Please be patient as the page loads
|
GBF1_HUMAN
|
||||||
| SwissProt Accessions | Q92538, Q5VXX3, Q96CK6, Q96HZ3, Q9H473 | Gene names | GBF1, KIAA0248 | |||
|
Domain Architecture |

|
|||||
| Description | Golgi-specific brefeldin A-resistance guanine nucleotide exchange factor 1 (BFA-resistant GEF 1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
GBF1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 142 | |
| SwissProt Accessions | Q92538, Q5VXX3, Q96CK6, Q96HZ3, Q9H473 | Gene names | GBF1, KIAA0248 | |||
|
Domain Architecture |

|
|||||
| Description | Golgi-specific brefeldin A-resistance guanine nucleotide exchange factor 1 (BFA-resistant GEF 1). | |||||
|
BIG2_HUMAN
|
||||||
| θ value | 4.10295e-61 (rank : 2) | NC score | 0.868836 (rank : 3) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9Y6D5, Q5TFT9, Q9NTS1 | Gene names | ARFGEF2, ARFGEP2, BIG2 | |||
|
Domain Architecture |

|
|||||
| Description | Brefeldin A-inhibited guanine nucleotide-exchange protein 2 (Brefeldin A-inhibited GEP 2). | |||||
|
BIG1_HUMAN
|
||||||
| θ value | 3.59435e-57 (rank : 3) | NC score | 0.869742 (rank : 2) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9Y6D6, Q9NV46, Q9UFV2, Q9UNL0 | Gene names | ARFGEF1, ARFGEP1, BIG1 | |||
|
Domain Architecture |

|
|||||
| Description | Brefeldin A-inhibited guanine nucleotide-exchange protein 1 (Brefeldin A-inhibited GEP 1) (p200 ARF-GEP1) (p200 ARF guanine nucleotide exchange factor). | |||||
|
CYH3_HUMAN
|
||||||
| θ value | 9.20422e-37 (rank : 4) | NC score | 0.813035 (rank : 9) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O43739 | Gene names | PSCD3, ARNO3, GRP1 | |||
|
Domain Architecture |

|
|||||
| Description | Cytohesin-3 (ARF nucleotide-binding site opener 3) (ARNO3 protein) (General receptor of phosphoinositides 1) (Grp1). | |||||
|
CYH3_MOUSE
|
||||||
| θ value | 9.20422e-37 (rank : 5) | NC score | 0.807238 (rank : 15) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O08967, Q8CI93 | Gene names | Pscd3, Grp1 | |||
|
Domain Architecture |

|
|||||
| Description | Cytohesin-3 (ARF nucleotide-binding site opener 3) (ARNO3 protein) (General receptor of phosphoinositides 1) (Grp1) (CLM-3). | |||||
|
CYH1_HUMAN
|
||||||
| θ value | 7.79185e-36 (rank : 6) | NC score | 0.808090 (rank : 14) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q15438, Q9P123, Q9P124 | Gene names | PSCD1, D17S811E | |||
|
Domain Architecture |

|
|||||
| Description | Cytohesin-1 (SEC7 homolog B2-1). | |||||
|
CYH1_MOUSE
|
||||||
| θ value | 7.79185e-36 (rank : 7) | NC score | 0.810310 (rank : 12) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9QX11, O88817 | Gene names | Pscd1 | |||
|
Domain Architecture |
|
|||||
| Description | Cytohesin-1 (CLM1). | |||||
|
CYH2_MOUSE
|
||||||
| θ value | 6.59618e-35 (rank : 8) | NC score | 0.813468 (rank : 8) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P63034, O89099, P97695, Q3UP39 | Gene names | Pscd2, Sec7b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytohesin-2 (CLM2) (SEC7 homolog B) (msec7-2). | |||||
|
CYH2_HUMAN
|
||||||
| θ value | 2.50655e-34 (rank : 9) | NC score | 0.812975 (rank : 10) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q99418, Q92958 | Gene names | PSCD2, ARNO | |||
|
Domain Architecture |
|
|||||
| Description | Cytohesin-2 (ARF nucleotide-binding site opener) (ARNO protein) (ARF exchange factor). | |||||
|
CYH4_MOUSE
|
||||||
| θ value | 1.6247e-33 (rank : 10) | NC score | 0.810636 (rank : 11) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q80YW0 | Gene names | Pscd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytohesin-4. | |||||
|
CYH4_HUMAN
|
||||||
| θ value | 4.72714e-33 (rank : 11) | NC score | 0.810015 (rank : 13) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UIA0, Q5R3F9, Q9UGT6 | Gene names | PSCD4, CYT4 | |||
|
Domain Architecture |

|
|||||
| Description | Cytohesin-4. | |||||
|
IQEC1_HUMAN
|
||||||
| θ value | 5.22648e-32 (rank : 12) | NC score | 0.818949 (rank : 7) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6DN90, O94863, Q96D85 | Gene names | IQSEC1, ARFGEP100, BRAG2, KIAA0763 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 1 (ADP-ribosylation factors guanine nucleotide-exchange protein 100) (ADP-ribosylation factors guanine nucleotide-exchange protein 2) (Brefeldin-resistant Arf-GEF 2 protein). | |||||
|
IQEC1_MOUSE
|
||||||
| θ value | 5.22648e-32 (rank : 13) | NC score | 0.821060 (rank : 5) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8R0S2, Q3TZC3, Q5DU15 | Gene names | Iqsec1, Kiaa0763 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 1. | |||||
|
IQEC2_MOUSE
|
||||||
| θ value | 4.42448e-31 (rank : 14) | NC score | 0.782323 (rank : 16) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q5DU25 | Gene names | Iqsec2, Kiaa0522 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 2. | |||||
|
IQEC2_HUMAN
|
||||||
| θ value | 9.8567e-31 (rank : 15) | NC score | 0.781531 (rank : 17) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q5JU85, O60275 | Gene names | IQSEC2, KIAA0522 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 2. | |||||
|
IQEC3_HUMAN
|
||||||
| θ value | 1.85889e-29 (rank : 16) | NC score | 0.824317 (rank : 4) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9UPP2, Q8TB43 | Gene names | IQSEC3, KIAA1110 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 3. | |||||
|
IQEC3_MOUSE
|
||||||
| θ value | 1.85889e-29 (rank : 17) | NC score | 0.819649 (rank : 6) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q3TES0, Q3UHP8, Q80TJ8 | Gene names | Iqsec3, Kiaa1110 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 3. | |||||
|
PSD3_HUMAN
|
||||||
| θ value | 1.5773e-20 (rank : 18) | NC score | 0.755649 (rank : 18) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9NYI0, Q9Y2F1 | Gene names | PSD3, EFA6R, HCA67, KIAA0942 | |||
|
Domain Architecture |

|
|||||
| Description | PH and SEC7 domain-containing protein 3 (Pleckstrin homology and SEC7 domain-containing protein 3) (Exchange factor for ADP-ribosylation factor guanine nucleotide factor 6) (Hepatocellular carcinoma- associated antigen 67). | |||||
|
PSD4_MOUSE
|
||||||
| θ value | 3.28887e-18 (rank : 19) | NC score | 0.720633 (rank : 19) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BLR5, Q3TE33, Q3TQF5, Q3UD59, Q3UFH9, Q80V44 | Gene names | Psd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH and SEC7 domain-containing protein 4 (Pleckstrin homology and SEC7 domain-containing protein 4). | |||||
|
PSD4_HUMAN
|
||||||
| θ value | 3.63628e-17 (rank : 20) | NC score | 0.714305 (rank : 21) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8NDX1, O95621, Q4ZG34, Q6GPH8, Q8IYP4 | Gene names | PSD4, EFA6B, TIC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH and SEC7 domain-containing protein 4 (Pleckstrin homology and SEC7 domain-containing protein 4) (Exchange factor for ADP-ribosylation factor guanine nucleotide factor 6B) (Telomeric of interleukin-1 cluster protein). | |||||
|
FBX8_MOUSE
|
||||||
| θ value | 5.26297e-08 (rank : 21) | NC score | 0.715186 (rank : 20) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9QZN3 | Gene names | Fbxo8, Fbx8 | |||
|
Domain Architecture |

|
|||||
| Description | F-box only protein 8. | |||||
|
FBX8_HUMAN
|
||||||
| θ value | 8.97725e-08 (rank : 22) | NC score | 0.712496 (rank : 22) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NRD0, Q6UWN4, Q9NRP5, Q9UKC4 | Gene names | FBXO8, FBS, FBX8 | |||
|
Domain Architecture |
|
|||||
| Description | F-box only protein 8 (F-box/SEC7 protein FBS). | |||||
|
FOG1_MOUSE
|
||||||
| θ value | 0.00298849 (rank : 23) | NC score | 0.026538 (rank : 100) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1237 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O35615 | Gene names | Zfpm1, Fog, Fog1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein ZFPM1 (Zinc finger protein multitype 1) (Friend of GATA protein 1) (Friend of GATA-1) (FOG-1). | |||||
|
OGFR_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 24) | NC score | 0.054628 (rank : 60) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
CUTL1_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 25) | NC score | 0.023492 (rank : 108) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P53564, O08994, P70301, Q571L6, Q91ZD2 | Gene names | Cutl1, Cux, Cux1, Kiaa4047 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP) (Homeobox protein Cux). | |||||
|
B4GT1_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 26) | NC score | 0.026154 (rank : 101) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P15291, Q12909, Q12910, Q12911, Q14456, Q14509, Q14523 | Gene names | B4GALT1, GGTB2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-1,4-galactosyltransferase 1 (EC 2.4.1.-) (Beta-1,4-GalTase 1) (Beta4Gal-T1) (b4Gal-T1) (UDP-galactose:beta-N-acetylglucosamine beta- 1,4-galactosyltransferase 1) (UDP-Gal:beta-GlcNAc beta-1,4- galactosyltransferase 1) [Includes: Lactose synthase A protein (EC 2.4.1.22); N-acetyllactosamine synthase (EC 2.4.1.90) (Nal synthetase); Beta-N-acetylglucosaminylglycopeptide beta-1,4- galactosyltransferase (EC 2.4.1.38); Beta-N-acetylglucosaminyl- glycolipid beta-1,4-galactosyltransferase (EC 2.4.1.-)]. | |||||
|
SON_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 27) | NC score | 0.040095 (rank : 75) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
RGS3_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 28) | NC score | 0.040852 (rank : 73) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
ZBT17_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 29) | NC score | 0.005203 (rank : 172) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 940 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q60821, Q60699 | Gene names | Zbtb17, Zfp100, Znf151 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger and BTB domain-containing protein 17 (Zinc finger protein 151) (Zinc finger protein 100) (Zfp-100) (Polyomavirus late initiator promoter-binding protein) (LP-1) (Zinc finger protein Z13). | |||||
|
OSBL2_HUMAN
|
||||||
| θ value | 0.125558 (rank : 30) | NC score | 0.022951 (rank : 113) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H1P3, Q9BZB1, Q9Y4B8 | Gene names | OSBPL2, KIAA0772, ORP2 | |||
|
Domain Architecture |
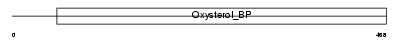
|
|||||
| Description | Oxysterol-binding protein-related protein 2 (OSBP-related protein 2) (ORP-2). | |||||
|
TSH3_MOUSE
|
||||||
| θ value | 0.125558 (rank : 31) | NC score | 0.026822 (rank : 98) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 590 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8CGV9, Q5DTX4 | Gene names | Tshz3, Kiaa1474, Tsh3, Zfp537, Znf537 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Teashirt homolog 3 (Zinc finger protein 537). | |||||
|
CSKI1_MOUSE
|
||||||
| θ value | 0.163984 (rank : 32) | NC score | 0.022954 (rank : 112) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1031 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q6P9K8, Q6ZPU2, Q8BWU2, Q8BX99, Q9CXH0 | Gene names | Caskin1, Kiaa1306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-1 (CASK-interacting protein 1). | |||||
|
LRCH2_MOUSE
|
||||||
| θ value | 0.163984 (rank : 33) | NC score | 0.018317 (rank : 122) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q3UMG5, Q7TNP9, Q80TC9 | Gene names | Lrch2, Kiaa1495 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 2. | |||||
|
MAGE1_HUMAN
|
||||||
| θ value | 0.163984 (rank : 34) | NC score | 0.036114 (rank : 79) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
CD248_HUMAN
|
||||||
| θ value | 0.21417 (rank : 35) | NC score | 0.031794 (rank : 91) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 679 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9HCU0, Q3SX55, Q96KB6 | Gene names | CD248, CD164L1, TEM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endosialin precursor (Tumor endothelial marker 1) (CD248 antigen). | |||||
|
ETV6_HUMAN
|
||||||
| θ value | 0.279714 (rank : 36) | NC score | 0.025014 (rank : 104) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P41212, Q9UMF6 | Gene names | ETV6, TEL, TEL1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor ETV6 (ETS-related protein Tel1) (Tel) (ETS translocation variant 6). | |||||
|
RGS3_MOUSE
|
||||||
| θ value | 0.279714 (rank : 37) | NC score | 0.043998 (rank : 71) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
GRIN1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 38) | NC score | 0.036052 (rank : 80) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q7Z2K8, Q8ND74, Q96PZ4 | Gene names | GPRIN1, KIAA1893 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
ZN667_HUMAN
|
||||||
| θ value | 0.47712 (rank : 39) | NC score | 0.002917 (rank : 180) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 773 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5HYK9, Q6B093, Q9H807 | Gene names | ZNF667 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 667. | |||||
|
MFAP1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 40) | NC score | 0.029688 (rank : 95) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 353 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9CQU1, Q3TU29, Q8CCL1, Q9CSJ5 | Gene names | Mfap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microfibrillar-associated protein 1. | |||||
|
MINT_MOUSE
|
||||||
| θ value | 0.62314 (rank : 41) | NC score | 0.029728 (rank : 94) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
NFAC1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 42) | NC score | 0.025484 (rank : 102) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O95644, Q12865, Q15793 | Gene names | NFATC1, NFAT2, NFATC | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor of activated T-cells, cytoplasmic 1 (NFAT transcription complex cytosolic component) (NF-ATc1) (NF-ATc). | |||||
|
SIX4_MOUSE
|
||||||
| θ value | 0.62314 (rank : 43) | NC score | 0.018071 (rank : 123) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61321, Q61322, Q61323 | Gene names | Six4, Arec3 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein SIX4 (Sine oculis homeobox homolog 4) (Skeletal muscle-specific ARE-binding protein AREC3). | |||||
|
CU024_HUMAN
|
||||||
| θ value | 0.813845 (rank : 44) | NC score | 0.056834 (rank : 54) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6XXX2 | Gene names | C21orf24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative uncharacterized protein C21orf24. | |||||
|
K1543_HUMAN
|
||||||
| θ value | 0.813845 (rank : 45) | NC score | 0.049820 (rank : 67) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9P1Y5, Q8NDF1 | Gene names | KIAA1543 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1543. | |||||
|
MKL1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 46) | NC score | 0.040671 (rank : 74) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 667 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8K4J6 | Gene names | Mkl1, Bsac | |||
|
Domain Architecture |

