NLVCF, URIM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 39S ribosomal protein L40, mitochondrial precursor (L40mt) (MRP-40) (Nuclear localization signal-containing protein deleted in velocardiofacial syndrome) (Up-regulated in metastasis). | |||||
|
RPO3D_MOUSE
|
||||||
| θ value | 8.99809 (rank : 146) | NC score | 0.010365 (rank : 141) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q91WD1, Q9CZ02 | Gene names | Polr3d, Bn51t | |||
|
Domain Architecture |

|
|||||
| Description | DNA-directed RNA polymerase III subunit D (EC 2.7.7.6) (DNA-directed RNA polymerase III 47 kDa polypeptide) (RNA polymerase C subunit 4) (RPC4). | |||||
|
RTN4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 147) | NC score | 0.024981 (rank : 101) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NQC3, O94962, Q9BXG5, Q9H212, Q9H3I3, Q9UQ42, Q9Y293, Q9Y2Y7, Q9Y5U6 | Gene names | RTN4, KIAA0886, NOGO | |||
|
Domain Architecture |

|
|||||
| Description | Reticulon-4 (Neurite outgrowth inhibitor) (Nogo protein) (Foocen) (Neuroendocrine-specific protein) (NSP) (Neuroendocrine-specific protein C homolog) (RTN-x) (Reticulon-5). | |||||
|
SFRS4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 148) | NC score | 0.022104 (rank : 111) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q08170, Q5VXP1, Q9BUA4, Q9UEB5 | Gene names | SFRS4, SRP75 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 4 (Pre-mRNA-splicing factor SRP75) (SRP001LB). | |||||
|
SIM2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 149) | NC score | 0.009817 (rank : 143) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q14190, O60766, Q15470, Q15471, Q15472, Q15473, Q16532 | Gene names | SIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2. | |||||
|
TBCD5_HUMAN
|
||||||
| θ value | 8.99809 (rank : 150) | NC score | 0.180778 (rank : 26) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q92609 | Gene names | TBC1D5, KIAA0210 | |||
|
Domain Architecture |
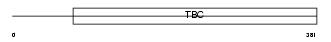
|
|||||
| Description | TBC1 domain family member 5. | |||||
|
THSD1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 151) | NC score | 0.008840 (rank : 148) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NS62, Q6P3U1, Q6UXZ2 | Gene names | THSD1, TMTSP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thrombospondin type-1 domain-containing protein 1 precursor (Transmembrane molecule with thrombospondin module). | |||||
|
TIP60_HUMAN
|
||||||
| θ value | 8.99809 (rank : 152) | NC score | 0.009718 (rank : 144) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q92993, O95624, Q13430, Q6GSE8, Q9BWK7 | Gene names | HTATIP, TIP60 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase HTATIP (EC 2.3.1.48) (EC 2.3.1.-) (60 kDa Tat interactive protein) (Tip60) (HIV-1 Tat interactive protein) (cPLA(2)-interacting protein). | |||||
|
TIP60_MOUSE
|
||||||
| θ value | 8.99809 (rank : 153) | NC score | 0.009884 (rank : 142) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8CHK4, Q8CGZ3, Q8CGZ4, Q8VIH0 | Gene names | Htatip, Tip60 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase HTATIP (EC 2.3.1.48) (EC 2.3.1.-) (60 kDa Tat interactive protein) (Tip60). | |||||
|
U2AFL_HUMAN
|
||||||
| θ value | 8.99809 (rank : 154) | NC score | 0.008946 (rank : 147) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 503 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q15695, Q13570 | Gene names | ZRSR1, U2AF1-RS1, U2AF1L1, U2AF1RS1, U2AFBPL | |||
|
Domain Architecture |
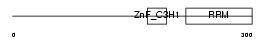
|
|||||
| Description | U2 small nuclear ribonucleoprotein auxiliary factor 35 kDa subunit- related protein 1 (U2(RNU2) small nuclear RNA auxiliary factor 1-like 1) (CCCH type zinc finger, RNA-binding motif and serine/arginine rich protein 1). | |||||
|
WASL_HUMAN
|
||||||
| θ value | 8.99809 (rank : 155) | NC score | 0.023496 (rank : 106) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O00401, Q7Z746 | Gene names | WASL | |||
|
Domain Architecture |
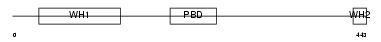
|
|||||
| Description | Neural Wiskott-Aldrich syndrome protein (N-WASP). | |||||
|
WDR47_HUMAN
|
||||||
| θ value | 8.99809 (rank : 156) | NC score | 0.001870 (rank : 159) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 295 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O94967, Q8IXT7, Q8IYU9 | Gene names | WDR47, KIAA0893 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 47. | |||||
|
ZDHC8_HUMAN
|
||||||
| θ value | 8.99809 (rank : 157) | NC score | 0.013816 (rank : 135) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9ULC8, Q6ICL1, Q6ZNF5, Q7Z6L9 | Gene names | ZDHHC8, KIAA1292, ZDHHCL1, ZNF378 | |||
|
Domain Architecture |
|
|||||
| Description | Probable palmitoyltransferase ZDHHC8 (EC 2.3.1.-) (Zinc finger DHHC domain-containing protein 8) (DHHC-8) (Zinc finger protein 378). | |||||
|
TBC13_HUMAN
|
||||||
| θ value | θ > 10 (rank : 158) | NC score | 0.102077 (rank : 31) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9NVG8 | Gene names | TBC1D13 | |||
|
Domain Architecture |
|
|||||
| Description | TBC1 domain family member 13. | |||||
|
TBC13_MOUSE
|
||||||
| θ value | θ > 10 (rank : 159) | NC score | 0.253226 (rank : 25) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8R3D1 | Gene names | Tbc1d13 | |||
|
Domain Architecture |
|
|||||
| Description | TBC1 domain family member 13. | |||||
|
TBC20_HUMAN
|
||||||
| θ value | θ > 10 (rank : 160) | NC score | 0.054884 (rank : 51) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96BZ9, Q5JWQ7, Q96NE1, Q9BYM7, Q9H140 | Gene names | TBC1D20, C20orf140 | |||
|
Domain Architecture |
|
|||||
| Description | TBC1 domain family member 20. | |||||
|
TBC20_MOUSE
|
||||||
| θ value | θ > 10 (rank : 161) | NC score | 0.052382 (rank : 55) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9D9I4, Q3TYW9, Q99LW2 | Gene names | Tbc1d20 | |||
|
Domain Architecture |
|
|||||
| Description | TBC1 domain family member 20. | |||||
|
TBC21_HUMAN
|
||||||
| θ value | θ > 10 (rank : 162) | NC score | 0.173354 (rank : 27) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8IYX1 | Gene names | TBC1D21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 21. | |||||
|
TBCD5_MOUSE
|
||||||
| θ value | θ > 10 (rank : 163) | NC score | 0.162470 (rank : 28) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q80XQ2 | Gene names | Tbc1d5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 5. | |||||
|
UBP32_HUMAN
|
||||||
| θ value | θ > 10 (rank : 164) | NC score | 0.092624 (rank : 34) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NFA0, Q9BX85, Q9Y591 | Gene names | USP32, USP10 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 32 (EC 3.1.2.15) (Ubiquitin thioesterase 32) (Ubiquitin-specific-processing protease 32) (Deubiquitinating enzyme 32) (NY-REN-60 antigen). | |||||
|
VCX2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 165) | NC score | 0.076085 (rank : 38) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H322, Q9P0H5 | Gene names | VCX2, VCX2R, VCXB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Variable charge X-linked protein 2 (VCX-B protein) (Variably charged protein X-B) (Variable charge protein on X with two repeats) (VCX-2r). | |||||
|
US6NL_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 157 | |
| SwissProt Accessions | Q80XC3, Q6ZQK8 | Gene names | Usp6nl, Kiaa0019 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USP6 N-terminal-like protein. | |||||
|
US6NL_HUMAN
|
||||||
| NC score | 0.958673 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q92738, Q15400, Q7L0K9 | Gene names | USP6NL, KIAA0019 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USP6 N-terminal-like protein (Related to the N-terminus of tre) (RN- tre). | |||||
|
TBCD3_HUMAN
|
||||||
| NC score | 0.870449 (rank : 3) | θ value | 3.03575e-64 (rank : 4) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8IZP1, Q9H0B9, Q9UDD4 | Gene names | TBC1D3, PRC17 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 3 (Rab GTPase-activating protein PRC17) (Prostate cancer gene 17 protein) (TRE17 alpha protein). | |||||
|
TBC10_HUMAN
|
||||||
