Please be patient as the page loads
|
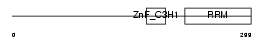
TE2IP_HUMAN
|
||||||
| SwissProt Accessions | Q9NYB0, Q8WYZ3, Q9NWR2 | Gene names | TERF2IP, RAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Telomeric repeat-binding factor 2-interacting protein 1 (TRF2- interacting telomeric protein Rap1) (hRap1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
TE2IP_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | Q9NYB0, Q8WYZ3, Q9NWR2 | Gene names | TERF2IP, RAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Telomeric repeat-binding factor 2-interacting protein 1 (TRF2- interacting telomeric protein Rap1) (hRap1). | |||||
|
TE2IP_MOUSE
|
||||||
| θ value | 2.94559e-168 (rank : 2) | NC score | 0.952938 (rank : 2) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q91VL8, Q543F4, Q9JJE8, Q9JJE9 | Gene names | Terf2ip, Rap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Telomeric repeat-binding factor 2-interacting protein 1 (TRF2- interacting telomeric protein Rap1). | |||||
|
NASP_HUMAN
|
||||||
| θ value | 0.00175202 (rank : 3) | NC score | 0.130440 (rank : 5) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P49321, Q96A69, Q9BTW2 | Gene names | NASP | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear autoantigenic sperm protein (NASP). | |||||
|
U2AFM_MOUSE
|
||||||
| θ value | 0.00509761 (rank : 4) | NC score | 0.130626 (rank : 4) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q62377 | Gene names | Zrsr2, U2af1-rs2, U2af1l2, U2af1rs2 | |||
|
Domain Architecture |
|
|||||
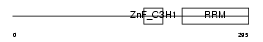
| Description | U2 small nuclear ribonucleoprotein auxiliary factor 35 kDa subunit- related protein 2 (U2(RNU2) small nuclear RNA auxiliary factor 1-like 2) (CCCH type zinc finger, RNA-binding motif and serine/arginine rich protein 2). | |||||
|
U2AFM_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 5) | NC score | 0.127031 (rank : 6) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 577 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q15696 | Gene names | ZRSR2, U2AF1-RS2, U2AF1L2, U2AF1RS2 | |||
|
Domain Architecture |
|
|||||
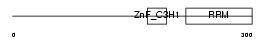
| Description | U2 small nuclear ribonucleoprotein auxiliary factor 35 kDa subunit- related protein 2 (U2(RNU2) small nuclear RNA auxiliary factor 1-like 2) (CCCH type zinc finger, RNA-binding motif and serine/arginine rich protein 2) (NY-REN-20 antigen). | |||||
|
RN168_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 6) | NC score | 0.065988 (rank : 16) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 440 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q80XJ2 | Gene names | Rnf168 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 168. | |||||
|
TXND2_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 7) | NC score | 0.069514 (rank : 13) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
U2AFL_MOUSE
|
||||||
| θ value | 0.125558 (rank : 8) | NC score | 0.132594 (rank : 3) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 389 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q64707 | Gene names | Zrsr1, Sp2, Sp2-7, U2af1-rs1, U2af1l1 | |||
|
Domain Architecture |

|
|||||
| Description | U2 small nuclear ribonucleoprotein auxiliary factor 35 kDa subunit- related protein 1 (U2(RNU2) small nuclear RNA auxiliary factor 1-like 1) (CCCH type zinc finger, RNA-binding motif and serine/arginine rich protein 1) (SP2). | |||||
|
NEST_HUMAN
|
||||||
| θ value | 0.163984 (rank : 9) | NC score | 0.056388 (rank : 21) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1305 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P48681, O00552 | Gene names | NES | |||
|
Domain Architecture |

|
|||||
| Description | Nestin. | |||||
|
U2AFL_HUMAN
|
||||||
| θ value | 0.163984 (rank : 10) | NC score | 0.124166 (rank : 7) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 503 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q15695, Q13570 | Gene names | ZRSR1, U2AF1-RS1, U2AF1L1, U2AF1RS1, U2AFBPL | |||
|
Domain Architecture |
|
|||||
| Description | U2 small nuclear ribonucleoprotein auxiliary factor 35 kDa subunit- related protein 1 (U2(RNU2) small nuclear RNA auxiliary factor 1-like 1) (CCCH type zinc finger, RNA-binding motif and serine/arginine rich protein 1). | |||||
|
CIR_HUMAN
|
||||||
| θ value | 0.365318 (rank : 11) | NC score | 0.081161 (rank : 12) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q86X95, O95367, Q12804, Q4G1B9, Q6PJI4, Q8IWI2 | Gene names | CIR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CBF1-interacting corepressor (Recepin). | |||||
|
MYST4_HUMAN
|
||||||
| θ value | 0.365318 (rank : 12) | NC score | 0.056670 (rank : 20) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8WYB5, O15087, Q86Y05, Q8WU81, Q9UKW2, Q9UKW3, Q9UKX0 | Gene names | MYST4, KIAA0383, MORF, MOZ2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Histone acetyltransferase MOZ2) (Monocytic leukemia zinc finger protein- related factor) (Histone acetyltransferase MORF). | |||||
|
CIR_MOUSE
|
||||||
| θ value | 0.47712 (rank : 13) | NC score | 0.082844 (rank : 11) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9DA19, Q3V2N9, Q4KL44, Q52KL9, Q5FW66 | Gene names | Cir | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CBF1-interacting corepressor. | |||||
|
CMGA_HUMAN
|
||||||
| θ value | 0.813845 (rank : 14) | NC score | 0.089787 (rank : 8) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P10645, Q96E84, Q96GL7, Q9BQB5 | Gene names | CHGA | |||
|
Domain Architecture |

