Please be patient as the page loads
|
AXN1_MOUSE
|
||||||
| SwissProt Accessions | O35625 | Gene names | Axin1, Axin, Fu | |||
|
Domain Architecture |

|
|||||
| Description | Axin-1 (Axis inhibition protein 1) (Fused protein). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
AXN1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.992127 (rank : 2) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | O15169, Q4TT26, Q4TT27, Q86YA7, Q8WVW6, Q96S28 | Gene names | AXIN1, AXIN | |||
|
Domain Architecture |

|
|||||
| Description | Axin-1 (Axis inhibition protein 1) (hAxin). | |||||
|
AXN1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 89 | |
| SwissProt Accessions | O35625 | Gene names | Axin1, Axin, Fu | |||
|
Domain Architecture |

|
|||||
| Description | Axin-1 (Axis inhibition protein 1) (Fused protein). | |||||
|
AXN2_MOUSE
|
||||||
| θ value | 3.7435e-147 (rank : 3) | NC score | 0.948949 (rank : 4) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O88566, Q9QXJ6 | Gene names | Axin2 | |||
|
Domain Architecture |

|
|||||
| Description | Axin-2 (Axis inhibition protein 2) (Conductin) (Axin-like protein) (Axil). | |||||
|
AXN2_HUMAN
|
||||||
| θ value | 6.38546e-147 (rank : 4) | NC score | 0.954450 (rank : 3) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9Y2T1, Q9H3M6, Q9UH84 | Gene names | AXIN2 | |||
|
Domain Architecture |