| NC score | 0.809232 (rank : 4) | θ value | 8.3442e-30 (rank : 6) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9BXI6, O76053, Q543A2 | Gene names | TBC1D10A, EPI64, TBC1D10 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 10A (EBP50-PDX interactor of 64 kDa) (EPI64 protein). | |||||
|
TBC10_MOUSE
|
||||||
| NC score | 0.805776 (rank : 5) | θ value | 9.8567e-31 (rank : 5) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P58802 | Gene names | Tbc1d10a, Epi64, Tbc1d10 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 10A (EBP50-PDX interactor of 64 kDa) (EPI64 protein). | |||||
|
TBC12_HUMAN
|
||||||
| NC score | 0.736634 (rank : 6) | θ value | 1.80048e-24 (rank : 11) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O60347, Q5VYA6, Q8WX26, Q8WX59, Q9UG83 | Gene names | TBC1D12, KIAA0608 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 12. | |||||
|
TBC14_HUMAN
|
||||||
| NC score | 0.707853 (rank : 7) | θ value | 5.79196e-23 (rank : 12) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9P2M4, Q8IW15 | Gene names | TBC1D14, KIAA1322 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 14. | |||||
|
TBC14_MOUSE
|
||||||
| NC score | 0.703514 (rank : 8) | θ value | 6.40375e-22 (rank : 13) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8CGA2, Q8CHA5 | Gene names | Tbc1d14, D5Ertd110e, Kiaa1322 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 14. | |||||
|
TBCD1_HUMAN
|
||||||
| NC score | 0.699563 (rank : 9) | θ value | 1.33217e-27 (rank : 7) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q86TI0, Q96K82, Q9UPP4 | Gene names | TBC1D1, KIAA1108 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 1. | |||||
|
TBCD1_MOUSE
|
||||||
| NC score | 0.693071 (rank : 10) | θ value | 1.33217e-27 (rank : 8) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q60949, Q80TJ9, Q923F8 | Gene names | Tbc1d1, Kiaa1108, Tbc1 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 1. | |||||
|
TBCD4_MOUSE
|
||||||
| NC score | 0.668202 (rank : 11) | θ value | 7.30988e-26 (rank : 9) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8BYJ6, Q5DU23, Q66JU2, Q6P2M2, Q8BMH6, Q8BXM2 | Gene names | Tbc1d4, Kiaa0603 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 4 (Akt substrate of 160 kDa) (AS160). | |||||
|
TBCD8_MOUSE
|
||||||
| NC score | 0.658989 (rank : 12) | θ value | 6.62687e-19 (rank : 16) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9Z1A9 | Gene names | Tbc1d8, Hblp1, Vrp | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 8 (Vascular Rab-GAP/TBC-containing protein) (BUB2-like protein 1). | |||||
|
TBCD4_HUMAN
|
||||||
| NC score | 0.658094 (rank : 13) | θ value | 1.37858e-24 (rank : 10) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O60343, Q5W0B9 | Gene names | TBC1D4, KIAA0603 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 4 (Akt substrate of 160 kDa) (AS160). | |||||
|
TBCD8_HUMAN
|
||||||
| NC score | 0.654078 (rank : 14) | θ value | 1.33526e-19 (rank : 15) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O95759, Q9UQ32 | Gene names | TBC1D8, VRP | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 8 (Vascular Rab-GAP/TBC-containing protein) (AD 3). | |||||
|
TBCD2_HUMAN
|
||||||
| NC score | 0.642058 (rank : 15) | θ value | 7.82807e-20 (rank : 14) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9BYX2 | Gene names | TBC1D2, PARIS1 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 2 (Prostate antigen recognized and identified by SEREX) (PARIS-1). | |||||
|
TBC16_HUMAN
|
||||||
| NC score | 0.607441 (rank : 16) | θ value | 1.68911e-14 (rank : 17) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8TBP0 | Gene names | TBC1D16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 16. | |||||
|
UBP6_HUMAN
|
||||||
| NC score | 0.520774 (rank : 17) | θ value | 2.09745e-73 (rank : 3) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P35125, Q15634, Q86WP6, Q8IWT4 | Gene names | USP6, TRE2 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 6 (EC 3.1.2.15) (Ubiquitin thioesterase 6) (Ubiquitin-specific-processing protease 6) (Deubiquitinating enzyme 6) (Proto-oncogene TRE-2). | |||||
|
TBC15_MOUSE
|
||||||
| NC score | 0.475562 (rank : 18) | θ value | 1.9326e-10 (rank : 20) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9CXF4, Q3UI41 | Gene names | Tbc1d15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 15. | |||||
|
TBC17_MOUSE
|
||||||
| NC score | 0.474905 (rank : 19) | θ value | 6.64225e-11 (rank : 18) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BYH7 | Gene names | Tbc1d17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 17. | |||||
|
TBC15_HUMAN
|
||||||
| NC score | 0.471316 (rank : 20) | θ value | 1.47974e-10 (rank : 19) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8TC07 | Gene names | TBC1D15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 15. | |||||
|
TBC17_HUMAN
|
||||||
| NC score | 0.463924 (rank : 21) | θ value | 1.25267e-09 (rank : 21) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9HA65 | Gene names | TBC1D17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 17. | |||||
|
TB22B_HUMAN
|
||||||
| NC score | 0.366128 (rank : 22) | θ value | 0.000158464 (rank : 24) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NU19, Q5VUK9, Q7Z6P7, Q9BPV6, Q9BUT5, Q9NXB6 | Gene names | TBC1D22B, C6orf197 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 22B. | |||||
|
TB22A_MOUSE
|
||||||
| NC score | 0.364189 (rank : 23) | θ value | 7.59969e-07 (rank : 23) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8R5A6, Q3U268, Q3U3T0, Q8CA49 | Gene names | Tbc1d22a, D15Ertd781e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 22A. | |||||
|
TB22A_HUMAN
|
||||||
| NC score | 0.361663 (rank : 24) | θ value | 3.41135e-07 (rank : 22) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8WUA7, Q5TE47, Q6ZUH2, Q92680, Q9BVD6, Q9UGG0, Q9UGT2, Q9UGU6, Q9UH25, Q9Y4W5 | Gene names | TBC1D22A, C22orf4 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 22A. | |||||
|
TBC13_MOUSE
|
||||||
| NC score | 0.253226 (rank : 25) | θ value | θ > 10 (rank : 159) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8R3D1 | Gene names | Tbc1d13 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 13. | |||||
|
TBCD5_HUMAN
|
||||||
| NC score | 0.180778 (rank : 26) | θ value | 8.99809 (rank : 150) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q92609 | Gene names | TBC1D5, KIAA0210 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 5. | |||||
|
TBC21_HUMAN
|
||||||
| NC score | 0.173354 (rank : 27) | θ value | θ > 10 (rank : 162) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8IYX1 | Gene names | TBC1D21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 21. | |||||
|
TBCD5_MOUSE
|
||||||
| NC score | 0.162470 (rank : 28) | θ value | θ > 10 (rank : 163) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q80XQ2 | Gene names | Tbc1d5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 5. | |||||
|
TBCD7_MOUSE
|
||||||
| NC score | 0.147914 (rank : 29) | θ value | 0.125558 (rank : 37) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9D0K0, Q3U0V0 | Gene names | Tbc1d7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 7. | |||||
|
TBCD7_HUMAN
|
||||||
| NC score | 0.124408 (rank : 30) | θ value | 2.36792 (rank : 89) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9P0N9 | Gene names | TBC1D7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 7. | |||||
|
TBC13_HUMAN
|
||||||
| NC score | 0.102077 (rank : 31) | θ value | θ > 10 (rank : 158) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9NVG8 | Gene names | TBC1D13 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 13. | |||||
|
RPB1_MOUSE
|
||||||
| NC score | 0.099401 (rank : 32) | θ value | 0.00228821 (rank : 28) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 363 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P08775 | Gene names | Polr2a, Rpii215, Rpo2-1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-directed RNA polymerase II largest subunit (EC 2.7.7.6) (RPB1). | |||||
|
RPB1_HUMAN
|
||||||
| NC score | 0.098576 (rank : 33) | θ value | 0.00228821 (rank : 27) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P24928 | Gene names | POLR2A | |||
|
Domain Architecture |

