Please be patient as the page loads
|
AXN2_MOUSE
|
||||||
| SwissProt Accessions | O88566, Q9QXJ6 | Gene names | Axin2 | |||
|
Domain Architecture |

|
|||||
| Description | Axin-2 (Axis inhibition protein 2) (Conductin) (Axin-like protein) (Axil). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
AXN2_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.989516 (rank : 2) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9Y2T1, Q9H3M6, Q9UH84 | Gene names | AXIN2 | |||
|
Domain Architecture |

|
|||||
| Description | Axin-2 (Axis inhibition protein 2) (Conductin) (Axin-like protein) (Axil). | |||||
|
AXN2_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 116 | |
| SwissProt Accessions | O88566, Q9QXJ6 | Gene names | Axin2 | |||
|
Domain Architecture |

|
|||||
| Description | Axin-2 (Axis inhibition protein 2) (Conductin) (Axin-like protein) (Axil). | |||||
|
AXN1_MOUSE
|
||||||
| θ value | 3.7435e-147 (rank : 3) | NC score | 0.948949 (rank : 3) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O35625 | Gene names | Axin1, Axin, Fu | |||
|
Domain Architecture |

|
|||||
| Description | Axin-1 (Axis inhibition protein 1) (Fused protein). | |||||
|
AXN1_HUMAN
|
||||||
| θ value | 2.77624e-142 (rank : 4) | NC score | 0.948670 (rank : 4) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O15169, Q4TT26, Q4TT27, Q86YA7, Q8WVW6, Q96S28 | Gene names | AXIN1, AXIN | |||
|
Domain Architecture |

|
|||||
| Description | Axin-1 (Axis inhibition protein 1) (hAxin). | |||||
|
RGS18_HUMAN
|
||||||
| θ value | 1.68911e-14 (rank : 5) | NC score | 0.677020 (rank : 6) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9NS28 | Gene names | RGS18, RGS13 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 18 (RGS18). | |||||
|
RGS18_MOUSE
|
||||||
| θ value | 1.09485e-13 (rank : 6) | NC score | 0.675483 (rank : 7) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q99PG4 | Gene names | Rgs18 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 18 (RGS18). | |||||
|
RGS3_HUMAN
|
||||||
| θ value | 4.16044e-13 (rank : 7) | NC score | 0.489779 (rank : 37) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
RGS3_MOUSE
|
||||||
| θ value | 1.02475e-11 (rank : 8) | NC score | 0.432395 (rank : 40) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
RGS2_HUMAN
|
||||||
| θ value | 2.98157e-11 (rank : 9) | NC score | 0.660500 (rank : 9) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P41220 | Gene names | RGS2, G0S8 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 2 (RGS2) (G0/G1 switch regulatory protein 8). | |||||
|
RGS10_HUMAN
|
||||||
| θ value | 5.08577e-11 (rank : 10) | NC score | 0.679279 (rank : 5) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O43665 | Gene names | RGS10 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 10 (RGS10). | |||||
|
RGS1_HUMAN
|
||||||
| θ value | 1.133e-10 (rank : 11) | NC score | 0.660094 (rank : 10) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q08116, Q07918, Q9H1W2 | Gene names | RGS1, 1R20, BL34 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 1 (RGS1) (Early response protein 1R20) (B-cell activation protein BL34). | |||||
|
RGS2_MOUSE
|
||||||
| θ value | 1.133e-10 (rank : 12) | NC score | 0.656380 (rank : 15) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O08849, Q544S7, Q91WX1, Q9JL24 | Gene names | Rgs2 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 2 (RGS2). | |||||
|
RGS1_MOUSE
|
||||||
| θ value | 1.47974e-10 (rank : 13) | NC score | 0.656473 (rank : 14) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9JL25, Q3TBD1, Q3TC18, Q3TD56 | Gene names | Rgs1 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 1 (RGS1). | |||||
|
RGS8_MOUSE
|
||||||
| θ value | 1.47974e-10 (rank : 14) | NC score | 0.657859 (rank : 12) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8BXT1 | Gene names | Rgs8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of G-protein signaling 8 (RGS8). | |||||
|
RGS8_HUMAN
|
||||||
| θ value | 1.9326e-10 (rank : 15) | NC score | 0.657068 (rank : 13) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P57771, Q3SYD2 | Gene names | RGS8 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 8 (RGS8). | |||||
|
RGS10_MOUSE
|
||||||
| θ value | 4.30538e-10 (rank : 16) | NC score | 0.671350 (rank : 8) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9CQE5, Q9D3L2 | Gene names | Rgs10 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 10 (RGS10). | |||||
|
RGS5_HUMAN
|
||||||
| θ value | 4.30538e-10 (rank : 17) | NC score | 0.659034 (rank : 11) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O15539 | Gene names | RGS5 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 5 (RGS5). | |||||
|
RGS4_HUMAN
|
||||||
| θ value | 5.62301e-10 (rank : 18) | NC score | 0.652517 (rank : 16) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P49798 | Gene names | RGS4 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 4 (RGS4) (RGP4). | |||||
|
RGS4_MOUSE
|
||||||
| θ value | 2.79066e-09 (rank : 19) | NC score | 0.651500 (rank : 18) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O08899, Q99L30 | Gene names | Rgs4 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 4 (RGS4). | |||||
|
RGS5_MOUSE
|
||||||
| θ value | 2.79066e-09 (rank : 20) | NC score | 0.652500 (rank : 17) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O08850, Q543B1, Q9D0Z2 | Gene names | Rgs5 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 5 (RGS5). | |||||
|
RGS13_HUMAN
|
||||||
| θ value | 1.06045e-08 (rank : 21) | NC score | 0.645154 (rank : 20) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O14921, Q6PGR2, Q8TD63, Q9BX45 | Gene names | RGS13 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 13 (RGS13). | |||||
|
RGS14_MOUSE
|
||||||
| θ value | 1.80886e-08 (rank : 22) | NC score | 0.606304 (rank : 29) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P97492, Q9DCD1 | Gene names | Rgs14 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 14 (RGS14) (RAP1/RAP2-interacting protein). | |||||
|
RGS16_HUMAN
|
||||||
| θ value | 2.36244e-08 (rank : 23) | NC score | 0.640992 (rank : 22) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O15492, Q99701 | Gene names | RGS16, RGSR | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 16 (RGS16) (Retinally abundant regulator of G-protein signaling) (RGS-R) (A28-RGS14P). | |||||
|
RGS19_MOUSE
|
||||||
| θ value | 4.0297e-08 (rank : 24) | NC score | 0.622748 (rank : 24) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9CX84, Q99L50 | Gene names | Rgs19 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 19 (RGS19). | |||||
|
RGS16_MOUSE
|
||||||
| θ value | 6.87365e-08 (rank : 25) | NC score | 0.643552 (rank : 21) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P97428, O09091, P97420, Q80V16 | Gene names | Rgs16, Rgsr | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 16 (RGS16) (Retinally abundant regulator of G-protein signaling) (RGS-R) (A28-RGS14P). | |||||
|
RGS13_MOUSE
|
||||||
| θ value | 8.97725e-08 (rank : 26) | NC score | 0.645921 (rank : 19) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8K443 | Gene names | Rgs13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of G-protein signaling 13 (RGS13). | |||||
|
RGS14_HUMAN
|
||||||
| θ value | 1.53129e-07 (rank : 27) | NC score | 0.596286 (rank : 30) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O43566, O43565, Q8TD62 | Gene names | RGS14 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 14 (RGS14). | |||||
|
RGS19_HUMAN
|
||||||
| θ value | 1.99992e-07 (rank : 28) | NC score | 0.620785 (rank : 25) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P49795, Q53XN0, Q8TD60 | Gene names | RGS19, GAIP, GNAI3IP | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 19 (RGS19) (G-alpha-interacting protein) (GAIP protein). | |||||
|
RGS20_MOUSE
|
||||||
| θ value | 4.45536e-07 (rank : 29) | NC score | 0.620762 (rank : 26) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9QZB1, Q3TY63, Q9CUV8, Q9QZB2 | Gene names | Rgs20, Rgsz1 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 20 (RGS20) (Regulator of G-protein signaling Z1). | |||||
|
RGS17_MOUSE
|
||||||
| θ value | 1.29631e-06 (rank : 30) | NC score | 0.623291 (rank : 23) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9QZB0, Q543T9 | Gene names | Rgs17, Rgsz2 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 17 (RGS17) (Regulator of Gz-selective protein signaling 2). | |||||
|
RGS11_HUMAN
|
||||||
| θ value | 2.88788e-06 (rank : 31) | NC score | 0.551682 (rank : 31) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O94810, O75883 | Gene names | RGS11 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 11 (RGS11). | |||||
|
RGS17_HUMAN
|
||||||
| θ value | 2.88788e-06 (rank : 32) | NC score | 0.614581 (rank : 27) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9UGC6, Q5TF49, Q8TD61, Q9UJS8 | Gene names | RGS17 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 17 (RGS17). | |||||
|
RGS20_HUMAN
|
||||||
| θ value | 4.92598e-06 (rank : 33) | NC score | 0.608154 (rank : 28) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O76081, Q96BG9 | Gene names | RGS20, RGSZ1, ZGAP1 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 20 (RGS20) (Regulator of Gz-selective protein signaling 1) (Gz-selective GTPase-activating protein) (G(z)GAP). | |||||
|
DVL2_HUMAN
|
||||||
| θ value | 6.43352e-06 (rank : 34) | NC score | 0.246146 (rank : 44) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O14641 | Gene names | DVL2 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-2 (Dishevelled-2) (DSH homolog 2). | |||||
|
DVL2_MOUSE
|
||||||
| θ value | 6.43352e-06 (rank : 35) | NC score | 0.246155 (rank : 43) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q60838 | Gene names | Dvl2 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-2 (Dishevelled-2) (DSH homolog 2). | |||||
|
DVL3_HUMAN
|
||||||
| θ value | 6.43352e-06 (rank : 36) | NC score | 0.253483 (rank : 42) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q92997, O14642, Q13531, Q8N5E9, Q92607 | Gene names | DVL3, KIAA0208 | |||
|
Domain Architecture |
|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-3 (Dishevelled-3) (DSH homolog 3). | |||||
|
DVL3_MOUSE
|
||||||
| θ value | 6.43352e-06 (rank : 37) | NC score | 0.253796 (rank : 41) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q61062 | Gene names | Dvl3 | |||
|
Domain Architecture |
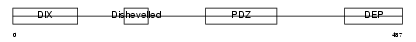
|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-3 (Dishevelled-3) (DSH homolog 3). | |||||
|
RGS9_HUMAN
|
||||||
| θ value | 2.44474e-05 (rank : 38) | NC score | 0.530641 (rank : 33) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O75916, O75573, Q696R2, Q8TD64, Q8TD65, Q9HC32, Q9HC33 | Gene names | RGS9 | |||
|
Domain Architecture |
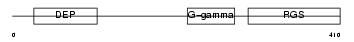
|
|||||
| Description | Regulator of G-protein signaling 9 (RGS9). | |||||
|
RGS9_MOUSE
|
||||||
| θ value | 2.44474e-05 (rank : 39) | NC score | 0.531204 (rank : 32) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O54828, Q9Z0S0 | Gene names | Rgs9 | |||
|
Domain Architecture |
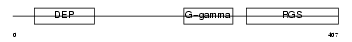
|
|||||
| Description | Regulator of G-protein signaling 9 (RGS9). | |||||
|
RGS12_HUMAN
|
||||||
| θ value | 5.44631e-05 (rank : 40) | NC score | 0.502127 (rank : 34) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | O14924, O14922, O14923, O43510, O75338 | Gene names | RGS12 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 12 (RGS12). | |||||
|
DVL1L_HUMAN
|
||||||
| θ value | 0.00020696 (rank : 41) | NC score | 0.229242 (rank : 45) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P54792 | Gene names | DVL1L1, DVL, DVL1 | |||
|
Domain Architecture |
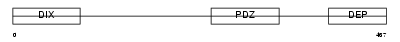
|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1-like (Dishevelled- 1-like) (DSH homolog 1-like). | |||||
|
DVL1_MOUSE
|
||||||
| θ value | 0.00020696 (rank : 42) | NC score | 0.226719 (rank : 46) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P51141, Q60868 | Gene names | Dvl1, Dvl | |||
|
Domain Architecture |
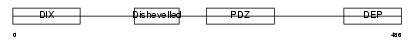
|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1 (Dishevelled-1) (DSH homolog 1). | |||||
|
DVL1_HUMAN
|
||||||
| θ value | 0.000270298 (rank : 43) | NC score | 0.226299 (rank : 47) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O14640 | Gene names | DVL1 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1 (Dishevelled-1) (DSH homolog 1). | |||||
|
RGS6_MOUSE
|
||||||
| θ value | 0.000270298 (rank : 44) | NC score | 0.487156 (rank : 38) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9Z2H2, Q8BGI8 | Gene names | Rgs6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of G-protein signaling 6 (RGS6). | |||||
|
RGS7_HUMAN
|
||||||
| θ value | 0.00035302 (rank : 45) | NC score | 0.500667 (rank : 35) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P49802, Q8TD66, Q8TD67, Q9UNU7, Q9Y6B9 | Gene names | RGS7 | |||
|
Domain Architecture |
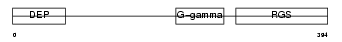
|
|||||
| Description | Regulator of G-protein signaling 7 (RGS7). | |||||
|
PO6F2_MOUSE
|
||||||
| θ value | 0.000786445 (rank : 46) | NC score | 0.048084 (rank : 57) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 639 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8BJI4 | Gene names | Pou6f2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | POU domain, class 6, transcription factor 2. | |||||
|
RGS7_MOUSE
|
||||||
| θ value | 0.000786445 (rank : 47) | NC score | 0.493758 (rank : 36) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O54829 | Gene names | Rgs7 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 7 (RGS7). | |||||
|
RGS6_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 48) | NC score | 0.459423 (rank : 39) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P49758, O75576, O75577, Q8TE13, Q8TE14, Q8TE15, Q8TE16, Q8TE17, Q8TE18, Q8TE19, Q8TE20, Q8TE21, Q8TE22, Q9Y245, Q9Y647 | Gene names | RGS6 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 6 (RGS6) (S914). | |||||
|
MSH3_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 49) | NC score | 0.029051 (rank : 63) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P20585, Q6PJT5, Q86UQ6, Q92867 | Gene names | MSH3, DUC1, DUG | |||
|
Domain Architecture |