|
|||||
| Description | Axin-2 (Axis inhibition protein 2) (Conductin) (Axin-like protein) (Axil). | |||||
|
RGS10_HUMAN
|
||||||
| θ value | 3.40345e-15 (rank : 5) | NC score | 0.726629 (rank : 5) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O43665 | Gene names | RGS10 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 10 (RGS10). | |||||
|
RGS10_MOUSE
|
||||||
| θ value | 2.88119e-14 (rank : 6) | NC score | 0.719653 (rank : 6) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9CQE5, Q9D3L2 | Gene names | Rgs10 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 10 (RGS10). | |||||
|
RGS3_HUMAN
|
||||||
| θ value | 8.38298e-14 (rank : 7) | NC score | 0.506526 (rank : 37) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
RGS5_HUMAN
|
||||||
| θ value | 1.86753e-13 (rank : 8) | NC score | 0.703449 (rank : 9) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O15539 | Gene names | RGS5 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 5 (RGS5). | |||||
|
RGS18_MOUSE
|
||||||
| θ value | 4.16044e-13 (rank : 9) | NC score | 0.706835 (rank : 8) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q99PG4 | Gene names | Rgs18 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 18 (RGS18). | |||||
|
RGS1_HUMAN
|
||||||
| θ value | 4.16044e-13 (rank : 10) | NC score | 0.703380 (rank : 10) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q08116, Q07918, Q9H1W2 | Gene names | RGS1, 1R20, BL34 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 1 (RGS1) (Early response protein 1R20) (B-cell activation protein BL34). | |||||
|
RGS3_MOUSE
|
||||||
| θ value | 4.16044e-13 (rank : 11) | NC score | 0.447518 (rank : 40) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
RGS8_MOUSE
|
||||||
| θ value | 5.43371e-13 (rank : 12) | NC score | 0.699550 (rank : 11) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8BXT1 | Gene names | Rgs8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of G-protein signaling 8 (RGS8). | |||||
|
RGS18_HUMAN
|
||||||
| θ value | 7.09661e-13 (rank : 13) | NC score | 0.706844 (rank : 7) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9NS28 | Gene names | RGS18, RGS13 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 18 (RGS18). | |||||
|
RGS2_HUMAN
|
||||||
| θ value | 9.26847e-13 (rank : 14) | NC score | 0.698753 (rank : 12) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P41220 | Gene names | RGS2, G0S8 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 2 (RGS2) (G0/G1 switch regulatory protein 8). | |||||
|
RGS8_HUMAN
|
||||||
| θ value | 9.26847e-13 (rank : 15) | NC score | 0.698234 (rank : 13) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P57771, Q3SYD2 | Gene names | RGS8 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 8 (RGS8). | |||||
|
RGS2_MOUSE
|
||||||
| θ value | 1.2105e-12 (rank : 16) | NC score | 0.695416 (rank : 16) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O08849, Q544S7, Q91WX1, Q9JL24 | Gene names | Rgs2 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 2 (RGS2). | |||||
|
RGS5_MOUSE
|
||||||
| θ value | 1.2105e-12 (rank : 17) | NC score | 0.697454 (rank : 15) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O08850, Q543B1, Q9D0Z2 | Gene names | Rgs5 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 5 (RGS5). | |||||
|
RGS1_MOUSE
|
||||||
| θ value | 3.52202e-12 (rank : 18) | NC score | 0.697665 (rank : 14) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9JL25, Q3TBD1, Q3TC18, Q3TD56 | Gene names | Rgs1 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 1 (RGS1). | |||||
|
RGS4_MOUSE
|
||||||
| θ value | 4.59992e-12 (rank : 19) | NC score | 0.694192 (rank : 17) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O08899, Q99L30 | Gene names | Rgs4 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 4 (RGS4). | |||||
|
RGS4_HUMAN
|
||||||
| θ value | 6.00763e-12 (rank : 20) | NC score | 0.693996 (rank : 18) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P49798 | Gene names | RGS4 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 4 (RGS4) (RGP4). | |||||
|
RGS16_MOUSE
|
||||||
| θ value | 7.84624e-12 (rank : 21) | NC score | 0.687323 (rank : 19) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P97428, O09091, P97420, Q80V16 | Gene names | Rgs16, Rgsr | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 16 (RGS16) (Retinally abundant regulator of G-protein signaling) (RGS-R) (A28-RGS14P). | |||||
|
RGS17_MOUSE
|
||||||
| θ value | 2.98157e-11 (rank : 22) | NC score | 0.673806 (rank : 23) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9QZB0, Q543T9 | Gene names | Rgs17, Rgsz2 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 17 (RGS17) (Regulator of Gz-selective protein signaling 2). | |||||
|
RGS14_MOUSE
|
||||||
| θ value | 3.89403e-11 (rank : 23) | NC score | 0.646791 (rank : 29) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P97492, Q9DCD1 | Gene names | Rgs14 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 14 (RGS14) (RAP1/RAP2-interacting protein). | |||||
|
RGS14_HUMAN
|
||||||
| θ value | 1.133e-10 (rank : 24) | NC score | 0.640013 (rank : 30) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O43566, O43565, Q8TD62 | Gene names | RGS14 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 14 (RGS14). | |||||
|
RGS20_MOUSE
|
||||||
| θ value | 1.133e-10 (rank : 25) | NC score | 0.669473 (rank : 24) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9QZB1, Q3TY63, Q9CUV8, Q9QZB2 | Gene names | Rgs20, Rgsz1 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 20 (RGS20) (Regulator of G-protein signaling Z1). | |||||
|
RGS16_HUMAN
|
||||||
| θ value | 3.29651e-10 (rank : 26) | NC score | 0.680943 (rank : 22) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O15492, Q99701 | Gene names | RGS16, RGSR | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 16 (RGS16) (Retinally abundant regulator of G-protein signaling) (RGS-R) (A28-RGS14P). | |||||
|
RGS20_HUMAN
|
||||||
| θ value | 5.62301e-10 (rank : 27) | NC score | 0.658163 (rank : 28) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O76081, Q96BG9 | Gene names | RGS20, RGSZ1, ZGAP1 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 20 (RGS20) (Regulator of Gz-selective protein signaling 1) (Gz-selective GTPase-activating protein) (G(z)GAP). | |||||
|
RGS13_MOUSE
|
||||||
| θ value | 9.59137e-10 (rank : 28) | NC score | 0.686555 (rank : 20) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8K443 | Gene names | Rgs13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of G-protein signaling 13 (RGS13). | |||||
|
RGS19_HUMAN
|
||||||
| θ value | 9.59137e-10 (rank : 29) | NC score | 0.665039 (rank : 26) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P49795, Q53XN0, Q8TD60 | Gene names | RGS19, GAIP, GNAI3IP | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 19 (RGS19) (G-alpha-interacting protein) (GAIP protein). | |||||
|
RGS17_HUMAN
|
||||||
| θ value | 1.25267e-09 (rank : 30) | NC score | 0.663469 (rank : 27) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9UGC6, Q5TF49, Q8TD61, Q9UJS8 | Gene names | RGS17 | |||
|
Domain Architecture |
|
|||||
| Description | Regulator of G-protein signaling 17 (RGS17). | |||||
|
RGS13_HUMAN
|
||||||
| θ value | 2.13673e-09 (rank : 31) | NC score | 0.685035 (rank : 21) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O14921, Q6PGR2, Q8TD63, Q9BX45 | Gene names | RGS13 | |||
|
Domain Architecture |
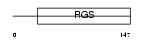
|
|||||
| Description | Regulator of G-protein signaling 13 (RGS13). | |||||
|
RGS19_MOUSE
|
||||||
| θ value | 6.21693e-09 (rank : 32) | NC score | 0.665500 (rank : 25) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9CX84, Q99L50 | Gene names | Rgs19 | |||
|
Domain Architecture |
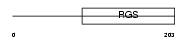
|
|||||
| Description | Regulator of G-protein signaling 19 (RGS19). | |||||
|
RGS12_HUMAN
|
||||||
| θ value | 1.99992e-07 (rank : 33) | NC score | 0.543099 (rank : 34) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | O14924, O14922, O14923, O43510, O75338 | Gene names | RGS12 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 12 (RGS12). | |||||
|
RGS11_HUMAN
|
||||||
| θ value | 2.88788e-06 (rank : 34) | NC score | 0.569785 (rank : 31) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | O94810, O75883 | Gene names | RGS11 | |||
|
Domain Architecture |
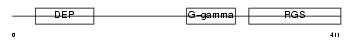
|
|||||
| Description | Regulator of G-protein signaling 11 (RGS11). | |||||
|
RGS9_MOUSE
|
||||||
| θ value | 1.43324e-05 (rank : 35) | NC score | 0.546502 (rank : 32) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O54828, Q9Z0S0 | Gene names | Rgs9 | |||
|
Domain Architecture |
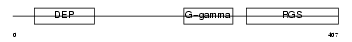
|
|||||
| Description | Regulator of G-protein signaling 9 (RGS9). | |||||
|
RGS9_HUMAN
|
||||||
| θ value | 2.44474e-05 (rank : 36) | NC score | 0.545452 (rank : 33) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O75916, O75573, Q696R2, Q8TD64, Q8TD65, Q9HC32, Q9HC33 | Gene names | RGS9 | |||
|
Domain Architecture |
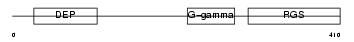
|
|||||
| Description | Regulator of G-protein signaling 9 (RGS9). | |||||
|
DVL2_MOUSE
|
||||||
| θ value | 3.19293e-05 (rank : 37) | NC score | 0.214745 (rank : 48) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q60838 | Gene names | Dvl2 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-2 (Dishevelled-2) (DSH homolog 2). | |||||
|
DVL2_HUMAN
|
||||||
| θ value | 9.29e-05 (rank : 38) | NC score | 0.214809 (rank : 47) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O14641 | Gene names | DVL2 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-2 (Dishevelled-2) (DSH homolog 2). | |||||
|
DVL3_HUMAN
|
||||||
| θ value | 0.000121331 (rank : 39) | NC score | 0.222113 (rank : 46) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q92997, O14642, Q13531, Q8N5E9, Q92607 | Gene names | DVL3, KIAA0208 | |||
|
Domain Architecture |
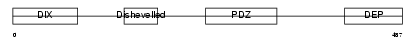
|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-3 (Dishevelled-3) (DSH homolog 3). | |||||
|
DVL3_MOUSE
|
||||||
| θ value | 0.000121331 (rank : 40) | NC score | 0.222224 (rank : 45) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q61062 | Gene names | Dvl3 | |||
|
Domain Architecture |
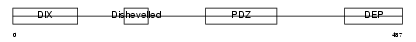
|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-3 (Dishevelled-3) (DSH homolog 3). | |||||
|
SNX14_HUMAN
|
||||||
| θ value | 0.000158464 (rank : 41) | NC score | 0.167967 (rank : 53) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Y5W7, Q9BSD1 | Gene names | SNX14 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-14. | |||||
|
SNX14_MOUSE
|
||||||
| θ value | 0.000158464 (rank : 42) | NC score | 0.168925 (rank : 52) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8BHY8 | Gene names | Snx14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sorting nexin-14. | |||||
|
RGS7_HUMAN
|
||||||
| θ value | 0.000786445 (rank : 43) | NC score | 0.516986 (rank : 35) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P49802, Q8TD66, Q8TD67, Q9UNU7, Q9Y6B9 | Gene names | RGS7 | |||
|
Domain Architecture |
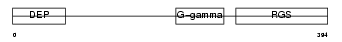
|
|||||
| Description | Regulator of G-protein signaling 7 (RGS7). | |||||
|
RGS7_MOUSE
|
||||||
| θ value | 0.00298849 (rank : 44) | NC score | 0.509671 (rank : 36) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O54829 | Gene names | Rgs7 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 7 (RGS7). | |||||
|
ARBK1_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 45) | NC score | 0.026366 (rank : 62) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 859 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P25098, Q13837 | Gene names | ADRBK1, BARK, BARK1, GRK2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-adrenergic receptor kinase 1 (EC 2.7.11.15) (Beta-ARK-1) (G- protein coupled receptor kinase 2). | |||||
|
ARBK1_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 46) | NC score | 0.023258 (rank : 64) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 849 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q99MK8, Q99LL8 | Gene names | Adrbk1, Grk2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-adrenergic receptor kinase 1 (EC 2.7.11.15) (Beta-ARK-1) (G- protein-coupled receptor kinase 2). | |||||
|
DVL1L_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 47) | NC score | 0.197148 (rank : 49) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P54792 | Gene names | DVL1L1, DVL, DVL1 | |||
|
Domain Architecture |
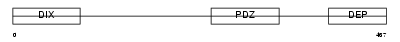
|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1-like (Dishevelled- 1-like) (DSH homolog 1-like). | |||||
|
DVL1_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 48) | NC score | 0.194771 (rank : 51) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O14640 | Gene names | DVL1 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1 (Dishevelled-1) (DSH homolog 1). | |||||
|
DVL1_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 49) | NC score | 0.194929 (rank : 50) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P51141, Q60868 | Gene names | Dvl1, Dvl | |||
|
Domain Architecture |
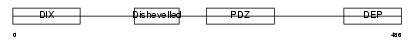
|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1 (Dishevelled-1) (DSH homolog 1). | |||||
|
BAZ1B_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 50) | NC score | 0.040015 (rank : 55) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 701 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UIG0, O95039, O95247, O95277 | Gene names | BAZ1B, WBSC10, WBSCR9, WSTF | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein) (Williams syndrome transcription factor) (hWALP2). | |||||
|
SNX13_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 51) | NC score | 0.241655 (rank : 41) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9Y5W8, O94821, Q8WVZ2, Q8WXH8 | Gene names | SNX13, KIAA0713 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-13 (RGS domain- and PHOX domain-containing protein) (RGS-PX1). | |||||
|
SNX13_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 52) | NC score | 0.241552 (rank : 42) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q6PHS6 | Gene names | Snx13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sorting nexin-13. | |||||
|
RGS6_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 53) | NC score | 0.496551 (rank : 38) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9Z2H2, Q8BGI8 | Gene names | Rgs6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of G-protein signaling 6 (RGS6). | |||||
|
ARBK2_HUMAN
|
||||||
| θ value | 0.163984 (rank : 54) | NC score | 0.022451 (rank : 65) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 849 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P35626, Q9UGW9 | Gene names | ADRBK2, BARK2, GRK3 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-adrenergic receptor kinase 2 (EC 2.7.11.15) (Beta-ARK-2) (G- protein-coupled receptor kinase 3). | |||||
|
DDX49_HUMAN
|
||||||
| θ value | 0.47712 (rank : 55) | NC score | 0.009922 (rank : 76) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y6V7, Q53FJ1 | Gene names | DDX49 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX49 (EC 3.6.1.-) (DEAD box protein 49). | |||||
|
SNX25_HUMAN
|
||||||
| θ value | 0.47712 (rank : 56) | NC score | 0.134747 (rank : 54) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9H3E2 | Gene names | SNX25 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-25. | |||||
|
TCF20_MOUSE
|
||||||
| θ value | 0.47712 (rank : 57) | NC score | 0.027965 (rank : 60) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 396 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9EPQ8, Q60792 | Gene names | Tcf20, Spbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor 20 (Stromelysin 1 PDGF-responsive element-binding protein) (SPRE-binding protein) (Nuclear factor SPBP). | |||||
|
AKA10_HUMAN
|
||||||
| θ value | 0.62314 (rank : 58) | NC score | 0.227760 (rank : 43) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O43572, Q96AJ7 | Gene names | AKAP10 | |||
|
Domain Architecture |