|
|||||
| Description | DNA-directed RNA polymerase II largest subunit (EC 2.7.7.6) (RPB1). | |||||
|
UBP32_HUMAN
|
||||||
| NC score | 0.092624 (rank : 34) | θ value | θ > 10 (rank : 164) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NFA0, Q9BX85, Q9Y591 | Gene names | USP32, USP10 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 32 (EC 3.1.2.15) (Ubiquitin thioesterase 32) (Ubiquitin-specific-processing protease 32) (Deubiquitinating enzyme 32) (NY-REN-60 antigen). | |||||
|
NUCKS_HUMAN
|
||||||
| NC score | 0.088169 (rank : 35) | θ value | 0.47712 (rank : 49) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9H1E3, Q9H1D6, Q9H723 | Gene names | NUCKS1, NUCKS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (P1). | |||||
|
SRRM1_HUMAN
|
||||||
| NC score | 0.087942 (rank : 36) | θ value | 1.81305 (rank : 81) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
SRRM1_MOUSE
|
||||||
| NC score | 0.080786 (rank : 37) | θ value | 0.813845 (rank : 57) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
VCX2_HUMAN
|
||||||
| NC score | 0.076085 (rank : 38) | θ value | θ > 10 (rank : 165) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H322, Q9P0H5 | Gene names | VCX2, VCX2R, VCXB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Variable charge X-linked protein 2 (VCX-B protein) (Variably charged protein X-B) (Variable charge protein on X with two repeats) (VCX-2r). | |||||
|
NUCKS_MOUSE
|
||||||
| NC score | 0.074625 (rank : 39) | θ value | 0.813845 (rank : 56) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q80XU3, Q8BVD8 | Gene names | Nucks1, Nucks | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (JC7). | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.074440 (rank : 40) | θ value | 0.813845 (rank : 59) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.074252 (rank : 41) | θ value | 0.813845 (rank : 58) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
DAZP2_HUMAN
|
||||||
| NC score | 0.065875 (rank : 42) | θ value | 0.47712 (rank : 47) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q15038 | Gene names | DAZAP2, KIAA0058 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DAZ-associated protein 2 (Deleted in azoospermia-associated protein 2). | |||||
|
SSXT_MOUSE
|
||||||
| NC score | 0.065779 (rank : 43) | θ value | 0.0252991 (rank : 29) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q62280 | Gene names | Ss18, Ssxt, Syt | |||
|
Domain Architecture |

|
|||||
| Description | SSXT protein (SYT protein) (Synovial sarcoma-associated Ss18-alpha). | |||||
|
DAZP2_MOUSE
|
||||||
| NC score | 0.065766 (rank : 44) | θ value | 0.47712 (rank : 48) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9DCP9, O88675, Q3UVI4 | Gene names | Dazap2, Prtb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DAZ-associated protein 2 (Deleted in azoospermia-associated protein 2) (Proline-rich protein expressed in brain). | |||||
|
SSXT_HUMAN
|
||||||
| NC score | 0.065337 (rank : 45) | θ value | 0.0330416 (rank : 31) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q15532, Q16404, Q9BXC6 | Gene names | SS18, SSXT, SYT | |||
|
Domain Architecture |

|
|||||
| Description | SSXT protein (Synovial sarcoma, translocated to X chromosome) (SYT protein). | |||||
|
NELFB_MOUSE
|
||||||
| NC score | 0.061032 (rank : 46) | θ value | 0.163984 (rank : 39) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8C4Y3, Q99J41, Q9JJA5 | Gene names | Nelfb | |||
|
Domain Architecture |

|
|||||
| Description | Negative elongation factor B (NELF-B). | |||||
|
NELFB_HUMAN
|
||||||
| NC score | 0.060839 (rank : 47) | θ value | 0.163984 (rank : 38) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8WX92, Q96EW5, Q9H9R4, Q9ULN8, Q9Y3W0 | Gene names | NELFB, COBRA1, KIAA1182 | |||
|
Domain Architecture |

|
|||||
| Description | Negative elongation factor B (NELF-B) (Cofactor of BRCA1). | |||||
|
TR150_HUMAN
|
||||||
| NC score | 0.060093 (rank : 48) | θ value | 0.163984 (rank : 41) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9Y2W1, Q5VTK6 | Gene names | THRAP3, TRAP150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid hormone receptor-associated protein 3 (Thyroid hormone receptor-associated protein complex 150 kDa component) (Trap150). | |||||
|
TARA_HUMAN
|
||||||
| NC score | 0.059624 (rank : 49) | θ value | 0.000602161 (rank : 25) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9H2D6, O94797, Q2PZW8, Q2Q3Z9, Q2Q400, Q5R3M6, Q96DW1, Q9BT77, Q9BTL7, Q9BY98, Q9Y3L4 | Gene names | TRIOBP, KIAA1662, TARA | |||
|
Domain Architecture |

