Please be patient as the page loads
|
IQEC3_HUMAN
|
||||||
| SwissProt Accessions | Q9UPP2, Q8TB43 | Gene names | IQSEC3, KIAA1110 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 3. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
IQEC3_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 125 | |
| SwissProt Accessions | Q9UPP2, Q8TB43 | Gene names | IQSEC3, KIAA1110 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 3. | |||||
|
IQEC3_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.968732 (rank : 2) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q3TES0, Q3UHP8, Q80TJ8 | Gene names | Iqsec3, Kiaa1110 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 3. | |||||
|
IQEC1_HUMAN
|
||||||
| θ value | 1.12714e-143 (rank : 3) | NC score | 0.965990 (rank : 4) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6DN90, O94863, Q96D85 | Gene names | IQSEC1, ARFGEP100, BRAG2, KIAA0763 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 1 (ADP-ribosylation factors guanine nucleotide-exchange protein 100) (ADP-ribosylation factors guanine nucleotide-exchange protein 2) (Brefeldin-resistant Arf-GEF 2 protein). | |||||
|
IQEC1_MOUSE
|
||||||
| θ value | 1.12714e-143 (rank : 4) | NC score | 0.966666 (rank : 3) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8R0S2, Q3TZC3, Q5DU15 | Gene names | Iqsec1, Kiaa0763 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 1. | |||||
|
IQEC2_HUMAN
|
||||||
| θ value | 2.60451e-132 (rank : 5) | NC score | 0.924435 (rank : 5) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q5JU85, O60275 | Gene names | IQSEC2, KIAA0522 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 2. | |||||
|
IQEC2_MOUSE
|
||||||
| θ value | 7.57797e-132 (rank : 6) | NC score | 0.923416 (rank : 6) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q5DU25 | Gene names | Iqsec2, Kiaa0522 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 2. | |||||
|
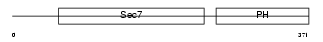
CYH2_HUMAN
|
||||||
| θ value | 4.41421e-39 (rank : 7) | NC score | 0.808988 (rank : 10) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q99418, Q92958 | Gene names | PSCD2, ARNO | |||
|
Domain Architecture |
|
|||||
| Description | Cytohesin-2 (ARF nucleotide-binding site opener) (ARNO protein) (ARF exchange factor). | |||||
|
CYH2_MOUSE
|
||||||
| θ value | 5.76516e-39 (rank : 8) | NC score | 0.809385 (rank : 9) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P63034, O89099, P97695, Q3UP39 | Gene names | Pscd2, Sec7b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytohesin-2 (CLM2) (SEC7 homolog B) (msec7-2). | |||||
|
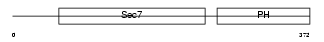
CYH1_MOUSE
|
||||||
| θ value | 2.19075e-38 (rank : 9) | NC score | 0.801783 (rank : 13) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9QX11, O88817 | Gene names | Pscd1 | |||
|
Domain Architecture |
|
|||||
| Description | Cytohesin-1 (CLM1). | |||||
|
CYH3_HUMAN
|
||||||
| θ value | 3.73686e-38 (rank : 10) | NC score | 0.805286 (rank : 12) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O43739 | Gene names | PSCD3, ARNO3, GRP1 | |||
|
Domain Architecture |

|
|||||
| Description | Cytohesin-3 (ARF nucleotide-binding site opener 3) (ARNO3 protein) (General receptor of phosphoinositides 1) (Grp1). | |||||
|
CYH1_HUMAN
|
||||||
| θ value | 4.88048e-38 (rank : 11) | NC score | 0.799134 (rank : 16) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q15438, Q9P123, Q9P124 | Gene names | PSCD1, D17S811E | |||
|
Domain Architecture |

|
|||||
| Description | Cytohesin-1 (SEC7 homolog B2-1). | |||||
|
CYH3_MOUSE
|
||||||
| θ value | 6.37413e-38 (rank : 12) | NC score | 0.800355 (rank : 14) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O08967, Q8CI93 | Gene names | Pscd3, Grp1 | |||
|
Domain Architecture |

|
|||||
| Description | Cytohesin-3 (ARF nucleotide-binding site opener 3) (ARNO3 protein) (General receptor of phosphoinositides 1) (Grp1) (CLM-3). | |||||
|
BIG1_HUMAN
|
||||||
| θ value | 3.4976e-36 (rank : 13) | NC score | 0.822834 (rank : 8) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9Y6D6, Q9NV46, Q9UFV2, Q9UNL0 | Gene names | ARFGEF1, ARFGEP1, BIG1 | |||
|
Domain Architecture |

|
|||||
| Description | Brefeldin A-inhibited guanine nucleotide-exchange protein 1 (Brefeldin A-inhibited GEP 1) (p200 ARF-GEP1) (p200 ARF guanine nucleotide exchange factor). | |||||
|
CYH4_MOUSE
|
||||||
| θ value | 8.06329e-33 (rank : 14) | NC score | 0.799366 (rank : 15) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q80YW0 | Gene names | Pscd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytohesin-4. | |||||
|
CYH4_HUMAN
|
||||||
| θ value | 8.91499e-32 (rank : 15) | NC score | 0.798601 (rank : 17) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9UIA0, Q5R3F9, Q9UGT6 | Gene names | PSCD4, CYT4 | |||
|
Domain Architecture |

|
|||||
| Description | Cytohesin-4. | |||||
|
BIG2_HUMAN
|
||||||
| θ value | 4.42448e-31 (rank : 16) | NC score | 0.805995 (rank : 11) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9Y6D5, Q5TFT9, Q9NTS1 | Gene names | ARFGEF2, ARFGEP2, BIG2 | |||
|
Domain Architecture |

|
|||||
| Description | Brefeldin A-inhibited guanine nucleotide-exchange protein 2 (Brefeldin A-inhibited GEP 2). | |||||
|
GBF1_HUMAN
|
||||||
| θ value | 1.85889e-29 (rank : 17) | NC score | 0.824317 (rank : 7) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q92538, Q5VXX3, Q96CK6, Q96HZ3, Q9H473 | Gene names | GBF1, KIAA0248 | |||
|
Domain Architecture |

|
|||||
| Description | Golgi-specific brefeldin A-resistance guanine nucleotide exchange factor 1 (BFA-resistant GEF 1). | |||||
|
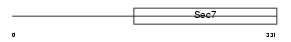
FBX8_HUMAN
|
||||||
| θ value | 1.24977e-17 (rank : 18) | NC score | 0.784027 (rank : 19) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NRD0, Q6UWN4, Q9NRP5, Q9UKC4 | Gene names | FBXO8, FBS, FBX8 | |||
|
Domain Architecture |
|
|||||
| Description | F-box only protein 8 (F-box/SEC7 protein FBS). | |||||
|
PSD3_HUMAN
|
||||||
| θ value | 1.63225e-17 (rank : 19) | NC score | 0.736920 (rank : 20) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9NYI0, Q9Y2F1 | Gene names | PSD3, EFA6R, HCA67, KIAA0942 | |||
|
Domain Architecture |

|
|||||
| Description | PH and SEC7 domain-containing protein 3 (Pleckstrin homology and SEC7 domain-containing protein 3) (Exchange factor for ADP-ribosylation factor guanine nucleotide factor 6) (Hepatocellular carcinoma- associated antigen 67). | |||||
|
PSD4_HUMAN
|
||||||
| θ value | 1.63225e-17 (rank : 20) | NC score | 0.700195 (rank : 22) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8NDX1, O95621, Q4ZG34, Q6GPH8, Q8IYP4 | Gene names | PSD4, EFA6B, TIC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH and SEC7 domain-containing protein 4 (Pleckstrin homology and SEC7 domain-containing protein 4) (Exchange factor for ADP-ribosylation factor guanine nucleotide factor 6B) (Telomeric of interleukin-1 cluster protein). | |||||
|
PSD4_MOUSE
|
||||||
| θ value | 2.7842e-17 (rank : 21) | NC score | 0.705578 (rank : 21) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8BLR5, Q3TE33, Q3TQF5, Q3UD59, Q3UFH9, Q80V44 | Gene names | Psd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH and SEC7 domain-containing protein 4 (Pleckstrin homology and SEC7 domain-containing protein 4). | |||||
|
FBX8_MOUSE
|
||||||
| θ value | 3.63628e-17 (rank : 22) | NC score | 0.785143 (rank : 18) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9QZN3 | Gene names | Fbxo8, Fbx8 | |||
|
Domain Architecture |

|
|||||
| Description | F-box only protein 8. | |||||
|
FARP1_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 23) | NC score | 0.065112 (rank : 48) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9Y4F1, Q6IQ29 | Gene names | FARP1, CDEP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FERM, RhoGEF and pleckstrin domain-containing protein 1 (Chondrocyte- derived ezrin-like protein). | |||||
|
FGD4_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 24) | NC score | 0.060622 (rank : 52) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q96M96, Q6ULS2, Q8TCP6 | Gene names | FGD4, FRABP, ZFYVE6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 4 (Actin filament- binding protein frabin) (FGD1-related F-actin-binding protein) (Zinc finger FYVE domain-containing protein 6). | |||||
|
ATS13_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 25) | NC score | 0.017901 (rank : 104) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q76LX8, Q6UY16, Q710F6, Q711T8, Q96L37, Q9H0G3 | Gene names | ADAMTS13, C9orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAMTS-13 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase with thrombospondin motifs 13) (ADAM-TS 13) (ADAM-TS13) (von Willebrand factor-cleaving protease) (vWF-cleaving protease) (vWF-CP). | |||||
|
CAC1G_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 26) | NC score | 0.037014 (rank : 70) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 587 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O43497, O43498, O94770, Q9NYU4, Q9NYU5, Q9NYU6, Q9NYU7, Q9NYU8, Q9NYU9, Q9NYV0, Q9NYV1, Q9UHN9, Q9UHP0, Q9ULU6, Q9UNG7, Q9Y5T2, Q9Y5T3 | Gene names | CACNA1G, KIAA1123 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1G (Voltage- gated calcium channel subunit alpha Cav3.1) (Cav3.1c) (NBR13). | |||||
|
FGD4_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 27) | NC score | 0.059286 (rank : 54) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q91ZT5, Q3UEB6, Q8BW60, Q8BZI7, Q91ZT3, Q91ZT4 | Gene names | Fgd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 4 (Actin filament- binding protein frabin) (FGD1-related F-actin-binding protein). | |||||
|
RGS3_HUMAN
|
||||||
| θ value | 0.125558 (rank : 28) | NC score | 0.031564 (rank : 77) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
ZN142_HUMAN
|
||||||
| θ value | 0.125558 (rank : 29) | NC score | 0.003140 (rank : 151) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1007 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P52746, Q92510 | Gene names | ZNF142, KIAA0236 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 142 (HA4654). | |||||
|
CJ047_HUMAN
|
||||||
| θ value | 0.163984 (rank : 30) | NC score | 0.043387 (rank : 66) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q86WR7, Q5W0J9, Q5W0K0, Q5W0K1, Q5W0K2, Q6PJC8, Q8N317 | Gene names | C10orf47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf47. | |||||
|
PGCA_MOUSE
|
||||||
| θ value | 0.163984 (rank : 31) | NC score | 0.015327 (rank : 116) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 643 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q61282, Q64021 | Gene names | Agc1, Agc | |||
|
Domain Architecture |

|
|||||
| Description | Aggrecan core protein precursor (Cartilage-specific proteoglycan core protein) (CSPCP). | |||||
|
EP300_HUMAN
|
||||||
| θ value | 0.21417 (rank : 32) | NC score | 0.040919 (rank : 68) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q09472 | Gene names | EP300, P300 | |||
|
Domain Architecture |

|
|||||
| Description | E1A-associated protein p300 (EC 2.3.1.48). | |||||
|
RFFL_HUMAN
|
||||||
| θ value | 0.21417 (rank : 33) | NC score | 0.023313 (rank : 92) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8WZ73, Q8NHW0, Q8TBY7, Q96BE6 | Gene names | RFFL, RNF189, RNF34L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rififylin (RING finger and FYVE-like domain-containing protein 1) (FYVE-RING finger protein Sakura) (Fring) (Caspases-8 and -10- associated RING finger protein 2) (CARP-2) (Caspase regulator CARP2) (RING finger protein 189) (RING finger protein 34-like). | |||||
|
NFH_MOUSE
|
||||||
| θ value | 0.279714 (rank : 34) | NC score | 0.028283 (rank : 84) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
FGD6_HUMAN
|
||||||
| θ value | 0.365318 (rank : 35) | NC score | 0.058421 (rank : 55) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 237 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q6ZV73, Q6ZR53, Q7Z2Z7, Q96D44, Q9NUR8, Q9P2I5 | Gene names | FGD6, KIAA1362, ZFYVE24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 6 (Zinc finger FYVE domain-containing protein 24). | |||||
|
FGD6_MOUSE
|
||||||
| θ value | 0.365318 (rank : 36) | NC score | 0.057272 (rank : 56) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q69ZL1, Q8C8W5, Q8K3B0, Q9D3Y7 | Gene names | Fgd6, Kiaa1362 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 6. | |||||
|
ZEP1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 37) | NC score | 0.012204 (rank : 127) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1036 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P15822, Q14122 | Gene names | HIVEP1, ZNF40 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 40 (Human immunodeficiency virus type I enhancer- binding protein 1) (HIV-EP1) (Major histocompatibility complex-binding protein 1) (MBP-1) (Positive regulatory domain II-binding factor 1) (PRDII-BF1). | |||||
|
AKAP8_HUMAN
|
||||||
| θ value | 0.47712 (rank : 38) | NC score | 0.023609 (rank : 89) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O43823 | Gene names | AKAP8, AKAP95 | |||
|
Domain Architecture |