|
|||||
| Description | A kinase anchor protein 10, mitochondrial precursor (Protein kinase A- anchoring protein 10) (PRKA10) (Dual specificity A kinase-anchoring protein 2) (D-AKAP-2). | |||||
|
AKA10_MOUSE
|
||||||
| θ value | 0.62314 (rank : 59) | NC score | 0.227169 (rank : 44) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O88845, Q5SUB5, Q5SUB7, Q7TPE7, Q8BQL6 | Gene names | Akap10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A kinase anchor protein 10, mitochondrial precursor (Protein kinase A- anchoring protein 10) (PRKA10) (Dual specificity A kinase-anchoring protein 2) (D-AKAP-2). | |||||
|
APC_HUMAN
|
||||||
| θ value | 0.62314 (rank : 60) | NC score | 0.035282 (rank : 57) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 603 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P25054, Q15162, Q15163, Q93042 | Gene names | APC, DP2.5 | |||
|
Domain Architecture |

|
|||||
| Description | Adenomatous polyposis coli protein (Protein APC). | |||||
|
CTDP1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 61) | NC score | 0.035612 (rank : 56) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Y5B0, Q7Z644, Q96BZ1, Q9Y6F5 | Gene names | CTDP1, FCP1 | |||
|
Domain Architecture |

|
|||||
| Description | RNA polymerase II subunit A C-terminal domain phosphatase (EC 3.1.3.16) (TFIIF-associating CTD phosphatase). | |||||
|
RGS6_HUMAN
|
||||||
| θ value | 1.06291 (rank : 62) | NC score | 0.465921 (rank : 39) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P49758, O75576, O75577, Q8TE13, Q8TE14, Q8TE15, Q8TE16, Q8TE17, Q8TE18, Q8TE19, Q8TE20, Q8TE21, Q8TE22, Q9Y245, Q9Y647 | Gene names | RGS6 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 6 (RGS6) (S914). | |||||
|
ZN184_HUMAN
|
||||||
| θ value | 1.06291 (rank : 63) | NC score | -0.002314 (rank : 87) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 811 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q99676, O60792, Q8TBA9 | Gene names | ZNF184 | |||
|
Domain Architecture |
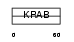
|
|||||
| Description | Zinc finger protein 184. | |||||
|
MAST4_HUMAN
|
||||||
| θ value | 1.38821 (rank : 64) | NC score | 0.005324 (rank : 79) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1804 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O15021, Q6ZN07, Q96LY3 | Gene names | MAST4, KIAA0303 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase 4 (EC 2.7.11.1). | |||||
|
RUFY3_HUMAN
|
||||||
| θ value | 1.38821 (rank : 65) | NC score | 0.007381 (rank : 78) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 594 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7L099, O94948, Q9UI00 | Gene names | RUFY3, KIAA0871 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein RUFY3 (Rap2-interacting protein x) (RIPx). | |||||
|
TEX14_MOUSE
|
||||||
| θ value | 1.38821 (rank : 66) | NC score | 0.003372 (rank : 81) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1333 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q7M6U3, Q3UT36, Q5NC10, Q5NC11, Q8CGK1, Q8CGK2, Q99MV8 | Gene names | Tex14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-expressed protein 14 (Testis-expressed sequence 14). | |||||
|
AP3B2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 67) | NC score | 0.012350 (rank : 75) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9JME5, Q6QR53, Q8R1E5 | Gene names | Ap3b2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP-3 complex subunit beta-2 (Adapter-related protein complex 3 beta-2 subunit) (Beta3B-adaptin) (Adaptor protein complex AP-3 beta-2 subunit) (Clathrin assembly protein complex 3 beta-2 large chain). | |||||
|
RNPS1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 68) | NC score | 0.019617 (rank : 67) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q15287, O75308, Q8WY42, Q9NYG3 | Gene names | RNPS1, LDC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein with serine-rich domain 1 (SR-related protein LDC2). | |||||
|
RNPS1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 69) | NC score | 0.019617 (rank : 68) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99M28, Q3TMJ1, Q922H8 | Gene names | Rnps1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein with serine-rich domain 1. | |||||
|
APC_MOUSE
|
||||||
| θ value | 2.36792 (rank : 70) | NC score | 0.029245 (rank : 59) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q61315, Q62044 | Gene names | Apc | |||
|
Domain Architecture |

