Please be patient as the page loads
|
SNIP1_HUMAN
|
||||||
| SwissProt Accessions | Q8TAD8, Q96SP9, Q9H9T7 | Gene names | SNIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Smad nuclear-interacting protein 1. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
SNIP1_HUMAN
|
||||||
| θ value | 3.82926e-184 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | Q8TAD8, Q96SP9, Q9H9T7 | Gene names | SNIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Smad nuclear-interacting protein 1. | |||||
|
SNIP1_MOUSE
|
||||||
| θ value | 3.06238e-149 (rank : 2) | NC score | 0.937560 (rank : 2) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q8BIZ6, Q3V106, Q8BIZ4 | Gene names | Snip1 | |||
|
Domain Architecture |

|
|||||
| Description | Smad nuclear-interacting protein 1. | |||||
|
PP1R8_HUMAN
|
||||||
| θ value | 5.62301e-10 (rank : 3) | NC score | 0.483403 (rank : 3) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q12972, Q5TIF2, Q6PKF6, Q9UBH1, Q9UBZ0 | Gene names | PPP1R8, ARD1, NIPP1 | |||
|
Domain Architecture |
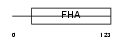
|
|||||
| Description | Nuclear inhibitor of protein phosphatase 1 (NIPP-1) (Protein phosphatase 1 regulatory inhibitor subunit 8) [Includes: Activator of RNA decay (EC 3.1.4.-) (ARD-1)]. | |||||
|
PP1R8_MOUSE
|
||||||
| θ value | 5.62301e-10 (rank : 4) | NC score | 0.483189 (rank : 4) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8R3G1, Q8C087 | Gene names | Ppp1r8, Nipp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear inhibitor of protein phosphatase 1 (NIPP-1) (Protein phosphatase 1 regulatory inhibitor subunit 8). | |||||
|
NADAP_HUMAN
|
||||||
| θ value | 9.59137e-10 (rank : 5) | NC score | 0.431715 (rank : 5) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9BWU0, Q4KMT1, Q4KMX0, Q7Z5Q9, Q9NVN2 | Gene names | SLC4A1AP, HLC3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kanadaptin (Kidney anion exchanger adapter protein) (Solute carrier family 4 anion exchanger member 1 adapter protein) (Lung cancer oncogene 3 protein). | |||||
|
RPTN_MOUSE
|
||||||
| θ value | 9.29e-05 (rank : 6) | NC score | 0.163743 (rank : 9) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P97347 | Gene names | Rptn | |||
|
Domain Architecture |

|
|||||
| Description | Repetin. | |||||
|
CCD55_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 7) | NC score | 0.198085 (rank : 7) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9H0G5, Q6FI71 | Gene names | CCDC55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 55. | |||||
|
NGEF_MOUSE
|
||||||
| θ value | 0.00869519 (rank : 8) | NC score | 0.050560 (rank : 90) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8CHT1, Q8R204, Q923H2, Q9JHT9 | Gene names | Ngef | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ephexin-1 (Eph-interacting exchange protein) (Neuronal guanine nucleotide exchange factor). | |||||
|
TARA_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 9) | NC score | 0.095332 (rank : 23) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9H2D6, O94797, Q2PZW8, Q2Q3Z9, Q2Q400, Q5R3M6, Q96DW1, Q9BT77, Q9BTL7, Q9BY98, Q9Y3L4 | Gene names | TRIOBP, KIAA1662, TARA | |||
|
Domain Architecture |

|
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
CCD55_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 10) | NC score | 0.191315 (rank : 8) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q5NCR9 | Gene names | Ccdc55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 55. | |||||
|
NIPBL_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 11) | NC score | 0.143635 (rank : 13) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
PPIG_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 12) | NC score | 0.108800 (rank : 21) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q13427, O00706, Q96DG9 | Gene names | PPIG | |||
|
Domain Architecture |

|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase G (EC 5.2.1.8) (Peptidyl-prolyl isomerase G) (PPIase G) (Rotamase G) (Cyclophilin G) (Clk-associating RS-cyclophilin) (CARS-cyclophilin) (CARS-Cyp) (SR-cyclophilin) (SRcyp) (SR-cyp) (CASP10). | |||||
|
RP3A_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 13) | NC score | 0.036814 (rank : 106) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 510 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P47708 | Gene names | Rph3a | |||
|
Domain Architecture |

|
|||||
| Description | Rabphilin-3A (Exophilin-1). | |||||
|
RPTN_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 14) | NC score | 0.153333 (rank : 11) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q6XPR3 | Gene names | RPTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Repetin. | |||||
|
CHD1_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 15) | NC score | 0.063206 (rank : 57) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P40201 | Gene names | Chd1, Chd-1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
CD2L5_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 16) | NC score | 0.021149 (rank : 118) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 987 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q14004, Q6DKQ9, Q75MH4, Q75MH5, Q96JN4, Q9H4A0, Q9H4A1, Q9UDR4 | Gene names | CDC2L5, CDC2L, CHED, KIAA1791 | |||
|
Domain Architecture |
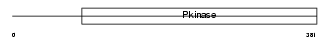
|
|||||
| Description | Cell division cycle 2-like protein kinase 5 (EC 2.7.11.22) (CDC2- related protein kinase 5) (Cholinesterase-related cell division controller). | |||||
|
FILA_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 17) | NC score | 0.159460 (rank : 10) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 685 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P20930, Q01720, Q5T583 | Gene names | FLG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filaggrin. | |||||
|
RPGF1_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 18) | NC score | 0.050889 (rank : 88) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13905 | Gene names | RAPGEF1, GRF2 | |||
|
Domain Architecture |

|
|||||
| Description | Rap guanine nucleotide exchange factor 1 (Guanine nucleotide-releasing factor 2) (C3G protein) (CRK SH3-binding GNRP). | |||||
|
T2_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 19) | NC score | 0.213098 (rank : 6) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q06666 | Gene names | Srst, T2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Octapeptide-repeat protein T2. | |||||
|
ANK2_HUMAN
|
||||||
| θ value | 0.125558 (rank : 20) | NC score | 0.031093 (rank : 110) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
CHD1_HUMAN
|
||||||
| θ value | 0.125558 (rank : 21) | NC score | 0.066354 (rank : 50) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O14646 | Gene names | CHD1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
NPBL_MOUSE
|
||||||
| θ value | 0.125558 (rank : 22) | NC score | 0.138979 (rank : 15) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
FBXW7_HUMAN
|
||||||
| θ value | 0.163984 (rank : 23) | NC score | 0.020386 (rank : 119) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q969H0, Q96A16, Q96LE0, Q96RI2, Q9NUX6 | Gene names | FBXW7, FBW7, FBX30, SEL10 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/WD repeat protein 7 (F-box and WD-40 domain protein 7) (F-box protein FBX30) (hCdc4) (Archipelago homolog) (hAgo) (SEL-10). | |||||
|
TGON2_HUMAN
|
||||||
| θ value | 0.163984 (rank : 24) | NC score | 0.072082 (rank : 43) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
OTUD3_HUMAN
|
||||||
| θ value | 0.365318 (rank : 25) | NC score | 0.078507 (rank : 32) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5T2D3, O75047 | Gene names | OTUD3, KIAA0459 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | OTU domain-containing protein 3. | |||||
|
RBBP6_MOUSE
|
||||||
| θ value | 0.62314 (rank : 26) | NC score | 0.089814 (rank : 24) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
SFRIP_HUMAN
|
||||||
| θ value | 0.62314 (rank : 27) | NC score | 0.113926 (rank : 18) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 530 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q99590, Q8IW59 | Gene names | SFRS2IP, CASP11, SIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SFRS2-interacting protein (Splicing factor, arginine/serine-rich 2- interacting protein) (SC35-interacting protein 1) (CTD-associated SR protein 11) (Splicing regulatory protein 129) (SRrp129) (NY-REN-40 antigen). | |||||
|
SPRL1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 28) | NC score | 0.056761 (rank : 72) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P70663, P97810, Q99L82 | Gene names | Sparcl1, Ecm2, Sc1 | |||
|
Domain Architecture |

|
|||||
| Description | SPARC-like protein 1 precursor (Matrix glycoprotein Sc1) (Extracellular matrix protein 2). | |||||
|
TRHY_HUMAN
|
||||||
| θ value | 0.62314 (rank : 29) | NC score | 0.056423 (rank : 73) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1385 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q07283 | Gene names | THH, THL, TRHY | |||
|
Domain Architecture |