|
|||||
| Description | Chromogranin A precursor (CgA) (Pituitary secretory protein I) (SP-I) [Contains: Vasostatin-1 (Vasostatin I); Vasostatin-2 (Vasostatin II); EA-92; ES-43; Pancreastatin; SS-18; WA-8; WE-14; LF-19; AL-11; GV-19; GR-44; ER-37]. | |||||
|
SFR17_HUMAN
|
||||||
| θ value | 0.813845 (rank : 15) | NC score | 0.059236 (rank : 18) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8TF01, Q5T064, Q6P2B4, Q6P2N4, Q6PJ93, Q6PK36, Q7Z640, Q8TF00, Q96K10, Q96SM5, Q9P076, Q9P0C0, Q9Y4N3 | Gene names | SRRP130, C6orf111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 130 (Serine-arginine-rich- splicing regulatory protein 130) (SRrp130) (SR-rich protein) (SR- related protein). | |||||
|
TTBK1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 16) | NC score | 0.017083 (rank : 62) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1058 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q5TCY1, Q2L6C6, Q8N444, Q96JH2 | Gene names | TTBK1, BDTK, KIAA1855 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tau-tubulin kinase 1 (EC 2.7.11.1) (Brain-derived tau kinase). | |||||
|
DDIT3_MOUSE
|
||||||
| θ value | 1.06291 (rank : 17) | NC score | 0.066443 (rank : 15) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P35639, Q91YW9 | Gene names | Ddit3, Chop, Gadd153 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA damage-inducible transcript 3 (DDIT-3) (Growth arrest and DNA- damage-inducible protein GADD153) (C/EBP-homologous protein) (CHOP). | |||||
|
IQGA2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 18) | NC score | 0.031078 (rank : 45) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q13576 | Gene names | IQGAP2 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP2. | |||||
|
AFF4_HUMAN
|
||||||
| θ value | 1.38821 (rank : 19) | NC score | 0.044693 (rank : 31) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 611 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9UHB7, Q498B2, Q59FB3, Q6P592, Q8TDR1, Q9P0E4 | Gene names | AFF4, AF5Q31, MCEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4 (ALL1-fused gene from chromosome 5q31) (Major CDK9 elongation factor-associated protein). | |||||
|
CCD45_HUMAN
|
||||||
| θ value | 1.38821 (rank : 20) | NC score | 0.038704 (rank : 35) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 525 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q96GE4, Q96M81 | Gene names | CCDC45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 45. | |||||
|
CHD1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 21) | NC score | 0.019590 (rank : 57) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O14646 | Gene names | CHD1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
DEK_MOUSE
|
||||||
| θ value | 1.38821 (rank : 22) | NC score | 0.066540 (rank : 14) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q7TNV0, Q80VC5, Q8BZV6 | Gene names | Dek | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DEK. | |||||
|
GLU2B_HUMAN
|
||||||
| θ value | 1.38821 (rank : 23) | NC score | 0.055255 (rank : 22) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 377 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P14314, Q96BU9, Q96D06, Q9P0W9 | Gene names | PRKCSH, G19P1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glucosidase 2 subunit beta precursor (Glucosidase II subunit beta) (Protein kinase C substrate, 60.1 kDa protein, heavy chain) (PKCSH) (80K-H protein). | |||||
|
MAP1B_HUMAN
|
||||||
| θ value | 1.38821 (rank : 24) | NC score | 0.049749 (rank : 26) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P46821 | Gene names | MAP1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) [Contains: MAP1 light chain LC1]. | |||||
|
CNGA1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 25) | NC score | 0.022142 (rank : 53) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P29974, Q60776 | Gene names | Cnga1, Cncg, Cncg1 | |||
|
Domain Architecture |

|
|||||
| Description | cGMP-gated cation channel alpha 1 (CNG channel alpha 1) (CNG-1) (CNG1) (Cyclic nucleotide-gated channel alpha 1) (Cyclic nucleotide-gated channel, photoreceptor) (Cyclic nucleotide-gated cation channel 1) (Rod photoreceptor cGMP-gated channel subunit alpha). | |||||
|
NFH_HUMAN
|
||||||
| θ value | 1.81305 (rank : 26) | NC score | 0.027096 (rank : 46) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
GBF1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 27) | NC score | 0.032799 (rank : 42) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q92538, Q5VXX3, Q96CK6, Q96HZ3, Q9H473 | Gene names | GBF1, KIAA0248 | |||
|
Domain Architecture |

|
|||||
| Description | Golgi-specific brefeldin A-resistance guanine nucleotide exchange factor 1 (BFA-resistant GEF 1). | |||||
|
MYT1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 28) | NC score | 0.036415 (rank : 38) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 475 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q01538, O94922, Q9UPV2 | Gene names | MYT1, KIAA0835, KIAA1050, MTF1, MYTI, PLPB1 | |||
|
Domain Architecture |

|
|||||
| Description | Myelin transcription factor 1 (MyT1) (MyTI) (Proteolipid protein- binding protein) (PLPB1). | |||||
|
MYT1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 29) | NC score | 0.044569 (rank : 32) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8CFC2, O08995, Q8CFH1 | Gene names | Myt1, Kiaa0835, Nzf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1 (MyT1) (Neural zinc finger factor 2) (NZF-2). | |||||
|
PELP1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 30) | NC score | 0.054308 (rank : 24) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 596 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8IZL8, O15450, Q5EGN3, Q6NTE6, Q96FT1, Q9BU60 | Gene names | PELP1, HMX3, MNAR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor) (Transcription factor HMX3). | |||||
|
DDX24_MOUSE
|
||||||
| θ value | 3.0926 (rank : 31) | NC score | 0.012271 (rank : 73) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9ESV0, Q61119 | Gene names | Ddx24 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent RNA helicase DDX24 (EC 3.6.1.-) (DEAD box protein 24). | |||||
|
EMIL2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 32) | NC score | 0.019590 (rank : 58) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 485 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8K482 | Gene names | Emilin2 | |||
|
Domain Architecture |

|
|||||
| Description | EMILIN-2 precursor (Elastin microfibril interface-located protein 2) (Elastin microfibril interfacer 2) (Basilin). | |||||
|
NIN_MOUSE
|
||||||
| θ value | 3.0926 (rank : 33) | NC score | 0.026013 (rank : 49) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q61043, Q6ZPM7 | Gene names | Nin, Kiaa1565 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ninein. | |||||
|
YTDC1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 34) | NC score | 0.054449 (rank : 23) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q96MU7, Q7Z622, Q8TF35 | Gene names | YTHDC1, KIAA1966, YT521 | |||
|
Domain Architecture |

|
|||||
| Description | YTH domain-containing protein 1 (Putative splicing factor YT521). | |||||
|
ANLN_MOUSE
|
||||||
| θ value | 4.03905 (rank : 35) | NC score | 0.031601 (rank : 43) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 297 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8K298, Q8BL79, Q8BLB3, Q8K2N0, Q9CUY0 | Gene names | Anln | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Actin-binding protein anillin. | |||||
|
ATBF1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 36) | NC score | 0.011470 (rank : 74) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1841 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q61329 | Gene names | Atbf1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-fetoprotein enhancer-binding protein (AT motif-binding factor) (AT-binding transcription factor 1). | |||||
|
CD2L1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 37) | NC score | 0.007090 (rank : 76) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1467 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P21127, O95265, Q12817, Q12818, Q12819, Q12820, Q12822, Q8N530, Q9NZS5, Q9UBJ0, Q9UBQ1, Q9UBR0, Q9UNY2, Q9UP57, Q9UP58, Q9UP59 | Gene names | CDC2L1, PITSLREA, PK58 | |||
|
Domain Architecture |