|
|||||
| Description | Adenomatous polyposis coli protein (Protein APC) (mAPC). | |||||
|
FILA_HUMAN
|
||||||
| θ value | 2.36792 (rank : 71) | NC score | 0.021656 (rank : 66) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 685 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P20930, Q01720, Q5T583 | Gene names | FLG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filaggrin. | |||||
|
TCAL3_MOUSE
|
||||||
| θ value | 2.36792 (rank : 72) | NC score | 0.014363 (rank : 73) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8R0A5, Q9CYK5, Q9D1S4 | Gene names | Tceal3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 3 (TCEA-like protein 3) (Transcription elongation factor S-II protein-like 3). | |||||
|
SPAG1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 73) | NC score | 0.013607 (rank : 74) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80ZX8, Q8CCK7, Q9QZJ3, Q9QZJ4 | Gene names | Spag1, Tpis | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-associated antigen 1 (Infertility-related sperm protein Spag-1) (TPR-containing protein involved in spermatogenesis) (TPIS). | |||||
|
CD2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 74) | NC score | 0.031034 (rank : 58) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P06729, Q96TE5 | Gene names | CD2 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell surface antigen CD2 precursor (T-cell surface antigen T11/Leu- 5) (LFA-2) (LFA-3 receptor) (Erythrocyte receptor) (Rosette receptor). | |||||
|
OSBL5_HUMAN
|
||||||
| θ value | 4.03905 (rank : 75) | NC score | 0.008446 (rank : 77) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9H0X9, Q9BZB0, Q9P1Z4 | Gene names | OSBPL5, KIAA1534, ORP5 | |||
|
Domain Architecture |

|
|||||
| Description | Oxysterol-binding protein-related protein 5 (OSBP-related protein 5) (ORP-5). | |||||
|
PAPD1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 76) | NC score | 0.014684 (rank : 72) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9D0D3, Q3UXJ1, Q8C651 | Gene names | Papd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly(A) RNA polymerase, mitochondrial precursor (EC 2.7.7.19) (PAP) (Polynucleotide adenylyltransferase) (PAP-associated domain-containing protein 1). | |||||
|
PININ_MOUSE
|
||||||
| θ value | 4.03905 (rank : 77) | NC score | 0.016981 (rank : 69) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 777 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O35691, Q8CD89, Q8CGU3 | Gene names | Pnn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pinin. | |||||
|
WHRN_MOUSE
|
||||||
| θ value | 4.03905 (rank : 78) | NC score | 0.027647 (rank : 61) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q80VW5, Q80TC2, Q80VW4 | Gene names | Whrn, Kiaa1526 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Whirlin. | |||||
|
PK3C3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 79) | NC score | 0.015978 (rank : 71) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NEB9, Q15134 | Gene names | PIK3C3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol 3-kinase catalytic subunit type 3 (EC 2.7.1.137) (PtdIns-3-kinase type 3) (PI3-kinase type 3) (PI3K type 3) (Phosphoinositide-3-kinase class 3) (Phosphatidylinositol 3-kinase p100 subunit). | |||||
|
PK3C3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 80) | NC score | 0.015987 (rank : 70) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6PF93, Q8R3S8 | Gene names | Pik3c3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol 3-kinase catalytic subunit type 3 (EC 2.7.1.137) (PtdIns-3-kinase type 3) (PI3-kinase type 3) (PI3K type 3) (Phosphoinositide-3-kinase class 3). | |||||
|
RABE1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 81) | NC score | 0.000295 (rank : 84) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1105 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O35551, Q99L08, Q9CRP3, Q9EQF9, Q9JI94 | Gene names | Rabep1, Rab5ep, Rabpt5, Rabpt5a | |||
|
Domain Architecture |

|
|||||
| Description | Rab GTPase-binding effector protein 1 (Rabaptin-5) (Rabaptin-5alpha). | |||||
|
TROAP_HUMAN
|
||||||
| θ value | 5.27518 (rank : 82) | NC score | 0.024459 (rank : 63) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q12815 | Gene names | TROAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trophinin-associated protein (Tastin) (Trophinin-assisting protein). | |||||
|
ANK2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 83) | NC score | 0.000914 (rank : 83) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
LTBP3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 84) | NC score | -0.000115 (rank : 85) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 502 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NS15, O15107, Q96HB9, Q9H7K2, Q9UFN4 | Gene names | LTBP3 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein 3 precursor (LTBP-3). | |||||
|
ZN264_HUMAN
|
||||||
| θ value | 6.88961 (rank : 85) | NC score | -0.003045 (rank : 89) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 770 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O43296, Q9P1V0 | Gene names | ZNF264, KIAA0412 | |||
|
Domain Architecture |
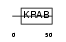
|
|||||
| Description | Zinc finger protein 264. | |||||
|
ZN579_MOUSE
|
||||||
| θ value | 6.88961 (rank : 86) | NC score | -0.002176 (rank : 86) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 719 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80VM4 | Gene names | Znf579, Zfp579 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 579. | |||||
|
DUOX1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 87) | NC score | 0.003390 (rank : 80) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NRD9, Q6ZMB3, Q6ZR09, Q9NZC1 | Gene names | DUOX1, DUOX, LNOX1, THOX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dual oxidase 1 precursor (EC 1.6.3.1) (EC 1.11.1.-) (NADPH thyroid oxidase 1) (Thyroid oxidase 1) (Large NOX 1) (Long NOX 1). | |||||
|
ULK2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 88) | NC score | 0.001606 (rank : 82) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 943 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9QY01, Q80TV7, Q9WTP4 | Gene names | Ulk2, Kiaa0623 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase ULK2 (EC 2.7.11.1) (Unc-51-like kinase 2) (Serine/threonine-protein kinase Unc51.2). | |||||
|
ZN426_HUMAN
|
||||||
| θ value | 8.99809 (rank : 89) | NC score | -0.002757 (rank : 88) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 747 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9BUY5 | Gene names | ZNF426 | |||
|
Domain Architecture |
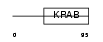
|
|||||
| Description | Zinc finger protein 426. | |||||
|
AXN1_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 89 | |
| SwissProt Accessions | O35625 | Gene names | Axin1, Axin, Fu | |||
|
Domain Architecture |

