Please be patient as the page loads
|
MEF2D_MOUSE
|
||||||
| SwissProt Accessions | Q63943, Q63944 | Gene names | Mef2d | |||
|
Domain Architecture |
|
|||||
| Description | Myocyte-specific enhancer factor 2D. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
MEF2D_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.981474 (rank : 2) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 102 | |
| SwissProt Accessions | Q14814, Q14815 | Gene names | MEF2D | |||
|
Domain Architecture |
|
|||||
| Description | Myocyte-specific enhancer factor 2D. | |||||
|
MEF2D_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 146 | |
| SwissProt Accessions | Q63943, Q63944 | Gene names | Mef2d | |||
|
Domain Architecture |

|
|||||
| Description | Myocyte-specific enhancer factor 2D. | |||||
|
MEF2C_MOUSE
|
||||||
| θ value | 1.09934e-114 (rank : 3) | NC score | 0.945284 (rank : 5) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8CFN5, Q8R0H1, Q9D7L0, Q9QW20 | Gene names | Mef2c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myocyte-specific enhancer factor 2C. | |||||
|
MEF2C_HUMAN
|
||||||
| θ value | 1.8752e-114 (rank : 4) | NC score | 0.942816 (rank : 6) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q06413 | Gene names | MEF2C | |||
|
Domain Architecture |
|
|||||
| Description | Myocyte-specific enhancer factor 2C. | |||||
|
MEF2A_MOUSE
|
||||||
| θ value | 2.1455e-110 (rank : 5) | NC score | 0.957010 (rank : 3) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q60929 | Gene names | Mef2a | |||
|
Domain Architecture |
|
|||||
| Description | Myocyte-specific enhancer factor 2A. | |||||
|
MEF2A_HUMAN
|
||||||
| θ value | 2.6227e-108 (rank : 6) | NC score | 0.954101 (rank : 4) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q02078, O43814, Q14223, Q14224, Q96D14 | Gene names | MEF2A, MEF2 | |||
|
Domain Architecture |
|
|||||
| Description | Myocyte-specific enhancer factor 2A (Serum response factor-like protein 1). | |||||
|
MEF2B_HUMAN
|
||||||
| θ value | 1.01529e-43 (rank : 7) | NC score | 0.884881 (rank : 7) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q02080 | Gene names | MEF2B, XMEF2 | |||
|
Domain Architecture |
|
|||||
| Description | Myocyte-specific enhancer factor 2B (Serum response factor-like protein 2) (XMEF2) (RSRFR2). | |||||
|
MEF2B_MOUSE
|
||||||
| θ value | 5.57106e-42 (rank : 8) | NC score | 0.838483 (rank : 8) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O55087, O55088, O55089, O55090, O55231, Q61843 | Gene names | Mef2b | |||
|
Domain Architecture |
|
|||||
| Description | Myocyte-specific enhancer factor 2B. | |||||
|
SRF_HUMAN
|
||||||
| θ value | 7.59969e-07 (rank : 9) | NC score | 0.457295 (rank : 10) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P11831 | Gene names | SRF | |||
|
Domain Architecture |

|
|||||
| Description | Serum response factor (SRF). | |||||
|
SRF_MOUSE
|
||||||
| θ value | 7.59969e-07 (rank : 10) | NC score | 0.460674 (rank : 9) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9JM73 | Gene names | Srf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serum response factor (SRF). | |||||
|
MLL2_HUMAN
|
||||||
| θ value | 0.00035302 (rank : 11) | NC score | 0.089701 (rank : 12) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 87 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
ADA12_MOUSE
|
||||||
| θ value | 0.000461057 (rank : 12) | NC score | 0.032071 (rank : 75) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 474 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q61824 | Gene names | Adam12, Mltna | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 12 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 12) (Meltrin alpha). | |||||
|
K1802_HUMAN
|
||||||
| θ value | 0.000786445 (rank : 13) | NC score | 0.055584 (rank : 32) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1378 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q96JM3, Q6P181, Q8NC88, Q9BST0 | Gene names | KIAA1802, C13orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
ATX7_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 14) | NC score | 0.089852 (rank : 11) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O15265, O75328, O75329, Q9Y6P8 | Gene names | ATXN7, SCA7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-7 (Spinocerebellar ataxia type 7 protein). | |||||
|
EVI2B_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 15) | NC score | 0.088460 (rank : 13) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8VD58, Q3U1A3 | Gene names | Evi2b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EVI2B protein precursor (Ecotropic viral integration site 2B protein). | |||||
|
MLL3_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 16) | NC score | 0.055991 (rank : 29) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
IRF7_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 17) | NC score | 0.030845 (rank : 79) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P70434 | Gene names | Irf7 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon regulatory factor 7 (IRF-7). | |||||
|
TARSH_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 18) | NC score | 0.068229 (rank : 21) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 768 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q7Z7G0, Q6ZW20, Q6ZW22, Q9C082, Q9UFI6 | Gene names | ABI3BP, NESHBP, TARSH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Target of Nesh-SH3 precursor (Tarsh) (Nesh-binding protein) (NeshBP) (ABI gene family member 3-binding protein). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | 0.125558 (rank : 19) | NC score | 0.060710 (rank : 23) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 86 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
HCN2_HUMAN
|
||||||
| θ value | 0.163984 (rank : 20) | NC score | 0.052337 (rank : 37) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 369 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9UL51, O60742, O60743, O75267, Q9UBS2 | Gene names | HCN2, BCNG2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 2 (Brain cyclic nucleotide-gated channel 2) (BCNG-2). | |||||
|
ZHX2_MOUSE
|
||||||
| θ value | 0.163984 (rank : 21) | NC score | 0.027115 (rank : 93) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8C0C0 | Gene names | Zhx2, Afr1, Raf | |||
|
Domain Architecture |

|
|||||
| Description | Zinc fingers and homeoboxes protein 2 (Zinc finger and homeodomain protein 2) (Alpha-fetoprotein regulator 1) (AFP regulator 1) (Regulator of AFP). | |||||
|
NFRKB_MOUSE
|
||||||
| θ value | 0.21417 (rank : 22) | NC score | 0.069914 (rank : 19) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 393 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q6PIJ4, Q8BWV5, Q8K0X6 | Gene names | Nfrkb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear factor related to kappa-B-binding protein (DNA-binding protein R kappa-B). | |||||
|
PCLO_HUMAN
|
||||||
| θ value | 0.21417 (rank : 23) | NC score | 0.060273 (rank : 24) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 91 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
SSBP4_HUMAN
|
||||||
| θ value | 0.21417 (rank : 24) | NC score | 0.051315 (rank : 41) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 753 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9BWG4 | Gene names | SSBP4 | |||
|
Domain Architecture |

|
|||||
| Description | Single-stranded DNA-binding protein 4. | |||||
|
SOX6_MOUSE
|
||||||
| θ value | 0.279714 (rank : 25) | NC score | 0.035340 (rank : 68) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P40645, Q62250, Q9QWS5 | Gene names | Sox6, Sox-6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-6 (SOX-LZ). | |||||
|
CD248_HUMAN
|
||||||
| θ value | 0.365318 (rank : 26) | NC score | 0.039647 (rank : 61) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 679 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9HCU0, Q3SX55, Q96KB6 | Gene names | CD248, CD164L1, TEM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endosialin precursor (Tumor endothelial marker 1) (CD248 antigen). | |||||
|
GP152_HUMAN
|
||||||
| θ value | 0.365318 (rank : 27) | NC score | 0.020021 (rank : 115) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 636 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8TDT2, Q86SM0 | Gene names | GPR152, PGR5 | |||
|
Domain Architecture |

|
|||||
| Description | Probable G-protein coupled receptor 152 (G-protein coupled receptor PGR5). | |||||
|
GPR98_HUMAN
|
||||||
| θ value | 0.365318 (rank : 28) | NC score | 0.029324 (rank : 81) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8WXG9, O75171, Q9H0X5, Q9UL61 | Gene names | GPR98, KIAA0686, MASS1, VLGR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G-protein coupled receptor 98 precursor (Monogenic audiogenic seizure susceptibility protein 1 homolog) (Very large G-protein coupled receptor 1) (Usher syndrome type-2C protein). | |||||
|
SPEG_MOUSE
|
||||||
| θ value | 0.365318 (rank : 29) | NC score | 0.011798 (rank : 137) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
SPHK2_MOUSE
|
||||||
| θ value | 0.365318 (rank : 30) | NC score | 0.044910 (rank : 53) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9JIA7, Q91VA9, Q9DBH6 | Gene names | Sphk2 | |||
|
Domain Architecture |

|
|||||
| Description | Sphingosine kinase 2 (EC 2.7.1.-) (SK 2) (SPK 2). | |||||
|
FOXP4_HUMAN
|
||||||
| θ value | 0.47712 (rank : 31) | NC score | 0.027411 (rank : 91) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8IVH2, Q96E19 | Gene names | FOXP4, FKHLA | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P4 (Fork head-related protein-like A). | |||||
|
HCN4_MOUSE
|
||||||
| θ value | 0.47712 (rank : 32) | NC score | 0.046328 (rank : 51) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 284 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O70507 | Gene names | Hcn4, Bcng3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 4 (Brain cyclic nucleotide-gated channel 3) (BCNG-3). | |||||
|
NCOA3_MOUSE
|
||||||
| θ value | 0.47712 (rank : 33) | NC score | 0.040449 (rank : 60) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O09000, Q9CRD5 | Gene names | Ncoa3, Aib1, Pcip, Rac3, Tram1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein homolog) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP- interacting protein) (p/CIP) (pCIP). | |||||
|
PCBP4_HUMAN
|
||||||
| θ value | 0.62314 (rank : 34) | NC score | 0.017659 (rank : 119) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P57723, Q96AH7 | Gene names | PCBP4 | |||
|
Domain Architecture |
|
|||||
| Description | Poly(rC)-binding protein 4 (Alpha-CP4). | |||||
|
PCLO_MOUSE
|
||||||
| θ value | 0.62314 (rank : 35) | NC score | 0.055851 (rank : 30) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 90 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
PRB3_HUMAN
|
||||||
| θ value | 0.62314 (rank : 36) | NC score | 0.051058 (rank : 43) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q04118, Q15188, Q7M4M9 | Gene names | PRB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 3 precursor (Parotid salivary glycoprotein G1) (Proline-rich protein G1). | |||||
|
UBQL3_HUMAN
|
||||||
| θ value | 0.62314 (rank : 37) | NC score | 0.022992 (rank : 108) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9H347, Q9NRE0 | Gene names | UBQLN3 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquilin-3. | |||||
|
AFF2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 38) | NC score | 0.041744 (rank : 56) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P51816, O43786, O60215, P78407, Q13521, Q14323, Q9UNA5 | Gene names | AFF2, FMR2, OX19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation 2 protein) (Protein FMR-2) (FMR2P) (Protein Ox19) (Fragile X E mental retardation syndrome protein). | |||||
|
BORG1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 39) | NC score | 0.025671 (rank : 102) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O14613, Q9UNS0 | Gene names | CDC42EP2, BORG1, CEP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cdc42 effector protein 2 (Binder of Rho GTPases 1). | |||||
|
BSN_MOUSE
|
||||||
| θ value | 0.813845 (rank : 40) | NC score | 0.051432 (rank : 40) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 91 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
ES8L3_HUMAN
|
||||||
| θ value | 0.813845 (rank : 41) | NC score | 0.018368 (rank : 118) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8TE67, Q5T8Q6, Q5T8Q7, Q5T8Q8, Q96E47, Q9H719 | Gene names | EPS8L3, EPS8R3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Epidermal growth factor receptor kinase substrate 8-like protein 3 (Epidermal growth factor receptor pathway substrate 8-related protein 3) (EPS8-like protein 3). | |||||
|
FBN2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 42) | NC score | 0.009552 (rank : 143) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P35556 | Gene names | FBN2 | |||
|
Domain Architecture |

