Please be patient as the page loads
|
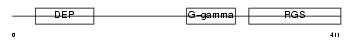
RGS11_HUMAN
|
||||||
| SwissProt Accessions | O94810, O75883 | Gene names | RGS11 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 11 (RGS11). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
RGS11_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | O94810, O75883 | Gene names | RGS11 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 11 (RGS11). | |||||
|
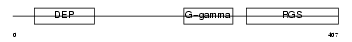
RGS9_MOUSE
|
||||||
| θ value | 2.97304e-136 (rank : 2) | NC score | 0.981415 (rank : 2) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O54828, Q9Z0S0 | Gene names | Rgs9 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 9 (RGS9). | |||||
|
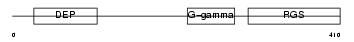
RGS9_HUMAN
|
||||||
| θ value | 3.28708e-135 (rank : 3) | NC score | 0.981321 (rank : 3) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | O75916, O75573, Q696R2, Q8TD64, Q8TD65, Q9HC32, Q9HC33 | Gene names | RGS9 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 9 (RGS9). | |||||
|
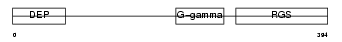
RGS7_HUMAN
|
||||||
| θ value | 1.89267e-82 (rank : 4) | NC score | 0.964253 (rank : 4) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P49802, Q8TD66, Q8TD67, Q9UNU7, Q9Y6B9 | Gene names | RGS7 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 7 (RGS7). | |||||
|
RGS7_MOUSE
|
||||||
| θ value | 2.47191e-82 (rank : 5) | NC score | 0.963996 (rank : 5) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | O54829 | Gene names | Rgs7 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 7 (RGS7). | |||||
|
RGS6_MOUSE
|
||||||
| θ value | 4.67263e-73 (rank : 6) | NC score | 0.957923 (rank : 6) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9Z2H2, Q8BGI8 | Gene names | Rgs6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of G-protein signaling 6 (RGS6). | |||||
|
RGS6_HUMAN
|
||||||
| θ value | 1.27248e-70 (rank : 7) | NC score | 0.948344 (rank : 7) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P49758, O75576, O75577, Q8TE13, Q8TE14, Q8TE15, Q8TE16, Q8TE17, Q8TE18, Q8TE19, Q8TE20, Q8TE21, Q8TE22, Q9Y245, Q9Y647 | Gene names | RGS6 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 6 (RGS6) (S914). | |||||
|
RGS19_HUMAN
|
||||||
| θ value | 2.20094e-22 (rank : 8) | NC score | 0.868230 (rank : 13) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P49795, Q53XN0, Q8TD60 | Gene names | RGS19, GAIP, GNAI3IP | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 19 (RGS19) (G-alpha-interacting protein) (GAIP protein). | |||||
|
RGS20_HUMAN
|
||||||
| θ value | 2.87452e-22 (rank : 9) | NC score | 0.869944 (rank : 10) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O76081, Q96BG9 | Gene names | RGS20, RGSZ1, ZGAP1 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 20 (RGS20) (Regulator of Gz-selective protein signaling 1) (Gz-selective GTPase-activating protein) (G(z)GAP). | |||||
|
RGS1_HUMAN
|
||||||
| θ value | 1.86321e-21 (rank : 10) | NC score | 0.868811 (rank : 12) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q08116, Q07918, Q9H1W2 | Gene names | RGS1, 1R20, BL34 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 1 (RGS1) (Early response protein 1R20) (B-cell activation protein BL34). | |||||
|
RGS19_MOUSE
|
||||||
| θ value | 2.43343e-21 (rank : 11) | NC score | 0.867916 (rank : 14) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9CX84, Q99L50 | Gene names | Rgs19 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 19 (RGS19). | |||||
|
RGS20_MOUSE
|
||||||
| θ value | 2.43343e-21 (rank : 12) | NC score | 0.869888 (rank : 11) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9QZB1, Q3TY63, Q9CUV8, Q9QZB2 | Gene names | Rgs20, Rgsz1 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 20 (RGS20) (Regulator of G-protein signaling Z1). | |||||
|
RGS12_HUMAN
|
||||||
| θ value | 3.51386e-20 (rank : 13) | NC score | 0.737060 (rank : 34) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O14924, O14922, O14923, O43510, O75338 | Gene names | RGS12 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 12 (RGS12). | |||||
|
RGS14_HUMAN
|
||||||
| θ value | 5.99374e-20 (rank : 14) | NC score | 0.817982 (rank : 32) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O43566, O43565, Q8TD62 | Gene names | RGS14 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 14 (RGS14). | |||||
|
RGS8_MOUSE
|
||||||
| θ value | 5.99374e-20 (rank : 15) | NC score | 0.854671 (rank : 22) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8BXT1 | Gene names | Rgs8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of G-protein signaling 8 (RGS8). | |||||
|
RGS3_HUMAN
|
||||||
| θ value | 1.02238e-19 (rank : 16) | NC score | 0.602781 (rank : 35) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
RGS8_HUMAN
|
||||||
| θ value | 1.02238e-19 (rank : 17) | NC score | 0.855389 (rank : 21) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P57771, Q3SYD2 | Gene names | RGS8 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 8 (RGS8). | |||||
|
RGS18_HUMAN
|
||||||
