Please be patient as the page loads
|
TIP60_HUMAN
|
||||||
| SwissProt Accessions | Q92993, O95624, Q13430, Q6GSE8, Q9BWK7 | Gene names | HTATIP, TIP60 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase HTATIP (EC 2.3.1.48) (EC 2.3.1.-) (60 kDa Tat interactive protein) (Tip60) (HIV-1 Tat interactive protein) (cPLA(2)-interacting protein). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
TIP60_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 90 | |
| SwissProt Accessions | Q92993, O95624, Q13430, Q6GSE8, Q9BWK7 | Gene names | HTATIP, TIP60 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase HTATIP (EC 2.3.1.48) (EC 2.3.1.-) (60 kDa Tat interactive protein) (Tip60) (HIV-1 Tat interactive protein) (cPLA(2)-interacting protein). | |||||
|
TIP60_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999444 (rank : 2) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 88 | |
| SwissProt Accessions | Q8CHK4, Q8CGZ3, Q8CGZ4, Q8VIH0 | Gene names | Htatip, Tip60 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase HTATIP (EC 2.3.1.48) (EC 2.3.1.-) (60 kDa Tat interactive protein) (Tip60). | |||||
|
MYST1_HUMAN
|
||||||
| θ value | 5.14231e-88 (rank : 3) | NC score | 0.952888 (rank : 3) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9H7Z6, Q659G0, Q7LC17, Q8IY59, Q8WYB4, Q8WZ14, Q9HAC5, Q9NR35 | Gene names | MYST1, MOF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable histone acetyltransferase MYST1 (EC 2.3.1.48) (MYST protein 1) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 1) (hMOF). | |||||
|
MYST1_MOUSE
|
||||||
| θ value | 5.14231e-88 (rank : 4) | NC score | 0.952429 (rank : 4) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9D1P2, Q3UIY0, Q8BJ69, Q8BJ76, Q8CI73 | Gene names | Myst1, Mof | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable histone acetyltransferase MYST1 (EC 2.3.1.48) (MYST protein 1) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 1). | |||||
|
MYST2_HUMAN
|
||||||
| θ value | 1.95407e-87 (rank : 5) | NC score | 0.927248 (rank : 5) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O95251 | Gene names | MYST2, HBO1, HBOa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST2 (EC 2.3.1.48) (MYST protein 2) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 2) (Histone acetyltransferase binding to ORC1). | |||||
|
MYST2_MOUSE
|
||||||
| θ value | 1.95407e-87 (rank : 6) | NC score | 0.927023 (rank : 6) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q5SVQ0, Q5SVQ1, Q5SVQ2, Q5SVQ3, Q5SVQ7, Q6PGC6, Q80Y65 | Gene names | Myst2, Hbo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST2 (EC 2.3.1.48) (MYST protein 2) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 2) (Histone acetyltransferase binding to ORC1). | |||||
|
MYST3_HUMAN
|
||||||
| θ value | 2.47191e-82 (rank : 7) | NC score | 0.719642 (rank : 10) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q92794 | Gene names | MYST3, MOZ, RUNXBP2, ZNF220 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Runt-related transcription factor-binding protein 2) (Monocytic leukemia zinc finger protein) (Zinc finger protein 220). | |||||
|
MYST3_MOUSE
|
||||||
| θ value | 3.22841e-82 (rank : 8) | NC score | 0.733006 (rank : 8) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 688 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8BZ21, Q8BW52, Q8BYH1, Q8C1F3 | Gene names | Myst3, Moz | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Monocytic leukemia zinc finger protein) (Monocytic leukemia zinc finger homolog). | |||||
|
MYST4_MOUSE
|
||||||
| θ value | 2.02684e-76 (rank : 9) | NC score | 0.766123 (rank : 7) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8BRB7, Q7TNW5, Q8BG35, Q8C441, Q9JKX5 | Gene names | Myst4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Querkopf protein). | |||||
|
MYST4_HUMAN
|
||||||
| θ value | 4.99229e-75 (rank : 10) | NC score | 0.727395 (rank : 9) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8WYB5, O15087, Q86Y05, Q8WU81, Q9UKW2, Q9UKW3, Q9UKX0 | Gene names | MYST4, KIAA0383, MORF, MOZ2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Histone acetyltransferase MOZ2) (Monocytic leukemia zinc finger protein- related factor) (Histone acetyltransferase MORF). | |||||
|
PCLO_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 11) | NC score | 0.054137 (rank : 75) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
ARI4A_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 12) | NC score | 0.086835 (rank : 30) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 434 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P29374, Q15991, Q15992, Q15993 | Gene names | ARID4A, RBBP1, RBP1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 4A (ARID domain- containing protein 4A) (Retinoblastoma-binding protein 1) (RBBP-1). | |||||
|
MUC7_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 13) | NC score | 0.062704 (rank : 54) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 538 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8TAX7 | Gene names | MUC7, MG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-7 precursor (MUC-7) (Salivary mucin-7) (Apo-MG2). | |||||
|
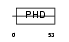
AF17_HUMAN
|
||||||
| θ value | 0.21417 (rank : 14) | NC score | 0.049488 (rank : 90) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P55198, Q9UF49 | Gene names | MLLT6, AF17 | |||
|
Domain Architecture |
|
|||||
| Description | Protein AF-17. | |||||
|
MDC1_HUMAN
|
||||||
| θ value | 0.21417 (rank : 15) | NC score | 0.072083 (rank : 43) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
TAF4_HUMAN
|
||||||
| θ value | 0.279714 (rank : 16) | NC score | 0.052599 (rank : 80) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 687 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O00268, Q5TBP6, Q99721, Q9BR40, Q9BX42 | Gene names | TAF4, TAF2C, TAF2C1, TAF4A, TAFII130, TAFII135 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 4 (TBP-associated factor 4) (Transcription initiation factor TFIID 135 kDa subunit) (TAF(II)135) (TAFII-135) (TAFII135) (TAFII-130) (TAFII130). | |||||
|
MINT_MOUSE
|
||||||
| θ value | 0.365318 (rank : 17) | NC score | 0.048181 (rank : 92) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 0.365318 (rank : 18) | NC score | 0.059157 (rank : 65) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
TCOF_MOUSE
|
||||||
| θ value | 0.365318 (rank : 19) | NC score | 0.053273 (rank : 77) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O08784, O08857 | Gene names | Tcof1 | |||
|
Domain Architecture |