|
|||||
| Description | MKL/myocardin-like protein 1 (Myocardin-related transcription factor A) (MRTF-A) (Megakaryoblastic leukemia 1 protein homolog) (Basic SAP coiled-coil transcription activator). | |||||
|
SEPT9_HUMAN
|
||||||
| θ value | 0.813845 (rank : 47) | NC score | 0.024503 (rank : 106) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 706 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9UHD8, Q96QF3, Q96QF4, Q96QF5, Q9HA04, Q9UG40, Q9Y5W4 | Gene names | SEPT9, KIAA0991, MSF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Septin-9 (MLL septin-like fusion protein) (MLL septin-like fusion protein MSF-A) (Ovarian/Breast septin) (Ov/Br septin) (Septin D1). | |||||
|
ZBT17_HUMAN
|
||||||
| θ value | 0.813845 (rank : 48) | NC score | 0.004318 (rank : 175) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 925 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q13105, Q15932, Q5JYB2, Q9NUC9 | Gene names | ZBTB17, MIZ1, ZNF151 | |||
|
Domain Architecture |
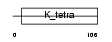
|
|||||
| Description | Zinc finger and BTB domain-containing protein 17 (Zinc finger protein 151) (Myc-interacting zinc finger protein) (Miz-1 protein). | |||||
|
ENPL_HUMAN
|
||||||
| θ value | 1.06291 (rank : 49) | NC score | 0.017613 (rank : 126) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P14625, Q96A97 | Gene names | HSP90B1, TRA1 | |||
|
Domain Architecture |

|
|||||
| Description | Endoplasmin precursor (Heat shock protein 90 kDa beta member 1) (94 kDa glucose-regulated protein) (GRP94) (gp96 homolog) (Tumor rejection antigen 1). | |||||
|
ERCC5_MOUSE
|
||||||
| θ value | 1.06291 (rank : 50) | NC score | 0.032716 (rank : 89) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P35689, Q61528, Q64248 | Gene names | Ercc5, Ercc-5, Xpg | |||
|
Domain Architecture |

|
|||||
| Description | DNA-repair protein complementing XP-G cells homolog (Xeroderma pigmentosum group G-complementing protein homolog) (DNA excision repair protein ERCC-5). | |||||
|
FGD3_HUMAN
|
||||||
| θ value | 1.06291 (rank : 51) | NC score | 0.010288 (rank : 152) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5JSP0, Q7Z7D9, Q8N5G1 | Gene names | FGD3, ZFYVE5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 3 (Zinc finger FYVE domain-containing protein 5). | |||||
|
NALDL_MOUSE
|
||||||
| θ value | 1.06291 (rank : 52) | NC score | 0.018471 (rank : 121) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7M758 | Gene names | Naaladl1, Naaladasel | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylated-alpha-linked acidic dipeptidase-like protein (EC 3.4.17.21) (NAALADase L). | |||||
|
SRRM1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 53) | NC score | 0.046968 (rank : 68) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
BAZ2A_MOUSE
|
||||||
| θ value | 1.38821 (rank : 54) | NC score | 0.021124 (rank : 117) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q91YE5 | Gene names | Baz2a, Tip5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5). | |||||
|
BPAEA_HUMAN
|
||||||
| θ value | 1.38821 (rank : 55) | NC score | 0.008850 (rank : 159) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1369 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O94833, Q5TBT0, Q8N1T8, Q8N8J3, Q8WXK9, Q96AK9, Q96DQ5, Q9H555 | Gene names | DST, BPAG1, DMH, DT, KIAA0728 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 6/9/10 (Trabeculin-beta) (Bullous pemphigoid antigen) (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
F100B_MOUSE
|
||||||
| θ value | 1.38821 (rank : 56) | NC score | 0.044286 (rank : 70) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BQH4, Q3UMH5 | Gene names | Fam100b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM100B. | |||||
|
NF2L1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 57) | NC score | 0.016998 (rank : 128) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q14494, Q12877, Q96FN6 | Gene names | NFE2L1, HBZ17, NRF1, TCF11 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor erythroid 2-related factor 1 (NF-E2-related factor 1) (NFE2-related factor 1) (Nuclear factor, erythroid derived 2, like 1) (Transcription factor 11) (Transcription factor HBZ17) (Transcription factor LCR-F1) (Locus control region-factor 1). | |||||
|
SC24C_HUMAN
|
||||||
| θ value | 1.38821 (rank : 58) | NC score | 0.031476 (rank : 92) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 662 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P53992, Q8WV25 | Gene names | SEC24C, KIAA0079 | |||
|
Domain Architecture |

|
|||||
| Description | Protein transport protein Sec24C (SEC24-related protein C). | |||||
|
TM108_MOUSE
|
||||||
| θ value | 1.38821 (rank : 59) | NC score | 0.034632 (rank : 84) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 336 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8BHE4, Q80WR9 | Gene names | Tmem108 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 108 precursor. | |||||
|
ZN509_MOUSE
|
||||||
| θ value | 1.38821 (rank : 60) | NC score | 0.003700 (rank : 179) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 990 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8BXX2, Q8CID4, Q8K2Z6 | Gene names | Znf509, Zfp509 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 509. | |||||
|
BAT3_HUMAN
|
||||||
| θ value | 1.81305 (rank : 61) | NC score | 0.041399 (rank : 72) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 490 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P46379, O95874, Q5SQ35, Q5SQ36, Q5SQ37, Q5SRP8, Q96SA6, Q9BCN4 | Gene names | BAT3, G3 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT3 (HLA-B-associated transcript 3) (Protein G3). | |||||
|
BAT3_MOUSE
|
||||||
| θ value | 1.81305 (rank : 62) | NC score | 0.046232 (rank : 69) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 518 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9Z1R2, Q8SNA3 | Gene names | Bat3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT3 (HLA-B-associated transcript 3). | |||||
|
BCL6_HUMAN
|
||||||
| θ value | 1.81305 (rank : 63) | NC score | 0.003920 (rank : 177) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 922 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P41182 | Gene names | BCL6, BCL5, LAZ3, ZBTB27, ZNF51 | |||
|
Domain Architecture |
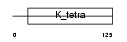
|
|||||
| Description | B-cell lymphoma 6 protein (BCL-6) (Zinc finger protein 51) (LAZ-3 protein) (BCL-5) (Zinc finger and BTB domain-containing protein 27). | |||||
|
IWS1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 64) | NC score | 0.023229 (rank : 111) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
K1802_MOUSE
|
||||||
| θ value | 1.81305 (rank : 65) | NC score | 0.038835 (rank : 77) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
MLL3_HUMAN
|
||||||
| θ value | 1.81305 (rank : 66) | NC score | 0.033915 (rank : 85) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
P66A_MOUSE
|
||||||
| θ value | 1.81305 (rank : 67) | NC score | 0.022293 (rank : 115) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CHY6, Q8BTQ2, Q8VEC9 | Gene names | Gatad2a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcriptional repressor p66 alpha (GATA zinc finger domain- containing protein 2A). | |||||
|
TAU_HUMAN
|
||||||
| θ value | 1.81305 (rank : 68) | NC score | 0.033358 (rank : 87) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 448 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P10636, P18518, Q14799, Q15549, Q15550, Q15551, Q9UDJ3, Q9UMH0, Q9UQ96 | Gene names | MAPT, MTBT1, TAU | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein tau (Neurofibrillary tangle protein) (Paired helical filament-tau) (PHF-tau). | |||||
|
ZN688_HUMAN
|
||||||
| θ value | 1.81305 (rank : 69) | NC score | 0.003738 (rank : 178) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 873 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8WV14, O75701, Q8IW91, Q96MN0 | Gene names | ZNF688 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 688. | |||||
|
CF010_HUMAN
|
||||||
| θ value | 2.36792 (rank : 70) | NC score | 0.021378 (rank : 116) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q5SRN2, Q5SPI9, Q5SPJ0, Q5SPK9, Q5SPL0, Q5SRN3, Q5TG25, Q5TG26, Q8N4B6 | Gene names | C6orf10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C6orf10. | |||||
|
MILK1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 71) | NC score | 0.035013 (rank : 82) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 788 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8BGT6, Q8BJ60 | Gene names | Mirab13, D15Mit260, Kiaa1668 | |||
|
Domain Architecture |

|
|||||
| Description | Molecule interacting with Rab13 (MIRab13) (MICAL-like protein 1). | |||||
|
MK15_HUMAN
|
||||||
| θ value | 2.36792 (rank : 72) | NC score | -0.001318 (rank : 183) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1038 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8TD08, Q2TCF9, Q8N362 | Gene names | MAPK15, ERK8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase 15 (EC 2.7.11.24) (Extracellular signal-regulated kinase 8). | |||||
|
PCDA5_HUMAN
|
||||||
| θ value | 2.36792 (rank : 73) | NC score | 0.002000 (rank : 181) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y5H7, O75284, Q8N4R3 | Gene names | PCDHA5 | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin alpha 5 precursor (PCDH-alpha5). | |||||
|
PER3_HUMAN
|
||||||
| θ value | 2.36792 (rank : 74) | NC score | 0.023353 (rank : 109) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P56645, Q5H8X4, Q969K6, Q96S77, Q96S78, Q9C0J3, Q9NSP9, Q9UGU8 | Gene names | PER3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (hPER3). | |||||
|
RC3H1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 75) | NC score | 0.014310 (rank : 136) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5TC82, Q5W180, Q5W181, Q8IVE6, Q8N9V1 | Gene names | RC3H1, KIAA2025 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Roquin (RING finger and C3H zinc finger protein 1). | |||||
|
TE2IP_HUMAN
|
||||||
| θ value | 2.36792 (rank : 76) | NC score | 0.032799 (rank : 88) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NYB0, Q8WYZ3, Q9NWR2 | Gene names | TERF2IP, RAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Telomeric repeat-binding factor 2-interacting protein 1 (TRF2- interacting telomeric protein Rap1) (hRap1). | |||||
|
TE2IP_MOUSE
|
||||||
| θ value | 2.36792 (rank : 77) | NC score | 0.033397 (rank : 86) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q91VL8, Q543F4, Q9JJE8, Q9JJE9 | Gene names | Terf2ip, Rap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Telomeric repeat-binding factor 2-interacting protein 1 (TRF2- interacting telomeric protein Rap1). | |||||
|
CAC1B_MOUSE
|
||||||
| θ value | 3.0926 (rank : 78) | NC score | 0.008006 (rank : 163) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O55017, Q60609 | Gene names | Cacna1b, Cach5, Cacnl1a5, Cchn1a | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent N-type calcium channel subunit alpha-1B (Voltage- gated calcium channel subunit alpha Cav2.2) (Calcium channel, L type, alpha-1 polypeptide isoform 5) (Brain calcium channel III) (BIII). | |||||
|
CEBPE_HUMAN
|
||||||
| θ value | 3.0926 (rank : 79) | NC score | 0.016027 (rank : 132) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q15744, Q15745, Q8IYI2, Q99803 | Gene names | CEBPE | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein epsilon (C/EBP epsilon). | |||||
|
CTND2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 80) | NC score | 0.017977 (rank : 124) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9UQB3, O00379, O15390, O43206, O43840, Q13589, Q9UM66, Q9UPM3 | Gene names | CTNND2, NPRAP | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-2 (Delta-catenin) (Neural plakophilin-related ARM-repeat protein) (NPRAP) (Neurojungin) (GT24). | |||||
|
FOXM1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 81) | NC score | 0.011562 (rank : 145) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q08050, O43258, O43259, O43260, Q4ZGG7, Q9BRL2 | Gene names | FOXM1, FKHL16, HFH11, MPP2, WIN | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein M1 (Forkhead-related protein FKHL16) (Hepatocyte nuclear factor 3 forkhead homolog 11) (HNF-3/fork-head homolog 11) (HFH-11) (Winged helix factor from INS-1 cells) (M-phase phosphoprotein 2) (MPM-2 reactive phosphoprotein 2) (Transcription factor Trident). | |||||
|
FRS3_MOUSE
|
||||||
| θ value | 3.0926 (rank : 82) | NC score | 0.015806 (rank : 134) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q91WJ0 | Gene names | Frs3, Frs2b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fibroblast growth factor receptor substrate 3 (FGFR substrate 3) (FRS2-beta). | |||||
|
HAP1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 83) | NC score | 0.012402 (rank : 142) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 606 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P54257, O75358, Q9H4G3, Q9HA98, Q9NY90 | Gene names | HAP1, HLP1 | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin-associated protein 1 (HAP-1) (Neuroan 1). | |||||
|
KPCL_HUMAN
|
||||||
| θ value | 3.0926 (rank : 84) | NC score | -0.002588 (rank : 184) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 887 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P24723, Q16246 | Gene names | PRKCH, PKCL | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C eta type (EC 2.7.11.13) (nPKC-eta) (PKC-L). | |||||
|
NPAS4_MOUSE
|
||||||
| θ value | 3.0926 (rank : 85) | NC score | 0.016984 (rank : 129) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8BGD7, Q3V3U3 | Gene names | Npas4, Nxf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF) (Limbic-enhanced PAS protein) (LE-PAS). | |||||
|
SM1L2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 86) | NC score | 0.006496 (rank : 169) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1019 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8NDV3, Q5TIC3, Q9Y3G5 | Gene names | SMC1L2 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 2 protein (SMC1beta protein). | |||||
|
AL2S8_MOUSE
|
||||||
| θ value | 4.03905 (rank : 87) | NC score | 0.017551 (rank : 127) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8VHI4 | Gene names | Als2cr8, Carf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyotrophic lateral sclerosis 2 chromosomal region candidate gene 8 protein homolog (Calcium-response factor) (CaRF). | |||||
|
BPA1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 88) | NC score | 0.006821 (rank : 167) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q91ZU6, Q91ZU7 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 1/2/3/4 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
INOC1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 89) | NC score | 0.010801 (rank : 150) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9ULG1, Q9NTG6 | Gene names | INOC1, KIAA1259 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative DNA helicase INO80 complex homolog 1 (EC 3.6.1.-) (hINO80). | |||||
|
INOC1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 90) | NC score | 0.010898 (rank : 149) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6ZPV2, Q6P7V0, Q6PCP1, Q8C9T7 | Gene names | Inoc1, Kiaa1259 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative DNA helicase INO80 complex homolog 1 (EC 3.6.1.-). | |||||
|
K1802_HUMAN
|
||||||
| θ value | 4.03905 (rank : 91) | NC score | 0.034763 (rank : 83) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1378 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q96JM3, Q6P181, Q8NC88, Q9BST0 | Gene names | KIAA1802, C13orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
NKTR_HUMAN
|
||||||
| θ value | 4.03905 (rank : 92) | NC score | 0.009390 (rank : 156) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 658 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P30414 | Gene names | NKTR | |||
|
Domain Architecture |