|
|||||
| Description | DNA mismatch repair protein Msh3 (Divergent upstream protein) (DUP) (Mismatch repair protein 1) (MRP1). | |||||
|
AHTF1_HUMAN
|
||||||
| θ value | 0.163984 (rank : 50) | NC score | 0.049266 (rank : 56) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8WYP5, Q7Z4E3, Q8IZA4, Q96EH9, Q9Y4Q6 | Gene names | AHCTF1, ELYS, TMBS62 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-hook-containing transcription factor 1 (Embryonic large molecule derived from yolk sac). | |||||
|
K0310_HUMAN
|
||||||
| θ value | 0.163984 (rank : 51) | NC score | 0.063321 (rank : 55) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O15027, Q5SXP0, Q5SXP1, Q8N347, Q96HP1 | Gene names | KIAA0310 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0310. | |||||
|
JIP3_HUMAN
|
||||||
| θ value | 0.47712 (rank : 52) | NC score | 0.025811 (rank : 67) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1039 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9UPT6, Q96RY4, Q9H4I4, Q9H7P1, Q9NUG0 | Gene names | MAPK8IP3, JIP3, KIAA1066 | |||
|
Domain Architecture |

|
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 3 (JNK-interacting protein 3) (JIP-3) (JNK MAP kinase scaffold protein 3) (Mitogen- activated protein kinase 8-interacting protein 3). | |||||
|
CT112_HUMAN
|
||||||
| θ value | 0.62314 (rank : 53) | NC score | 0.041901 (rank : 58) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96MY1, Q5JYB7, Q6P0Y4, Q9BR34, Q9NQF6 | Gene names | C20orf112 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C20orf112. | |||||
|
NDF2_HUMAN
|
||||||
| θ value | 0.62314 (rank : 54) | NC score | 0.015440 (rank : 87) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q15784, Q8TBI7, Q9UQC6 | Gene names | NEUROD2, NDRF | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic differentiation factor 2 (NeuroD2) (NeuroD-related factor) (NDRF). | |||||
|
SCG2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 55) | NC score | 0.039717 (rank : 60) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P13521, Q8TBH3 | Gene names | SCG2, CHGC | |||
|
Domain Architecture |

|
|||||
| Description | Secretogranin-2 precursor (Secretogranin II) (SgII) (Chromogranin C) [Contains: Secretoneurin (SN)]. | |||||
|
VAPB_MOUSE
|
||||||
| θ value | 0.813845 (rank : 56) | NC score | 0.025194 (rank : 68) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9QY76 | Gene names | Vapb | |||
|
Domain Architecture |

|
|||||
| Description | Vesicle-associated membrane protein-associated protein B (VAMP- associated protein B) (VAMP-associated protein 33b) (VAMP-B) (VAP-B). | |||||
|
IWS1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 57) | NC score | 0.020684 (rank : 79) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
COBA2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 58) | NC score | 0.022522 (rank : 73) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P13942, Q07751, Q13271, Q13272, Q13273, Q99866, Q9UIP9 | Gene names | COL11A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(XI) chain precursor. | |||||
|
CV106_HUMAN
|
||||||
| θ value | 1.38821 (rank : 59) | NC score | 0.040360 (rank : 59) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6P0N0, Q86V14, Q96PY4, Q9NUR5, Q9Y4X9 | Gene names | C14orf106, KIAA1903 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf106 (P243). | |||||
|
CXA1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 60) | NC score | 0.011485 (rank : 96) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P17302, Q9Y5I8 | Gene names | GJA1, GJAL | |||
|
Domain Architecture |