|
|||||
| Description | Treacle protein (Treacher Collins syndrome protein homolog). | |||||
|
CBP_MOUSE
|
||||||
| θ value | 0.47712 (rank : 20) | NC score | 0.059707 (rank : 61) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
CSKI1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 21) | NC score | 0.022171 (rank : 118) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1031 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q6P9K8, Q6ZPU2, Q8BWU2, Q8BX99, Q9CXH0 | Gene names | Caskin1, Kiaa1306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-1 (CASK-interacting protein 1). | |||||
|
SET1A_HUMAN
|
||||||
| θ value | 0.47712 (rank : 22) | NC score | 0.051061 (rank : 84) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O15047, Q6PIF3, Q8TAJ6 | Gene names | SETD1A, KIAA0339, SET1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET1 (EC 2.1.1.43) (Set1/Ash2 histone methyltransferase complex subunit SET1) (SET-domain-containing protein 1A). | |||||
|
SORC1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 23) | NC score | 0.022768 (rank : 116) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8WY21, Q59GG7, Q5JVT7, Q5JVT8, Q5VY14, Q86WQ1, Q86WQ2, Q9H1Y1, Q9H1Y2 | Gene names | SORCS1, SORCS | |||
|
Domain Architecture |

|
|||||
| Description | VPS10 domain-containing receptor SorCS1 precursor (hSorCS). | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | 0.62314 (rank : 24) | NC score | 0.055985 (rank : 72) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
ST5_HUMAN
|
||||||
| θ value | 0.62314 (rank : 25) | NC score | 0.025785 (rank : 113) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P78524, P78523, Q16492, Q7KYY2, Q7KZ12, Q8NE12, Q9BQQ6 | Gene names | ST5, DENND2B, HTS1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppression of tumorigenicity 5 (HeLa tumor suppression 1) (DENN domain-containing protein 2B). | |||||
|
UBP31_HUMAN
|
||||||
| θ value | 0.62314 (rank : 26) | NC score | 0.025841 (rank : 112) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 293 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q70CQ4, Q6AW97, Q6ZTC0, Q6ZTN2, Q9ULL7 | Gene names | USP31, KIAA1203 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 31 (EC 3.1.2.15) (Ubiquitin thioesterase 31) (Ubiquitin-specific-processing protease 31) (Deubiquitinating enzyme 31). | |||||
|
ZMYM3_HUMAN
|
||||||
| θ value | 0.62314 (rank : 27) | NC score | 0.032233 (rank : 109) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q14202, O15089 | Gene names | ZMYM3, DXS6673E, KIAA0385, ZNF261 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger MYM-type protein 3 (Zinc finger protein 261). | |||||
|
CBP_HUMAN
|
||||||
| θ value | 0.813845 (rank : 28) | NC score | 0.042720 (rank : 99) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
FYB_HUMAN
|
||||||
| θ value | 0.813845 (rank : 29) | NC score | 0.052095 (rank : 82) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 379 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O15117, O00359 | Gene names | FYB, SLAP130 | |||
|
Domain Architecture |