| θ value | 1.74391e-19 (rank : 18) | NC score | 0.857787 (rank : 19) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9NS28 | Gene names | RGS18, RGS13 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 18 (RGS18). | |||||
|
RGS17_MOUSE
|
||||||
| θ value | 5.07402e-19 (rank : 19) | NC score | 0.864513 (rank : 17) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9QZB0, Q543T9 | Gene names | Rgs17, Rgsz2 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 17 (RGS17) (Regulator of Gz-selective protein signaling 2). | |||||
|
RGS17_HUMAN
|
||||||
| θ value | 6.62687e-19 (rank : 20) | NC score | 0.865608 (rank : 16) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9UGC6, Q5TF49, Q8TD61, Q9UJS8 | Gene names | RGS17 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 17 (RGS17). | |||||
|
RGS3_MOUSE
|
||||||
| θ value | 8.65492e-19 (rank : 21) | NC score | 0.537198 (rank : 40) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
RGS14_MOUSE
|
||||||
| θ value | 1.13037e-18 (rank : 22) | NC score | 0.817883 (rank : 33) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P97492, Q9DCD1 | Gene names | Rgs14 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 14 (RGS14) (RAP1/RAP2-interacting protein). | |||||
|
RGS1_MOUSE
|
||||||
| θ value | 2.5182e-18 (rank : 23) | NC score | 0.866421 (rank : 15) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9JL25, Q3TBD1, Q3TC18, Q3TD56 | Gene names | Rgs1 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 1 (RGS1). | |||||
|
RGS18_MOUSE
|
||||||
| θ value | 3.28887e-18 (rank : 24) | NC score | 0.852239 (rank : 23) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q99PG4 | Gene names | Rgs18 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 18 (RGS18). | |||||
|
RGS5_MOUSE
|
||||||
| θ value | 7.32683e-18 (rank : 25) | NC score | 0.863064 (rank : 18) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O08850, Q543B1, Q9D0Z2 | Gene names | Rgs5 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 5 (RGS5). | |||||
|
RGS5_HUMAN
|
||||||
| θ value | 3.63628e-17 (rank : 26) | NC score | 0.856326 (rank : 20) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O15539 | Gene names | RGS5 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 5 (RGS5). | |||||
|
RGS10_HUMAN
|
||||||
| θ value | 1.058e-16 (rank : 27) | NC score | 0.870100 (rank : 9) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O43665 | Gene names | RGS10 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 10 (RGS10). | |||||
|
RGS10_MOUSE
|
||||||
| θ value | 1.38178e-16 (rank : 28) | NC score | 0.870717 (rank : 8) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9CQE5, Q9D3L2 | Gene names | Rgs10 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 10 (RGS10). | |||||
|
RGS4_HUMAN
|
||||||
| θ value | 1.38178e-16 (rank : 29) | NC score | 0.847024 (rank : 26) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P49798 | Gene names | RGS4 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 4 (RGS4) (RGP4). | |||||
|
RGS4_MOUSE
|
||||||
| θ value | 2.35696e-16 (rank : 30) | NC score | 0.849714 (rank : 25) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O08899, Q99L30 | Gene names | Rgs4 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 4 (RGS4). | |||||
|
RGS13_HUMAN
|
||||||
| θ value | 3.07829e-16 (rank : 31) | NC score | 0.851429 (rank : 24) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O14921, Q6PGR2, Q8TD63, Q9BX45 | Gene names | RGS13 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 13 (RGS13). | |||||
|
RGS16_HUMAN
|
||||||
| θ value | 5.25075e-16 (rank : 32) | NC score | 0.844977 (rank : 28) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O15492, Q99701 | Gene names | RGS16, RGSR | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 16 (RGS16) (Retinally abundant regulator of G-protein signaling) (RGS-R) (A28-RGS14P). | |||||
|
RGS13_MOUSE
|
||||||
| θ value | 1.52774e-15 (rank : 33) | NC score | 0.846602 (rank : 27) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8K443 | Gene names | Rgs13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of G-protein signaling 13 (RGS13). | |||||
|
RGS16_MOUSE
|
||||||
| θ value | 4.44505e-15 (rank : 34) | NC score | 0.839533 (rank : 30) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P97428, O09091, P97420, Q80V16 | Gene names | Rgs16, Rgsr | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 16 (RGS16) (Retinally abundant regulator of G-protein signaling) (RGS-R) (A28-RGS14P). | |||||
|
RGS2_MOUSE
|
||||||
| θ value | 1.29331e-14 (rank : 35) | NC score | 0.838842 (rank : 31) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O08849, Q544S7, Q91WX1, Q9JL24 | Gene names | Rgs2 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 2 (RGS2). | |||||
|
RGS2_HUMAN
|
||||||
| θ value | 2.88119e-14 (rank : 36) | NC score | 0.839864 (rank : 29) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P41220 | Gene names | RGS2, G0S8 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 2 (RGS2) (G0/G1 switch regulatory protein 8). | |||||
|
AXN2_HUMAN
|
||||||
| θ value | 1.69304e-06 (rank : 37) | NC score | 0.557248 (rank : 37) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9Y2T1, Q9H3M6, Q9UH84 | Gene names | AXIN2 | |||
|
Domain Architecture |