|
|||||
| Description | PITSLRE serine/threonine-protein kinase CDC2L1 (EC 2.7.11.22) (Galactosyltransferase-associated protein kinase p58/GTA) (Cell division cycle 2-like protein kinase 1) (CLK-1) (CDK11) (p58 CLK-1). | |||||
|
CROP_HUMAN
|
||||||
| θ value | 4.03905 (rank : 38) | NC score | 0.036194 (rank : 39) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O95232, Q6PHR9, Q9NUY0, Q9P2S7 | Gene names | CROP, CREAP1, O48 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cisplatin resistance-associated overexpressed protein (cAMP regulatory element-associated protein 1) (CRE-associated protein 1) (CREAP-1) (Luc7A) (Okadaic acid-inducible phosphoprotein OA48-18). | |||||
|
CROP_MOUSE
|
||||||
| θ value | 4.03905 (rank : 39) | NC score | 0.037856 (rank : 36) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q5SUF2, Q3U9D5, Q8BUJ5, Q921Z3, Q9CRS7, Q9CTY4 | Gene names | Crop | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cisplatin resistance-associated overexpressed protein. | |||||
|
DDIT3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 40) | NC score | 0.060835 (rank : 17) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P35638 | Gene names | DDIT3, CHOP, GADD153 | |||
|
Domain Architecture |
|
|||||
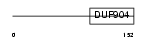
| Description | DNA damage-inducible transcript 3 (DDIT-3) (Growth arrest and DNA- damage-inducible protein GADD153) (C/EBP-homologous protein) (CHOP). | |||||
|
DEK_HUMAN
|
||||||
| θ value | 4.03905 (rank : 41) | NC score | 0.057992 (rank : 19) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P35659, Q5TGV4, Q5TGV5 | Gene names | DEK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DEK. | |||||
|
FGD5_HUMAN
|
||||||
| θ value | 4.03905 (rank : 42) | NC score | 0.014308 (rank : 69) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6ZNL6, Q6MZY1, Q7Z303, Q8IYP3, Q8N861, Q8N8G4 | Gene names | FGD5, ZFYVE23 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 5 (Zinc finger FYVE domain-containing protein 23). | |||||
|
PPWD1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 43) | NC score | 0.016425 (rank : 66) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8CEC6 | Gene names | Ppwd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peptidylprolyl isomerase domain and WD repeat-containing protein 1 (EC 5.2.1.8). | |||||
|
RBBP6_MOUSE
|
||||||
| θ value | 4.03905 (rank : 44) | NC score | 0.045406 (rank : 29) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
RPGR1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 45) | NC score | 0.031516 (rank : 44) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 713 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9EPQ2, Q8CAC2, Q8CDJ9, Q91WE0, Q9CUK6, Q9D5Q1 | Gene names | Rpgrip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator-interacting protein 1 (RPGR-interacting protein 1). | |||||
|
U3IP2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 46) | NC score | 0.010523 (rank : 75) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O43818, Q8IZ30 | Gene names | RNU3IP2, U355K | |||
|
Domain Architecture |

|
|||||
| Description | U3 small nucleolar RNA-interacting protein 2 (U3 small nucleolar ribonucleoprotein-associated 55 kDa protein) (U3 snoRNP-associated 55 kDa protein) (U3-55K). | |||||
|
CENPE_HUMAN
|
||||||
| θ value | 5.27518 (rank : 47) | NC score | 0.018596 (rank : 61) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 2060 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q02224 | Gene names | CENPE | |||
|
Domain Architecture |

|
|||||
| Description | Centromeric protein E (CENP-E). | |||||
|
CP250_HUMAN
|
||||||
| θ value | 5.27518 (rank : 48) | NC score | 0.020495 (rank : 55) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1805 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9BV73, O14812, O60588, Q9H450 | Gene names | CEP250, CEP2, CNAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Centrosome-associated protein CEP250 (Centrosomal protein 2) (Centrosomal Nek2-associated protein 1) (C-Nap1). | |||||
|
DHX57_MOUSE
|
||||||
| θ value | 5.27518 (rank : 49) | NC score | 0.012974 (rank : 71) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6P5D3, Q3TS93, Q6NZK4, Q6P1B4, Q8BI63, Q8BIA2, Q8R360 | Gene names | Dhx57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative ATP-dependent RNA helicase DHX57 (EC 3.6.1.-) (DEAH box protein 57). | |||||
|
GLU2B_MOUSE
|
||||||
| θ value | 5.27518 (rank : 50) | NC score | 0.047432 (rank : 27) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O08795, Q921X2 | Gene names | Prkcsh | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glucosidase 2 subunit beta precursor (Glucosidase II subunit beta) (Protein kinase C substrate, 60.1 kDa protein, heavy chain) (PKCSH) (80K-H protein). | |||||
|
LONM_HUMAN
|
||||||
| θ value | 5.27518 (rank : 51) | NC score | 0.026028 (rank : 48) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P36776, P36777, Q9UQ95 | Gene names | PRSS15 | |||
|
Domain Architecture |

|
|||||
| Description | Lon protease homolog, mitochondrial precursor (EC 3.4.21.-) (Lon protease-like protein) (LONP) (LONHs). | |||||
|
NFH_MOUSE
|
||||||
| θ value | 5.27518 (rank : 52) | NC score | 0.026399 (rank : 47) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
PARC_MOUSE
|
||||||
| θ value | 5.27518 (rank : 53) | NC score | 0.014400 (rank : 68) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80TT8, Q8BKL9, Q8CGC0, Q8R363 | Gene names | Parc, Kiaa0708 | |||
|
Domain Architecture |

|
|||||
| Description | p53-associated parkin-like cytoplasmic protein. | |||||
|
PELP1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 54) | NC score | 0.045267 (rank : 30) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9DBD5, Q5F2E2, Q6PEM0, Q91YM9 | Gene names | Pelp1, Mnar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor). | |||||
|
SPRL1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 55) | NC score | 0.046395 (rank : 28) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P70663, P97810, Q99L82 | Gene names | Sparcl1, Ecm2, Sc1 | |||
|
Domain Architecture |

|
|||||
| Description | SPARC-like protein 1 precursor (Matrix glycoprotein Sc1) (Extracellular matrix protein 2). | |||||
|
SYNE1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 56) | NC score | 0.019312 (rank : 59) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1484 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8NF91, O94890, Q5JV19, Q5JV22, Q8N9P7, Q8TCP1, Q8WWW6, Q8WWW7, Q8WXF6, Q96N17, Q9C0A7, Q9H525, Q9H526, Q9NS36, Q9NU50, Q9UJ06, Q9UJ07, Q9ULF8 | Gene names | SYNE1, KIAA0796, KIAA1262, KIAA1756, MYNE1 | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-1 (Nuclear envelope spectrin repeat protein 1) (Synaptic nuclear envelope protein 1) (Syne-1) (Myocyte nuclear envelope protein 1) (Myne-1) (Enaptin). | |||||
|
TATD2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 57) | NC score | 0.035774 (rank : 40) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q93075 | Gene names | TATDN2, KIAA0218 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TatD DNase domain-containing deoxyribonuclease 2 (EC 3.1.21.-). | |||||
|
CCD27_HUMAN
|
||||||
| θ value | 6.88961 (rank : 58) | NC score | 0.018666 (rank : 60) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 594 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q2M243, Q5TBV3, Q96M50 | Gene names | CCDC27 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 27. | |||||
|
GA2L2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 59) | NC score | 0.016564 (rank : 64) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8NHY3, Q8NHY4 | Gene names | GAS2L2, GAR17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GAS2-like protein 2 (Growth arrest-specific 2-like 2) (GAS2-related protein on chromosome 17) (GAR17 protein). | |||||
|
HUWE1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 60) | NC score | 0.012715 (rank : 72) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q7Z6Z7, O15029, Q4G2Z2, Q5H961, Q6P4D0, Q8NG67, Q9BUI0, Q9HCJ4, Q9NSL6, Q9P0A9 | Gene names | HUWE1, KIAA0312, KIAA1578, UREB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (Mcl-1 ubiquitin ligase E3) (Mule) (ARF-binding protein 1) (ARF-BP1). | |||||
|
MYST4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 61) | NC score | 0.037800 (rank : 37) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8BRB7, Q7TNW5, Q8BG35, Q8C441, Q9JKX5 | Gene names | Myst4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Querkopf protein). | |||||
|
NKTR_HUMAN
|
||||||
| θ value | 6.88961 (rank : 62) | NC score | 0.035673 (rank : 41) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 658 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P30414 | Gene names | NKTR | |||
|
Domain Architecture |