|
|||||
| Description | FYN-binding protein (FYN-T-binding protein) (FYB-120/130) (p120/p130) (SLP-76-associated phosphoprotein) (SLAP-130). | |||||
|
RGS3_MOUSE
|
||||||
| θ value | 0.813845 (rank : 30) | NC score | 0.046227 (rank : 95) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
SYNJ1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 31) | NC score | 0.033958 (rank : 107) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8CHC4 | Gene names | Synj1, Kiaa0910 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptojanin-1 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 1). | |||||
|
ZBT38_HUMAN
|
||||||
| θ value | 0.813845 (rank : 32) | NC score | 0.005741 (rank : 146) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 885 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8NAP3 | Gene names | ZBTB38 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger and BTB domain-containing protein 38. | |||||
|
GDF5_HUMAN
|
||||||
| θ value | 1.06291 (rank : 33) | NC score | 0.016168 (rank : 127) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P43026, Q96SB1 | Gene names | GDF5, CDMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Growth/differentiation factor 5 precursor (GDF-5) (Cartilage-derived morphogenetic protein 1) (CDMP-1) (Radotermin). | |||||
|
SYNPO_HUMAN
|
||||||
| θ value | 1.06291 (rank : 34) | NC score | 0.040126 (rank : 102) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8N3V7, O15271, Q9UPX1 | Gene names | SYNPO, KIAA1029 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptopodin. | |||||
|
TAF6_HUMAN
|
||||||
| θ value | 1.06291 (rank : 35) | NC score | 0.047313 (rank : 94) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P49848 | Gene names | TAF6, TAF2E, TAFII70 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 6 (Transcription initiation factor TFIID 70 kDa subunit) (TAF(II)70) (TAFII-70) (TAFII- 80) (TAFII80). | |||||
|
ATX2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 36) | NC score | 0.043611 (rank : 98) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O70305, P97421 | Gene names | Atxn2, Atx2, Sca2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2 (Spinocerebellar ataxia type 2 protein homolog). | |||||
|
PO6F2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 37) | NC score | 0.018141 (rank : 124) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 639 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8BJI4 | Gene names | Pou6f2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | POU domain, class 6, transcription factor 2. | |||||
|
RKHD1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 38) | NC score | 0.040267 (rank : 101) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q86XN8 | Gene names | RKHD1, KIAA2031 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger and KH domain-containing protein 1. | |||||
|
PAPOA_MOUSE
|
||||||
| θ value | 1.81305 (rank : 39) | NC score | 0.021353 (rank : 121) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q61183, Q61208, Q61209, Q8K4X2 | Gene names | Papola, Pap, Plap | |||
|
Domain Architecture |

|
|||||
| Description | Poly(A) polymerase alpha (EC 2.7.7.19) (PAP) (Polynucleotide adenylyltransferase). | |||||
|
PCLO_HUMAN
|
||||||
| θ value | 1.81305 (rank : 40) | NC score | 0.047925 (rank : 93) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
SHAN1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 41) | NC score | 0.015581 (rank : 130) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9Y566, Q9NYW9 | Gene names | SHANK1 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 1 (Shank1) (Somatostatin receptor-interacting protein) (SSTR-interacting protein) (SSTRIP). | |||||
|
TACC1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 42) | NC score | 0.020957 (rank : 122) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 667 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q6Y685, Q6Y686 | Gene names | Tacc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transforming acidic coiled-coil-containing protein 1. | |||||
|
TAF6_MOUSE
|
||||||
| θ value | 1.81305 (rank : 43) | NC score | 0.042328 (rank : 100) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q62311 | Gene names | Taf6, Taf2e | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 6 (Transcription initiation factor TFIID 70 kDa subunit) (TAF(II)70) (TAFII-70) (TAFII- 80) (TAFII80) (p80). | |||||
|
CS016_HUMAN
|
||||||
| θ value | 2.36792 (rank : 44) | NC score | 0.038980 (rank : 103) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8ND99 | Gene names | C19orf16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C19orf16. | |||||
|
DDEF2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 45) | NC score | 0.015583 (rank : 129) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 625 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q7SIG6, Q501K1, Q66JN2 | Gene names | Ddef2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Development and differentiation-enhancing factor 2 (Pyk2 C-terminus- associated protein) (PAP) (Paxillin-associated protein with ARFGAP activity 3) (PAG3). | |||||
|
LRIG2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 46) | NC score | 0.004519 (rank : 149) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 595 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O94898, Q9NSN2 | Gene names | LRIG2, KIAA0806, LIG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats and immunoglobulin-like domains protein 2 precursor (LIG-2). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | 2.36792 (rank : 47) | NC score | 0.045408 (rank : 96) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
RHG26_HUMAN
|
||||||
| θ value | 2.36792 (rank : 48) | NC score | 0.013303 (rank : 134) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UNA1, O75117, Q9BYS6, Q9BYS7, Q9UJ00 | Gene names | ARHGAP26, GRAF, KIAA0621, OPHN1L | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 26 (Oligophrenin-1-like protein) (GTPase regulator associated with focal adhesion kinase). | |||||
|
TREF1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 49) | NC score | 0.021868 (rank : 119) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8BXJ2, Q6PCQ4, Q80X25, Q810H8, Q8BY31 | Gene names | Trerf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcriptional-regulating factor 1 (Transcriptional-regulating protein 132) (Zinc finger transcription factor TReP-132). | |||||
|
ADDB_HUMAN
|
||||||
| θ value | 3.0926 (rank : 50) | NC score | 0.027975 (rank : 110) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P35612, Q13482, Q16412, Q59G82, Q5U5P4, Q6P0P2, Q6PGQ4, Q7Z688, Q7Z689, Q7Z690, Q7Z691 | Gene names | ADD2, ADDB | |||
|
Domain Architecture |