|
|||||
| Description | Fibrillin-2 precursor. | |||||
|
GNAS1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 43) | NC score | 0.026236 (rank : 96) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 651 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q6R0H7, Q6R0H4, Q6R0H5, Q6R2J5, Q9JJX0, Q9Z1N8 | Gene names | Gnas, Gnas1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Guanine nucleotide-binding protein G(s) subunit alpha isoforms XLas (Adenylate cyclase-stimulating G alpha protein) (Extra large alphas protein) (XLalphas). | |||||
|
IPMK_HUMAN
|
||||||
| θ value | 0.813845 (rank : 44) | NC score | 0.035486 (rank : 65) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NFU5 | Gene names | IPMK, IMPK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol polyphosphate multikinase (EC 2.7.1.151) (Inositol 1,3,4,6- tetrakisphosphate 5-kinase). | |||||
|
SMR2C_MOUSE
|
||||||
| θ value | 0.813845 (rank : 45) | NC score | 0.053233 (rank : 36) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O35985 | Gene names | Smr2, Msg2, Vcs2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Submaxillary gland androgen-regulated protein 2, isoform gamma precursor (Salivary protein MSG2, isoform gamma). | |||||
|
SMR2D_MOUSE
|
||||||
| θ value | 0.813845 (rank : 46) | NC score | 0.054200 (rank : 33) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O35979 | Gene names | Smr2, Msg2, Vcs2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Submaxillary gland androgen-regulated protein 2, isoform delta precursor (Salivary protein MSG2, isoform delta). | |||||
|
CN004_HUMAN
|
||||||
| θ value | 1.06291 (rank : 47) | NC score | 0.047259 (rank : 50) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9H1B7, Q8NDQ2, Q96JG2, Q9H3I7 | Gene names | C14orf4, KIAA1865 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf4. | |||||
|
CUTL1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 48) | NC score | 0.021068 (rank : 113) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P53564, O08994, P70301, Q571L6, Q91ZD2 | Gene names | Cutl1, Cux, Cux1, Kiaa4047 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP) (Homeobox protein Cux). | |||||
|
FOXC1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 49) | NC score | 0.023463 (rank : 107) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q12948, Q86UP7, Q9BYM1, Q9NUE5, Q9UDD0, Q9UP06 | Gene names | FOXC1, FKHL7, FREAC3 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein C1 (Forkhead-related protein FKHL7) (Forkhead- related transcription factor 3) (FREAC-3). | |||||
|
FOXC1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 50) | NC score | 0.024997 (rank : 105) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q61572, O88409, Q61582 | Gene names | Foxc1, Fkh1, Fkhl7, Freac3, Mf1 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein C1 (Forkhead-related protein FKHL7) (Forkhead- related transcription factor 3) (FREAC-3) (Transcription factor FKH-1) (Mesoderm/mesenchyme forkhead 1) (MF-1). | |||||
|
FOXN4_HUMAN
|
||||||
| θ value | 1.06291 (rank : 51) | NC score | 0.018372 (rank : 117) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q96NZ1, Q6ZMR4, Q96NZ0 | Gene names | FOXN4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein N4. | |||||
|
HCN2_MOUSE
|
||||||
| θ value | 1.06291 (rank : 52) | NC score | 0.041729 (rank : 57) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O88703, O70506 | Gene names | Hcn2, Bcng2, Hac1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 2 (Brain cyclic nucleotide-gated channel 2) (BCNG-2) (Hyperpolarization-activated cation channel 1) (HAC-1). | |||||
|
IF4G1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 53) | NC score | 0.038392 (rank : 62) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 534 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q6NZJ6, Q6NZN8, Q8BW99 | Gene names | Eif4g1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1). | |||||
|
UTX_HUMAN
|
||||||
| θ value | 1.06291 (rank : 54) | NC score | 0.035450 (rank : 66) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O15550 | Gene names | UTX | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitously transcribed X chromosome tetratricopeptide repeat protein (Ubiquitously transcribed TPR protein on the X chromosome). | |||||
|
ABCA7_HUMAN
|
||||||
| θ value | 1.38821 (rank : 55) | NC score | 0.008361 (rank : 147) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IZY2, Q96S58, Q9BZC4, Q9NR73, Q9UKP8 | Gene names | ABCA7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-binding cassette sub-family A member 7 (Macrophage ABC transporter) (Autoantigen SS-N) (ABCA-SSN). | |||||
|
ARRD1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 56) | NC score | 0.034718 (rank : 70) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99KN1, Q3TDT6, Q8VEG7 | Gene names | Arrdc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arrestin domain-containing protein 1. | |||||
|
ECM1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 57) | NC score | 0.044924 (rank : 52) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 182 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q16610 | Gene names | ECM1 | |||
|
Domain Architecture |

|
|||||
| Description | Extracellular matrix protein 1 precursor (Secretory component p85). | |||||
|
GP179_HUMAN
|
||||||
| θ value | 1.38821 (rank : 58) | NC score | 0.031700 (rank : 76) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 372 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6PRD1 | Gene names | GPR179, GPR158L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 179 precursor (Probable G-protein coupled receptor 158-like 1). | |||||
|
MINT_HUMAN
|
||||||
| θ value | 1.38821 (rank : 59) | NC score | 0.034476 (rank : 71) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1291 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q96T58, Q9H9A8, Q9NWH5, Q9UQ01, Q9Y556 | Gene names | SPEN, KIAA0929, MINT, SHARP | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
PKN3_MOUSE
|
||||||
| θ value | 1.38821 (rank : 60) | NC score | 0.000959 (rank : 157) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1009 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8K045 | Gene names | Pkn3, Pknbeta | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase N3 (EC 2.7.11.13) (Protein kinase PKN- beta) (Protein-kinase C-related kinase 3). | |||||
|
PRP1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 61) | NC score | 0.066692 (rank : 22) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P04280, Q08805, Q15186, Q15187, Q15214, Q15215, Q16038 | Gene names | PRB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 1 precursor (Salivary proline-rich protein) [Contains: Basic peptide IB-6; Peptide P-H]. | |||||
|
PSL1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 62) | NC score | 0.043651 (rank : 54) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8TCT7, O60365, Q567S3, Q8IUH9, Q9BUY6, Q9H3M4, Q9NPN2, Q9P1Z6 | Gene names | SPPL2B, KIAA1532, PSL1 | |||
|
Domain Architecture |

|
|||||
| Description | Signal peptide peptidase-like 2B (EC 3.4.23.-) (Protein SPP-like 2B) (Protein SPPL2b) (Intramembrane protease 4) (IMP4) (Presenilin-like protein 1). | |||||
|
RIF1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 63) | NC score | 0.019999 (rank : 116) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6PR54, Q6T8D9, Q7TPD8, Q8BTU2, Q8C408, Q9CS77 | Gene names | Rif1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Telomere-associated protein RIF1 (Rap1-interacting factor 1 homolog) (mRif1). | |||||
|
SEM6C_MOUSE
|
||||||
| θ value | 1.38821 (rank : 64) | NC score | 0.012754 (rank : 135) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9WTM3 | Gene names | Sema6c, Semay | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6C precursor (Semaphorin Y) (Sema Y). | |||||
|
BAT2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 65) | NC score | 0.083269 (rank : 15) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
CUTL1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 66) | NC score | 0.016497 (rank : 125) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1574 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P39880, Q6NYH4, Q9UEV5 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP). | |||||
|
FARP2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 67) | NC score | 0.011741 (rank : 138) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O94887, Q53QM5, Q8WU27, Q9UFE7 | Gene names | FARP2, KIAA0793 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FERM, RhoGEF and pleckstrin domain-containing protein 2 (FERM domain including RhoGEF) (FIR). | |||||
|
HNF1A_MOUSE
|
||||||
| θ value | 1.81305 (rank : 68) | NC score | 0.047321 (rank : 49) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P22361 | Gene names | Tcf1, Hnf-1, Hnf-1a, Hnf1a | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 1-alpha (HNF-1A) (Liver-specific transcription factor LF-B1) (LFB1). | |||||
|
JUND_MOUSE
|
||||||
| θ value | 1.81305 (rank : 69) | NC score | 0.025878 (rank : 100) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P15066 | Gene names | Jund, Jun-d, Jund1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor jun-D. | |||||
|
LPP_HUMAN
|
||||||
| θ value | 1.81305 (rank : 70) | NC score | 0.025922 (rank : 99) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 775 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q93052, Q8NFX5 | Gene names | LPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipoma-preferred partner (LIM domain-containing preferred translocation partner in lipoma). | |||||
|
SOX13_MOUSE
|
||||||
| θ value | 1.81305 (rank : 71) | NC score | 0.025025 (rank : 104) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 495 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q04891, Q922L3 | Gene names | Sox13, Sox-13 | |||
|
Domain Architecture |

|
|||||
| Description | SOX-13 protein. | |||||
|
CRTC1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 72) | NC score | 0.042775 (rank : 55) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q68ED7, Q6ZQ85 | Gene names | Crtc1, Kiaa0616, Mect1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CREB-regulated transcription coactivator 1 (Mucoepidermoid carcinoma translocated protein 1 homolog). | |||||
|
GAK7_HUMAN
|
||||||
| θ value | 2.36792 (rank : 73) | NC score | 0.027462 (rank : 90) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q96PI4, Q9QC08 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_1q23.3 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K18 Gag protein) (HERV-K110 Gag protein) (HERV-K(C1a) Gag protein) [Contains: Matrix protein]. | |||||
|
K0552_HUMAN
|
||||||
| θ value | 2.36792 (rank : 74) | NC score | 0.025777 (rank : 101) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 442 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O60299 | Gene names | KIAA0552 | |||
|
Domain Architecture |

|
|||||
| Description | Protein KIAA0552. | |||||
|
LBN_MOUSE
|
||||||
| θ value | 2.36792 (rank : 75) | NC score | 0.022662 (rank : 109) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8K1G2, Q8BRF3 | Gene names | Evc2, Lbn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Limbin. | |||||
|
PO2F2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 76) | NC score | 0.012487 (rank : 136) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 379 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P09086, Q16648, Q9BRS4 | Gene names | POU2F2, OCT2, OTF2 | |||
|
Domain Architecture |
|
|||||
| Description | POU domain, class 2, transcription factor 2 (Octamer-binding transcription factor 2) (Oct-2) (OTF-2) (Lymphoid-restricted immunoglobulin octamer-binding protein NF-A2). | |||||
|
RAPH1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 77) | NC score | 0.059858 (rank : 25) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q70E73, Q96Q37, Q9C0I2 | Gene names | RAPH1, ALS2CR9, KIAA1681, LPD, PREL2, RMO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras-associated and pleckstrin homology domains-containing protein 1 (RAPH1) (Lamellipodin) (Proline-rich EVH1 ligand 2) (PREL-2) (Protein RMO1) (Amyotrophic lateral sclerosis 2 chromosomal region candidate 9 gene protein). | |||||
|
RX_HUMAN
|
||||||
| θ value | 2.36792 (rank : 78) | NC score | 0.011320 (rank : 141) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 423 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9Y2V3 | Gene names | RAX, RX | |||
|
Domain Architecture |