|
|||||
| Description | NK-tumor recognition protein (Natural-killer cells cyclophilin-related protein) (NK-TR protein). | |||||
|
SMRCD_MOUSE
|
||||||
| θ value | 6.88961 (rank : 63) | NC score | 0.013180 (rank : 70) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q04692 | Gene names | Smarcad1, Etl1, Kiaa1122 | |||
|
Domain Architecture |

|
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A containing DEAD/H box 1 (EC 3.6.1.-) (ATP- dependent helicase SMARCAD1) (Enhancer trap locus homolog 1) (Etl-1). | |||||
|
TRHY_HUMAN
|
||||||
| θ value | 6.88961 (rank : 64) | NC score | 0.016560 (rank : 65) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1385 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q07283 | Gene names | THH, THL, TRHY | |||
|
Domain Architecture |

|
|||||
| Description | Trichohyalin. | |||||
|
CD2L1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 65) | NC score | 0.006486 (rank : 77) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1463 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P24788, Q3UI03, Q61399, Q7TST4, Q8BP53 | Gene names | Cdc2l1 | |||
|
Domain Architecture |

|
|||||
| Description | PITSLRE serine/threonine-protein kinase CDC2L1 (EC 2.7.11.22) (Galactosyltransferase-associated protein kinase p58/GTA) (Cell division cycle 2-like protein kinase 1). | |||||
|
CHIC1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 66) | NC score | 0.014686 (rank : 67) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5VXU3 | Gene names | CHIC1, BRX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cysteine-rich hydrophobic domain 1 protein (Brain X-linked protein). | |||||
|
CNGB1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 67) | NC score | 0.020643 (rank : 54) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14028, Q14029 | Gene names | CNGB1, CNCG4 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic nucleotide-gated cation channel 4 (CNG channel 4) (CNG-4) (CNG4) (Cyclic nucleotide-gated cation channel modulatory subunit). | |||||
|
DYHC_HUMAN
|
||||||
| θ value | 8.99809 (rank : 68) | NC score | 0.024148 (rank : 51) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 733 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q14204, Q6DKQ7, Q8WU28, Q92814, Q9Y4G5 | Gene names | DYNC1H1, DNCH1, DNECL, KIAA0325 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dynein heavy chain, cytosolic (DYHC) (Cytoplasmic dynein heavy chain 1) (DHC1) (Dynein heavy chain 1, cytoplasmic 1). | |||||
|
DYHC_MOUSE
|
||||||
| θ value | 8.99809 (rank : 69) | NC score | 0.024018 (rank : 52) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 729 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9JHU4 | Gene names | Dync1h1, Dnch1, Dnchc1 | |||
|
Domain Architecture |

|
|||||
| Description | Dynein heavy chain, cytosolic (DYHC) (Cytoplasmic dynein heavy chain 1) (DHC1) (Dynein heavy chain 1, cytoplasmic 1). | |||||
|
MYST3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 70) | NC score | 0.040733 (rank : 33) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q92794 | Gene names | MYST3, MOZ, RUNXBP2, ZNF220 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Runt-related transcription factor-binding protein 2) (Monocytic leukemia zinc finger protein) (Zinc finger protein 220). | |||||
|
MYST3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 71) | NC score | 0.040502 (rank : 34) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 688 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8BZ21, Q8BW52, Q8BYH1, Q8C1F3 | Gene names | Myst3, Moz | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Monocytic leukemia zinc finger protein) (Monocytic leukemia zinc finger homolog). | |||||
|
NFL_HUMAN
|
||||||
| θ value | 8.99809 (rank : 72) | NC score | 0.017036 (rank : 63) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 666 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P07196, Q16154, Q8IU72 | Gene names | NEFL, NF68, NFL | |||
|
Domain Architecture |
|
|||||
| Description | Neurofilament triplet L protein (68 kDa neurofilament protein) (Neurofilament light polypeptide) (NF-L). | |||||
|
RBM13_MOUSE
|
||||||
| θ value | 8.99809 (rank : 73) | NC score | 0.020371 (rank : 56) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BGS0, Q921E0, Q9D5G6 | Gene names | Rbm13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MAK16-like protein RBM13 (RNA-binding motif protein 13). | |||||
|
SRCA_HUMAN
|
||||||
| θ value | 8.99809 (rank : 74) | NC score | 0.025644 (rank : 50) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q86TD4 | Gene names | SRL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcalumenin precursor. | |||||
|
ZN202_HUMAN
|
||||||
| θ value | 8.99809 (rank : 75) | NC score | 0.003318 (rank : 78) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 756 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O95125, Q4JG21, Q9H1B9, Q9NSM4 | Gene names | ZNF202 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 202. | |||||
|
TXND2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 76) | NC score | 0.052495 (rank : 25) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
U2AF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 77) | NC score | 0.083690 (rank : 9) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q01081 | Gene names | U2AF1, U2AF35 | |||
|
Domain Architecture |
|
|||||
| Description | Splicing factor U2AF 35 kDa subunit (U2 auxiliary factor 35 kDa subunit) (U2 snRNP auxiliary factor small subunit). | |||||
|
U2AF1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 78) | NC score | 0.083666 (rank : 10) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D883, Q564E4, Q99LX2, Q9CZ98 | Gene names | U2af1 | |||
|
Domain Architecture |
|
|||||
| Description | Splicing factor U2AF 35 kDa subunit (U2 auxiliary factor 35 kDa subunit) (U2 snRNP auxiliary factor small subunit). | |||||
|
TE2IP_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | Q9NYB0, Q8WYZ3, Q9NWR2 | Gene names | TERF2IP, RAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Telomeric repeat-binding factor 2-interacting protein 1 (TRF2- interacting telomeric protein Rap1) (hRap1). | |||||
|
TE2IP_MOUSE
|
||||||
| NC score | 0.952938 (rank : 2) | θ value | 2.94559e-168 (rank : 2) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q91VL8, Q543F4, Q9JJE8, Q9JJE9 | Gene names | Terf2ip, Rap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Telomeric repeat-binding factor 2-interacting protein 1 (TRF2- interacting telomeric protein Rap1). | |||||
|
U2AFL_MOUSE
|
||||||
| NC score | 0.132594 (rank : 3) | θ value | 0.125558 (rank : 8) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 389 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q64707 | Gene names | Zrsr1, Sp2, Sp2-7, U2af1-rs1, U2af1l1 | |||
|
Domain Architecture |