|
|||||
| Description | Axin-2 (Axis inhibition protein 2) (Conductin) (Axin-like protein) (Axil). | |||||
|
AXN1_MOUSE
|
||||||
| θ value | 2.88788e-06 (rank : 38) | NC score | 0.569785 (rank : 36) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | O35625 | Gene names | Axin1, Axin, Fu | |||
|
Domain Architecture |

|
|||||
| Description | Axin-1 (Axis inhibition protein 1) (Fused protein). | |||||
|
AXN2_MOUSE
|
||||||
| θ value | 2.88788e-06 (rank : 39) | NC score | 0.551682 (rank : 39) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O88566, Q9QXJ6 | Gene names | Axin2 | |||
|
Domain Architecture |

|
|||||
| Description | Axin-2 (Axis inhibition protein 2) (Conductin) (Axin-like protein) (Axil). | |||||
|
SNX13_HUMAN
|
||||||
| θ value | 4.92598e-06 (rank : 40) | NC score | 0.313304 (rank : 42) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9Y5W8, O94821, Q8WVZ2, Q8WXH8 | Gene names | SNX13, KIAA0713 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-13 (RGS domain- and PHOX domain-containing protein) (RGS-PX1). | |||||
|
SNX13_MOUSE
|
||||||
| θ value | 1.43324e-05 (rank : 41) | NC score | 0.314744 (rank : 41) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q6PHS6 | Gene names | Snx13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sorting nexin-13. | |||||
|
PLEK_MOUSE
|
||||||
| θ value | 3.19293e-05 (rank : 42) | NC score | 0.114629 (rank : 53) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9JHK5, Q9ERI9 | Gene names | Plek | |||
|
Domain Architecture |
|
|||||
| Description | Pleckstrin. | |||||
|
AXN1_HUMAN
|
||||||
| θ value | 5.44631e-05 (rank : 43) | NC score | 0.553728 (rank : 38) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | O15169, Q4TT26, Q4TT27, Q86YA7, Q8WVW6, Q96S28 | Gene names | AXIN1, AXIN | |||
|
Domain Architecture |

|
|||||
| Description | Axin-1 (Axis inhibition protein 1) (hAxin). | |||||
|
DVL1L_HUMAN
|
||||||
| θ value | 0.000602161 (rank : 44) | NC score | 0.125051 (rank : 48) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P54792 | Gene names | DVL1L1, DVL, DVL1 | |||
|
Domain Architecture |
|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1-like (Dishevelled- 1-like) (DSH homolog 1-like). | |||||
|
SNX25_HUMAN
|
||||||
| θ value | 0.000786445 (rank : 45) | NC score | 0.236135 (rank : 43) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9H3E2 | Gene names | SNX25 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-25. | |||||
|
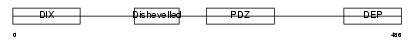
DVL1_MOUSE
|
||||||
| θ value | 0.00298849 (rank : 46) | NC score | 0.124330 (rank : 49) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P51141, Q60868 | Gene names | Dvl1, Dvl | |||
|
Domain Architecture |
|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1 (Dishevelled-1) (DSH homolog 1). | |||||
|
DVL1_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 47) | NC score | 0.124282 (rank : 50) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O14640 | Gene names | DVL1 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1 (Dishevelled-1) (DSH homolog 1). | |||||
|
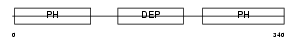
PLEK_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 48) | NC score | 0.088106 (rank : 54) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P08567, Q53SU8, Q6FGM8, Q8WV81 | Gene names | PLEK, P47 | |||
|
Domain Architecture |
|
|||||
| Description | Pleckstrin (Platelet p47 protein). | |||||
|
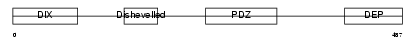
DVL3_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 49) | NC score | 0.144997 (rank : 47) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q92997, O14642, Q13531, Q8N5E9, Q92607 | Gene names | DVL3, KIAA0208 | |||
|
Domain Architecture |
|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-3 (Dishevelled-3) (DSH homolog 3). | |||||
|
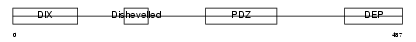
DVL3_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 50) | NC score | 0.145032 (rank : 46) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q61062 | Gene names | Dvl3 | |||
|
Domain Architecture |
|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-3 (Dishevelled-3) (DSH homolog 3). | |||||
|
DVL2_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 51) | NC score | 0.122897 (rank : 51) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O14641 | Gene names | DVL2 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-2 (Dishevelled-2) (DSH homolog 2). | |||||
|
DVL2_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 52) | NC score | 0.122002 (rank : 52) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q60838 | Gene names | Dvl2 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-2 (Dishevelled-2) (DSH homolog 2). | |||||
|
PREX1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 53) | NC score | 0.042264 (rank : 57) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8TCU6, Q5JS95, Q5JS96, Q69YL2, Q7Z2L9, Q9BQH0, Q9BX55, Q9H4Q6, Q9P2D2, Q9UGQ4 | Gene names | PREX1, KIAA1415 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 3,4,5-trisphosphate-dependent Rac exchanger 1 protein (P-Rex1 protein). | |||||
|
VRK3_MOUSE
|
||||||
| θ value | 1.06291 (rank : 54) | NC score | 0.011271 (rank : 64) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8K3G5, Q921W6 | Gene names | Vrk3 | |||
|
Domain Architecture |
|
|||||
| Description | Serine/threonine-protein kinase VRK3 (EC 2.7.11.1) (Vaccinia-related kinase 3). | |||||
|
AMGO1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 55) | NC score | 0.006615 (rank : 67) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 353 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q86WK6, Q8IW71, Q9ULQ7 | Gene names | AMIGO1, ALI2, AMIGO, KIAA1163 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amphoterin-induced protein 1 precursor (AMIGO-1) (Alivin-2). | |||||
|
GBG5_HUMAN
|
||||||
| θ value | 1.81305 (rank : 56) | NC score | 0.020655 (rank : 58) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P63218, P30670, Q5VX54, Q61015 | Gene names | GNG5, GNGT5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Guanine nucleotide-binding protein G(I)/G(S)/G(O) gamma-5 subunit precursor. | |||||
|
GBG5_MOUSE
|
||||||
| θ value | 1.81305 (rank : 57) | NC score | 0.020528 (rank : 59) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80SZ7, P30670, Q5I0W4, Q61015 | Gene names | Gng5, Gngt5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Guanine nucleotide-binding protein G(I)/G(S)/G(O) gamma-5 subunit precursor. | |||||
|
AMGO1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 58) | NC score | 0.006167 (rank : 69) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80ZD8, Q69ZQ0, Q8R5D4, Q921U9 | Gene names | Amigo1, Ali2, Amigo, Kiaa1163 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amphoterin-induced protein 1 precursor (AMIGO-1) (Alivin-2). | |||||
|
ZN638_HUMAN
|
||||||
| θ value | 3.0926 (rank : 59) | NC score | 0.010352 (rank : 65) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14966, Q53R34, Q5XJ05, Q68DP3, Q6P2H2, Q7Z3T7, Q8NF92, Q8TCA1, Q9H2G1, Q9NP37 | Gene names | ZNF638, NP220, ZFML | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 638 (Nuclear protein 220) (Zinc-finger matrin-like protein) (Cutaneous T-cell lymphoma-associated antigen se33-1) (CTCL tumor antigen se33-1). | |||||
|
HIC1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 60) | NC score | -0.001829 (rank : 72) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 863 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9R1Y5, Q9R1Y6, Q9R2B0 | Gene names | Hic1 | |||
|
Domain Architecture |
|
|||||
| Description | Hypermethylated in cancer 1 protein (Hic-1). | |||||
|
CXXC6_HUMAN
|
||||||
| θ value | 5.27518 (rank : 61) | NC score | 0.014183 (rank : 60) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NFU7, Q5VUP7, Q7Z6B6, Q8TCR1, Q9C0I7 | Gene names | CXXC6, KIAA1676, LCX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CXXC-type zinc finger protein 6 (Leukemia-associated protein with a CXXC domain). | |||||
|
GBG11_HUMAN
|
||||||
| θ value | 6.88961 (rank : 62) | NC score | 0.013118 (rank : 61) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P61952, P50152 | Gene names | GNG11, GNGT11 | |||
|
Domain Architecture |