|
|||||
| Description | Gap junction alpha-1 protein (Connexin-43) (Cx43) (Gap junction 43 kDa heart protein). | |||||
|
GP156_HUMAN
|
||||||
| θ value | 1.38821 (rank : 61) | NC score | 0.022397 (rank : 74) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NFN8, Q86SN6 | Gene names | GPR156, GABABL, PGR28 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 156 (GABAB-related G-protein coupled receptor) (G-protein coupled receptor PGR28). | |||||
|
PCLO_HUMAN
|
||||||
| θ value | 1.38821 (rank : 62) | NC score | 0.019387 (rank : 82) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
ABLM1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 63) | NC score | 0.006009 (rank : 109) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8K4G5, Q80U86, Q8BIR9, Q8K4G3, Q8K4G4 | Gene names | Ablim1, Ablim, Kiaa0059 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Actin-binding LIM protein 1 (Actin-binding LIM protein family member 1) (abLIM-1). | |||||
|
AKA10_HUMAN
|
||||||
| θ value | 1.81305 (rank : 64) | NC score | 0.193397 (rank : 48) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O43572, Q96AJ7 | Gene names | AKAP10 | |||
|
Domain Architecture |

|
|||||
| Description | A kinase anchor protein 10, mitochondrial precursor (Protein kinase A- anchoring protein 10) (PRKA10) (Dual specificity A kinase-anchoring protein 2) (D-AKAP-2). | |||||
|
AKA10_MOUSE
|
||||||
| θ value | 1.81305 (rank : 65) | NC score | 0.192497 (rank : 49) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O88845, Q5SUB5, Q5SUB7, Q7TPE7, Q8BQL6 | Gene names | Akap10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A kinase anchor protein 10, mitochondrial precursor (Protein kinase A- anchoring protein 10) (PRKA10) (Dual specificity A kinase-anchoring protein 2) (D-AKAP-2). | |||||
|
CO8A2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 66) | NC score | 0.023555 (rank : 71) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 299 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P25067, Q8TEJ5 | Gene names | COL8A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(VIII) chain precursor (Endothelial collagen). | |||||
|
CT112_MOUSE
|
||||||
| θ value | 1.81305 (rank : 67) | NC score | 0.030996 (rank : 62) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6DIB4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C20orf112 homolog. | |||||
|
FURIN_MOUSE
|
||||||
| θ value | 1.81305 (rank : 68) | NC score | 0.010176 (rank : 101) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P23188 | Gene names | Furin, Fur, Pcsk3 | |||
|
Domain Architecture |

|
|||||
| Description | Furin precursor (EC 3.4.21.75) (Paired basic amino acid residue cleaving enzyme) (PACE) (Dibasic-processing enzyme) (Prohormone convertase 3). | |||||
|
K0562_HUMAN
|
||||||
| θ value | 1.81305 (rank : 69) | NC score | 0.020337 (rank : 81) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O60308, Q5JSQ3, Q5SR24, Q5SR25, Q6PKF5, Q86W32, Q86X14 | Gene names | KIAA0562 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0562. | |||||
|
K1949_HUMAN
|
||||||
| θ value | 1.81305 (rank : 70) | NC score | 0.035385 (rank : 61) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q6NYC8, Q68CK8, Q6ZUJ6, Q8NDQ4, Q8TF52, Q9BRL9 | Gene names | KIAA1949 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1949. | |||||
|
NFH_MOUSE
|
||||||
| θ value | 1.81305 (rank : 71) | NC score | 0.012634 (rank : 93) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
TNC6B_HUMAN
|
||||||
| θ value | 1.81305 (rank : 72) | NC score | 0.026081 (rank : 66) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UPQ9, Q5TH52, Q8TBX2 | Gene names | TNRC6B, KIAA1093 | |||
|
Domain Architecture |

|
|||||
| Description | Trinucleotide repeat-containing 6B protein. | |||||
|
IASPP_MOUSE
|
||||||
| θ value | 2.36792 (rank : 73) | NC score | 0.015847 (rank : 86) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1024 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q5I1X5 | Gene names | Ppp1r13l, Nkip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RelA-associated inhibitor (Inhibitor of ASPP protein) (Protein iASPP) (PPP1R13B-like protein) (NFkB-interacting protein 1). | |||||
|
ULK2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 74) | NC score | 0.000849 (rank : 116) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 979 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8IYT8, O75119 | Gene names | ULK2, KIAA0623 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase ULK2 (EC 2.7.11.1) (Unc-51-like kinase 2). | |||||
|
ARHGG_HUMAN
|
||||||
| θ value | 3.0926 (rank : 75) | NC score | 0.009874 (rank : 102) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5VV41, Q86TF0, Q99434 | Gene names | ARHGEF16, NBR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 16. | |||||
|
CO1A2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 76) | NC score | 0.020688 (rank : 78) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P08123, P02464, Q9UEB6, Q9UPH0 | Gene names | COL1A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(I) chain precursor. | |||||
|
CO4A5_HUMAN
|
||||||
| θ value | 3.0926 (rank : 77) | NC score | 0.028189 (rank : 64) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 383 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P29400, Q16006, Q16126, Q6LD84 | Gene names | COL4A5 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-5(IV) chain precursor. | |||||
|
OBSCN_HUMAN
|
||||||
| θ value | 3.0926 (rank : 78) | NC score | 0.000399 (rank : 117) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1931 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q5VST9, Q2A664, Q5T7G8, Q5T7G9, Q5VSU2, Q86YC7, Q8NHN0, Q8NHN1, Q8NHN2, Q8NHN4, Q8NHN5, Q8NHN6, Q8NHN7, Q8NHN8, Q8NHN9, Q96AA2, Q9HCD3, Q9HCL6 | Gene names | OBSCN, KIAA1556, KIAA1639 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Obscurin (Obscurin-myosin light chain kinase) (Obscurin-MLCK) (Obscurin-RhoGEF). | |||||
|
PUNC_HUMAN
|
||||||
| θ value | 3.0926 (rank : 79) | NC score | 0.006032 (rank : 108) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 449 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8IVU1, O95215 | Gene names | PUNC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative neuronal cell adhesion molecule precursor. | |||||
|
CO4A3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 80) | NC score | 0.022105 (rank : 75) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 421 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q01955, Q9BQT2 | Gene names | COL4A3 | |||
|
Domain Architecture |
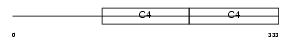
|
|||||
| Description | Collagen alpha-3(IV) chain precursor (Goodpasture antigen). | |||||
|
CO9A2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 81) | NC score | 0.023476 (rank : 72) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q07643 | Gene names | Col9a2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-2(IX) chain precursor. | |||||
|
ERBB3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 82) | NC score | -0.000263 (rank : 118) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 898 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61526, Q3KQR1, Q68J64, Q810U8, Q8K317 | Gene names | Erbb3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Receptor tyrosine-protein kinase erbB-3 precursor (EC 2.7.10.1) (c- erbB3) (Glial growth factor receptor). | |||||
|
SET1A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 83) | NC score | 0.026742 (rank : 65) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O15047, Q6PIF3, Q8TAJ6 | Gene names | SETD1A, KIAA0339, SET1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET1 (EC 2.1.1.43) (Set1/Ash2 histone methyltransferase complex subunit SET1) (SET-domain-containing protein 1A). | |||||
|
TCEA2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 84) | NC score | 0.010498 (rank : 99) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q15560, Q8TD37, Q8TD38 | Gene names | TCEA2 | |||
|
Domain Architecture |
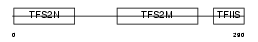
|
|||||
| Description | Transcription elongation factor A protein 2 (Transcription elongation factor S-II protein 2) (Testis-specific S-II) (Transcription elongation factor TFIIS.l). | |||||
|
ANGL3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 85) | NC score | 0.004330 (rank : 115) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 306 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9R182 | Gene names | Angptl3 | |||
|
Domain Architecture |
|
|||||
| Description | Angiopoietin-related protein 3 precursor (Angiopoietin-like 3). | |||||
|
CO4A4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 86) | NC score | 0.021775 (rank : 76) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 440 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P53420 | Gene names | COL4A4 | |||
|
Domain Architecture |
|
|||||
| Description | Collagen alpha-4(IV) chain precursor. | |||||
|
CXA1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 87) | NC score | 0.009038 (rank : 104) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P23242 | Gene names | Gja1, Cxn-43 | |||
|
Domain Architecture |

|
|||||
| Description | Gap junction alpha-1 protein (Connexin-43) (Cx43) (Gap junction 43 kDa heart protein). | |||||
|
HS90A_HUMAN
|
||||||
| θ value | 5.27518 (rank : 88) | NC score | 0.011809 (rank : 95) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P07900, Q2PP14, Q9BVQ5 | Gene names | HSP90AA1, HSP90A, HSPC1, HSPCA | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock protein HSP 90-alpha (HSP 86) (NY-REN-38 antigen). | |||||
|
HS90A_MOUSE
|
||||||
| θ value | 5.27518 (rank : 89) | NC score | 0.011328 (rank : 97) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P07901 | Gene names | Hsp90aa1, Hsp86, Hsp86-1, Hspca | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock protein HSP 90-alpha (HSP 86) (Tumor-specific transplantation 86 kDa antigen) (TSTA). | |||||
|
IQEC3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 90) | NC score | 0.012419 (rank : 94) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UPP2, Q8TB43 | Gene names | IQSEC3, KIAA1110 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 3. | |||||
|
RNF39_HUMAN
|
||||||
| θ value | 5.27518 (rank : 91) | NC score | 0.005204 (rank : 112) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9H2S5, Q5SPM8, Q5SPM9, Q5SPN0, Q5SRJ9, Q5SRK1, Q5SS29, Q9H2S3, Q9H2S4 | Gene names | RNF39, HZFW | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 39 (Protein HZFw). | |||||
|
ZN692_HUMAN
|
||||||
| θ value | 5.27518 (rank : 92) | NC score | -0.001519 (rank : 121) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 808 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9BU19, Q5SRA5, Q5SRA6, Q9HBC9, Q9NW93, Q9NWY6, Q9UF97 | Gene names | ZNF692 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 692. | |||||
|
CCD16_MOUSE
|
||||||
| θ value | 6.88961 (rank : 93) | NC score | 0.010453 (rank : 100) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8R1N0, Q3TR52, Q9CWV9, Q9CYI6 | Gene names | Ccdc16, Omcg1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 16 (Ovus mutant candidate gene 1 protein). | |||||
|
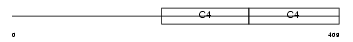
CO4A1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 94) | NC score | 0.020366 (rank : 80) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P02462 | Gene names | COL4A1 | |||
|
Domain Architecture |
|
|||||
| Description | Collagen alpha-1(IV) chain precursor. | |||||
|
CO9A1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 95) | NC score | 0.018583 (rank : 83) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 477 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q05722, Q61269, Q61270, Q61433, Q61940 | Gene names | Col9a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(IX) chain precursor. | |||||
|
ERBB3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 96) | NC score | -0.000707 (rank : 119) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 918 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P21860 | Gene names | ERBB3, HER3 | |||
|
Domain Architecture |