|
|||||
| Description | Beta-adducin (Erythrocyte adducin subunit beta). | |||||
|
CROCC_HUMAN
|
||||||
| θ value | 3.0926 (rank : 51) | NC score | 0.005261 (rank : 147) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1409 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q5TZA2, Q2VHY3, Q66GT7, Q7Z2L4, Q7Z5D7 | Gene names | CROCC, KIAA0445 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rootletin (Ciliary rootlet coiled-coil protein). | |||||
|
K1802_HUMAN
|
||||||
| θ value | 3.0926 (rank : 52) | NC score | 0.026864 (rank : 111) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1378 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q96JM3, Q6P181, Q8NC88, Q9BST0 | Gene names | KIAA1802, C13orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
MLL2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 53) | NC score | 0.110088 (rank : 18) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
PININ_MOUSE
|
||||||
| θ value | 3.0926 (rank : 54) | NC score | 0.023346 (rank : 115) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 777 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O35691, Q8CD89, Q8CGU3 | Gene names | Pnn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pinin. | |||||
|
SRBS1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 55) | NC score | 0.019639 (rank : 123) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q62417, Q80TF8, Q8BZI3, Q8K3Y2, Q921F8, Q9Z0Z8, Q9Z0Z9 | Gene names | Sorbs1, Kiaa1296, Sh3d5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sorbin and SH3 domain-containing protein 1 (Ponsin) (c-Cbl-associated protein) (CAP) (SH3 domain protein 5) (SH3P12). | |||||
|
TCOF_HUMAN
|
||||||
| θ value | 3.0926 (rank : 56) | NC score | 0.037895 (rank : 105) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q13428, Q99408, Q99860 | Gene names | TCOF1 | |||
|
Domain Architecture |

|
|||||
| Description | Treacle protein (Treacher Collins syndrome protein). | |||||
|
TXND2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 57) | NC score | 0.090829 (rank : 28) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
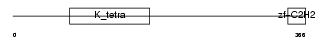
ZBT26_HUMAN
|
||||||
| θ value | 3.0926 (rank : 58) | NC score | 0.002957 (rank : 150) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 784 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9HCK0, Q8WTR1 | Gene names | ZBTB26, KIAA1572, ZNF481 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger and BTB domain-containing protein 26 (Zinc finger protein 481) (Zinc finger protein Bioref). | |||||
|
ZN555_HUMAN
|
||||||
| θ value | 3.0926 (rank : 59) | NC score | 0.001004 (rank : 153) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 741 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8NEP9, Q8NA46, Q96MP1 | Gene names | ZNF555 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 555. | |||||
|
CTDP1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 60) | NC score | 0.037898 (rank : 104) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9Y5B0, Q7Z644, Q96BZ1, Q9Y6F5 | Gene names | CTDP1, FCP1 | |||
|
Domain Architecture |