|
|||||
| Description | Guanine nucleotide-binding protein G(I)/G(S)/G(O) gamma-11 subunit precursor. | |||||
|
GBG11_MOUSE
|
||||||
| θ value | 6.88961 (rank : 63) | NC score | 0.013118 (rank : 62) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P61953, P50152, Q4QRN2 | Gene names | Gng11, Gngt11 | |||
|
Domain Architecture |

|
|||||
| Description | Guanine nucleotide-binding protein G(I)/G(S)/G(O) gamma-11 subunit precursor. | |||||
|
TTLL4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 64) | NC score | 0.005773 (rank : 70) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q14679, Q8WW29 | Gene names | TTLL4, KIAA0173 | |||
|
Domain Architecture |

|
|||||
| Description | Tubulin--tyrosine ligase-like protein 4. | |||||
|
WN10A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 65) | NC score | 0.001946 (rank : 71) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9GZT5, Q96TA7, Q9H7S8 | Gene names | WNT10A | |||
|
Domain Architecture |
|
|||||
| Description | Protein Wnt-10a precursor. | |||||
|
AKAP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 66) | NC score | 0.009798 (rank : 66) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q92667, Q13320, Q9BW14 | Gene names | AKAP1, AKAP149 | |||
|
Domain Architecture |

|
|||||
| Description | A kinase anchor protein 1, mitochondrial precursor (Protein kinase A- anchoring protein 1) (PRKA1) (A-kinase anchor protein 149 kDa) (AKAP 149) (Dual specificity A-kinase-anchoring protein 1) (D-AKAP-1) (Spermatid A-kinase anchor protein 84) (S-AKAP84). | |||||
|
AKAP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 67) | NC score | 0.012196 (rank : 63) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O08715, O08714, P97488 | Gene names | Akap1, Akap | |||
|
Domain Architecture |

|
|||||
| Description | A kinase anchor protein 1, mitochondrial precursor (Protein kinase A- anchoring protein 1) (PRKA1) (Dual specificity A-kinase-anchoring protein 1) (D-AKAP-1) (Spermatid A-kinase anchor protein) (S-AKAP). | |||||
|
HES2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 68) | NC score | 0.006318 (rank : 68) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y543, Q96EN4, Q9Y542 | Gene names | HES2 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor HES-2 (Hairy and enhancer of split 2). | |||||
|
AKA10_HUMAN
|
||||||
| θ value | θ > 10 (rank : 69) | NC score | 0.206760 (rank : 44) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O43572, Q96AJ7 | Gene names | AKAP10 | |||
|
Domain Architecture |