|
|||||
| Description | NK-tumor recognition protein (Natural-killer cells cyclophilin-related protein) (NK-TR protein). | |||||
|
CC28B_HUMAN
|
||||||
| θ value | 5.27518 (rank : 93) | NC score | 0.022895 (rank : 114) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BUN5, Q8TBV8 | Gene names | CCDC28B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 28B. | |||||
|
CC28B_MOUSE
|
||||||
| θ value | 5.27518 (rank : 94) | NC score | 0.023291 (rank : 110) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CEG5, Q8K132 | Gene names | Ccdc28b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 28B. | |||||
|
DOK3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 95) | NC score | 0.013568 (rank : 139) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9QZK7 | Gene names | Dok3, Dokl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Docking protein 3 (Downstream of tyrosine kinase 3) (P62(dok)-like protein) (DOK-L). | |||||
|
HDGR3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 96) | NC score | 0.012296 (rank : 143) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y3E1 | Gene names | HDGFRP3, HDGF2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatoma-derived growth factor-related protein 3 (HRP-3) (Hepatoma- derived growth factor 2). | |||||
|
LZTS1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 97) | NC score | 0.011540 (rank : 147) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 540 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P60853 | Gene names | Lzts1, Fez1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine zipper putative tumor suppressor 1 (F37/Esophageal cancer- related gene-coding leucine-zipper motif) (Fez1). | |||||
|
MDC1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 98) | NC score | 0.027328 (rank : 97) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
NADE1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 99) | NC score | 0.014727 (rank : 135) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q711T7, Q711P6, Q8CFY6, Q9D280 | Gene names | Nadsyn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamine-dependent NAD(+) synthetase (EC 6.3.5.1) (NAD(+) synthase [glutamine-hydrolyzing]) (NAD(+) synthetase 1) (NH3-dependent NAD(+) synthetase-like protein). | |||||
|
S3TC2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 100) | NC score | 0.011555 (rank : 146) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80VA5, Q8BWH2, Q8BXB8, Q8C0G6 | Gene names | Sh3tc2, Kiaa1985 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 domain and tetratricopeptide repeats-containing protein 2. | |||||
|
SRRM1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 101) | NC score | 0.037382 (rank : 78) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
ZC3H6_HUMAN
|
||||||
| θ value | 5.27518 (rank : 102) | NC score | 0.014104 (rank : 138) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P61129 | Gene names | ZC3H6, KIAA2035, ZC3HDC6 | |||
|
Domain Architecture |
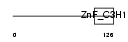
|
|||||
| Description | Zinc finger CCCH domain-containing protein 6. | |||||
|
ARI1A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 103) | NC score | 0.023526 (rank : 107) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 787 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O14497, Q53FK9, Q5T0W1, Q5T0W2, Q5T0W3, Q8NFD6, Q96T89, Q9BY33, Q9HBJ5, Q9UPZ1 | Gene names | ARID1A, C1orf4, OSA1, SMARCF1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 1A (ARID domain- containing protein 1A) (SWI/SNF-related, matrix-associated, actin- dependent regulator of chromatin subfamily F member 1) (SWI-SNF complex protein p270) (B120) (SWI-like protein) (Osa homolog 1) (hOSA1) (hELD) (BRG1-associated factor 250) (BAF250) (BRG1-associated factor 250a) (BAF250A). | |||||
|
BCN1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 104) | NC score | 0.009276 (rank : 157) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q14457, O75595, Q9UNA8 | Gene names | BECN1, GT197 | |||
|
Domain Architecture |

|
|||||
| Description | Beclin-1 (Coiled-coil myosin-like BCL2-interacting protein) (Protein GT197). | |||||
|
BRDT_MOUSE
|
||||||
| θ value | 6.88961 (rank : 105) | NC score | 0.009394 (rank : 155) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q91Y44, Q59HJ4 | Gene names | Brdt, Fsrg3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain testis-specific protein (RING3-like protein) (Bromodomain- containing female sterile homeotic-like protein). | |||||
|
BSN_MOUSE
|
||||||
| θ value | 6.88961 (rank : 106) | NC score | 0.035882 (rank : 81) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
CD248_MOUSE
|
||||||
| θ value | 6.88961 (rank : 107) | NC score | 0.019676 (rank : 119) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 578 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q91V98, Q3UMV6, Q91ZV1 | Gene names | Cd248, Tem1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endosialin precursor (Tumor endothelial marker 1) (CD248 antigen). | |||||
|
DLGP1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 108) | NC score | 0.008034 (rank : 161) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O14490, O14489, P78335 | Gene names | DLGAP1, DAP1, GKAP | |||
|
Domain Architecture |

|
|||||
| Description | Disks large-associated protein 1 (DAP-1) (Guanylate kinase-associated protein) (hGKAP) (SAP90/PSD-95-associated protein 1) (SAPAP1) (PSD- 95/SAP90-binding protein 1). | |||||
|
DMRTD_HUMAN
|
||||||
| θ value | 6.88961 (rank : 109) | NC score | 0.016512 (rank : 131) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8IXT2, Q8N6Q2, Q96M39, Q96SD4 | Gene names | DMRTC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Doublesex- and mab-3-related transcription factor C2. | |||||
|
FOXC2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 110) | NC score | 0.004000 (rank : 176) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q61850, P97948, Q63869, Q8C694 | Gene names | Foxc2, Fkh14, Fkhl14, Mfh1 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein C2 (Forkhead-related protein FKHL14) (Mesenchyme fork head protein 1) (MFH-1 protein) (Transcription factor FKH-14). | |||||
|
GLU2B_HUMAN
|
||||||
| θ value | 6.88961 (rank : 111) | NC score | 0.011470 (rank : 148) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 377 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P14314, Q96BU9, Q96D06, Q9P0W9 | Gene names | PRKCSH, G19P1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glucosidase 2 subunit beta precursor (Glucosidase II subunit beta) (Protein kinase C substrate, 60.1 kDa protein, heavy chain) (PKCSH) (80K-H protein). | |||||
|
MAP1A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 112) | NC score | 0.019099 (rank : 120) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P78559, O95643, Q12973, Q15882, Q9UJT4 | Gene names | MAP1A | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 1A (MAP 1A) (Proliferation-related protein p80) [Contains: MAP1 light chain LC2]. | |||||
|
MTSS1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 113) | NC score | 0.017868 (rank : 125) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O43312, Q8TCA2, Q96RX2 | Gene names | MTSS1, KIAA0429, MIM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metastasis suppressor protein 1 (Missing in metastasis protein) (Metastasis suppressor YGL-1). | |||||
|
MUG2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 114) | NC score | 0.006239 (rank : 170) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P28666 | Gene names | Mug2, Mug-2 | |||
|
Domain Architecture |

|
|||||
| Description | Murinoglobulin-2 precursor (MuG2). | |||||
|
PI5PA_MOUSE
|
||||||
| θ value | 6.88961 (rank : 115) | NC score | 0.026685 (rank : 99) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P59644 | Gene names | Pib5pa | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 4,5-bisphosphate 5-phosphatase A (EC 3.1.3.56). | |||||
|
PRKDC_HUMAN
|
||||||
| θ value | 6.88961 (rank : 116) | NC score | 0.007673 (rank : 165) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P78527, P78528, Q13327, Q13337, Q14175, Q59H99, Q7Z611, Q96SE6, Q9UME3 | Gene names | PRKDC | |||
|
Domain Architecture |

|
|||||
| Description | DNA-dependent protein kinase catalytic subunit (EC 2.7.11.1) (DNA-PK catalytic subunit) (DNA-PKcs) (DNPK1) (p460). | |||||
|
RBY1A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 117) | NC score | 0.005544 (rank : 171) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15414, Q15377, Q8NHR0 | Gene names | RBMY1A1, RBM1, YRRM1 | |||
|
Domain Architecture |
|
|||||
| Description | RNA-binding motif protein, Y chromosome, family 1 member A1 (RNA- binding motif protein 1). | |||||
|
RT28_HUMAN
|
||||||
| θ value | 6.88961 (rank : 118) | NC score | 0.016944 (rank : 130) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 4 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y2Q9, Q96Q21 | Gene names | MRPS28, MRPS35 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitochondrial 28S ribosomal protein S28 (S28mt) (MRP-S28) (MRP-S35). | |||||
|
SFRS4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 119) | NC score | 0.007389 (rank : 166) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q08170, Q5VXP1, Q9BUA4, Q9UEB5 | Gene names | SFRS4, SRP75 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 4 (Pre-mRNA-splicing factor SRP75) (SRP001LB). | |||||
|
SMOO_HUMAN
|
||||||
| θ value | 6.88961 (rank : 120) | NC score | 0.025143 (rank : 103) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 391 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P53814, O00569, O95769, O95937, Q9P1S8, Q9UIT1, Q9UIT2 | Gene names | SMTN, SMSMO | |||
|
Domain Architecture |

|
|||||
| Description | Smoothelin. | |||||
|
ARI4A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 121) | NC score | 0.012693 (rank : 141) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 434 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P29374, Q15991, Q15992, Q15993 | Gene names | ARID4A, RBBP1, RBP1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 4A (ARID domain- containing protein 4A) (Retinoblastoma-binding protein 1) (RBBP-1). | |||||
|
BAG3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 122) | NC score | 0.030973 (rank : 93) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9JLV1, Q9JJC7 | Gene names | Bag3, Bis | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 3 (BCL-2-binding athanogene- 3) (BAG-3) (Bcl-2-binding protein Bis). | |||||
|
BCL9_HUMAN
|
||||||
| θ value | 8.99809 (rank : 123) | NC score | 0.032502 (rank : 90) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1008 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O00512 | Gene names | BCL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9) (Legless homolog). | |||||
|
CUTL1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 124) | NC score | 0.014203 (rank : 137) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1574 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P39880, Q6NYH4, Q9UEV5 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP). | |||||
|
DDX51_HUMAN
|
||||||
| θ value | 8.99809 (rank : 125) | NC score | 0.008022 (rank : 162) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8N8A6, Q5CZ71, Q8IXK5, Q96ED1 | Gene names | DDX51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX51 (EC 3.6.1.-) (DEAD box protein 51). | |||||
|
GNAS1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 126) | NC score | 0.008930 (rank : 158) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q5JWF2, O75684, O75685, Q5JW67, Q5JWF1, Q9NY42 | Gene names | GNAS, GNAS1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Guanine nucleotide-binding protein G(s) subunit alpha isoforms XLas (Adenylate cyclase-stimulating G alpha protein) (Extra large alphas protein) (XLalphas). | |||||
|
LPP_HUMAN
|
||||||
| θ value | 8.99809 (rank : 127) | NC score | 0.012934 (rank : 140) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 775 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q93052, Q8NFX5 | Gene names | LPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipoma-preferred partner (LIM domain-containing preferred translocation partner in lipoma). | |||||
|
LRCH2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 128) | NC score | 0.009987 (rank : 153) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5VUJ6, Q9HA88, Q9P233 | Gene names | LRCH2, KIAA1495 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 2. | |||||
|
MCP_MOUSE
|
||||||
| θ value | 8.99809 (rank : 129) | NC score | 0.004787 (rank : 173) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O88174, Q9R0R9 | Gene names | Cd46, Mcp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane cofactor protein precursor (CD46 antigen). | |||||
|
MEF2D_HUMAN
|
||||||
| θ value | 8.99809 (rank : 130) | NC score | 0.008740 (rank : 160) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q14814, Q14815 | Gene names | MEF2D | |||
|
Domain Architecture |
|
|||||
| Description | Myocyte-specific enhancer factor 2D. | |||||
|
MFAP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 131) | NC score | 0.020369 (rank : 118) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P55081 | Gene names | MFAP1 | |||
|
Domain Architecture |
|
|||||
| Description | Microfibrillar-associated protein 1. | |||||
|
PARD3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 132) | NC score | 0.004321 (rank : 174) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8TEW0, Q8TEW1, Q8TEW2, Q8TEW3, Q96K28, Q96RM6, Q96RM7, Q9BY57, Q9BY58, Q9HC48, Q9NWL4, Q9NYE6 | Gene names | PARD3, PAR3, PAR3A | |||
|
Domain Architecture |

|
|||||
| Description | Partitioning-defective 3 homolog (PARD-3) (PAR-3) (Atypical PKC isotype-specific-interacting protein) (ASIP) (CTCL tumor antigen se2- 5) (PAR3-alpha). | |||||
|
PCLO_MOUSE
|
||||||
| θ value | 8.99809 (rank : 133) | NC score | 0.024870 (rank : 105) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
RNPL1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 134) | NC score | 0.006540 (rank : 168) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9HAU8, Q96AC9, Q9H033, Q9NVD0 | Gene names | RNPEPL1 | |||
|
Domain Architecture |

|
|||||
| Description | Arginyl aminopeptidase-like 1 (EC 3.4.11.-) (RNPEP-like protein). | |||||
|
SGOL2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 135) | NC score | 0.009858 (rank : 154) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7TSY8, Q811I4, Q8C9C4, Q8CGJ4, Q9CS55 | Gene names | Sgol2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Shugoshin-like 2. | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 136) | NC score | 0.029002 (rank : 96) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
SYF2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 137) | NC score | 0.012199 (rank : 144) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D198 | Gene names | Syf2, Cbpin, Gcipip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pre-mRNA-splicing factor SYF2 (CCNDBP1-interactor) (p29) (mp29). | |||||
|
TACC2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 138) | NC score | 0.010673 (rank : 151) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 774 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9JJG0 | Gene names | Tacc2 | |||
|
Domain Architecture |

|
|||||
| Description | Transforming acidic coiled-coil-containing protein 2. | |||||
|
TARA_HUMAN
|
||||||
| θ value | 8.99809 (rank : 139) | NC score | 0.039278 (rank : 76) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q9H2D6, O94797, Q2PZW8, Q2Q3Z9, Q2Q400, Q5R3M6, Q96DW1, Q9BT77, Q9BTL7, Q9BY98, Q9Y3L4 | Gene names | TRIOBP, KIAA1662, TARA | |||
|
Domain Architecture |

|
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
TS13_MOUSE
|
||||||
| θ value | 8.99809 (rank : 140) | NC score | 0.015809 (rank : 133) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q01755 | Gene names | Tcp11 | |||
|
Domain Architecture |

|
|||||
| Description | Testis-specific protein PBS13. | |||||
|
YTHD2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 141) | NC score | 0.007698 (rank : 164) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y5A9, Q5VSZ9, Q8TDH0, Q9BUJ5 | Gene names | YTHDF2, HGRG8 | |||
|
Domain Architecture |