|
|||||
| Description | Axin-1 (Axis inhibition protein 1) (Fused protein). | |||||
|
AXN1_HUMAN
|
||||||
| NC score | 0.992127 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | O15169, Q4TT26, Q4TT27, Q86YA7, Q8WVW6, Q96S28 | Gene names | AXIN1, AXIN | |||
|
Domain Architecture |

|
|||||
| Description | Axin-1 (Axis inhibition protein 1) (hAxin). | |||||
|
AXN2_HUMAN
|
||||||
| NC score | 0.954450 (rank : 3) | θ value | 6.38546e-147 (rank : 4) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9Y2T1, Q9H3M6, Q9UH84 | Gene names | AXIN2 | |||
|
Domain Architecture |

|
|||||
| Description | Axin-2 (Axis inhibition protein 2) (Conductin) (Axin-like protein) (Axil). | |||||
|
AXN2_MOUSE
|
||||||
| NC score | 0.948949 (rank : 4) | θ value | 3.7435e-147 (rank : 3) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O88566, Q9QXJ6 | Gene names | Axin2 | |||
|
Domain Architecture |

|
|||||
| Description | Axin-2 (Axis inhibition protein 2) (Conductin) (Axin-like protein) (Axil). | |||||
|
RGS10_HUMAN
|
||||||
| NC score | 0.726629 (rank : 5) | θ value | 3.40345e-15 (rank : 5) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O43665 | Gene names | RGS10 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 10 (RGS10). | |||||
|
RGS10_MOUSE
|
||||||
| NC score | 0.719653 (rank : 6) | θ value | 2.88119e-14 (rank : 6) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9CQE5, Q9D3L2 | Gene names | Rgs10 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 10 (RGS10). | |||||
|
RGS18_HUMAN
|
||||||
| NC score | 0.706844 (rank : 7) | θ value | 7.09661e-13 (rank : 13) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9NS28 | Gene names | RGS18, RGS13 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 18 (RGS18). | |||||
|
RGS18_MOUSE
|
||||||
| NC score | 0.706835 (rank : 8) | θ value | 4.16044e-13 (rank : 9) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q99PG4 | Gene names | Rgs18 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 18 (RGS18). | |||||
|
RGS5_HUMAN
|
||||||
| NC score | 0.703449 (rank : 9) | θ value | 1.86753e-13 (rank : 8) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O15539 | Gene names | RGS5 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 5 (RGS5). | |||||
|
RGS1_HUMAN
|
||||||
| NC score | 0.703380 (rank : 10) | θ value | 4.16044e-13 (rank : 10) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q08116, Q07918, Q9H1W2 | Gene names | RGS1, 1R20, BL34 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 1 (RGS1) (Early response protein 1R20) (B-cell activation protein BL34). | |||||
|
RGS8_MOUSE
|
||||||
| NC score | 0.699550 (rank : 11) | θ value | 5.43371e-13 (rank : 12) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8BXT1 | Gene names | Rgs8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of G-protein signaling 8 (RGS8). | |||||
|
RGS2_HUMAN
|
||||||
| NC score | 0.698753 (rank : 12) | θ value | 9.26847e-13 (rank : 14) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P41220 | Gene names | RGS2, G0S8 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 2 (RGS2) (G0/G1 switch regulatory protein 8). | |||||
|
RGS8_HUMAN
|
||||||
| NC score | 0.698234 (rank : 13) | θ value | 9.26847e-13 (rank : 15) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P57771, Q3SYD2 | Gene names | RGS8 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 8 (RGS8). | |||||
|
RGS1_MOUSE
|
||||||
| NC score | 0.697665 (rank : 14) | θ value | 3.52202e-12 (rank : 18) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9JL25, Q3TBD1, Q3TC18, Q3TD56 | Gene names | Rgs1 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 1 (RGS1). | |||||
|
RGS5_MOUSE
|
||||||
| NC score | 0.697454 (rank : 15) | θ value | 1.2105e-12 (rank : 17) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O08850, Q543B1, Q9D0Z2 | Gene names | Rgs5 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 5 (RGS5). | |||||
|
RGS2_MOUSE
|
||||||
| NC score | 0.695416 (rank : 16) | θ value | 1.2105e-12 (rank : 16) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O08849, Q544S7, Q91WX1, Q9JL24 | Gene names | Rgs2 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 2 (RGS2). | |||||
|
RGS4_MOUSE
|
||||||
| NC score | 0.694192 (rank : 17) | θ value | 4.59992e-12 (rank : 19) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O08899, Q99L30 | Gene names | Rgs4 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 4 (RGS4). | |||||
|
RGS4_HUMAN
|
||||||
| NC score | 0.693996 (rank : 18) | θ value | 6.00763e-12 (rank : 20) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P49798 | Gene names | RGS4 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 4 (RGS4) (RGP4). | |||||
|
RGS16_MOUSE
|
||||||
| NC score | 0.687323 (rank : 19) | θ value | 7.84624e-12 (rank : 21) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P97428, O09091, P97420, Q80V16 | Gene names | Rgs16, Rgsr | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 16 (RGS16) (Retinally abundant regulator of G-protein signaling) (RGS-R) (A28-RGS14P). | |||||
|
RGS13_MOUSE
|
||||||
| NC score | 0.686555 (rank : 20) | θ value | 9.59137e-10 (rank : 28) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8K443 | Gene names | Rgs13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of G-protein signaling 13 (RGS13). | |||||
|
RGS13_HUMAN
|
||||||
| NC score | 0.685035 (rank : 21) | θ value | 2.13673e-09 (rank : 31) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O14921, Q6PGR2, Q8TD63, Q9BX45 | Gene names | RGS13 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 13 (RGS13). | |||||
|
RGS16_HUMAN
|
||||||
| NC score | 0.680943 (rank : 22) | θ value | 3.29651e-10 (rank : 26) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O15492, Q99701 | Gene names | RGS16, RGSR | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 16 (RGS16) (Retinally abundant regulator of G-protein signaling) (RGS-R) (A28-RGS14P). | |||||
|
RGS17_MOUSE
|
||||||
| NC score | 0.673806 (rank : 23) | θ value | 2.98157e-11 (rank : 22) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9QZB0, Q543T9 | Gene names | Rgs17, Rgsz2 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 17 (RGS17) (Regulator of Gz-selective protein signaling 2). | |||||
|
RGS20_MOUSE
|
||||||
| NC score | 0.669473 (rank : 24) | θ value | 1.133e-10 (rank : 25) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9QZB1, Q3TY63, Q9CUV8, Q9QZB2 | Gene names | Rgs20, Rgsz1 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 20 (RGS20) (Regulator of G-protein signaling Z1). | |||||
|
RGS19_MOUSE
|
||||||
| NC score | 0.665500 (rank : 25) | θ value | 6.21693e-09 (rank : 32) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9CX84, Q99L50 | Gene names | Rgs19 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 19 (RGS19). | |||||
|
RGS19_HUMAN
|
||||||
| NC score | 0.665039 (rank : 26) | θ value | 9.59137e-10 (rank : 29) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P49795, Q53XN0, Q8TD60 | Gene names | RGS19, GAIP, GNAI3IP | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 19 (RGS19) (G-alpha-interacting protein) (GAIP protein). | |||||
|
RGS17_HUMAN
|
||||||
| NC score | 0.663469 (rank : 27) | θ value | 1.25267e-09 (rank : 30) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9UGC6, Q5TF49, Q8TD61, Q9UJS8 | Gene names | RGS17 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 17 (RGS17). | |||||
|
RGS20_HUMAN
|
||||||
| NC score | 0.658163 (rank : 28) | θ value | 5.62301e-10 (rank : 27) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O76081, Q96BG9 | Gene names | RGS20, RGSZ1, ZGAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 20 (RGS20) (Regulator of Gz-selective protein signaling 1) (Gz-selective GTPase-activating protein) (G(z)GAP). | |||||
|
RGS14_MOUSE
|
||||||
| NC score | 0.646791 (rank : 29) | θ value | 3.89403e-11 (rank : 23) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P97492, Q9DCD1 | Gene names | Rgs14 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 14 (RGS14) (RAP1/RAP2-interacting protein). | |||||
|
RGS14_HUMAN
|
||||||
| NC score | 0.640013 (rank : 30) | θ value | 1.133e-10 (rank : 24) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O43566, O43565, Q8TD62 | Gene names | RGS14 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 14 (RGS14). | |||||
|
RGS11_HUMAN
|
||||||
| NC score | 0.569785 (rank : 31) | θ value | 2.88788e-06 (rank : 34) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | O94810, O75883 | Gene names | RGS11 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 11 (RGS11). | |||||
|
RGS9_MOUSE
|
||||||
| NC score | 0.546502 (rank : 32) | θ value | 1.43324e-05 (rank : 35) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O54828, Q9Z0S0 | Gene names | Rgs9 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 9 (RGS9). | |||||
|
RGS9_HUMAN
|
||||||
| NC score | 0.545452 (rank : 33) | θ value | 2.44474e-05 (rank : 36) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O75916, O75573, Q696R2, Q8TD64, Q8TD65, Q9HC32, Q9HC33 | Gene names | RGS9 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 9 (RGS9). | |||||
|
RGS12_HUMAN
|
||||||
| NC score | 0.543099 (rank : 34) | θ value | 1.99992e-07 (rank : 33) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | O14924, O14922, O14923, O43510, O75338 | Gene names | RGS12 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 12 (RGS12). | |||||
|
RGS7_HUMAN
|
||||||
| NC score | 0.516986 (rank : 35) | θ value | 0.000786445 (rank : 43) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P49802, Q8TD66, Q8TD67, Q9UNU7, Q9Y6B9 | Gene names | RGS7 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 7 (RGS7). | |||||
|
RGS7_MOUSE
|
||||||
| NC score | 0.509671 (rank : 36) | θ value | 0.00298849 (rank : 44) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O54829 | Gene names | Rgs7 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 7 (RGS7). | |||||
|
RGS3_HUMAN
|
||||||
| NC score | 0.506526 (rank : 37) | θ value | 8.38298e-14 (rank : 7) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
RGS6_MOUSE
|
||||||
| NC score | 0.496551 (rank : 38) | θ value | 0.0563607 (rank : 53) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9Z2H2, Q8BGI8 | Gene names | Rgs6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of G-protein signaling 6 (RGS6). | |||||
|
RGS6_HUMAN
|
||||||
| NC score | 0.465921 (rank : 39) | θ value | 1.06291 (rank : 62) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P49758, O75576, O75577, Q8TE13, Q8TE14, Q8TE15, Q8TE16, Q8TE17, Q8TE18, Q8TE19, Q8TE20, Q8TE21, Q8TE22, Q9Y245, Q9Y647 | Gene names | RGS6 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 6 (RGS6) (S914). | |||||
|
RGS3_MOUSE
|
||||||
| NC score | 0.447518 (rank : 40) | θ value | 4.16044e-13 (rank : 11) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
SNX13_HUMAN
|
||||||
| NC score | 0.241655 (rank : 41) | θ value | 0.0431538 (rank : 51) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9Y5W8, O94821, Q8WVZ2, Q8WXH8 | Gene names | SNX13, KIAA0713 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-13 (RGS domain- and PHOX domain-containing protein) (RGS-PX1). | |||||
|
SNX13_MOUSE
|
||||||
| NC score | 0.241552 (rank : 42) | θ value | 0.0431538 (rank : 52) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q6PHS6 | Gene names | Snx13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sorting nexin-13. | |||||
|
AKA10_HUMAN
|
||||||
| NC score | 0.227760 (rank : 43) | θ value | 0.62314 (rank : 58) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O43572, Q96AJ7 | Gene names | AKAP10 | |||
|
Domain Architecture |