|
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
TBC19_HUMAN
|
||||||
| NC score | 0.055206 (rank : 50) | θ value | 4.03905 (rank : 119) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8N5T2 | Gene names | TBC1D19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 19. | |||||
|
TBC20_HUMAN
|
||||||
| NC score | 0.054884 (rank : 51) | θ value | θ > 10 (rank : 160) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96BZ9, Q5JWQ7, Q96NE1, Q9BYM7, Q9H140 | Gene names | TBC1D20, C20orf140 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 20. | |||||
|
MBD5_HUMAN
|
||||||
| NC score | 0.054558 (rank : 52) | θ value | 0.125558 (rank : 36) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9P267, Q9NUV6 | Gene names | MBD5, KIAA1461 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 5 (Methyl-CpG-binding protein MBD5). | |||||
|
TR150_MOUSE
|
||||||
| NC score | 0.054471 (rank : 53) | θ value | 0.163984 (rank : 42) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 468 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q569Z6 | Gene names | Thrap3, Trap150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid hormone receptor-associated protein 3 (Thyroid hormone receptor-associated protein complex 150 kDa component) (Trap150). | |||||
|
ATN1_HUMAN
|
||||||
| NC score | 0.053728 (rank : 54) | θ value | 0.0563607 (rank : 32) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
TBC20_MOUSE
|
||||||
| NC score | 0.052382 (rank : 55) | θ value | θ > 10 (rank : 161) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9D9I4, Q3TYW9, Q99LW2 | Gene names | Tbc1d20 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 20. | |||||
|
SNIP1_HUMAN
|
||||||
| NC score | 0.048955 (rank : 56) | θ value | 1.06291 (rank : 64) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8TAD8, Q96SP9, Q9H9T7 | Gene names | SNIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Smad nuclear-interacting protein 1. | |||||
|
WASIP_HUMAN
|
||||||
| NC score | 0.046386 (rank : 57) | θ value | 1.38821 (rank : 76) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O43516, Q15220, Q6MZU9, Q9BU37, Q9UNP1 | Gene names | WASPIP, WIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein) (PRPL-2 protein). | |||||
|
YLPM1_HUMAN
|
||||||
| NC score | 0.045791 (rank : 58) | θ value | 0.62314 (rank : 53) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P49750, P49752, Q96I64, Q9P1V7 | Gene names | YLPM1, C14orf170, ZAP3 | |||
|
Domain Architecture |

|
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3) (ZAP113). | |||||
|
RB15B_MOUSE
|
||||||
| NC score | 0.045725 (rank : 59) | θ value | 0.163984 (rank : 40) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 284 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q6PHZ5, Q8C6G2 | Gene names | Rbm15b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative RNA-binding protein 15B (RNA-binding motif protein 15B). | |||||
|
SFR11_HUMAN
|
||||||
| NC score | 0.043111 (rank : 60) | θ value | 5.27518 (rank : 123) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q05519 | Gene names | SFRS11 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor arginine/serine-rich 11 (Arginine-rich 54 kDa nuclear protein) (p54). | |||||
|
K0280_HUMAN
|
||||||
| NC score | 0.042111 (rank : 61) | θ value | 1.38821 (rank : 69) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q92567, Q86UG2 | Gene names | KIAA0280 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0280. | |||||
|
K0280_MOUSE
|
||||||
| NC score | 0.042111 (rank : 62) | θ value | 1.38821 (rank : 70) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BGZ2, Q3UVC2, Q80U50, Q8BGN7 | Gene names | Kiaa0280 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0280. | |||||
|
BSN_MOUSE
|
||||||
| NC score | 0.041659 (rank : 63) | θ value | 1.06291 (rank : 60) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
TB182_HUMAN
|
||||||
| NC score | 0.040578 (rank : 64) | θ value | 0.62314 (rank : 52) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9C0C2 | Gene names | TNKS1BP1, KIAA1741, TAB182 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 182 kDa tankyrase 1-binding protein. | |||||
|
RB15B_HUMAN
|
||||||
| NC score | 0.038920 (rank : 65) | θ value | 0.279714 (rank : 45) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8NDT2, Q6QE19, Q9BV96 | Gene names | RBM15B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative RNA-binding protein 15B (RNA-binding motif protein 15B). | |||||
|
CNOT3_MOUSE
|
||||||
| NC score | 0.038400 (rank : 66) | θ value | 0.62314 (rank : 50) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8K0V4 | Gene names | Cnot3, Not3 | |||
|
Domain Architecture |

|
|||||
| Description | CCR4-NOT transcription complex subunit 3 (CCR4-associated factor 3). | |||||
|
FILA_HUMAN
|
||||||
| NC score | 0.037480 (rank : 67) | θ value | 0.62314 (rank : 51) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 685 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P20930, Q01720, Q5T583 | Gene names | FLG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filaggrin. | |||||
|
PRR8_HUMAN
|
||||||
| NC score | 0.036639 (rank : 68) | θ value | 5.27518 (rank : 122) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9NSV0 | Gene names | PRR8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 8. | |||||
|
DEDD2_HUMAN
|
||||||
| NC score | 0.035291 (rank : 69) | θ value | 3.0926 (rank : 92) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8WXF8, Q8NBR2, Q8NES1, Q8TAA8, Q96D35 | Gene names | DEDD2, FLAME3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA-binding death effector domain-containing protein 2 (FADD-like anti-apoptotic molecule 3) (DED-containing protein FLAME-3). | |||||
|
NIPBL_HUMAN
|
||||||
| NC score | 0.035035 (rank : 70) | θ value | 1.38821 (rank : 72) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
MBB1A_HUMAN
|
||||||
| NC score | 0.034807 (rank : 71) | θ value | 1.81305 (rank : 79) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 665 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9BQG0, Q86VM3, Q9BW49, Q9P0V5, Q9UF99 | Gene names | MYBBP1A, P160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myb-binding protein 1A. | |||||
|
LCP2_HUMAN
|
||||||
| NC score | 0.033955 (rank : 72) | θ value | 6.88961 (rank : 131) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 531 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q13094 | Gene names | LCP2 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphocyte cytosolic protein 2 (SH2 domain-containing leukocyte protein of 76 kDa) (SLP-76 tyrosine phosphoprotein) (SLP76). | |||||
|
SOX30_MOUSE
|
||||||
| NC score | 0.033649 (rank : 73) | θ value | 1.38821 (rank : 74) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8CGW4 | Gene names | Sox30 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-30. | |||||
|
MAGE1_HUMAN
|
||||||
| NC score | 0.032480 (rank : 74) | θ value | 1.06291 (rank : 62) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
CCNL1_MOUSE
|
||||||
| NC score | 0.031319 (rank : 75) | θ value | 4.03905 (rank : 108) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q52KE7, Q8BQ75, Q8R5H9, Q922K0, Q9CSZ3, Q9WV44 | Gene names | Ccnl1, Ania6a, Ccn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-L1 (Cyclin-L) (Cyclin Ania-6a). | |||||
|
E41L1_MOUSE
|
||||||
| NC score | 0.031109 (rank : 76) | θ value | 0.0563607 (rank : 33) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Z2H5 | Gene names | Epb41l1, Epb4, Epb4.1l1 | |||
|
Domain Architecture |

|
|||||
| Description | Band 4.1-like protein 1 (Neuronal protein 4.1) (4.1N). | |||||
|
CCNL2_HUMAN
|
||||||
| NC score | 0.030840 (rank : 77) | θ value | 4.03905 (rank : 109) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q96S94, Q5T2N5, Q5T2N6, Q6IQ12, Q7Z4Z8, Q8N3C9, Q8N3D5, Q8NHE3, Q8TEL0, Q96B00 | Gene names | CCNL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-L2 (Paneth cell-enhanced expression protein). | |||||
|
E41L1_HUMAN
|
||||||
| NC score | 0.030271 (rank : 78) | θ value | 0.0961366 (rank : 35) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9H4G0, O15046, Q4VXN4 | Gene names | EPB41L1, KIAA0338 | |||
|
Domain Architecture |