|
|||||
| Description | Receptor tyrosine-protein kinase erbB-3 precursor (EC 2.7.10.1) (c- erbB3) (Tyrosine kinase-type cell surface receptor HER3). | |||||
|
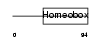
HXB3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 97) | NC score | 0.005406 (rank : 111) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 493 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P09026, P10285, Q4PJ06, Q61680 | Gene names | Hoxb3, Hox-2.7, Hoxb-3 | |||
|
Domain Architecture |
|
|||||
| Description | Homeobox protein Hox-B3 (Hox-2.7) (Homeobox protein MH-23). | |||||
|
HXD3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 98) | NC score | 0.006893 (rank : 106) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P31249, Q99955, Q9BSC5 | Gene names | HOXD3, HOX4A | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-D3 (Hox-4A). | |||||
|
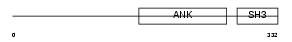
IASPP_HUMAN
|
||||||
| θ value | 6.88961 (rank : 99) | NC score | 0.013663 (rank : 90) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 688 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8WUF5, Q2PNZ9, Q5DU71, Q5I1X4, Q6P1R7, Q6PKF8, Q9Y290 | Gene names | PPP1R13L, IASPP, NKIP1, PPP1R13BL, RAI | |||
|
Domain Architecture |
|
|||||
| Description | RelA-associated inhibitor (Inhibitor of ASPP protein) (Protein iASPP) (PPP1R13B-like protein) (NFkB-interacting protein 1). | |||||
|
ITPR3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 100) | NC score | 0.008385 (rank : 105) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P70227, Q5DWM4, Q8CED5, Q91Z08 | Gene names | Itpr3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol 1,4,5-trisphosphate receptor type 3 (Type 3 inositol 1,4,5- trisphosphate receptor) (Type 3 InsP3 receptor) (IP3 receptor isoform 3) (InsP3R3). | |||||
|
SAM68_HUMAN
|
||||||
| θ value | 6.88961 (rank : 101) | NC score | 0.011022 (rank : 98) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q07666, Q6PJX7, Q8NB97, Q99760 | Gene names | KHDRBS1, SAM68 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | KH domain-containing, RNA-binding, signal transduction-associated protein 1 (p21 Ras GTPase-activating protein-associated p62) (GAP- associated tyrosine phosphoprotein p62) (Src-associated in mitosis 68 kDa protein) (Sam68) (p68). | |||||
|
SCG2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 102) | NC score | 0.024599 (rank : 70) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q03517, Q80Y79, Q9CW80 | Gene names | Scg2, Chgc, Scg-2 | |||
|
Domain Architecture |

|
|||||
| Description | Secretogranin-2 precursor (Secretogranin II) (SgII) (Chromogranin C) [Contains: Secretoneurin (SN)]. | |||||
|
ZN516_MOUSE
|
||||||
| θ value | 6.88961 (rank : 103) | NC score | -0.001197 (rank : 120) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1197 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q7TSH3 | Gene names | Znf516, Kiaa0222, Zfp516 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 516. | |||||
|
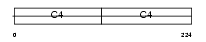
CO4A1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 104) | NC score | 0.021198 (rank : 77) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 450 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P02463 | Gene names | Col4a1 | |||
|
Domain Architecture |
|
|||||
| Description | Collagen alpha-1(IV) chain precursor. | |||||
|
CO9A1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 105) | NC score | 0.017831 (rank : 84) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P20849, Q13699, Q13700, Q5TF52, Q6P467, Q96BM8, Q99225, Q9H151, Q9H152, Q9Y6P2, Q9Y6P3 | Gene names | COL9A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(IX) chain precursor. | |||||
|
HXB3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 106) | NC score | 0.005413 (rank : 110) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 559 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P14651, O95615, P17484 | Gene names | HOXB3, HOX2G | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-B3 (Hox-2G) (Hox-2.7). | |||||
|
HXD3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 107) | NC score | 0.004418 (rank : 114) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P09027, Q3UUD3 | Gene names | Hoxd3, Hox-4.1, Hoxd-3 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-D3 (Hox-4.1) (Homeobox protein MH-19). | |||||
|
IQEC3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 108) | NC score | 0.015300 (rank : 88) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q3TES0, Q3UHP8, Q80TJ8 | Gene names | Iqsec3, Kiaa1110 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 3. | |||||
|
KIF1A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 109) | NC score | 0.004908 (rank : 113) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P33173, Q61770 | Gene names | Kif1a, Atsv, Kif1 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF1A (Axonal transporter of synaptic vesicles). | |||||
|
NFH_HUMAN
|
||||||
| θ value | 8.99809 (rank : 110) | NC score | 0.013467 (rank : 91) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
SHAN1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 111) | NC score | 0.009207 (rank : 103) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9Y566, Q9NYW9 | Gene names | SHANK1 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 1 (Shank1) (Somatostatin receptor-interacting protein) (SSTR-interacting protein) (SSTRIP). | |||||
|
T22D2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 112) | NC score | 0.024743 (rank : 69) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O75157, Q6PI50, Q9H2Z6, Q9H2Z7, Q9H2Z8 | Gene names | TSC22D2, KIAA0669, TILZ4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TSC22 domain family protein 2 (TSC22-related-inducible leucine zipper protein 4). | |||||
|
TRRAP_HUMAN
|
||||||
| θ value | 8.99809 (rank : 113) | NC score | 0.016281 (rank : 85) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y4A5, O75218, Q9Y631, Q9Y6H4 | Gene names | TRRAP, PAF400 | |||
|
Domain Architecture |

|
|||||
| Description | Transformation/transcription domain-associated protein (350/400 kDa PCAF-associated factor) (PAF350/400) (STAF40) (Tra1 homolog). | |||||
|
TRRAP_MOUSE
|
||||||
| θ value | 8.99809 (rank : 114) | NC score | 0.014914 (rank : 89) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80YV3, Q8C0Z5, Q8K104 | Gene names | Trrap | |||
|
Domain Architecture |

|
|||||
| Description | Transformation/transcription domain-associated protein (Tra1 homolog). | |||||
|
UBP42_HUMAN
|
||||||
| θ value | 8.99809 (rank : 115) | NC score | 0.012896 (rank : 92) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9H9J4, Q3C166, Q6P9B4 | Gene names | USP42 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 42 (EC 3.1.2.15) (Ubiquitin thioesterase 42) (Ubiquitin-specific-processing protease 42) (Deubiquitinating enzyme 42). | |||||
|
US6NL_MOUSE
|
||||||
| θ value | 8.99809 (rank : 116) | NC score | 0.006516 (rank : 107) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80XC3, Q6ZQK8 | Gene names | Usp6nl, Kiaa0019 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USP6 N-terminal-like protein. | |||||
|
SNX13_HUMAN
|
||||||
| θ value | θ > 10 (rank : 117) | NC score | 0.157868 (rank : 51) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9Y5W8, O94821, Q8WVZ2, Q8WXH8 | Gene names | SNX13, KIAA0713 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-13 (RGS domain- and PHOX domain-containing protein) (RGS-PX1). | |||||
|
SNX13_MOUSE
|
||||||
| θ value | θ > 10 (rank : 118) | NC score | 0.160708 (rank : 50) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q6PHS6 | Gene names | Snx13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sorting nexin-13. | |||||
|
SNX14_HUMAN
|
||||||
| θ value | θ > 10 (rank : 119) | NC score | 0.073765 (rank : 52) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y5W7, Q9BSD1 | Gene names | SNX14 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-14. | |||||
|
SNX14_MOUSE
|
||||||
| θ value | θ > 10 (rank : 120) | NC score | 0.072766 (rank : 53) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8BHY8 | Gene names | Snx14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sorting nexin-14. | |||||
|
SNX25_HUMAN
|
||||||
| θ value | θ > 10 (rank : 121) | NC score | 0.071898 (rank : 54) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9H3E2 | Gene names | SNX25 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-25. | |||||
|
AXN2_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 116 | |
| SwissProt Accessions | O88566, Q9QXJ6 | Gene names | Axin2 | |||
|
Domain Architecture |