|
|||||
| Description | Retinal homeobox protein Rx (Retina and anterior neural fold homeobox protein). | |||||
|
ATBF1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 79) | NC score | 0.016517 (rank : 124) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1841 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q61329 | Gene names | Atbf1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-fetoprotein enhancer-binding protein (AT motif-binding factor) (AT-binding transcription factor 1). | |||||
|
BCL9_MOUSE
|
||||||
| θ value | 3.0926 (rank : 80) | NC score | 0.047490 (rank : 48) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 770 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9D219, Q67FX9, Q8BUJ8, Q8VE74 | Gene names | Bcl9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9). | |||||
|
CD2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 81) | NC score | 0.031477 (rank : 77) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P06729, Q96TE5 | Gene names | CD2 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell surface antigen CD2 precursor (T-cell surface antigen T11/Leu- 5) (LFA-2) (LFA-3 receptor) (Erythrocyte receptor) (Rosette receptor). | |||||
|
FA47B_HUMAN
|
||||||
| θ value | 3.0926 (rank : 82) | NC score | 0.034441 (rank : 72) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 431 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8NA70, Q5JQN5, Q6PIG3 | Gene names | FAM47B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM47B. | |||||
|
IF4G2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 83) | NC score | 0.034855 (rank : 69) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P78344, O60877, P78404, Q53EU1, Q58EZ2, Q8NI71, Q96C16 | Gene names | EIF4G2, DAP5 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 2 (eIF-4-gamma 2) (eIF-4G 2) (eIF4G 2) (p97) (Death-associated protein 5) (DAP-5). | |||||
|
IF4G2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 84) | NC score | 0.034019 (rank : 74) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q62448, P97867, Q921U3 | Gene names | Eif4g2, Nat1 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 2 (eIF-4-gamma 2) (eIF-4G 2) (eIF4G 2) (p97) (Novel APOBEC-1 target 1) (Translation repressor NAT1). | |||||
|
MAP1B_MOUSE
|
||||||
| θ value | 3.0926 (rank : 85) | NC score | 0.024019 (rank : 106) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 977 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P14873 | Gene names | Map1b, Mtap1b, Mtap5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) (MAP1.2) (MAP1(X)) [Contains: MAP1 light chain LC1]. | |||||
|
SOX6_HUMAN
|
||||||
| θ value | 3.0926 (rank : 86) | NC score | 0.027028 (rank : 95) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P35712, Q9BXQ3, Q9BXQ4, Q9BXQ5, Q9H0I8 | Gene names | SOX6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-6. | |||||
|
US6NL_MOUSE
|
||||||
| θ value | 3.0926 (rank : 87) | NC score | 0.016733 (rank : 123) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q80XC3, Q6ZQK8 | Gene names | Usp6nl, Kiaa0019 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USP6 N-terminal-like protein. | |||||
|
BCL9_HUMAN
|
||||||
| θ value | 4.03905 (rank : 88) | NC score | 0.050582 (rank : 46) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1008 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | O00512 | Gene names | BCL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9) (Legless homolog). | |||||
|
CEL_HUMAN
|
||||||
| θ value | 4.03905 (rank : 89) | NC score | 0.029375 (rank : 80) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P19835, Q16398 | Gene names | CEL, BAL | |||
|
Domain Architecture |

|
|||||
| Description | Bile salt-activated lipase precursor (EC 3.1.1.3) (EC 3.1.1.13) (BAL) (Bile salt-stimulated lipase) (BSSL) (Carboxyl ester lipase) (Sterol esterase) (Cholesterol esterase) (Pancreatic lysophospholipase). | |||||
|
EPN4_MOUSE
|
||||||
| θ value | 4.03905 (rank : 90) | NC score | 0.027150 (rank : 92) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99KN9, Q8CFH4 | Gene names | Clint1, Enth, Epn4, Epnr, Kiaa0171 | |||
|
Domain Architecture |

|
|||||
| Description | Clathrin interactor 1 (Epsin-4) (Epsin-related protein) (EpsinR) (Enthoprotin). | |||||
|
ETV5_MOUSE
|
||||||
| θ value | 4.03905 (rank : 91) | NC score | 0.017449 (rank : 121) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9CXC9, Q3TG49, Q8C0F3, Q9JHB1 | Gene names | Etv5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 5. | |||||
|
EVI2B_HUMAN
|
||||||
| θ value | 4.03905 (rank : 92) | NC score | 0.040791 (rank : 59) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P34910 | Gene names | EVI2B, EVDB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EVI2B protein precursor (Ecotropic viral integration site 2B protein). | |||||
|
GLI2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 93) | NC score | 0.002164 (rank : 155) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 971 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P10070, O60252, O60253, O60254, O60255, Q15590, Q15591 | Gene names | GLI2, THP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein GLI2 (Tax helper protein). | |||||
|
HCN4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 94) | NC score | 0.035343 (rank : 67) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9Y3Q4, Q9UMQ7 | Gene names | HCN4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 4. | |||||
|
KBTB9_MOUSE
|
||||||
| θ value | 4.03905 (rank : 95) | NC score | 0.006659 (rank : 149) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q80T74 | Gene names | Kbtbd9, Kiaa1921 | |||
|
Domain Architecture |

|
|||||
| Description | Kelch repeat and BTB domain-containing protein 9. | |||||
|
MEGF9_MOUSE
|
||||||
| θ value | 4.03905 (rank : 96) | NC score | 0.015500 (rank : 128) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8BH27, Q8BWI4 | Gene names | Megf9, Egfl5, Kiaa0818 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 9 precursor (EGF-like domain-containing protein 5) (Multiple EGF-like domain protein 5). | |||||
|
MYO15_HUMAN
|
||||||
| θ value | 4.03905 (rank : 97) | NC score | 0.013768 (rank : 132) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9UKN7 | Gene names | MYO15A, MYO15 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-15 (Myosin XV) (Unconventional myosin-15). | |||||
|
PI5PA_HUMAN
|
||||||
| θ value | 4.03905 (rank : 98) | NC score | 0.034151 (rank : 73) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 749 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q15735, Q32M61, Q6ZTH6, Q8N902, Q9UDT9 | Gene names | PIB5PA, PIPP | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 4,5-bisphosphate 5-phosphatase A (EC 3.1.3.56). | |||||
|
TAF4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 99) | NC score | 0.056658 (rank : 28) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 687 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | O00268, Q5TBP6, Q99721, Q9BR40, Q9BX42 | Gene names | TAF4, TAF2C, TAF2C1, TAF4A, TAFII130, TAFII135 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 4 (TBP-associated factor 4) (Transcription initiation factor TFIID 135 kDa subunit) (TAF(II)135) (TAFII-135) (TAFII135) (TAFII-130) (TAFII130). | |||||
|
TAU_HUMAN
|
||||||
| θ value | 4.03905 (rank : 100) | NC score | 0.028615 (rank : 83) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 448 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P10636, P18518, Q14799, Q15549, Q15550, Q15551, Q9UDJ3, Q9UMH0, Q9UQ96 | Gene names | MAPT, MTBT1, TAU | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein tau (Neurofibrillary tangle protein) (Paired helical filament-tau) (PHF-tau). | |||||
|
TENS4_MOUSE
|
||||||
| θ value | 4.03905 (rank : 101) | NC score | 0.015412 (rank : 129) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8BZ33, Q3TCM8, Q7TNR5 | Gene names | Tns4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tensin-4 precursor. | |||||
|
TM108_MOUSE
|
||||||
| θ value | 4.03905 (rank : 102) | NC score | 0.041360 (rank : 58) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 336 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8BHE4, Q80WR9 | Gene names | Tmem108 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 108 precursor. | |||||
|
WEE1B_HUMAN
|
||||||
| θ value | 4.03905 (rank : 103) | NC score | 0.008594 (rank : 146) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 836 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P0C1S8 | Gene names | WEE1B, WEE2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wee1-like protein kinase 1B (EC 2.7.10.2) (Wee1B kinase). | |||||
|
WNK1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 104) | NC score | 0.009414 (rank : 144) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1158 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9H4A3, O15052, Q86WL5, Q8N673, Q9P1S9 | Gene names | WNK1, KDP, KIAA0344, PRKWNK1 | |||
|
Domain Architecture |
|
|||||
| Description | Serine/threonine-protein kinase WNK1 (EC 2.7.11.1) (Protein kinase with no lysine 1) (Protein kinase, lysine-deficient 1) (Kinase deficient protein). | |||||
|
ZN414_HUMAN
|
||||||
| θ value | 4.03905 (rank : 105) | NC score | 0.021179 (rank : 112) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 548 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q96IQ9 | Gene names | ZNF414 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 414. | |||||
|
4ET_HUMAN
|
||||||
| θ value | 5.27518 (rank : 106) | NC score | 0.015218 (rank : 130) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NRA8, Q8NCF2, Q9H708 | Gene names | EIF4ENIF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4E transporter (eIF4E transporter) (4E-T) (Eukaryotic translation initiation factor 4E nuclear import factor 1). | |||||
|
ATX1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 107) | NC score | 0.028398 (rank : 86) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P54254 | Gene names | Atxn1, Sca1 | |||
|
Domain Architecture |

|
|||||
| Description | Ataxin-1 (Spinocerebellar ataxia type 1 protein homolog). | |||||
|
BOLL_HUMAN
|
||||||
| θ value | 5.27518 (rank : 108) | NC score | 0.012804 (rank : 133) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8N9W6, Q969U3 | Gene names | BOLL, BOULE | |||
|
Domain Architecture |
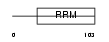
|
|||||
| Description | Protein boule-like. | |||||
|
CIZ1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 109) | NC score | 0.037784 (rank : 63) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9ULV3, Q9NYM8, Q9UHK4, Q9Y3F9, Q9Y3G0 | Gene names | CIZ1, LSFR1, NP94 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cip1-interacting zinc finger protein (Nuclear protein NP94). | |||||
|
CN037_MOUSE
|
||||||
| θ value | 5.27518 (rank : 110) | NC score | 0.027108 (rank : 94) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 179 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8BT18, Q8BFQ7, Q8R1Q5, Q9D6C5 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C14orf37 homolog precursor. | |||||
|
GAK11_HUMAN
|
||||||
| θ value | 5.27518 (rank : 111) | NC score | 0.028537 (rank : 85) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P63145, Q9UKI1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_22q11.21 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K101 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 112) | NC score | 0.028588 (rank : 84) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 415 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P63126, Q9UKH4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_6q14.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K109 Gag protein) (HERV-K(C6) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK5_HUMAN
|
||||||
| θ value | 5.27518 (rank : 113) | NC score | 0.031204 (rank : 78) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 429 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P62684 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19p13.11 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K113 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
LAMP3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 114) | NC score | 0.020318 (rank : 114) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7TST5, Q8C1F6 | Gene names | Lamp3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lysosome-associated membrane glycoprotein 3 precursor (LAMP-3) (Lysosomal-associated membrane protein 3) (DC-lysosome-associated membrane glycoprotein) (DC LAMP) (CD208 antigen). | |||||
|
MAGC1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 115) | NC score | 0.028138 (rank : 88) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 431 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O60732, O75451, Q8TCV4 | Gene names | MAGEC1 | |||
|
Domain Architecture |

|
|||||
| Description | Melanoma-associated antigen C1 (MAGE-C1 antigen) (Cancer-testis antigen CT7). | |||||
|
NCOA6_MOUSE
|
||||||
| θ value | 5.27518 (rank : 116) | NC score | 0.053683 (rank : 34) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1076 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9JL19, Q9JLT9 | Gene names | Ncoa6, Aib3, Prip, Rap250, Trbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC) (Thyroid hormone receptor-binding protein). | |||||
|
OGFR_MOUSE
|
||||||
| θ value | 5.27518 (rank : 117) | NC score | 0.025176 (rank : 103) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 584 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q99PG2, Q99L33 | Gene names | Ogfr | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor). | |||||
|
PENK_HUMAN
|
||||||
| θ value | 5.27518 (rank : 118) | NC score | 0.022229 (rank : 110) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P01210 | Gene names | PENK | |||
|
Domain Architecture |

|
|||||
| Description | Proenkephalin A precursor [Contains: Synenkephalin; Met-enkephalin (Opioid growth factor) (OGF); Met-enkephalin-Arg-Gly-Leu; Leu- enkephalin; Met-enkephalin-Arg-Phe]. | |||||
|
PRD10_HUMAN
|
||||||
| θ value | 5.27518 (rank : 119) | NC score | -0.001095 (rank : 158) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 769 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NQV6, Q9ULI9 | Gene names | PRDM10, KIAA1231, PFM7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PR domain zinc finger protein 10 (PR domain-containing protein 10). | |||||
|
ROBO3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 120) | NC score | 0.005736 (rank : 152) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 524 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Z2I4 | Gene names | Robo3, Rbig1 | |||
|
Domain Architecture |