|
|||||
| Description | A kinase anchor protein 10, mitochondrial precursor (Protein kinase A- anchoring protein 10) (PRKA10) (Dual specificity A kinase-anchoring protein 2) (D-AKAP-2). | |||||
|
AKA10_MOUSE
|
||||||
| θ value | θ > 10 (rank : 70) | NC score | 0.206377 (rank : 45) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O88845, Q5SUB5, Q5SUB7, Q7TPE7, Q8BQL6 | Gene names | Akap10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A kinase anchor protein 10, mitochondrial precursor (Protein kinase A- anchoring protein 10) (PRKA10) (Dual specificity A kinase-anchoring protein 2) (D-AKAP-2). | |||||
|
SNX14_HUMAN
|
||||||
| θ value | θ > 10 (rank : 71) | NC score | 0.050603 (rank : 56) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Y5W7, Q9BSD1 | Gene names | SNX14 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-14. | |||||
|
SNX14_MOUSE
|
||||||
| θ value | θ > 10 (rank : 72) | NC score | 0.061325 (rank : 55) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BHY8 | Gene names | Snx14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sorting nexin-14. | |||||
|
RGS11_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | O94810, O75883 | Gene names | RGS11 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 11 (RGS11). | |||||
|
RGS9_MOUSE
|
||||||
| NC score | 0.981415 (rank : 2) | θ value | 2.97304e-136 (rank : 2) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O54828, Q9Z0S0 | Gene names | Rgs9 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 9 (RGS9). | |||||
|
RGS9_HUMAN
|
||||||
| NC score | 0.981321 (rank : 3) | θ value | 3.28708e-135 (rank : 3) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | O75916, O75573, Q696R2, Q8TD64, Q8TD65, Q9HC32, Q9HC33 | Gene names | RGS9 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 9 (RGS9). | |||||
|
RGS7_HUMAN
|
||||||
| NC score | 0.964253 (rank : 4) | θ value | 1.89267e-82 (rank : 4) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P49802, Q8TD66, Q8TD67, Q9UNU7, Q9Y6B9 | Gene names | RGS7 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 7 (RGS7). | |||||
|
RGS7_MOUSE
|
||||||
| NC score | 0.963996 (rank : 5) | θ value | 2.47191e-82 (rank : 5) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | O54829 | Gene names | Rgs7 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 7 (RGS7). | |||||
|
RGS6_MOUSE
|
||||||
| NC score | 0.957923 (rank : 6) | θ value | 4.67263e-73 (rank : 6) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9Z2H2, Q8BGI8 | Gene names | Rgs6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of G-protein signaling 6 (RGS6). | |||||
|
RGS6_HUMAN
|
||||||
| NC score | 0.948344 (rank : 7) | θ value | 1.27248e-70 (rank : 7) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P49758, O75576, O75577, Q8TE13, Q8TE14, Q8TE15, Q8TE16, Q8TE17, Q8TE18, Q8TE19, Q8TE20, Q8TE21, Q8TE22, Q9Y245, Q9Y647 | Gene names | RGS6 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 6 (RGS6) (S914). | |||||
|
RGS10_MOUSE
|
||||||
| NC score | 0.870717 (rank : 8) | θ value | 1.38178e-16 (rank : 28) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9CQE5, Q9D3L2 | Gene names | Rgs10 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 10 (RGS10). | |||||
|
RGS10_HUMAN
|
||||||
| NC score | 0.870100 (rank : 9) | θ value | 1.058e-16 (rank : 27) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O43665 | Gene names | RGS10 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 10 (RGS10). | |||||
|
RGS20_HUMAN
|
||||||
| NC score | 0.869944 (rank : 10) | θ value | 2.87452e-22 (rank : 9) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O76081, Q96BG9 | Gene names | RGS20, RGSZ1, ZGAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 20 (RGS20) (Regulator of Gz-selective protein signaling 1) (Gz-selective GTPase-activating protein) (G(z)GAP). | |||||
|
RGS20_MOUSE
|
||||||
| NC score | 0.869888 (rank : 11) | θ value | 2.43343e-21 (rank : 12) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9QZB1, Q3TY63, Q9CUV8, Q9QZB2 | Gene names | Rgs20, Rgsz1 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 20 (RGS20) (Regulator of G-protein signaling Z1). | |||||
|
RGS1_HUMAN
|
||||||
| NC score | 0.868811 (rank : 12) | θ value | 1.86321e-21 (rank : 10) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q08116, Q07918, Q9H1W2 | Gene names | RGS1, 1R20, BL34 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 1 (RGS1) (Early response protein 1R20) (B-cell activation protein BL34). | |||||
|
RGS19_HUMAN
|
||||||
| NC score | 0.868230 (rank : 13) | θ value | 2.20094e-22 (rank : 8) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P49795, Q53XN0, Q8TD60 | Gene names | RGS19, GAIP, GNAI3IP | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 19 (RGS19) (G-alpha-interacting protein) (GAIP protein). | |||||
|
RGS19_MOUSE
|
||||||
| NC score | 0.867916 (rank : 14) | θ value | 2.43343e-21 (rank : 11) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9CX84, Q99L50 | Gene names | Rgs19 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 19 (RGS19). | |||||
|
RGS1_MOUSE
|
||||||
| NC score | 0.866421 (rank : 15) | θ value | 2.5182e-18 (rank : 23) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9JL25, Q3TBD1, Q3TC18, Q3TD56 | Gene names | Rgs1 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 1 (RGS1). | |||||
|
RGS17_HUMAN
|
||||||