|
|||||
| Description | YTH domain family protein 2 (High-glucose-regulated protein 8) (NY- REN-2 antigen) (CLL-associated antigen KW-14). | |||||
|
ZN621_HUMAN
|
||||||
| θ value | 8.99809 (rank : 142) | NC score | 0.001567 (rank : 182) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 737 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6ZSS3, Q8TE91 | Gene names | ZNF621 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 621. | |||||
|
3BP2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 143) | NC score | 0.111470 (rank : 36) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P78314, O00500, O15373, P78315 | Gene names | SH3BP2, 3BP2 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 domain-binding protein 2 (3BP-2). | |||||
|
3BP2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 144) | NC score | 0.112613 (rank : 35) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q06649 | Gene names | Sh3bp2, 3bp2 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 domain-binding protein 2 (3BP-2). | |||||
|
AB1IP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 145) | NC score | 0.054722 (rank : 59) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q7Z5R6, Q8IWS8, Q8IZZ7 | Gene names | APBB1IP, PREL1, RARP1, RIAM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 1- interacting protein (APBB1-interacting protein 1) (Rap1-GTP- interacting adapter molecule) (RIAM) (Proline-rich EVH1 ligand 1) (PREL-1) (Proline-rich protein 73) (Retinoic acid-responsive proline- rich protein 1) (RARP-1). | |||||
|
ANLN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 146) | NC score | 0.115852 (rank : 33) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9NQW6, Q5CZ78, Q6NSK5, Q9H8Y4, Q9NVN9, Q9NVP0 | Gene names | ANLN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Actin-binding protein anillin. | |||||
|
ANLN_MOUSE
|
||||||
| θ value | θ > 10 (rank : 147) | NC score | 0.094305 (rank : 44) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 297 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8K298, Q8BL79, Q8BLB3, Q8K2N0, Q9CUY0 | Gene names | Anln | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Actin-binding protein anillin. | |||||
|
CENB2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 148) | NC score | 0.066065 (rank : 49) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q15057, Q9UQR3 | Gene names | CENTB2, KIAA0041 | |||
|
Domain Architecture |

|
|||||
| Description | Centaurin-beta 2 (Cnt-b2). | |||||
|
CENB5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 149) | NC score | 0.060317 (rank : 51) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q96P50, Q9BSR9, Q9C0E7 | Gene names | CENTB5, KIAA1716 | |||
|
Domain Architecture |

|
|||||
| Description | Centaurin-beta 5 (Cnt-b5). | |||||
|
CNKR1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 150) | NC score | 0.192006 (rank : 23) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q969H4, O95381 | Gene names | CNKSR1, CNK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Connector enhancer of kinase suppressor of ras 1 (Connector enhancer of KSR1) (hCNK1) (Connector enhancer of KSR-like) (CNK homolog protein 1). | |||||
|
CNKR2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 151) | NC score | 0.097952 (rank : 42) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8WXI2, O94976, Q8WXI1 | Gene names | CNKSR2, CNK2, KIAA0902, KSR2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Connector enhancer of kinase suppressor of ras 2 (Connector enhancer of KSR2) (CNK2). | |||||
|
CNKR2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 152) | NC score | 0.098038 (rank : 41) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q80YA9, Q80TP2 | Gene names | Cnksr2, Kiaa0902 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Connector enhancer of kinase suppressor of ras 2 (Connector enhancer of KSR2) (CNK2). | |||||
|
DAPP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 153) | NC score | 0.109210 (rank : 37) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UN19, Q8TCK5, Q9UHF2 | Gene names | DAPP1, BAM32 | |||
|
Domain Architecture |
|
|||||
| Description | Dual adapter for phosphotyrosine and 3-phosphotyrosine and 3- phosphoinositide (hDAPP1) (B cell adapter molecule of 32 kDa) (B lymphocyte adapter protein Bam32). | |||||
|
DAPP1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 154) | NC score | 0.114917 (rank : 34) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9QXT1, Q9R178 | Gene names | Dapp1, Bam32 | |||
|
Domain Architecture |
|
|||||
| Description | Dual adapter for phosphotyrosine and 3-phosphotyrosine and 3- phosphoinositide (mDAPP1) (B cell adapter molecule of 32 kDa) (B lymphocyte adapter protein Bam32). | |||||
|
DYN2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 155) | NC score | 0.056489 (rank : 55) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P50570, Q7Z5S3, Q9UPH4 | Gene names | DNM2, DYN2 | |||
|
Domain Architecture |

|
|||||
| Description | Dynamin-2 (EC 3.6.5.5). | |||||
|
DYN2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 156) | NC score | 0.055731 (rank : 57) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P39054, Q9DBE1 | Gene names | Dnm2, Dyn2 | |||
|
Domain Architecture |

|
|||||
| Description | Dynamin-2 (EC 3.6.5.5) (Dynamin UDNM). | |||||
|
DYN3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 157) | NC score | 0.058854 (rank : 52) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UQ16, O14982, O95555, Q5W129, Q9H0P3, Q9H548, Q9NQ68, Q9NQN6 | Gene names | DNM3, KIAA0820 | |||
|
Domain Architecture |

|
|||||
| Description | Dynamin-3 (EC 3.6.5.5) (Dynamin, testicular) (T-dynamin). | |||||
|
GNRP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 158) | NC score | 0.056993 (rank : 53) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q13972, Q16027 | Gene names | RASGRF1, CDC25 | |||
|
Domain Architecture |

|
|||||
| Description | Guanine nucleotide-releasing protein (GNRP) (Ras-specific nucleotide exchange factor CDC25) (Ras-specific guanine nucleotide-releasing factor). | |||||
|
GNRP_MOUSE
|
||||||
| θ value | θ > 10 (rank : 159) | NC score | 0.054064 (rank : 61) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P27671 | Gene names | Rasgrf1, Cdc25, Grf1 | |||
|
Domain Architecture |

|
|||||
| Description | Guanine nucleotide-releasing protein (GNRP) (Ras-specific nucleotide exchange factor CDC25) (CDC25Mm). | |||||
|
MTMR5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 160) | NC score | 0.053140 (rank : 62) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O95248, O60228, Q5JXD8, Q5PPM2, Q96GR9, Q9UGB8 | Gene names | SBF1, MTMR5 | |||
|
Domain Architecture |

|
|||||
| Description | SET-binding factor 1 (Sbf1) (Myotubularin-related protein 5). | |||||
|
OSBL8_HUMAN
|
||||||
| θ value | θ > 10 (rank : 161) | NC score | 0.053091 (rank : 63) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9BZF1, Q8WXP8, Q9P277 | Gene names | OSBPL8, KIAA1451, ORP8, OSBP10 | |||
|
Domain Architecture |

|
|||||
| Description | Oxysterol-binding protein-related protein 8 (OSBP-related protein 8) (ORP-8). | |||||
|
PHLB1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 162) | NC score | 0.051230 (rank : 66) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1038 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q86UU1, O75133, Q4KMF8, Q8TEQ2 | Gene names | PHLDB1, DLNB07, KIAA0638, LL5A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family B member 1 (Protein LL5-alpha). | |||||
|
PKHA1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 163) | NC score | 0.133234 (rank : 26) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9HB21, Q9BVK0 | Gene names | PLEKHA1, TAPP1 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 1 (Tandem PH domain-containing protein 1) (TAPP-1). | |||||
|
PKHA1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 164) | NC score | 0.132768 (rank : 27) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BUL6, Q8BK96, Q8BXU1, Q8VE13 | Gene names | Plekha1, Tapp1 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 1 (Tandem PH domain-containing protein 1) (TAPP-1). | |||||
|
PKHA2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 165) | NC score | 0.104431 (rank : 38) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9HB19 | Gene names | PLEKHA2, TAPP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family A member 2 (Tandem PH domain-containing protein 2) (TAPP-2). | |||||
|
PKHA2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 166) | NC score | 0.118445 (rank : 32) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9ERS5, Q8BY29 | Gene names | Plekha2, Tapp2 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 2 (Tandem PH domain-containing protein 2) (TAPP-2) (PH domain-containing adaptor PHAD47). | |||||
|
PKHA4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 167) | NC score | 0.118900 (rank : 31) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9H4M7, Q8N4M8, Q8N658 | Gene names | PLEKHA4, PEPP1 | |||
|
Domain Architecture |
|
|||||
| Description | Pleckstrin homology domain-containing family A member 4 (Phosphoinositol 3-phosphate-binding protein 1) (PEPP-1). | |||||
|
PKHA4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 168) | NC score | 0.123373 (rank : 30) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8VC98, Q8R3N3 | Gene names | Plekha4, Pepp1 | |||
|
Domain Architecture |
|
|||||
| Description | Pleckstrin homology domain-containing family A member 4 (Phosphoinositol 3-phosphate-binding protein 1) (PEPP-1). | |||||
|
PKHA5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 169) | NC score | 0.096758 (rank : 43) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9HAU0, Q8N3K6, Q96DY9, Q9BVR4, Q9C0H7, Q9H924, Q9NVK8 | Gene names | PLEKHA5, KIAA1686, PEPP2 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 5 (Phosphoinositol 3-phosphate-binding protein 2) (PEPP-2). | |||||
|
PKHA6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 170) | NC score | 0.136106 (rank : 24) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9Y2H5 | Gene names | PLEKHA6, KIAA0969, PEPP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family A member 6 (Phosphoinositol 3-phosphate-binding protein 3) (PEPP-3). | |||||
|
PKHA6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 171) | NC score | 0.130300 (rank : 29) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 591 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q7TQG1, Q8K0J5 | Gene names | Plekha6, Pepp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 6 (Phosphoinositol 3-phosphate-binding protein 3) (PEPP-3). | |||||
|
PKHB1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 172) | NC score | 0.051348 (rank : 65) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UF11, Q9UBF5, Q9UI37, Q9UI44 | Gene names | PLEKHB1, KPL1, PHR1 | |||
|
Domain Architecture |
|
|||||
| Description | Pleckstrin homology domain-containing family B member 1 (Pleckstrin homology domain retinal protein 1) (PH domain-containing protein in retina 1) (PHRET1). | |||||
|
PKHB2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 173) | NC score | 0.056318 (rank : 56) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q96CS7, Q86W37, Q9BV75, Q9NWK1 | Gene names | PLEKHB2, EVT2 | |||
|
Domain Architecture |
|
|||||
| Description | Pleckstrin homology domain-containing family B member 2 (Evectin-2). | |||||
|
PKHB2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 174) | NC score | 0.054867 (rank : 58) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9QZC7, Q3UDH6, Q8C4I4, Q8CA27 | Gene names | Plekhb2, Evt2 | |||
|
Domain Architecture |
|
|||||
| Description | Pleckstrin homology domain-containing family B member 2 (Evectin-2). | |||||
|
PLEK2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 175) | NC score | 0.099184 (rank : 40) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NYT0, Q96JT0 | Gene names | PLEK2 | |||
|
Domain Architecture |
|
|||||
| Description | Pleckstrin-2. | |||||
|
PLEK2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 176) | NC score | 0.101095 (rank : 39) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9WV52 | Gene names | Plek2 | |||
|
Domain Architecture |
|
|||||
| Description | Pleckstrin-2. | |||||
|
PLEK_HUMAN
|
||||||
| θ value | θ > 10 (rank : 177) | NC score | 0.130642 (rank : 28) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P08567, Q53SU8, Q6FGM8, Q8WV81 | Gene names | PLEK, P47 | |||
|
Domain Architecture |
|
|||||
| Description | Pleckstrin (Platelet p47 protein). | |||||
|
PLEK_MOUSE
|
||||||
| θ value | θ > 10 (rank : 178) | NC score | 0.133417 (rank : 25) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9JHK5, Q9ERI9 | Gene names | Plek | |||
|
Domain Architecture |
|
|||||
| Description | Pleckstrin. | |||||
|
RASA1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 179) | NC score | 0.077785 (rank : 45) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P20936 | Gene names | RASA1, RASA | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 1 (GTPase-activating protein) (GAP) (Ras p21 protein activator) (p120GAP) (RasGAP). | |||||
|
RHG25_HUMAN
|
||||||
| θ value | θ > 10 (rank : 180) | NC score | 0.073744 (rank : 47) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 550 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P42331, Q8IXQ2 | Gene names | ARHGAP25, KIAA0053 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 25. | |||||
|
RHG25_MOUSE
|
||||||
| θ value | θ > 10 (rank : 181) | NC score | 0.077066 (rank : 46) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 492 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8BYW1, Q2VPR2, Q3TE04, Q3TVX0, Q3UVB7, Q6A0E0, Q8BX98 | Gene names | Arhgap25, Kiaa0053 | |||
|
Domain Architecture |
|
|||||
| Description | Rho-GTPase-activating protein 25. | |||||
|
SWP70_HUMAN
|
||||||
| θ value | θ > 10 (rank : 182) | NC score | 0.068988 (rank : 48) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 973 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9UH65, O75135, Q7LCY6, Q9P061, Q9P0Z8 | Gene names | SWAP70, KIAA0640 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Switch-associated protein 70 (SWAP-70). | |||||
|
SWP70_MOUSE
|
||||||
| θ value | θ > 10 (rank : 183) | NC score | 0.060496 (rank : 50) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1097 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6A028, O88443, Q3TQR6, Q3UCA3, Q6P1D0 | Gene names | Swap70, Kiaa0640 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Switch-associated protein 70 (SWAP-70). | |||||
|
TBCD2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 184) | NC score | 0.051916 (rank : 64) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9BYX2 | Gene names | TBC1D2, PARIS1 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 2 (Prostate antigen recognized and identified by SEREX) (PARIS-1). | |||||
|
GBF1_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 142 | |
| SwissProt Accessions | Q92538, Q5VXX3, Q96CK6, Q96HZ3, Q9H473 | Gene names | GBF1, KIAA0248 | |||
|
Domain Architecture |

|
|||||
| Description | Golgi-specific brefeldin A-resistance guanine nucleotide exchange factor 1 (BFA-resistant GEF 1). | |||||
|
BIG1_HUMAN
|
||||||
| NC score | 0.869742 (rank : 2) | θ value | 3.59435e-57 (rank : 3) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9Y6D6, Q9NV46, Q9UFV2, Q9UNL0 | Gene names | ARFGEF1, ARFGEP1, BIG1 | |||
|
Domain Architecture |

|
|||||
| Description | Brefeldin A-inhibited guanine nucleotide-exchange protein 1 (Brefeldin A-inhibited GEP 1) (p200 ARF-GEP1) (p200 ARF guanine nucleotide exchange factor). | |||||
|
BIG2_HUMAN
|
||||||
| NC score | 0.868836 (rank : 3) | θ value | 4.10295e-61 (rank : 2) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9Y6D5, Q5TFT9, Q9NTS1 | Gene names | ARFGEF2, ARFGEP2, BIG2 | |||
|
Domain Architecture |