|
|||||
| Description | Trichohyalin. | |||||
|
DVL1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 30) | NC score | 0.023580 (rank : 116) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P51141, Q60868 | Gene names | Dvl1, Dvl | |||
|
Domain Architecture |
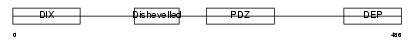
|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1 (Dishevelled-1) (DSH homolog 1). | |||||
|
RU17_MOUSE
|
||||||
| θ value | 0.813845 (rank : 31) | NC score | 0.068534 (rank : 46) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q62376, Q3UIW4 | Gene names | Snrp70 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U1 small nuclear ribonucleoprotein 70 kDa (U1 SNRNP 70 kDa) (snRNP70). | |||||
|
SFR12_MOUSE
|
||||||
| θ value | 0.813845 (rank : 32) | NC score | 0.127277 (rank : 16) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8BZX4, Q8BLM8, Q8BWK7, Q8BYK0, Q8BYZ4 | Gene names | Sfrs12, Srrp86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86). | |||||
|
ZKSC1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 33) | NC score | 0.003666 (rank : 130) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 949 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P17029, P52745, Q8TBW5, Q8TEK7 | Gene names | ZKSCAN1, KOX18, ZNF139, ZNF36 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger with KRAB and SCAN domain-containing protein 1 (Zinc finger protein 36) (Zinc finger protein KOX18). | |||||
|
CAC1A_HUMAN
|
||||||
| θ value | 1.06291 (rank : 34) | NC score | 0.030855 (rank : 111) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O00555, P78510, P78511, Q16290, Q92690, Q99790, Q99791, Q99792, Q99793 | Gene names | CACNA1A, CACH4, CACN3, CACNL1A4 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent P/Q-type calcium channel subunit alpha-1A (Voltage- gated calcium channel subunit alpha Cav2.1) (Calcium channel, L type, alpha-1 polypeptide isoform 4) (Brain calcium channel I) (BI). | |||||
|
CROP_HUMAN
|
||||||
| θ value | 1.06291 (rank : 35) | NC score | 0.109250 (rank : 19) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O95232, Q6PHR9, Q9NUY0, Q9P2S7 | Gene names | CROP, CREAP1, O48 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cisplatin resistance-associated overexpressed protein (cAMP regulatory element-associated protein 1) (CRE-associated protein 1) (CREAP-1) (Luc7A) (Okadaic acid-inducible phosphoprotein OA48-18). | |||||
|
MRCKB_HUMAN
|
||||||
| θ value | 1.06291 (rank : 36) | NC score | 0.015298 (rank : 123) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 2239 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9Y5S2, Q2L7A5, Q86TJ1, Q9ULU5 | Gene names | CDC42BPB, KIAA1124 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK beta (EC 2.7.11.1) (CDC42-binding protein kinase beta) (Myotonic dystrophy kinase-related CDC42-binding kinase beta) (Myotonic dystrophy protein kinase-like beta) (MRCK beta) (DMPK-like beta). | |||||
|
US6NL_MOUSE
|
||||||
| θ value | 1.06291 (rank : 37) | NC score | 0.048955 (rank : 95) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q80XC3, Q6ZQK8 | Gene names | Usp6nl, Kiaa0019 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USP6 N-terminal-like protein. | |||||
|
BSND_MOUSE
|
||||||
| θ value | 1.38821 (rank : 38) | NC score | 0.046438 (rank : 97) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8VIM4, Q8C740, Q8CHY0 | Gene names | Bsnd | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Barttin. | |||||
|
CK035_MOUSE
|
||||||
| θ value | 1.38821 (rank : 39) | NC score | 0.057722 (rank : 68) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q0VET5, Q9CW10 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C11orf35 homolog. | |||||
|
DBNL_HUMAN
|
||||||
| θ value | 1.38821 (rank : 40) | NC score | 0.038750 (rank : 105) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 244 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UJU6, P84070, Q6IAI8, Q96F30, Q96K74, Q9HBN8, Q9NR72 | Gene names | DBNL, CMAP, SH3P7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Drebrin-like protein (SH3 domain-containing protein 7) (Drebrin F) (Cervical SH3P7) (HPK1-interacting protein of 55 kDa) (HIP-55) (Cervical mucin-associated protein). | |||||
|
LA_MOUSE
|
||||||
| θ value | 1.38821 (rank : 41) | NC score | 0.041985 (rank : 102) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P32067 | Gene names | Ssb, Ss-b | |||
|
Domain Architecture |
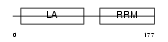
|
|||||
| Description | Lupus La protein homolog (La ribonucleoprotein) (La autoantigen homolog). | |||||
|
SRY_MOUSE
|
||||||
| θ value | 1.38821 (rank : 42) | NC score | 0.043122 (rank : 101) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q05738 | Gene names | Sry, Tdf, Tdy | |||
|
Domain Architecture |

|
|||||
| Description | Sex-determining region Y protein (Testis-determining factor). | |||||
|
US6NL_HUMAN
|
||||||
| θ value | 1.38821 (rank : 43) | NC score | 0.048679 (rank : 96) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q92738, Q15400, Q7L0K9 | Gene names | USP6NL, KIAA0019 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USP6 N-terminal-like protein (Related to the N-terminus of tre) (RN- tre). | |||||
|
CROP_MOUSE
|
||||||
| θ value | 1.81305 (rank : 44) | NC score | 0.108895 (rank : 20) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q5SUF2, Q3U9D5, Q8BUJ5, Q921Z3, Q9CRS7, Q9CTY4 | Gene names | Crop | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cisplatin resistance-associated overexpressed protein. | |||||
|
ELOA1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 45) | NC score | 0.065725 (rank : 51) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q14241, Q8IXH1 | Gene names | TCEB3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription elongation factor B polypeptide 3 (RNA polymerase II transcription factor SIII subunit A1) (SIII p110) (Elongin-A) (EloA) (Elongin 110 kDa subunit). | |||||
|
ERCC6_HUMAN
|
||||||
| θ value | 1.81305 (rank : 46) | NC score | 0.034717 (rank : 107) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q03468 | Gene names | ERCC6, CSB | |||
|
Domain Architecture |

|
|||||
| Description | DNA excision repair protein ERCC-6 (EC 3.6.1.-) (ATP-dependent helicase ERCC6) (Cockayne syndrome protein CSB). | |||||
|
RIMS2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 47) | NC score | 0.038908 (rank : 104) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 376 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9EQZ7, Q8C433, Q8CCK2 | Gene names | Rims2, Rab3ip2, Rim2 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 2 (Rab3-interacting molecule 2) (RIM 2) (Rab3-interacting protein 2). | |||||
|
RNF8_HUMAN
|
||||||
| θ value | 1.81305 (rank : 48) | NC score | 0.033832 (rank : 108) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 622 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O76064 | Gene names | RNF8, KIAA0646 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin ligase protein RNF8 (EC 6.3.2.-) (RING finger protein 8). | |||||
|
ZC313_HUMAN
|
||||||
| θ value | 1.81305 (rank : 49) | NC score | 0.145471 (rank : 12) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q5T200, O94936, Q5T1Z9, Q7Z7J3, Q8NDT6, Q9H0L6 | Gene names | ZC3H13, KIAA0853 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 13. | |||||
|
DDX46_MOUSE
|
||||||
| θ value | 2.36792 (rank : 50) | NC score | 0.032365 (rank : 109) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 358 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q569Z5, Q6ZQ42, Q8R0R6 | Gene names | Ddx46, Kiaa0801 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX46 (EC 3.6.1.-) (DEAD box protein 46). | |||||
|
KIF3B_HUMAN
|
||||||
| θ value | 2.36792 (rank : 51) | NC score | 0.013900 (rank : 125) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 540 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O15066 | Gene names | KIF3B, KIAA0359 | |||
|
Domain Architecture |
|
|||||
| Description | Kinesin-like protein KIF3B (Microtubule plus end-directed kinesin motor 3B) (HH0048). | |||||
|
KIF3B_MOUSE
|
||||||
| θ value | 2.36792 (rank : 52) | NC score | 0.013960 (rank : 124) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 533 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q61771 | Gene names | Kif3b | |||
|
Domain Architecture |
|
|||||
| Description | Kinesin-like protein KIF3B (Microtubule plus end-directed kinesin motor 3B). | |||||
|
MDC1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 53) | NC score | 0.043209 (rank : 100) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
SFR12_HUMAN
|
||||||
| θ value | 2.36792 (rank : 54) | NC score | 0.141250 (rank : 14) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 553 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8WXA9, Q86X37 | Gene names | SFRS12, SRRP86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86) (Splicing regulatory protein 508) (SRrp508). | |||||
|
CBX8_HUMAN
|
||||||
| θ value | 3.0926 (rank : 55) | NC score | 0.049806 (rank : 94) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9HC52, Q96H39, Q9NR07 | Gene names | CBX8, PC3, RC1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 8 (Polycomb 3 homolog) (Pc3) (hPc3) (Rectachrome 1). | |||||
|
MRCKB_MOUSE
|
||||||
| θ value | 3.0926 (rank : 56) | NC score | 0.013655 (rank : 126) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 2180 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q7TT50, Q69ZR5, Q80W33 | Gene names | Cdc42bpb, Kiaa1124 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK beta (EC 2.7.11.1) (CDC42-binding protein kinase beta) (Myotonic dystrophy kinase-related CDC42-binding kinase beta) (Myotonic dystrophy protein kinase-like beta) (MRCK beta) (DMPK-like beta). | |||||
|
P52K_HUMAN
|
||||||
| θ value | 3.0926 (rank : 57) | NC score | 0.027282 (rank : 113) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O43422, Q8WTW1, Q9Y3Z4 | Gene names | PRKRIR, DAP4, P52RIPK, THAP0 | |||
|
Domain Architecture |

|
|||||
| Description | 52 kDa repressor of the inhibitor of the protein kinase (p58IPK- interacting protein) (58 kDa interferon-induced protein kinase- interacting protein) (P52rIPK) (Death-associated protein 4) (THAP domain-containing protein 0). | |||||
|
GASP1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 58) | NC score | 0.027697 (rank : 112) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q5U4C1, Q8BKR8, Q8BUN4, Q8BYK9, Q8CHF4, Q8R095 | Gene names | Gprasp1, Kiaa0443 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G-protein coupled receptor-associated sorting protein 1 (GASP-1). | |||||
|
INCE_HUMAN
|
||||||
| θ value | 4.03905 (rank : 59) | NC score | 0.044829 (rank : 99) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 978 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9NQS7, Q5Y192 | Gene names | INCENP | |||
|
Domain Architecture |