|
|||||
| Description | U2 small nuclear ribonucleoprotein auxiliary factor 35 kDa subunit- related protein 1 (U2(RNU2) small nuclear RNA auxiliary factor 1-like 1) (CCCH type zinc finger, RNA-binding motif and serine/arginine rich protein 1) (SP2). | |||||
|
U2AFM_MOUSE
|
||||||
| NC score | 0.130626 (rank : 4) | θ value | 0.00509761 (rank : 4) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q62377 | Gene names | Zrsr2, U2af1-rs2, U2af1l2, U2af1rs2 | |||
|
Domain Architecture |

|
|||||
| Description | U2 small nuclear ribonucleoprotein auxiliary factor 35 kDa subunit- related protein 2 (U2(RNU2) small nuclear RNA auxiliary factor 1-like 2) (CCCH type zinc finger, RNA-binding motif and serine/arginine rich protein 2). | |||||
|
NASP_HUMAN
|
||||||
| NC score | 0.130440 (rank : 5) | θ value | 0.00175202 (rank : 3) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P49321, Q96A69, Q9BTW2 | Gene names | NASP | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear autoantigenic sperm protein (NASP). | |||||
|
U2AFM_HUMAN
|
||||||
| NC score | 0.127031 (rank : 6) | θ value | 0.00869519 (rank : 5) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 577 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q15696 | Gene names | ZRSR2, U2AF1-RS2, U2AF1L2, U2AF1RS2 | |||
|
Domain Architecture |

|
|||||
| Description | U2 small nuclear ribonucleoprotein auxiliary factor 35 kDa subunit- related protein 2 (U2(RNU2) small nuclear RNA auxiliary factor 1-like 2) (CCCH type zinc finger, RNA-binding motif and serine/arginine rich protein 2) (NY-REN-20 antigen). | |||||
|
U2AFL_HUMAN
|
||||||
| NC score | 0.124166 (rank : 7) | θ value | 0.163984 (rank : 10) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 503 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q15695, Q13570 | Gene names | ZRSR1, U2AF1-RS1, U2AF1L1, U2AF1RS1, U2AFBPL | |||
|
Domain Architecture |

|
|||||
| Description | U2 small nuclear ribonucleoprotein auxiliary factor 35 kDa subunit- related protein 1 (U2(RNU2) small nuclear RNA auxiliary factor 1-like 1) (CCCH type zinc finger, RNA-binding motif and serine/arginine rich protein 1). | |||||
|
CMGA_HUMAN
|
||||||
| NC score | 0.089787 (rank : 8) | θ value | 0.813845 (rank : 14) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P10645, Q96E84, Q96GL7, Q9BQB5 | Gene names | CHGA | |||
|
Domain Architecture |

|
|||||
| Description | Chromogranin A precursor (CgA) (Pituitary secretory protein I) (SP-I) [Contains: Vasostatin-1 (Vasostatin I); Vasostatin-2 (Vasostatin II); EA-92; ES-43; Pancreastatin; SS-18; WA-8; WE-14; LF-19; AL-11; GV-19; GR-44; ER-37]. | |||||
|
U2AF1_HUMAN
|
||||||
| NC score | 0.083690 (rank : 9) | θ value | θ > 10 (rank : 77) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q01081 | Gene names | U2AF1, U2AF35 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor U2AF 35 kDa subunit (U2 auxiliary factor 35 kDa subunit) (U2 snRNP auxiliary factor small subunit). | |||||
|
U2AF1_MOUSE
|
||||||
| NC score | 0.083666 (rank : 10) | θ value | θ > 10 (rank : 78) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D883, Q564E4, Q99LX2, Q9CZ98 | Gene names | U2af1 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor U2AF 35 kDa subunit (U2 auxiliary factor 35 kDa subunit) (U2 snRNP auxiliary factor small subunit). | |||||
|
CIR_MOUSE
|
||||||
| NC score | 0.082844 (rank : 11) | θ value | 0.47712 (rank : 13) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9DA19, Q3V2N9, Q4KL44, Q52KL9, Q5FW66 | Gene names | Cir | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CBF1-interacting corepressor. | |||||
|
CIR_HUMAN
|
||||||
| NC score | 0.081161 (rank : 12) | θ value | 0.365318 (rank : 11) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q86X95, O95367, Q12804, Q4G1B9, Q6PJI4, Q8IWI2 | Gene names | CIR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CBF1-interacting corepressor (Recepin). | |||||
|
TXND2_HUMAN
|
||||||
| NC score | 0.069514 (rank : 13) | θ value | 0.0431538 (rank : 7) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
DEK_MOUSE
|
||||||
| NC score | 0.066540 (rank : 14) | θ value | 1.38821 (rank : 22) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q7TNV0, Q80VC5, Q8BZV6 | Gene names | Dek | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DEK. | |||||
|
DDIT3_MOUSE
|
||||||
| NC score | 0.066443 (rank : 15) | θ value | 1.06291 (rank : 17) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P35639, Q91YW9 | Gene names | Ddit3, Chop, Gadd153 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA damage-inducible transcript 3 (DDIT-3) (Growth arrest and DNA- damage-inducible protein GADD153) (C/EBP-homologous protein) (CHOP). | |||||
|
RN168_MOUSE
|
||||||
| NC score | 0.065988 (rank : 16) | θ value | 0.0252991 (rank : 6) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 440 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q80XJ2 | Gene names | Rnf168 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 168. | |||||
|
DDIT3_HUMAN
|
||||||
| NC score | 0.060835 (rank : 17) | θ value | 4.03905 (rank : 40) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P35638 | Gene names | DDIT3, CHOP, GADD153 | |||
|
Domain Architecture |

|
|||||
| Description | DNA damage-inducible transcript 3 (DDIT-3) (Growth arrest and DNA- damage-inducible protein GADD153) (C/EBP-homologous protein) (CHOP). | |||||
|
SFR17_HUMAN
|
||||||
| NC score | 0.059236 (rank : 18) | θ value | 0.813845 (rank : 15) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8TF01, Q5T064, Q6P2B4, Q6P2N4, Q6PJ93, Q6PK36, Q7Z640, Q8TF00, Q96K10, Q96SM5, Q9P076, Q9P0C0, Q9Y4N3 | Gene names | SRRP130, C6orf111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 130 (Serine-arginine-rich- splicing regulatory protein 130) (SRrp130) (SR-rich protein) (SR- related protein). | |||||
|
DEK_HUMAN
|
||||||
| NC score | 0.057992 (rank : 19) | θ value | 4.03905 (rank : 41) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P35659, Q5TGV4, Q5TGV5 | Gene names | DEK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DEK. | |||||
|
MYST4_HUMAN
|
||||||
| NC score | 0.056670 (rank : 20) | θ value | 0.365318 (rank : 12) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8WYB5, O15087, Q86Y05, Q8WU81, Q9UKW2, Q9UKW3, Q9UKX0 | Gene names | MYST4, KIAA0383, MORF, MOZ2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Histone acetyltransferase MOZ2) (Monocytic leukemia zinc finger protein- related factor) (Histone acetyltransferase MORF). | |||||
|
NEST_HUMAN
|
||||||
| NC score | 0.056388 (rank : 21) | θ value | 0.163984 (rank : 9) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1305 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P48681, O00552 | Gene names | NES | |||
|
Domain Architecture |