|
|||||
| Description | Band 4.1-like protein 1 (Neuronal protein 4.1) (4.1N). | |||||
|
SPR1A_MOUSE
|
||||||
| NC score | 0.029754 (rank : 79) | θ value | 0.365318 (rank : 46) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q62266 | Gene names | Sprr1a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cornifin A (Small proline-rich protein 1A) (SPR1A) (SPR1 A). | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.029676 (rank : 80) | θ value | 1.38821 (rank : 68) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
ANK2_HUMAN
|
||||||
| NC score | 0.029511 (rank : 81) | θ value | 0.00228821 (rank : 26) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
CCNL2_MOUSE
|
||||||
| NC score | 0.029470 (rank : 82) | θ value | 2.36792 (rank : 84) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9JJA7, Q5XK66, Q60995, Q8C136, Q8CIJ8, Q99L73, Q9QXH5 | Gene names | Ccnl2, Ania6b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-L2 (Cyclin Ania-6b) (Paneth cell-enhanced expression protein) (PCEE). | |||||
|
CCNL1_HUMAN
|
||||||
| NC score | 0.028550 (rank : 83) | θ value | 6.88961 (rank : 127) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UK58, Q6NVY9, Q6UWS7, Q8NI48, Q96QT0, Q9NZF3 | Gene names | CCNL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-L1 (Cyclin-L). | |||||
|
RTN4_MOUSE
|
||||||
| NC score | 0.028423 (rank : 84) | θ value | 4.03905 (rank : 116) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q99P72, Q5DTK9, Q7TNB7, Q80W95, Q8BGK7, Q8BGM9, Q8K290, Q8K3G8, Q9CTE3 | Gene names | Rtn4, Kiaa0886, Nogo | |||
|
Domain Architecture |

|
|||||
| Description | Reticulon-4 (Neurite outgrowth inhibitor) (Nogo protein). | |||||
|
EWS_MOUSE
|
||||||
| NC score | 0.028206 (rank : 85) | θ value | 4.03905 (rank : 111) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q61545 | Gene names | Ewsr1, Ews, Ewsh | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein EWS. | |||||
|
RL1D1_HUMAN
|
||||||
| NC score | 0.028138 (rank : 86) | θ value | 8.99809 (rank : 144) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O76021, Q6PL22, Q8IWS7, Q8WUZ1, Q9HDA9, Q9Y3Z9 | Gene names | RSL1D1, CATX11, CSIG, PBK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ribosomal L1 domain-containing protein 1 (Cellular senescence- inhibited gene protein) (Protein PBK1) (CATX-11). | |||||
|
TAB3_MOUSE
|
||||||
| NC score | 0.028036 (rank : 87) | θ value | 3.0926 (rank : 106) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q571K4 | Gene names | Map3k7ip3, Kiaa4135, Tab3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase 7-interacting protein 3 (TAK1-binding protein 3). | |||||
|
CPSF6_HUMAN
|
||||||
| NC score | 0.027735 (rank : 88) | θ value | 6.88961 (rank : 128) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q16630, Q53ES1, Q9BSJ7, Q9BW18 | Gene names | CPSF6, CFIM68 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage and polyadenylation specificity factor 6 (Cleavage and polyadenylation specificity factor 68 kDa subunit) (CPSF 68 kDa subunit) (Pre-mRNA cleavage factor Im 68 kDa subunit) (Protein HPBRII- 4/7). | |||||
|
CPSF6_MOUSE
|
||||||
| NC score | 0.027511 (rank : 89) | θ value | 6.88961 (rank : 129) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6NVF9, Q8BX86, Q8BXI8 | Gene names | Cpsf6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage and polyadenylation specificity factor 6. | |||||
|
K1210_HUMAN
|
||||||
| NC score | 0.027481 (rank : 90) | θ value | 4.03905 (rank : 113) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9ULL0, Q5JPN4 | Gene names | KIAA1210 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1210. | |||||
|
SFPQ_HUMAN
|
||||||
| NC score | 0.026986 (rank : 91) | θ value | 3.0926 (rank : 104) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 986 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P23246, P30808 | Gene names | SFPQ, PSF | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, proline- and glutamine-rich (Polypyrimidine tract- binding protein-associated-splicing factor) (PTB-associated-splicing factor) (PSF) (DNA-binding p52/p100 complex, 100 kDa subunit) (100 kDa DNA-pairing protein) (hPOMp100). | |||||
|
HTSF1_MOUSE
|
||||||
| NC score | 0.026929 (rank : 92) | θ value | 1.38821 (rank : 67) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8BGC0, Q1WWK0, Q9CT41, Q9DAU3 | Gene names | Htatsf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIV Tat-specific factor 1 homolog. | |||||
|
FYB_HUMAN
|
||||||
| NC score | 0.026763 (rank : 93) | θ value | 4.03905 (rank : 112) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 379 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O15117, O00359 | Gene names | FYB, SLAP130 | |||
|
Domain Architecture |

|
|||||
| Description | FYN-binding protein (FYN-T-binding protein) (FYB-120/130) (p120/p130) (SLP-76-associated phosphoprotein) (SLAP-130). | |||||
|
SPR1B_MOUSE
|
||||||
| NC score | 0.026564 (rank : 94) | θ value | 1.81305 (rank : 80) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q62267 | Gene names | Sprr1b | |||
|
Domain Architecture |

|
|||||
| Description | Cornifin B (Small proline-rich protein 1B) (SPR1B) (SPR1 B). | |||||
|
TSSC4_MOUSE
|
||||||
| NC score | 0.026404 (rank : 95) | θ value | 2.36792 (rank : 90) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9JHE7 | Gene names | Tssc4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein TSSC4. | |||||
|
RGS3_MOUSE
|
||||||
| NC score | 0.026150 (rank : 96) | θ value | 3.0926 (rank : 101) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
IASPP_MOUSE
|
||||||
| NC score | 0.026138 (rank : 97) | θ value | 1.06291 (rank : 61) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 1024 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q5I1X5 | Gene names | Ppp1r13l, Nkip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RelA-associated inhibitor (Inhibitor of ASPP protein) (Protein iASPP) (PPP1R13B-like protein) (NFkB-interacting protein 1). | |||||
|
THAP7_MOUSE
|
||||||
| NC score | 0.025717 (rank : 98) | θ value | 1.06291 (rank : 65) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8VCZ3 | Gene names | Thap7 | |||
|
Domain Architecture |

|
|||||
| Description | THAP domain-containing protein 7. | |||||
|
GGN_MOUSE
|
||||||
| NC score | 0.025687 (rank : 99) | θ value | 3.0926 (rank : 96) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q80WJ1, Q5EBP4, Q80WI9, Q80WJ0 | Gene names | Ggn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin. | |||||
|
K1802_MOUSE
|
||||||
| NC score | 0.025312 (rank : 100) | θ value | 1.38821 (rank : 71) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
RTN4_HUMAN
|
||||||
| NC score | 0.024981 (rank : 101) | θ value | 8.99809 (rank : 147) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NQC3, O94962, Q9BXG5, Q9H212, Q9H3I3, Q9UQ42, Q9Y293, Q9Y2Y7, Q9Y5U6 | Gene names | RTN4, KIAA0886, NOGO | |||
|
Domain Architecture |