|
|||||
| Description | A-kinase anchor protein 8 (A-kinase anchor protein 95 kDa) (AKAP 95). | |||||
|
NSMA2_MOUSE
|
||||||
| θ value | 0.47712 (rank : 39) | NC score | 0.036437 (rank : 71) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9JJY3, Q3UTQ5 | Gene names | smpd3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sphingomyelin phosphodiesterase 3 (EC 3.1.4.12) (Neutral sphingomyelinase 2) (Neutral sphingomyelinase II) (nSMase2) (nSMase- 2). | |||||
|
OXR1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 40) | NC score | 0.020713 (rank : 97) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q4KMM3, Q5FWW1, Q99L06, Q99MK1, Q99MP4 | Gene names | Oxr1, C7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oxidation resistance protein 1 (Protein C7). | |||||
|
PRRT3_HUMAN
|
||||||
| θ value | 0.47712 (rank : 41) | NC score | 0.045145 (rank : 65) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q5FWE3, Q49AD0, Q6UXY6, Q8NBC9 | Gene names | PRRT3, UNQ5823/PRO19642 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
FOXP4_MOUSE
|
||||||
| θ value | 0.62314 (rank : 42) | NC score | 0.015400 (rank : 114) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9DBY0, Q80V92, Q8CG10, Q8CIS1 | Gene names | Foxp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein P4 (Fork head-related protein like A) (mFKHLA). | |||||
|
GCC2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 43) | NC score | 0.005841 (rank : 143) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1619 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8CHG3, Q8BR44, Q8R2Q5, Q9CT45 | Gene names | Gcc2, Kiaa0336 | |||
|
Domain Architecture |

|
|||||
| Description | GRIP and coiled-coil domain-containing protein 2 (Golgi coiled coil protein GCC185). | |||||
|
DDX51_HUMAN
|
||||||
| θ value | 0.813845 (rank : 44) | NC score | 0.012279 (rank : 126) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8N8A6, Q5CZ71, Q8IXK5, Q96ED1 | Gene names | DDX51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX51 (EC 3.6.1.-) (DEAD box protein 51). | |||||
|
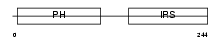
DOK2_MOUSE
|
||||||
| θ value | 0.813845 (rank : 45) | NC score | 0.018636 (rank : 103) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 244 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O70469, O70272, Q99KL1 | Gene names | Dok2, Frip | |||
|
Domain Architecture |
|
|||||
| Description | Docking protein 2 (Downstream of tyrosine kinase 2) (p56(dok-2)) (Dok- related protein) (Dok-R) (IL-four receptor-interacting protein) (FRIP). | |||||
|
FARP2_MOUSE
|
||||||
| θ value | 0.813845 (rank : 46) | NC score | 0.051687 (rank : 61) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q91VS8, Q3TAP2, Q69ZZ0 | Gene names | Farp2, Kiaa0793 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FERM, RhoGEF and pleckstrin domain-containing protein 2 (FERM domain including RhoGEF) (FIR). | |||||
|
MLL3_HUMAN
|
||||||
| θ value | 0.813845 (rank : 47) | NC score | 0.031658 (rank : 75) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
SPEG_MOUSE
|
||||||
| θ value | 0.813845 (rank : 48) | NC score | 0.004918 (rank : 146) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
ASPX_HUMAN
|
||||||
| θ value | 1.06291 (rank : 49) | NC score | 0.029175 (rank : 82) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P26436 | Gene names | ACRV1 | |||
|
Domain Architecture |

|
|||||
| Description | Acrosomal protein SP-10 precursor (Acrosomal vesicle protein 1). | |||||
|
HSF2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 50) | NC score | 0.032602 (rank : 74) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q03933, Q9H445 | Gene names | HSF2, HSTF2 | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock factor protein 2 (HSF 2) (Heat shock transcription factor 2) (HSTF 2). | |||||
|
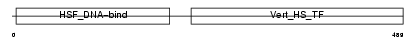
HSF2_MOUSE
|
||||||
| θ value | 1.06291 (rank : 51) | NC score | 0.032904 (rank : 73) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P38533, Q64157, Q8VEJ0, Q9Z2L8 | Gene names | Hsf2 | |||
|
Domain Architecture |
|
|||||
| Description | Heat shock factor protein 2 (HSF 2) (Heat shock transcription factor 2) (HSTF 2). | |||||
|
DHX16_HUMAN
|
||||||
| θ value | 1.38821 (rank : 52) | NC score | 0.008030 (rank : 138) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O60231, O60322, Q969X7, Q96QC1 | Gene names | DHX16, DBP2, DDX16, KIAA0577 | |||
|
Domain Architecture |

|
|||||
| Description | Putative pre-mRNA-splicing factor ATP-dependent RNA helicase DHX16 (EC 3.6.1.-) (DEAH-box protein 16) (ATP-dependent RNA helicase #3). | |||||
|
TP53B_MOUSE
|
||||||
| θ value | 1.38821 (rank : 53) | NC score | 0.030998 (rank : 79) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 644 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P70399, Q68FD0, Q8CI97, Q91YC9 | Gene names | Tp53bp1, Trp53bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor suppressor p53-binding protein 1 (p53-binding protein 1) (p53BP1) (53BP1). | |||||
|
AN30A_HUMAN
|
||||||
| θ value | 1.81305 (rank : 54) | NC score | 0.006317 (rank : 142) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1119 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9BXX3, Q5W025 | Gene names | ANKRD30A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 30A (NY-BR-1 antigen). | |||||
|
ASPM_MOUSE
|
||||||
| θ value | 1.81305 (rank : 55) | NC score | 0.017552 (rank : 106) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CJ27, O88482, Q8BJI8, Q8BKT4 | Gene names | Aspm, Calmbp1, Sha1 | |||
|
Domain Architecture |

|
|||||
| Description | Abnormal spindle-like microcephaly-associated protein homolog (Calmodulin-binding protein 1) (Spindle and hydroxyurea checkpoint abnormal protein) (Calmodulin-binding protein Sha1). | |||||
|
BSN_HUMAN
|
||||||
| θ value | 1.81305 (rank : 56) | NC score | 0.033640 (rank : 72) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
MUC1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 57) | NC score | 0.031056 (rank : 78) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
OGFR_HUMAN
|
||||||
| θ value | 1.81305 (rank : 58) | NC score | 0.040974 (rank : 67) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
TAF1C_MOUSE
|
||||||
| θ value | 1.81305 (rank : 59) | NC score | 0.025179 (rank : 87) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6PDZ2, P97359, Q3U3B6, Q8BN54 | Gene names | Taf1c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TATA box-binding protein-associated factor, RNA polymerase I, subunit C (TATA box-binding protein-associated factor 1C) (TBP-associated factor 1C) (TBP-associated factor RNA polymerase I 95 kDa) (TAFI95). | |||||
|
CHSS2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 60) | NC score | 0.012535 (rank : 123) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6IQX7 | Gene names | Css2, D1Bwg1363e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate synthase 2 (EC 2.4.1.175) (Glucuronosyl-N- acetylgalactosaminyl-proteoglycan 4-beta-N- acetylgalactosaminyltransferase II) (N-acetylgalactosaminyl- proteoglycan 3-beta-glucuronosyltransferase II) (EC 2.4.1.226) (Chondroitin glucuronyltransferase II) (N- acetylgalactosaminyltransferase). | |||||
|
CLIC6_HUMAN
|
||||||
| θ value | 2.36792 (rank : 61) | NC score | 0.023463 (rank : 91) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q96NY7, Q8IX31 | Gene names | CLIC6, CLIC1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride intracellular channel 6. | |||||
|
ERCC5_MOUSE
|
||||||
| θ value | 2.36792 (rank : 62) | NC score | 0.028112 (rank : 85) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P35689, Q61528, Q64248 | Gene names | Ercc5, Ercc-5, Xpg | |||
|
Domain Architecture |

|
|||||
| Description | DNA-repair protein complementing XP-G cells homolog (Xeroderma pigmentosum group G-complementing protein homolog) (DNA excision repair protein ERCC-5). | |||||
|
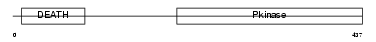
IRAK2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 63) | NC score | 0.003212 (rank : 150) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 714 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O43187 | Gene names | IRAK2 | |||
|
Domain Architecture |
|
|||||
| Description | Interleukin-1 receptor-associated kinase-like 2 (IRAK-2). | |||||
|
MYO3A_HUMAN
|
||||||
| θ value | 2.36792 (rank : 64) | NC score | -0.000209 (rank : 154) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1033 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8NEV4, Q8WX17, Q9NYS8 | Gene names | MYO3A | |||
|
Domain Architecture |

|
|||||
| Description | Myosin IIIA (EC 2.7.11.1). | |||||
|
NALDL_HUMAN
|
||||||
| θ value | 2.36792 (rank : 65) | NC score | 0.017173 (rank : 107) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UQQ1, O43176 | Gene names | NAALADL1, NAALADASEL, NAALADL | |||
|
Domain Architecture |

|
|||||
| Description | N-acetylated-alpha-linked acidic dipeptidase-like protein (EC 3.4.17.21) (NAALADase L) (Ileal dipeptidylpeptidase) (100 kDa ileum brush border membrane protein) (I100). | |||||
|
PAWR_HUMAN
|
||||||
| θ value | 2.36792 (rank : 66) | NC score | 0.029698 (rank : 81) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 252 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96IZ0, O75796, Q6FHY9, Q8N700 | Gene names | PAWR, PAR4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PRKC apoptosis WT1 regulator protein (Prostate apoptosis response 4 protein) (Par-4). | |||||
|
PKD1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 67) | NC score | 0.016195 (rank : 111) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P98161, Q15140, Q15141 | Gene names | PKD1 | |||
|
Domain Architecture |

|
|||||
| Description | Polycystin-1 precursor (Autosomal dominant polycystic kidney disease protein 1). | |||||
|
PTN23_HUMAN
|
||||||
| θ value | 2.36792 (rank : 68) | NC score | 0.017564 (rank : 105) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 646 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9H3S7, Q7KZF8, Q8N6Z5, Q9BSR5, Q9P257, Q9UG03, Q9UMZ4 | Gene names | PTPN23, KIAA1471 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 23 (EC 3.1.3.48) (His- domain-containing protein tyrosine phosphatase) (HD-PTP) (Protein tyrosine phosphatase TD14) (PTP-TD14). | |||||
|
RIMB1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 69) | NC score | 0.020689 (rank : 98) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1479 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O95153, O75111, Q8N5W3 | Gene names | BZRAP1, KIAA0612, RBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Peripheral-type benzodiazepine receptor-associated protein 1 (PRAX-1) (Peripheral benzodiazepine receptor-interacting protein) (PBR-IP) (RIM-binding protein 1) (RIM-BP1). | |||||
|
WDR81_MOUSE
|
||||||
| θ value | 2.36792 (rank : 70) | NC score | 0.011713 (rank : 129) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5ND34, Q3TA90, Q3TUS9, Q6KAR1 | Gene names | Wdr81 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 81. | |||||
|
CD2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 71) | NC score | 0.022235 (rank : 95) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P06729, Q96TE5 | Gene names | CD2 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell surface antigen CD2 precursor (T-cell surface antigen T11/Leu- 5) (LFA-2) (LFA-3 receptor) (Erythrocyte receptor) (Rosette receptor). | |||||
|
CI082_MOUSE
|
||||||
| θ value | 3.0926 (rank : 72) | NC score | 0.031572 (rank : 76) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8VDY9 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C9orf82 homolog. | |||||
|
HIP1R_MOUSE
|
||||||
| θ value | 3.0926 (rank : 73) | NC score | 0.007416 (rank : 140) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 959 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9JKY5 | Gene names | Hip1r | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin-interacting protein 1-related protein (Hip1-related). | |||||
|
K1802_MOUSE
|
||||||
| θ value | 3.0926 (rank : 74) | NC score | 0.030822 (rank : 80) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
MDC1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 75) | NC score | 0.017036 (rank : 109) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
ZN592_HUMAN
|
||||||
| θ value | 3.0926 (rank : 76) | NC score | 0.002465 (rank : 152) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 762 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q92610 | Gene names | ZNF592, KIAA0211 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 592. | |||||
|
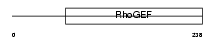
ARHGA_HUMAN
|
||||||
| θ value | 4.03905 (rank : 77) | NC score | 0.024138 (rank : 88) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O15013, O14665, Q68D55, Q8IWD9, Q8IY77 | Gene names | ARHGEF10, KIAA0294 | |||
|
Domain Architecture |
|
|||||
| Description | Rho guanine nucleotide exchange factor 10. | |||||
|
CS021_HUMAN
|
||||||
| θ value | 4.03905 (rank : 78) | NC score | 0.019384 (rank : 99) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8IVT2 | Gene names | C19orf21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C19orf21. | |||||
|
DPOG1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 79) | NC score | 0.012685 (rank : 121) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P54099, Q9JI28 | Gene names | Polg, Polg1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase subunit gamma 1 (EC 2.7.7.7) (Mitochondrial DNA polymerase catalytic subunit) (PolG-alpha). | |||||
|
FOXN1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 80) | NC score | 0.015651 (rank : 113) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 377 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O15353, O15352 | Gene names | FOXN1, WHN | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein N1 (Transcription factor winged-helix nude). | |||||
|
FZD6_MOUSE
|
||||||
| θ value | 4.03905 (rank : 81) | NC score | 0.004667 (rank : 147) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q61089 | Gene names | Fzd6 | |||
|
Domain Architecture |