|
|||||
| Description | Inner centromere protein. | |||||
|
C8AP2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 60) | NC score | 0.064747 (rank : 55) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 548 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9UKL3, Q6PH76, Q7LCQ7, Q86YD9, Q9NUQ4, Q9NZV9, Q9P2N1, Q9Y563 | Gene names | CASP8AP2, FLASH, KIAA1315, RIP25 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
CD2L5_MOUSE
|
||||||
| θ value | 5.27518 (rank : 61) | NC score | 0.016851 (rank : 121) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1011 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q69ZA1, Q80V11, Q8BZG1, Q8K0A4 | Gene names | Cdc2l5, Kiaa1791 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle 2-like protein kinase 5 (EC 2.7.11.22) (CDC2- related protein kinase 5). | |||||
|
MK07_HUMAN
|
||||||
| θ value | 5.27518 (rank : 62) | NC score | 0.007221 (rank : 129) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1087 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q13164, Q16634 | Gene names | MAPK7, ERK4, ERK5, PRKM7 | |||
|
Domain Architecture |
|
|||||
| Description | Mitogen-activated protein kinase 7 (EC 2.7.11.24) (Extracellular signal-regulated kinase 5) (ERK-5) (ERK4) (BMK1 kinase). | |||||
|
PHF20_MOUSE
|
||||||
| θ value | 5.27518 (rank : 63) | NC score | 0.053955 (rank : 79) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8BLG0, Q8BMA2, Q8BYR4, Q921N1 | Gene names | Phf20, Hca58 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58 homolog). | |||||
|
PPHLN_HUMAN
|
||||||
| θ value | 5.27518 (rank : 64) | NC score | 0.046412 (rank : 98) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8NEY8, Q86YT2, Q8IXN3, Q8TB09, Q96NB9, Q9NXL4, Q9P0P6, Q9P0R9 | Gene names | PPHLN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Periphilin-1 (Gastric cancer antigen Ga50). | |||||
|
RB15B_MOUSE
|
||||||
| θ value | 5.27518 (rank : 65) | NC score | 0.039783 (rank : 103) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 284 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6PHZ5, Q8C6G2 | Gene names | Rbm15b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative RNA-binding protein 15B (RNA-binding motif protein 15B). | |||||
|
UTY_HUMAN
|
||||||
| θ value | 5.27518 (rank : 66) | NC score | 0.017378 (rank : 120) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O14607, O14608 | Gene names | UTY | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitously transcribed Y chromosome tetratricopeptide repeat protein (Ubiquitously transcribed TPR protein on the Y chromosome). | |||||
|
ZNFX1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 67) | NC score | 0.015331 (rank : 122) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9P2E3, Q9BQM7, Q9BQM8, Q9H8C1, Q9H9S2, Q9NUM1, Q9NWW1 | Gene names | ZNFX1, KIAA1404 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NFX1-type zinc finger-containing protein 1. | |||||
|
ACINU_HUMAN
|
||||||
| θ value | 6.88961 (rank : 68) | NC score | 0.073149 (rank : 39) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9UKV3, O75158, Q9UG91, Q9UKV1, Q9UKV2 | Gene names | ACIN1, ACINUS, KIAA0670 | |||
|
Domain Architecture |

|
|||||
| Description | Apoptotic chromatin condensation inducer in the nucleus (Acinus). | |||||
|
NCOA6_MOUSE
|
||||||
| θ value | 6.88961 (rank : 69) | NC score | 0.026417 (rank : 114) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1076 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9JL19, Q9JLT9 | Gene names | Ncoa6, Aib3, Prip, Rap250, Trbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC) (Thyroid hormone receptor-binding protein). | |||||
|
RU17_HUMAN
|
||||||
| θ value | 6.88961 (rank : 70) | NC score | 0.075005 (rank : 36) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P08621, P78493, P78494, Q15364, Q15686, Q15687, Q15689, Q99377, Q9UE45, Q9UE46, Q9UE47, Q9UE48, Q9UFQ6 | Gene names | SNRP70, RPU1, U1AP1 | |||
|
Domain Architecture |

|
|||||
| Description | U1 small nuclear ribonucleoprotein 70 kDa (U1 snRNP 70 kDa) (snRNP70) (U1-70K). | |||||
|
WDR60_HUMAN
|
||||||
| θ value | 6.88961 (rank : 71) | NC score | 0.062556 (rank : 58) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8WVS4, Q9NW58 | Gene names | WDR60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 60. | |||||
|
CAC1F_HUMAN
|
||||||
| θ value | 8.99809 (rank : 72) | NC score | 0.010809 (rank : 128) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O60840, O43901 | Gene names | CACNA1F, CACNAF1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1F (Voltage- gated calcium channel subunit alpha Cav1.4). | |||||
|
DBNL_MOUSE
|
||||||
| θ value | 8.99809 (rank : 73) | NC score | 0.024510 (rank : 115) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 306 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q62418, Q80WP1, Q8BH56 | Gene names | Dbnl, Abp1, Sh3p7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Drebrin-like protein (SH3 domain-containing protein 7) (Actin-binding protein 1). | |||||
|
HES6_MOUSE
|
||||||
| θ value | 8.99809 (rank : 74) | NC score | 0.013481 (rank : 127) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JHE6, Q3U2D7 | Gene names | Hes6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription cofactor HES-6 (Hairy and enhancer of split 6). | |||||
|
IF2P_HUMAN
|
||||||
| θ value | 8.99809 (rank : 75) | NC score | 0.053820 (rank : 81) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 889 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O60841, O95805, Q9UF81, Q9UMN7 | Gene names | EIF5B, IF2, KIAA0741 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 5B (eIF-5B) (Translation initiation factor IF-2). | |||||
|
LEO1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 76) | NC score | 0.084239 (rank : 29) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
NCOA6_HUMAN
|
||||||
| θ value | 8.99809 (rank : 77) | NC score | 0.021383 (rank : 117) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1202 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q14686, Q9NTZ9, Q9UH74, Q9UK86 | Gene names | NCOA6, AIB3, KIAA0181, RAP250, TRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (PRIP) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC RAP250) (Thyroid hormone receptor-binding protein). | |||||
|
NKTR_HUMAN
|
||||||
| θ value | 8.99809 (rank : 78) | NC score | 0.073183 (rank : 38) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 658 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P30414 | Gene names | NKTR | |||
|
Domain Architecture |

|
|||||
| Description | NK-tumor recognition protein (Natural-killer cells cyclophilin-related protein) (NK-TR protein). | |||||
|
RBBP6_HUMAN
|
||||||
| θ value | 8.99809 (rank : 79) | NC score | 0.089100 (rank : 25) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
SCG1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 80) | NC score | 0.065297 (rank : 52) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P05060, Q9BQV6, Q9UJA6 | Gene names | CHGB, SCG1 | |||
|
Domain Architecture |

|
|||||
| Description | Secretogranin-1 precursor (Secretogranin I) (SgI) (Chromogranin B) (CgB) [Contains: GAWK peptide; CCB peptide]. | |||||
|
ACINU_MOUSE
|
||||||
| θ value | θ > 10 (rank : 81) | NC score | 0.058198 (rank : 66) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 712 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9JIX8, Q9CSN7, Q9CSR9, Q9CSX7, Q9R046, Q9R047 | Gene names | Acin1, Acinus | |||
|
Domain Architecture |

|
|||||
| Description | Apoptotic chromatin condensation inducer in the nucleus (Acinus). | |||||
|
AFF4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 82) | NC score | 0.054806 (rank : 76) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 611 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9UHB7, Q498B2, Q59FB3, Q6P592, Q8TDR1, Q9P0E4 | Gene names | AFF4, AF5Q31, MCEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4 (ALL1-fused gene from chromosome 5q31) (Major CDK9 elongation factor-associated protein). | |||||
|
AFF4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 83) | NC score | 0.051991 (rank : 86) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9ESC8, Q8C6K4, Q8C6W3, Q8CCH3 | Gene names | Aff4, Alf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4. | |||||
|
C8AP2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 84) | NC score | 0.053440 (rank : 82) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 405 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9WUF3, Q9CSW2 | Gene names | Casp8ap2, Flash | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
CIR_HUMAN
|
||||||
| θ value | θ > 10 (rank : 85) | NC score | 0.073031 (rank : 41) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q86X95, O95367, Q12804, Q4G1B9, Q6PJI4, Q8IWI2 | Gene names | CIR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CBF1-interacting corepressor (Recepin). | |||||
|
CIR_MOUSE
|
||||||
| θ value | θ > 10 (rank : 86) | NC score | 0.060360 (rank : 61) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9DA19, Q3V2N9, Q4KL44, Q52KL9, Q5FW66 | Gene names | Cir | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CBF1-interacting corepressor. | |||||
|
CYLC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 87) | NC score | 0.050042 (rank : 93) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P35663, Q5JQQ9 | Gene names | CYLC1, CYL, CYL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-1 (Cylicin I) (Multiple-band polypeptide I). | |||||
|
CYLC2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 88) | NC score | 0.051663 (rank : 87) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q14093 | Gene names | CYLC2, CYL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-2 (Cylicin II) (Multiple-band polypeptide II). | |||||
|
DMP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 89) | NC score | 0.081710 (rank : 31) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q13316, O43265 | Gene names | DMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1). | |||||
|
DMP1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 90) | NC score | 0.074410 (rank : 37) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O55188 | Gene names | Dmp1, Dmp | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1) (AG1). | |||||
|
DSPP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 91) | NC score | 0.068315 (rank : 48) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
DSPP_MOUSE
|
||||||
| θ value | θ > 10 (rank : 92) | NC score | 0.075955 (rank : 34) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
ENL_HUMAN
|
||||||
| θ value | θ > 10 (rank : 93) | NC score | 0.057460 (rank : 69) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q03111, Q14768 | Gene names | MLLT1, ENL, LTG19, YEATS1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein ENL (YEATS domain-containing protein 1). | |||||
|
GPTC8_HUMAN
|
||||||
| θ value | θ > 10 (rank : 94) | NC score | 0.100281 (rank : 22) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 366 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9UKJ3, O60300 | Gene names | GPATC8, KIAA0553 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 8. | |||||
|
HIRP3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 95) | NC score | 0.059007 (rank : 65) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9BW71, O75707, O75708 | Gene names | HIRIP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIRA-interacting protein 3. | |||||
|
HORN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 96) | NC score | 0.088805 (rank : 26) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q86YZ3 | Gene names | HRNR, S100A18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hornerin. | |||||
|
HORN_MOUSE
|
||||||
| θ value | θ > 10 (rank : 97) | NC score | 0.082113 (rank : 30) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8VHD8 | Gene names | Hrnr | |||
|
Domain Architecture |