|
|||||
| Description | Reticulon-4 (Neurite outgrowth inhibitor) (Nogo protein) (Foocen) (Neuroendocrine-specific protein) (NSP) (Neuroendocrine-specific protein C homolog) (RTN-x) (Reticulon-5). | |||||
|
REXO1_HUMAN
|
||||||
| NC score | 0.024971 (rank : 102) | θ value | 6.88961 (rank : 132) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8N1G1, Q9ULT2 | Gene names | REXO1, ELOABP1, KIAA1138, TCEB3BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA exonuclease 1 homolog (EC 3.1.-.-) (Elongin A-binding protein 1) (EloA-BP1) (Transcription elongation factor B polypeptide 3-binding protein 1). | |||||
|
SYCP2_HUMAN
|
||||||
| NC score | 0.024744 (rank : 103) | θ value | 2.36792 (rank : 88) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BX26, O75763, Q5JX11, Q9NTX8, Q9UG27 | Gene names | SYCP2, SCP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptonemal complex protein 2 (SCP-2) (Synaptonemal complex lateral element protein) (hsSCP2). | |||||
|
DC1L2_MOUSE
|
||||||
| NC score | 0.024262 (rank : 104) | θ value | 0.813845 (rank : 54) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6PDL0 | Gene names | Dync1li2, Dncli2, Dnclic2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoplasmic dynein 1 light intermediate chain 2 (Dynein light intermediate chain 2, cytosolic). | |||||
|
AFF4_HUMAN
|
||||||
| NC score | 0.023553 (rank : 105) | θ value | 6.88961 (rank : 125) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 611 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9UHB7, Q498B2, Q59FB3, Q6P592, Q8TDR1, Q9P0E4 | Gene names | AFF4, AF5Q31, MCEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4 (ALL1-fused gene from chromosome 5q31) (Major CDK9 elongation factor-associated protein). | |||||
|
WASL_HUMAN
|
||||||
| NC score | 0.023496 (rank : 106) | θ value | 8.99809 (rank : 155) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O00401, Q7Z746 | Gene names | WASL | |||
|
Domain Architecture |

|
|||||
| Description | Neural Wiskott-Aldrich syndrome protein (N-WASP). | |||||
|
SATB1_HUMAN
|
||||||
| NC score | 0.022966 (rank : 107) | θ value | 3.0926 (rank : 102) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q01826 | Gene names | SATB1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-binding protein SATB1 (Special AT-rich sequence-binding protein 1). | |||||
|
SFPQ_MOUSE
|
||||||
| NC score | 0.022884 (rank : 108) | θ value | 3.0926 (rank : 105) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8VIJ6, Q9ERW2 | Gene names | Sfpq | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, proline- and glutamine-rich (Polypyrimidine tract- binding protein-associated-splicing factor) (PTB-associated-splicing factor) (PSF) (DNA-binding p52/p100 complex, 100 kDa subunit). | |||||
|
RAI1_MOUSE
|
||||||
| NC score | 0.022831 (rank : 109) | θ value | 4.03905 (rank : 115) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q61818, Q6ZPH7, Q8CEV1 | Gene names | Rai1, Kiaa1820 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoic acid-induced protein 1. | |||||
|
NFIB_MOUSE
|
||||||
| NC score | 0.022425 (rank : 110) | θ value | 0.279714 (rank : 43) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P97863, P70252, P70253, P70254, Q9R1G4 | Gene names | Nfib | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor 1 B-type (Nuclear factor 1/B) (NF1-B) (NFI-B) (NF-I/B) (CCAAT-box-binding transcription factor) (CTF) (TGGCA-binding protein). | |||||
|
SFRS4_HUMAN
|
||||||
| NC score | 0.022104 (rank : 111) | θ value | 8.99809 (rank : 148) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q08170, Q5VXP1, Q9BUA4, Q9UEB5 | Gene names | SFRS4, SRP75 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 4 (Pre-mRNA-splicing factor SRP75) (SRP001LB). | |||||
|
SATB1_MOUSE
|
||||||
| NC score | 0.022084 (rank : 112) | θ value | 3.0926 (rank : 103) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q60611 | Gene names | Satb1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-binding protein SATB1 (Special AT-rich sequence-binding protein 1). | |||||
|
STAT6_HUMAN
|
||||||
| NC score | 0.021791 (rank : 113) | θ value | 1.38821 (rank : 75) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P42226 | Gene names | STAT6 | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 6 (IL-4 Stat). | |||||
|
PTN21_HUMAN
|
||||||
| NC score | 0.021612 (rank : 114) | θ value | 0.279714 (rank : 44) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q16825 | Gene names | PTPN21, PTPD1 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 21 (EC 3.1.3.48) (Protein-tyrosine phosphatase D1). | |||||
|
SYN1_MOUSE
|
||||||
| NC score | 0.021563 (rank : 115) | θ value | 4.03905 (rank : 118) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 363 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O88935, Q62279, Q8QZT8 | Gene names | Syn1, Syn-1 | |||
|
Domain Architecture |

|
|||||
| Description | Synapsin-1 (Synapsin I). | |||||
|
NKTR_HUMAN
|
||||||
| NC score | 0.020269 (rank : 116) | θ value | 3.0926 (rank : 100) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 658 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P30414 | Gene names | NKTR | |||
|
Domain Architecture |

|
|||||
| Description | NK-tumor recognition protein (Natural-killer cells cyclophilin-related protein) (NK-TR protein). | |||||
|
FANCJ_HUMAN
|
||||||
| NC score | 0.019596 (rank : 117) | θ value | 3.0926 (rank : 94) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BX63, Q8NCI5 | Gene names | BRIP1, BACH1, FANCJ | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fanconi anemia group J protein (EC 3.6.1.-) (ATP-dependent RNA helicase BRIP1) (Protein FACJ) (BRCA1-interacting protein C-terminal helicase 1) (BRCA1-interacting protein 1) (BRCA1-associated C-terminal helicase 1). | |||||
|
CF055_MOUSE
|
||||||
| NC score | 0.019359 (rank : 118) | θ value | 4.03905 (rank : 110) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9CR26 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | Protein C6orf55 homolog. | |||||
|
REXO1_MOUSE
|
||||||
| NC score | 0.019148 (rank : 119) | θ value | 6.88961 (rank : 133) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q7TT28, Q3UMP1, Q69ZR0, Q6NSQ6, Q6PI95, Q9DA29 | Gene names | Rexo1, Kiaa1138, Tceb3bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA exonuclease 1 homolog (EC 3.1.-.-) (Transcription elongation factor B polypeptide 3-binding protein 1). | |||||
|
XE7_HUMAN
|
||||||
| NC score | 0.018691 (rank : 120) | θ value | 1.38821 (rank : 77) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 738 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q02040, Q02832 | Gene names | CXYorf3, DXYS155E, XE7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-lymphocyte antigen precursor (B-lymphocyte surface antigen) (721P) (Protein XE7). | |||||
|
RFX1_MOUSE
|
||||||
| NC score | 0.018689 (rank : 121) | θ value | 1.06291 (rank : 63) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P48377 | Gene names | Rfx1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-binding protein RFX1. | |||||
|
CREB5_HUMAN
|
||||||
| NC score | 0.018585 (rank : 122) | θ value | 1.81305 (rank : 78) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q02930, Q05886, Q06246 | Gene names | CREB5, CREBPA | |||
|
Domain Architecture |

|
|||||
| Description | cAMP response element-binding protein 5 (CRE-BPa). | |||||
|
SMCA2_HUMAN
|
||||||
| NC score | 0.018352 (rank : 123) | θ value | 0.0736092 (rank : 34) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 503 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P51531 | Gene names | SMARCA2, BRM, SNF2A, SNF2L2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L2 (EC 3.6.1.-) (ATP- dependent helicase SMARCA2) (SNF2-alpha) (SWI/SNF-related matrix- associated actin-dependent regulator of chromatin subfamily A member 2) (hBRM). | |||||
|
DCC_MOUSE
|
||||||
| NC score | 0.017723 (rank : 124) | θ value | 0.0330416 (rank : 30) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P70211 | Gene names | Dcc | |||
|
Domain Architecture |