| NC score | 0.865608 (rank : 16) | θ value | 6.62687e-19 (rank : 20) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9UGC6, Q5TF49, Q8TD61, Q9UJS8 | Gene names | RGS17 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 17 (RGS17). | |||||
|
RGS17_MOUSE
|
||||||
| NC score | 0.864513 (rank : 17) | θ value | 5.07402e-19 (rank : 19) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9QZB0, Q543T9 | Gene names | Rgs17, Rgsz2 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 17 (RGS17) (Regulator of Gz-selective protein signaling 2). | |||||
|
RGS5_MOUSE
|
||||||
| NC score | 0.863064 (rank : 18) | θ value | 7.32683e-18 (rank : 25) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O08850, Q543B1, Q9D0Z2 | Gene names | Rgs5 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 5 (RGS5). | |||||
|
RGS18_HUMAN
|
||||||
| NC score | 0.857787 (rank : 19) | θ value | 1.74391e-19 (rank : 18) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9NS28 | Gene names | RGS18, RGS13 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 18 (RGS18). | |||||
|
RGS5_HUMAN
|
||||||
| NC score | 0.856326 (rank : 20) | θ value | 3.63628e-17 (rank : 26) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O15539 | Gene names | RGS5 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 5 (RGS5). | |||||
|
RGS8_HUMAN
|
||||||
| NC score | 0.855389 (rank : 21) | θ value | 1.02238e-19 (rank : 17) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P57771, Q3SYD2 | Gene names | RGS8 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 8 (RGS8). | |||||
|
RGS8_MOUSE
|
||||||
| NC score | 0.854671 (rank : 22) | θ value | 5.99374e-20 (rank : 15) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8BXT1 | Gene names | Rgs8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of G-protein signaling 8 (RGS8). | |||||
|
RGS18_MOUSE
|
||||||
| NC score | 0.852239 (rank : 23) | θ value | 3.28887e-18 (rank : 24) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q99PG4 | Gene names | Rgs18 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 18 (RGS18). | |||||
|
RGS13_HUMAN
|
||||||
| NC score | 0.851429 (rank : 24) | θ value | 3.07829e-16 (rank : 31) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O14921, Q6PGR2, Q8TD63, Q9BX45 | Gene names | RGS13 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 13 (RGS13). | |||||
|
RGS4_MOUSE
|
||||||
| NC score | 0.849714 (rank : 25) | θ value | 2.35696e-16 (rank : 30) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O08899, Q99L30 | Gene names | Rgs4 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 4 (RGS4). | |||||
|
RGS4_HUMAN
|
||||||
| NC score | 0.847024 (rank : 26) | θ value | 1.38178e-16 (rank : 29) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P49798 | Gene names | RGS4 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 4 (RGS4) (RGP4). | |||||
|
RGS13_MOUSE
|
||||||
| NC score | 0.846602 (rank : 27) | θ value | 1.52774e-15 (rank : 33) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8K443 | Gene names | Rgs13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of G-protein signaling 13 (RGS13). | |||||
|
RGS16_HUMAN
|
||||||
| NC score | 0.844977 (rank : 28) | θ value | 5.25075e-16 (rank : 32) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O15492, Q99701 | Gene names | RGS16, RGSR | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 16 (RGS16) (Retinally abundant regulator of G-protein signaling) (RGS-R) (A28-RGS14P). | |||||
|
RGS2_HUMAN
|
||||||
| NC score | 0.839864 (rank : 29) | θ value | 2.88119e-14 (rank : 36) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P41220 | Gene names | RGS2, G0S8 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 2 (RGS2) (G0/G1 switch regulatory protein 8). | |||||
|
RGS16_MOUSE
|
||||||
| NC score | 0.839533 (rank : 30) | θ value | 4.44505e-15 (rank : 34) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P97428, O09091, P97420, Q80V16 | Gene names | Rgs16, Rgsr | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 16 (RGS16) (Retinally abundant regulator of G-protein signaling) (RGS-R) (A28-RGS14P). | |||||
|
RGS2_MOUSE
|
||||||
| NC score | 0.838842 (rank : 31) | θ value | 1.29331e-14 (rank : 35) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O08849, Q544S7, Q91WX1, Q9JL24 | Gene names | Rgs2 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 2 (RGS2). | |||||
|
RGS14_HUMAN
|
||||||
| NC score | 0.817982 (rank : 32) | θ value | 5.99374e-20 (rank : 14) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O43566, O43565, Q8TD62 | Gene names | RGS14 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 14 (RGS14). | |||||
|
RGS14_MOUSE
|
||||||
| NC score | 0.817883 (rank : 33) | θ value | 1.13037e-18 (rank : 22) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P97492, Q9DCD1 | Gene names | Rgs14 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 14 (RGS14) (RAP1/RAP2-interacting protein). | |||||
|
RGS12_HUMAN
|
||||||
| NC score | 0.737060 (rank : 34) | θ value | 3.51386e-20 (rank : 13) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O14924, O14922, O14923, O43510, O75338 | Gene names | RGS12 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 12 (RGS12). | |||||
|
RGS3_HUMAN
|
||||||
| NC score | 0.602781 (rank : 35) | θ value | 1.02238e-19 (rank : 16) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
AXN1_MOUSE
|
||||||
| NC score | 0.569785 (rank : 36) | θ value | 2.88788e-06 (rank : 38) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | O35625 | Gene names | Axin1, Axin, Fu | |||
|
Domain Architecture |