|
|||||
| Description | RNA polymerase II subunit A C-terminal domain phosphatase (EC 3.1.3.16) (TFIIF-associating CTD phosphatase). | |||||
|
CTGE2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 61) | NC score | 0.009168 (rank : 140) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 594 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96RT6 | Gene names | CTAGE1, CTAGE2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein cTAGE-2. | |||||
|
GNAS1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 62) | NC score | 0.009599 (rank : 139) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q5JWF2, O75684, O75685, Q5JW67, Q5JWF1, Q9NY42 | Gene names | GNAS, GNAS1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Guanine nucleotide-binding protein G(s) subunit alpha isoforms XLas (Adenylate cyclase-stimulating G alpha protein) (Extra large alphas protein) (XLalphas). | |||||
|
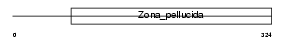
GP2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 63) | NC score | 0.015245 (rank : 131) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P55259, Q13338, Q9UIF1 | Gene names | GP2 | |||
|
Domain Architecture |
|
|||||
| Description | Pancreatic secretory granule membrane major glycoprotein GP2 precursor (Pancreatic zymogen granule membrane protein GP-2) (ZAP75). | |||||
|
K1802_MOUSE
|
||||||
| θ value | 4.03905 (rank : 64) | NC score | 0.032981 (rank : 108) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
DB125_HUMAN
|
||||||
| θ value | 5.27518 (rank : 65) | NC score | 0.036452 (rank : 106) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8N687, Q7Z7B9 | Gene names | DEFB125, DEFB25 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-defensin 125 precursor (Defensin, beta 125) (Beta-defensin 25) (DEFB-25). | |||||
|
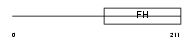
FOXQ1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 66) | NC score | 0.008520 (rank : 142) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9C009, Q9NS06 | Gene names | FOXQ1, HFH1 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein Q1 (Hepatocyte nuclear factor 3 forkhead homolog 1) (HNF-3/forkhead-like protein 1) (HFH-1). | |||||
|
MAP1B_HUMAN
|
||||||
| θ value | 5.27518 (rank : 67) | NC score | 0.048402 (rank : 91) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P46821 | Gene names | MAP1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) [Contains: MAP1 light chain LC1]. | |||||
|
SYP2L_HUMAN
|
||||||
| θ value | 5.27518 (rank : 68) | NC score | 0.021696 (rank : 120) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 400 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9H987 | Gene names | SYNPO2L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptopodin 2-like protein. | |||||
|
ZAN_HUMAN
|
||||||
| θ value | 5.27518 (rank : 69) | NC score | 0.024013 (rank : 114) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 679 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9Y493, O00218, Q96L85, Q96L86, Q96L87, Q96L88, Q96L89, Q96L90, Q9BXN9, Q9BZ83, Q9BZ84, Q9BZ85, Q9BZ86, Q9BZ87, Q9BZ88 | Gene names | ZAN | |||
|
Domain Architecture |

|
|||||
| Description | Zonadhesin precursor. | |||||
|
ZAN_MOUSE
|
||||||
| θ value | 5.27518 (rank : 70) | NC score | 0.012978 (rank : 135) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 808 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O88799, O08647 | Gene names | Zan | |||
|
Domain Architecture |

|
|||||
| Description | Zonadhesin precursor. | |||||
|
ADDB_MOUSE
|
||||||
| θ value | 6.88961 (rank : 71) | NC score | 0.050694 (rank : 86) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 574 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9QYB8, Q3U0E1, Q80VH9, Q8C1C4, Q9CXE3, Q9JLE5, Q9QYB7 | Gene names | Add2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-adducin (Erythrocyte adducin subunit beta) (Add97). | |||||
|
CCNK_MOUSE
|
||||||
| θ value | 6.88961 (rank : 72) | NC score | 0.022462 (rank : 117) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O88874, Q8R068 | Gene names | Ccnk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-K. | |||||
|
CCNL2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 73) | NC score | 0.015758 (rank : 128) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9JJA7, Q5XK66, Q60995, Q8C136, Q8CIJ8, Q99L73, Q9QXH5 | Gene names | Ccnl2, Ania6b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-L2 (Cyclin Ania-6b) (Paneth cell-enhanced expression protein) (PCEE). | |||||
|
CHD5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 74) | NC score | 0.050682 (rank : 87) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8TDI0, O75032, Q5TG89, Q9UFR9 | Gene names | CHD5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain helicase-DNA-binding protein 5 (EC 3.6.1.-) (ATP- dependent helicase CHD5) (CHD-5). | |||||
|
CN092_MOUSE
|
||||||
| θ value | 6.88961 (rank : 75) | NC score | 0.017598 (rank : 125) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8BU11, Q80UI2, Q99PN9, Q9CS16 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | Epidermal Langerhans cell protein LCP1. | |||||
|
GGA2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 76) | NC score | 0.011362 (rank : 137) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UJY4, O14564, Q9NYN2, Q9UPS2 | Gene names | GGA2, KIAA1080 | |||
|
Domain Architecture |