|
|||||
| Description | Nestin. | |||||
|
GLU2B_HUMAN
|
||||||
| NC score | 0.055255 (rank : 22) | θ value | 1.38821 (rank : 23) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 377 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P14314, Q96BU9, Q96D06, Q9P0W9 | Gene names | PRKCSH, G19P1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glucosidase 2 subunit beta precursor (Glucosidase II subunit beta) (Protein kinase C substrate, 60.1 kDa protein, heavy chain) (PKCSH) (80K-H protein). | |||||
|
YTDC1_HUMAN
|
||||||
| NC score | 0.054449 (rank : 23) | θ value | 3.0926 (rank : 34) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q96MU7, Q7Z622, Q8TF35 | Gene names | YTHDC1, KIAA1966, YT521 | |||
|
Domain Architecture |

|
|||||
| Description | YTH domain-containing protein 1 (Putative splicing factor YT521). | |||||
|
PELP1_HUMAN
|
||||||
| NC score | 0.054308 (rank : 24) | θ value | 2.36792 (rank : 30) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 596 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8IZL8, O15450, Q5EGN3, Q6NTE6, Q96FT1, Q9BU60 | Gene names | PELP1, HMX3, MNAR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor) (Transcription factor HMX3). | |||||
|
TXND2_MOUSE
|
||||||
| NC score | 0.052495 (rank : 25) | θ value | θ > 10 (rank : 76) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
MAP1B_HUMAN
|
||||||
| NC score | 0.049749 (rank : 26) | θ value | 1.38821 (rank : 24) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P46821 | Gene names | MAP1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) [Contains: MAP1 light chain LC1]. | |||||
|
GLU2B_MOUSE
|
||||||
| NC score | 0.047432 (rank : 27) | θ value | 5.27518 (rank : 50) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O08795, Q921X2 | Gene names | Prkcsh | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glucosidase 2 subunit beta precursor (Glucosidase II subunit beta) (Protein kinase C substrate, 60.1 kDa protein, heavy chain) (PKCSH) (80K-H protein). | |||||
|
SPRL1_MOUSE
|
||||||
| NC score | 0.046395 (rank : 28) | θ value | 5.27518 (rank : 55) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P70663, P97810, Q99L82 | Gene names | Sparcl1, Ecm2, Sc1 | |||
|
Domain Architecture |

|
|||||
| Description | SPARC-like protein 1 precursor (Matrix glycoprotein Sc1) (Extracellular matrix protein 2). | |||||
|
RBBP6_MOUSE
|
||||||
| NC score | 0.045406 (rank : 29) | θ value | 4.03905 (rank : 44) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
PELP1_MOUSE
|
||||||
| NC score | 0.045267 (rank : 30) | θ value | 5.27518 (rank : 54) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9DBD5, Q5F2E2, Q6PEM0, Q91YM9 | Gene names | Pelp1, Mnar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor). | |||||
|
AFF4_HUMAN
|
||||||
| NC score | 0.044693 (rank : 31) | θ value | 1.38821 (rank : 19) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 611 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9UHB7, Q498B2, Q59FB3, Q6P592, Q8TDR1, Q9P0E4 | Gene names | AFF4, AF5Q31, MCEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4 (ALL1-fused gene from chromosome 5q31) (Major CDK9 elongation factor-associated protein). | |||||
|
MYT1_MOUSE
|
||||||
| NC score | 0.044569 (rank : 32) | θ value | 2.36792 (rank : 29) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8CFC2, O08995, Q8CFH1 | Gene names | Myt1, Kiaa0835, Nzf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1 (MyT1) (Neural zinc finger factor 2) (NZF-2). | |||||
|
MYST3_HUMAN
|
||||||
| NC score | 0.040733 (rank : 33) | θ value | 8.99809 (rank : 70) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q92794 | Gene names | MYST3, MOZ, RUNXBP2, ZNF220 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Runt-related transcription factor-binding protein 2) (Monocytic leukemia zinc finger protein) (Zinc finger protein 220). | |||||
|
MYST3_MOUSE
|
||||||
| NC score | 0.040502 (rank : 34) | θ value | 8.99809 (rank : 71) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 688 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8BZ21, Q8BW52, Q8BYH1, Q8C1F3 | Gene names | Myst3, Moz | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Monocytic leukemia zinc finger protein) (Monocytic leukemia zinc finger homolog). | |||||
|
CCD45_HUMAN
|
||||||
| NC score | 0.038704 (rank : 35) | θ value | 1.38821 (rank : 20) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 525 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q96GE4, Q96M81 | Gene names | CCDC45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 45. | |||||
|
CROP_MOUSE
|
||||||
| NC score | 0.037856 (rank : 36) | θ value | 4.03905 (rank : 39) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q5SUF2, Q3U9D5, Q8BUJ5, Q921Z3, Q9CRS7, Q9CTY4 | Gene names | Crop | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cisplatin resistance-associated overexpressed protein. | |||||
|
MYST4_MOUSE
|
||||||
| NC score | 0.037800 (rank : 37) | θ value | 6.88961 (rank : 61) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8BRB7, Q7TNW5, Q8BG35, Q8C441, Q9JKX5 | Gene names | Myst4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Querkopf protein). | |||||
|
MYT1_HUMAN
|
||||||
| NC score | 0.036415 (rank : 38) | θ value | 2.36792 (rank : 28) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 475 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q01538, O94922, Q9UPV2 | Gene names | MYT1, KIAA0835, KIAA1050, MTF1, MYTI, PLPB1 | |||
|
Domain Architecture |

|
|||||
| Description | Myelin transcription factor 1 (MyT1) (MyTI) (Proteolipid protein- binding protein) (PLPB1). | |||||
|
CROP_HUMAN
|
||||||
| NC score | 0.036194 (rank : 39) | θ value | 4.03905 (rank : 38) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O95232, Q6PHR9, Q9NUY0, Q9P2S7 | Gene names | CROP, CREAP1, O48 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cisplatin resistance-associated overexpressed protein (cAMP regulatory element-associated protein 1) (CRE-associated protein 1) (CREAP-1) (Luc7A) (Okadaic acid-inducible phosphoprotein OA48-18). | |||||
|
TATD2_HUMAN
|
||||||
| NC score | 0.035774 (rank : 40) | θ value | 5.27518 (rank : 57) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q93075 | Gene names | TATDN2, KIAA0218 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TatD DNase domain-containing deoxyribonuclease 2 (EC 3.1.21.-). | |||||
|
NKTR_HUMAN
|
||||||
| NC score | 0.035673 (rank : 41) | θ value | 6.88961 (rank : 62) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 658 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P30414 | Gene names | NKTR | |||
|
Domain Architecture |