|
|||||
| Description | Brefeldin A-inhibited guanine nucleotide-exchange protein 2 (Brefeldin A-inhibited GEP 2). | |||||
|
IQEC3_HUMAN
|
||||||
| NC score | 0.824317 (rank : 4) | θ value | 1.85889e-29 (rank : 16) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9UPP2, Q8TB43 | Gene names | IQSEC3, KIAA1110 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 3. | |||||
|
IQEC1_MOUSE
|
||||||
| NC score | 0.821060 (rank : 5) | θ value | 5.22648e-32 (rank : 13) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8R0S2, Q3TZC3, Q5DU15 | Gene names | Iqsec1, Kiaa0763 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 1. | |||||
|
IQEC3_MOUSE
|
||||||
| NC score | 0.819649 (rank : 6) | θ value | 1.85889e-29 (rank : 17) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q3TES0, Q3UHP8, Q80TJ8 | Gene names | Iqsec3, Kiaa1110 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 3. | |||||
|
IQEC1_HUMAN
|
||||||
| NC score | 0.818949 (rank : 7) | θ value | 5.22648e-32 (rank : 12) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6DN90, O94863, Q96D85 | Gene names | IQSEC1, ARFGEP100, BRAG2, KIAA0763 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 1 (ADP-ribosylation factors guanine nucleotide-exchange protein 100) (ADP-ribosylation factors guanine nucleotide-exchange protein 2) (Brefeldin-resistant Arf-GEF 2 protein). | |||||
|
CYH2_MOUSE
|
||||||
| NC score | 0.813468 (rank : 8) | θ value | 6.59618e-35 (rank : 8) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P63034, O89099, P97695, Q3UP39 | Gene names | Pscd2, Sec7b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytohesin-2 (CLM2) (SEC7 homolog B) (msec7-2). | |||||
|
CYH3_HUMAN
|
||||||
| NC score | 0.813035 (rank : 9) | θ value | 9.20422e-37 (rank : 4) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O43739 | Gene names | PSCD3, ARNO3, GRP1 | |||
|
Domain Architecture |

|
|||||
| Description | Cytohesin-3 (ARF nucleotide-binding site opener 3) (ARNO3 protein) (General receptor of phosphoinositides 1) (Grp1). | |||||
|
CYH2_HUMAN
|
||||||
| NC score | 0.812975 (rank : 10) | θ value | 2.50655e-34 (rank : 9) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q99418, Q92958 | Gene names | PSCD2, ARNO | |||
|
Domain Architecture |

|
|||||
| Description | Cytohesin-2 (ARF nucleotide-binding site opener) (ARNO protein) (ARF exchange factor). | |||||
|
CYH4_MOUSE
|
||||||
| NC score | 0.810636 (rank : 11) | θ value | 1.6247e-33 (rank : 10) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q80YW0 | Gene names | Pscd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytohesin-4. | |||||
|
CYH1_MOUSE
|
||||||
| NC score | 0.810310 (rank : 12) | θ value | 7.79185e-36 (rank : 7) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9QX11, O88817 | Gene names | Pscd1 | |||
|
Domain Architecture |

|
|||||
| Description | Cytohesin-1 (CLM1). | |||||
|
CYH4_HUMAN
|
||||||
| NC score | 0.810015 (rank : 13) | θ value | 4.72714e-33 (rank : 11) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UIA0, Q5R3F9, Q9UGT6 | Gene names | PSCD4, CYT4 | |||
|
Domain Architecture |

|
|||||
| Description | Cytohesin-4. | |||||
|
CYH1_HUMAN
|
||||||
| NC score | 0.808090 (rank : 14) | θ value | 7.79185e-36 (rank : 6) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q15438, Q9P123, Q9P124 | Gene names | PSCD1, D17S811E | |||
|
Domain Architecture |

|
|||||
| Description | Cytohesin-1 (SEC7 homolog B2-1). | |||||
|
CYH3_MOUSE
|
||||||
| NC score | 0.807238 (rank : 15) | θ value | 9.20422e-37 (rank : 5) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O08967, Q8CI93 | Gene names | Pscd3, Grp1 | |||
|
Domain Architecture |

|
|||||
| Description | Cytohesin-3 (ARF nucleotide-binding site opener 3) (ARNO3 protein) (General receptor of phosphoinositides 1) (Grp1) (CLM-3). | |||||
|
IQEC2_MOUSE
|
||||||
| NC score | 0.782323 (rank : 16) | θ value | 4.42448e-31 (rank : 14) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q5DU25 | Gene names | Iqsec2, Kiaa0522 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 2. | |||||
|
IQEC2_HUMAN
|
||||||
| NC score | 0.781531 (rank : 17) | θ value | 9.8567e-31 (rank : 15) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q5JU85, O60275 | Gene names | IQSEC2, KIAA0522 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 2. | |||||
|
PSD3_HUMAN
|
||||||
| NC score | 0.755649 (rank : 18) | θ value | 1.5773e-20 (rank : 18) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9NYI0, Q9Y2F1 | Gene names | PSD3, EFA6R, HCA67, KIAA0942 | |||
|
Domain Architecture |

|
|||||
| Description | PH and SEC7 domain-containing protein 3 (Pleckstrin homology and SEC7 domain-containing protein 3) (Exchange factor for ADP-ribosylation factor guanine nucleotide factor 6) (Hepatocellular carcinoma- associated antigen 67). | |||||
|
PSD4_MOUSE
|
||||||
| NC score | 0.720633 (rank : 19) | θ value | 3.28887e-18 (rank : 19) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BLR5, Q3TE33, Q3TQF5, Q3UD59, Q3UFH9, Q80V44 | Gene names | Psd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH and SEC7 domain-containing protein 4 (Pleckstrin homology and SEC7 domain-containing protein 4). | |||||
|
FBX8_MOUSE
|
||||||
| NC score | 0.715186 (rank : 20) | θ value | 5.26297e-08 (rank : 21) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9QZN3 | Gene names | Fbxo8, Fbx8 | |||
|
Domain Architecture |

|
|||||
| Description | F-box only protein 8. | |||||
|
PSD4_HUMAN
|
||||||
| NC score | 0.714305 (rank : 21) | θ value | 3.63628e-17 (rank : 20) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8NDX1, O95621, Q4ZG34, Q6GPH8, Q8IYP4 | Gene names | PSD4, EFA6B, TIC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH and SEC7 domain-containing protein 4 (Pleckstrin homology and SEC7 domain-containing protein 4) (Exchange factor for ADP-ribosylation factor guanine nucleotide factor 6B) (Telomeric of interleukin-1 cluster protein). | |||||
|
FBX8_HUMAN
|
||||||
| NC score | 0.712496 (rank : 22) | θ value | 8.97725e-08 (rank : 22) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NRD0, Q6UWN4, Q9NRP5, Q9UKC4 | Gene names | FBXO8, FBS, FBX8 | |||
|
Domain Architecture |

|
|||||
| Description | F-box only protein 8 (F-box/SEC7 protein FBS). | |||||
|
CNKR1_HUMAN
|
||||||
| NC score | 0.192006 (rank : 23) | θ value | θ > 10 (rank : 150) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q969H4, O95381 | Gene names | CNKSR1, CNK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Connector enhancer of kinase suppressor of ras 1 (Connector enhancer of KSR1) (hCNK1) (Connector enhancer of KSR-like) (CNK homolog protein 1). | |||||
|
PKHA6_HUMAN
|
||||||
| NC score | 0.136106 (rank : 24) | θ value | θ > 10 (rank : 170) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9Y2H5 | Gene names | PLEKHA6, KIAA0969, PEPP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family A member 6 (Phosphoinositol 3-phosphate-binding protein 3) (PEPP-3). | |||||
|
PLEK_MOUSE
|
||||||
| NC score | 0.133417 (rank : 25) | θ value | θ > 10 (rank : 178) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9JHK5, Q9ERI9 | Gene names | Plek | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin. | |||||
|
PKHA1_HUMAN
|
||||||
| NC score | 0.133234 (rank : 26) | θ value | θ > 10 (rank : 163) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9HB21, Q9BVK0 | Gene names | PLEKHA1, TAPP1 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 1 (Tandem PH domain-containing protein 1) (TAPP-1). | |||||
|
PKHA1_MOUSE
|
||||||
| NC score | 0.132768 (rank : 27) | θ value | θ > 10 (rank : 164) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BUL6, Q8BK96, Q8BXU1, Q8VE13 | Gene names | Plekha1, Tapp1 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 1 (Tandem PH domain-containing protein 1) (TAPP-1). | |||||
|
PLEK_HUMAN
|
||||||
| NC score | 0.130642 (rank : 28) | θ value | θ > 10 (rank : 177) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P08567, Q53SU8, Q6FGM8, Q8WV81 | Gene names | PLEK, P47 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin (Platelet p47 protein). | |||||
|
PKHA6_MOUSE
|
||||||
| NC score | 0.130300 (rank : 29) | θ value | θ > 10 (rank : 171) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 591 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q7TQG1, Q8K0J5 | Gene names | Plekha6, Pepp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 6 (Phosphoinositol 3-phosphate-binding protein 3) (PEPP-3). | |||||
|
PKHA4_MOUSE
|
||||||
| NC score | 0.123373 (rank : 30) | θ value | θ > 10 (rank : 168) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8VC98, Q8R3N3 | Gene names | Plekha4, Pepp1 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 4 (Phosphoinositol 3-phosphate-binding protein 1) (PEPP-1). | |||||
|
PKHA4_HUMAN
|
||||||
| NC score | 0.118900 (rank : 31) | θ value | θ > 10 (rank : 167) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9H4M7, Q8N4M8, Q8N658 | Gene names | PLEKHA4, PEPP1 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 4 (Phosphoinositol 3-phosphate-binding protein 1) (PEPP-1). | |||||
|
PKHA2_MOUSE
|
||||||
| NC score | 0.118445 (rank : 32) | θ value | θ > 10 (rank : 166) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9ERS5, Q8BY29 | Gene names | Plekha2, Tapp2 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 2 (Tandem PH domain-containing protein 2) (TAPP-2) (PH domain-containing adaptor PHAD47). | |||||
|
ANLN_HUMAN
|
||||||
| NC score | 0.115852 (rank : 33) | θ value | θ > 10 (rank : 146) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9NQW6, Q5CZ78, Q6NSK5, Q9H8Y4, Q9NVN9, Q9NVP0 | Gene names | ANLN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Actin-binding protein anillin. | |||||
|
DAPP1_MOUSE
|
||||||
| NC score | 0.114917 (rank : 34) | θ value | θ > 10 (rank : 154) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9QXT1, Q9R178 | Gene names | Dapp1, Bam32 | |||
|
Domain Architecture |

|
|||||
| Description | Dual adapter for phosphotyrosine and 3-phosphotyrosine and 3- phosphoinositide (mDAPP1) (B cell adapter molecule of 32 kDa) (B lymphocyte adapter protein Bam32). | |||||
|
3BP2_MOUSE
|
||||||
| NC score | 0.112613 (rank : 35) | θ value | θ > 10 (rank : 144) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q06649 | Gene names | Sh3bp2, 3bp2 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 domain-binding protein 2 (3BP-2). | |||||
|
3BP2_HUMAN
|
||||||
| NC score | 0.111470 (rank : 36) | θ value | θ > 10 (rank : 143) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P78314, O00500, O15373, P78315 | Gene names | SH3BP2, 3BP2 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 domain-binding protein 2 (3BP-2). | |||||
|
DAPP1_HUMAN
|
||||||
| NC score | 0.109210 (rank : 37) | θ value | θ > 10 (rank : 153) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UN19, Q8TCK5, Q9UHF2 | Gene names | DAPP1, BAM32 | |||
|
Domain Architecture |

|
|||||
| Description | Dual adapter for phosphotyrosine and 3-phosphotyrosine and 3- phosphoinositide (hDAPP1) (B cell adapter molecule of 32 kDa) (B lymphocyte adapter protein Bam32). | |||||
|
PKHA2_HUMAN
|
||||||
| NC score | 0.104431 (rank : 38) | θ value | θ > 10 (rank : 165) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9HB19 | Gene names | PLEKHA2, TAPP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family A member 2 (Tandem PH domain-containing protein 2) (TAPP-2). | |||||
|
PLEK2_MOUSE
|
||||||
| NC score | 0.101095 (rank : 39) | θ value | θ > 10 (rank : 176) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9WV52 | Gene names | Plek2 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin-2. | |||||
|
PLEK2_HUMAN
|
||||||
| NC score | 0.099184 (rank : 40) | θ value | θ > 10 (rank : 175) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NYT0, Q96JT0 | Gene names | PLEK2 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin-2. | |||||
|
CNKR2_MOUSE
|
||||||
| NC score | 0.098038 (rank : 41) | θ value | θ > 10 (rank : 152) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q80YA9, Q80TP2 | Gene names | Cnksr2, Kiaa0902 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Connector enhancer of kinase suppressor of ras 2 (Connector enhancer of KSR2) (CNK2). | |||||
|
CNKR2_HUMAN
|
||||||
| NC score | 0.097952 (rank : 42) | θ value | θ > 10 (rank : 151) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8WXI2, O94976, Q8WXI1 | Gene names | CNKSR2, CNK2, KIAA0902, KSR2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Connector enhancer of kinase suppressor of ras 2 (Connector enhancer of KSR2) (CNK2). | |||||
|
PKHA5_HUMAN
|
||||||
| NC score | 0.096758 (rank : 43) | θ value | θ > 10 (rank : 169) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9HAU0, Q8N3K6, Q96DY9, Q9BVR4, Q9C0H7, Q9H924, Q9NVK8 | Gene names | PLEKHA5, KIAA1686, PEPP2 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 5 (Phosphoinositol 3-phosphate-binding protein 2) (PEPP-2). | |||||
|
ANLN_MOUSE
|
||||||
| NC score | 0.094305 (rank : 44) | θ value | θ > 10 (rank : 147) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 297 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8K298, Q8BL79, Q8BLB3, Q8K2N0, Q9CUY0 | Gene names | Anln | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Actin-binding protein anillin. | |||||
|
RASA1_HUMAN
|
||||||
| NC score | 0.077785 (rank : 45) | θ value | θ > 10 (rank : 179) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P20936 | Gene names | RASA1, RASA | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 1 (GTPase-activating protein) (GAP) (Ras p21 protein activator) (p120GAP) (RasGAP). | |||||
|
RHG25_MOUSE
|
||||||
| NC score | 0.077066 (rank : 46) | θ value | θ > 10 (rank : 181) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 492 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8BYW1, Q2VPR2, Q3TE04, Q3TVX0, Q3UVB7, Q6A0E0, Q8BX98 | Gene names | Arhgap25, Kiaa0053 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 25. | |||||
|
RHG25_HUMAN
|
||||||
| NC score | 0.073744 (rank : 47) | θ value | θ > 10 (rank : 180) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 550 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P42331, Q8IXQ2 | Gene names | ARHGAP25, KIAA0053 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 25. | |||||
|
SWP70_HUMAN
|
||||||
| NC score | 0.068988 (rank : 48) | θ value | θ > 10 (rank : 182) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 973 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9UH65, O75135, Q7LCY6, Q9P061, Q9P0Z8 | Gene names | SWAP70, KIAA0640 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Switch-associated protein 70 (SWAP-70). | |||||
|
CENB2_HUMAN
|
||||||
| NC score | 0.066065 (rank : 49) | θ value | θ > 10 (rank : 148) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q15057, Q9UQR3 | Gene names | CENTB2, KIAA0041 | |||
|
Domain Architecture |