|
|||||
| Description | Frizzled-6 precursor (Fz-6) (mFz6). | |||||
|
GPTC8_HUMAN
|
||||||
| θ value | 4.03905 (rank : 82) | NC score | 0.018948 (rank : 101) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 366 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9UKJ3, O60300 | Gene names | GPATC8, KIAA0553 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 8. | |||||
|
KLF15_HUMAN
|
||||||
| θ value | 4.03905 (rank : 83) | NC score | -0.000392 (rank : 155) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 720 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UIH9 | Gene names | KLF15, KKLF | |||
|
Domain Architecture |

|
|||||
| Description | Krueppel-like factor 15 (Kidney-enriched krueppel-like factor). | |||||
|
LPP_HUMAN
|
||||||
| θ value | 4.03905 (rank : 84) | NC score | 0.015212 (rank : 118) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 775 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q93052, Q8NFX5 | Gene names | LPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipoma-preferred partner (LIM domain-containing preferred translocation partner in lipoma). | |||||
|
MAP1A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 85) | NC score | 0.016691 (rank : 110) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P78559, O95643, Q12973, Q15882, Q9UJT4 | Gene names | MAP1A | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 1A (MAP 1A) (Proliferation-related protein p80) [Contains: MAP1 light chain LC2]. | |||||
|
NALDL_MOUSE
|
||||||
| θ value | 4.03905 (rank : 86) | NC score | 0.015358 (rank : 115) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7M758 | Gene names | Naaladl1, Naaladasel | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylated-alpha-linked acidic dipeptidase-like protein (EC 3.4.17.21) (NAALADase L). | |||||
|
SALL2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 87) | NC score | 0.005199 (rank : 145) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 835 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9Y467, Q9Y4G1 | Gene names | SALL2, KIAA0360, SAL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sal-like protein 2 (Zinc finger protein SALL2) (HSal2). | |||||
|
SAPS1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 88) | NC score | 0.023565 (rank : 90) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9UPN7, Q504V2, Q6NVJ6, Q9BU97 | Gene names | SAPS1, KIAA1115 | |||
|
Domain Architecture |

|
|||||
| Description | SAPS domain family member 1. | |||||
|
SFR14_MOUSE
|
||||||
| θ value | 4.03905 (rank : 89) | NC score | 0.019106 (rank : 100) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8CH09, Q6PG19, Q80UY8, Q8BY32, Q8CFM0 | Gene names | Sfrs14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative splicing factor, arginine/serine-rich 14 (Arginine/serine- rich-splicing factor 14). | |||||
|
TROAP_HUMAN
|
||||||
| θ value | 4.03905 (rank : 90) | NC score | 0.021931 (rank : 96) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q12815 | Gene names | TROAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trophinin-associated protein (Tastin) (Trophinin-assisting protein). | |||||
|
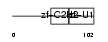
ZEP1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 91) | NC score | 0.009298 (rank : 134) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1063 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q03172 | Gene names | Hivep1, Cryabp1, Znf40 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger protein 40 (Transcription factor alphaA-CRYBP1) (Alpha A- crystallin-binding protein I) (Alpha A-CRYBP1). | |||||
|
ARD1H_HUMAN
|
||||||
| θ value | 5.27518 (rank : 92) | NC score | 0.012605 (rank : 122) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P41227 | Gene names | ARD1A, ARD1, TE2 | |||
|
Domain Architecture |
|
|||||
| Description | N-terminal acetyltransferase complex ARD1 subunit homolog A (EC 2.3.1.88) (EC 2.3.1.-). | |||||
|
ATX7_HUMAN
|
||||||
| θ value | 5.27518 (rank : 93) | NC score | 0.048896 (rank : 63) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O15265, O75328, O75329, Q9Y6P8 | Gene names | ATXN7, SCA7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-7 (Spinocerebellar ataxia type 7 protein). | |||||
|
AXN2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 94) | NC score | 0.012419 (rank : 124) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O88566, Q9QXJ6 | Gene names | Axin2 | |||
|
Domain Architecture |

|
|||||
| Description | Axin-2 (Axis inhibition protein 2) (Conductin) (Axin-like protein) (Axil). | |||||
|
DEN4C_HUMAN
|
||||||
| θ value | 5.27518 (rank : 95) | NC score | 0.009641 (rank : 133) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5VZ89, Q6AI48, Q6ZUB3, Q8NCY7, Q9H6N4, Q9NUT1, Q9NWA5 | Gene names | DENND4C, C9orf55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 4C. | |||||
|
F120A_HUMAN
|
||||||
| θ value | 5.27518 (rank : 96) | NC score | 0.010602 (rank : 130) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NZB2, O60649, Q14688, Q4VXF4, Q4VXF5, Q4VXG2, Q96I21, Q9NZB1 | Gene names | FAM120A, C9orf10, KIAA0183 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UPF0318 protein FAM120A. | |||||
|
FA47A_HUMAN
|
||||||
| θ value | 5.27518 (rank : 97) | NC score | 0.029001 (rank : 83) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 451 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q5JRC9, Q8TAA0 | Gene names | FAM47A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM47A. | |||||
|
FARP2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 98) | NC score | 0.047405 (rank : 64) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O94887, Q53QM5, Q8WU27, Q9UFE7 | Gene names | FARP2, KIAA0793 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FERM, RhoGEF and pleckstrin domain-containing protein 2 (FERM domain including RhoGEF) (FIR). | |||||
|
IWS1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 99) | NC score | 0.018636 (rank : 102) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
MUS81_MOUSE
|
||||||
| θ value | 5.27518 (rank : 100) | NC score | 0.012112 (rank : 128) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q91ZJ0, Q8R356 | Gene names | Mus81 | |||
|
Domain Architecture |

|
|||||
| Description | Crossover junction endonuclease MUS81 (EC 3.1.22.-). | |||||
|
PDZD2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 101) | NC score | 0.009161 (rank : 135) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O15018, Q9BXD4 | Gene names | PDZD2, AIPC, KIAA0300, PDZK3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 2 (PDZ domain-containing protein 3) (Activated in prostate cancer protein). | |||||
|
RBP2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 102) | NC score | 0.004531 (rank : 148) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P49792, Q13074, Q15280, Q53TE2, Q59FH7 | Gene names | RANBP2, NUP358 | |||
|
Domain Architecture |

|
|||||
| Description | Ran-binding protein 2 (RanBP2) (Nuclear pore complex protein Nup358) (Nucleoporin Nup358) (358 kDa nucleoporin) (P270). | |||||
|
BSN_MOUSE
|
||||||
| θ value | 6.88961 (rank : 103) | NC score | 0.037941 (rank : 69) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
CI082_HUMAN
|
||||||
| θ value | 6.88961 (rank : 104) | NC score | 0.026512 (rank : 86) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H8G2, Q5VY32, Q6IPE6, Q96C59 | Gene names | C9orf82 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C9orf82. | |||||
|
CS021_MOUSE
|
||||||
| θ value | 6.88961 (rank : 105) | NC score | 0.017048 (rank : 108) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9D279, Q3UI04 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C19orf21 homolog. | |||||
|
CUTL1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 106) | NC score | 0.008142 (rank : 137) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P53564, O08994, P70301, Q571L6, Q91ZD2 | Gene names | Cutl1, Cux, Cux1, Kiaa4047 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP) (Homeobox protein Cux). | |||||
|
GON4L_MOUSE
|
||||||
| θ value | 6.88961 (rank : 107) | NC score | 0.013085 (rank : 120) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9DB00, Q80TB4, Q91YI9 | Gene names | Gon4l, Gon4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
MSH3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 108) | NC score | 0.015302 (rank : 117) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P20585, Q6PJT5, Q86UQ6, Q92867 | Gene names | MSH3, DUC1, DUG | |||
|
Domain Architecture |

|
|||||
| Description | DNA mismatch repair protein Msh3 (Divergent upstream protein) (DUP) (Mismatch repair protein 1) (MRP1). | |||||
|
MY18B_HUMAN
|
||||||
| θ value | 6.88961 (rank : 109) | NC score | 0.006467 (rank : 141) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1299 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8IUG5, Q8NDI8, Q8TE65, Q8WWS0, Q96KH2, Q96KR8, Q96KR9 | Gene names | MYO18B | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18B (Myosin XVIIIb). | |||||
|
NPBL_MOUSE
|
||||||
| θ value | 6.88961 (rank : 110) | NC score | 0.013355 (rank : 119) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
OXRP_HUMAN
|
||||||
| θ value | 6.88961 (rank : 111) | NC score | 0.009845 (rank : 131) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 544 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9Y4L1 | Gene names | HYOU1, ORP150 | |||
|
Domain Architecture |

|
|||||
| Description | 150 kDa oxygen-regulated protein precursor (Orp150) (Hypoxia up- regulated 1). | |||||
|
SIRT6_HUMAN
|
||||||
| θ value | 6.88961 (rank : 112) | NC score | 0.008943 (rank : 136) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N6T7, O75291, Q6IAF5, Q6PK99, Q8NCD2, Q9BSI5, Q9BWP3, Q9NRC7, Q9UQD1 | Gene names | SIRT6, SIR2L6 | |||
|
Domain Architecture |
|
|||||
| Description | Mono-ADP-ribosyltransferase sirtuin-6 (EC 2.4.2.31) (SIR2-like protein 6). | |||||
|
SON_HUMAN
|
||||||
| θ value | 6.88961 (rank : 113) | NC score | 0.023113 (rank : 93) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
ZBT37_HUMAN
|
||||||
| θ value | 6.88961 (rank : 114) | NC score | -0.000691 (rank : 156) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 815 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5TC79, Q5TC80, Q96M87, Q9BQ88 | Gene names | ZBTB37 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger and BTB domain-containing protein 37. | |||||
|
ZN668_HUMAN
|
||||||
| θ value | 6.88961 (rank : 115) | NC score | -0.001921 (rank : 158) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 761 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q96K58, Q59EV1, Q8N669, Q9H8L4 | Gene names | ZNF668 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 668. | |||||
|
AFAD_HUMAN
|
||||||
| θ value | 8.99809 (rank : 116) | NC score | 0.007889 (rank : 139) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 689 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P55196, O75087, O75088, O75089, Q5TIG6, Q9NSN7, Q9NU92 | Gene names | MLLT4, AF6 | |||
|
Domain Architecture |