|
|||||
| Description | ADP-ribosylation factor-binding protein GGA2 (Golgi-localized, gamma ear-containing, ARF-binding protein 2) (Gamma-adaptin-related protein 2) (VHS domain and ear domain of gamma-adaptin) (Vear). | |||||
|
NGAP_HUMAN
|
||||||
| θ value | 6.88961 (rank : 77) | NC score | 0.012585 (rank : 136) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 517 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UJF2, O95174, Q5TFU9 | Gene names | RASAL2, NGAP | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein nGAP (RAS protein activator-like 1). | |||||
|
PALB2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 78) | NC score | 0.015234 (rank : 132) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q86YC2, Q8N7Y6, Q8ND31, Q9H6W1 | Gene names | PALB2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Partner and localizer of BRCA2. | |||||
|
REPS2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 79) | NC score | 0.016689 (rank : 126) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8NFH8, O43428, Q8NFI5 | Gene names | REPS2, POB1 | |||
|
Domain Architecture |

|
|||||
| Description | RalBP1-associated Eps domain-containing protein 2 (RalBP1-interacting protein 2) (Partner of RalBP1). | |||||
|
TITIN_HUMAN
|
||||||
| θ value | 6.88961 (rank : 80) | NC score | 0.004767 (rank : 148) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 2923 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q8WZ42, Q10465, Q10466, Q15598, Q2XUS3, Q32Q60, Q4U1Z6, Q6NSG0, Q6PDB1, Q6PJP0, Q7KYM2, Q7KYN4, Q7KYN5, Q7LDM3, Q7Z2X3, Q8TCG8, Q8WZ51, Q8WZ52, Q8WZ53, Q8WZB3, Q92761, Q92762, Q9UD97, Q9UP84, Q9Y6L9 | Gene names | TTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Titin (EC 2.7.11.1) (Connectin) (Rhabdomyosarcoma antigen MU-RMS- 40.14). | |||||
|
ZN580_MOUSE
|
||||||
| θ value | 6.88961 (rank : 81) | NC score | 0.001642 (rank : 151) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 711 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9DB38 | Gene names | Znf580, Zfp580 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 580. | |||||
|
FOG1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 82) | NC score | 0.008905 (rank : 141) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1237 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O35615 | Gene names | Zfpm1, Fog, Fog1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein ZFPM1 (Zinc finger protein multitype 1) (Friend of GATA protein 1) (Friend of GATA-1) (FOG-1). | |||||
|
FOXB1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 83) | NC score | 0.006336 (rank : 144) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q99853, O60652, O75917 | Gene names | FOXB1, FKH5 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein B1 (Transcription factor FKH-5). | |||||
|
FOXB1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 84) | NC score | 0.006418 (rank : 143) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q64732, O08753 | Gene names | Foxb1, Fkh5, Foxb1a, Foxb1b, Mf3 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein B1 (Transcription factor FKH-5). | |||||
|
IF4G1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 85) | NC score | 0.013459 (rank : 133) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q04637, O43177, O95066, Q5HYG0, Q6ZN21, Q8N102 | Gene names | EIF4G1, EIF4G, EIF4GI | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1) (p220). | |||||
|
LATS1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 86) | NC score | 0.001282 (rank : 152) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1027 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8BYR2, Q9Z0W4 | Gene names | Lats1, Warts | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase LATS1 (EC 2.7.11.1) (Large tumor suppressor homolog 1) (WARTS protein kinase). | |||||
|
RBP2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 87) | NC score | 0.005912 (rank : 145) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P49792, Q13074, Q15280, Q53TE2, Q59FH7 | Gene names | RANBP2, NUP358 | |||
|
Domain Architecture |

|
|||||
| Description | Ran-binding protein 2 (RanBP2) (Nuclear pore complex protein Nup358) (Nucleoporin Nup358) (358 kDa nucleoporin) (P270). | |||||
|
SRRM1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 88) | NC score | 0.044773 (rank : 97) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
US6NL_MOUSE
|
||||||
| θ value | 8.99809 (rank : 89) | NC score | 0.009718 (rank : 138) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q80XC3, Q6ZQK8 | Gene names | Usp6nl, Kiaa0019 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USP6 N-terminal-like protein. | |||||
|
ZN324_HUMAN
|
||||||
| θ value | 8.99809 (rank : 90) | NC score | -0.000452 (rank : 154) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 765 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O75467 | Gene names | ZNF324 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 324 (Zinc finger protein ZF5128). | |||||
|
AIRE_HUMAN
|
||||||
| θ value | θ > 10 (rank : 91) | NC score | 0.092211 (rank : 26) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O43918, O43922, O43932, O75745 | Gene names | AIRE, APECED | |||
|
Domain Architecture |

|
|||||
| Description | Autoimmune regulator (Autoimmune polyendocrinopathy candidiasis ectodermal dystrophy protein) (APECED protein). | |||||
|
AIRE_MOUSE
|
||||||
| θ value | θ > 10 (rank : 92) | NC score | 0.091553 (rank : 27) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9Z0E3, Q9JLW0, Q9JLW1, Q9JLW2, Q9JLW3, Q9JLW4, Q9JLW5, Q9JLW6, Q9JLW7, Q9JLW8, Q9JLW9, Q9JLX0 | Gene names | Aire | |||
|
Domain Architecture |