|
|||||
| Description | Centaurin-beta 2 (Cnt-b2). | |||||
|
SWP70_MOUSE
|
||||||
| NC score | 0.060496 (rank : 50) | θ value | θ > 10 (rank : 183) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1097 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6A028, O88443, Q3TQR6, Q3UCA3, Q6P1D0 | Gene names | Swap70, Kiaa0640 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Switch-associated protein 70 (SWAP-70). | |||||
|
CENB5_HUMAN
|
||||||
| NC score | 0.060317 (rank : 51) | θ value | θ > 10 (rank : 149) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q96P50, Q9BSR9, Q9C0E7 | Gene names | CENTB5, KIAA1716 | |||
|
Domain Architecture |

|
|||||
| Description | Centaurin-beta 5 (Cnt-b5). | |||||
|
DYN3_HUMAN
|
||||||
| NC score | 0.058854 (rank : 52) | θ value | θ > 10 (rank : 157) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UQ16, O14982, O95555, Q5W129, Q9H0P3, Q9H548, Q9NQ68, Q9NQN6 | Gene names | DNM3, KIAA0820 | |||
|
Domain Architecture |

|
|||||
| Description | Dynamin-3 (EC 3.6.5.5) (Dynamin, testicular) (T-dynamin). | |||||
|
GNRP_HUMAN
|
||||||
| NC score | 0.056993 (rank : 53) | θ value | θ > 10 (rank : 158) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q13972, Q16027 | Gene names | RASGRF1, CDC25 | |||
|
Domain Architecture |

|
|||||
| Description | Guanine nucleotide-releasing protein (GNRP) (Ras-specific nucleotide exchange factor CDC25) (Ras-specific guanine nucleotide-releasing factor). | |||||
|
CU024_HUMAN
|
||||||
| NC score | 0.056834 (rank : 54) | θ value | 0.813845 (rank : 44) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6XXX2 | Gene names | C21orf24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative uncharacterized protein C21orf24. | |||||
|
DYN2_HUMAN
|
||||||
| NC score | 0.056489 (rank : 55) | θ value | θ > 10 (rank : 155) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P50570, Q7Z5S3, Q9UPH4 | Gene names | DNM2, DYN2 | |||
|
Domain Architecture |

|
|||||
| Description | Dynamin-2 (EC 3.6.5.5). | |||||
|
PKHB2_HUMAN
|
||||||
| NC score | 0.056318 (rank : 56) | θ value | θ > 10 (rank : 173) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q96CS7, Q86W37, Q9BV75, Q9NWK1 | Gene names | PLEKHB2, EVT2 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family B member 2 (Evectin-2). | |||||
|
DYN2_MOUSE
|
||||||
| NC score | 0.055731 (rank : 57) | θ value | θ > 10 (rank : 156) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P39054, Q9DBE1 | Gene names | Dnm2, Dyn2 | |||
|
Domain Architecture |

|
|||||
| Description | Dynamin-2 (EC 3.6.5.5) (Dynamin UDNM). | |||||
|
PKHB2_MOUSE
|
||||||
| NC score | 0.054867 (rank : 58) | θ value | θ > 10 (rank : 174) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9QZC7, Q3UDH6, Q8C4I4, Q8CA27 | Gene names | Plekhb2, Evt2 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family B member 2 (Evectin-2). | |||||
|
AB1IP_HUMAN
|
||||||
| NC score | 0.054722 (rank : 59) | θ value | θ > 10 (rank : 145) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q7Z5R6, Q8IWS8, Q8IZZ7 | Gene names | APBB1IP, PREL1, RARP1, RIAM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 1- interacting protein (APBB1-interacting protein 1) (Rap1-GTP- interacting adapter molecule) (RIAM) (Proline-rich EVH1 ligand 1) (PREL-1) (Proline-rich protein 73) (Retinoic acid-responsive proline- rich protein 1) (RARP-1). | |||||
|
OGFR_HUMAN
|
||||||
| NC score | 0.054628 (rank : 60) | θ value | 0.0113563 (rank : 24) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
GNRP_MOUSE
|
||||||
| NC score | 0.054064 (rank : 61) | θ value | θ > 10 (rank : 159) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P27671 | Gene names | Rasgrf1, Cdc25, Grf1 | |||
|
Domain Architecture |

|
|||||
| Description | Guanine nucleotide-releasing protein (GNRP) (Ras-specific nucleotide exchange factor CDC25) (CDC25Mm). | |||||
|
MTMR5_HUMAN
|
||||||
| NC score | 0.053140 (rank : 62) | θ value | θ > 10 (rank : 160) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O95248, O60228, Q5JXD8, Q5PPM2, Q96GR9, Q9UGB8 | Gene names | SBF1, MTMR5 | |||
|
Domain Architecture |

|
|||||
| Description | SET-binding factor 1 (Sbf1) (Myotubularin-related protein 5). | |||||
|
OSBL8_HUMAN
|
||||||
| NC score | 0.053091 (rank : 63) | θ value | θ > 10 (rank : 161) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9BZF1, Q8WXP8, Q9P277 | Gene names | OSBPL8, KIAA1451, ORP8, OSBP10 | |||
|
Domain Architecture |

|
|||||
| Description | Oxysterol-binding protein-related protein 8 (OSBP-related protein 8) (ORP-8). | |||||
|
TBCD2_HUMAN
|
||||||
| NC score | 0.051916 (rank : 64) | θ value | θ > 10 (rank : 184) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9BYX2 | Gene names | TBC1D2, PARIS1 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 2 (Prostate antigen recognized and identified by SEREX) (PARIS-1). | |||||
|
PKHB1_HUMAN
|
||||||
| NC score | 0.051348 (rank : 65) | θ value | θ > 10 (rank : 172) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UF11, Q9UBF5, Q9UI37, Q9UI44 | Gene names | PLEKHB1, KPL1, PHR1 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family B member 1 (Pleckstrin homology domain retinal protein 1) (PH domain-containing protein in retina 1) (PHRET1). | |||||
|
PHLB1_HUMAN
|
||||||
| NC score | 0.051230 (rank : 66) | θ value | θ > 10 (rank : 162) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1038 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q86UU1, O75133, Q4KMF8, Q8TEQ2 | Gene names | PHLDB1, DLNB07, KIAA0638, LL5A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family B member 1 (Protein LL5-alpha). | |||||
|
K1543_HUMAN
|
||||||
| NC score | 0.049820 (rank : 67) | θ value | 0.813845 (rank : 45) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9P1Y5, Q8NDF1 | Gene names | KIAA1543 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1543. | |||||
|
SRRM1_MOUSE
|
||||||
| NC score | 0.046968 (rank : 68) | θ value | 1.06291 (rank : 53) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
BAT3_MOUSE
|
||||||
| NC score | 0.046232 (rank : 69) | θ value | 1.81305 (rank : 62) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 518 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9Z1R2, Q8SNA3 | Gene names | Bat3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT3 (HLA-B-associated transcript 3). | |||||
|
F100B_MOUSE
|
||||||
| NC score | 0.044286 (rank : 70) | θ value | 1.38821 (rank : 56) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BQH4, Q3UMH5 | Gene names | Fam100b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM100B. | |||||
|
RGS3_MOUSE
|
||||||
| NC score | 0.043998 (rank : 71) | θ value | 0.279714 (rank : 37) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
BAT3_HUMAN
|
||||||
| NC score | 0.041399 (rank : 72) | θ value | 1.81305 (rank : 61) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 490 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P46379, O95874, Q5SQ35, Q5SQ36, Q5SQ37, Q5SRP8, Q96SA6, Q9BCN4 | Gene names | BAT3, G3 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT3 (HLA-B-associated transcript 3) (Protein G3). | |||||
|
RGS3_HUMAN
|
||||||
| NC score | 0.040852 (rank : 73) | θ value | 0.0961366 (rank : 28) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
MKL1_MOUSE
|
||||||
| NC score | 0.040671 (rank : 74) | θ value | 0.813845 (rank : 46) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 667 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8K4J6 | Gene names | Mkl1, Bsac | |||
|
Domain Architecture |

|
|||||
| Description | MKL/myocardin-like protein 1 (Myocardin-related transcription factor A) (MRTF-A) (Megakaryoblastic leukemia 1 protein homolog) (Basic SAP coiled-coil transcription activator). | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.040095 (rank : 75) | θ value | 0.0736092 (rank : 27) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
TARA_HUMAN
|
||||||
| NC score | 0.039278 (rank : 76) | θ value | 8.99809 (rank : 139) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q9H2D6, O94797, Q2PZW8, Q2Q3Z9, Q2Q400, Q5R3M6, Q96DW1, Q9BT77, Q9BTL7, Q9BY98, Q9Y3L4 | Gene names | TRIOBP, KIAA1662, TARA | |||
|
Domain Architecture |

|
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
K1802_MOUSE
|
||||||
| NC score | 0.038835 (rank : 77) | θ value | 1.81305 (rank : 65) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
SRRM1_HUMAN
|
||||||
| NC score | 0.037382 (rank : 78) | θ value | 5.27518 (rank : 101) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
MAGE1_HUMAN
|
||||||
| NC score | 0.036114 (rank : 79) | θ value | 0.163984 (rank : 34) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
GRIN1_HUMAN
|
||||||
| NC score | 0.036052 (rank : 80) | θ value | 0.365318 (rank : 38) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q7Z2K8, Q8ND74, Q96PZ4 | Gene names | GPRIN1, KIAA1893 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
BSN_MOUSE
|
||||||
| NC score | 0.035882 (rank : 81) | θ value | 6.88961 (rank : 106) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
MILK1_MOUSE
|
||||||
| NC score | 0.035013 (rank : 82) | θ value | 2.36792 (rank : 71) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 788 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8BGT6, Q8BJ60 | Gene names | Mirab13, D15Mit260, Kiaa1668 | |||
|
Domain Architecture |

|
|||||
| Description | Molecule interacting with Rab13 (MIRab13) (MICAL-like protein 1). | |||||
|
K1802_HUMAN
|
||||||
| NC score | 0.034763 (rank : 83) | θ value | 4.03905 (rank : 91) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1378 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q96JM3, Q6P181, Q8NC88, Q9BST0 | Gene names | KIAA1802, C13orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
TM108_MOUSE
|
||||||
| NC score | 0.034632 (rank : 84) | θ value | 1.38821 (rank : 59) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 336 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8BHE4, Q80WR9 | Gene names | Tmem108 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 108 precursor. | |||||
|
MLL3_HUMAN
|
||||||
| NC score | 0.033915 (rank : 85) | θ value | 1.81305 (rank : 66) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
TE2IP_MOUSE
|
||||||
| NC score | 0.033397 (rank : 86) | θ value | 2.36792 (rank : 77) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q91VL8, Q543F4, Q9JJE8, Q9JJE9 | Gene names | Terf2ip, Rap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Telomeric repeat-binding factor 2-interacting protein 1 (TRF2- interacting telomeric protein Rap1). | |||||
|
TAU_HUMAN
|
||||||
| NC score | 0.033358 (rank : 87) | θ value | 1.81305 (rank : 68) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 448 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P10636, P18518, Q14799, Q15549, Q15550, Q15551, Q9UDJ3, Q9UMH0, Q9UQ96 | Gene names | MAPT, MTBT1, TAU | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein tau (Neurofibrillary tangle protein) (Paired helical filament-tau) (PHF-tau). | |||||
|
TE2IP_HUMAN
|
||||||
| NC score | 0.032799 (rank : 88) | θ value | 2.36792 (rank : 76) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NYB0, Q8WYZ3, Q9NWR2 | Gene names | TERF2IP, RAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Telomeric repeat-binding factor 2-interacting protein 1 (TRF2- interacting telomeric protein Rap1) (hRap1). | |||||
|
ERCC5_MOUSE
|
||||||
| NC score | 0.032716 (rank : 89) | θ value | 1.06291 (rank : 50) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P35689, Q61528, Q64248 | Gene names | Ercc5, Ercc-5, Xpg | |||
|
Domain Architecture |

|
|||||
| Description | DNA-repair protein complementing XP-G cells homolog (Xeroderma pigmentosum group G-complementing protein homolog) (DNA excision repair protein ERCC-5). | |||||
|
BCL9_HUMAN
|
||||||
| NC score | 0.032502 (rank : 90) | θ value | 8.99809 (rank : 123) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1008 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O00512 | Gene names | BCL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9) (Legless homolog). | |||||
|
CD248_HUMAN
|
||||||
| NC score | 0.031794 (rank : 91) | θ value | 0.21417 (rank : 35) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 679 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9HCU0, Q3SX55, Q96KB6 | Gene names | CD248, CD164L1, TEM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endosialin precursor (Tumor endothelial marker 1) (CD248 antigen). | |||||
|
SC24C_HUMAN
|
||||||
| NC score | 0.031476 (rank : 92) | θ value | 1.38821 (rank : 58) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 662 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P53992, Q8WV25 | Gene names | SEC24C, KIAA0079 | |||
|
Domain Architecture |

|
|||||
| Description | Protein transport protein Sec24C (SEC24-related protein C). | |||||
|
BAG3_MOUSE
|
||||||
| NC score | 0.030973 (rank : 93) | θ value | 8.99809 (rank : 122) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9JLV1, Q9JJC7 | Gene names | Bag3, Bis | |||
|
Domain Architecture |

|
|||||
| Description | BAG family molecular chaperone regulator 3 (BCL-2-binding athanogene- 3) (BAG-3) (Bcl-2-binding protein Bis). | |||||
|
MINT_MOUSE
|
||||||
| NC score | 0.029728 (rank : 94) | θ value | 0.62314 (rank : 41) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
MFAP1_MOUSE
|
||||||
| NC score | 0.029688 (rank : 95) | θ value | 0.62314 (rank : 40) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 353 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9CQU1, Q3TU29, Q8CCL1, Q9CSJ5 | Gene names | Mfap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microfibrillar-associated protein 1. | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.029002 (rank : 96) | θ value | 8.99809 (rank : 136) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
MDC1_MOUSE
|
||||||
| NC score | 0.027328 (rank : 97) | θ value | 5.27518 (rank : 98) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
TSH3_MOUSE
|
||||||
| NC score | 0.026822 (rank : 98) | θ value | 0.125558 (rank : 31) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 590 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8CGV9, Q5DTX4 | Gene names | Tshz3, Kiaa1474, Tsh3, Zfp537, Znf537 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Teashirt homolog 3 (Zinc finger protein 537). | |||||
|
PI5PA_MOUSE
|
||||||
| NC score | 0.026685 (rank : 99) | θ value | 6.88961 (rank : 115) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P59644 | Gene names | Pib5pa | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 4,5-bisphosphate 5-phosphatase A (EC 3.1.3.56). | |||||
|
FOG1_MOUSE
|
||||||
| NC score | 0.026538 (rank : 100) | θ value | 0.00298849 (rank : 23) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1237 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O35615 | Gene names | Zfpm1, Fog, Fog1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein ZFPM1 (Zinc finger protein multitype 1) (Friend of GATA protein 1) (Friend of GATA-1) (FOG-1). | |||||
|
B4GT1_HUMAN
|
||||||
| NC score | 0.026154 (rank : 101) | θ value | 0.0736092 (rank : 26) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P15291, Q12909, Q12910, Q12911, Q14456, Q14509, Q14523 | Gene names | B4GALT1, GGTB2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-1,4-galactosyltransferase 1 (EC 2.4.1.-) (Beta-1,4-GalTase 1) (Beta4Gal-T1) (b4Gal-T1) (UDP-galactose:beta-N-acetylglucosamine beta- 1,4-galactosyltransferase 1) (UDP-Gal:beta-GlcNAc beta-1,4- galactosyltransferase 1) [Includes: Lactose synthase A protein (EC 2.4.1.22); N-acetyllactosamine synthase (EC 2.4.1.90) (Nal synthetase); Beta-N-acetylglucosaminylglycopeptide beta-1,4- galactosyltransferase (EC 2.4.1.38); Beta-N-acetylglucosaminyl- glycolipid beta-1,4-galactosyltransferase (EC 2.4.1.-)]. | |||||
|
NFAC1_HUMAN
|
||||||
| NC score | 0.025484 (rank : 102) | θ value | 0.62314 (rank : 42) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O95644, Q12865, Q15793 | Gene names | NFATC1, NFAT2, NFATC | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor of activated T-cells, cytoplasmic 1 (NFAT transcription complex cytosolic component) (NF-ATc1) (NF-ATc). | |||||
|
SMOO_HUMAN
|
||||||
| NC score | 0.025143 (rank : 103) | θ value | 6.88961 (rank : 120) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 391 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P53814, O00569, O95769, O95937, Q9P1S8, Q9UIT1, Q9UIT2 | Gene names | SMTN, SMSMO | |||
|
Domain Architecture |