|
|||||
| Description | Afadin (Protein AF-6). | |||||
|
ANKS3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 117) | NC score | 0.001026 (rank : 153) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 355 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6ZW76, Q8TF25 | Gene names | ANKS3, KIAA1977 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat and SAM domain-containing protein 3. | |||||
|
BCAS1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 118) | NC score | 0.012325 (rank : 125) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q80YN3, Q9CVA1 | Gene names | Bcas1, Nabc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Breast carcinoma amplified sequence 1 homolog (Novel amplified in breast cancer 1 homolog). | |||||
|
CA156_HUMAN
|
||||||
| θ value | 8.99809 (rank : 119) | NC score | 0.009803 (rank : 132) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O95568 | Gene names | C1orf156, ASTP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C1orf156 (Arsenic-transactivated protein 2) (AsTP2). | |||||
|
CIC_HUMAN
|
||||||
| θ value | 8.99809 (rank : 120) | NC score | 0.015784 (rank : 112) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 992 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q96RK0, Q7LGI1, Q9UEG5, Q9Y6T1 | Gene names | CIC, KIAA0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
F10A1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 121) | NC score | 0.004386 (rank : 149) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P50502, O14999 | Gene names | ST13, FAM10A1, HIP, SNC6 | |||
|
Domain Architecture |
|
|||||
| Description | Hsc70-interacting protein (Hip) (Suppression of tumorigenicity protein 13) (Putative tumor suppressor ST13) (Protein FAM10A1) (Progesterone receptor-associated p48 protein) (NY-REN-33 antigen). | |||||
|
MAGE1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 122) | NC score | 0.022446 (rank : 94) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
PAK4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 123) | NC score | -0.002390 (rank : 159) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1098 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8BTW9, Q6ZPX0, Q80Z97, Q9CS71 | Gene names | Pak4, Kiaa1142 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase PAK 4 (EC 2.7.11.1) (p21-activated kinase 4) (PAK-4). | |||||
|
RB15B_HUMAN
|
||||||
| θ value | 8.99809 (rank : 124) | NC score | 0.005829 (rank : 144) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8NDT2, Q6QE19, Q9BV96 | Gene names | RBM15B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative RNA-binding protein 15B (RNA-binding motif protein 15B). | |||||
|
ZBT16_HUMAN
|
||||||
| θ value | 8.99809 (rank : 125) | NC score | -0.001913 (rank : 157) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1010 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q05516, Q8TAL4 | Gene names | ZBTB16, PLZF, ZNF145 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger and BTB domain-containing protein 16 (Zinc finger protein PLZF) (Promyelocytic leukemia zinc finger protein) (Zinc finger protein 145). | |||||
|
3BP2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 126) | NC score | 0.106322 (rank : 34) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P78314, O00500, O15373, P78315 | Gene names | SH3BP2, 3BP2 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 domain-binding protein 2 (3BP-2). | |||||
|
3BP2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 127) | NC score | 0.108574 (rank : 32) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q06649 | Gene names | Sh3bp2, 3bp2 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 domain-binding protein 2 (3BP-2). | |||||
|
ANLN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 128) | NC score | 0.103558 (rank : 35) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9NQW6, Q5CZ78, Q6NSK5, Q9H8Y4, Q9NVN9, Q9NVP0 | Gene names | ANLN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Actin-binding protein anillin. | |||||
|
ANLN_MOUSE
|
||||||
| θ value | θ > 10 (rank : 129) | NC score | 0.084721 (rank : 44) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 297 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8K298, Q8BL79, Q8BLB3, Q8K2N0, Q9CUY0 | Gene names | Anln | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Actin-binding protein anillin. | |||||
|
BASP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 130) | NC score | 0.051938 (rank : 60) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
BASP_MOUSE
|
||||||
| θ value | θ > 10 (rank : 131) | NC score | 0.053060 (rank : 59) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
CENB2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 132) | NC score | 0.062082 (rank : 50) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q15057, Q9UQR3 | Gene names | CENTB2, KIAA0041 | |||
|
Domain Architecture |

|
|||||
| Description | Centaurin-beta 2 (Cnt-b2). | |||||
|
CENB5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 133) | NC score | 0.053594 (rank : 58) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q96P50, Q9BSR9, Q9C0E7 | Gene names | CENTB5, KIAA1716 | |||
|
Domain Architecture |

|
|||||
| Description | Centaurin-beta 5 (Cnt-b5). | |||||
|
CNKR1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 134) | NC score | 0.173293 (rank : 23) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q969H4, O95381 | Gene names | CNKSR1, CNK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Connector enhancer of kinase suppressor of ras 1 (Connector enhancer of KSR1) (hCNK1) (Connector enhancer of KSR-like) (CNK homolog protein 1). | |||||
|
CNKR2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 135) | NC score | 0.090848 (rank : 41) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8WXI2, O94976, Q8WXI1 | Gene names | CNKSR2, CNK2, KIAA0902, KSR2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Connector enhancer of kinase suppressor of ras 2 (Connector enhancer of KSR2) (CNK2). | |||||
|
CNKR2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 136) | NC score | 0.090928 (rank : 40) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q80YA9, Q80TP2 | Gene names | Cnksr2, Kiaa0902 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Connector enhancer of kinase suppressor of ras 2 (Connector enhancer of KSR2) (CNK2). | |||||
|
DAPP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 137) | NC score | 0.096591 (rank : 38) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UN19, Q8TCK5, Q9UHF2 | Gene names | DAPP1, BAM32 | |||
|
Domain Architecture |
|
|||||
| Description | Dual adapter for phosphotyrosine and 3-phosphotyrosine and 3- phosphoinositide (hDAPP1) (B cell adapter molecule of 32 kDa) (B lymphocyte adapter protein Bam32). | |||||
|
DAPP1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 138) | NC score | 0.101683 (rank : 36) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9QXT1, Q9R178 | Gene names | Dapp1, Bam32 | |||
|
Domain Architecture |
|
|||||
| Description | Dual adapter for phosphotyrosine and 3-phosphotyrosine and 3- phosphoinositide (mDAPP1) (B cell adapter molecule of 32 kDa) (B lymphocyte adapter protein Bam32). | |||||
|
GNRP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 139) | NC score | 0.061602 (rank : 51) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q13972, Q16027 | Gene names | RASGRF1, CDC25 | |||
|
Domain Architecture |

|
|||||
| Description | Guanine nucleotide-releasing protein (GNRP) (Ras-specific nucleotide exchange factor CDC25) (Ras-specific guanine nucleotide-releasing factor). | |||||
|
GNRP_MOUSE
|
||||||
| θ value | θ > 10 (rank : 140) | NC score | 0.060216 (rank : 53) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P27671 | Gene names | Rasgrf1, Cdc25, Grf1 | |||
|
Domain Architecture |

|
|||||
| Description | Guanine nucleotide-releasing protein (GNRP) (Ras-specific nucleotide exchange factor CDC25) (CDC25Mm). | |||||
|
MTMR5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 141) | NC score | 0.050158 (rank : 62) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O95248, O60228, Q5JXD8, Q5PPM2, Q96GR9, Q9UGB8 | Gene names | SBF1, MTMR5 | |||
|
Domain Architecture |

|
|||||
| Description | SET-binding factor 1 (Sbf1) (Myotubularin-related protein 5). | |||||
|
PKHA1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 142) | NC score | 0.120216 (rank : 27) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9HB21, Q9BVK0 | Gene names | PLEKHA1, TAPP1 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 1 (Tandem PH domain-containing protein 1) (TAPP-1). | |||||
|
PKHA1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 143) | NC score | 0.119740 (rank : 29) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8BUL6, Q8BK96, Q8BXU1, Q8VE13 | Gene names | Plekha1, Tapp1 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 1 (Tandem PH domain-containing protein 1) (TAPP-1). | |||||
|
PKHA2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 144) | NC score | 0.097251 (rank : 37) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9HB19 | Gene names | PLEKHA2, TAPP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family A member 2 (Tandem PH domain-containing protein 2) (TAPP-2). | |||||
|
PKHA2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 145) | NC score | 0.110538 (rank : 31) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9ERS5, Q8BY29 | Gene names | Plekha2, Tapp2 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 2 (Tandem PH domain-containing protein 2) (TAPP-2) (PH domain-containing adaptor PHAD47). | |||||
|
PKHA4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 146) | NC score | 0.107470 (rank : 33) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9H4M7, Q8N4M8, Q8N658 | Gene names | PLEKHA4, PEPP1 | |||
|
Domain Architecture |
|
|||||
| Description | Pleckstrin homology domain-containing family A member 4 (Phosphoinositol 3-phosphate-binding protein 1) (PEPP-1). | |||||
|
PKHA4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 147) | NC score | 0.115252 (rank : 30) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8VC98, Q8R3N3 | Gene names | Plekha4, Pepp1 | |||
|
Domain Architecture |
|
|||||
| Description | Pleckstrin homology domain-containing family A member 4 (Phosphoinositol 3-phosphate-binding protein 1) (PEPP-1). | |||||
|
PKHA5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 148) | NC score | 0.091354 (rank : 39) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9HAU0, Q8N3K6, Q96DY9, Q9BVR4, Q9C0H7, Q9H924, Q9NVK8 | Gene names | PLEKHA5, KIAA1686, PEPP2 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 5 (Phosphoinositol 3-phosphate-binding protein 2) (PEPP-2). | |||||
|
PKHA6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 149) | NC score | 0.130866 (rank : 24) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9Y2H5 | Gene names | PLEKHA6, KIAA0969, PEPP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family A member 6 (Phosphoinositol 3-phosphate-binding protein 3) (PEPP-3). | |||||
|
PKHA6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 150) | NC score | 0.126440 (rank : 25) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 591 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q7TQG1, Q8K0J5 | Gene names | Plekha6, Pepp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 6 (Phosphoinositol 3-phosphate-binding protein 3) (PEPP-3). | |||||
|
PLEK2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 151) | NC score | 0.088483 (rank : 43) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NYT0, Q96JT0 | Gene names | PLEK2 | |||
|
Domain Architecture |
|
|||||
| Description | Pleckstrin-2. | |||||
|
PLEK2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 152) | NC score | 0.090014 (rank : 42) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9WV52 | Gene names | Plek2 | |||
|
Domain Architecture |
|
|||||
| Description | Pleckstrin-2. | |||||
|
PLEK_HUMAN
|
||||||
| θ value | θ > 10 (rank : 153) | NC score | 0.120212 (rank : 28) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P08567, Q53SU8, Q6FGM8, Q8WV81 | Gene names | PLEK, P47 | |||
|
Domain Architecture |
|
|||||
| Description | Pleckstrin (Platelet p47 protein). | |||||
|
PLEK_MOUSE
|
||||||
| θ value | θ > 10 (rank : 154) | NC score | 0.122589 (rank : 26) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9JHK5, Q9ERI9 | Gene names | Plek | |||
|
Domain Architecture |
|
|||||
| Description | Pleckstrin. | |||||
|
RASA1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 155) | NC score | 0.069474 (rank : 45) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P20936 | Gene names | RASA1, RASA | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 1 (GTPase-activating protein) (GAP) (Ras p21 protein activator) (p120GAP) (RasGAP). | |||||
|
RHG25_HUMAN
|
||||||
| θ value | θ > 10 (rank : 156) | NC score | 0.065419 (rank : 47) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 550 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P42331, Q8IXQ2 | Gene names | ARHGAP25, KIAA0053 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 25. | |||||
|
RHG25_MOUSE
|
||||||
| θ value | θ > 10 (rank : 157) | NC score | 0.068010 (rank : 46) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 492 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8BYW1, Q2VPR2, Q3TE04, Q3TVX0, Q3UVB7, Q6A0E0, Q8BX98 | Gene names | Arhgap25, Kiaa0053 | |||
|
Domain Architecture |
|
|||||
| Description | Rho-GTPase-activating protein 25. | |||||
|
SWP70_HUMAN
|
||||||
| θ value | θ > 10 (rank : 158) | NC score | 0.063582 (rank : 49) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 973 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9UH65, O75135, Q7LCY6, Q9P061, Q9P0Z8 | Gene names | SWAP70, KIAA0640 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Switch-associated protein 70 (SWAP-70). | |||||
|
SWP70_MOUSE
|
||||||
| θ value | θ > 10 (rank : 159) | NC score | 0.053657 (rank : 57) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1097 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6A028, O88443, Q3TQR6, Q3UCA3, Q6P1D0 | Gene names | Swap70, Kiaa0640 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Switch-associated protein 70 (SWAP-70). | |||||
|
IQEC3_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 125 | |
| SwissProt Accessions | Q9UPP2, Q8TB43 | Gene names | IQSEC3, KIAA1110 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 3. | |||||
|
IQEC3_MOUSE
|
||||||
| NC score | 0.968732 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q3TES0, Q3UHP8, Q80TJ8 | Gene names | Iqsec3, Kiaa1110 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 3. | |||||
|
IQEC1_MOUSE
|
||||||
| NC score | 0.966666 (rank : 3) | θ value | 1.12714e-143 (rank : 4) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8R0S2, Q3TZC3, Q5DU15 | Gene names | Iqsec1, Kiaa0763 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 1. | |||||
|
IQEC1_HUMAN
|
||||||
| NC score | 0.965990 (rank : 4) | θ value | 1.12714e-143 (rank : 3) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6DN90, O94863, Q96D85 | Gene names | IQSEC1, ARFGEP100, BRAG2, KIAA0763 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 1 (ADP-ribosylation factors guanine nucleotide-exchange protein 100) (ADP-ribosylation factors guanine nucleotide-exchange protein 2) (Brefeldin-resistant Arf-GEF 2 protein). | |||||
|
IQEC2_HUMAN
|
||||||
| NC score | 0.924435 (rank : 5) | θ value | 2.60451e-132 (rank : 5) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q5JU85, O60275 | Gene names | IQSEC2, KIAA0522 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 2. | |||||
|
IQEC2_MOUSE
|
||||||
| NC score | 0.923416 (rank : 6) | θ value | 7.57797e-132 (rank : 6) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q5DU25 | Gene names | Iqsec2, Kiaa0522 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 2. | |||||
|
GBF1_HUMAN
|
||||||
| NC score | 0.824317 (rank : 7) | θ value | 1.85889e-29 (rank : 17) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q92538, Q5VXX3, Q96CK6, Q96HZ3, Q9H473 | Gene names | GBF1, KIAA0248 | |||
|
Domain Architecture |