|
|||||
| Description | Axin-1 (Axis inhibition protein 1) (Fused protein). | |||||
|
AXN2_HUMAN
|
||||||
| NC score | 0.557248 (rank : 37) | θ value | 1.69304e-06 (rank : 37) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9Y2T1, Q9H3M6, Q9UH84 | Gene names | AXIN2 | |||
|
Domain Architecture |

|
|||||
| Description | Axin-2 (Axis inhibition protein 2) (Conductin) (Axin-like protein) (Axil). | |||||
|
AXN1_HUMAN
|
||||||
| NC score | 0.553728 (rank : 38) | θ value | 5.44631e-05 (rank : 43) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | O15169, Q4TT26, Q4TT27, Q86YA7, Q8WVW6, Q96S28 | Gene names | AXIN1, AXIN | |||
|
Domain Architecture |

|
|||||
| Description | Axin-1 (Axis inhibition protein 1) (hAxin). | |||||
|
AXN2_MOUSE
|
||||||
| NC score | 0.551682 (rank : 39) | θ value | 2.88788e-06 (rank : 39) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O88566, Q9QXJ6 | Gene names | Axin2 | |||
|
Domain Architecture |

|
|||||
| Description | Axin-2 (Axis inhibition protein 2) (Conductin) (Axin-like protein) (Axil). | |||||
|
RGS3_MOUSE
|
||||||
| NC score | 0.537198 (rank : 40) | θ value | 8.65492e-19 (rank : 21) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
SNX13_MOUSE
|
||||||
| NC score | 0.314744 (rank : 41) | θ value | 1.43324e-05 (rank : 41) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q6PHS6 | Gene names | Snx13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sorting nexin-13. | |||||
|
SNX13_HUMAN
|
||||||
| NC score | 0.313304 (rank : 42) | θ value | 4.92598e-06 (rank : 40) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9Y5W8, O94821, Q8WVZ2, Q8WXH8 | Gene names | SNX13, KIAA0713 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-13 (RGS domain- and PHOX domain-containing protein) (RGS-PX1). | |||||
|
SNX25_HUMAN
|
||||||
| NC score | 0.236135 (rank : 43) | θ value | 0.000786445 (rank : 45) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9H3E2 | Gene names | SNX25 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-25. | |||||
|
AKA10_HUMAN
|
||||||
| NC score | 0.206760 (rank : 44) | θ value | θ > 10 (rank : 69) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O43572, Q96AJ7 | Gene names | AKAP10 | |||
|
Domain Architecture |

|
|||||
| Description | A kinase anchor protein 10, mitochondrial precursor (Protein kinase A- anchoring protein 10) (PRKA10) (Dual specificity A kinase-anchoring protein 2) (D-AKAP-2). | |||||
|
AKA10_MOUSE
|
||||||
| NC score | 0.206377 (rank : 45) | θ value | θ > 10 (rank : 70) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O88845, Q5SUB5, Q5SUB7, Q7TPE7, Q8BQL6 | Gene names | Akap10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A kinase anchor protein 10, mitochondrial precursor (Protein kinase A- anchoring protein 10) (PRKA10) (Dual specificity A kinase-anchoring protein 2) (D-AKAP-2). | |||||
|
DVL3_MOUSE
|
||||||
| NC score | 0.145032 (rank : 46) | θ value | 0.0113563 (rank : 50) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q61062 | Gene names | Dvl3 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-3 (Dishevelled-3) (DSH homolog 3). | |||||
|
DVL3_HUMAN
|
||||||
| NC score | 0.144997 (rank : 47) | θ value | 0.0113563 (rank : 49) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q92997, O14642, Q13531, Q8N5E9, Q92607 | Gene names | DVL3, KIAA0208 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-3 (Dishevelled-3) (DSH homolog 3). | |||||
|
DVL1L_HUMAN
|
||||||
| NC score | 0.125051 (rank : 48) | θ value | 0.000602161 (rank : 44) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P54792 | Gene names | DVL1L1, DVL, DVL1 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1-like (Dishevelled- 1-like) (DSH homolog 1-like). | |||||
|
DVL1_MOUSE
|
||||||
| NC score | 0.124330 (rank : 49) | θ value | 0.00298849 (rank : 46) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P51141, Q60868 | Gene names | Dvl1, Dvl | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1 (Dishevelled-1) (DSH homolog 1). | |||||
|
DVL1_HUMAN
|
||||||
| NC score | 0.124282 (rank : 50) | θ value | 0.00390308 (rank : 47) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O14640 | Gene names | DVL1 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1 (Dishevelled-1) (DSH homolog 1). | |||||
|
DVL2_HUMAN
|
||||||
| NC score | 0.122897 (rank : 51) | θ value | 0.0736092 (rank : 51) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O14641 | Gene names | DVL2 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-2 (Dishevelled-2) (DSH homolog 2). | |||||
|
DVL2_MOUSE
|
||||||
| NC score | 0.122002 (rank : 52) | θ value | 0.0736092 (rank : 52) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q60838 | Gene names | Dvl2 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-2 (Dishevelled-2) (DSH homolog 2). | |||||
|
PLEK_MOUSE
|
||||||
| NC score | 0.114629 (rank : 53) | θ value | 3.19293e-05 (rank : 42) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9JHK5, Q9ERI9 | Gene names | Plek | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin. | |||||
|
PLEK_HUMAN
|
||||||
| NC score | 0.088106 (rank : 54) | θ value | 0.00869519 (rank : 48) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P08567, Q53SU8, Q6FGM8, Q8WV81 | Gene names | PLEK, P47 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin (Platelet p47 protein). | |||||
|
SNX14_MOUSE
|
||||||
| NC score | 0.061325 (rank : 55) | θ value | θ > 10 (rank : 72) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BHY8 | Gene names | Snx14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sorting nexin-14. | |||||
|
SNX14_HUMAN
|
||||||
| NC score | 0.050603 (rank : 56) | θ value | θ > 10 (rank : 71) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Y5W7, Q9BSD1 | Gene names | SNX14 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-14. | |||||
|
PREX1_HUMAN
|
||||||
| NC score | 0.042264 (rank : 57) | θ value | 0.813845 (rank : 53) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8TCU6, Q5JS95, Q5JS96, Q69YL2, Q7Z2L9, Q9BQH0, Q9BX55, Q9H4Q6, Q9P2D2, Q9UGQ4 | Gene names | PREX1, KIAA1415 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 3,4,5-trisphosphate-dependent Rac exchanger 1 protein (P-Rex1 protein). | |||||
|
GBG5_HUMAN
|
||||||
| NC score | 0.020655 (rank : 58) | θ value | 1.81305 (rank : 56) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P63218, P30670, Q5VX54, Q61015 | Gene names | GNG5, GNGT5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Guanine nucleotide-binding protein G(I)/G(S)/G(O) gamma-5 subunit precursor. | |||||
|
GBG5_MOUSE
|
||||||
| NC score | 0.020528 (rank : 59) | θ value | 1.81305 (rank : 57) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80SZ7, P30670, Q5I0W4, Q61015 | Gene names | Gng5, Gngt5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Guanine nucleotide-binding protein G(I)/G(S)/G(O) gamma-5 subunit precursor. | |||||
|
CXXC6_HUMAN
|
||||||
| NC score | 0.014183 (rank : 60) | θ value | 5.27518 (rank : 61) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NFU7, Q5VUP7, Q7Z6B6, Q8TCR1, Q9C0I7 | Gene names | CXXC6, KIAA1676, LCX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CXXC-type zinc finger protein 6 (Leukemia-associated protein with a CXXC domain). | |||||
|
GBG11_HUMAN
|
||||||
| NC score | 0.013118 (rank : 61) | θ value | 6.88961 (rank : 62) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P61952, P50152 | Gene names | GNG11, GNGT11 | |||
|
Domain Architecture |