|
|||||
| Description | Autoimmune regulator (Autoimmune polyendocrinopathy candidiasis ectodermal dystrophy protein homolog) (APECED protein homolog). | |||||
|
AKA12_HUMAN
|
||||||
| θ value | θ > 10 (rank : 93) | NC score | 0.066672 (rank : 49) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q02952, O00310, O00498, Q99970 | Gene names | AKAP12, AKAP250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 12 (A-kinase anchor protein 250 kDa) (AKAP 250) (Myasthenia gravis autoantigen gravin). | |||||
|
ASPH_HUMAN
|
||||||
| θ value | θ > 10 (rank : 94) | NC score | 0.052382 (rank : 81) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q12797 | Gene names | ASPH | |||
|
Domain Architecture |

|
|||||
| Description | Aspartyl/asparaginyl beta-hydroxylase (EC 1.14.11.16) (Aspartate beta- hydroxylase) (ASP beta-hydroxylase) (Peptide-aspartate beta- dioxygenase). | |||||
|
BAZ1A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 95) | NC score | 0.111998 (rank : 17) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 693 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9NRL2, Q9NZ15, Q9P065, Q9UIG1, Q9Y3V3 | Gene names | BAZ1A, ACF1, WCRF180 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1A (ATP-utilizing chromatin assembly and remodeling factor 1) (hACF1) (ATP-dependent chromatin remodelling protein) (Williams syndrome transcription factor-related chromatin remodeling factor 180) (WCRF180) (hWALp1) (CHRAC subunit ACF1). | |||||
|
BAZ1B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 96) | NC score | 0.074296 (rank : 41) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 701 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9UIG0, O95039, O95247, O95277 | Gene names | BAZ1B, WBSC10, WBSCR9, WSTF | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein) (Williams syndrome transcription factor) (hWALP2). | |||||
|
BAZ1B_MOUSE
|
||||||
| θ value | θ > 10 (rank : 97) | NC score | 0.081527 (rank : 35) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9Z277, Q9CU68 | Gene names | Baz1b, Wbscr9 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein homolog) (WBRS9). | |||||
|
BAZ2A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 98) | NC score | 0.084517 (rank : 32) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 780 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9UIF9, O00536, O15030, Q96H26 | Gene names | BAZ2A, KIAA0314, TIP5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5) (hWALp3). | |||||
|
BAZ2A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 99) | NC score | 0.085524 (rank : 31) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q91YE5 | Gene names | Baz2a, Tip5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5). | |||||
|
BAZ2B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.078116 (rank : 37) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 803 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9UIF8, Q96EA1, Q96SQ8, Q9P252, Q9Y4N8 | Gene names | BAZ2B, KIAA1476 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2B (hWALp4). | |||||
|
BPTF_HUMAN
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.064109 (rank : 53) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 786 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q12830, Q6NX67, Q7Z7D6, Q9UIG2 | Gene names | FALZ, BPTF, FAC1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleosome remodeling factor subunit BPTF (Bromodomain and PHD finger- containing transcription factor) (Fetal Alzheimer antigen) (Fetal Alz- 50 clone 1 protein). | |||||
|
CD2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.050131 (rank : 89) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P06729, Q96TE5 | Gene names | CD2 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell surface antigen CD2 precursor (T-cell surface antigen T11/Leu- 5) (LFA-2) (LFA-3 receptor) (Erythrocyte receptor) (Rosette receptor). | |||||
|
CJ012_HUMAN
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.061351 (rank : 58) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8N655, Q9H945, Q9Y457 | Gene names | C10orf12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf12. | |||||
|
DPF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.194398 (rank : 13) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q92782 | Gene names | DPF1, NEUD4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein neuro-d4 (D4, zinc and double PHD fingers family 1). | |||||
|
DPF1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.169903 (rank : 16) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9QX66, Q9QX65, Q9QYA4 | Gene names | Dpf1, Neud4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein neuro-d4 (D4, zinc and double PHD fingers family 1). | |||||
|
DPF3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 106) | NC score | 0.076857 (rank : 38) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P58269 | Gene names | Dpf3, Cerd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc-finger protein Dpf3 (cer-d4). | |||||
|
ERIC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 107) | NC score | 0.068930 (rank : 45) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q86X53, Q9P063 | Gene names | ERICH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate-rich protein 1. | |||||
|
HRX_HUMAN
|
||||||
| θ value | θ > 10 (rank : 108) | NC score | 0.073579 (rank : 42) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
IWS1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 109) | NC score | 0.082946 (rank : 34) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
IWS1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 110) | NC score | 0.075868 (rank : 39) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
JAD1A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 111) | NC score | 0.058697 (rank : 66) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P29375 | Gene names | JARID1A, RBBP2, RBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1A (Retinoblastoma-binding protein 2) (RBBP-2). | |||||
|
JAD1C_HUMAN
|
||||||
| θ value | θ > 10 (rank : 112) | NC score | 0.067836 (rank : 46) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P41229 | Gene names | SMCX, DXS1272E, JARID1C, XE169 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1C (Protein SmcX) (Protein Xe169). | |||||
|
JAD1C_MOUSE
|
||||||
| θ value | θ > 10 (rank : 113) | NC score | 0.067551 (rank : 47) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P41230, O54995, Q9CVI4, Q9D0C3, Q9QVR8, Q9R039 | Gene names | Smcx, Jarid1c, Xe169 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1C (SmcX protein) (Xe169 protein). | |||||
|
JAD1D_HUMAN
|
||||||
| θ value | θ > 10 (rank : 114) | NC score | 0.062320 (rank : 55) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BY66, Q92509, Q92809, Q9HCU1 | Gene names | SMCY, HY, HYA, JARID1D, KIAA0234 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1D (SmcY protein) (Histocompatibility Y antigen) (H-Y). | |||||
|
JAD1D_MOUSE
|
||||||
| θ value | θ > 10 (rank : 115) | NC score | 0.059568 (rank : 62) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q62240, Q9QVR9, Q9R040 | Gene names | Smcy, Hya, Jarid1d | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1D (SmcY protein) (Histocompatibility Y antigen) (H-Y). | |||||
|
JADE2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 116) | NC score | 0.050163 (rank : 88) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q6ZQF7, Q3UHD5, Q6IE83 | Gene names | Phf15, Jade2, Kiaa0239 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-2 (PHD finger protein 15). | |||||
|
KI67_HUMAN
|
||||||
| θ value | θ > 10 (rank : 117) | NC score | 0.053319 (rank : 76) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P46013 | Gene names | MKI67 | |||
|
Domain Architecture |