|
|||||
| Description | Roundabout homolog 3 precursor (Retinoblastoma-inhibiting gene 1) (Rig-1). | |||||
|
SC6A5_MOUSE
|
||||||
| θ value | 5.27518 (rank : 121) | NC score | 0.006339 (rank : 150) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q761V0, Q8CFM5, Q91ZQ2 | Gene names | Slc6a5, Glyt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium- and chloride-dependent glycine transporter 2 (GlyT2) (GlyT-2) (Solute carrier family 6 member 5). | |||||
|
TAB3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 122) | NC score | 0.028369 (rank : 87) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q571K4 | Gene names | Map3k7ip3, Kiaa4135, Tab3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase 7-interacting protein 3 (TAK1-binding protein 3). | |||||
|
TTC28_HUMAN
|
||||||
| θ value | 5.27518 (rank : 123) | NC score | 0.014496 (rank : 131) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q96AY4, O95928, O95929, Q9NTE4, Q9UG31, Q9UGG5, Q9UPV8, Q9Y3S5 | Gene names | TTC28, KIAA1043 | |||
|
Domain Architecture |
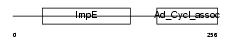
|
|||||
| Description | Tetratricopeptide repeat protein 28 (TPR repeat protein 28). | |||||
|
WNK1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 124) | NC score | 0.005256 (rank : 153) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1179 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P83741 | Gene names | Wnk1, Prkwnk1 | |||
|
Domain Architecture |
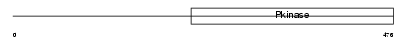
|
|||||
| Description | Serine/threonine-protein kinase WNK1 (EC 2.7.11.1) (Protein kinase with no lysine 1) (Protein kinase, lysine-deficient 1). | |||||
|
AP1G1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 125) | NC score | 0.011426 (rank : 140) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P22892 | Gene names | Ap1g1, Adtg, Clapg1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-1 complex subunit gamma-1 (Adapter-related protein complex 1 gamma- 1 subunit) (Gamma-adaptin) (Adaptor protein complex AP-1 gamma-1 subunit) (Golgi adaptor HA1/AP1 adaptin subunit gamma-1) (Clathrin assembly protein complex 1 gamma-1 large chain). | |||||
|
BAT2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 126) | NC score | 0.087283 (rank : 14) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q7TSC1, Q923A9, Q9Z1R1 | Gene names | Bat2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
GTSE1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 127) | NC score | 0.036776 (rank : 64) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8R080, O89015, Q9CSG9 | Gene names | Gtse1, B99 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G2 and S phase expressed protein 1 (Gtse-1) (B99 protein). | |||||
|
IQEC3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 128) | NC score | 0.012778 (rank : 134) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q3TES0, Q3UHP8, Q80TJ8 | Gene names | Iqsec3, Kiaa1110 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 3. | |||||
|
MB3L2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 129) | NC score | 0.021515 (rank : 111) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8NHZ7 | Gene names | MBD3L2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 3-like 2 (MBD3-like 2). | |||||
|
NCOA6_HUMAN
|
||||||
| θ value | 6.88961 (rank : 130) | NC score | 0.070863 (rank : 18) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1202 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | Q14686, Q9NTZ9, Q9UH74, Q9UK86 | Gene names | NCOA6, AIB3, KIAA0181, RAP250, TRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (PRIP) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC RAP250) (Thyroid hormone receptor-binding protein). | |||||
|
RAD9A_MOUSE
|
||||||
| θ value | 6.88961 (rank : 131) | NC score | 0.011735 (rank : 139) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Z0F6, Q8QZZ3 | Gene names | Rad9a, Rad9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell cycle checkpoint control protein RAD9A (EC 3.1.11.2) (DNA repair exonuclease rad9 homolog A) (Rad9-like protein) (mRAD9). | |||||
|
TCF8_MOUSE
|
||||||
| θ value | 6.88961 (rank : 132) | NC score | 0.001139 (rank : 156) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1067 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q64318, Q62519 | Gene names | Tcf8, Zfhx1a, Zfx1a, Zfx1ha | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 8 (Zinc finger homeobox protein 1a) (MEB1) (Delta EF1). | |||||
|
AP1G1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 133) | NC score | 0.010513 (rank : 142) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O43747, O75709, O75842, Q9UG09, Q9Y3U4 | Gene names | AP1G1, ADTG, CLAPG1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-1 complex subunit gamma-1 (Adapter-related protein complex 1 gamma- 1 subunit) (Gamma-adaptin) (Adaptor protein complex AP-1 gamma-1 subunit) (Golgi adaptor HA1/AP1 adaptin subunit gamma-1) (Clathrin assembly protein complex 1 gamma-1 large chain). | |||||
|
CAC1B_HUMAN
|
||||||
| θ value | 8.99809 (rank : 134) | NC score | 0.006309 (rank : 151) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q00975 | Gene names | CACNA1B, CACH5, CACNL1A5 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent N-type calcium channel subunit alpha-1B (Voltage- gated calcium channel subunit alpha Cav2.2) (Calcium channel, L type, alpha-1 polypeptide isoform 5) (Brain calcium channel III) (BIII). | |||||
|
DCP1A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 135) | NC score | 0.017573 (rank : 120) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q91YD3, Q6NZE3 | Gene names | Dcp1a, Mitc1, Smif | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | mRNA decapping enzyme 1A (EC 3.-.-.-) (Transcription factor SMIF) (MAD homolog 4-interacting transcription coactivator 1) (Smad4-interacting transcriptional co-activator). | |||||
|
GAK12_HUMAN
|
||||||
| θ value | 8.99809 (rank : 136) | NC score | 0.027818 (rank : 89) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P63130, Q9UKI0 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_1q22 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K102 Gag protein) (HERV-K(III) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 137) | NC score | 0.026187 (rank : 98) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q7LDI9, Q9UKH5, Q9Y6I1, Q9YNA6, Q9YNB0 | Gene names | ERVK6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_7p22.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K(HML-2.HOM) Gag protein) (HERV-K108 Gag protein) (HERV-K(C7) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK6_HUMAN
|
||||||
| θ value | 8.99809 (rank : 138) | NC score | 0.026199 (rank : 97) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P62685 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_8p23.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K115 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
HME1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 139) | NC score | 0.009399 (rank : 145) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P09065 | Gene names | En1, En-1 | |||
|
Domain Architecture |
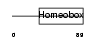
|
|||||
| Description | Homeobox protein engrailed-1 (Mo-En-1). | |||||
|
LR37A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 140) | NC score | 0.028810 (rank : 82) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 720 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O60309, Q49A01, Q49A80, Q8NB33 | Gene names | LRRC37A, KIAA0563 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 37A. | |||||
|
MACF4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 141) | NC score | 0.002319 (rank : 154) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1321 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q96PK2, Q8WXY1 | Gene names | MACF1, ABP620, ACF7, KIAA0465, KIAA1251 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1, isoform 4. | |||||
|
MAGE1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 142) | NC score | 0.050015 (rank : 47) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
PRRT3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 143) | NC score | 0.016163 (rank : 126) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q6PE13 | Gene names | Prrt3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
TCF7_MOUSE
|
||||||
| θ value | 8.99809 (rank : 144) | NC score | 0.017113 (rank : 122) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q00417 | Gene names | Tcf7, Tcf-1, Tcf1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 7 (T-cell-specific transcription factor 1) (TCF- 1) (T-cell factor 1). | |||||
|
VINEX_MOUSE
|
||||||
| θ value | 8.99809 (rank : 145) | NC score | 0.006718 (rank : 148) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9R1Z8, Q62423 | Gene names | Sorbs3, Scam1, Sh3d4 | |||
|
Domain Architecture |

|
|||||
| Description | Vinexin (Sorbin and SH3 domain-containing protein 3) (SH3-containing adapter molecule 1) (SCAM-1) (SH3 domain-containing protein SH3P3). | |||||
|
ZC3H3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 146) | NC score | 0.016048 (rank : 127) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8CHP0, Q80X78 | Gene names | Zc3h3, Kiaa0150, Zc3hdc3, Zfp623 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 3. | |||||
|
ATN1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 147) | NC score | 0.068624 (rank : 20) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
CNOT3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 148) | NC score | 0.051106 (rank : 42) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O75175, Q9NZN7, Q9UF76 | Gene names | CNOT3, KIAA0691, NOT3 | |||
|
Domain Architecture |

|
|||||
| Description | CCR4-NOT transcription complex subunit 3 (CCR4-associated factor 3). | |||||
|
CNOT3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 149) | NC score | 0.050730 (rank : 45) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8K0V4 | Gene names | Cnot3, Not3 | |||
|
Domain Architecture |

|
|||||
| Description | CCR4-NOT transcription complex subunit 3 (CCR4-associated factor 3). | |||||
|
PRP2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 150) | NC score | 0.058431 (rank : 26) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P05142 | Gene names | Prh1, Prp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein MP-2 precursor. | |||||
|
PRR12_HUMAN
|
||||||
| θ value | θ > 10 (rank : 151) | NC score | 0.050932 (rank : 44) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q9ULL5, Q8N4J6 | Gene names | PRR12, KIAA1205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 12. | |||||
|
SMR1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 152) | NC score | 0.051907 (rank : 38) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q61900 | Gene names | Smr1, Msg1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Submaxillary gland androgen-regulated protein 1 precursor (Salivary protein MSG1). | |||||
|
WASIP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 153) | NC score | 0.072207 (rank : 17) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | O43516, Q15220, Q6MZU9, Q9BU37, Q9UNP1 | Gene names | WASPIP, WIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein) (PRPL-2 protein). | |||||
|
WASIP_MOUSE
|
||||||
| θ value | θ > 10 (rank : 154) | NC score | 0.073913 (rank : 16) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q8K1I7, Q3U0U8 | Gene names | Waspip, Wip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein). | |||||
|
WBP11_HUMAN
|
||||||
| θ value | θ > 10 (rank : 155) | NC score | 0.057407 (rank : 27) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9Y2W2, Q96AY8 | Gene names | WBP11, NPWBP, SNP70 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 11 (WBP-11) (SH3 domain-binding protein SNP70) (Npw38-binding protein) (NpwBP). | |||||
|
WBP11_MOUSE
|
||||||
| θ value | θ > 10 (rank : 156) | NC score | 0.055690 (rank : 31) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q923D5, O88539, Q3UMD8, Q8VDI0 | Gene names | Wbp11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 11 (WBP-11). | |||||
|
WIRE_HUMAN
|
||||||
| θ value | θ > 10 (rank : 157) | NC score | 0.053567 (rank : 35) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q8TF74, Q658J8, Q71RE1, Q8TE44 | Gene names | WIRE, WICH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WIP-related protein (WASP-interacting protein-related protein) (WIP- and CR16-homologous protein). | |||||
|
YLPM1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 158) | NC score | 0.051627 (rank : 39) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
MEF2D_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 146 | |
| SwissProt Accessions | Q63943, Q63944 | Gene names | Mef2d | |||
|
Domain Architecture |

|
|||||
| Description | Myocyte-specific enhancer factor 2D. | |||||
|
MEF2D_HUMAN
|
||||||
| NC score | 0.981474 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 102 | |
| SwissProt Accessions | Q14814, Q14815 | Gene names | MEF2D | |||
|
Domain Architecture |

|
|||||
| Description | Myocyte-specific enhancer factor 2D. | |||||
|
MEF2A_MOUSE
|
||||||
| NC score | 0.957010 (rank : 3) | θ value | 2.1455e-110 (rank : 5) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q60929 | Gene names | Mef2a | |||
|
Domain Architecture |

|
|||||
| Description | Myocyte-specific enhancer factor 2A. | |||||
|
MEF2A_HUMAN
|
||||||
| NC score | 0.954101 (rank : 4) | θ value | 2.6227e-108 (rank : 6) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q02078, O43814, Q14223, Q14224, Q96D14 | Gene names | MEF2A, MEF2 | |||
|
Domain Architecture |

|
|||||
| Description | Myocyte-specific enhancer factor 2A (Serum response factor-like protein 1). | |||||
|
MEF2C_MOUSE
|
||||||
| NC score | 0.945284 (rank : 5) | θ value | 1.09934e-114 (rank : 3) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8CFN5, Q8R0H1, Q9D7L0, Q9QW20 | Gene names | Mef2c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myocyte-specific enhancer factor 2C. | |||||
|
MEF2C_HUMAN
|
||||||
| NC score | 0.942816 (rank : 6) | θ value | 1.8752e-114 (rank : 4) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q06413 | Gene names | MEF2C | |||
|
Domain Architecture |