|
|||||
| Description | Golgi-specific brefeldin A-resistance guanine nucleotide exchange factor 1 (BFA-resistant GEF 1). | |||||
|
BIG1_HUMAN
|
||||||
| NC score | 0.822834 (rank : 8) | θ value | 3.4976e-36 (rank : 13) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9Y6D6, Q9NV46, Q9UFV2, Q9UNL0 | Gene names | ARFGEF1, ARFGEP1, BIG1 | |||
|
Domain Architecture |

|
|||||
| Description | Brefeldin A-inhibited guanine nucleotide-exchange protein 1 (Brefeldin A-inhibited GEP 1) (p200 ARF-GEP1) (p200 ARF guanine nucleotide exchange factor). | |||||
|
CYH2_MOUSE
|
||||||
| NC score | 0.809385 (rank : 9) | θ value | 5.76516e-39 (rank : 8) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P63034, O89099, P97695, Q3UP39 | Gene names | Pscd2, Sec7b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytohesin-2 (CLM2) (SEC7 homolog B) (msec7-2). | |||||
|
CYH2_HUMAN
|
||||||
| NC score | 0.808988 (rank : 10) | θ value | 4.41421e-39 (rank : 7) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q99418, Q92958 | Gene names | PSCD2, ARNO | |||
|
Domain Architecture |

|
|||||
| Description | Cytohesin-2 (ARF nucleotide-binding site opener) (ARNO protein) (ARF exchange factor). | |||||
|
BIG2_HUMAN
|
||||||
| NC score | 0.805995 (rank : 11) | θ value | 4.42448e-31 (rank : 16) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9Y6D5, Q5TFT9, Q9NTS1 | Gene names | ARFGEF2, ARFGEP2, BIG2 | |||
|
Domain Architecture |

|
|||||
| Description | Brefeldin A-inhibited guanine nucleotide-exchange protein 2 (Brefeldin A-inhibited GEP 2). | |||||
|
CYH3_HUMAN
|
||||||
| NC score | 0.805286 (rank : 12) | θ value | 3.73686e-38 (rank : 10) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O43739 | Gene names | PSCD3, ARNO3, GRP1 | |||
|
Domain Architecture |

|
|||||
| Description | Cytohesin-3 (ARF nucleotide-binding site opener 3) (ARNO3 protein) (General receptor of phosphoinositides 1) (Grp1). | |||||
|
CYH1_MOUSE
|
||||||
| NC score | 0.801783 (rank : 13) | θ value | 2.19075e-38 (rank : 9) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9QX11, O88817 | Gene names | Pscd1 | |||
|
Domain Architecture |

|
|||||
| Description | Cytohesin-1 (CLM1). | |||||
|
CYH3_MOUSE
|
||||||
| NC score | 0.800355 (rank : 14) | θ value | 6.37413e-38 (rank : 12) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O08967, Q8CI93 | Gene names | Pscd3, Grp1 | |||
|
Domain Architecture |

|
|||||
| Description | Cytohesin-3 (ARF nucleotide-binding site opener 3) (ARNO3 protein) (General receptor of phosphoinositides 1) (Grp1) (CLM-3). | |||||
|
CYH4_MOUSE
|
||||||
| NC score | 0.799366 (rank : 15) | θ value | 8.06329e-33 (rank : 14) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q80YW0 | Gene names | Pscd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytohesin-4. | |||||
|
CYH1_HUMAN
|
||||||
| NC score | 0.799134 (rank : 16) | θ value | 4.88048e-38 (rank : 11) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q15438, Q9P123, Q9P124 | Gene names | PSCD1, D17S811E | |||
|
Domain Architecture |

|
|||||
| Description | Cytohesin-1 (SEC7 homolog B2-1). | |||||
|
CYH4_HUMAN
|
||||||
| NC score | 0.798601 (rank : 17) | θ value | 8.91499e-32 (rank : 15) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9UIA0, Q5R3F9, Q9UGT6 | Gene names | PSCD4, CYT4 | |||
|
Domain Architecture |

|
|||||
| Description | Cytohesin-4. | |||||
|
FBX8_MOUSE
|
||||||
| NC score | 0.785143 (rank : 18) | θ value | 3.63628e-17 (rank : 22) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9QZN3 | Gene names | Fbxo8, Fbx8 | |||
|
Domain Architecture |

|
|||||
| Description | F-box only protein 8. | |||||
|
FBX8_HUMAN
|
||||||
| NC score | 0.784027 (rank : 19) | θ value | 1.24977e-17 (rank : 18) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NRD0, Q6UWN4, Q9NRP5, Q9UKC4 | Gene names | FBXO8, FBS, FBX8 | |||
|
Domain Architecture |

|
|||||
| Description | F-box only protein 8 (F-box/SEC7 protein FBS). | |||||
|
PSD3_HUMAN
|
||||||
| NC score | 0.736920 (rank : 20) | θ value | 1.63225e-17 (rank : 19) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9NYI0, Q9Y2F1 | Gene names | PSD3, EFA6R, HCA67, KIAA0942 | |||
|
Domain Architecture |

|
|||||
| Description | PH and SEC7 domain-containing protein 3 (Pleckstrin homology and SEC7 domain-containing protein 3) (Exchange factor for ADP-ribosylation factor guanine nucleotide factor 6) (Hepatocellular carcinoma- associated antigen 67). | |||||
|
PSD4_MOUSE
|
||||||
| NC score | 0.705578 (rank : 21) | θ value | 2.7842e-17 (rank : 21) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8BLR5, Q3TE33, Q3TQF5, Q3UD59, Q3UFH9, Q80V44 | Gene names | Psd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH and SEC7 domain-containing protein 4 (Pleckstrin homology and SEC7 domain-containing protein 4). | |||||
|
PSD4_HUMAN
|
||||||
| NC score | 0.700195 (rank : 22) | θ value | 1.63225e-17 (rank : 20) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8NDX1, O95621, Q4ZG34, Q6GPH8, Q8IYP4 | Gene names | PSD4, EFA6B, TIC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH and SEC7 domain-containing protein 4 (Pleckstrin homology and SEC7 domain-containing protein 4) (Exchange factor for ADP-ribosylation factor guanine nucleotide factor 6B) (Telomeric of interleukin-1 cluster protein). | |||||
|
CNKR1_HUMAN
|
||||||
| NC score | 0.173293 (rank : 23) | θ value | θ > 10 (rank : 134) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q969H4, O95381 | Gene names | CNKSR1, CNK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Connector enhancer of kinase suppressor of ras 1 (Connector enhancer of KSR1) (hCNK1) (Connector enhancer of KSR-like) (CNK homolog protein 1). | |||||
|
PKHA6_HUMAN
|
||||||
| NC score | 0.130866 (rank : 24) | θ value | θ > 10 (rank : 149) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9Y2H5 | Gene names | PLEKHA6, KIAA0969, PEPP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family A member 6 (Phosphoinositol 3-phosphate-binding protein 3) (PEPP-3). | |||||
|
PKHA6_MOUSE
|
||||||
| NC score | 0.126440 (rank : 25) | θ value | θ > 10 (rank : 150) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 591 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q7TQG1, Q8K0J5 | Gene names | Plekha6, Pepp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 6 (Phosphoinositol 3-phosphate-binding protein 3) (PEPP-3). | |||||
|
PLEK_MOUSE
|
||||||
| NC score | 0.122589 (rank : 26) | θ value | θ > 10 (rank : 154) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9JHK5, Q9ERI9 | Gene names | Plek | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin. | |||||
|
PKHA1_HUMAN
|
||||||
| NC score | 0.120216 (rank : 27) | θ value | θ > 10 (rank : 142) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9HB21, Q9BVK0 | Gene names | PLEKHA1, TAPP1 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 1 (Tandem PH domain-containing protein 1) (TAPP-1). | |||||
|
PLEK_HUMAN
|
||||||
| NC score | 0.120212 (rank : 28) | θ value | θ > 10 (rank : 153) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P08567, Q53SU8, Q6FGM8, Q8WV81 | Gene names | PLEK, P47 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin (Platelet p47 protein). | |||||
|
PKHA1_MOUSE
|
||||||
| NC score | 0.119740 (rank : 29) | θ value | θ > 10 (rank : 143) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8BUL6, Q8BK96, Q8BXU1, Q8VE13 | Gene names | Plekha1, Tapp1 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 1 (Tandem PH domain-containing protein 1) (TAPP-1). | |||||
|
PKHA4_MOUSE
|
||||||
| NC score | 0.115252 (rank : 30) | θ value | θ > 10 (rank : 147) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8VC98, Q8R3N3 | Gene names | Plekha4, Pepp1 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 4 (Phosphoinositol 3-phosphate-binding protein 1) (PEPP-1). | |||||
|
PKHA2_MOUSE
|
||||||
| NC score | 0.110538 (rank : 31) | θ value | θ > 10 (rank : 145) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9ERS5, Q8BY29 | Gene names | Plekha2, Tapp2 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 2 (Tandem PH domain-containing protein 2) (TAPP-2) (PH domain-containing adaptor PHAD47). | |||||
|
3BP2_MOUSE
|
||||||
| NC score | 0.108574 (rank : 32) | θ value | θ > 10 (rank : 127) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q06649 | Gene names | Sh3bp2, 3bp2 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 domain-binding protein 2 (3BP-2). | |||||
|
PKHA4_HUMAN
|
||||||
| NC score | 0.107470 (rank : 33) | θ value | θ > 10 (rank : 146) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9H4M7, Q8N4M8, Q8N658 | Gene names | PLEKHA4, PEPP1 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 4 (Phosphoinositol 3-phosphate-binding protein 1) (PEPP-1). | |||||
|
3BP2_HUMAN
|
||||||
| NC score | 0.106322 (rank : 34) | θ value | θ > 10 (rank : 126) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P78314, O00500, O15373, P78315 | Gene names | SH3BP2, 3BP2 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 domain-binding protein 2 (3BP-2). | |||||
|
ANLN_HUMAN
|
||||||
| NC score | 0.103558 (rank : 35) | θ value | θ > 10 (rank : 128) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9NQW6, Q5CZ78, Q6NSK5, Q9H8Y4, Q9NVN9, Q9NVP0 | Gene names | ANLN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Actin-binding protein anillin. | |||||
|
DAPP1_MOUSE
|
||||||
| NC score | 0.101683 (rank : 36) | θ value | θ > 10 (rank : 138) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9QXT1, Q9R178 | Gene names | Dapp1, Bam32 | |||
|
Domain Architecture |

|
|||||
| Description | Dual adapter for phosphotyrosine and 3-phosphotyrosine and 3- phosphoinositide (mDAPP1) (B cell adapter molecule of 32 kDa) (B lymphocyte adapter protein Bam32). | |||||
|
PKHA2_HUMAN
|
||||||
| NC score | 0.097251 (rank : 37) | θ value | θ > 10 (rank : 144) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9HB19 | Gene names | PLEKHA2, TAPP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family A member 2 (Tandem PH domain-containing protein 2) (TAPP-2). | |||||
|
DAPP1_HUMAN
|
||||||
| NC score | 0.096591 (rank : 38) | θ value | θ > 10 (rank : 137) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UN19, Q8TCK5, Q9UHF2 | Gene names | DAPP1, BAM32 | |||
|
Domain Architecture |