|
|||||
| Description | Netrin receptor DCC precursor (Tumor suppressor protein DCC). | |||||
|
PHC3_MOUSE
|
||||||
| NC score | 0.017501 (rank : 125) | θ value | 4.03905 (rank : 114) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8CHP6, Q3TLV5 | Gene names | Phc3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyhomeotic-like protein 3. | |||||
|
MEF2D_HUMAN
|
||||||
| NC score | 0.017145 (rank : 126) | θ value | 3.0926 (rank : 97) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q14814, Q14815 | Gene names | MEF2D | |||
|
Domain Architecture |

|
|||||
| Description | Myocyte-specific enhancer factor 2D. | |||||
|
PTN21_MOUSE
|
||||||
| NC score | 0.016858 (rank : 127) | θ value | 2.36792 (rank : 87) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q62136 | Gene names | Ptpn21 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 21 (EC 3.1.3.48) (Protein-tyrosine phosphatase PTP-RL10). | |||||
|
MEF2D_MOUSE
|
||||||
| NC score | 0.016733 (rank : 128) | θ value | 3.0926 (rank : 98) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q63943, Q63944 | Gene names | Mef2d | |||
|
Domain Architecture |

|
|||||
| Description | Myocyte-specific enhancer factor 2D. | |||||
|
IF4B_MOUSE
|
||||||
| NC score | 0.016381 (rank : 129) | θ value | 6.88961 (rank : 130) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BGD9, Q3TD64 | Gene names | Eif4b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4B (eIF-4B). | |||||
|
RM40_HUMAN
|
||||||
| NC score | 0.016119 (rank : 130) | θ value | 8.99809 (rank : 145) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NQ50, O95134 | Gene names | MRPL40, NLVCF, URIM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 39S ribosomal protein L40, mitochondrial precursor (L40mt) (MRP-40) (Nuclear localization signal-containing protein deleted in velocardiofacial syndrome) (Up-regulated in metastasis). | |||||
|
SURF6_MOUSE
|
||||||
| NC score | 0.015507 (rank : 131) | θ value | 4.03905 (rank : 117) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 195 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P70279, Q8BPI4, Q8BRL7, Q8R0Q7 | Gene names | Surf6, Surf-6 | |||
|
Domain Architecture |

|
|||||
| Description | Surfeit locus protein 6. | |||||
|
CA106_HUMAN
|
||||||
| NC score | 0.014150 (rank : 132) | θ value | 6.88961 (rank : 126) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q3KP66, Q9NV65, Q9NVI0 | Gene names | C1orf106 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C1orf106. | |||||
|
PDZ11_MOUSE
|
||||||
| NC score | 0.013976 (rank : 133) | θ value | 2.36792 (rank : 86) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9CZG9 | Gene names | Pdzd11, Pdzk11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 11. | |||||
|
DTX2_HUMAN
|
||||||
| NC score | 0.013822 (rank : 134) | θ value | 3.0926 (rank : 93) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q86UW9, Q96H69, Q9H890, Q9P200 | Gene names | DTX2, KIAA1528 | |||
|
Domain Architecture |

|
|||||
| Description | Protein deltex-2 (Deltex-2) (Deltex2) (hDTX2). | |||||
|
ZDHC8_HUMAN
|
||||||
| NC score | 0.013816 (rank : 135) | θ value | 8.99809 (rank : 157) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9ULC8, Q6ICL1, Q6ZNF5, Q7Z6L9 | Gene names | ZDHHC8, KIAA1292, ZDHHCL1, ZNF378 | |||
|
Domain Architecture |

|
|||||
| Description | Probable palmitoyltransferase ZDHHC8 (EC 2.3.1.-) (Zinc finger DHHC domain-containing protein 8) (DHHC-8) (Zinc finger protein 378). | |||||
|
XYLT1_HUMAN
|
||||||
| NC score | 0.013337 (rank : 136) | θ value | 3.0926 (rank : 107) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q86Y38, Q9H1B6 | Gene names | XYLT1, XT1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Xylosyltransferase 1 (EC 2.4.2.26) (Xylosyltransferase I) (XylT-I) (XT-I) (Peptide O-xylosyltransferase 1). | |||||
|
CSKI1_MOUSE
|
||||||
| NC score | 0.012457 (rank : 137) | θ value | 5.27518 (rank : 121) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 1031 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q6P9K8, Q6ZPU2, Q8BWU2, Q8BX99, Q9CXH0 | Gene names | Caskin1, Kiaa1306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-1 (CASK-interacting protein 1). | |||||
|
ADA19_HUMAN
|
||||||
| NC score | 0.012170 (rank : 138) | θ value | 2.36792 (rank : 82) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9H013, Q9BZL5, Q9UHP2 | Gene names | ADAM19, MLTNB | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 19 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 19) (Meltrin beta) (Metalloprotease and disintegrin dentritic antigen marker) (MADDAM). | |||||
|
SPG20_MOUSE
|
||||||
| NC score | 0.011337 (rank : 139) | θ value | 6.88961 (rank : 135) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8R1X6, Q6ZQ87, Q8BJD3, Q8BM37, Q8BZ63 | Gene names | Spg20, Kiaa0610 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Spartin. | |||||
|
SMAD9_HUMAN
|
||||||
| NC score | 0.010962 (rank : 140) | θ value | 1.38821 (rank : 73) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O15198, O14989 | Gene names | SMAD9, MADH6, MADH9 | |||
|
Domain Architecture |

|
|||||
| Description | Mothers against decapentaplegic homolog 9 (SMAD 9) (Mothers against DPP homolog 9) (Smad9) (Madh6). | |||||
|
RPO3D_MOUSE
|
||||||
| NC score | 0.010365 (rank : 141) | θ value | 8.99809 (rank : 146) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q91WD1, Q9CZ02 | Gene names | Polr3d, Bn51t | |||
|
Domain Architecture |

|
|||||
| Description | DNA-directed RNA polymerase III subunit D (EC 2.7.7.6) (DNA-directed RNA polymerase III 47 kDa polypeptide) (RNA polymerase C subunit 4) (RPC4). | |||||
|
TIP60_MOUSE
|
||||||
| NC score | 0.009884 (rank : 142) | θ value | 8.99809 (rank : 153) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8CHK4, Q8CGZ3, Q8CGZ4, Q8VIH0 | Gene names | Htatip, Tip60 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase HTATIP (EC 2.3.1.48) (EC 2.3.1.-) (60 kDa Tat interactive protein) (Tip60). | |||||
|
SIM2_HUMAN
|
||||||
| NC score | 0.009817 (rank : 143) | θ value | 8.99809 (rank : 149) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q14190, O60766, Q15470, Q15471, Q15472, Q15473, Q16532 | Gene names | SIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2. | |||||
|
TIP60_HUMAN
|
||||||
| NC score | 0.009718 (rank : 144) | θ value | 8.99809 (rank : 152) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q92993, O95624, Q13430, Q6GSE8, Q9BWK7 | Gene names | HTATIP, TIP60 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase HTATIP (EC 2.3.1.48) (EC 2.3.1.-) (60 kDa Tat interactive protein) (Tip60) (HIV-1 Tat interactive protein) (cPLA(2)-interacting protein). | |||||
|
PDZ11_HUMAN
|
||||||
| NC score | 0.009616 (rank : 145) | θ value | 8.99809 (rank : 143) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5EBL8, Q6UWE1, Q9P0Q1 | Gene names | PDZD11, PDZK11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 11. | |||||
|
JARD2_HUMAN
|
||||||
| NC score | 0.009593 (rank : 146) | θ value | 8.99809 (rank : 142) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q92833, Q5U5L5 | Gene names | JARID2, JMJ | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji protein (Jumonji/ARID domain-containing protein 2). | |||||
|
U2AFL_HUMAN
|
||||||
| NC score | 0.008946 (rank : 147) | θ value | 8.99809 (rank : 154) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 503 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q15695, Q13570 | Gene names | ZRSR1, U2AF1-RS1, U2AF1L1, U2AF1RS1, U2AFBPL | |||
|
Domain Architecture |