|
|||||
| Description | Myocyte-specific enhancer factor 2C. | |||||
|
MEF2B_HUMAN
|
||||||
| NC score | 0.884881 (rank : 7) | θ value | 1.01529e-43 (rank : 7) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q02080 | Gene names | MEF2B, XMEF2 | |||
|
Domain Architecture |

|
|||||
| Description | Myocyte-specific enhancer factor 2B (Serum response factor-like protein 2) (XMEF2) (RSRFR2). | |||||
|
MEF2B_MOUSE
|
||||||
| NC score | 0.838483 (rank : 8) | θ value | 5.57106e-42 (rank : 8) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O55087, O55088, O55089, O55090, O55231, Q61843 | Gene names | Mef2b | |||
|
Domain Architecture |

|
|||||
| Description | Myocyte-specific enhancer factor 2B. | |||||
|
SRF_MOUSE
|
||||||
| NC score | 0.460674 (rank : 9) | θ value | 7.59969e-07 (rank : 10) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9JM73 | Gene names | Srf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serum response factor (SRF). | |||||
|
SRF_HUMAN
|
||||||
| NC score | 0.457295 (rank : 10) | θ value | 7.59969e-07 (rank : 9) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P11831 | Gene names | SRF | |||
|
Domain Architecture |

|
|||||
| Description | Serum response factor (SRF). | |||||
|
ATX7_HUMAN
|
||||||
| NC score | 0.089852 (rank : 11) | θ value | 0.0193708 (rank : 14) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O15265, O75328, O75329, Q9Y6P8 | Gene names | ATXN7, SCA7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-7 (Spinocerebellar ataxia type 7 protein). | |||||
|
MLL2_HUMAN
|
||||||
| NC score | 0.089701 (rank : 12) | θ value | 0.00035302 (rank : 11) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 87 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
EVI2B_MOUSE
|
||||||
| NC score | 0.088460 (rank : 13) | θ value | 0.0330416 (rank : 15) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8VD58, Q3U1A3 | Gene names | Evi2b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EVI2B protein precursor (Ecotropic viral integration site 2B protein). | |||||
|
BAT2_MOUSE
|
||||||
| NC score | 0.087283 (rank : 14) | θ value | 6.88961 (rank : 126) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q7TSC1, Q923A9, Q9Z1R1 | Gene names | Bat2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
BAT2_HUMAN
|
||||||
| NC score | 0.083269 (rank : 15) | θ value | 1.81305 (rank : 65) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
WASIP_MOUSE
|
||||||
| NC score | 0.073913 (rank : 16) | θ value | θ > 10 (rank : 154) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q8K1I7, Q3U0U8 | Gene names | Waspip, Wip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein). | |||||
|
WASIP_HUMAN
|
||||||
| NC score | 0.072207 (rank : 17) | θ value | θ > 10 (rank : 153) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | O43516, Q15220, Q6MZU9, Q9BU37, Q9UNP1 | Gene names | WASPIP, WIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein) (PRPL-2 protein). | |||||
|
NCOA6_HUMAN
|
||||||
| NC score | 0.070863 (rank : 18) | θ value | 6.88961 (rank : 130) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1202 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | Q14686, Q9NTZ9, Q9UH74, Q9UK86 | Gene names | NCOA6, AIB3, KIAA0181, RAP250, TRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (PRIP) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC RAP250) (Thyroid hormone receptor-binding protein). | |||||
|
NFRKB_MOUSE
|
||||||
| NC score | 0.069914 (rank : 19) | θ value | 0.21417 (rank : 22) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 393 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q6PIJ4, Q8BWV5, Q8K0X6 | Gene names | Nfrkb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear factor related to kappa-B-binding protein (DNA-binding protein R kappa-B). | |||||
|
ATN1_HUMAN
|
||||||
| NC score | 0.068624 (rank : 20) | θ value | θ > 10 (rank : 147) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
TARSH_HUMAN
|
||||||
| NC score | 0.068229 (rank : 21) | θ value | 0.0961366 (rank : 18) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 768 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q7Z7G0, Q6ZW20, Q6ZW22, Q9C082, Q9UFI6 | Gene names | ABI3BP, NESHBP, TARSH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Target of Nesh-SH3 precursor (Tarsh) (Nesh-binding protein) (NeshBP) (ABI gene family member 3-binding protein). | |||||
|
PRP1_HUMAN
|
||||||
| NC score | 0.066692 (rank : 22) | θ value | 1.38821 (rank : 61) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P04280, Q08805, Q15186, Q15187, Q15214, Q15215, Q16038 | Gene names | PRB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 1 precursor (Salivary proline-rich protein) [Contains: Basic peptide IB-6; Peptide P-H]. | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.060710 (rank : 23) | θ value | 0.125558 (rank : 19) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 86 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
PCLO_HUMAN
|
||||||
| NC score | 0.060273 (rank : 24) | θ value | 0.21417 (rank : 23) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 91 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
RAPH1_HUMAN
|
||||||
| NC score | 0.059858 (rank : 25) | θ value | 2.36792 (rank : 77) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q70E73, Q96Q37, Q9C0I2 | Gene names | RAPH1, ALS2CR9, KIAA1681, LPD, PREL2, RMO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras-associated and pleckstrin homology domains-containing protein 1 (RAPH1) (Lamellipodin) (Proline-rich EVH1 ligand 2) (PREL-2) (Protein RMO1) (Amyotrophic lateral sclerosis 2 chromosomal region candidate 9 gene protein). | |||||
|
PRP2_MOUSE
|
||||||
| NC score | 0.058431 (rank : 26) | θ value | θ > 10 (rank : 150) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P05142 | Gene names | Prh1, Prp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein MP-2 precursor. | |||||
|
WBP11_HUMAN
|
||||||
| NC score | 0.057407 (rank : 27) | θ value | θ > 10 (rank : 155) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9Y2W2, Q96AY8 | Gene names | WBP11, NPWBP, SNP70 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 11 (WBP-11) (SH3 domain-binding protein SNP70) (Npw38-binding protein) (NpwBP). | |||||
|
TAF4_HUMAN
|
||||||
| NC score | 0.056658 (rank : 28) | θ value | 4.03905 (rank : 99) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 687 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | O00268, Q5TBP6, Q99721, Q9BR40, Q9BX42 | Gene names | TAF4, TAF2C, TAF2C1, TAF4A, TAFII130, TAFII135 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 4 (TBP-associated factor 4) (Transcription initiation factor TFIID 135 kDa subunit) (TAF(II)135) (TAFII-135) (TAFII135) (TAFII-130) (TAFII130). | |||||
|
MLL3_HUMAN
|
||||||
| NC score | 0.055991 (rank : 29) | θ value | 0.0330416 (rank : 16) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
PCLO_MOUSE
|
||||||
| NC score | 0.055851 (rank : 30) | θ value | 0.62314 (rank : 35) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 90 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
WBP11_MOUSE
|
||||||
| NC score | 0.055690 (rank : 31) | θ value | θ > 10 (rank : 156) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q923D5, O88539, Q3UMD8, Q8VDI0 | Gene names | Wbp11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 11 (WBP-11). | |||||
|
K1802_HUMAN
|
||||||
| NC score | 0.055584 (rank : 32) | θ value | 0.000786445 (rank : 13) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1378 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q96JM3, Q6P181, Q8NC88, Q9BST0 | Gene names | KIAA1802, C13orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
SMR2D_MOUSE
|
||||||
| NC score | 0.054200 (rank : 33) | θ value | 0.813845 (rank : 46) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O35979 | Gene names | Smr2, Msg2, Vcs2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Submaxillary gland androgen-regulated protein 2, isoform delta precursor (Salivary protein MSG2, isoform delta). | |||||
|
NCOA6_MOUSE
|
||||||
| NC score | 0.053683 (rank : 34) | θ value | 5.27518 (rank : 116) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1076 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9JL19, Q9JLT9 | Gene names | Ncoa6, Aib3, Prip, Rap250, Trbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC) (Thyroid hormone receptor-binding protein). | |||||
|
WIRE_HUMAN
|
||||||
| NC score | 0.053567 (rank : 35) | θ value | θ > 10 (rank : 157) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q8TF74, Q658J8, Q71RE1, Q8TE44 | Gene names | WIRE, WICH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WIP-related protein (WASP-interacting protein-related protein) (WIP- and CR16-homologous protein). | |||||
|
SMR2C_MOUSE
|
||||||
| NC score | 0.053233 (rank : 36) | θ value | 0.813845 (rank : 45) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O35985 | Gene names | Smr2, Msg2, Vcs2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Submaxillary gland androgen-regulated protein 2, isoform gamma precursor (Salivary protein MSG2, isoform gamma). | |||||
|
HCN2_HUMAN
|
||||||
| NC score | 0.052337 (rank : 37) | θ value | 0.163984 (rank : 20) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 369 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9UL51, O60742, O60743, O75267, Q9UBS2 | Gene names | HCN2, BCNG2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 2 (Brain cyclic nucleotide-gated channel 2) (BCNG-2). | |||||
|
SMR1_MOUSE
|
||||||
| NC score | 0.051907 (rank : 38) | θ value | θ > 10 (rank : 152) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q61900 | Gene names | Smr1, Msg1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Submaxillary gland androgen-regulated protein 1 precursor (Salivary protein MSG1). | |||||
|
YLPM1_MOUSE
|
||||||
| NC score | 0.051627 (rank : 39) | θ value | θ > 10 (rank : 158) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
BSN_MOUSE
|
||||||
| NC score | 0.051432 (rank : 40) | θ value | 0.813845 (rank : 40) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 91 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
SSBP4_HUMAN
|
||||||
| NC score | 0.051315 (rank : 41) | θ value | 0.21417 (rank : 24) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 753 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9BWG4 | Gene names | SSBP4 | |||
|
Domain Architecture |

|
|||||
| Description | Single-stranded DNA-binding protein 4. | |||||
|
CNOT3_HUMAN
|
||||||
| NC score | 0.051106 (rank : 42) | θ value | θ > 10 (rank : 148) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O75175, Q9NZN7, Q9UF76 | Gene names | CNOT3, KIAA0691, NOT3 | |||
|
Domain Architecture |

|
|||||
| Description | CCR4-NOT transcription complex subunit 3 (CCR4-associated factor 3). | |||||
|
PRB3_HUMAN
|
||||||
| NC score | 0.051058 (rank : 43) | θ value | 0.62314 (rank : 36) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q04118, Q15188, Q7M4M9 | Gene names | PRB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 3 precursor (Parotid salivary glycoprotein G1) (Proline-rich protein G1). | |||||
|
PRR12_HUMAN
|
||||||
| NC score | 0.050932 (rank : 44) | θ value | θ > 10 (rank : 151) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q9ULL5, Q8N4J6 | Gene names | PRR12, KIAA1205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 12. | |||||
|
CNOT3_MOUSE
|
||||||
| NC score | 0.050730 (rank : 45) | θ value | θ > 10 (rank : 149) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8K0V4 | Gene names | Cnot3, Not3 | |||
|
Domain Architecture |

|
|||||
| Description | CCR4-NOT transcription complex subunit 3 (CCR4-associated factor 3). | |||||
|
BCL9_HUMAN
|
||||||
| NC score | 0.050582 (rank : 46) | θ value | 4.03905 (rank : 88) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1008 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | O00512 | Gene names | BCL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9) (Legless homolog). | |||||
|
MAGE1_HUMAN
|
||||||
| NC score | 0.050015 (rank : 47) | θ value | 8.99809 (rank : 142) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
BCL9_MOUSE
|
||||||
| NC score | 0.047490 (rank : 48) | θ value | 3.0926 (rank : 80) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 770 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9D219, Q67FX9, Q8BUJ8, Q8VE74 | Gene names | Bcl9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9). | |||||
|
HNF1A_MOUSE
|
||||||
| NC score | 0.047321 (rank : 49) | θ value | 1.81305 (rank : 68) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P22361 | Gene names | Tcf1, Hnf-1, Hnf-1a, Hnf1a | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 1-alpha (HNF-1A) (Liver-specific transcription factor LF-B1) (LFB1). | |||||
|
CN004_HUMAN
|
||||||
| NC score | 0.047259 (rank : 50) | θ value | 1.06291 (rank : 47) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9H1B7, Q8NDQ2, Q96JG2, Q9H3I7 | Gene names | C14orf4, KIAA1865 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf4. | |||||
|
HCN4_MOUSE
|
||||||
| NC score | 0.046328 (rank : 51) | θ value | 0.47712 (rank : 32) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 284 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O70507 | Gene names | Hcn4, Bcng3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 4 (Brain cyclic nucleotide-gated channel 3) (BCNG-3). | |||||
|
ECM1_HUMAN
|
||||||
| NC score | 0.044924 (rank : 52) | θ value | 1.38821 (rank : 57) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 182 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q16610 | Gene names | ECM1 | |||
|
Domain Architecture |