|
|||||
| Description | Dual adapter for phosphotyrosine and 3-phosphotyrosine and 3- phosphoinositide (hDAPP1) (B cell adapter molecule of 32 kDa) (B lymphocyte adapter protein Bam32). | |||||
|
PKHA5_HUMAN
|
||||||
| NC score | 0.091354 (rank : 39) | θ value | θ > 10 (rank : 148) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9HAU0, Q8N3K6, Q96DY9, Q9BVR4, Q9C0H7, Q9H924, Q9NVK8 | Gene names | PLEKHA5, KIAA1686, PEPP2 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 5 (Phosphoinositol 3-phosphate-binding protein 2) (PEPP-2). | |||||
|
CNKR2_MOUSE
|
||||||
| NC score | 0.090928 (rank : 40) | θ value | θ > 10 (rank : 136) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q80YA9, Q80TP2 | Gene names | Cnksr2, Kiaa0902 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Connector enhancer of kinase suppressor of ras 2 (Connector enhancer of KSR2) (CNK2). | |||||
|
CNKR2_HUMAN
|
||||||
| NC score | 0.090848 (rank : 41) | θ value | θ > 10 (rank : 135) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8WXI2, O94976, Q8WXI1 | Gene names | CNKSR2, CNK2, KIAA0902, KSR2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Connector enhancer of kinase suppressor of ras 2 (Connector enhancer of KSR2) (CNK2). | |||||
|
PLEK2_MOUSE
|
||||||
| NC score | 0.090014 (rank : 42) | θ value | θ > 10 (rank : 152) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9WV52 | Gene names | Plek2 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin-2. | |||||
|
PLEK2_HUMAN
|
||||||
| NC score | 0.088483 (rank : 43) | θ value | θ > 10 (rank : 151) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NYT0, Q96JT0 | Gene names | PLEK2 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin-2. | |||||
|
ANLN_MOUSE
|
||||||
| NC score | 0.084721 (rank : 44) | θ value | θ > 10 (rank : 129) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 297 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8K298, Q8BL79, Q8BLB3, Q8K2N0, Q9CUY0 | Gene names | Anln | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Actin-binding protein anillin. | |||||
|
RASA1_HUMAN
|
||||||
| NC score | 0.069474 (rank : 45) | θ value | θ > 10 (rank : 155) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P20936 | Gene names | RASA1, RASA | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 1 (GTPase-activating protein) (GAP) (Ras p21 protein activator) (p120GAP) (RasGAP). | |||||
|
RHG25_MOUSE
|
||||||
| NC score | 0.068010 (rank : 46) | θ value | θ > 10 (rank : 157) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 492 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8BYW1, Q2VPR2, Q3TE04, Q3TVX0, Q3UVB7, Q6A0E0, Q8BX98 | Gene names | Arhgap25, Kiaa0053 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 25. | |||||
|
RHG25_HUMAN
|
||||||
| NC score | 0.065419 (rank : 47) | θ value | θ > 10 (rank : 156) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 550 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P42331, Q8IXQ2 | Gene names | ARHGAP25, KIAA0053 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 25. | |||||
|
FARP1_HUMAN
|
||||||
| NC score | 0.065112 (rank : 48) | θ value | 0.00390308 (rank : 23) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9Y4F1, Q6IQ29 | Gene names | FARP1, CDEP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FERM, RhoGEF and pleckstrin domain-containing protein 1 (Chondrocyte- derived ezrin-like protein). | |||||
|
SWP70_HUMAN
|
||||||
| NC score | 0.063582 (rank : 49) | θ value | θ > 10 (rank : 158) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 973 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9UH65, O75135, Q7LCY6, Q9P061, Q9P0Z8 | Gene names | SWAP70, KIAA0640 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Switch-associated protein 70 (SWAP-70). | |||||
|
CENB2_HUMAN
|
||||||
| NC score | 0.062082 (rank : 50) | θ value | θ > 10 (rank : 132) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q15057, Q9UQR3 | Gene names | CENTB2, KIAA0041 | |||
|
Domain Architecture |

|
|||||
| Description | Centaurin-beta 2 (Cnt-b2). | |||||
|
GNRP_HUMAN
|
||||||
| NC score | 0.061602 (rank : 51) | θ value | θ > 10 (rank : 139) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q13972, Q16027 | Gene names | RASGRF1, CDC25 | |||
|
Domain Architecture |

|
|||||
| Description | Guanine nucleotide-releasing protein (GNRP) (Ras-specific nucleotide exchange factor CDC25) (Ras-specific guanine nucleotide-releasing factor). | |||||
|
FGD4_HUMAN
|
||||||
| NC score | 0.060622 (rank : 52) | θ value | 0.0563607 (rank : 24) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q96M96, Q6ULS2, Q8TCP6 | Gene names | FGD4, FRABP, ZFYVE6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 4 (Actin filament- binding protein frabin) (FGD1-related F-actin-binding protein) (Zinc finger FYVE domain-containing protein 6). | |||||
|
GNRP_MOUSE
|
||||||
| NC score | 0.060216 (rank : 53) | θ value | θ > 10 (rank : 140) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P27671 | Gene names | Rasgrf1, Cdc25, Grf1 | |||
|
Domain Architecture |

|
|||||
| Description | Guanine nucleotide-releasing protein (GNRP) (Ras-specific nucleotide exchange factor CDC25) (CDC25Mm). | |||||
|
FGD4_MOUSE
|
||||||
| NC score | 0.059286 (rank : 54) | θ value | 0.0961366 (rank : 27) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q91ZT5, Q3UEB6, Q8BW60, Q8BZI7, Q91ZT3, Q91ZT4 | Gene names | Fgd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 4 (Actin filament- binding protein frabin) (FGD1-related F-actin-binding protein). | |||||
|
FGD6_HUMAN
|
||||||
| NC score | 0.058421 (rank : 55) | θ value | 0.365318 (rank : 35) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 237 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q6ZV73, Q6ZR53, Q7Z2Z7, Q96D44, Q9NUR8, Q9P2I5 | Gene names | FGD6, KIAA1362, ZFYVE24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 6 (Zinc finger FYVE domain-containing protein 24). | |||||
|
FGD6_MOUSE
|
||||||
| NC score | 0.057272 (rank : 56) | θ value | 0.365318 (rank : 36) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q69ZL1, Q8C8W5, Q8K3B0, Q9D3Y7 | Gene names | Fgd6, Kiaa1362 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 6. | |||||
|
SWP70_MOUSE
|
||||||
| NC score | 0.053657 (rank : 57) | θ value | θ > 10 (rank : 159) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1097 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6A028, O88443, Q3TQR6, Q3UCA3, Q6P1D0 | Gene names | Swap70, Kiaa0640 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Switch-associated protein 70 (SWAP-70). | |||||
|
CENB5_HUMAN
|
||||||
| NC score | 0.053594 (rank : 58) | θ value | θ > 10 (rank : 133) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q96P50, Q9BSR9, Q9C0E7 | Gene names | CENTB5, KIAA1716 | |||
|
Domain Architecture |

|
|||||
| Description | Centaurin-beta 5 (Cnt-b5). | |||||
|
BASP_MOUSE
|
||||||
| NC score | 0.053060 (rank : 59) | θ value | θ > 10 (rank : 131) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
BASP_HUMAN
|
||||||
| NC score | 0.051938 (rank : 60) | θ value | θ > 10 (rank : 130) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
FARP2_MOUSE
|
||||||
| NC score | 0.051687 (rank : 61) | θ value | 0.813845 (rank : 46) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q91VS8, Q3TAP2, Q69ZZ0 | Gene names | Farp2, Kiaa0793 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FERM, RhoGEF and pleckstrin domain-containing protein 2 (FERM domain including RhoGEF) (FIR). | |||||
|
MTMR5_HUMAN
|
||||||
| NC score | 0.050158 (rank : 62) | θ value | θ > 10 (rank : 141) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O95248, O60228, Q5JXD8, Q5PPM2, Q96GR9, Q9UGB8 | Gene names | SBF1, MTMR5 | |||
|
Domain Architecture |

|
|||||
| Description | SET-binding factor 1 (Sbf1) (Myotubularin-related protein 5). | |||||
|
ATX7_HUMAN
|
||||||
| NC score | 0.048896 (rank : 63) | θ value | 5.27518 (rank : 93) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O15265, O75328, O75329, Q9Y6P8 | Gene names | ATXN7, SCA7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-7 (Spinocerebellar ataxia type 7 protein). | |||||
|
FARP2_HUMAN
|
||||||
| NC score | 0.047405 (rank : 64) | θ value | 5.27518 (rank : 98) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O94887, Q53QM5, Q8WU27, Q9UFE7 | Gene names | FARP2, KIAA0793 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FERM, RhoGEF and pleckstrin domain-containing protein 2 (FERM domain including RhoGEF) (FIR). | |||||
|
PRRT3_HUMAN
|
||||||
| NC score | 0.045145 (rank : 65) | θ value | 0.47712 (rank : 41) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q5FWE3, Q49AD0, Q6UXY6, Q8NBC9 | Gene names | PRRT3, UNQ5823/PRO19642 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
CJ047_HUMAN
|
||||||
| NC score | 0.043387 (rank : 66) | θ value | 0.163984 (rank : 30) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q86WR7, Q5W0J9, Q5W0K0, Q5W0K1, Q5W0K2, Q6PJC8, Q8N317 | Gene names | C10orf47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf47. | |||||
|
OGFR_HUMAN
|
||||||
| NC score | 0.040974 (rank : 67) | θ value | 1.81305 (rank : 58) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
EP300_HUMAN
|
||||||
| NC score | 0.040919 (rank : 68) | θ value | 0.21417 (rank : 32) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q09472 | Gene names | EP300, P300 | |||
|
Domain Architecture |

|
|||||
| Description | E1A-associated protein p300 (EC 2.3.1.48). | |||||
|
BSN_MOUSE
|
||||||
| NC score | 0.037941 (rank : 69) | θ value | 6.88961 (rank : 103) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
CAC1G_HUMAN
|
||||||
| NC score | 0.037014 (rank : 70) | θ value | 0.0961366 (rank : 26) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 587 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O43497, O43498, O94770, Q9NYU4, Q9NYU5, Q9NYU6, Q9NYU7, Q9NYU8, Q9NYU9, Q9NYV0, Q9NYV1, Q9UHN9, Q9UHP0, Q9ULU6, Q9UNG7, Q9Y5T2, Q9Y5T3 | Gene names | CACNA1G, KIAA1123 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent T-type calcium channel subunit alpha-1G (Voltage- gated calcium channel subunit alpha Cav3.1) (Cav3.1c) (NBR13). | |||||
|
NSMA2_MOUSE
|
||||||
| NC score | 0.036437 (rank : 71) | θ value | 0.47712 (rank : 39) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9JJY3, Q3UTQ5 | Gene names | smpd3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sphingomyelin phosphodiesterase 3 (EC 3.1.4.12) (Neutral sphingomyelinase 2) (Neutral sphingomyelinase II) (nSMase2) (nSMase- 2). | |||||
|
BSN_HUMAN
|
||||||
| NC score | 0.033640 (rank : 72) | θ value | 1.81305 (rank : 56) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
HSF2_MOUSE
|
||||||
| NC score | 0.032904 (rank : 73) | θ value | 1.06291 (rank : 51) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P38533, Q64157, Q8VEJ0, Q9Z2L8 | Gene names | Hsf2 | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock factor protein 2 (HSF 2) (Heat shock transcription factor 2) (HSTF 2). | |||||
|
HSF2_HUMAN
|
||||||
| NC score | 0.032602 (rank : 74) | θ value | 1.06291 (rank : 50) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q03933, Q9H445 | Gene names | HSF2, HSTF2 | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock factor protein 2 (HSF 2) (Heat shock transcription factor 2) (HSTF 2). | |||||
|
MLL3_HUMAN
|
||||||
| NC score | 0.031658 (rank : 75) | θ value | 0.813845 (rank : 47) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
CI082_MOUSE
|
||||||
| NC score | 0.031572 (rank : 76) | θ value | 3.0926 (rank : 72) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8VDY9 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C9orf82 homolog. | |||||
|
RGS3_HUMAN
|
||||||
| NC score | 0.031564 (rank : 77) | θ value | 0.125558 (rank : 28) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
MUC1_HUMAN
|
||||||
| NC score | 0.031056 (rank : 78) | θ value | 1.81305 (rank : 57) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
TP53B_MOUSE
|
||||||
| NC score | 0.030998 (rank : 79) | θ value | 1.38821 (rank : 53) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 644 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P70399, Q68FD0, Q8CI97, Q91YC9 | Gene names | Tp53bp1, Trp53bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor suppressor p53-binding protein 1 (p53-binding protein 1) (p53BP1) (53BP1). | |||||
|
K1802_MOUSE
|
||||||
| NC score | 0.030822 (rank : 80) | θ value | 3.0926 (rank : 74) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
PAWR_HUMAN
|
||||||
| NC score | 0.029698 (rank : 81) | θ value | 2.36792 (rank : 66) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 252 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96IZ0, O75796, Q6FHY9, Q8N700 | Gene names | PAWR, PAR4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PRKC apoptosis WT1 regulator protein (Prostate apoptosis response 4 protein) (Par-4). | |||||
|
ASPX_HUMAN
|
||||||
| NC score | 0.029175 (rank : 82) | θ value | 1.06291 (rank : 49) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P26436 | Gene names | ACRV1 | |||
|
Domain Architecture |