|
|||||
| Description | Hornerin. | |||||
|
IWS1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 98) | NC score | 0.071725 (rank : 44) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
IWS1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 99) | NC score | 0.063822 (rank : 56) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
LC7L2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.061574 (rank : 59) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9Y383, Q8IUP9, Q9NVL3, Q9NVN7, Q9UQN1 | Gene names | LUC7L2 | |||
|
Domain Architecture |
|
|||||
| Description | Putative RNA-binding protein Luc7-like 2. | |||||
|
LC7L2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.065213 (rank : 53) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 244 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q7TNC4, Q99LM5, Q99PC3 | Gene names | Luc7l2 | |||
|
Domain Architecture |
|
|||||
| Description | Putative RNA-binding protein Luc7-like 2 (CGI-74 homolog). | |||||
|
LEO1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.077959 (rank : 33) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
NKTR_MOUSE
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.056957 (rank : 70) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P30415 | Gene names | Nktr | |||
|
Domain Architecture |

|
|||||
| Description | NK-tumor recognition protein (Natural-killer cells cyclophilin-related protein) (NK-TR protein). | |||||
|
NOLC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.054445 (rank : 77) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q14978, Q15030 | Gene names | NOLC1, KIAA0035 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar phosphoprotein p130 (Nucleolar 130 kDa protein) (140 kDa nucleolar phosphoprotein) (Nopp140) (Nucleolar and coiled-body phosphoprotein 1). | |||||
|
NSBP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.057944 (rank : 67) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P82970 | Gene names | NSBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1. | |||||
|
NSBP1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 106) | NC score | 0.050234 (rank : 92) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
NUCKS_HUMAN
|
||||||
| θ value | θ > 10 (rank : 107) | NC score | 0.060027 (rank : 64) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9H1E3, Q9H1D6, Q9H723 | Gene names | NUCKS1, NUCKS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (P1). | |||||
|
NUCKS_MOUSE
|
||||||
| θ value | θ > 10 (rank : 108) | NC score | 0.072396 (rank : 42) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q80XU3, Q8BVD8 | Gene names | Nucks1, Nucks | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (JC7). | |||||
|
PININ_MOUSE
|
||||||
| θ value | θ > 10 (rank : 109) | NC score | 0.050458 (rank : 91) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 777 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O35691, Q8CD89, Q8CGU3 | Gene names | Pnn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pinin. | |||||
|
RBM25_HUMAN
|
||||||
| θ value | θ > 10 (rank : 110) | NC score | 0.056792 (rank : 71) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P49756, Q9UEQ5, Q9UIE9 | Gene names | RBM25, RNPC7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable RNA-binding protein 25 (RNA-binding motif protein 25) (RNA- binding region-containing protein 7) (Protein S164). | |||||
|
RSRC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 111) | NC score | 0.055343 (rank : 75) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q96IZ7, Q96QK2, Q9NZE5 | Gene names | RSRC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine/serine-rich coiled coil protein 1. | |||||
|
RSRC1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 112) | NC score | 0.056032 (rank : 74) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9DBU6, Q3TF06 | Gene names | Rsrc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine/serine-rich coiled coil protein 1. | |||||
|
SCG1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 113) | NC score | 0.052761 (rank : 84) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P16014 | Gene names | Chgb, Scg-1, Scg1 | |||
|
Domain Architecture |

|
|||||
| Description | Secretogranin-1 precursor (Secretogranin I) (SgI) (Chromogranin B) (CgB). | |||||
|
SEMG2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 114) | NC score | 0.054347 (rank : 78) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q02383, Q6X2M6 | Gene names | SEMG2 | |||
|
Domain Architecture |

|
|||||
| Description | Semenogelin-2 precursor (Semenogelin II) (SGII). | |||||
|
SFR11_HUMAN
|
||||||
| θ value | θ > 10 (rank : 115) | NC score | 0.114529 (rank : 17) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q05519 | Gene names | SFRS11 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor arginine/serine-rich 11 (Arginine-rich 54 kDa nuclear protein) (p54). | |||||
|
SFR15_HUMAN
|
||||||
| θ value | θ > 10 (rank : 116) | NC score | 0.086998 (rank : 27) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O95104, Q6P1M5, Q8N3I8, Q9UFM1, Q9ULP8 | Gene names | SFRS15, KIAA1172 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 15 (CTD-binding SR-like protein RA4). | |||||
|
SFR16_HUMAN
|
||||||
| θ value | θ > 10 (rank : 117) | NC score | 0.050821 (rank : 89) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8N2M8, O96026, Q6UW71, Q96DX2 | Gene names | SFRS16, SWAP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 16 (Suppressor of white-apricot homolog 2). | |||||
|
SFR17_HUMAN
|
||||||
| θ value | θ > 10 (rank : 118) | NC score | 0.085239 (rank : 28) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8TF01, Q5T064, Q6P2B4, Q6P2N4, Q6PJ93, Q6PK36, Q7Z640, Q8TF00, Q96K10, Q96SM5, Q9P076, Q9P0C0, Q9Y4N3 | Gene names | SRRP130, C6orf111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 130 (Serine-arginine-rich- splicing regulatory protein 130) (SRrp130) (SR-rich protein) (SR- related protein). | |||||
|
SFRS4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 119) | NC score | 0.066679 (rank : 49) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q08170, Q5VXP1, Q9BUA4, Q9UEB5 | Gene names | SFRS4, SRP75 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 4 (Pre-mRNA-splicing factor SRP75) (SRP001LB). | |||||
|
SFRS4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 120) | NC score | 0.068423 (rank : 47) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8VE97, Q9JJC3 | Gene names | Sfrs4 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 4. | |||||
|
SNUT3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 121) | NC score | 0.052493 (rank : 85) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8WVK2, Q15410 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U4/U6.U5 tri-snRNP-associated protein 3 (U4/U6.U5 tri-snRNP-associated 27 kDa protein) (27K) (Nucleic acid-binding protein RY-1). | |||||
|
SRCH_HUMAN
|
||||||
| θ value | θ > 10 (rank : 122) | NC score | 0.075811 (rank : 35) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P23327, Q504Y6 | Gene names | HRC, HCP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcoplasmic reticulum histidine-rich calcium-binding protein precursor. | |||||
|
SRRM1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 123) | NC score | 0.073103 (rank : 40) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
SRRM1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 124) | NC score | 0.070796 (rank : 45) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 125) | NC score | 0.064891 (rank : 54) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 126) | NC score | 0.060302 (rank : 62) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
TARA_MOUSE
|
||||||
| θ value | θ > 10 (rank : 127) | NC score | 0.061500 (rank : 60) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1133 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q99KW3, Q2PZW9, Q2Q402, Q2Q403, Q2Q404, Q6ZPK4, Q8C6T3, Q8CG90 | Gene names | Triobp, Kiaa1662, Tara | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
THOC2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 128) | NC score | 0.053843 (rank : 80) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8NI27, Q9H8I6 | Gene names | THOC2, CXorf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | THO complex subunit 2 (Tho2). | |||||
|
TR150_MOUSE
|
||||||
| θ value | θ > 10 (rank : 129) | NC score | 0.053245 (rank : 83) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 468 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q569Z6 | Gene names | Thrap3, Trap150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid hormone receptor-associated protein 3 (Thyroid hormone receptor-associated protein complex 150 kDa component) (Trap150). | |||||
|
YTDC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 130) | NC score | 0.060289 (rank : 63) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q96MU7, Q7Z622, Q8TF35 | Gene names | YTHDC1, KIAA1966, YT521 | |||
|
Domain Architecture |

|
|||||
| Description | YTH domain-containing protein 1 (Putative splicing factor YT521). | |||||
|
SNIP1_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 3.82926e-184 (rank : 1) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | Q8TAD8, Q96SP9, Q9H9T7 | Gene names | SNIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Smad nuclear-interacting protein 1. | |||||
|
SNIP1_MOUSE
|
||||||
| NC score | 0.937560 (rank : 2) | θ value | 3.06238e-149 (rank : 2) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q8BIZ6, Q3V106, Q8BIZ4 | Gene names | Snip1 | |||
|
Domain Architecture |