|
|||||
| Description | Axin-2 (Axis inhibition protein 2) (Conductin) (Axin-like protein) (Axil). | |||||
|
AXN2_HUMAN
|
||||||
| NC score | 0.989516 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9Y2T1, Q9H3M6, Q9UH84 | Gene names | AXIN2 | |||
|
Domain Architecture |

|
|||||
| Description | Axin-2 (Axis inhibition protein 2) (Conductin) (Axin-like protein) (Axil). | |||||
|
AXN1_MOUSE
|
||||||
| NC score | 0.948949 (rank : 3) | θ value | 3.7435e-147 (rank : 3) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O35625 | Gene names | Axin1, Axin, Fu | |||
|
Domain Architecture |

|
|||||
| Description | Axin-1 (Axis inhibition protein 1) (Fused protein). | |||||
|
AXN1_HUMAN
|
||||||
| NC score | 0.948670 (rank : 4) | θ value | 2.77624e-142 (rank : 4) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O15169, Q4TT26, Q4TT27, Q86YA7, Q8WVW6, Q96S28 | Gene names | AXIN1, AXIN | |||
|
Domain Architecture |

|
|||||
| Description | Axin-1 (Axis inhibition protein 1) (hAxin). | |||||
|
RGS10_HUMAN
|
||||||
| NC score | 0.679279 (rank : 5) | θ value | 5.08577e-11 (rank : 10) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O43665 | Gene names | RGS10 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 10 (RGS10). | |||||
|
RGS18_HUMAN
|
||||||
| NC score | 0.677020 (rank : 6) | θ value | 1.68911e-14 (rank : 5) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9NS28 | Gene names | RGS18, RGS13 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 18 (RGS18). | |||||
|
RGS18_MOUSE
|
||||||
| NC score | 0.675483 (rank : 7) | θ value | 1.09485e-13 (rank : 6) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q99PG4 | Gene names | Rgs18 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 18 (RGS18). | |||||
|
RGS10_MOUSE
|
||||||
| NC score | 0.671350 (rank : 8) | θ value | 4.30538e-10 (rank : 16) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9CQE5, Q9D3L2 | Gene names | Rgs10 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 10 (RGS10). | |||||
|
RGS2_HUMAN
|
||||||
| NC score | 0.660500 (rank : 9) | θ value | 2.98157e-11 (rank : 9) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P41220 | Gene names | RGS2, G0S8 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 2 (RGS2) (G0/G1 switch regulatory protein 8). | |||||
|
RGS1_HUMAN
|
||||||
| NC score | 0.660094 (rank : 10) | θ value | 1.133e-10 (rank : 11) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q08116, Q07918, Q9H1W2 | Gene names | RGS1, 1R20, BL34 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 1 (RGS1) (Early response protein 1R20) (B-cell activation protein BL34). | |||||
|
RGS5_HUMAN
|
||||||
| NC score | 0.659034 (rank : 11) | θ value | 4.30538e-10 (rank : 17) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O15539 | Gene names | RGS5 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 5 (RGS5). | |||||
|
RGS8_MOUSE
|
||||||
| NC score | 0.657859 (rank : 12) | θ value | 1.47974e-10 (rank : 14) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8BXT1 | Gene names | Rgs8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of G-protein signaling 8 (RGS8). | |||||
|
RGS8_HUMAN
|
||||||
| NC score | 0.657068 (rank : 13) | θ value | 1.9326e-10 (rank : 15) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P57771, Q3SYD2 | Gene names | RGS8 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 8 (RGS8). | |||||
|
RGS1_MOUSE
|
||||||
| NC score | 0.656473 (rank : 14) | θ value | 1.47974e-10 (rank : 13) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9JL25, Q3TBD1, Q3TC18, Q3TD56 | Gene names | Rgs1 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 1 (RGS1). | |||||
|
RGS2_MOUSE
|
||||||
| NC score | 0.656380 (rank : 15) | θ value | 1.133e-10 (rank : 12) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O08849, Q544S7, Q91WX1, Q9JL24 | Gene names | Rgs2 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 2 (RGS2). | |||||
|
RGS4_HUMAN
|
||||||
| NC score | 0.652517 (rank : 16) | θ value | 5.62301e-10 (rank : 18) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P49798 | Gene names | RGS4 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 4 (RGS4) (RGP4). | |||||
|
RGS5_MOUSE
|
||||||
| NC score | 0.652500 (rank : 17) | θ value | 2.79066e-09 (rank : 20) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O08850, Q543B1, Q9D0Z2 | Gene names | Rgs5 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 5 (RGS5). | |||||
|
RGS4_MOUSE
|
||||||
| NC score | 0.651500 (rank : 18) | θ value | 2.79066e-09 (rank : 19) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O08899, Q99L30 | Gene names | Rgs4 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 4 (RGS4). | |||||
|
RGS13_MOUSE
|
||||||
| NC score | 0.645921 (rank : 19) | θ value | 8.97725e-08 (rank : 26) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8K443 | Gene names | Rgs13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of G-protein signaling 13 (RGS13). | |||||
|
RGS13_HUMAN
|
||||||
| NC score | 0.645154 (rank : 20) | θ value | 1.06045e-08 (rank : 21) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O14921, Q6PGR2, Q8TD63, Q9BX45 | Gene names | RGS13 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 13 (RGS13). | |||||
|
RGS16_MOUSE
|
||||||
| NC score | 0.643552 (rank : 21) | θ value | 6.87365e-08 (rank : 25) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P97428, O09091, P97420, Q80V16 | Gene names | Rgs16, Rgsr | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 16 (RGS16) (Retinally abundant regulator of G-protein signaling) (RGS-R) (A28-RGS14P). | |||||
|
RGS16_HUMAN
|
||||||
| NC score | 0.640992 (rank : 22) | θ value | 2.36244e-08 (rank : 23) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O15492, Q99701 | Gene names | RGS16, RGSR | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 16 (RGS16) (Retinally abundant regulator of G-protein signaling) (RGS-R) (A28-RGS14P). | |||||
|
RGS17_MOUSE
|
||||||
| NC score | 0.623291 (rank : 23) | θ value | 1.29631e-06 (rank : 30) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9QZB0, Q543T9 | Gene names | Rgs17, Rgsz2 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 17 (RGS17) (Regulator of Gz-selective protein signaling 2). | |||||
|
RGS19_MOUSE
|
||||||
| NC score | 0.622748 (rank : 24) | θ value | 4.0297e-08 (rank : 24) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9CX84, Q99L50 | Gene names | Rgs19 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 19 (RGS19). | |||||
|
RGS19_HUMAN
|
||||||
| NC score | 0.620785 (rank : 25) | θ value | 1.99992e-07 (rank : 28) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P49795, Q53XN0, Q8TD60 | Gene names | RGS19, GAIP, GNAI3IP | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 19 (RGS19) (G-alpha-interacting protein) (GAIP protein). | |||||
|
RGS20_MOUSE
|
||||||
| NC score | 0.620762 (rank : 26) | θ value | 4.45536e-07 (rank : 29) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9QZB1, Q3TY63, Q9CUV8, Q9QZB2 | Gene names | Rgs20, Rgsz1 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 20 (RGS20) (Regulator of G-protein signaling Z1). | |||||
|
RGS17_HUMAN
|
||||||
| NC score | 0.614581 (rank : 27) | θ value | 2.88788e-06 (rank : 32) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9UGC6, Q5TF49, Q8TD61, Q9UJS8 | Gene names | RGS17 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 17 (RGS17). | |||||
|
RGS20_HUMAN
|
||||||
| NC score | 0.608154 (rank : 28) | θ value | 4.92598e-06 (rank : 33) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O76081, Q96BG9 | Gene names | RGS20, RGSZ1, ZGAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 20 (RGS20) (Regulator of Gz-selective protein signaling 1) (Gz-selective GTPase-activating protein) (G(z)GAP). | |||||
|
RGS14_MOUSE
|
||||||
| NC score | 0.606304 (rank : 29) | θ value | 1.80886e-08 (rank : 22) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P97492, Q9DCD1 | Gene names | Rgs14 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 14 (RGS14) (RAP1/RAP2-interacting protein). | |||||
|
RGS14_HUMAN
|
||||||
| NC score | 0.596286 (rank : 30) | θ value | 1.53129e-07 (rank : 27) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O43566, O43565, Q8TD62 | Gene names | RGS14 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 14 (RGS14). | |||||
|
RGS11_HUMAN
|
||||||
| NC score | 0.551682 (rank : 31) | θ value | 2.88788e-06 (rank : 31) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O94810, O75883 | Gene names | RGS11 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 11 (RGS11). | |||||
|
RGS9_MOUSE
|
||||||
| NC score | 0.531204 (rank : 32) | θ value | 2.44474e-05 (rank : 39) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O54828, Q9Z0S0 | Gene names | Rgs9 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 9 (RGS9). | |||||
|
RGS9_HUMAN
|
||||||
| NC score | 0.530641 (rank : 33) | θ value | 2.44474e-05 (rank : 38) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O75916, O75573, Q696R2, Q8TD64, Q8TD65, Q9HC32, Q9HC33 | Gene names | RGS9 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 9 (RGS9). | |||||
|
RGS12_HUMAN
|
||||||
| NC score | 0.502127 (rank : 34) | θ value | 5.44631e-05 (rank : 40) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | O14924, O14922, O14923, O43510, O75338 | Gene names | RGS12 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 12 (RGS12). | |||||
|
RGS7_HUMAN
|
||||||
| NC score | 0.500667 (rank : 35) | θ value | 0.00035302 (rank : 45) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P49802, Q8TD66, Q8TD67, Q9UNU7, Q9Y6B9 | Gene names | RGS7 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 7 (RGS7). | |||||
|
RGS7_MOUSE
|
||||||
| NC score | 0.493758 (rank : 36) | θ value | 0.000786445 (rank : 47) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O54829 | Gene names | Rgs7 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 7 (RGS7). | |||||
|
RGS3_HUMAN
|
||||||
| NC score | 0.489779 (rank : 37) | θ value | 4.16044e-13 (rank : 7) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
RGS6_MOUSE
|
||||||
| NC score | 0.487156 (rank : 38) | θ value | 0.000270298 (rank : 44) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9Z2H2, Q8BGI8 | Gene names | Rgs6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of G-protein signaling 6 (RGS6). | |||||
|
RGS6_HUMAN
|
||||||
| NC score | 0.459423 (rank : 39) | θ value | 0.00509761 (rank : 48) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P49758, O75576, O75577, Q8TE13, Q8TE14, Q8TE15, Q8TE16, Q8TE17, Q8TE18, Q8TE19, Q8TE20, Q8TE21, Q8TE22, Q9Y245, Q9Y647 | Gene names | RGS6 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 6 (RGS6) (S914). | |||||
|
RGS3_MOUSE
|
||||||
| NC score | 0.432395 (rank : 40) | θ value | 1.02475e-11 (rank : 8) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
DVL3_MOUSE
|
||||||
| NC score | 0.253796 (rank : 41) | θ value | 6.43352e-06 (rank : 37) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q61062 | Gene names | Dvl3 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-3 (Dishevelled-3) (DSH homolog 3). | |||||
|
DVL3_HUMAN
|
||||||
| NC score | 0.253483 (rank : 42) | θ value | 6.43352e-06 (rank : 36) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q92997, O14642, Q13531, Q8N5E9, Q92607 | Gene names | DVL3, KIAA0208 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-3 (Dishevelled-3) (DSH homolog 3). | |||||
|
DVL2_MOUSE
|
||||||
| NC score | 0.246155 (rank : 43) | θ value | 6.43352e-06 (rank : 35) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q60838 | Gene names | Dvl2 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-2 (Dishevelled-2) (DSH homolog 2). | |||||
|
DVL2_HUMAN
|
||||||
| NC score | 0.246146 (rank : 44) | θ value | 6.43352e-06 (rank : 34) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O14641 | Gene names | DVL2 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-2 (Dishevelled-2) (DSH homolog 2). | |||||
|
DVL1L_HUMAN
|
||||||
| NC score | 0.229242 (rank : 45) | θ value | 0.00020696 (rank : 41) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P54792 | Gene names | DVL1L1, DVL, DVL1 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1-like (Dishevelled- 1-like) (DSH homolog 1-like). | |||||
|
DVL1_MOUSE
|
||||||
| NC score | 0.226719 (rank : 46) | θ value | 0.00020696 (rank : 42) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P51141, Q60868 | Gene names | Dvl1, Dvl | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1 (Dishevelled-1) (DSH homolog 1). | |||||
|
DVL1_HUMAN
|
||||||
| NC score | 0.226299 (rank : 47) | θ value | 0.000270298 (rank : 43) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O14640 | Gene names | DVL1 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1 (Dishevelled-1) (DSH homolog 1). | |||||
|
AKA10_HUMAN
|
||||||
| NC score | 0.193397 (rank : 48) | θ value | 1.81305 (rank : 64) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O43572, Q96AJ7 | Gene names | AKAP10 | |||
|
Domain Architecture |