|
|||||
| Description | Acrosomal protein SP-10 precursor (Acrosomal vesicle protein 1). | |||||
|
FA47A_HUMAN
|
||||||
| NC score | 0.029001 (rank : 83) | θ value | 5.27518 (rank : 97) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 451 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q5JRC9, Q8TAA0 | Gene names | FAM47A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM47A. | |||||
|
NFH_MOUSE
|
||||||
| NC score | 0.028283 (rank : 84) | θ value | 0.279714 (rank : 34) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
ERCC5_MOUSE
|
||||||
| NC score | 0.028112 (rank : 85) | θ value | 2.36792 (rank : 62) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P35689, Q61528, Q64248 | Gene names | Ercc5, Ercc-5, Xpg | |||
|
Domain Architecture |

|
|||||
| Description | DNA-repair protein complementing XP-G cells homolog (Xeroderma pigmentosum group G-complementing protein homolog) (DNA excision repair protein ERCC-5). | |||||
|
CI082_HUMAN
|
||||||
| NC score | 0.026512 (rank : 86) | θ value | 6.88961 (rank : 104) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H8G2, Q5VY32, Q6IPE6, Q96C59 | Gene names | C9orf82 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C9orf82. | |||||
|
TAF1C_MOUSE
|
||||||
| NC score | 0.025179 (rank : 87) | θ value | 1.81305 (rank : 59) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6PDZ2, P97359, Q3U3B6, Q8BN54 | Gene names | Taf1c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TATA box-binding protein-associated factor, RNA polymerase I, subunit C (TATA box-binding protein-associated factor 1C) (TBP-associated factor 1C) (TBP-associated factor RNA polymerase I 95 kDa) (TAFI95). | |||||
|
ARHGA_HUMAN
|
||||||
| NC score | 0.024138 (rank : 88) | θ value | 4.03905 (rank : 77) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O15013, O14665, Q68D55, Q8IWD9, Q8IY77 | Gene names | ARHGEF10, KIAA0294 | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 10. | |||||
|
AKAP8_HUMAN
|
||||||
| NC score | 0.023609 (rank : 89) | θ value | 0.47712 (rank : 38) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O43823 | Gene names | AKAP8, AKAP95 | |||
|
Domain Architecture |

|
|||||
| Description | A-kinase anchor protein 8 (A-kinase anchor protein 95 kDa) (AKAP 95). | |||||
|
SAPS1_HUMAN
|
||||||
| NC score | 0.023565 (rank : 90) | θ value | 4.03905 (rank : 88) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9UPN7, Q504V2, Q6NVJ6, Q9BU97 | Gene names | SAPS1, KIAA1115 | |||
|
Domain Architecture |

|
|||||
| Description | SAPS domain family member 1. | |||||
|
CLIC6_HUMAN
|
||||||
| NC score | 0.023463 (rank : 91) | θ value | 2.36792 (rank : 61) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q96NY7, Q8IX31 | Gene names | CLIC6, CLIC1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride intracellular channel 6. | |||||
|
RFFL_HUMAN
|
||||||
| NC score | 0.023313 (rank : 92) | θ value | 0.21417 (rank : 33) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8WZ73, Q8NHW0, Q8TBY7, Q96BE6 | Gene names | RFFL, RNF189, RNF34L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rififylin (RING finger and FYVE-like domain-containing protein 1) (FYVE-RING finger protein Sakura) (Fring) (Caspases-8 and -10- associated RING finger protein 2) (CARP-2) (Caspase regulator CARP2) (RING finger protein 189) (RING finger protein 34-like). | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.023113 (rank : 93) | θ value | 6.88961 (rank : 113) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
MAGE1_HUMAN
|
||||||
| NC score | 0.022446 (rank : 94) | θ value | 8.99809 (rank : 122) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
CD2_HUMAN
|
||||||
| NC score | 0.022235 (rank : 95) | θ value | 3.0926 (rank : 71) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P06729, Q96TE5 | Gene names | CD2 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell surface antigen CD2 precursor (T-cell surface antigen T11/Leu- 5) (LFA-2) (LFA-3 receptor) (Erythrocyte receptor) (Rosette receptor). | |||||
|
TROAP_HUMAN
|
||||||
| NC score | 0.021931 (rank : 96) | θ value | 4.03905 (rank : 90) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q12815 | Gene names | TROAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trophinin-associated protein (Tastin) (Trophinin-assisting protein). | |||||
|
OXR1_MOUSE
|
||||||
| NC score | 0.020713 (rank : 97) | θ value | 0.47712 (rank : 40) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q4KMM3, Q5FWW1, Q99L06, Q99MK1, Q99MP4 | Gene names | Oxr1, C7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oxidation resistance protein 1 (Protein C7). | |||||
|
RIMB1_HUMAN
|
||||||
| NC score | 0.020689 (rank : 98) | θ value | 2.36792 (rank : 69) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1479 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O95153, O75111, Q8N5W3 | Gene names | BZRAP1, KIAA0612, RBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Peripheral-type benzodiazepine receptor-associated protein 1 (PRAX-1) (Peripheral benzodiazepine receptor-interacting protein) (PBR-IP) (RIM-binding protein 1) (RIM-BP1). | |||||
|
CS021_HUMAN
|
||||||
| NC score | 0.019384 (rank : 99) | θ value | 4.03905 (rank : 78) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8IVT2 | Gene names | C19orf21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C19orf21. | |||||
|
SFR14_MOUSE
|
||||||
| NC score | 0.019106 (rank : 100) | θ value | 4.03905 (rank : 89) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8CH09, Q6PG19, Q80UY8, Q8BY32, Q8CFM0 | Gene names | Sfrs14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative splicing factor, arginine/serine-rich 14 (Arginine/serine- rich-splicing factor 14). | |||||
|
GPTC8_HUMAN
|
||||||
| NC score | 0.018948 (rank : 101) | θ value | 4.03905 (rank : 82) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 366 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9UKJ3, O60300 | Gene names | GPATC8, KIAA0553 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 8. | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.018636 (rank : 102) | θ value | 5.27518 (rank : 99) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
DOK2_MOUSE
|
||||||
| NC score | 0.018636 (rank : 103) | θ value | 0.813845 (rank : 45) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 244 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O70469, O70272, Q99KL1 | Gene names | Dok2, Frip | |||
|
Domain Architecture |

|
|||||
| Description | Docking protein 2 (Downstream of tyrosine kinase 2) (p56(dok-2)) (Dok- related protein) (Dok-R) (IL-four receptor-interacting protein) (FRIP). | |||||
|
ATS13_HUMAN
|
||||||
| NC score | 0.017901 (rank : 104) | θ value | 0.0961366 (rank : 25) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q76LX8, Q6UY16, Q710F6, Q711T8, Q96L37, Q9H0G3 | Gene names | ADAMTS13, C9orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAMTS-13 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase with thrombospondin motifs 13) (ADAM-TS 13) (ADAM-TS13) (von Willebrand factor-cleaving protease) (vWF-cleaving protease) (vWF-CP). | |||||
|
PTN23_HUMAN
|
||||||
| NC score | 0.017564 (rank : 105) | θ value | 2.36792 (rank : 68) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 646 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9H3S7, Q7KZF8, Q8N6Z5, Q9BSR5, Q9P257, Q9UG03, Q9UMZ4 | Gene names | PTPN23, KIAA1471 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 23 (EC 3.1.3.48) (His- domain-containing protein tyrosine phosphatase) (HD-PTP) (Protein tyrosine phosphatase TD14) (PTP-TD14). | |||||
|
ASPM_MOUSE
|
||||||
| NC score | 0.017552 (rank : 106) | θ value | 1.81305 (rank : 55) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CJ27, O88482, Q8BJI8, Q8BKT4 | Gene names | Aspm, Calmbp1, Sha1 | |||
|
Domain Architecture |

|
|||||
| Description | Abnormal spindle-like microcephaly-associated protein homolog (Calmodulin-binding protein 1) (Spindle and hydroxyurea checkpoint abnormal protein) (Calmodulin-binding protein Sha1). | |||||
|
NALDL_HUMAN
|
||||||
| NC score | 0.017173 (rank : 107) | θ value | 2.36792 (rank : 65) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UQQ1, O43176 | Gene names | NAALADL1, NAALADASEL, NAALADL | |||
|
Domain Architecture |

|
|||||
| Description | N-acetylated-alpha-linked acidic dipeptidase-like protein (EC 3.4.17.21) (NAALADase L) (Ileal dipeptidylpeptidase) (100 kDa ileum brush border membrane protein) (I100). | |||||
|
CS021_MOUSE
|
||||||
| NC score | 0.017048 (rank : 108) | θ value | 6.88961 (rank : 105) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9D279, Q3UI04 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C19orf21 homolog. | |||||
|
MDC1_MOUSE
|
||||||
| NC score | 0.017036 (rank : 109) | θ value | 3.0926 (rank : 75) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
MAP1A_HUMAN
|
||||||
| NC score | 0.016691 (rank : 110) | θ value | 4.03905 (rank : 85) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P78559, O95643, Q12973, Q15882, Q9UJT4 | Gene names | MAP1A | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 1A (MAP 1A) (Proliferation-related protein p80) [Contains: MAP1 light chain LC2]. | |||||
|
PKD1_HUMAN
|
||||||
| NC score | 0.016195 (rank : 111) | θ value | 2.36792 (rank : 67) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P98161, Q15140, Q15141 | Gene names | PKD1 | |||
|
Domain Architecture |

|
|||||
| Description | Polycystin-1 precursor (Autosomal dominant polycystic kidney disease protein 1). | |||||
|
CIC_HUMAN
|
||||||
| NC score | 0.015784 (rank : 112) | θ value | 8.99809 (rank : 120) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 992 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q96RK0, Q7LGI1, Q9UEG5, Q9Y6T1 | Gene names | CIC, KIAA0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
FOXN1_HUMAN
|
||||||
| NC score | 0.015651 (rank : 113) | θ value | 4.03905 (rank : 80) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 377 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O15353, O15352 | Gene names | FOXN1, WHN | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein N1 (Transcription factor winged-helix nude). | |||||
|
FOXP4_MOUSE
|
||||||
| NC score | 0.015400 (rank : 114) | θ value | 0.62314 (rank : 42) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9DBY0, Q80V92, Q8CG10, Q8CIS1 | Gene names | Foxp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein P4 (Fork head-related protein like A) (mFKHLA). | |||||
|
NALDL_MOUSE
|
||||||
| NC score | 0.015358 (rank : 115) | θ value | 4.03905 (rank : 86) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7M758 | Gene names | Naaladl1, Naaladasel | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylated-alpha-linked acidic dipeptidase-like protein (EC 3.4.17.21) (NAALADase L). | |||||
|
PGCA_MOUSE
|
||||||
| NC score | 0.015327 (rank : 116) | θ value | 0.163984 (rank : 31) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 643 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q61282, Q64021 | Gene names | Agc1, Agc | |||
|
Domain Architecture |

|
|||||
| Description | Aggrecan core protein precursor (Cartilage-specific proteoglycan core protein) (CSPCP). | |||||
|
MSH3_HUMAN
|
||||||
| NC score | 0.015302 (rank : 117) | θ value | 6.88961 (rank : 108) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P20585, Q6PJT5, Q86UQ6, Q92867 | Gene names | MSH3, DUC1, DUG | |||
|
Domain Architecture |

|
|||||
| Description | DNA mismatch repair protein Msh3 (Divergent upstream protein) (DUP) (Mismatch repair protein 1) (MRP1). | |||||
|
LPP_HUMAN
|
||||||
| NC score | 0.015212 (rank : 118) | θ value | 4.03905 (rank : 84) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 775 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q93052, Q8NFX5 | Gene names | LPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipoma-preferred partner (LIM domain-containing preferred translocation partner in lipoma). | |||||
|
NPBL_MOUSE
|
||||||
| NC score | 0.013355 (rank : 119) | θ value | 6.88961 (rank : 110) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
GON4L_MOUSE
|
||||||
| NC score | 0.013085 (rank : 120) | θ value | 6.88961 (rank : 107) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9DB00, Q80TB4, Q91YI9 | Gene names | Gon4l, Gon4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
DPOG1_MOUSE
|
||||||
| NC score | 0.012685 (rank : 121) | θ value | 4.03905 (rank : 79) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P54099, Q9JI28 | Gene names | Polg, Polg1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase subunit gamma 1 (EC 2.7.7.7) (Mitochondrial DNA polymerase catalytic subunit) (PolG-alpha). | |||||
|
ARD1H_HUMAN
|
||||||
| NC score | 0.012605 (rank : 122) | θ value | 5.27518 (rank : 92) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P41227 | Gene names | ARD1A, ARD1, TE2 | |||
|
Domain Architecture |