|
|||||
| Description | A kinase anchor protein 10, mitochondrial precursor (Protein kinase A- anchoring protein 10) (PRKA10) (Dual specificity A kinase-anchoring protein 2) (D-AKAP-2). | |||||
|
AKA10_MOUSE
|
||||||
| NC score | 0.227169 (rank : 44) | θ value | 0.62314 (rank : 59) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O88845, Q5SUB5, Q5SUB7, Q7TPE7, Q8BQL6 | Gene names | Akap10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A kinase anchor protein 10, mitochondrial precursor (Protein kinase A- anchoring protein 10) (PRKA10) (Dual specificity A kinase-anchoring protein 2) (D-AKAP-2). | |||||
|
DVL3_MOUSE
|
||||||
| NC score | 0.222224 (rank : 45) | θ value | 0.000121331 (rank : 40) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q61062 | Gene names | Dvl3 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-3 (Dishevelled-3) (DSH homolog 3). | |||||
|
DVL3_HUMAN
|
||||||
| NC score | 0.222113 (rank : 46) | θ value | 0.000121331 (rank : 39) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q92997, O14642, Q13531, Q8N5E9, Q92607 | Gene names | DVL3, KIAA0208 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-3 (Dishevelled-3) (DSH homolog 3). | |||||
|
DVL2_HUMAN
|
||||||
| NC score | 0.214809 (rank : 47) | θ value | 9.29e-05 (rank : 38) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O14641 | Gene names | DVL2 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-2 (Dishevelled-2) (DSH homolog 2). | |||||
|
DVL2_MOUSE
|
||||||
| NC score | 0.214745 (rank : 48) | θ value | 3.19293e-05 (rank : 37) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q60838 | Gene names | Dvl2 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-2 (Dishevelled-2) (DSH homolog 2). | |||||
|
DVL1L_HUMAN
|
||||||
| NC score | 0.197148 (rank : 49) | θ value | 0.00390308 (rank : 47) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P54792 | Gene names | DVL1L1, DVL, DVL1 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1-like (Dishevelled- 1-like) (DSH homolog 1-like). | |||||
|
DVL1_MOUSE
|
||||||
| NC score | 0.194929 (rank : 50) | θ value | 0.00390308 (rank : 49) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P51141, Q60868 | Gene names | Dvl1, Dvl | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1 (Dishevelled-1) (DSH homolog 1). | |||||
|
DVL1_HUMAN
|
||||||
| NC score | 0.194771 (rank : 51) | θ value | 0.00390308 (rank : 48) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O14640 | Gene names | DVL1 | |||
|
Domain Architecture |