|
|||||
| Description | A kinase anchor protein 10, mitochondrial precursor (Protein kinase A- anchoring protein 10) (PRKA10) (Dual specificity A kinase-anchoring protein 2) (D-AKAP-2). | |||||
|
AKA10_MOUSE
|
||||||
| NC score | 0.192497 (rank : 49) | θ value | 1.81305 (rank : 65) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O88845, Q5SUB5, Q5SUB7, Q7TPE7, Q8BQL6 | Gene names | Akap10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A kinase anchor protein 10, mitochondrial precursor (Protein kinase A- anchoring protein 10) (PRKA10) (Dual specificity A kinase-anchoring protein 2) (D-AKAP-2). | |||||
|
SNX13_MOUSE
|
||||||
| NC score | 0.160708 (rank : 50) | θ value | θ > 10 (rank : 118) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q6PHS6 | Gene names | Snx13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sorting nexin-13. | |||||
|
SNX13_HUMAN
|
||||||
| NC score | 0.157868 (rank : 51) | θ value | θ > 10 (rank : 117) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9Y5W8, O94821, Q8WVZ2, Q8WXH8 | Gene names | SNX13, KIAA0713 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-13 (RGS domain- and PHOX domain-containing protein) (RGS-PX1). | |||||
|
SNX14_HUMAN
|
||||||
| NC score | 0.073765 (rank : 52) | θ value | θ > 10 (rank : 119) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y5W7, Q9BSD1 | Gene names | SNX14 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-14. | |||||
|
SNX14_MOUSE
|
||||||
| NC score | 0.072766 (rank : 53) | θ value | θ > 10 (rank : 120) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8BHY8 | Gene names | Snx14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sorting nexin-14. | |||||
|
SNX25_HUMAN
|
||||||
| NC score | 0.071898 (rank : 54) | θ value | θ > 10 (rank : 121) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9H3E2 | Gene names | SNX25 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-25. | |||||
|
K0310_HUMAN
|
||||||
| NC score | 0.063321 (rank : 55) | θ value | 0.163984 (rank : 51) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O15027, Q5SXP0, Q5SXP1, Q8N347, Q96HP1 | Gene names | KIAA0310 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0310. | |||||
|
AHTF1_HUMAN
|
||||||
| NC score | 0.049266 (rank : 56) | θ value | 0.163984 (rank : 50) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8WYP5, Q7Z4E3, Q8IZA4, Q96EH9, Q9Y4Q6 | Gene names | AHCTF1, ELYS, TMBS62 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-hook-containing transcription factor 1 (Embryonic large molecule derived from yolk sac). | |||||
|
PO6F2_MOUSE
|
||||||
| NC score | 0.048084 (rank : 57) | θ value | 0.000786445 (rank : 46) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 639 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8BJI4 | Gene names | Pou6f2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | POU domain, class 6, transcription factor 2. | |||||
|
CT112_HUMAN
|
||||||
| NC score | 0.041901 (rank : 58) | θ value | 0.62314 (rank : 53) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96MY1, Q5JYB7, Q6P0Y4, Q9BR34, Q9NQF6 | Gene names | C20orf112 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C20orf112. | |||||
|
CV106_HUMAN
|
||||||
| NC score | 0.040360 (rank : 59) | θ value | 1.38821 (rank : 59) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6P0N0, Q86V14, Q96PY4, Q9NUR5, Q9Y4X9 | Gene names | C14orf106, KIAA1903 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf106 (P243). | |||||
|
SCG2_HUMAN
|
||||||
| NC score | 0.039717 (rank : 60) | θ value | 0.813845 (rank : 55) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P13521, Q8TBH3 | Gene names | SCG2, CHGC | |||
|
Domain Architecture |

|
|||||
| Description | Secretogranin-2 precursor (Secretogranin II) (SgII) (Chromogranin C) [Contains: Secretoneurin (SN)]. | |||||
|
K1949_HUMAN
|
||||||
| NC score | 0.035385 (rank : 61) | θ value | 1.81305 (rank : 70) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q6NYC8, Q68CK8, Q6ZUJ6, Q8NDQ4, Q8TF52, Q9BRL9 | Gene names | KIAA1949 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1949. | |||||
|
CT112_MOUSE
|
||||||
| NC score | 0.030996 (rank : 62) | θ value | 1.81305 (rank : 67) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6DIB4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C20orf112 homolog. | |||||
|
MSH3_HUMAN
|
||||||
| NC score | 0.029051 (rank : 63) | θ value | 0.0961366 (rank : 49) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P20585, Q6PJT5, Q86UQ6, Q92867 | Gene names | MSH3, DUC1, DUG | |||
|
Domain Architecture |

|
|||||
| Description | DNA mismatch repair protein Msh3 (Divergent upstream protein) (DUP) (Mismatch repair protein 1) (MRP1). | |||||
|
CO4A5_HUMAN
|
||||||
| NC score | 0.028189 (rank : 64) | θ value | 3.0926 (rank : 77) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 383 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P29400, Q16006, Q16126, Q6LD84 | Gene names | COL4A5 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-5(IV) chain precursor. | |||||
|
SET1A_HUMAN
|
||||||
| NC score | 0.026742 (rank : 65) | θ value | 4.03905 (rank : 83) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O15047, Q6PIF3, Q8TAJ6 | Gene names | SETD1A, KIAA0339, SET1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET1 (EC 2.1.1.43) (Set1/Ash2 histone methyltransferase complex subunit SET1) (SET-domain-containing protein 1A). | |||||
|
TNC6B_HUMAN
|
||||||
| NC score | 0.026081 (rank : 66) | θ value | 1.81305 (rank : 72) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UPQ9, Q5TH52, Q8TBX2 | Gene names | TNRC6B, KIAA1093 | |||
|
Domain Architecture |