|
|||||
| Description | N-terminal acetyltransferase complex ARD1 subunit homolog A (EC 2.3.1.88) (EC 2.3.1.-). | |||||
|
CHSS2_MOUSE
|
||||||
| NC score | 0.012535 (rank : 123) | θ value | 2.36792 (rank : 60) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6IQX7 | Gene names | Css2, D1Bwg1363e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate synthase 2 (EC 2.4.1.175) (Glucuronosyl-N- acetylgalactosaminyl-proteoglycan 4-beta-N- acetylgalactosaminyltransferase II) (N-acetylgalactosaminyl- proteoglycan 3-beta-glucuronosyltransferase II) (EC 2.4.1.226) (Chondroitin glucuronyltransferase II) (N- acetylgalactosaminyltransferase). | |||||
|
AXN2_MOUSE
|
||||||
| NC score | 0.012419 (rank : 124) | θ value | 5.27518 (rank : 94) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O88566, Q9QXJ6 | Gene names | Axin2 | |||
|
Domain Architecture |

|
|||||
| Description | Axin-2 (Axis inhibition protein 2) (Conductin) (Axin-like protein) (Axil). | |||||
|
BCAS1_MOUSE
|
||||||
| NC score | 0.012325 (rank : 125) | θ value | 8.99809 (rank : 118) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q80YN3, Q9CVA1 | Gene names | Bcas1, Nabc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Breast carcinoma amplified sequence 1 homolog (Novel amplified in breast cancer 1 homolog). | |||||
|
DDX51_HUMAN
|
||||||
| NC score | 0.012279 (rank : 126) | θ value | 0.813845 (rank : 44) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8N8A6, Q5CZ71, Q8IXK5, Q96ED1 | Gene names | DDX51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX51 (EC 3.6.1.-) (DEAD box protein 51). | |||||
|
ZEP1_HUMAN
|
||||||
| NC score | 0.012204 (rank : 127) | θ value | 0.365318 (rank : 37) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1036 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P15822, Q14122 | Gene names | HIVEP1, ZNF40 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 40 (Human immunodeficiency virus type I enhancer- binding protein 1) (HIV-EP1) (Major histocompatibility complex-binding protein 1) (MBP-1) (Positive regulatory domain II-binding factor 1) (PRDII-BF1). | |||||
|
MUS81_MOUSE
|
||||||
| NC score | 0.012112 (rank : 128) | θ value | 5.27518 (rank : 100) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q91ZJ0, Q8R356 | Gene names | Mus81 | |||
|
Domain Architecture |

|
|||||
| Description | Crossover junction endonuclease MUS81 (EC 3.1.22.-). | |||||
|
WDR81_MOUSE
|
||||||
| NC score | 0.011713 (rank : 129) | θ value | 2.36792 (rank : 70) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5ND34, Q3TA90, Q3TUS9, Q6KAR1 | Gene names | Wdr81 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 81. | |||||
|
F120A_HUMAN
|
||||||
| NC score | 0.010602 (rank : 130) | θ value | 5.27518 (rank : 96) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NZB2, O60649, Q14688, Q4VXF4, Q4VXF5, Q4VXG2, Q96I21, Q9NZB1 | Gene names | FAM120A, C9orf10, KIAA0183 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UPF0318 protein FAM120A. | |||||
|
OXRP_HUMAN
|
||||||
| NC score | 0.009845 (rank : 131) | θ value | 6.88961 (rank : 111) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 544 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9Y4L1 | Gene names | HYOU1, ORP150 | |||
|
Domain Architecture |

|
|||||
| Description | 150 kDa oxygen-regulated protein precursor (Orp150) (Hypoxia up- regulated 1). | |||||
|
CA156_HUMAN
|
||||||
| NC score | 0.009803 (rank : 132) | θ value | 8.99809 (rank : 119) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O95568 | Gene names | C1orf156, ASTP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C1orf156 (Arsenic-transactivated protein 2) (AsTP2). | |||||
|
DEN4C_HUMAN
|
||||||
| NC score | 0.009641 (rank : 133) | θ value | 5.27518 (rank : 95) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5VZ89, Q6AI48, Q6ZUB3, Q8NCY7, Q9H6N4, Q9NUT1, Q9NWA5 | Gene names | DENND4C, C9orf55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 4C. | |||||
|
ZEP1_MOUSE
|
||||||
| NC score | 0.009298 (rank : 134) | θ value | 4.03905 (rank : 91) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1063 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q03172 | Gene names | Hivep1, Cryabp1, Znf40 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 40 (Transcription factor alphaA-CRYBP1) (Alpha A- crystallin-binding protein I) (Alpha A-CRYBP1). | |||||
|
PDZD2_HUMAN
|
||||||
| NC score | 0.009161 (rank : 135) | θ value | 5.27518 (rank : 101) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O15018, Q9BXD4 | Gene names | PDZD2, AIPC, KIAA0300, PDZK3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 2 (PDZ domain-containing protein 3) (Activated in prostate cancer protein). | |||||
|
SIRT6_HUMAN
|
||||||
| NC score | 0.008943 (rank : 136) | θ value | 6.88961 (rank : 112) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N6T7, O75291, Q6IAF5, Q6PK99, Q8NCD2, Q9BSI5, Q9BWP3, Q9NRC7, Q9UQD1 | Gene names | SIRT6, SIR2L6 | |||
|
Domain Architecture |

|
|||||
| Description | Mono-ADP-ribosyltransferase sirtuin-6 (EC 2.4.2.31) (SIR2-like protein 6). | |||||
|
CUTL1_MOUSE
|
||||||
| NC score | 0.008142 (rank : 137) | θ value | 6.88961 (rank : 106) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P53564, O08994, P70301, Q571L6, Q91ZD2 | Gene names | Cutl1, Cux, Cux1, Kiaa4047 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP) (Homeobox protein Cux). | |||||
|
DHX16_HUMAN
|
||||||
| NC score | 0.008030 (rank : 138) | θ value | 1.38821 (rank : 52) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O60231, O60322, Q969X7, Q96QC1 | Gene names | DHX16, DBP2, DDX16, KIAA0577 | |||
|
Domain Architecture |

|
|||||
| Description | Putative pre-mRNA-splicing factor ATP-dependent RNA helicase DHX16 (EC 3.6.1.-) (DEAH-box protein 16) (ATP-dependent RNA helicase #3). | |||||
|
AFAD_HUMAN
|
||||||
| NC score | 0.007889 (rank : 139) | θ value | 8.99809 (rank : 116) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 689 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P55196, O75087, O75088, O75089, Q5TIG6, Q9NSN7, Q9NU92 | Gene names | MLLT4, AF6 | |||
|
Domain Architecture |

|
|||||
| Description | Afadin (Protein AF-6). | |||||
|
HIP1R_MOUSE
|
||||||
| NC score | 0.007416 (rank : 140) | θ value | 3.0926 (rank : 73) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 959 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9JKY5 | Gene names | Hip1r | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin-interacting protein 1-related protein (Hip1-related). | |||||
|
MY18B_HUMAN
|
||||||
| NC score | 0.006467 (rank : 141) | θ value | 6.88961 (rank : 109) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1299 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8IUG5, Q8NDI8, Q8TE65, Q8WWS0, Q96KH2, Q96KR8, Q96KR9 | Gene names | MYO18B | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18B (Myosin XVIIIb). | |||||
|
AN30A_HUMAN
|
||||||
| NC score | 0.006317 (rank : 142) | θ value | 1.81305 (rank : 54) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1119 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9BXX3, Q5W025 | Gene names | ANKRD30A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 30A (NY-BR-1 antigen). | |||||
|
GCC2_MOUSE
|
||||||
| NC score | 0.005841 (rank : 143) | θ value | 0.62314 (rank : 43) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1619 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8CHG3, Q8BR44, Q8R2Q5, Q9CT45 | Gene names | Gcc2, Kiaa0336 | |||
|
Domain Architecture |

|
|||||
| Description | GRIP and coiled-coil domain-containing protein 2 (Golgi coiled coil protein GCC185). | |||||
|
RB15B_HUMAN
|
||||||
| NC score | 0.005829 (rank : 144) | θ value | 8.99809 (rank : 124) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8NDT2, Q6QE19, Q9BV96 | Gene names | RBM15B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative RNA-binding protein 15B (RNA-binding motif protein 15B). | |||||
|
SALL2_HUMAN
|
||||||
| NC score | 0.005199 (rank : 145) | θ value | 4.03905 (rank : 87) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 835 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9Y467, Q9Y4G1 | Gene names | SALL2, KIAA0360, SAL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sal-like protein 2 (Zinc finger protein SALL2) (HSal2). | |||||
|
SPEG_MOUSE
|
||||||
| NC score | 0.004918 (rank : 146) | θ value | 0.813845 (rank : 48) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
FZD6_MOUSE
|
||||||
| NC score | 0.004667 (rank : 147) | θ value | 4.03905 (rank : 81) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q61089 | Gene names | Fzd6 | |||
|
Domain Architecture |

|
|||||
| Description | Frizzled-6 precursor (Fz-6) (mFz6). | |||||
|
RBP2_HUMAN
|
||||||
| NC score | 0.004531 (rank : 148) | θ value | 5.27518 (rank : 102) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P49792, Q13074, Q15280, Q53TE2, Q59FH7 | Gene names | RANBP2, NUP358 | |||
|
Domain Architecture |

|
|||||
| Description | Ran-binding protein 2 (RanBP2) (Nuclear pore complex protein Nup358) (Nucleoporin Nup358) (358 kDa nucleoporin) (P270). | |||||
|
F10A1_HUMAN
|
||||||
| NC score | 0.004386 (rank : 149) | θ value | 8.99809 (rank : 121) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P50502, O14999 | Gene names | ST13, FAM10A1, HIP, SNC6 | |||
|
Domain Architecture |

|
|||||
| Description | Hsc70-interacting protein (Hip) (Suppression of tumorigenicity protein 13) (Putative tumor suppressor ST13) (Protein FAM10A1) (Progesterone receptor-associated p48 protein) (NY-REN-33 antigen). | |||||
|
IRAK2_HUMAN
|
||||||
| NC score | 0.003212 (rank : 150) | θ value | 2.36792 (rank : 63) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 714 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O43187 | Gene names | IRAK2 | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin-1 receptor-associated kinase-like 2 (IRAK-2). | |||||
|
ZN142_HUMAN
|
||||||
| NC score | 0.003140 (rank : 151) | θ value | 0.125558 (rank : 29) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1007 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P52746, Q92510 | Gene names | ZNF142, KIAA0236 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 142 (HA4654). | |||||
|
ZN592_HUMAN
|
||||||
| NC score | 0.002465 (rank : 152) | θ value | 3.0926 (rank : 76) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 762 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q92610 | Gene names | ZNF592, KIAA0211 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 592. | |||||
|
ANKS3_HUMAN
|
||||||
| NC score | 0.001026 (rank : 153) | θ value | 8.99809 (rank : 117) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 355 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6ZW76, Q8TF25 | Gene names | ANKS3, KIAA1977 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat and SAM domain-containing protein 3. | |||||
|
MYO3A_HUMAN
|
||||||
| NC score | -0.000209 (rank : 154) | θ value | 2.36792 (rank : 64) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1033 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8NEV4, Q8WX17, Q9NYS8 | Gene names | MYO3A | |||
|
Domain Architecture |

|
|||||
| Description | Myosin IIIA (EC 2.7.11.1). | |||||
|
KLF15_HUMAN
|
||||||
| NC score | -0.000392 (rank : 155) | θ value | 4.03905 (rank : 83) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 720 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UIH9 | Gene names | KLF15, KKLF | |||
|
Domain Architecture |

|
|||||
| Description | Krueppel-like factor 15 (Kidney-enriched krueppel-like factor). | |||||
|
ZBT37_HUMAN
|
||||||
| NC score | -0.000691 (rank : 156) | θ value | 6.88961 (rank : 114) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 815 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5TC79, Q5TC80, Q96M87, Q9BQ88 | Gene names | ZBTB37 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger and BTB domain-containing protein 37. | |||||
|
ZBT16_HUMAN
|
||||||
| NC score | -0.001913 (rank : 157) | θ value | 8.99809 (rank : 125) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1010 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q05516, Q8TAL4 | Gene names | ZBTB16, PLZF, ZNF145 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 16 (Zinc finger protein PLZF) (Promyelocytic leukemia zinc finger protein) (Zinc finger protein 145). | |||||
|
ZN668_HUMAN
|
||||||
| NC score | -0.001921 (rank : 158) | θ value | 6.88961 (rank : 115) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 761 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q96K58, Q59EV1, Q8N669, Q9H8L4 | Gene names | ZNF668 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 668. | |||||
|
PAK4_MOUSE
|
||||||
| NC score | -0.002390 (rank : 159) | θ value | 8.99809 (rank : 123) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1098 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8BTW9, Q6ZPX0, Q80Z97, Q9CS71 | Gene names | Pak4, Kiaa1142 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase PAK 4 (EC 2.7.11.1) (p21-activated kinase 4) (PAK-4). | |||||