|
|||||
| Description | Segment polarity protein dishevelled homolog DVL-1 (Dishevelled-1) (DSH homolog 1). | |||||
|
SNX14_MOUSE
|
||||||
| NC score | 0.168925 (rank : 52) | θ value | 0.000158464 (rank : 42) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8BHY8 | Gene names | Snx14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sorting nexin-14. | |||||
|
SNX14_HUMAN
|
||||||
| NC score | 0.167967 (rank : 53) | θ value | 0.000158464 (rank : 41) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Y5W7, Q9BSD1 | Gene names | SNX14 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-14. | |||||
|
SNX25_HUMAN
|
||||||
| NC score | 0.134747 (rank : 54) | θ value | 0.47712 (rank : 56) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9H3E2 | Gene names | SNX25 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-25. | |||||
|
BAZ1B_HUMAN
|
||||||
| NC score | 0.040015 (rank : 55) | θ value | 0.0148317 (rank : 50) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 701 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UIG0, O95039, O95247, O95277 | Gene names | BAZ1B, WBSC10, WBSCR9, WSTF | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein) (Williams syndrome transcription factor) (hWALP2). | |||||
|
CTDP1_HUMAN
|
||||||
| NC score | 0.035612 (rank : 56) | θ value | 0.62314 (rank : 61) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Y5B0, Q7Z644, Q96BZ1, Q9Y6F5 | Gene names | CTDP1, FCP1 | |||
|
Domain Architecture |

|
|||||
| Description | RNA polymerase II subunit A C-terminal domain phosphatase (EC 3.1.3.16) (TFIIF-associating CTD phosphatase). | |||||
|
APC_HUMAN
|
||||||
| NC score | 0.035282 (rank : 57) | θ value | 0.62314 (rank : 60) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 603 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P25054, Q15162, Q15163, Q93042 | Gene names | APC, DP2.5 | |||
|
Domain Architecture |

|
|||||
| Description | Adenomatous polyposis coli protein (Protein APC). | |||||
|
CD2_HUMAN
|
||||||
| NC score | 0.031034 (rank : 58) | θ value | 4.03905 (rank : 74) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P06729, Q96TE5 | Gene names | CD2 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell surface antigen CD2 precursor (T-cell surface antigen T11/Leu- 5) (LFA-2) (LFA-3 receptor) (Erythrocyte receptor) (Rosette receptor). | |||||
|
APC_MOUSE
|
||||||
| NC score | 0.029245 (rank : 59) | θ value | 2.36792 (rank : 70) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q61315, Q62044 | Gene names | Apc | |||
|
Domain Architecture |

|
|||||
| Description | Adenomatous polyposis coli protein (Protein APC) (mAPC). | |||||
|
TCF20_MOUSE
|
||||||
| NC score | 0.027965 (rank : 60) | θ value | 0.47712 (rank : 57) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 396 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9EPQ8, Q60792 | Gene names | Tcf20, Spbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor 20 (Stromelysin 1 PDGF-responsive element-binding protein) (SPRE-binding protein) (Nuclear factor SPBP). | |||||
|
WHRN_MOUSE
|
||||||
| NC score | 0.027647 (rank : 61) | θ value | 4.03905 (rank : 78) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q80VW5, Q80TC2, Q80VW4 | Gene names | Whrn, Kiaa1526 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Whirlin. | |||||
|
ARBK1_HUMAN
|
||||||
| NC score | 0.026366 (rank : 62) | θ value | 0.00390308 (rank : 45) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 859 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P25098, Q13837 | Gene names | ADRBK1, BARK, BARK1, GRK2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-adrenergic receptor kinase 1 (EC 2.7.11.15) (Beta-ARK-1) (G- protein coupled receptor kinase 2). | |||||
|
TROAP_HUMAN
|
||||||
| NC score | 0.024459 (rank : 63) | θ value | 5.27518 (rank : 82) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q12815 | Gene names | TROAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trophinin-associated protein (Tastin) (Trophinin-assisting protein). | |||||
|
ARBK1_MOUSE
|
||||||
| NC score | 0.023258 (rank : 64) | θ value | 0.00390308 (rank : 46) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 849 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q99MK8, Q99LL8 | Gene names | Adrbk1, Grk2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-adrenergic receptor kinase 1 (EC 2.7.11.15) (Beta-ARK-1) (G- protein-coupled receptor kinase 2). | |||||
|
ARBK2_HUMAN
|
||||||
| NC score | 0.022451 (rank : 65) | θ value | 0.163984 (rank : 54) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 849 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P35626, Q9UGW9 | Gene names | ADRBK2, BARK2, GRK3 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-adrenergic receptor kinase 2 (EC 2.7.11.15) (Beta-ARK-2) (G- protein-coupled receptor kinase 3). | |||||
|
FILA_HUMAN
|
||||||
| NC score | 0.021656 (rank : 66) | θ value | 2.36792 (rank : 71) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 685 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P20930, Q01720, Q5T583 | Gene names | FLG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filaggrin. | |||||
|
RNPS1_HUMAN
|
||||||
| NC score | 0.019617 (rank : 67) | θ value | 1.81305 (rank : 68) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q15287, O75308, Q8WY42, Q9NYG3 | Gene names | RNPS1, LDC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein with serine-rich domain 1 (SR-related protein LDC2). | |||||
|
RNPS1_MOUSE
|
||||||
| NC score | 0.019617 (rank : 68) | θ value | 1.81305 (rank : 69) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99M28, Q3TMJ1, Q922H8 | Gene names | Rnps1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein with serine-rich domain 1. | |||||
|
PININ_MOUSE
|
||||||
| NC score | 0.016981 (rank : 69) | θ value | 4.03905 (rank : 77) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 777 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O35691, Q8CD89, Q8CGU3 | Gene names | Pnn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pinin. | |||||
|
PK3C3_MOUSE
|
||||||
| NC score | 0.015987 (rank : 70) | θ value | 5.27518 (rank : 80) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6PF93, Q8R3S8 | Gene names | Pik3c3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol 3-kinase catalytic subunit type 3 (EC 2.7.1.137) (PtdIns-3-kinase type 3) (PI3-kinase type 3) (PI3K type 3) (Phosphoinositide-3-kinase class 3). | |||||
|
PK3C3_HUMAN
|
||||||
| NC score | 0.015978 (rank : 71) | θ value | 5.27518 (rank : 79) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NEB9, Q15134 | Gene names | PIK3C3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol 3-kinase catalytic subunit type 3 (EC 2.7.1.137) (PtdIns-3-kinase type 3) (PI3-kinase type 3) (PI3K type 3) (Phosphoinositide-3-kinase class 3) (Phosphatidylinositol 3-kinase p100 subunit). | |||||
|
PAPD1_MOUSE
|
||||||
| NC score | 0.014684 (rank : 72) | θ value | 4.03905 (rank : 76) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9D0D3, Q3UXJ1, Q8C651 | Gene names | Papd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly(A) RNA polymerase, mitochondrial precursor (EC 2.7.7.19) (PAP) (Polynucleotide adenylyltransferase) (PAP-associated domain-containing protein 1). | |||||
|
TCAL3_MOUSE
|
||||||
| NC score | 0.014363 (rank : 73) | θ value | 2.36792 (rank : 72) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8R0A5, Q9CYK5, Q9D1S4 | Gene names | Tceal3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 3 (TCEA-like protein 3) (Transcription elongation factor S-II protein-like 3). | |||||
|
SPAG1_MOUSE
|
||||||
| NC score | 0.013607 (rank : 74) | θ value | 3.0926 (rank : 73) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80ZX8, Q8CCK7, Q9QZJ3, Q9QZJ4 | Gene names | Spag1, Tpis | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-associated antigen 1 (Infertility-related sperm protein Spag-1) (TPR-containing protein involved in spermatogenesis) (TPIS). | |||||
|
AP3B2_MOUSE
|
||||||
| NC score | 0.012350 (rank : 75) | θ value | 1.81305 (rank : 67) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9JME5, Q6QR53, Q8R1E5 | Gene names | Ap3b2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP-3 complex subunit beta-2 (Adapter-related protein complex 3 beta-2 subunit) (Beta3B-adaptin) (Adaptor protein complex AP-3 beta-2 subunit) (Clathrin assembly protein complex 3 beta-2 large chain). | |||||
|
DDX49_HUMAN
|
||||||
| NC score | 0.009922 (rank : 76) | θ value | 0.47712 (rank : 55) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y6V7, Q53FJ1 | Gene names | DDX49 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX49 (EC 3.6.1.-) (DEAD box protein 49). | |||||
|
OSBL5_HUMAN
|
||||||
| NC score | 0.008446 (rank : 77) | θ value | 4.03905 (rank : 75) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9H0X9, Q9BZB0, Q9P1Z4 | Gene names | OSBPL5, KIAA1534, ORP5 | |||
|
Domain Architecture |