|
|||||
| Description | Trinucleotide repeat-containing 6B protein. | |||||
|
JIP3_HUMAN
|
||||||
| NC score | 0.025811 (rank : 67) | θ value | 0.47712 (rank : 52) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1039 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9UPT6, Q96RY4, Q9H4I4, Q9H7P1, Q9NUG0 | Gene names | MAPK8IP3, JIP3, KIAA1066 | |||
|
Domain Architecture |

|
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 3 (JNK-interacting protein 3) (JIP-3) (JNK MAP kinase scaffold protein 3) (Mitogen- activated protein kinase 8-interacting protein 3). | |||||
|
VAPB_MOUSE
|
||||||
| NC score | 0.025194 (rank : 68) | θ value | 0.813845 (rank : 56) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9QY76 | Gene names | Vapb | |||
|
Domain Architecture |

|
|||||
| Description | Vesicle-associated membrane protein-associated protein B (VAMP- associated protein B) (VAMP-associated protein 33b) (VAMP-B) (VAP-B). | |||||
|
T22D2_HUMAN
|
||||||
| NC score | 0.024743 (rank : 69) | θ value | 8.99809 (rank : 112) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O75157, Q6PI50, Q9H2Z6, Q9H2Z7, Q9H2Z8 | Gene names | TSC22D2, KIAA0669, TILZ4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TSC22 domain family protein 2 (TSC22-related-inducible leucine zipper protein 4). | |||||
|
SCG2_MOUSE
|
||||||
| NC score | 0.024599 (rank : 70) | θ value | 6.88961 (rank : 102) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q03517, Q80Y79, Q9CW80 | Gene names | Scg2, Chgc, Scg-2 | |||
|
Domain Architecture |

|
|||||
| Description | Secretogranin-2 precursor (Secretogranin II) (SgII) (Chromogranin C) [Contains: Secretoneurin (SN)]. | |||||
|
CO8A2_HUMAN
|
||||||
| NC score | 0.023555 (rank : 71) | θ value | 1.81305 (rank : 66) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 299 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P25067, Q8TEJ5 | Gene names | COL8A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(VIII) chain precursor (Endothelial collagen). | |||||
|
CO9A2_MOUSE
|
||||||
| NC score | 0.023476 (rank : 72) | θ value | 4.03905 (rank : 81) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q07643 | Gene names | Col9a2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-2(IX) chain precursor. | |||||
|
COBA2_HUMAN
|
||||||
| NC score | 0.022522 (rank : 73) | θ value | 1.38821 (rank : 58) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P13942, Q07751, Q13271, Q13272, Q13273, Q99866, Q9UIP9 | Gene names | COL11A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(XI) chain precursor. | |||||
|
GP156_HUMAN
|
||||||
| NC score | 0.022397 (rank : 74) | θ value | 1.38821 (rank : 61) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NFN8, Q86SN6 | Gene names | GPR156, GABABL, PGR28 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 156 (GABAB-related G-protein coupled receptor) (G-protein coupled receptor PGR28). | |||||
|
CO4A3_HUMAN
|
||||||
| NC score | 0.022105 (rank : 75) | θ value | 4.03905 (rank : 80) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 421 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q01955, Q9BQT2 | Gene names | COL4A3 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-3(IV) chain precursor (Goodpasture antigen). | |||||
|
CO4A4_HUMAN
|
||||||
| NC score | 0.021775 (rank : 76) | θ value | 5.27518 (rank : 86) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 440 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P53420 | Gene names | COL4A4 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-4(IV) chain precursor. | |||||
|
CO4A1_MOUSE
|
||||||
| NC score | 0.021198 (rank : 77) | θ value | 8.99809 (rank : 104) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 450 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P02463 | Gene names | Col4a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(IV) chain precursor. | |||||
|
CO1A2_HUMAN
|
||||||
| NC score | 0.020688 (rank : 78) | θ value | 3.0926 (rank : 76) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P08123, P02464, Q9UEB6, Q9UPH0 | Gene names | COL1A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(I) chain precursor. | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.020684 (rank : 79) | θ value | 1.06291 (rank : 57) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
CO4A1_HUMAN
|
||||||
| NC score | 0.020366 (rank : 80) | θ value | 6.88961 (rank : 94) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P02462 | Gene names | COL4A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(IV) chain precursor. | |||||
|
K0562_HUMAN
|
||||||
| NC score | 0.020337 (rank : 81) | θ value | 1.81305 (rank : 69) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O60308, Q5JSQ3, Q5SR24, Q5SR25, Q6PKF5, Q86W32, Q86X14 | Gene names | KIAA0562 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0562. | |||||
|
PCLO_HUMAN
|
||||||
| NC score | 0.019387 (rank : 82) | θ value | 1.38821 (rank : 62) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
CO9A1_MOUSE
|
||||||
| NC score | 0.018583 (rank : 83) | θ value | 6.88961 (rank : 95) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 477 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q05722, Q61269, Q61270, Q61433, Q61940 | Gene names | Col9a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(IX) chain precursor. | |||||
|
CO9A1_HUMAN
|
||||||
| NC score | 0.017831 (rank : 84) | θ value | 8.99809 (rank : 105) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P20849, Q13699, Q13700, Q5TF52, Q6P467, Q96BM8, Q99225, Q9H151, Q9H152, Q9Y6P2, Q9Y6P3 | Gene names | COL9A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(IX) chain precursor. | |||||
|
TRRAP_HUMAN
|
||||||
| NC score | 0.016281 (rank : 85) | θ value | 8.99809 (rank : 113) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y4A5, O75218, Q9Y631, Q9Y6H4 | Gene names | TRRAP, PAF400 | |||
|
Domain Architecture |

|
|||||
| Description | Transformation/transcription domain-associated protein (350/400 kDa PCAF-associated factor) (PAF350/400) (STAF40) (Tra1 homolog). | |||||
|
IASPP_MOUSE
|
||||||
| NC score | 0.015847 (rank : 86) | θ value | 2.36792 (rank : 73) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1024 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q5I1X5 | Gene names | Ppp1r13l, Nkip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RelA-associated inhibitor (Inhibitor of ASPP protein) (Protein iASPP) (PPP1R13B-like protein) (NFkB-interacting protein 1). | |||||
|
NDF2_HUMAN
|
||||||
| NC score | 0.015440 (rank : 87) | θ value | 0.62314 (rank : 54) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q15784, Q8TBI7, Q9UQC6 | Gene names | NEUROD2, NDRF | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic differentiation factor 2 (NeuroD2) (NeuroD-related factor) (NDRF). | |||||
|
IQEC3_MOUSE
|
||||||
| NC score | 0.015300 (rank : 88) | θ value | 8.99809 (rank : 108) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q3TES0, Q3UHP8, Q80TJ8 | Gene names | Iqsec3, Kiaa1110 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 3. | |||||
|
TRRAP_MOUSE
|
||||||
| NC score | 0.014914 (rank : 89) | θ value | 8.99809 (rank : 114) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80YV3, Q8C0Z5, Q8K104 | Gene names | Trrap | |||
|
Domain Architecture |

|
|||||
| Description | Transformation/transcription domain-associated protein (Tra1 homolog). | |||||
|
IASPP_HUMAN
|
||||||
| NC score | 0.013663 (rank : 90) | θ value | 6.88961 (rank : 99) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 688 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8WUF5, Q2PNZ9, Q5DU71, Q5I1X4, Q6P1R7, Q6PKF8, Q9Y290 | Gene names | PPP1R13L, IASPP, NKIP1, PPP1R13BL, RAI | |||
|
Domain Architecture |

|
|||||
| Description | RelA-associated inhibitor (Inhibitor of ASPP protein) (Protein iASPP) (PPP1R13B-like protein) (NFkB-interacting protein 1). | |||||
|
NFH_HUMAN
|
||||||
| NC score | 0.013467 (rank : 91) | θ value | 8.99809 (rank : 110) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
UBP42_HUMAN
|
||||||
| NC score | 0.012896 (rank : 92) | θ value | 8.99809 (rank : 115) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9H9J4, Q3C166, Q6P9B4 | Gene names | USP42 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 42 (EC 3.1.2.15) (Ubiquitin thioesterase 42) (Ubiquitin-specific-processing protease 42) (Deubiquitinating enzyme 42). | |||||
|
NFH_MOUSE
|
||||||
| NC score | 0.012634 (rank : 93) | θ value | 1.81305 (rank : 71) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
IQEC3_HUMAN
|
||||||
| NC score | 0.012419 (rank : 94) | θ value | 5.27518 (rank : 90) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UPP2, Q8TB43 | Gene names | IQSEC3, KIAA1110 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 3. | |||||
|
HS90A_HUMAN
|
||||||
| NC score | 0.011809 (rank : 95) | θ value | 5.27518 (rank : 88) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P07900, Q2PP14, Q9BVQ5 | Gene names | HSP90AA1, HSP90A, HSPC1, HSPCA | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock protein HSP 90-alpha (HSP 86) (NY-REN-38 antigen). | |||||
|
CXA1_HUMAN
|
||||||
| NC score | 0.011485 (rank : 96) | θ value | 1.38821 (rank : 60) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P17302, Q9Y5I8 | Gene names | GJA1, GJAL | |||
|
Domain Architecture |