|
|||||
| Description | Smad nuclear-interacting protein 1. | |||||
|
PP1R8_HUMAN
|
||||||
| NC score | 0.483403 (rank : 3) | θ value | 5.62301e-10 (rank : 3) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q12972, Q5TIF2, Q6PKF6, Q9UBH1, Q9UBZ0 | Gene names | PPP1R8, ARD1, NIPP1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear inhibitor of protein phosphatase 1 (NIPP-1) (Protein phosphatase 1 regulatory inhibitor subunit 8) [Includes: Activator of RNA decay (EC 3.1.4.-) (ARD-1)]. | |||||
|
PP1R8_MOUSE
|
||||||
| NC score | 0.483189 (rank : 4) | θ value | 5.62301e-10 (rank : 4) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8R3G1, Q8C087 | Gene names | Ppp1r8, Nipp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear inhibitor of protein phosphatase 1 (NIPP-1) (Protein phosphatase 1 regulatory inhibitor subunit 8). | |||||
|
NADAP_HUMAN
|
||||||
| NC score | 0.431715 (rank : 5) | θ value | 9.59137e-10 (rank : 5) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9BWU0, Q4KMT1, Q4KMX0, Q7Z5Q9, Q9NVN2 | Gene names | SLC4A1AP, HLC3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kanadaptin (Kidney anion exchanger adapter protein) (Solute carrier family 4 anion exchanger member 1 adapter protein) (Lung cancer oncogene 3 protein). | |||||
|
T2_MOUSE
|
||||||
| NC score | 0.213098 (rank : 6) | θ value | 0.0961366 (rank : 19) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q06666 | Gene names | Srst, T2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Octapeptide-repeat protein T2. | |||||
|
CCD55_HUMAN
|
||||||
| NC score | 0.198085 (rank : 7) | θ value | 0.00869519 (rank : 7) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9H0G5, Q6FI71 | Gene names | CCDC55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 55. | |||||
|
CCD55_MOUSE
|
||||||
| NC score | 0.191315 (rank : 8) | θ value | 0.0113563 (rank : 10) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q5NCR9 | Gene names | Ccdc55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 55. | |||||
|
RPTN_MOUSE
|
||||||
| NC score | 0.163743 (rank : 9) | θ value | 9.29e-05 (rank : 6) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P97347 | Gene names | Rptn | |||
|
Domain Architecture |

|
|||||
| Description | Repetin. | |||||
|
FILA_HUMAN
|
||||||
| NC score | 0.159460 (rank : 10) | θ value | 0.0736092 (rank : 17) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 685 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P20930, Q01720, Q5T583 | Gene names | FLG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filaggrin. | |||||
|
RPTN_HUMAN
|
||||||
| NC score | 0.153333 (rank : 11) | θ value | 0.0431538 (rank : 14) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q6XPR3 | Gene names | RPTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Repetin. | |||||
|
ZC313_HUMAN
|
||||||
| NC score | 0.145471 (rank : 12) | θ value | 1.81305 (rank : 49) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q5T200, O94936, Q5T1Z9, Q7Z7J3, Q8NDT6, Q9H0L6 | Gene names | ZC3H13, KIAA0853 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 13. | |||||
|
NIPBL_HUMAN
|
||||||
| NC score | 0.143635 (rank : 13) | θ value | 0.0113563 (rank : 11) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
SFR12_HUMAN
|
||||||
| NC score | 0.141250 (rank : 14) | θ value | 2.36792 (rank : 54) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 553 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8WXA9, Q86X37 | Gene names | SFRS12, SRRP86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86) (Splicing regulatory protein 508) (SRrp508). | |||||
|
NPBL_MOUSE
|
||||||
| NC score | 0.138979 (rank : 15) | θ value | 0.125558 (rank : 22) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
SFR12_MOUSE
|
||||||
| NC score | 0.127277 (rank : 16) | θ value | 0.813845 (rank : 32) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8BZX4, Q8BLM8, Q8BWK7, Q8BYK0, Q8BYZ4 | Gene names | Sfrs12, Srrp86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86). | |||||
|
SFR11_HUMAN
|
||||||
| NC score | 0.114529 (rank : 17) | θ value | θ > 10 (rank : 115) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q05519 | Gene names | SFRS11 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor arginine/serine-rich 11 (Arginine-rich 54 kDa nuclear protein) (p54). | |||||
|
SFRIP_HUMAN
|
||||||
| NC score | 0.113926 (rank : 18) | θ value | 0.62314 (rank : 27) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 530 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q99590, Q8IW59 | Gene names | SFRS2IP, CASP11, SIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SFRS2-interacting protein (Splicing factor, arginine/serine-rich 2- interacting protein) (SC35-interacting protein 1) (CTD-associated SR protein 11) (Splicing regulatory protein 129) (SRrp129) (NY-REN-40 antigen). | |||||
|
CROP_HUMAN
|
||||||
| NC score | 0.109250 (rank : 19) | θ value | 1.06291 (rank : 35) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O95232, Q6PHR9, Q9NUY0, Q9P2S7 | Gene names | CROP, CREAP1, O48 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cisplatin resistance-associated overexpressed protein (cAMP regulatory element-associated protein 1) (CRE-associated protein 1) (CREAP-1) (Luc7A) (Okadaic acid-inducible phosphoprotein OA48-18). | |||||
|
CROP_MOUSE
|
||||||
| NC score | 0.108895 (rank : 20) | θ value | 1.81305 (rank : 44) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q5SUF2, Q3U9D5, Q8BUJ5, Q921Z3, Q9CRS7, Q9CTY4 | Gene names | Crop | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cisplatin resistance-associated overexpressed protein. | |||||
|
PPIG_HUMAN
|
||||||
| NC score | 0.108800 (rank : 21) | θ value | 0.0148317 (rank : 12) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q13427, O00706, Q96DG9 | Gene names | PPIG | |||
|
Domain Architecture |

|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase G (EC 5.2.1.8) (Peptidyl-prolyl isomerase G) (PPIase G) (Rotamase G) (Cyclophilin G) (Clk-associating RS-cyclophilin) (CARS-cyclophilin) (CARS-Cyp) (SR-cyclophilin) (SRcyp) (SR-cyp) (CASP10). | |||||
|
GPTC8_HUMAN
|
||||||
| NC score | 0.100281 (rank : 22) | θ value | θ > 10 (rank : 94) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 366 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9UKJ3, O60300 | Gene names | GPATC8, KIAA0553 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 8. | |||||
|
TARA_HUMAN
|
||||||
| NC score | 0.095332 (rank : 23) | θ value | 0.00869519 (rank : 9) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9H2D6, O94797, Q2PZW8, Q2Q3Z9, Q2Q400, Q5R3M6, Q96DW1, Q9BT77, Q9BTL7, Q9BY98, Q9Y3L4 | Gene names | TRIOBP, KIAA1662, TARA | |||
|
Domain Architecture |

|
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
RBBP6_MOUSE
|
||||||
| NC score | 0.089814 (rank : 24) | θ value | 0.62314 (rank : 26) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
RBBP6_HUMAN
|
||||||
| NC score | 0.089100 (rank : 25) | θ value | 8.99809 (rank : 79) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
HORN_HUMAN
|
||||||
| NC score | 0.088805 (rank : 26) | θ value | θ > 10 (rank : 96) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q86YZ3 | Gene names | HRNR, S100A18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hornerin. | |||||
|
SFR15_HUMAN
|
||||||
| NC score | 0.086998 (rank : 27) | θ value | θ > 10 (rank : 116) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O95104, Q6P1M5, Q8N3I8, Q9UFM1, Q9ULP8 | Gene names | SFRS15, KIAA1172 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 15 (CTD-binding SR-like protein RA4). | |||||
|
SFR17_HUMAN
|
||||||
| NC score | 0.085239 (rank : 28) | θ value | θ > 10 (rank : 118) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8TF01, Q5T064, Q6P2B4, Q6P2N4, Q6PJ93, Q6PK36, Q7Z640, Q8TF00, Q96K10, Q96SM5, Q9P076, Q9P0C0, Q9Y4N3 | Gene names | SRRP130, C6orf111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 130 (Serine-arginine-rich- splicing regulatory protein 130) (SRrp130) (SR-rich protein) (SR- related protein). | |||||
|
LEO1_HUMAN
|
||||||
| NC score | 0.084239 (rank : 29) | θ value | 8.99809 (rank : 76) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
HORN_MOUSE
|
||||||
| NC score | 0.082113 (rank : 30) | θ value | θ > 10 (rank : 97) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8VHD8 | Gene names | Hrnr | |||
|
Domain Architecture |

|
|||||
| Description | Hornerin. | |||||
|
DMP1_HUMAN
|
||||||
| NC score | 0.081710 (rank : 31) | θ value | θ > 10 (rank : 89) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q13316, O43265 | Gene names | DMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1). | |||||
|
OTUD3_HUMAN
|
||||||
| NC score | 0.078507 (rank : 32) | θ value | 0.365318 (rank : 25) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5T2D3, O75047 | Gene names | OTUD3, KIAA0459 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | OTU domain-containing protein 3. | |||||
|
LEO1_MOUSE
|
||||||
| NC score | 0.077959 (rank : 33) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
DSPP_MOUSE
|
||||||
| NC score | 0.075955 (rank : 34) | θ value | θ > 10 (rank : 92) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
SRCH_HUMAN
|
||||||
| NC score | 0.075811 (rank : 35) | θ value | θ > 10 (rank : 122) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P23327, Q504Y6 | Gene names | HRC, HCP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcoplasmic reticulum histidine-rich calcium-binding protein precursor. | |||||
|
RU17_HUMAN
|
||||||
| NC score | 0.075005 (rank : 36) | θ value | 6.88961 (rank : 70) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P08621, P78493, P78494, Q15364, Q15686, Q15687, Q15689, Q99377, Q9UE45, Q9UE46, Q9UE47, Q9UE48, Q9UFQ6 | Gene names | SNRP70, RPU1, U1AP1 | |||
|
Domain Architecture |

|
|||||
| Description | U1 small nuclear ribonucleoprotein 70 kDa (U1 snRNP 70 kDa) (snRNP70) (U1-70K). | |||||
|
DMP1_MOUSE
|
||||||
| NC score | 0.074410 (rank : 37) | θ value | θ > 10 (rank : 90) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O55188 | Gene names | Dmp1, Dmp | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1) (AG1). | |||||
|
NKTR_HUMAN
|
||||||
| NC score | 0.073183 (rank : 38) | θ value | 8.99809 (rank : 78) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 658 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P30414 | Gene names | NKTR | |||
|
Domain Architecture |