|
|||||
| Description | Oxysterol-binding protein-related protein 5 (OSBP-related protein 5) (ORP-5). | |||||
|
RUFY3_HUMAN
|
||||||
| NC score | 0.007381 (rank : 78) | θ value | 1.38821 (rank : 65) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 594 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7L099, O94948, Q9UI00 | Gene names | RUFY3, KIAA0871 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein RUFY3 (Rap2-interacting protein x) (RIPx). | |||||
|
MAST4_HUMAN
|
||||||
| NC score | 0.005324 (rank : 79) | θ value | 1.38821 (rank : 64) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1804 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O15021, Q6ZN07, Q96LY3 | Gene names | MAST4, KIAA0303 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase 4 (EC 2.7.11.1). | |||||
|
DUOX1_HUMAN
|
||||||
| NC score | 0.003390 (rank : 80) | θ value | 8.99809 (rank : 87) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NRD9, Q6ZMB3, Q6ZR09, Q9NZC1 | Gene names | DUOX1, DUOX, LNOX1, THOX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dual oxidase 1 precursor (EC 1.6.3.1) (EC 1.11.1.-) (NADPH thyroid oxidase 1) (Thyroid oxidase 1) (Large NOX 1) (Long NOX 1). | |||||
|
TEX14_MOUSE
|
||||||
| NC score | 0.003372 (rank : 81) | θ value | 1.38821 (rank : 66) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1333 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q7M6U3, Q3UT36, Q5NC10, Q5NC11, Q8CGK1, Q8CGK2, Q99MV8 | Gene names | Tex14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-expressed protein 14 (Testis-expressed sequence 14). | |||||
|
ULK2_MOUSE
|
||||||
| NC score | 0.001606 (rank : 82) | θ value | 8.99809 (rank : 88) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 943 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9QY01, Q80TV7, Q9WTP4 | Gene names | Ulk2, Kiaa0623 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase ULK2 (EC 2.7.11.1) (Unc-51-like kinase 2) (Serine/threonine-protein kinase Unc51.2). | |||||
|
ANK2_HUMAN
|
||||||
| NC score | 0.000914 (rank : 83) | θ value | 6.88961 (rank : 83) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
RABE1_MOUSE
|
||||||
| NC score | 0.000295 (rank : 84) | θ value | 5.27518 (rank : 81) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 1105 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O35551, Q99L08, Q9CRP3, Q9EQF9, Q9JI94 | Gene names | Rabep1, Rab5ep, Rabpt5, Rabpt5a | |||
|
Domain Architecture |

|
|||||
| Description | Rab GTPase-binding effector protein 1 (Rabaptin-5) (Rabaptin-5alpha). | |||||
|
LTBP3_HUMAN
|
||||||
| NC score | -0.000115 (rank : 85) | θ value | 6.88961 (rank : 84) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 502 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NS15, O15107, Q96HB9, Q9H7K2, Q9UFN4 | Gene names | LTBP3 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein 3 precursor (LTBP-3). | |||||
|
ZN579_MOUSE
|
||||||
| NC score | -0.002176 (rank : 86) | θ value | 6.88961 (rank : 86) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 719 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80VM4 | Gene names | Znf579, Zfp579 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 579. | |||||
|
ZN184_HUMAN
|
||||||
| NC score | -0.002314 (rank : 87) | θ value | 1.06291 (rank : 63) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 811 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q99676, O60792, Q8TBA9 | Gene names | ZNF184 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 184. | |||||
|
ZN426_HUMAN
|
||||||
| NC score | -0.002757 (rank : 88) | θ value | 8.99809 (rank : 89) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 747 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9BUY5 | Gene names | ZNF426 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 426. | |||||
|
ZN264_HUMAN
|
||||||
| NC score | -0.003045 (rank : 89) | θ value | 6.88961 (rank : 85) | |||
| Query Neighborhood Hits | 89 | Target Neighborhood Hits | 770 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O43296, Q9P1V0 | Gene names | ZNF264, KIAA0412 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 264. | |||||