|
|||||
| Description | Extracellular matrix protein 1 precursor (Secretory component p85). | |||||
|
SPHK2_MOUSE
|
||||||
| NC score | 0.044910 (rank : 53) | θ value | 0.365318 (rank : 30) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9JIA7, Q91VA9, Q9DBH6 | Gene names | Sphk2 | |||
|
Domain Architecture |

|
|||||
| Description | Sphingosine kinase 2 (EC 2.7.1.-) (SK 2) (SPK 2). | |||||
|
PSL1_HUMAN
|
||||||
| NC score | 0.043651 (rank : 54) | θ value | 1.38821 (rank : 62) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8TCT7, O60365, Q567S3, Q8IUH9, Q9BUY6, Q9H3M4, Q9NPN2, Q9P1Z6 | Gene names | SPPL2B, KIAA1532, PSL1 | |||
|
Domain Architecture |

|
|||||
| Description | Signal peptide peptidase-like 2B (EC 3.4.23.-) (Protein SPP-like 2B) (Protein SPPL2b) (Intramembrane protease 4) (IMP4) (Presenilin-like protein 1). | |||||
|
CRTC1_MOUSE
|
||||||
| NC score | 0.042775 (rank : 55) | θ value | 2.36792 (rank : 72) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q68ED7, Q6ZQ85 | Gene names | Crtc1, Kiaa0616, Mect1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CREB-regulated transcription coactivator 1 (Mucoepidermoid carcinoma translocated protein 1 homolog). | |||||
|
AFF2_HUMAN
|
||||||
| NC score | 0.041744 (rank : 56) | θ value | 0.813845 (rank : 38) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P51816, O43786, O60215, P78407, Q13521, Q14323, Q9UNA5 | Gene names | AFF2, FMR2, OX19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation 2 protein) (Protein FMR-2) (FMR2P) (Protein Ox19) (Fragile X E mental retardation syndrome protein). | |||||
|
HCN2_MOUSE
|
||||||
| NC score | 0.041729 (rank : 57) | θ value | 1.06291 (rank : 52) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O88703, O70506 | Gene names | Hcn2, Bcng2, Hac1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 2 (Brain cyclic nucleotide-gated channel 2) (BCNG-2) (Hyperpolarization-activated cation channel 1) (HAC-1). | |||||
|
TM108_MOUSE
|
||||||
| NC score | 0.041360 (rank : 58) | θ value | 4.03905 (rank : 102) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 336 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8BHE4, Q80WR9 | Gene names | Tmem108 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 108 precursor. | |||||
|
EVI2B_HUMAN
|
||||||
| NC score | 0.040791 (rank : 59) | θ value | 4.03905 (rank : 92) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P34910 | Gene names | EVI2B, EVDB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EVI2B protein precursor (Ecotropic viral integration site 2B protein). | |||||
|
NCOA3_MOUSE
|
||||||
| NC score | 0.040449 (rank : 60) | θ value | 0.47712 (rank : 33) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O09000, Q9CRD5 | Gene names | Ncoa3, Aib1, Pcip, Rac3, Tram1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein homolog) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP- interacting protein) (p/CIP) (pCIP). | |||||
|
CD248_HUMAN
|
||||||
| NC score | 0.039647 (rank : 61) | θ value | 0.365318 (rank : 26) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 679 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9HCU0, Q3SX55, Q96KB6 | Gene names | CD248, CD164L1, TEM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endosialin precursor (Tumor endothelial marker 1) (CD248 antigen). | |||||
|
IF4G1_MOUSE
|
||||||
| NC score | 0.038392 (rank : 62) | θ value | 1.06291 (rank : 53) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 534 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q6NZJ6, Q6NZN8, Q8BW99 | Gene names | Eif4g1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1). | |||||
|
CIZ1_HUMAN
|
||||||
| NC score | 0.037784 (rank : 63) | θ value | 5.27518 (rank : 109) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9ULV3, Q9NYM8, Q9UHK4, Q9Y3F9, Q9Y3G0 | Gene names | CIZ1, LSFR1, NP94 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cip1-interacting zinc finger protein (Nuclear protein NP94). | |||||
|
GTSE1_MOUSE
|
||||||
| NC score | 0.036776 (rank : 64) | θ value | 6.88961 (rank : 127) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8R080, O89015, Q9CSG9 | Gene names | Gtse1, B99 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G2 and S phase expressed protein 1 (Gtse-1) (B99 protein). | |||||
|
IPMK_HUMAN
|
||||||
| NC score | 0.035486 (rank : 65) | θ value | 0.813845 (rank : 44) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NFU5 | Gene names | IPMK, IMPK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol polyphosphate multikinase (EC 2.7.1.151) (Inositol 1,3,4,6- tetrakisphosphate 5-kinase). | |||||
|
UTX_HUMAN
|
||||||
| NC score | 0.035450 (rank : 66) | θ value | 1.06291 (rank : 54) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O15550 | Gene names | UTX | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitously transcribed X chromosome tetratricopeptide repeat protein (Ubiquitously transcribed TPR protein on the X chromosome). | |||||
|
HCN4_HUMAN
|
||||||
| NC score | 0.035343 (rank : 67) | θ value | 4.03905 (rank : 94) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9Y3Q4, Q9UMQ7 | Gene names | HCN4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 4. | |||||
|
SOX6_MOUSE
|
||||||
| NC score | 0.035340 (rank : 68) | θ value | 0.279714 (rank : 25) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P40645, Q62250, Q9QWS5 | Gene names | Sox6, Sox-6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-6 (SOX-LZ). | |||||
|
IF4G2_HUMAN
|
||||||
| NC score | 0.034855 (rank : 69) | θ value | 3.0926 (rank : 83) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P78344, O60877, P78404, Q53EU1, Q58EZ2, Q8NI71, Q96C16 | Gene names | EIF4G2, DAP5 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 2 (eIF-4-gamma 2) (eIF-4G 2) (eIF4G 2) (p97) (Death-associated protein 5) (DAP-5). | |||||
|
ARRD1_MOUSE
|
||||||
| NC score | 0.034718 (rank : 70) | θ value | 1.38821 (rank : 56) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99KN1, Q3TDT6, Q8VEG7 | Gene names | Arrdc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arrestin domain-containing protein 1. | |||||
|
MINT_HUMAN
|
||||||
| NC score | 0.034476 (rank : 71) | θ value | 1.38821 (rank : 59) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1291 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q96T58, Q9H9A8, Q9NWH5, Q9UQ01, Q9Y556 | Gene names | SPEN, KIAA0929, MINT, SHARP | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
FA47B_HUMAN
|
||||||
| NC score | 0.034441 (rank : 72) | θ value | 3.0926 (rank : 82) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 431 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8NA70, Q5JQN5, Q6PIG3 | Gene names | FAM47B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM47B. | |||||
|
PI5PA_HUMAN
|
||||||
| NC score | 0.034151 (rank : 73) | θ value | 4.03905 (rank : 98) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 749 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q15735, Q32M61, Q6ZTH6, Q8N902, Q9UDT9 | Gene names | PIB5PA, PIPP | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 4,5-bisphosphate 5-phosphatase A (EC 3.1.3.56). | |||||
|
IF4G2_MOUSE
|
||||||
| NC score | 0.034019 (rank : 74) | θ value | 3.0926 (rank : 84) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q62448, P97867, Q921U3 | Gene names | Eif4g2, Nat1 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 2 (eIF-4-gamma 2) (eIF-4G 2) (eIF4G 2) (p97) (Novel APOBEC-1 target 1) (Translation repressor NAT1). | |||||
|
ADA12_MOUSE
|
||||||
| NC score | 0.032071 (rank : 75) | θ value | 0.000461057 (rank : 12) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 474 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q61824 | Gene names | Adam12, Mltna | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 12 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 12) (Meltrin alpha). | |||||
|
GP179_HUMAN
|
||||||
| NC score | 0.031700 (rank : 76) | θ value | 1.38821 (rank : 58) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 372 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6PRD1 | Gene names | GPR179, GPR158L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 179 precursor (Probable G-protein coupled receptor 158-like 1). | |||||
|
CD2_HUMAN
|
||||||
| NC score | 0.031477 (rank : 77) | θ value | 3.0926 (rank : 81) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P06729, Q96TE5 | Gene names | CD2 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell surface antigen CD2 precursor (T-cell surface antigen T11/Leu- 5) (LFA-2) (LFA-3 receptor) (Erythrocyte receptor) (Rosette receptor). | |||||
|
GAK5_HUMAN
|
||||||
| NC score | 0.031204 (rank : 78) | θ value | 5.27518 (rank : 113) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 429 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P62684 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19p13.11 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K113 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
IRF7_MOUSE
|
||||||
| NC score | 0.030845 (rank : 79) | θ value | 0.0736092 (rank : 17) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P70434 | Gene names | Irf7 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon regulatory factor 7 (IRF-7). | |||||
|
CEL_HUMAN
|
||||||
| NC score | 0.029375 (rank : 80) | θ value | 4.03905 (rank : 89) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P19835, Q16398 | Gene names | CEL, BAL | |||
|
Domain Architecture |

|
|||||
| Description | Bile salt-activated lipase precursor (EC 3.1.1.3) (EC 3.1.1.13) (BAL) (Bile salt-stimulated lipase) (BSSL) (Carboxyl ester lipase) (Sterol esterase) (Cholesterol esterase) (Pancreatic lysophospholipase). | |||||
|
GPR98_HUMAN
|
||||||
| NC score | 0.029324 (rank : 81) | θ value | 0.365318 (rank : 28) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8WXG9, O75171, Q9H0X5, Q9UL61 | Gene names | GPR98, KIAA0686, MASS1, VLGR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G-protein coupled receptor 98 precursor (Monogenic audiogenic seizure susceptibility protein 1 homolog) (Very large G-protein coupled receptor 1) (Usher syndrome type-2C protein). | |||||
|
LR37A_HUMAN
|
||||||
| NC score | 0.028810 (rank : 82) | θ value | 8.99809 (rank : 140) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 720 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O60309, Q49A01, Q49A80, Q8NB33 | Gene names | LRRC37A, KIAA0563 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 37A. | |||||
|
TAU_HUMAN
|
||||||
| NC score | 0.028615 (rank : 83) | θ value | 4.03905 (rank : 100) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 448 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P10636, P18518, Q14799, Q15549, Q15550, Q15551, Q9UDJ3, Q9UMH0, Q9UQ96 | Gene names | MAPT, MTBT1, TAU | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein tau (Neurofibrillary tangle protein) (Paired helical filament-tau) (PHF-tau). | |||||
|
GAK4_HUMAN
|
||||||
| NC score | 0.028588 (rank : 84) | θ value | 5.27518 (rank : 112) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 415 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P63126, Q9UKH4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_6q14.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K109 Gag protein) (HERV-K(C6) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK11_HUMAN
|
||||||
| NC score | 0.028537 (rank : 85) | θ value | 5.27518 (rank : 111) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P63145, Q9UKI1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_22q11.21 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K101 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
ATX1_MOUSE
|
||||||
| NC score | 0.028398 (rank : 86) | θ value | 5.27518 (rank : 107) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P54254 | Gene names | Atxn1, Sca1 | |||
|
Domain Architecture |