|
|||||
| Description | U2 small nuclear ribonucleoprotein auxiliary factor 35 kDa subunit- related protein 1 (U2(RNU2) small nuclear RNA auxiliary factor 1-like 1) (CCCH type zinc finger, RNA-binding motif and serine/arginine rich protein 1). | |||||
|
THSD1_HUMAN
|
||||||
| NC score | 0.008840 (rank : 148) | θ value | 8.99809 (rank : 151) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NS62, Q6P3U1, Q6UXZ2 | Gene names | THSD1, TMTSP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thrombospondin type-1 domain-containing protein 1 precursor (Transmembrane molecule with thrombospondin module). | |||||
|
USH1C_MOUSE
|
||||||
| NC score | 0.008733 (rank : 149) | θ value | 5.27518 (rank : 124) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 557 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9ES64, Q91XD1, Q9CVG7, Q9ES65 | Gene names | Ush1c | |||
|
Domain Architecture |

|
|||||
| Description | Harmonin (Usher syndrome type-1C protein homolog) (PDZ domain- containing protein). | |||||
|
AAKG2_HUMAN
|
||||||
| NC score | 0.008254 (rank : 150) | θ value | 8.99809 (rank : 136) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UGJ0, Q53Y07, Q6NUI0, Q75MP4, Q9NUZ9, Q9UDN8, Q9ULX8 | Gene names | PRKAG2 | |||
|
Domain Architecture |

|
|||||
| Description | 5'-AMP-activated protein kinase subunit gamma-2 (AMPK gamma-2 chain) (AMPK gamma2) (H91620p). | |||||
|
ADA19_MOUSE
|
||||||
| NC score | 0.007698 (rank : 151) | θ value | 5.27518 (rank : 120) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O35674 | Gene names | Adam19, Mltnb | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 19 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 19) (Meltrin beta). | |||||
|
ADAM8_MOUSE
|
||||||
| NC score | 0.007541 (rank : 152) | θ value | 3.0926 (rank : 91) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 326 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q05910 | Gene names | Adam8, Ms2 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 8 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 8) (Cell surface antigen MS2) (Macrophage cysteine-rich glycoprotein) (CD156 antigen). | |||||
|
AXN2_MOUSE
|
||||||
| NC score | 0.006516 (rank : 153) | θ value | 8.99809 (rank : 137) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O88566, Q9QXJ6 | Gene names | Axin2 | |||
|
Domain Architecture |

|
|||||
| Description | Axin-2 (Axis inhibition protein 2) (Conductin) (Axin-like protein) (Axil). | |||||
|
SOX10_HUMAN
|
||||||
| NC score | 0.005782 (rank : 154) | θ value | 6.88961 (rank : 134) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P56693 | Gene names | SOX10 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-10. | |||||
|
LTB1L_HUMAN
|
||||||
| NC score | 0.005684 (rank : 155) | θ value | 0.813845 (rank : 55) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 553 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q14766 | Gene names | LTBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein, isoform 1L precursor (LTBP-1) (Transforming growth factor beta-1-binding protein 1) (TGF-beta1-BP-1). | |||||
|
CCD39_HUMAN
|
||||||
| NC score | 0.005504 (rank : 156) | θ value | 2.36792 (rank : 83) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 868 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UFE4 | Gene names | CCDC39 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 39. | |||||
|
CAC1A_MOUSE
|
||||||
| NC score | 0.004090 (rank : 157) | θ value | 8.99809 (rank : 138) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P97445 | Gene names | Cacna1a, Cach4, Cacn3, Cacnl1a4, Ccha1a | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent P/Q-type calcium channel subunit alpha-1A (Voltage- gated calcium channel subunit alpha Cav2.1) (Calcium channel, L type, alpha-1 polypeptide isoform 4) (Brain calcium channel I) (BI). | |||||
|
FAT2_HUMAN
|
||||||
| NC score | 0.002274 (rank : 158) | θ value | 3.0926 (rank : 95) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9NYQ8, O75091, Q9NSR7 | Gene names | FAT2, CDHF8, KIAA0811, MEGF1 | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin Fat 2 precursor (hFat2) (Multiple epidermal growth factor-like domains 1). | |||||
|
WDR47_HUMAN
|
||||||
| NC score | 0.001870 (rank : 159) | θ value | 8.99809 (rank : 156) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 295 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O94967, Q8IXT7, Q8IYU9 | Gene names | WDR47, KIAA0893 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 47. | |||||
|
CORO7_MOUSE
|
||||||
| NC score | 0.001800 (rank : 160) | θ value | 8.99809 (rank : 140) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9D2V7, Q3UDJ4, Q6P8Y8, Q8C9V7 | Gene names | Coro7 | |||
|
Domain Architecture |

|
|||||
| Description | Coronin-7 (70 kDa WD repeat tumor rejection antigen homolog). | |||||
|
HXA3_MOUSE
|
||||||
| NC score | 0.001360 (rank : 161) | θ value | 8.99809 (rank : 141) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 558 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P02831, Q4V9Z8, Q61197 | Gene names | Hoxa3, Hox-1.5, Hoxa-3 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-A3 (Hox-1.5) (MO-10). | |||||
|
ZBTB5_HUMAN
|
||||||
| NC score | 0.000792 (rank : 162) | θ value | 1.06291 (rank : 66) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 811 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O15062 | Gene names | ZBTB5, KIAA0354 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 5. | |||||
|
CAD15_HUMAN
|
||||||
| NC score | 0.000552 (rank : 163) | θ value | 8.99809 (rank : 139) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P55291 | Gene names | CDH15, CDH14, CDH3 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin-15 precursor (Muscle-cadherin) (M-cadherin) (Cadherin-14). | |||||
|
MARK2_HUMAN
|
||||||
| NC score | -0.000990 (rank : 164) | θ value | 2.36792 (rank : 85) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 1056 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q7KZI7, Q15449, Q15524, Q5XGA3, Q68A18, Q96HB3, Q96RG0 | Gene names | MARK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MARK2 (EC 2.7.11.1) (MAP/microtubule affinity-regulating kinase 2) (ELKL motif kinase) (EMK1) (PAR1 homolog). | |||||
|
MK15_HUMAN
|
||||||
| NC score | -0.001270 (rank : 165) | θ value | 3.0926 (rank : 99) | |||
| Query Neighborhood Hits | 157 | Target Neighborhood Hits | 1038 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8TD08, Q2TCF9, Q8N362 | Gene names | MAPK15, ERK8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase 15 (EC 2.7.11.24) (Extracellular signal-regulated kinase 8). | |||||