|
|||||
| Description | Smoothelin. | |||||
|
ETV6_HUMAN
|
||||||
| NC score | 0.025014 (rank : 104) | θ value | 0.279714 (rank : 36) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P41212, Q9UMF6 | Gene names | ETV6, TEL, TEL1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor ETV6 (ETS-related protein Tel1) (Tel) (ETS translocation variant 6). | |||||
|
PCLO_MOUSE
|
||||||
| NC score | 0.024870 (rank : 105) | θ value | 8.99809 (rank : 133) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
SEPT9_HUMAN
|
||||||
| NC score | 0.024503 (rank : 106) | θ value | 0.813845 (rank : 47) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 706 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9UHD8, Q96QF3, Q96QF4, Q96QF5, Q9HA04, Q9UG40, Q9Y5W4 | Gene names | SEPT9, KIAA0991, MSF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Septin-9 (MLL septin-like fusion protein) (MLL septin-like fusion protein MSF-A) (Ovarian/Breast septin) (Ov/Br septin) (Septin D1). | |||||
|
ARI1A_HUMAN
|
||||||
| NC score | 0.023526 (rank : 107) | θ value | 6.88961 (rank : 103) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 787 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O14497, Q53FK9, Q5T0W1, Q5T0W2, Q5T0W3, Q8NFD6, Q96T89, Q9BY33, Q9HBJ5, Q9UPZ1 | Gene names | ARID1A, C1orf4, OSA1, SMARCF1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 1A (ARID domain- containing protein 1A) (SWI/SNF-related, matrix-associated, actin- dependent regulator of chromatin subfamily F member 1) (SWI-SNF complex protein p270) (B120) (SWI-like protein) (Osa homolog 1) (hOSA1) (hELD) (BRG1-associated factor 250) (BAF250) (BRG1-associated factor 250a) (BAF250A). | |||||
|
CUTL1_MOUSE
|
||||||
| NC score | 0.023492 (rank : 108) | θ value | 0.0252991 (rank : 25) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P53564, O08994, P70301, Q571L6, Q91ZD2 | Gene names | Cutl1, Cux, Cux1, Kiaa4047 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP) (Homeobox protein Cux). | |||||
|
PER3_HUMAN
|
||||||
| NC score | 0.023353 (rank : 109) | θ value | 2.36792 (rank : 74) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P56645, Q5H8X4, Q969K6, Q96S77, Q96S78, Q9C0J3, Q9NSP9, Q9UGU8 | Gene names | PER3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (hPER3). | |||||
|
CC28B_MOUSE
|
||||||
| NC score | 0.023291 (rank : 110) | θ value | 5.27518 (rank : 94) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CEG5, Q8K132 | Gene names | Ccdc28b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 28B. | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.023229 (rank : 111) | θ value | 1.81305 (rank : 64) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
CSKI1_MOUSE
|
||||||
| NC score | 0.022954 (rank : 112) | θ value | 0.163984 (rank : 32) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1031 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q6P9K8, Q6ZPU2, Q8BWU2, Q8BX99, Q9CXH0 | Gene names | Caskin1, Kiaa1306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-1 (CASK-interacting protein 1). | |||||
|
OSBL2_HUMAN
|
||||||
| NC score | 0.022951 (rank : 113) | θ value | 0.125558 (rank : 30) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H1P3, Q9BZB1, Q9Y4B8 | Gene names | OSBPL2, KIAA0772, ORP2 | |||
|
Domain Architecture |

|
|||||
| Description | Oxysterol-binding protein-related protein 2 (OSBP-related protein 2) (ORP-2). | |||||
|
CC28B_HUMAN
|
||||||
| NC score | 0.022895 (rank : 114) | θ value | 5.27518 (rank : 93) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BUN5, Q8TBV8 | Gene names | CCDC28B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 28B. | |||||
|
P66A_MOUSE
|
||||||
| NC score | 0.022293 (rank : 115) | θ value | 1.81305 (rank : 67) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CHY6, Q8BTQ2, Q8VEC9 | Gene names | Gatad2a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcriptional repressor p66 alpha (GATA zinc finger domain- containing protein 2A). | |||||
|
CF010_HUMAN
|
||||||
| NC score | 0.021378 (rank : 116) | θ value | 2.36792 (rank : 70) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q5SRN2, Q5SPI9, Q5SPJ0, Q5SPK9, Q5SPL0, Q5SRN3, Q5TG25, Q5TG26, Q8N4B6 | Gene names | C6orf10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C6orf10. | |||||
|
BAZ2A_MOUSE
|
||||||
| NC score | 0.021124 (rank : 117) | θ value | 1.38821 (rank : 54) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q91YE5 | Gene names | Baz2a, Tip5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5). | |||||
|
MFAP1_HUMAN
|
||||||
| NC score | 0.020369 (rank : 118) | θ value | 8.99809 (rank : 131) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P55081 | Gene names | MFAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Microfibrillar-associated protein 1. | |||||
|
CD248_MOUSE
|
||||||
| NC score | 0.019676 (rank : 119) | θ value | 6.88961 (rank : 107) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 578 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q91V98, Q3UMV6, Q91ZV1 | Gene names | Cd248, Tem1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endosialin precursor (Tumor endothelial marker 1) (CD248 antigen). | |||||
|
MAP1A_HUMAN
|
||||||
| NC score | 0.019099 (rank : 120) | θ value | 6.88961 (rank : 112) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P78559, O95643, Q12973, Q15882, Q9UJT4 | Gene names | MAP1A | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 1A (MAP 1A) (Proliferation-related protein p80) [Contains: MAP1 light chain LC2]. | |||||
|
NALDL_MOUSE
|
||||||
| NC score | 0.018471 (rank : 121) | θ value | 1.06291 (rank : 52) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7M758 | Gene names | Naaladl1, Naaladasel | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylated-alpha-linked acidic dipeptidase-like protein (EC 3.4.17.21) (NAALADase L). | |||||
|
LRCH2_MOUSE
|
||||||
| NC score | 0.018317 (rank : 122) | θ value | 0.163984 (rank : 33) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q3UMG5, Q7TNP9, Q80TC9 | Gene names | Lrch2, Kiaa1495 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 2. | |||||
|
SIX4_MOUSE
|
||||||
| NC score | 0.018071 (rank : 123) | θ value | 0.62314 (rank : 43) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61321, Q61322, Q61323 | Gene names | Six4, Arec3 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein SIX4 (Sine oculis homeobox homolog 4) (Skeletal muscle-specific ARE-binding protein AREC3). | |||||
|
CTND2_HUMAN
|
||||||
| NC score | 0.017977 (rank : 124) | θ value | 3.0926 (rank : 80) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9UQB3, O00379, O15390, O43206, O43840, Q13589, Q9UM66, Q9UPM3 | Gene names | CTNND2, NPRAP | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-2 (Delta-catenin) (Neural plakophilin-related ARM-repeat protein) (NPRAP) (Neurojungin) (GT24). | |||||
|
MTSS1_HUMAN
|
||||||
| NC score | 0.017868 (rank : 125) | θ value | 6.88961 (rank : 113) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O43312, Q8TCA2, Q96RX2 | Gene names | MTSS1, KIAA0429, MIM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metastasis suppressor protein 1 (Missing in metastasis protein) (Metastasis suppressor YGL-1). | |||||
|
ENPL_HUMAN
|
||||||
| NC score | 0.017613 (rank : 126) | θ value | 1.06291 (rank : 49) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P14625, Q96A97 | Gene names | HSP90B1, TRA1 | |||
|
Domain Architecture |

|
|||||
| Description | Endoplasmin precursor (Heat shock protein 90 kDa beta member 1) (94 kDa glucose-regulated protein) (GRP94) (gp96 homolog) (Tumor rejection antigen 1). | |||||
|
AL2S8_MOUSE
|
||||||
| NC score | 0.017551 (rank : 127) | θ value | 4.03905 (rank : 87) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8VHI4 | Gene names | Als2cr8, Carf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyotrophic lateral sclerosis 2 chromosomal region candidate gene 8 protein homolog (Calcium-response factor) (CaRF). | |||||
|
NF2L1_HUMAN
|
||||||
| NC score | 0.016998 (rank : 128) | θ value | 1.38821 (rank : 57) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q14494, Q12877, Q96FN6 | Gene names | NFE2L1, HBZ17, NRF1, TCF11 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor erythroid 2-related factor 1 (NF-E2-related factor 1) (NFE2-related factor 1) (Nuclear factor, erythroid derived 2, like 1) (Transcription factor 11) (Transcription factor HBZ17) (Transcription factor LCR-F1) (Locus control region-factor 1). | |||||
|
NPAS4_MOUSE
|
||||||
| NC score | 0.016984 (rank : 129) | θ value | 3.0926 (rank : 85) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8BGD7, Q3V3U3 | Gene names | Npas4, Nxf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF) (Limbic-enhanced PAS protein) (LE-PAS). | |||||
|
RT28_HUMAN
|
||||||
| NC score | 0.016944 (rank : 130) | θ value | 6.88961 (rank : 118) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 4 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y2Q9, Q96Q21 | Gene names | MRPS28, MRPS35 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitochondrial 28S ribosomal protein S28 (S28mt) (MRP-S28) (MRP-S35). | |||||
|
DMRTD_HUMAN
|
||||||
| NC score | 0.016512 (rank : 131) | θ value | 6.88961 (rank : 109) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8IXT2, Q8N6Q2, Q96M39, Q96SD4 | Gene names | DMRTC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Doublesex- and mab-3-related transcription factor C2. | |||||
|
CEBPE_HUMAN
|
||||||
| NC score | 0.016027 (rank : 132) | θ value | 3.0926 (rank : 79) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q15744, Q15745, Q8IYI2, Q99803 | Gene names | CEBPE | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein epsilon (C/EBP epsilon). | |||||
|
TS13_MOUSE
|
||||||
| NC score | 0.015809 (rank : 133) | θ value | 8.99809 (rank : 140) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q01755 | Gene names | Tcp11 | |||
|
Domain Architecture |

|
|||||
| Description | Testis-specific protein PBS13. | |||||
|
FRS3_MOUSE
|
||||||
| NC score | 0.015806 (rank : 134) | θ value | 3.0926 (rank : 82) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q91WJ0 | Gene names | Frs3, Frs2b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fibroblast growth factor receptor substrate 3 (FGFR substrate 3) (FRS2-beta). | |||||
|
NADE1_MOUSE
|
||||||
| NC score | 0.014727 (rank : 135) | θ value | 5.27518 (rank : 99) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q711T7, Q711P6, Q8CFY6, Q9D280 | Gene names | Nadsyn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamine-dependent NAD(+) synthetase (EC 6.3.5.1) (NAD(+) synthase [glutamine-hydrolyzing]) (NAD(+) synthetase 1) (NH3-dependent NAD(+) synthetase-like protein). | |||||
|
RC3H1_HUMAN
|
||||||
| NC score | 0.014310 (rank : 136) | θ value | 2.36792 (rank : 75) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5TC82, Q5W180, Q5W181, Q8IVE6, Q8N9V1 | Gene names | RC3H1, KIAA2025 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Roquin (RING finger and C3H zinc finger protein 1). | |||||
|
CUTL1_HUMAN
|
||||||
| NC score | 0.014203 (rank : 137) | θ value | 8.99809 (rank : 124) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1574 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P39880, Q6NYH4, Q9UEV5 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP). | |||||
|
ZC3H6_HUMAN
|
||||||
| NC score | 0.014104 (rank : 138) | θ value | 5.27518 (rank : 102) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P61129 | Gene names | ZC3H6, KIAA2035, ZC3HDC6 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger CCCH domain-containing protein 6. | |||||
|
DOK3_MOUSE
|
||||||
| NC score | 0.013568 (rank : 139) | θ value | 5.27518 (rank : 95) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9QZK7 | Gene names | Dok3, Dokl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Docking protein 3 (Downstream of tyrosine kinase 3) (P62(dok)-like protein) (DOK-L). | |||||
|
LPP_HUMAN
|
||||||
| NC score | 0.012934 (rank : 140) | θ value | 8.99809 (rank : 127) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 775 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q93052, Q8NFX5 | Gene names | LPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipoma-preferred partner (LIM domain-containing preferred translocation partner in lipoma). | |||||
|
ARI4A_HUMAN
|
||||||
| NC score | 0.012693 (rank : 141) | θ value | 8.99809 (rank : 121) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 434 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P29374, Q15991, Q15992, Q15993 | Gene names | ARID4A, RBBP1, RBP1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 4A (ARID domain- containing protein 4A) (Retinoblastoma-binding protein 1) (RBBP-1). | |||||
|
HAP1_HUMAN
|
||||||
| NC score | 0.012402 (rank : 142) | θ value | 3.0926 (rank : 83) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 606 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P54257, O75358, Q9H4G3, Q9HA98, Q9NY90 | Gene names | HAP1, HLP1 | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin-associated protein 1 (HAP-1) (Neuroan 1). | |||||
|
HDGR3_HUMAN
|
||||||
| NC score | 0.012296 (rank : 143) | θ value | 5.27518 (rank : 96) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y3E1 | Gene names | HDGFRP3, HDGF2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatoma-derived growth factor-related protein 3 (HRP-3) (Hepatoma- derived growth factor 2). | |||||
|
SYF2_MOUSE
|
||||||
| NC score | 0.012199 (rank : 144) | θ value | 8.99809 (rank : 137) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D198 | Gene names | Syf2, Cbpin, Gcipip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pre-mRNA-splicing factor SYF2 (CCNDBP1-interactor) (p29) (mp29). | |||||
|
FOXM1_HUMAN
|
||||||
| NC score | 0.011562 (rank : 145) | θ value | 3.0926 (rank : 81) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q08050, O43258, O43259, O43260, Q4ZGG7, Q9BRL2 | Gene names | FOXM1, FKHL16, HFH11, MPP2, WIN | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein M1 (Forkhead-related protein FKHL16) (Hepatocyte nuclear factor 3 forkhead homolog 11) (HNF-3/fork-head homolog 11) (HFH-11) (Winged helix factor from INS-1 cells) (M-phase phosphoprotein 2) (MPM-2 reactive phosphoprotein 2) (Transcription factor Trident). | |||||
|
S3TC2_MOUSE
|
||||||
| NC score | 0.011555 (rank : 146) | θ value | 5.27518 (rank : 100) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80VA5, Q8BWH2, Q8BXB8, Q8C0G6 | Gene names | Sh3tc2, Kiaa1985 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 domain and tetratricopeptide repeats-containing protein 2. | |||||
|
LZTS1_MOUSE
|
||||||
| NC score | 0.011540 (rank : 147) | θ value | 5.27518 (rank : 97) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 540 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P60853 | Gene names | Lzts1, Fez1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine zipper putative tumor suppressor 1 (F37/Esophageal cancer- related gene-coding leucine-zipper motif) (Fez1). | |||||
|
GLU2B_HUMAN
|
||||||
| NC score | 0.011470 (rank : 148) | θ value | 6.88961 (rank : 111) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 377 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P14314, Q96BU9, Q96D06, Q9P0W9 | Gene names | PRKCSH, G19P1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glucosidase 2 subunit beta precursor (Glucosidase II subunit beta) (Protein kinase C substrate, 60.1 kDa protein, heavy chain) (PKCSH) (80K-H protein). | |||||
|
INOC1_MOUSE
|
||||||
| NC score | 0.010898 (rank : 149) | θ value | 4.03905 (rank : 90) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6ZPV2, Q6P7V0, Q6PCP1, Q8C9T7 | Gene names | Inoc1, Kiaa1259 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative DNA helicase INO80 complex homolog 1 (EC 3.6.1.-). | |||||
|
INOC1_HUMAN
|
||||||
| NC score | 0.010801 (rank : 150) | θ value | 4.03905 (rank : 89) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9ULG1, Q9NTG6 | Gene names | INOC1, KIAA1259 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative DNA helicase INO80 complex homolog 1 (EC 3.6.1.-) (hINO80). | |||||
|
TACC2_MOUSE
|
||||||
| NC score | 0.010673 (rank : 151) | θ value | 8.99809 (rank : 138) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 774 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9JJG0 | Gene names | Tacc2 | |||
|
Domain Architecture |