|
|||||
| Description | Ataxin-1 (Spinocerebellar ataxia type 1 protein homolog). | |||||
|
TAB3_MOUSE
|
||||||
| NC score | 0.028369 (rank : 87) | θ value | 5.27518 (rank : 122) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q571K4 | Gene names | Map3k7ip3, Kiaa4135, Tab3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase 7-interacting protein 3 (TAK1-binding protein 3). | |||||
|
MAGC1_HUMAN
|
||||||
| NC score | 0.028138 (rank : 88) | θ value | 5.27518 (rank : 115) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 431 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O60732, O75451, Q8TCV4 | Gene names | MAGEC1 | |||
|
Domain Architecture |

|
|||||
| Description | Melanoma-associated antigen C1 (MAGE-C1 antigen) (Cancer-testis antigen CT7). | |||||
|
GAK12_HUMAN
|
||||||
| NC score | 0.027818 (rank : 89) | θ value | 8.99809 (rank : 136) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P63130, Q9UKI0 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_1q22 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K102 Gag protein) (HERV-K(III) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK7_HUMAN
|
||||||
| NC score | 0.027462 (rank : 90) | θ value | 2.36792 (rank : 73) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q96PI4, Q9QC08 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_1q23.3 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K18 Gag protein) (HERV-K110 Gag protein) (HERV-K(C1a) Gag protein) [Contains: Matrix protein]. | |||||
|
FOXP4_HUMAN
|
||||||
| NC score | 0.027411 (rank : 91) | θ value | 0.47712 (rank : 31) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8IVH2, Q96E19 | Gene names | FOXP4, FKHLA | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P4 (Fork head-related protein-like A). | |||||
|
EPN4_MOUSE
|
||||||
| NC score | 0.027150 (rank : 92) | θ value | 4.03905 (rank : 90) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99KN9, Q8CFH4 | Gene names | Clint1, Enth, Epn4, Epnr, Kiaa0171 | |||
|
Domain Architecture |

|
|||||
| Description | Clathrin interactor 1 (Epsin-4) (Epsin-related protein) (EpsinR) (Enthoprotin). | |||||
|
ZHX2_MOUSE
|
||||||
| NC score | 0.027115 (rank : 93) | θ value | 0.163984 (rank : 21) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8C0C0 | Gene names | Zhx2, Afr1, Raf | |||
|
Domain Architecture |

|
|||||
| Description | Zinc fingers and homeoboxes protein 2 (Zinc finger and homeodomain protein 2) (Alpha-fetoprotein regulator 1) (AFP regulator 1) (Regulator of AFP). | |||||
|
CN037_MOUSE
|
||||||
| NC score | 0.027108 (rank : 94) | θ value | 5.27518 (rank : 110) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 179 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8BT18, Q8BFQ7, Q8R1Q5, Q9D6C5 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C14orf37 homolog precursor. | |||||
|
SOX6_HUMAN
|
||||||
| NC score | 0.027028 (rank : 95) | θ value | 3.0926 (rank : 86) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P35712, Q9BXQ3, Q9BXQ4, Q9BXQ5, Q9H0I8 | Gene names | SOX6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-6. | |||||
|
GNAS1_MOUSE
|
||||||
| NC score | 0.026236 (rank : 96) | θ value | 0.813845 (rank : 43) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 651 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q6R0H7, Q6R0H4, Q6R0H5, Q6R2J5, Q9JJX0, Q9Z1N8 | Gene names | Gnas, Gnas1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Guanine nucleotide-binding protein G(s) subunit alpha isoforms XLas (Adenylate cyclase-stimulating G alpha protein) (Extra large alphas protein) (XLalphas). | |||||
|
GAK6_HUMAN
|
||||||
| NC score | 0.026199 (rank : 97) | θ value | 8.99809 (rank : 138) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P62685 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_8p23.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K115 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK2_HUMAN
|
||||||
| NC score | 0.026187 (rank : 98) | θ value | 8.99809 (rank : 137) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q7LDI9, Q9UKH5, Q9Y6I1, Q9YNA6, Q9YNB0 | Gene names | ERVK6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_7p22.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K(HML-2.HOM) Gag protein) (HERV-K108 Gag protein) (HERV-K(C7) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
LPP_HUMAN
|
||||||
| NC score | 0.025922 (rank : 99) | θ value | 1.81305 (rank : 70) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 775 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q93052, Q8NFX5 | Gene names | LPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipoma-preferred partner (LIM domain-containing preferred translocation partner in lipoma). | |||||
|
JUND_MOUSE
|
||||||
| NC score | 0.025878 (rank : 100) | θ value | 1.81305 (rank : 69) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P15066 | Gene names | Jund, Jun-d, Jund1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor jun-D. | |||||
|
K0552_HUMAN
|
||||||
| NC score | 0.025777 (rank : 101) | θ value | 2.36792 (rank : 74) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 442 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O60299 | Gene names | KIAA0552 | |||
|
Domain Architecture |

|
|||||
| Description | Protein KIAA0552. | |||||
|
BORG1_HUMAN
|
||||||
| NC score | 0.025671 (rank : 102) | θ value | 0.813845 (rank : 39) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O14613, Q9UNS0 | Gene names | CDC42EP2, BORG1, CEP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cdc42 effector protein 2 (Binder of Rho GTPases 1). | |||||
|
OGFR_MOUSE
|
||||||
| NC score | 0.025176 (rank : 103) | θ value | 5.27518 (rank : 117) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 584 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q99PG2, Q99L33 | Gene names | Ogfr | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor). | |||||
|
SOX13_MOUSE
|
||||||
| NC score | 0.025025 (rank : 104) | θ value | 1.81305 (rank : 71) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 495 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q04891, Q922L3 | Gene names | Sox13, Sox-13 | |||
|
Domain Architecture |

|
|||||
| Description | SOX-13 protein. | |||||
|
FOXC1_MOUSE
|
||||||
| NC score | 0.024997 (rank : 105) | θ value | 1.06291 (rank : 50) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q61572, O88409, Q61582 | Gene names | Foxc1, Fkh1, Fkhl7, Freac3, Mf1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein C1 (Forkhead-related protein FKHL7) (Forkhead- related transcription factor 3) (FREAC-3) (Transcription factor FKH-1) (Mesoderm/mesenchyme forkhead 1) (MF-1). | |||||
|
MAP1B_MOUSE
|
||||||
| NC score | 0.024019 (rank : 106) | θ value | 3.0926 (rank : 85) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 977 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P14873 | Gene names | Map1b, Mtap1b, Mtap5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) (MAP1.2) (MAP1(X)) [Contains: MAP1 light chain LC1]. | |||||
|
FOXC1_HUMAN
|
||||||
| NC score | 0.023463 (rank : 107) | θ value | 1.06291 (rank : 49) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q12948, Q86UP7, Q9BYM1, Q9NUE5, Q9UDD0, Q9UP06 | Gene names | FOXC1, FKHL7, FREAC3 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein C1 (Forkhead-related protein FKHL7) (Forkhead- related transcription factor 3) (FREAC-3). | |||||
|
UBQL3_HUMAN
|
||||||
| NC score | 0.022992 (rank : 108) | θ value | 0.62314 (rank : 37) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9H347, Q9NRE0 | Gene names | UBQLN3 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquilin-3. | |||||
|
LBN_MOUSE
|
||||||
| NC score | 0.022662 (rank : 109) | θ value | 2.36792 (rank : 75) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8K1G2, Q8BRF3 | Gene names | Evc2, Lbn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Limbin. | |||||
|
PENK_HUMAN
|
||||||
| NC score | 0.022229 (rank : 110) | θ value | 5.27518 (rank : 118) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P01210 | Gene names | PENK | |||
|
Domain Architecture |

|
|||||
| Description | Proenkephalin A precursor [Contains: Synenkephalin; Met-enkephalin (Opioid growth factor) (OGF); Met-enkephalin-Arg-Gly-Leu; Leu- enkephalin; Met-enkephalin-Arg-Phe]. | |||||
|
MB3L2_HUMAN
|
||||||
| NC score | 0.021515 (rank : 111) | θ value | 6.88961 (rank : 129) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8NHZ7 | Gene names | MBD3L2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 3-like 2 (MBD3-like 2). | |||||
|
ZN414_HUMAN
|
||||||
| NC score | 0.021179 (rank : 112) | θ value | 4.03905 (rank : 105) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 548 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q96IQ9 | Gene names | ZNF414 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 414. | |||||
|
CUTL1_MOUSE
|
||||||
| NC score | 0.021068 (rank : 113) | θ value | 1.06291 (rank : 48) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P53564, O08994, P70301, Q571L6, Q91ZD2 | Gene names | Cutl1, Cux, Cux1, Kiaa4047 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP) (Homeobox protein Cux). | |||||
|
LAMP3_MOUSE
|
||||||
| NC score | 0.020318 (rank : 114) | θ value | 5.27518 (rank : 114) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7TST5, Q8C1F6 | Gene names | Lamp3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lysosome-associated membrane glycoprotein 3 precursor (LAMP-3) (Lysosomal-associated membrane protein 3) (DC-lysosome-associated membrane glycoprotein) (DC LAMP) (CD208 antigen). | |||||
|
GP152_HUMAN
|
||||||
| NC score | 0.020021 (rank : 115) | θ value | 0.365318 (rank : 27) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 636 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8TDT2, Q86SM0 | Gene names | GPR152, PGR5 | |||
|
Domain Architecture |

|
|||||
| Description | Probable G-protein coupled receptor 152 (G-protein coupled receptor PGR5). | |||||
|
RIF1_MOUSE
|
||||||
| NC score | 0.019999 (rank : 116) | θ value | 1.38821 (rank : 63) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6PR54, Q6T8D9, Q7TPD8, Q8BTU2, Q8C408, Q9CS77 | Gene names | Rif1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Telomere-associated protein RIF1 (Rap1-interacting factor 1 homolog) (mRif1). | |||||
|
FOXN4_HUMAN
|
||||||
| NC score | 0.018372 (rank : 117) | θ value | 1.06291 (rank : 51) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q96NZ1, Q6ZMR4, Q96NZ0 | Gene names | FOXN4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein N4. | |||||
|
ES8L3_HUMAN
|
||||||
| NC score | 0.018368 (rank : 118) | θ value | 0.813845 (rank : 41) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8TE67, Q5T8Q6, Q5T8Q7, Q5T8Q8, Q96E47, Q9H719 | Gene names | EPS8L3, EPS8R3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Epidermal growth factor receptor kinase substrate 8-like protein 3 (Epidermal growth factor receptor pathway substrate 8-related protein 3) (EPS8-like protein 3). | |||||
|
PCBP4_HUMAN
|
||||||
| NC score | 0.017659 (rank : 119) | θ value | 0.62314 (rank : 34) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P57723, Q96AH7 | Gene names | PCBP4 | |||
|
Domain Architecture |