|
|||||
| Description | NK-tumor recognition protein (Natural-killer cells cyclophilin-related protein) (NK-TR protein). | |||||
|
ACINU_HUMAN
|
||||||
| NC score | 0.073149 (rank : 39) | θ value | 6.88961 (rank : 68) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9UKV3, O75158, Q9UG91, Q9UKV1, Q9UKV2 | Gene names | ACIN1, ACINUS, KIAA0670 | |||
|
Domain Architecture |

|
|||||
| Description | Apoptotic chromatin condensation inducer in the nucleus (Acinus). | |||||
|
SRRM1_HUMAN
|
||||||
| NC score | 0.073103 (rank : 40) | θ value | θ > 10 (rank : 123) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
CIR_HUMAN
|
||||||
| NC score | 0.073031 (rank : 41) | θ value | θ > 10 (rank : 85) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q86X95, O95367, Q12804, Q4G1B9, Q6PJI4, Q8IWI2 | Gene names | CIR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CBF1-interacting corepressor (Recepin). | |||||
|
NUCKS_MOUSE
|
||||||
| NC score | 0.072396 (rank : 42) | θ value | θ > 10 (rank : 108) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q80XU3, Q8BVD8 | Gene names | Nucks1, Nucks | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (JC7). | |||||
|
TGON2_HUMAN
|
||||||
| NC score | 0.072082 (rank : 43) | θ value | 0.163984 (rank : 24) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.071725 (rank : 44) | θ value | θ > 10 (rank : 98) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
SRRM1_MOUSE
|
||||||
| NC score | 0.070796 (rank : 45) | θ value | θ > 10 (rank : 124) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
RU17_MOUSE
|
||||||
| NC score | 0.068534 (rank : 46) | θ value | 0.813845 (rank : 31) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q62376, Q3UIW4 | Gene names | Snrp70 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U1 small nuclear ribonucleoprotein 70 kDa (U1 SNRNP 70 kDa) (snRNP70). | |||||
|
SFRS4_MOUSE
|
||||||
| NC score | 0.068423 (rank : 47) | θ value | θ > 10 (rank : 120) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8VE97, Q9JJC3 | Gene names | Sfrs4 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 4. | |||||
|
DSPP_HUMAN
|
||||||
| NC score | 0.068315 (rank : 48) | θ value | θ > 10 (rank : 91) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
SFRS4_HUMAN
|
||||||
| NC score | 0.066679 (rank : 49) | θ value | θ > 10 (rank : 119) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q08170, Q5VXP1, Q9BUA4, Q9UEB5 | Gene names | SFRS4, SRP75 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 4 (Pre-mRNA-splicing factor SRP75) (SRP001LB). | |||||
|
CHD1_HUMAN
|
||||||
| NC score | 0.066354 (rank : 50) | θ value | 0.125558 (rank : 21) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O14646 | Gene names | CHD1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
ELOA1_HUMAN
|
||||||
| NC score | 0.065725 (rank : 51) | θ value | 1.81305 (rank : 45) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q14241, Q8IXH1 | Gene names | TCEB3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription elongation factor B polypeptide 3 (RNA polymerase II transcription factor SIII subunit A1) (SIII p110) (Elongin-A) (EloA) (Elongin 110 kDa subunit). | |||||
|
SCG1_HUMAN
|
||||||
| NC score | 0.065297 (rank : 52) | θ value | 8.99809 (rank : 80) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P05060, Q9BQV6, Q9UJA6 | Gene names | CHGB, SCG1 | |||
|
Domain Architecture |

|
|||||
| Description | Secretogranin-1 precursor (Secretogranin I) (SgI) (Chromogranin B) (CgB) [Contains: GAWK peptide; CCB peptide]. | |||||
|
LC7L2_MOUSE
|
||||||
| NC score | 0.065213 (rank : 53) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 244 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q7TNC4, Q99LM5, Q99PC3 | Gene names | Luc7l2 | |||
|
Domain Architecture |

|
|||||
| Description | Putative RNA-binding protein Luc7-like 2 (CGI-74 homolog). | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.064891 (rank : 54) | θ value | θ > 10 (rank : 125) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
C8AP2_HUMAN
|
||||||
| NC score | 0.064747 (rank : 55) | θ value | 5.27518 (rank : 60) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 548 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9UKL3, Q6PH76, Q7LCQ7, Q86YD9, Q9NUQ4, Q9NZV9, Q9P2N1, Q9Y563 | Gene names | CASP8AP2, FLASH, KIAA1315, RIP25 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.063822 (rank : 56) | θ value | θ > 10 (rank : 99) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
CHD1_MOUSE
|
||||||
| NC score | 0.063206 (rank : 57) | θ value | 0.0563607 (rank : 15) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P40201 | Gene names | Chd1, Chd-1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
WDR60_HUMAN
|
||||||
| NC score | 0.062556 (rank : 58) | θ value | 6.88961 (rank : 71) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8WVS4, Q9NW58 | Gene names | WDR60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 60. | |||||
|
LC7L2_HUMAN
|
||||||
| NC score | 0.061574 (rank : 59) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9Y383, Q8IUP9, Q9NVL3, Q9NVN7, Q9UQN1 | Gene names | LUC7L2 | |||
|
Domain Architecture |

|
|||||
| Description | Putative RNA-binding protein Luc7-like 2. | |||||
|
TARA_MOUSE
|
||||||
| NC score | 0.061500 (rank : 60) | θ value | θ > 10 (rank : 127) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1133 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q99KW3, Q2PZW9, Q2Q402, Q2Q403, Q2Q404, Q6ZPK4, Q8C6T3, Q8CG90 | Gene names | Triobp, Kiaa1662, Tara | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
CIR_MOUSE
|
||||||
| NC score | 0.060360 (rank : 61) | θ value | θ > 10 (rank : 86) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9DA19, Q3V2N9, Q4KL44, Q52KL9, Q5FW66 | Gene names | Cir | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CBF1-interacting corepressor. | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.060302 (rank : 62) | θ value | θ > 10 (rank : 126) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
YTDC1_HUMAN
|
||||||
| NC score | 0.060289 (rank : 63) | θ value | θ > 10 (rank : 130) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q96MU7, Q7Z622, Q8TF35 | Gene names | YTHDC1, KIAA1966, YT521 | |||
|
Domain Architecture |

|
|||||
| Description | YTH domain-containing protein 1 (Putative splicing factor YT521). | |||||
|
NUCKS_HUMAN
|
||||||
| NC score | 0.060027 (rank : 64) | θ value | θ > 10 (rank : 107) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9H1E3, Q9H1D6, Q9H723 | Gene names | NUCKS1, NUCKS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (P1). | |||||
|
HIRP3_HUMAN
|
||||||
| NC score | 0.059007 (rank : 65) | θ value | θ > 10 (rank : 95) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9BW71, O75707, O75708 | Gene names | HIRIP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIRA-interacting protein 3. | |||||
|
ACINU_MOUSE
|
||||||
| NC score | 0.058198 (rank : 66) | θ value | θ > 10 (rank : 81) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 712 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9JIX8, Q9CSN7, Q9CSR9, Q9CSX7, Q9R046, Q9R047 | Gene names | Acin1, Acinus | |||
|
Domain Architecture |

|
|||||
| Description | Apoptotic chromatin condensation inducer in the nucleus (Acinus). | |||||
|
NSBP1_HUMAN
|
||||||
| NC score | 0.057944 (rank : 67) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P82970 | Gene names | NSBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1. | |||||
|
CK035_MOUSE
|
||||||
| NC score | 0.057722 (rank : 68) | θ value | 1.38821 (rank : 39) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q0VET5, Q9CW10 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C11orf35 homolog. | |||||
|
ENL_HUMAN
|
||||||
| NC score | 0.057460 (rank : 69) | θ value | θ > 10 (rank : 93) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q03111, Q14768 | Gene names | MLLT1, ENL, LTG19, YEATS1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein ENL (YEATS domain-containing protein 1). | |||||
|
NKTR_MOUSE
|
||||||
| NC score | 0.056957 (rank : 70) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P30415 | Gene names | Nktr | |||
|
Domain Architecture |

|
|||||
| Description | NK-tumor recognition protein (Natural-killer cells cyclophilin-related protein) (NK-TR protein). | |||||
|
RBM25_HUMAN
|
||||||
| NC score | 0.056792 (rank : 71) | θ value | θ > 10 (rank : 110) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P49756, Q9UEQ5, Q9UIE9 | Gene names | RBM25, RNPC7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable RNA-binding protein 25 (RNA-binding motif protein 25) (RNA- binding region-containing protein 7) (Protein S164). | |||||
|
SPRL1_MOUSE
|
||||||
| NC score | 0.056761 (rank : 72) | θ value | 0.62314 (rank : 28) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P70663, P97810, Q99L82 | Gene names | Sparcl1, Ecm2, Sc1 | |||
|
Domain Architecture |

|
|||||
| Description | SPARC-like protein 1 precursor (Matrix glycoprotein Sc1) (Extracellular matrix protein 2). | |||||
|
TRHY_HUMAN
|
||||||
| NC score | 0.056423 (rank : 73) | θ value | 0.62314 (rank : 29) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1385 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q07283 | Gene names | THH, THL, TRHY | |||
|
Domain Architecture |