|
|||||
| Description | Transforming acidic coiled-coil-containing protein 2. | |||||
|
FGD3_HUMAN
|
||||||
| NC score | 0.010288 (rank : 152) | θ value | 1.06291 (rank : 51) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5JSP0, Q7Z7D9, Q8N5G1 | Gene names | FGD3, ZFYVE5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 3 (Zinc finger FYVE domain-containing protein 5). | |||||
|
LRCH2_HUMAN
|
||||||
| NC score | 0.009987 (rank : 153) | θ value | 8.99809 (rank : 128) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5VUJ6, Q9HA88, Q9P233 | Gene names | LRCH2, KIAA1495 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats and calponin homology domain-containing protein 2. | |||||
|
SGOL2_MOUSE
|
||||||
| NC score | 0.009858 (rank : 154) | θ value | 8.99809 (rank : 135) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7TSY8, Q811I4, Q8C9C4, Q8CGJ4, Q9CS55 | Gene names | Sgol2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Shugoshin-like 2. | |||||
|
BRDT_MOUSE
|
||||||
| NC score | 0.009394 (rank : 155) | θ value | 6.88961 (rank : 105) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q91Y44, Q59HJ4 | Gene names | Brdt, Fsrg3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain testis-specific protein (RING3-like protein) (Bromodomain- containing female sterile homeotic-like protein). | |||||
|
NKTR_HUMAN
|
||||||
| NC score | 0.009390 (rank : 156) | θ value | 4.03905 (rank : 92) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 658 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P30414 | Gene names | NKTR | |||
|
Domain Architecture |

|
|||||
| Description | NK-tumor recognition protein (Natural-killer cells cyclophilin-related protein) (NK-TR protein). | |||||
|
BCN1_HUMAN
|
||||||
| NC score | 0.009276 (rank : 157) | θ value | 6.88961 (rank : 104) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q14457, O75595, Q9UNA8 | Gene names | BECN1, GT197 | |||
|
Domain Architecture |

|
|||||
| Description | Beclin-1 (Coiled-coil myosin-like BCL2-interacting protein) (Protein GT197). | |||||
|
GNAS1_HUMAN
|
||||||
| NC score | 0.008930 (rank : 158) | θ value | 8.99809 (rank : 126) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q5JWF2, O75684, O75685, Q5JW67, Q5JWF1, Q9NY42 | Gene names | GNAS, GNAS1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Guanine nucleotide-binding protein G(s) subunit alpha isoforms XLas (Adenylate cyclase-stimulating G alpha protein) (Extra large alphas protein) (XLalphas). | |||||
|
BPAEA_HUMAN
|
||||||
| NC score | 0.008850 (rank : 159) | θ value | 1.38821 (rank : 55) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1369 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O94833, Q5TBT0, Q8N1T8, Q8N8J3, Q8WXK9, Q96AK9, Q96DQ5, Q9H555 | Gene names | DST, BPAG1, DMH, DT, KIAA0728 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 6/9/10 (Trabeculin-beta) (Bullous pemphigoid antigen) (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
MEF2D_HUMAN
|
||||||
| NC score | 0.008740 (rank : 160) | θ value | 8.99809 (rank : 130) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q14814, Q14815 | Gene names | MEF2D | |||
|
Domain Architecture |

|
|||||
| Description | Myocyte-specific enhancer factor 2D. | |||||
|
DLGP1_HUMAN
|
||||||
| NC score | 0.008034 (rank : 161) | θ value | 6.88961 (rank : 108) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O14490, O14489, P78335 | Gene names | DLGAP1, DAP1, GKAP | |||
|
Domain Architecture |

|
|||||
| Description | Disks large-associated protein 1 (DAP-1) (Guanylate kinase-associated protein) (hGKAP) (SAP90/PSD-95-associated protein 1) (SAPAP1) (PSD- 95/SAP90-binding protein 1). | |||||
|
DDX51_HUMAN
|
||||||
| NC score | 0.008022 (rank : 162) | θ value | 8.99809 (rank : 125) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8N8A6, Q5CZ71, Q8IXK5, Q96ED1 | Gene names | DDX51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX51 (EC 3.6.1.-) (DEAD box protein 51). | |||||
|
CAC1B_MOUSE
|
||||||
| NC score | 0.008006 (rank : 163) | θ value | 3.0926 (rank : 78) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O55017, Q60609 | Gene names | Cacna1b, Cach5, Cacnl1a5, Cchn1a | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent N-type calcium channel subunit alpha-1B (Voltage- gated calcium channel subunit alpha Cav2.2) (Calcium channel, L type, alpha-1 polypeptide isoform 5) (Brain calcium channel III) (BIII). | |||||
|
YTHD2_HUMAN
|
||||||
| NC score | 0.007698 (rank : 164) | θ value | 8.99809 (rank : 141) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y5A9, Q5VSZ9, Q8TDH0, Q9BUJ5 | Gene names | YTHDF2, HGRG8 | |||
|
Domain Architecture |

|
|||||
| Description | YTH domain family protein 2 (High-glucose-regulated protein 8) (NY- REN-2 antigen) (CLL-associated antigen KW-14). | |||||
|
PRKDC_HUMAN
|
||||||
| NC score | 0.007673 (rank : 165) | θ value | 6.88961 (rank : 116) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P78527, P78528, Q13327, Q13337, Q14175, Q59H99, Q7Z611, Q96SE6, Q9UME3 | Gene names | PRKDC | |||
|
Domain Architecture |

|
|||||
| Description | DNA-dependent protein kinase catalytic subunit (EC 2.7.11.1) (DNA-PK catalytic subunit) (DNA-PKcs) (DNPK1) (p460). | |||||
|
SFRS4_HUMAN
|
||||||
| NC score | 0.007389 (rank : 166) | θ value | 6.88961 (rank : 119) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q08170, Q5VXP1, Q9BUA4, Q9UEB5 | Gene names | SFRS4, SRP75 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 4 (Pre-mRNA-splicing factor SRP75) (SRP001LB). | |||||
|
BPA1_MOUSE
|
||||||
| NC score | 0.006821 (rank : 167) | θ value | 4.03905 (rank : 88) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q91ZU6, Q91ZU7 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 1/2/3/4 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
RNPL1_HUMAN
|
||||||
| NC score | 0.006540 (rank : 168) | θ value | 8.99809 (rank : 134) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9HAU8, Q96AC9, Q9H033, Q9NVD0 | Gene names | RNPEPL1 | |||
|
Domain Architecture |

|
|||||
| Description | Arginyl aminopeptidase-like 1 (EC 3.4.11.-) (RNPEP-like protein). | |||||
|
SM1L2_HUMAN
|
||||||
| NC score | 0.006496 (rank : 169) | θ value | 3.0926 (rank : 86) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1019 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8NDV3, Q5TIC3, Q9Y3G5 | Gene names | SMC1L2 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 2 protein (SMC1beta protein). | |||||
|
MUG2_MOUSE
|
||||||
| NC score | 0.006239 (rank : 170) | θ value | 6.88961 (rank : 114) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P28666 | Gene names | Mug2, Mug-2 | |||
|
Domain Architecture |

|
|||||
| Description | Murinoglobulin-2 precursor (MuG2). | |||||
|
RBY1A_HUMAN
|
||||||
| NC score | 0.005544 (rank : 171) | θ value | 6.88961 (rank : 117) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15414, Q15377, Q8NHR0 | Gene names | RBMY1A1, RBM1, YRRM1 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding motif protein, Y chromosome, family 1 member A1 (RNA- binding motif protein 1). | |||||
|
ZBT17_MOUSE
|
||||||
| NC score | 0.005203 (rank : 172) | θ value | 0.0961366 (rank : 29) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 940 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q60821, Q60699 | Gene names | Zbtb17, Zfp100, Znf151 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 17 (Zinc finger protein 151) (Zinc finger protein 100) (Zfp-100) (Polyomavirus late initiator promoter-binding protein) (LP-1) (Zinc finger protein Z13). | |||||
|
MCP_MOUSE
|
||||||
| NC score | 0.004787 (rank : 173) | θ value | 8.99809 (rank : 129) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O88174, Q9R0R9 | Gene names | Cd46, Mcp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane cofactor protein precursor (CD46 antigen). | |||||
|
PARD3_HUMAN
|
||||||
| NC score | 0.004321 (rank : 174) | θ value | 8.99809 (rank : 132) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8TEW0, Q8TEW1, Q8TEW2, Q8TEW3, Q96K28, Q96RM6, Q96RM7, Q9BY57, Q9BY58, Q9HC48, Q9NWL4, Q9NYE6 | Gene names | PARD3, PAR3, PAR3A | |||
|
Domain Architecture |

|
|||||
| Description | Partitioning-defective 3 homolog (PARD-3) (PAR-3) (Atypical PKC isotype-specific-interacting protein) (ASIP) (CTCL tumor antigen se2- 5) (PAR3-alpha). | |||||
|
ZBT17_HUMAN
|
||||||
| NC score | 0.004318 (rank : 175) | θ value | 0.813845 (rank : 48) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 925 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q13105, Q15932, Q5JYB2, Q9NUC9 | Gene names | ZBTB17, MIZ1, ZNF151 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 17 (Zinc finger protein 151) (Myc-interacting zinc finger protein) (Miz-1 protein). | |||||
|
FOXC2_MOUSE
|
||||||
| NC score | 0.004000 (rank : 176) | θ value | 6.88961 (rank : 110) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q61850, P97948, Q63869, Q8C694 | Gene names | Foxc2, Fkh14, Fkhl14, Mfh1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein C2 (Forkhead-related protein FKHL14) (Mesenchyme fork head protein 1) (MFH-1 protein) (Transcription factor FKH-14). | |||||
|
BCL6_HUMAN
|
||||||
| NC score | 0.003920 (rank : 177) | θ value | 1.81305 (rank : 63) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 922 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P41182 | Gene names | BCL6, BCL5, LAZ3, ZBTB27, ZNF51 | |||
|
Domain Architecture |

|
|||||
| Description | B-cell lymphoma 6 protein (BCL-6) (Zinc finger protein 51) (LAZ-3 protein) (BCL-5) (Zinc finger and BTB domain-containing protein 27). | |||||
|
ZN688_HUMAN
|
||||||
| NC score | 0.003738 (rank : 178) | θ value | 1.81305 (rank : 69) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 873 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8WV14, O75701, Q8IW91, Q96MN0 | Gene names | ZNF688 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 688. | |||||
|
ZN509_MOUSE
|
||||||
| NC score | 0.003700 (rank : 179) | θ value | 1.38821 (rank : 60) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 990 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8BXX2, Q8CID4, Q8K2Z6 | Gene names | Znf509, Zfp509 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 509. | |||||
|
ZN667_HUMAN
|
||||||
| NC score | 0.002917 (rank : 180) | θ value | 0.47712 (rank : 39) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 773 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5HYK9, Q6B093, Q9H807 | Gene names | ZNF667 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 667. | |||||
|
PCDA5_HUMAN
|
||||||
| NC score | 0.002000 (rank : 181) | θ value | 2.36792 (rank : 73) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y5H7, O75284, Q8N4R3 | Gene names | PCDHA5 | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin alpha 5 precursor (PCDH-alpha5). | |||||
|
ZN621_HUMAN
|
||||||
| NC score | 0.001567 (rank : 182) | θ value | 8.99809 (rank : 142) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 737 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6ZSS3, Q8TE91 | Gene names | ZNF621 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 621. | |||||
|
MK15_HUMAN
|
||||||
| NC score | -0.001318 (rank : 183) | θ value | 2.36792 (rank : 72) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 1038 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8TD08, Q2TCF9, Q8N362 | Gene names | MAPK15, ERK8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase 15 (EC 2.7.11.24) (Extracellular signal-regulated kinase 8). | |||||
|
KPCL_HUMAN
|
||||||
| NC score | -0.002588 (rank : 184) | θ value | 3.0926 (rank : 84) | |||
| Query Neighborhood Hits | 142 | Target Neighborhood Hits | 887 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P24723, Q16246 | Gene names | PRKCH, PKCL | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C eta type (EC 2.7.11.13) (nPKC-eta) (PKC-L). | |||||