|
|||||
| Description | NK-tumor recognition protein (Natural-killer cells cyclophilin-related protein) (NK-TR protein). | |||||
|
GBF1_HUMAN
|
||||||
| NC score | 0.032799 (rank : 42) | θ value | 2.36792 (rank : 27) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q92538, Q5VXX3, Q96CK6, Q96HZ3, Q9H473 | Gene names | GBF1, KIAA0248 | |||
|
Domain Architecture |

|
|||||
| Description | Golgi-specific brefeldin A-resistance guanine nucleotide exchange factor 1 (BFA-resistant GEF 1). | |||||
|
ANLN_MOUSE
|
||||||
| NC score | 0.031601 (rank : 43) | θ value | 4.03905 (rank : 35) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 297 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8K298, Q8BL79, Q8BLB3, Q8K2N0, Q9CUY0 | Gene names | Anln | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Actin-binding protein anillin. | |||||
|
RPGR1_MOUSE
|
||||||
| NC score | 0.031516 (rank : 44) | θ value | 4.03905 (rank : 45) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 713 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9EPQ2, Q8CAC2, Q8CDJ9, Q91WE0, Q9CUK6, Q9D5Q1 | Gene names | Rpgrip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator-interacting protein 1 (RPGR-interacting protein 1). | |||||
|
IQGA2_HUMAN
|
||||||
| NC score | 0.031078 (rank : 45) | θ value | 1.06291 (rank : 18) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q13576 | Gene names | IQGAP2 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP2. | |||||
|
NFH_HUMAN
|
||||||
| NC score | 0.027096 (rank : 46) | θ value | 1.81305 (rank : 26) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
NFH_MOUSE
|
||||||
| NC score | 0.026399 (rank : 47) | θ value | 5.27518 (rank : 52) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
LONM_HUMAN
|
||||||
| NC score | 0.026028 (rank : 48) | θ value | 5.27518 (rank : 51) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P36776, P36777, Q9UQ95 | Gene names | PRSS15 | |||
|
Domain Architecture |

|
|||||
| Description | Lon protease homolog, mitochondrial precursor (EC 3.4.21.-) (Lon protease-like protein) (LONP) (LONHs). | |||||
|
NIN_MOUSE
|
||||||
| NC score | 0.026013 (rank : 49) | θ value | 3.0926 (rank : 33) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q61043, Q6ZPM7 | Gene names | Nin, Kiaa1565 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ninein. | |||||
|
SRCA_HUMAN
|
||||||
| NC score | 0.025644 (rank : 50) | θ value | 8.99809 (rank : 74) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q86TD4 | Gene names | SRL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcalumenin precursor. | |||||
|
DYHC_HUMAN
|
||||||
| NC score | 0.024148 (rank : 51) | θ value | 8.99809 (rank : 68) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 733 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q14204, Q6DKQ7, Q8WU28, Q92814, Q9Y4G5 | Gene names | DYNC1H1, DNCH1, DNECL, KIAA0325 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dynein heavy chain, cytosolic (DYHC) (Cytoplasmic dynein heavy chain 1) (DHC1) (Dynein heavy chain 1, cytoplasmic 1). | |||||
|
DYHC_MOUSE
|
||||||
| NC score | 0.024018 (rank : 52) | θ value | 8.99809 (rank : 69) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 729 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9JHU4 | Gene names | Dync1h1, Dnch1, Dnchc1 | |||
|
Domain Architecture |

|
|||||
| Description | Dynein heavy chain, cytosolic (DYHC) (Cytoplasmic dynein heavy chain 1) (DHC1) (Dynein heavy chain 1, cytoplasmic 1). | |||||
|
CNGA1_MOUSE
|
||||||
| NC score | 0.022142 (rank : 53) | θ value | 1.81305 (rank : 25) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P29974, Q60776 | Gene names | Cnga1, Cncg, Cncg1 | |||
|
Domain Architecture |

|
|||||
| Description | cGMP-gated cation channel alpha 1 (CNG channel alpha 1) (CNG-1) (CNG1) (Cyclic nucleotide-gated channel alpha 1) (Cyclic nucleotide-gated channel, photoreceptor) (Cyclic nucleotide-gated cation channel 1) (Rod photoreceptor cGMP-gated channel subunit alpha). | |||||
|
CNGB1_HUMAN
|
||||||
| NC score | 0.020643 (rank : 54) | θ value | 8.99809 (rank : 67) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14028, Q14029 | Gene names | CNGB1, CNCG4 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic nucleotide-gated cation channel 4 (CNG channel 4) (CNG-4) (CNG4) (Cyclic nucleotide-gated cation channel modulatory subunit). | |||||
|
CP250_HUMAN
|
||||||
| NC score | 0.020495 (rank : 55) | θ value | 5.27518 (rank : 48) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1805 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9BV73, O14812, O60588, Q9H450 | Gene names | CEP250, CEP2, CNAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Centrosome-associated protein CEP250 (Centrosomal protein 2) (Centrosomal Nek2-associated protein 1) (C-Nap1). | |||||
|
RBM13_MOUSE
|
||||||
| NC score | 0.020371 (rank : 56) | θ value | 8.99809 (rank : 73) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BGS0, Q921E0, Q9D5G6 | Gene names | Rbm13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MAK16-like protein RBM13 (RNA-binding motif protein 13). | |||||
|
CHD1_HUMAN
|
||||||
| NC score | 0.019590 (rank : 57) | θ value | 1.38821 (rank : 21) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O14646 | Gene names | CHD1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
EMIL2_MOUSE
|
||||||
| NC score | 0.019590 (rank : 58) | θ value | 3.0926 (rank : 32) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 485 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8K482 | Gene names | Emilin2 | |||
|
Domain Architecture |

|
|||||
| Description | EMILIN-2 precursor (Elastin microfibril interface-located protein 2) (Elastin microfibril interfacer 2) (Basilin). | |||||
|
SYNE1_HUMAN
|
||||||
| NC score | 0.019312 (rank : 59) | θ value | 5.27518 (rank : 56) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1484 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8NF91, O94890, Q5JV19, Q5JV22, Q8N9P7, Q8TCP1, Q8WWW6, Q8WWW7, Q8WXF6, Q96N17, Q9C0A7, Q9H525, Q9H526, Q9NS36, Q9NU50, Q9UJ06, Q9UJ07, Q9ULF8 | Gene names | SYNE1, KIAA0796, KIAA1262, KIAA1756, MYNE1 | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-1 (Nuclear envelope spectrin repeat protein 1) (Synaptic nuclear envelope protein 1) (Syne-1) (Myocyte nuclear envelope protein 1) (Myne-1) (Enaptin). | |||||
|
CCD27_HUMAN
|
||||||
| NC score | 0.018666 (rank : 60) | θ value | 6.88961 (rank : 58) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 594 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q2M243, Q5TBV3, Q96M50 | Gene names | CCDC27 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 27. | |||||
|
CENPE_HUMAN
|
||||||
| NC score | 0.018596 (rank : 61) | θ value | 5.27518 (rank : 47) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 2060 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q02224 | Gene names | CENPE | |||
|
Domain Architecture |