|
|||||
| Description | Trichohyalin. | |||||
|
RSRC1_MOUSE
|
||||||
| NC score | 0.056032 (rank : 74) | θ value | θ > 10 (rank : 112) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9DBU6, Q3TF06 | Gene names | Rsrc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine/serine-rich coiled coil protein 1. | |||||
|
RSRC1_HUMAN
|
||||||
| NC score | 0.055343 (rank : 75) | θ value | θ > 10 (rank : 111) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q96IZ7, Q96QK2, Q9NZE5 | Gene names | RSRC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine/serine-rich coiled coil protein 1. | |||||
|
AFF4_HUMAN
|
||||||
| NC score | 0.054806 (rank : 76) | θ value | θ > 10 (rank : 82) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 611 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9UHB7, Q498B2, Q59FB3, Q6P592, Q8TDR1, Q9P0E4 | Gene names | AFF4, AF5Q31, MCEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4 (ALL1-fused gene from chromosome 5q31) (Major CDK9 elongation factor-associated protein). | |||||
|
NOLC1_HUMAN
|
||||||
| NC score | 0.054445 (rank : 77) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q14978, Q15030 | Gene names | NOLC1, KIAA0035 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar phosphoprotein p130 (Nucleolar 130 kDa protein) (140 kDa nucleolar phosphoprotein) (Nopp140) (Nucleolar and coiled-body phosphoprotein 1). | |||||
|
SEMG2_HUMAN
|
||||||
| NC score | 0.054347 (rank : 78) | θ value | θ > 10 (rank : 114) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q02383, Q6X2M6 | Gene names | SEMG2 | |||
|
Domain Architecture |

|
|||||
| Description | Semenogelin-2 precursor (Semenogelin II) (SGII). | |||||
|
PHF20_MOUSE
|
||||||
| NC score | 0.053955 (rank : 79) | θ value | 5.27518 (rank : 63) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8BLG0, Q8BMA2, Q8BYR4, Q921N1 | Gene names | Phf20, Hca58 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58 homolog). | |||||
|
THOC2_HUMAN
|
||||||
| NC score | 0.053843 (rank : 80) | θ value | θ > 10 (rank : 128) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8NI27, Q9H8I6 | Gene names | THOC2, CXorf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | THO complex subunit 2 (Tho2). | |||||
|
IF2P_HUMAN
|
||||||
| NC score | 0.053820 (rank : 81) | θ value | 8.99809 (rank : 75) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 889 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O60841, O95805, Q9UF81, Q9UMN7 | Gene names | EIF5B, IF2, KIAA0741 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 5B (eIF-5B) (Translation initiation factor IF-2). | |||||
|
C8AP2_MOUSE
|
||||||
| NC score | 0.053440 (rank : 82) | θ value | θ > 10 (rank : 84) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 405 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9WUF3, Q9CSW2 | Gene names | Casp8ap2, Flash | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
TR150_MOUSE
|
||||||
| NC score | 0.053245 (rank : 83) | θ value | θ > 10 (rank : 129) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 468 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q569Z6 | Gene names | Thrap3, Trap150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid hormone receptor-associated protein 3 (Thyroid hormone receptor-associated protein complex 150 kDa component) (Trap150). | |||||
|
SCG1_MOUSE
|
||||||
| NC score | 0.052761 (rank : 84) | θ value | θ > 10 (rank : 113) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P16014 | Gene names | Chgb, Scg-1, Scg1 | |||
|
Domain Architecture |

|
|||||
| Description | Secretogranin-1 precursor (Secretogranin I) (SgI) (Chromogranin B) (CgB). | |||||
|
SNUT3_HUMAN
|
||||||
| NC score | 0.052493 (rank : 85) | θ value | θ > 10 (rank : 121) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8WVK2, Q15410 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U4/U6.U5 tri-snRNP-associated protein 3 (U4/U6.U5 tri-snRNP-associated 27 kDa protein) (27K) (Nucleic acid-binding protein RY-1). | |||||
|
AFF4_MOUSE
|
||||||
| NC score | 0.051991 (rank : 86) | θ value | θ > 10 (rank : 83) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9ESC8, Q8C6K4, Q8C6W3, Q8CCH3 | Gene names | Aff4, Alf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4. | |||||
|
CYLC2_HUMAN
|
||||||
| NC score | 0.051663 (rank : 87) | θ value | θ > 10 (rank : 88) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q14093 | Gene names | CYLC2, CYL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-2 (Cylicin II) (Multiple-band polypeptide II). | |||||
|
RPGF1_HUMAN
|
||||||
| NC score | 0.050889 (rank : 88) | θ value | 0.0961366 (rank : 18) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13905 | Gene names | RAPGEF1, GRF2 | |||
|
Domain Architecture |

|
|||||
| Description | Rap guanine nucleotide exchange factor 1 (Guanine nucleotide-releasing factor 2) (C3G protein) (CRK SH3-binding GNRP). | |||||
|
SFR16_HUMAN
|
||||||
| NC score | 0.050821 (rank : 89) | θ value | θ > 10 (rank : 117) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8N2M8, O96026, Q6UW71, Q96DX2 | Gene names | SFRS16, SWAP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 16 (Suppressor of white-apricot homolog 2). | |||||
|
NGEF_MOUSE
|
||||||
| NC score | 0.050560 (rank : 90) | θ value | 0.00869519 (rank : 8) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8CHT1, Q8R204, Q923H2, Q9JHT9 | Gene names | Ngef | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ephexin-1 (Eph-interacting exchange protein) (Neuronal guanine nucleotide exchange factor). | |||||
|
PININ_MOUSE
|
||||||
| NC score | 0.050458 (rank : 91) | θ value | θ > 10 (rank : 109) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 777 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O35691, Q8CD89, Q8CGU3 | Gene names | Pnn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pinin. | |||||
|
NSBP1_MOUSE
|
||||||
| NC score | 0.050234 (rank : 92) | θ value | θ > 10 (rank : 106) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
CYLC1_HUMAN
|
||||||
| NC score | 0.050042 (rank : 93) | θ value | θ > 10 (rank : 87) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P35663, Q5JQQ9 | Gene names | CYLC1, CYL, CYL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-1 (Cylicin I) (Multiple-band polypeptide I). | |||||
|
CBX8_HUMAN
|
||||||
| NC score | 0.049806 (rank : 94) | θ value | 3.0926 (rank : 55) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9HC52, Q96H39, Q9NR07 | Gene names | CBX8, PC3, RC1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 8 (Polycomb 3 homolog) (Pc3) (hPc3) (Rectachrome 1). | |||||
|
US6NL_MOUSE
|
||||||
| NC score | 0.048955 (rank : 95) | θ value | 1.06291 (rank : 37) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q80XC3, Q6ZQK8 | Gene names | Usp6nl, Kiaa0019 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USP6 N-terminal-like protein. | |||||
|
US6NL_HUMAN
|
||||||
| NC score | 0.048679 (rank : 96) | θ value | 1.38821 (rank : 43) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q92738, Q15400, Q7L0K9 | Gene names | USP6NL, KIAA0019 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USP6 N-terminal-like protein (Related to the N-terminus of tre) (RN- tre). | |||||
|
BSND_MOUSE
|
||||||
| NC score | 0.046438 (rank : 97) | θ value | 1.38821 (rank : 38) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8VIM4, Q8C740, Q8CHY0 | Gene names | Bsnd | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Barttin. | |||||
|
PPHLN_HUMAN
|
||||||
| NC score | 0.046412 (rank : 98) | θ value | 5.27518 (rank : 64) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8NEY8, Q86YT2, Q8IXN3, Q8TB09, Q96NB9, Q9NXL4, Q9P0P6, Q9P0R9 | Gene names | PPHLN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Periphilin-1 (Gastric cancer antigen Ga50). | |||||
|
INCE_HUMAN
|
||||||
| NC score | 0.044829 (rank : 99) | θ value | 4.03905 (rank : 59) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 978 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9NQS7, Q5Y192 | Gene names | INCENP | |||
|
Domain Architecture |

|
|||||
| Description | Inner centromere protein. | |||||
|
MDC1_MOUSE
|
||||||
| NC score | 0.043209 (rank : 100) | θ value | 2.36792 (rank : 53) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
SRY_MOUSE
|
||||||
| NC score | 0.043122 (rank : 101) | θ value | 1.38821 (rank : 42) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q05738 | Gene names | Sry, Tdf, Tdy | |||
|
Domain Architecture |

|
|||||
| Description | Sex-determining region Y protein (Testis-determining factor). | |||||
|
LA_MOUSE
|
||||||
| NC score | 0.041985 (rank : 102) | θ value | 1.38821 (rank : 41) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P32067 | Gene names | Ssb, Ss-b | |||
|
Domain Architecture |

|
|||||
| Description | Lupus La protein homolog (La ribonucleoprotein) (La autoantigen homolog). | |||||
|
RB15B_MOUSE
|
||||||
| NC score | 0.039783 (rank : 103) | θ value | 5.27518 (rank : 65) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 284 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6PHZ5, Q8C6G2 | Gene names | Rbm15b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative RNA-binding protein 15B (RNA-binding motif protein 15B). | |||||
|
RIMS2_MOUSE
|
||||||
| NC score | 0.038908 (rank : 104) | θ value | 1.81305 (rank : 47) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 376 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9EQZ7, Q8C433, Q8CCK2 | Gene names | Rims2, Rab3ip2, Rim2 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 2 (Rab3-interacting molecule 2) (RIM 2) (Rab3-interacting protein 2). | |||||
|
DBNL_HUMAN
|
||||||
| NC score | 0.038750 (rank : 105) | θ value | 1.38821 (rank : 40) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 244 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UJU6, P84070, Q6IAI8, Q96F30, Q96K74, Q9HBN8, Q9NR72 | Gene names | DBNL, CMAP, SH3P7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Drebrin-like protein (SH3 domain-containing protein 7) (Drebrin F) (Cervical SH3P7) (HPK1-interacting protein of 55 kDa) (HIP-55) (Cervical mucin-associated protein). | |||||
|
RP3A_MOUSE
|
||||||
| NC score | 0.036814 (rank : 106) | θ value | 0.0431538 (rank : 13) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 510 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P47708 | Gene names | Rph3a | |||
|
Domain Architecture |