|
|||||
| Description | Poly(rC)-binding protein 4 (Alpha-CP4). | |||||
|
DCP1A_MOUSE
|
||||||
| NC score | 0.017573 (rank : 120) | θ value | 8.99809 (rank : 135) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q91YD3, Q6NZE3 | Gene names | Dcp1a, Mitc1, Smif | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | mRNA decapping enzyme 1A (EC 3.-.-.-) (Transcription factor SMIF) (MAD homolog 4-interacting transcription coactivator 1) (Smad4-interacting transcriptional co-activator). | |||||
|
ETV5_MOUSE
|
||||||
| NC score | 0.017449 (rank : 121) | θ value | 4.03905 (rank : 91) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9CXC9, Q3TG49, Q8C0F3, Q9JHB1 | Gene names | Etv5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 5. | |||||
|
TCF7_MOUSE
|
||||||
| NC score | 0.017113 (rank : 122) | θ value | 8.99809 (rank : 144) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q00417 | Gene names | Tcf7, Tcf-1, Tcf1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 7 (T-cell-specific transcription factor 1) (TCF- 1) (T-cell factor 1). | |||||
|
US6NL_MOUSE
|
||||||
| NC score | 0.016733 (rank : 123) | θ value | 3.0926 (rank : 87) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q80XC3, Q6ZQK8 | Gene names | Usp6nl, Kiaa0019 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USP6 N-terminal-like protein. | |||||
|
ATBF1_MOUSE
|
||||||
| NC score | 0.016517 (rank : 124) | θ value | 3.0926 (rank : 79) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1841 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q61329 | Gene names | Atbf1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-fetoprotein enhancer-binding protein (AT motif-binding factor) (AT-binding transcription factor 1). | |||||
|
CUTL1_HUMAN
|
||||||
| NC score | 0.016497 (rank : 125) | θ value | 1.81305 (rank : 66) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1574 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P39880, Q6NYH4, Q9UEV5 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP). | |||||
|
PRRT3_MOUSE
|
||||||
| NC score | 0.016163 (rank : 126) | θ value | 8.99809 (rank : 143) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q6PE13 | Gene names | Prrt3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
ZC3H3_MOUSE
|
||||||
| NC score | 0.016048 (rank : 127) | θ value | 8.99809 (rank : 146) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8CHP0, Q80X78 | Gene names | Zc3h3, Kiaa0150, Zc3hdc3, Zfp623 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 3. | |||||
|
MEGF9_MOUSE
|
||||||
| NC score | 0.015500 (rank : 128) | θ value | 4.03905 (rank : 96) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8BH27, Q8BWI4 | Gene names | Megf9, Egfl5, Kiaa0818 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 9 precursor (EGF-like domain-containing protein 5) (Multiple EGF-like domain protein 5). | |||||
|
TENS4_MOUSE
|
||||||
| NC score | 0.015412 (rank : 129) | θ value | 4.03905 (rank : 101) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8BZ33, Q3TCM8, Q7TNR5 | Gene names | Tns4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tensin-4 precursor. | |||||
|
4ET_HUMAN
|
||||||
| NC score | 0.015218 (rank : 130) | θ value | 5.27518 (rank : 106) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NRA8, Q8NCF2, Q9H708 | Gene names | EIF4ENIF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4E transporter (eIF4E transporter) (4E-T) (Eukaryotic translation initiation factor 4E nuclear import factor 1). | |||||
|
TTC28_HUMAN
|
||||||
| NC score | 0.014496 (rank : 131) | θ value | 5.27518 (rank : 123) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q96AY4, O95928, O95929, Q9NTE4, Q9UG31, Q9UGG5, Q9UPV8, Q9Y3S5 | Gene names | TTC28, KIAA1043 | |||
|
Domain Architecture |

|
|||||
| Description | Tetratricopeptide repeat protein 28 (TPR repeat protein 28). | |||||
|
MYO15_HUMAN
|
||||||
| NC score | 0.013768 (rank : 132) | θ value | 4.03905 (rank : 97) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9UKN7 | Gene names | MYO15A, MYO15 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-15 (Myosin XV) (Unconventional myosin-15). | |||||
|
BOLL_HUMAN
|
||||||
| NC score | 0.012804 (rank : 133) | θ value | 5.27518 (rank : 108) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8N9W6, Q969U3 | Gene names | BOLL, BOULE | |||
|
Domain Architecture |

|
|||||
| Description | Protein boule-like. | |||||
|
IQEC3_MOUSE
|
||||||
| NC score | 0.012778 (rank : 134) | θ value | 6.88961 (rank : 128) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q3TES0, Q3UHP8, Q80TJ8 | Gene names | Iqsec3, Kiaa1110 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 3. | |||||
|
SEM6C_MOUSE
|
||||||
| NC score | 0.012754 (rank : 135) | θ value | 1.38821 (rank : 64) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9WTM3 | Gene names | Sema6c, Semay | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-6C precursor (Semaphorin Y) (Sema Y). | |||||
|
PO2F2_HUMAN
|
||||||
| NC score | 0.012487 (rank : 136) | θ value | 2.36792 (rank : 76) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 379 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P09086, Q16648, Q9BRS4 | Gene names | POU2F2, OCT2, OTF2 | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 2, transcription factor 2 (Octamer-binding transcription factor 2) (Oct-2) (OTF-2) (Lymphoid-restricted immunoglobulin octamer-binding protein NF-A2). | |||||
|
SPEG_MOUSE
|
||||||
| NC score | 0.011798 (rank : 137) | θ value | 0.365318 (rank : 29) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
FARP2_HUMAN
|
||||||
| NC score | 0.011741 (rank : 138) | θ value | 1.81305 (rank : 67) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O94887, Q53QM5, Q8WU27, Q9UFE7 | Gene names | FARP2, KIAA0793 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FERM, RhoGEF and pleckstrin domain-containing protein 2 (FERM domain including RhoGEF) (FIR). | |||||
|
RAD9A_MOUSE
|
||||||
| NC score | 0.011735 (rank : 139) | θ value | 6.88961 (rank : 131) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Z0F6, Q8QZZ3 | Gene names | Rad9a, Rad9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell cycle checkpoint control protein RAD9A (EC 3.1.11.2) (DNA repair exonuclease rad9 homolog A) (Rad9-like protein) (mRAD9). | |||||
|
AP1G1_MOUSE
|
||||||
| NC score | 0.011426 (rank : 140) | θ value | 6.88961 (rank : 125) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P22892 | Gene names | Ap1g1, Adtg, Clapg1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-1 complex subunit gamma-1 (Adapter-related protein complex 1 gamma- 1 subunit) (Gamma-adaptin) (Adaptor protein complex AP-1 gamma-1 subunit) (Golgi adaptor HA1/AP1 adaptin subunit gamma-1) (Clathrin assembly protein complex 1 gamma-1 large chain). | |||||
|
RX_HUMAN
|
||||||
| NC score | 0.011320 (rank : 141) | θ value | 2.36792 (rank : 78) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 423 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9Y2V3 | Gene names | RAX, RX | |||
|
Domain Architecture |

|
|||||
| Description | Retinal homeobox protein Rx (Retina and anterior neural fold homeobox protein). | |||||
|
AP1G1_HUMAN
|
||||||
| NC score | 0.010513 (rank : 142) | θ value | 8.99809 (rank : 133) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O43747, O75709, O75842, Q9UG09, Q9Y3U4 | Gene names | AP1G1, ADTG, CLAPG1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-1 complex subunit gamma-1 (Adapter-related protein complex 1 gamma- 1 subunit) (Gamma-adaptin) (Adaptor protein complex AP-1 gamma-1 subunit) (Golgi adaptor HA1/AP1 adaptin subunit gamma-1) (Clathrin assembly protein complex 1 gamma-1 large chain). | |||||
|
FBN2_HUMAN
|
||||||
| NC score | 0.009552 (rank : 143) | θ value | 0.813845 (rank : 42) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P35556 | Gene names | FBN2 | |||
|
Domain Architecture |

|
|||||
| Description | Fibrillin-2 precursor. | |||||
|
WNK1_HUMAN
|
||||||
| NC score | 0.009414 (rank : 144) | θ value | 4.03905 (rank : 104) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1158 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9H4A3, O15052, Q86WL5, Q8N673, Q9P1S9 | Gene names | WNK1, KDP, KIAA0344, PRKWNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK1 (EC 2.7.11.1) (Protein kinase with no lysine 1) (Protein kinase, lysine-deficient 1) (Kinase deficient protein). | |||||
|
HME1_MOUSE
|
||||||
| NC score | 0.009399 (rank : 145) | θ value | 8.99809 (rank : 139) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P09065 | Gene names | En1, En-1 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein engrailed-1 (Mo-En-1). | |||||
|
WEE1B_HUMAN
|
||||||
| NC score | 0.008594 (rank : 146) | θ value | 4.03905 (rank : 103) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 836 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P0C1S8 | Gene names | WEE1B, WEE2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wee1-like protein kinase 1B (EC 2.7.10.2) (Wee1B kinase). | |||||
|
ABCA7_HUMAN
|
||||||
| NC score | 0.008361 (rank : 147) | θ value | 1.38821 (rank : 55) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IZY2, Q96S58, Q9BZC4, Q9NR73, Q9UKP8 | Gene names | ABCA7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-binding cassette sub-family A member 7 (Macrophage ABC transporter) (Autoantigen SS-N) (ABCA-SSN). | |||||
|
VINEX_MOUSE
|
||||||
| NC score | 0.006718 (rank : 148) | θ value | 8.99809 (rank : 145) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9R1Z8, Q62423 | Gene names | Sorbs3, Scam1, Sh3d4 | |||
|
Domain Architecture |

|
|||||
| Description | Vinexin (Sorbin and SH3 domain-containing protein 3) (SH3-containing adapter molecule 1) (SCAM-1) (SH3 domain-containing protein SH3P3). | |||||
|
KBTB9_MOUSE
|
||||||
| NC score | 0.006659 (rank : 149) | θ value | 4.03905 (rank : 95) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q80T74 | Gene names | Kbtbd9, Kiaa1921 | |||
|
Domain Architecture |

|
|||||
| Description | Kelch repeat and BTB domain-containing protein 9. | |||||
|
SC6A5_MOUSE
|
||||||
| NC score | 0.006339 (rank : 150) | θ value | 5.27518 (rank : 121) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q761V0, Q8CFM5, Q91ZQ2 | Gene names | Slc6a5, Glyt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium- and chloride-dependent glycine transporter 2 (GlyT2) (GlyT-2) (Solute carrier family 6 member 5). | |||||
|
CAC1B_HUMAN
|
||||||
| NC score | 0.006309 (rank : 151) | θ value | 8.99809 (rank : 134) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q00975 | Gene names | CACNA1B, CACH5, CACNL1A5 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent N-type calcium channel subunit alpha-1B (Voltage- gated calcium channel subunit alpha Cav2.2) (Calcium channel, L type, alpha-1 polypeptide isoform 5) (Brain calcium channel III) (BIII). | |||||
|
ROBO3_MOUSE
|
||||||
| NC score | 0.005736 (rank : 152) | θ value | 5.27518 (rank : 120) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 524 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Z2I4 | Gene names | Robo3, Rbig1 | |||
|
Domain Architecture |

|
|||||
| Description | Roundabout homolog 3 precursor (Retinoblastoma-inhibiting gene 1) (Rig-1). | |||||
|
WNK1_MOUSE
|
||||||
| NC score | 0.005256 (rank : 153) | θ value | 5.27518 (rank : 124) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1179 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P83741 | Gene names | Wnk1, Prkwnk1 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK1 (EC 2.7.11.1) (Protein kinase with no lysine 1) (Protein kinase, lysine-deficient 1). | |||||
|
MACF4_HUMAN
|
||||||
| NC score | 0.002319 (rank : 154) | θ value | 8.99809 (rank : 141) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1321 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q96PK2, Q8WXY1 | Gene names | MACF1, ABP620, ACF7, KIAA0465, KIAA1251 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1, isoform 4. | |||||
|
GLI2_HUMAN
|
||||||
| NC score | 0.002164 (rank : 155) | θ value | 4.03905 (rank : 93) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 971 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P10070, O60252, O60253, O60254, O60255, Q15590, Q15591 | Gene names | GLI2, THP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein GLI2 (Tax helper protein). | |||||
|
TCF8_MOUSE
|
||||||
| NC score | 0.001139 (rank : 156) | θ value | 6.88961 (rank : 132) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1067 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q64318, Q62519 | Gene names | Tcf8, Zfhx1a, Zfx1a, Zfx1ha | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 8 (Zinc finger homeobox protein 1a) (MEB1) (Delta EF1). | |||||
|
PKN3_MOUSE
|
||||||
| NC score | 0.000959 (rank : 157) | θ value | 1.38821 (rank : 60) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1009 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8K045 | Gene names | Pkn3, Pknbeta | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase N3 (EC 2.7.11.13) (Protein kinase PKN- beta) (Protein-kinase C-related kinase 3). | |||||
|
PRD10_HUMAN
|
||||||
| NC score | -0.001095 (rank : 158) | θ value | 5.27518 (rank : 119) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 769 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NQV6, Q9ULI9 | Gene names | PRDM10, KIAA1231, PFM7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PR domain zinc finger protein 10 (PR domain-containing protein 10). | |||||