|
|||||
| Description | Guanine nucleotide-binding protein G(I)/G(S)/G(O) gamma-11 subunit precursor. | |||||
|
GBG11_MOUSE
|
||||||
| NC score | 0.013118 (rank : 62) | θ value | 6.88961 (rank : 63) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P61953, P50152, Q4QRN2 | Gene names | Gng11, Gngt11 | |||
|
Domain Architecture |

|
|||||
| Description | Guanine nucleotide-binding protein G(I)/G(S)/G(O) gamma-11 subunit precursor. | |||||
|
AKAP1_MOUSE
|
||||||
| NC score | 0.012196 (rank : 63) | θ value | 8.99809 (rank : 67) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O08715, O08714, P97488 | Gene names | Akap1, Akap | |||
|
Domain Architecture |

|
|||||
| Description | A kinase anchor protein 1, mitochondrial precursor (Protein kinase A- anchoring protein 1) (PRKA1) (Dual specificity A-kinase-anchoring protein 1) (D-AKAP-1) (Spermatid A-kinase anchor protein) (S-AKAP). | |||||
|
VRK3_MOUSE
|
||||||
| NC score | 0.011271 (rank : 64) | θ value | 1.06291 (rank : 54) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8K3G5, Q921W6 | Gene names | Vrk3 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase VRK3 (EC 2.7.11.1) (Vaccinia-related kinase 3). | |||||
|
ZN638_HUMAN
|
||||||
| NC score | 0.010352 (rank : 65) | θ value | 3.0926 (rank : 59) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14966, Q53R34, Q5XJ05, Q68DP3, Q6P2H2, Q7Z3T7, Q8NF92, Q8TCA1, Q9H2G1, Q9NP37 | Gene names | ZNF638, NP220, ZFML | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 638 (Nuclear protein 220) (Zinc-finger matrin-like protein) (Cutaneous T-cell lymphoma-associated antigen se33-1) (CTCL tumor antigen se33-1). | |||||
|
AKAP1_HUMAN
|
||||||
| NC score | 0.009798 (rank : 66) | θ value | 8.99809 (rank : 66) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q92667, Q13320, Q9BW14 | Gene names | AKAP1, AKAP149 | |||
|
Domain Architecture |

|
|||||
| Description | A kinase anchor protein 1, mitochondrial precursor (Protein kinase A- anchoring protein 1) (PRKA1) (A-kinase anchor protein 149 kDa) (AKAP 149) (Dual specificity A-kinase-anchoring protein 1) (D-AKAP-1) (Spermatid A-kinase anchor protein 84) (S-AKAP84). | |||||
|
AMGO1_HUMAN
|
||||||
| NC score | 0.006615 (rank : 67) | θ value | 1.81305 (rank : 55) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 353 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q86WK6, Q8IW71, Q9ULQ7 | Gene names | AMIGO1, ALI2, AMIGO, KIAA1163 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amphoterin-induced protein 1 precursor (AMIGO-1) (Alivin-2). | |||||
|
HES2_HUMAN
|
||||||
| NC score | 0.006318 (rank : 68) | θ value | 8.99809 (rank : 68) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y543, Q96EN4, Q9Y542 | Gene names | HES2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-2 (Hairy and enhancer of split 2). | |||||
|
AMGO1_MOUSE
|
||||||
| NC score | 0.006167 (rank : 69) | θ value | 3.0926 (rank : 58) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80ZD8, Q69ZQ0, Q8R5D4, Q921U9 | Gene names | Amigo1, Ali2, Amigo, Kiaa1163 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amphoterin-induced protein 1 precursor (AMIGO-1) (Alivin-2). | |||||
|
TTLL4_HUMAN
|
||||||
| NC score | 0.005773 (rank : 70) | θ value | 6.88961 (rank : 64) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q14679, Q8WW29 | Gene names | TTLL4, KIAA0173 | |||
|
Domain Architecture |

|
|||||
| Description | Tubulin--tyrosine ligase-like protein 4. | |||||
|
WN10A_HUMAN
|
||||||
| NC score | 0.001946 (rank : 71) | θ value | 6.88961 (rank : 65) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9GZT5, Q96TA7, Q9H7S8 | Gene names | WNT10A | |||
|
Domain Architecture |

|
|||||
| Description | Protein Wnt-10a precursor. | |||||
|
HIC1_MOUSE
|
||||||
| NC score | -0.001829 (rank : 72) | θ value | 4.03905 (rank : 60) | |||
| Query Neighborhood Hits | 68 | Target Neighborhood Hits | 863 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9R1Y5, Q9R1Y6, Q9R2B0 | Gene names | Hic1 | |||
|
Domain Architecture |

|
|||||
| Description | Hypermethylated in cancer 1 protein (Hic-1). | |||||