|
|||||
| Description | Gap junction alpha-1 protein (Connexin-43) (Cx43) (Gap junction 43 kDa heart protein). | |||||
|
HS90A_MOUSE
|
||||||
| NC score | 0.011328 (rank : 97) | θ value | 5.27518 (rank : 89) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P07901 | Gene names | Hsp90aa1, Hsp86, Hsp86-1, Hspca | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock protein HSP 90-alpha (HSP 86) (Tumor-specific transplantation 86 kDa antigen) (TSTA). | |||||
|
SAM68_HUMAN
|
||||||
| NC score | 0.011022 (rank : 98) | θ value | 6.88961 (rank : 101) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q07666, Q6PJX7, Q8NB97, Q99760 | Gene names | KHDRBS1, SAM68 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | KH domain-containing, RNA-binding, signal transduction-associated protein 1 (p21 Ras GTPase-activating protein-associated p62) (GAP- associated tyrosine phosphoprotein p62) (Src-associated in mitosis 68 kDa protein) (Sam68) (p68). | |||||
|
TCEA2_HUMAN
|
||||||
| NC score | 0.010498 (rank : 99) | θ value | 4.03905 (rank : 84) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q15560, Q8TD37, Q8TD38 | Gene names | TCEA2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription elongation factor A protein 2 (Transcription elongation factor S-II protein 2) (Testis-specific S-II) (Transcription elongation factor TFIIS.l). | |||||
|
CCD16_MOUSE
|
||||||
| NC score | 0.010453 (rank : 100) | θ value | 6.88961 (rank : 93) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8R1N0, Q3TR52, Q9CWV9, Q9CYI6 | Gene names | Ccdc16, Omcg1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 16 (Ovus mutant candidate gene 1 protein). | |||||
|
FURIN_MOUSE
|
||||||
| NC score | 0.010176 (rank : 101) | θ value | 1.81305 (rank : 68) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P23188 | Gene names | Furin, Fur, Pcsk3 | |||
|
Domain Architecture |

|
|||||
| Description | Furin precursor (EC 3.4.21.75) (Paired basic amino acid residue cleaving enzyme) (PACE) (Dibasic-processing enzyme) (Prohormone convertase 3). | |||||
|
ARHGG_HUMAN
|
||||||
| NC score | 0.009874 (rank : 102) | θ value | 3.0926 (rank : 75) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5VV41, Q86TF0, Q99434 | Gene names | ARHGEF16, NBR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rho guanine nucleotide exchange factor 16. | |||||
|
SHAN1_HUMAN
|
||||||
| NC score | 0.009207 (rank : 103) | θ value | 8.99809 (rank : 111) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9Y566, Q9NYW9 | Gene names | SHANK1 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 1 (Shank1) (Somatostatin receptor-interacting protein) (SSTR-interacting protein) (SSTRIP). | |||||
|
CXA1_MOUSE
|
||||||
| NC score | 0.009038 (rank : 104) | θ value | 5.27518 (rank : 87) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P23242 | Gene names | Gja1, Cxn-43 | |||
|
Domain Architecture |

|
|||||
| Description | Gap junction alpha-1 protein (Connexin-43) (Cx43) (Gap junction 43 kDa heart protein). | |||||
|
ITPR3_MOUSE
|
||||||
| NC score | 0.008385 (rank : 105) | θ value | 6.88961 (rank : 100) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P70227, Q5DWM4, Q8CED5, Q91Z08 | Gene names | Itpr3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol 1,4,5-trisphosphate receptor type 3 (Type 3 inositol 1,4,5- trisphosphate receptor) (Type 3 InsP3 receptor) (IP3 receptor isoform 3) (InsP3R3). | |||||
|
HXD3_HUMAN
|
||||||
| NC score | 0.006893 (rank : 106) | θ value | 6.88961 (rank : 98) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P31249, Q99955, Q9BSC5 | Gene names | HOXD3, HOX4A | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-D3 (Hox-4A). | |||||
|
US6NL_MOUSE
|
||||||
| NC score | 0.006516 (rank : 107) | θ value | 8.99809 (rank : 116) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80XC3, Q6ZQK8 | Gene names | Usp6nl, Kiaa0019 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USP6 N-terminal-like protein. | |||||
|
PUNC_HUMAN
|
||||||
| NC score | 0.006032 (rank : 108) | θ value | 3.0926 (rank : 79) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 449 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8IVU1, O95215 | Gene names | PUNC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative neuronal cell adhesion molecule precursor. | |||||
|
ABLM1_MOUSE
|
||||||
| NC score | 0.006009 (rank : 109) | θ value | 1.81305 (rank : 63) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8K4G5, Q80U86, Q8BIR9, Q8K4G3, Q8K4G4 | Gene names | Ablim1, Ablim, Kiaa0059 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Actin-binding LIM protein 1 (Actin-binding LIM protein family member 1) (abLIM-1). | |||||
|
HXB3_HUMAN
|
||||||
| NC score | 0.005413 (rank : 110) | θ value | 8.99809 (rank : 106) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 559 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P14651, O95615, P17484 | Gene names | HOXB3, HOX2G | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-B3 (Hox-2G) (Hox-2.7). | |||||
|
HXB3_MOUSE
|
||||||
| NC score | 0.005406 (rank : 111) | θ value | 6.88961 (rank : 97) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 493 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P09026, P10285, Q4PJ06, Q61680 | Gene names | Hoxb3, Hox-2.7, Hoxb-3 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-B3 (Hox-2.7) (Homeobox protein MH-23). | |||||
|
RNF39_HUMAN
|
||||||
| NC score | 0.005204 (rank : 112) | θ value | 5.27518 (rank : 91) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9H2S5, Q5SPM8, Q5SPM9, Q5SPN0, Q5SRJ9, Q5SRK1, Q5SS29, Q9H2S3, Q9H2S4 | Gene names | RNF39, HZFW | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 39 (Protein HZFw). | |||||
|
KIF1A_MOUSE
|
||||||
| NC score | 0.004908 (rank : 113) | θ value | 8.99809 (rank : 109) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P33173, Q61770 | Gene names | Kif1a, Atsv, Kif1 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF1A (Axonal transporter of synaptic vesicles). | |||||
|
HXD3_MOUSE
|
||||||
| NC score | 0.004418 (rank : 114) | θ value | 8.99809 (rank : 107) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P09027, Q3UUD3 | Gene names | Hoxd3, Hox-4.1, Hoxd-3 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-D3 (Hox-4.1) (Homeobox protein MH-19). | |||||
|
ANGL3_MOUSE
|
||||||
| NC score | 0.004330 (rank : 115) | θ value | 5.27518 (rank : 85) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 306 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9R182 | Gene names | Angptl3 | |||
|
Domain Architecture |

|
|||||
| Description | Angiopoietin-related protein 3 precursor (Angiopoietin-like 3). | |||||
|
ULK2_HUMAN
|
||||||
| NC score | 0.000849 (rank : 116) | θ value | 2.36792 (rank : 74) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 979 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8IYT8, O75119 | Gene names | ULK2, KIAA0623 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase ULK2 (EC 2.7.11.1) (Unc-51-like kinase 2). | |||||
|
OBSCN_HUMAN
|
||||||
| NC score | 0.000399 (rank : 117) | θ value | 3.0926 (rank : 78) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1931 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q5VST9, Q2A664, Q5T7G8, Q5T7G9, Q5VSU2, Q86YC7, Q8NHN0, Q8NHN1, Q8NHN2, Q8NHN4, Q8NHN5, Q8NHN6, Q8NHN7, Q8NHN8, Q8NHN9, Q96AA2, Q9HCD3, Q9HCL6 | Gene names | OBSCN, KIAA1556, KIAA1639 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Obscurin (Obscurin-myosin light chain kinase) (Obscurin-MLCK) (Obscurin-RhoGEF). | |||||
|
ERBB3_MOUSE
|
||||||
| NC score | -0.000263 (rank : 118) | θ value | 4.03905 (rank : 82) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 898 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61526, Q3KQR1, Q68J64, Q810U8, Q8K317 | Gene names | Erbb3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Receptor tyrosine-protein kinase erbB-3 precursor (EC 2.7.10.1) (c- erbB3) (Glial growth factor receptor). | |||||
|
ERBB3_HUMAN
|
||||||
| NC score | -0.000707 (rank : 119) | θ value | 6.88961 (rank : 96) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 918 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P21860 | Gene names | ERBB3, HER3 | |||
|
Domain Architecture |

|
|||||
| Description | Receptor tyrosine-protein kinase erbB-3 precursor (EC 2.7.10.1) (c- erbB3) (Tyrosine kinase-type cell surface receptor HER3). | |||||
|
ZN516_MOUSE
|
||||||
| NC score | -0.001197 (rank : 120) | θ value | 6.88961 (rank : 103) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 1197 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q7TSH3 | Gene names | Znf516, Kiaa0222, Zfp516 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 516. | |||||
|
ZN692_HUMAN
|
||||||
| NC score | -0.001519 (rank : 121) | θ value | 5.27518 (rank : 92) | |||
| Query Neighborhood Hits | 116 | Target Neighborhood Hits | 808 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9BU19, Q5SRA5, Q5SRA6, Q9HBC9, Q9NW93, Q9NWY6, Q9UF97 | Gene names | ZNF692 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 692. | |||||