|
|||||
| Description | Rabphilin-3A (Exophilin-1). | |||||
|
ERCC6_HUMAN
|
||||||
| NC score | 0.034717 (rank : 107) | θ value | 1.81305 (rank : 46) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q03468 | Gene names | ERCC6, CSB | |||
|
Domain Architecture |

|
|||||
| Description | DNA excision repair protein ERCC-6 (EC 3.6.1.-) (ATP-dependent helicase ERCC6) (Cockayne syndrome protein CSB). | |||||
|
RNF8_HUMAN
|
||||||
| NC score | 0.033832 (rank : 108) | θ value | 1.81305 (rank : 48) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 622 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O76064 | Gene names | RNF8, KIAA0646 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin ligase protein RNF8 (EC 6.3.2.-) (RING finger protein 8). | |||||
|
DDX46_MOUSE
|
||||||
| NC score | 0.032365 (rank : 109) | θ value | 2.36792 (rank : 50) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 358 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q569Z5, Q6ZQ42, Q8R0R6 | Gene names | Ddx46, Kiaa0801 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX46 (EC 3.6.1.-) (DEAD box protein 46). | |||||
|
ANK2_HUMAN
|
||||||
| NC score | 0.031093 (rank : 110) | θ value | 0.125558 (rank : 20) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
CAC1A_HUMAN
|
||||||
| NC score | 0.030855 (rank : 111) | θ value | 1.06291 (rank : 34) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O00555, P78510, P78511, Q16290, Q92690, Q99790, Q99791, Q99792, Q99793 | Gene names | CACNA1A, CACH4, CACN3, CACNL1A4 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent P/Q-type calcium channel subunit alpha-1A (Voltage- gated calcium channel subunit alpha Cav2.1) (Calcium channel, L type, alpha-1 polypeptide isoform 4) (Brain calcium channel I) (BI). | |||||
|
GASP1_MOUSE
|
||||||
| NC score | 0.027697 (rank : 112) | θ value | 4.03905 (rank : 58) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q5U4C1, Q8BKR8, Q8BUN4, Q8BYK9, Q8CHF4, Q8R095 | Gene names | Gprasp1, Kiaa0443 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G-protein coupled receptor-associated sorting protein 1 (GASP-1). | |||||
|
P52K_HUMAN
|
||||||
| NC score | 0.027282 (rank : 113) | θ value | 3.0926 (rank : 57) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O43422, Q8WTW1, Q9Y3Z4 | Gene names | PRKRIR, DAP4, P52RIPK, THAP0 | |||
|
Domain Architecture |

|
|||||
| Description | 52 kDa repressor of the inhibitor of the protein kinase (p58IPK- interacting protein) (58 kDa interferon-induced protein kinase- interacting protein) (P52rIPK) (Death-associated protein 4) (THAP domain-containing protein 0). | |||||
|
NCOA6_MOUSE
|
||||||
| NC score | 0.026417 (rank : 114) | θ value | 6.88961 (rank : 69) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1076 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9JL19, Q9JLT9 | Gene names | Ncoa6, Aib3, Prip, Rap250, Trbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC) (Thyroid hormone receptor-binding protein). | |||||
|
DBNL_MOUSE
|
||||||
| NC score | 0.024510 (rank : 115) | θ value | 8.99809 (rank : 73) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 306 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q62418, Q80WP1, Q8BH56 | Gene names | Dbnl, Abp1, Sh3p7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Drebrin-like protein (SH3 domain-containing protein 7) (Actin-binding protein 1). | |||||
|
DVL1_MOUSE
|
||||||
| NC score | 0.023580 (rank : 116) | θ value | 0.813845 (rank : 30) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P51141, Q60868 | Gene names | Dvl1, Dvl | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1 (Dishevelled-1) (DSH homolog 1). | |||||
|
NCOA6_HUMAN
|
||||||
| NC score | 0.021383 (rank : 117) | θ value | 8.99809 (rank : 77) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1202 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q14686, Q9NTZ9, Q9UH74, Q9UK86 | Gene names | NCOA6, AIB3, KIAA0181, RAP250, TRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (PRIP) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC RAP250) (Thyroid hormone receptor-binding protein). | |||||
|
CD2L5_HUMAN
|
||||||
| NC score | 0.021149 (rank : 118) | θ value | 0.0736092 (rank : 16) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 987 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q14004, Q6DKQ9, Q75MH4, Q75MH5, Q96JN4, Q9H4A0, Q9H4A1, Q9UDR4 | Gene names | CDC2L5, CDC2L, CHED, KIAA1791 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle 2-like protein kinase 5 (EC 2.7.11.22) (CDC2- related protein kinase 5) (Cholinesterase-related cell division controller). | |||||
|
FBXW7_HUMAN
|
||||||
| NC score | 0.020386 (rank : 119) | θ value | 0.163984 (rank : 23) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q969H0, Q96A16, Q96LE0, Q96RI2, Q9NUX6 | Gene names | FBXW7, FBW7, FBX30, SEL10 | |||
|
Domain Architecture |

|
|||||
| Description | F-box/WD repeat protein 7 (F-box and WD-40 domain protein 7) (F-box protein FBX30) (hCdc4) (Archipelago homolog) (hAgo) (SEL-10). | |||||
|
UTY_HUMAN
|
||||||
| NC score | 0.017378 (rank : 120) | θ value | 5.27518 (rank : 66) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O14607, O14608 | Gene names | UTY | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitously transcribed Y chromosome tetratricopeptide repeat protein (Ubiquitously transcribed TPR protein on the Y chromosome). | |||||
|
CD2L5_MOUSE
|
||||||
| NC score | 0.016851 (rank : 121) | θ value | 5.27518 (rank : 61) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1011 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q69ZA1, Q80V11, Q8BZG1, Q8K0A4 | Gene names | Cdc2l5, Kiaa1791 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle 2-like protein kinase 5 (EC 2.7.11.22) (CDC2- related protein kinase 5). | |||||
|
ZNFX1_HUMAN
|
||||||
| NC score | 0.015331 (rank : 122) | θ value | 5.27518 (rank : 67) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9P2E3, Q9BQM7, Q9BQM8, Q9H8C1, Q9H9S2, Q9NUM1, Q9NWW1 | Gene names | ZNFX1, KIAA1404 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NFX1-type zinc finger-containing protein 1. | |||||
|
MRCKB_HUMAN
|
||||||
| NC score | 0.015298 (rank : 123) | θ value | 1.06291 (rank : 36) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 2239 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9Y5S2, Q2L7A5, Q86TJ1, Q9ULU5 | Gene names | CDC42BPB, KIAA1124 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK beta (EC 2.7.11.1) (CDC42-binding protein kinase beta) (Myotonic dystrophy kinase-related CDC42-binding kinase beta) (Myotonic dystrophy protein kinase-like beta) (MRCK beta) (DMPK-like beta). | |||||
|
KIF3B_MOUSE
|
||||||
| NC score | 0.013960 (rank : 124) | θ value | 2.36792 (rank : 52) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 533 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q61771 | Gene names | Kif3b | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF3B (Microtubule plus end-directed kinesin motor 3B). | |||||
|
KIF3B_HUMAN
|
||||||
| NC score | 0.013900 (rank : 125) | θ value | 2.36792 (rank : 51) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 540 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O15066 | Gene names | KIF3B, KIAA0359 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF3B (Microtubule plus end-directed kinesin motor 3B) (HH0048). | |||||
|
MRCKB_MOUSE
|
||||||
| NC score | 0.013655 (rank : 126) | θ value | 3.0926 (rank : 56) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 2180 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q7TT50, Q69ZR5, Q80W33 | Gene names | Cdc42bpb, Kiaa1124 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK beta (EC 2.7.11.1) (CDC42-binding protein kinase beta) (Myotonic dystrophy kinase-related CDC42-binding kinase beta) (Myotonic dystrophy protein kinase-like beta) (MRCK beta) (DMPK-like beta). | |||||
|
HES6_MOUSE
|
||||||
| NC score | 0.013481 (rank : 127) | θ value | 8.99809 (rank : 74) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JHE6, Q3U2D7 | Gene names | Hes6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription cofactor HES-6 (Hairy and enhancer of split 6). | |||||
|
CAC1F_HUMAN
|
||||||
| NC score | 0.010809 (rank : 128) | θ value | 8.99809 (rank : 72) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O60840, O43901 | Gene names | CACNA1F, CACNAF1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1F (Voltage- gated calcium channel subunit alpha Cav1.4). | |||||
|
MK07_HUMAN
|
||||||
| NC score | 0.007221 (rank : 129) | θ value | 5.27518 (rank : 62) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 1087 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q13164, Q16634 | Gene names | MAPK7, ERK4, ERK5, PRKM7 | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase 7 (EC 2.7.11.24) (Extracellular signal-regulated kinase 5) (ERK-5) (ERK4) (BMK1 kinase). | |||||
|
ZKSC1_HUMAN
|
||||||
| NC score | 0.003666 (rank : 130) | θ value | 0.813845 (rank : 33) | |||
| Query Neighborhood Hits | 80 | Target Neighborhood Hits | 949 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P17029, P52745, Q8TBW5, Q8TEK7 | Gene names | ZKSCAN1, KOX18, ZNF139, ZNF36 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger with KRAB and SCAN domain-containing protein 1 (Zinc finger protein 36) (Zinc finger protein KOX18). | |||||