|
|||||
| Description | Centromeric protein E (CENP-E). | |||||
|
TTBK1_HUMAN
|
||||||
| NC score | 0.017083 (rank : 62) | θ value | 0.813845 (rank : 16) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1058 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q5TCY1, Q2L6C6, Q8N444, Q96JH2 | Gene names | TTBK1, BDTK, KIAA1855 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tau-tubulin kinase 1 (EC 2.7.11.1) (Brain-derived tau kinase). | |||||
|
NFL_HUMAN
|
||||||
| NC score | 0.017036 (rank : 63) | θ value | 8.99809 (rank : 72) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 666 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P07196, Q16154, Q8IU72 | Gene names | NEFL, NF68, NFL | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet L protein (68 kDa neurofilament protein) (Neurofilament light polypeptide) (NF-L). | |||||
|
GA2L2_HUMAN
|
||||||
| NC score | 0.016564 (rank : 64) | θ value | 6.88961 (rank : 59) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8NHY3, Q8NHY4 | Gene names | GAS2L2, GAR17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GAS2-like protein 2 (Growth arrest-specific 2-like 2) (GAS2-related protein on chromosome 17) (GAR17 protein). | |||||
|
TRHY_HUMAN
|
||||||
| NC score | 0.016560 (rank : 65) | θ value | 6.88961 (rank : 64) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1385 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q07283 | Gene names | THH, THL, TRHY | |||
|
Domain Architecture |

|
|||||
| Description | Trichohyalin. | |||||
|
PPWD1_MOUSE
|
||||||
| NC score | 0.016425 (rank : 66) | θ value | 4.03905 (rank : 43) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8CEC6 | Gene names | Ppwd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peptidylprolyl isomerase domain and WD repeat-containing protein 1 (EC 5.2.1.8). | |||||
|
CHIC1_HUMAN
|
||||||
| NC score | 0.014686 (rank : 67) | θ value | 8.99809 (rank : 66) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5VXU3 | Gene names | CHIC1, BRX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cysteine-rich hydrophobic domain 1 protein (Brain X-linked protein). | |||||
|
PARC_MOUSE
|
||||||
| NC score | 0.014400 (rank : 68) | θ value | 5.27518 (rank : 53) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80TT8, Q8BKL9, Q8CGC0, Q8R363 | Gene names | Parc, Kiaa0708 | |||
|
Domain Architecture |

|
|||||
| Description | p53-associated parkin-like cytoplasmic protein. | |||||
|
FGD5_HUMAN
|
||||||
| NC score | 0.014308 (rank : 69) | θ value | 4.03905 (rank : 42) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6ZNL6, Q6MZY1, Q7Z303, Q8IYP3, Q8N861, Q8N8G4 | Gene names | FGD5, ZFYVE23 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 5 (Zinc finger FYVE domain-containing protein 23). | |||||
|
SMRCD_MOUSE
|
||||||
| NC score | 0.013180 (rank : 70) | θ value | 6.88961 (rank : 63) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q04692 | Gene names | Smarcad1, Etl1, Kiaa1122 | |||
|
Domain Architecture |

|
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A containing DEAD/H box 1 (EC 3.6.1.-) (ATP- dependent helicase SMARCAD1) (Enhancer trap locus homolog 1) (Etl-1). | |||||
|
DHX57_MOUSE
|
||||||
| NC score | 0.012974 (rank : 71) | θ value | 5.27518 (rank : 49) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6P5D3, Q3TS93, Q6NZK4, Q6P1B4, Q8BI63, Q8BIA2, Q8R360 | Gene names | Dhx57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative ATP-dependent RNA helicase DHX57 (EC 3.6.1.-) (DEAH box protein 57). | |||||
|
HUWE1_HUMAN
|
||||||
| NC score | 0.012715 (rank : 72) | θ value | 6.88961 (rank : 60) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q7Z6Z7, O15029, Q4G2Z2, Q5H961, Q6P4D0, Q8NG67, Q9BUI0, Q9HCJ4, Q9NSL6, Q9P0A9 | Gene names | HUWE1, KIAA0312, KIAA1578, UREB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (Mcl-1 ubiquitin ligase E3) (Mule) (ARF-binding protein 1) (ARF-BP1). | |||||
|
DDX24_MOUSE
|
||||||
| NC score | 0.012271 (rank : 73) | θ value | 3.0926 (rank : 31) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9ESV0, Q61119 | Gene names | Ddx24 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent RNA helicase DDX24 (EC 3.6.1.-) (DEAD box protein 24). | |||||
|
ATBF1_MOUSE
|
||||||
| NC score | 0.011470 (rank : 74) | θ value | 4.03905 (rank : 36) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1841 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q61329 | Gene names | Atbf1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-fetoprotein enhancer-binding protein (AT motif-binding factor) (AT-binding transcription factor 1). | |||||
|
U3IP2_HUMAN
|
||||||
| NC score | 0.010523 (rank : 75) | θ value | 4.03905 (rank : 46) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O43818, Q8IZ30 | Gene names | RNU3IP2, U355K | |||
|
Domain Architecture |

|
|||||
| Description | U3 small nucleolar RNA-interacting protein 2 (U3 small nucleolar ribonucleoprotein-associated 55 kDa protein) (U3 snoRNP-associated 55 kDa protein) (U3-55K). | |||||
|
CD2L1_HUMAN
|
||||||
| NC score | 0.007090 (rank : 76) | θ value | 4.03905 (rank : 37) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1467 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P21127, O95265, Q12817, Q12818, Q12819, Q12820, Q12822, Q8N530, Q9NZS5, Q9UBJ0, Q9UBQ1, Q9UBR0, Q9UNY2, Q9UP57, Q9UP58, Q9UP59 | Gene names | CDC2L1, PITSLREA, PK58 | |||
|
Domain Architecture |

|
|||||
| Description | PITSLRE serine/threonine-protein kinase CDC2L1 (EC 2.7.11.22) (Galactosyltransferase-associated protein kinase p58/GTA) (Cell division cycle 2-like protein kinase 1) (CLK-1) (CDK11) (p58 CLK-1). | |||||
|
CD2L1_MOUSE
|
||||||
| NC score | 0.006486 (rank : 77) | θ value | 8.99809 (rank : 65) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1463 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P24788, Q3UI03, Q61399, Q7TST4, Q8BP53 | Gene names | Cdc2l1 | |||
|
Domain Architecture |

|
|||||
| Description | PITSLRE serine/threonine-protein kinase CDC2L1 (EC 2.7.11.22) (Galactosyltransferase-associated protein kinase p58/GTA) (Cell division cycle 2-like protein kinase 1). | |||||
|
ZN202_HUMAN
|
||||||
| NC score | 0.003318 (rank : 78) | θ value | 8.99809 (rank : 75) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 756 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O95125, Q4JG21, Q9H1B9, Q9NSM4 | Gene names | ZNF202 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 202. | |||||