|
|||||
| Description | Antigen KI-67. | |||||
|
LEO1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 118) | NC score | 0.057753 (rank : 68) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
LEO1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 119) | NC score | 0.060385 (rank : 59) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
MLL3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 120) | NC score | 0.096971 (rank : 24) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
MLL3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 121) | NC score | 0.097073 (rank : 23) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
MLL4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 122) | NC score | 0.079713 (rank : 36) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9UMN6, O15022, O95836, Q96GP2, Q96IP3, Q9UK25, Q9Y668, Q9Y669 | Gene names | MLL4, HRX2, KIAA0304, MLL2, TRX2 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 4 (Trithorax homolog 2). | |||||
|
MYT1L_HUMAN
|
||||||
| θ value | θ > 10 (rank : 123) | NC score | 0.055643 (rank : 73) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UL68, Q6IQ17, Q9UPP6 | Gene names | MYT1L, KIAA1106 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1-like protein (MyT1L protein) (MyT1-L). | |||||
|
NASP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 124) | NC score | 0.066054 (rank : 51) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P49321, Q96A69, Q9BTW2 | Gene names | NASP | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear autoantigenic sperm protein (NASP). | |||||
|
NCOA6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 125) | NC score | 0.059350 (rank : 64) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1202 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q14686, Q9NTZ9, Q9UH74, Q9UK86 | Gene names | NCOA6, AIB3, KIAA0181, RAP250, TRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (PRIP) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC RAP250) (Thyroid hormone receptor-binding protein). | |||||
|
NIPBL_HUMAN
|
||||||
| θ value | θ > 10 (rank : 126) | NC score | 0.056600 (rank : 71) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
NSBP1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 127) | NC score | 0.057247 (rank : 69) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
NSD1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 128) | NC score | 0.059418 (rank : 63) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q96L73, Q96PD8, Q96RN7 | Gene names | NSD1, ARA267 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein) (Androgen receptor-associated coregulator 267). | |||||
|
NSD1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 129) | NC score | 0.066068 (rank : 50) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O88491, Q8C480, Q9CT70 | Gene names | Nsd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein). | |||||
|
OGFR_MOUSE
|
||||||
| θ value | θ > 10 (rank : 130) | NC score | 0.053041 (rank : 79) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 584 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q99PG2, Q99L33 | Gene names | Ogfr | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor). | |||||
|
PF21A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 131) | NC score | 0.108109 (rank : 19) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q96BD5, Q6AWA2, Q9C0G7, Q9H8V9, Q9HAK6, Q9NZE9 | Gene names | PHF21A, BHC80, KIAA1696 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (BHC80a). | |||||
|
PF21A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 132) | NC score | 0.106067 (rank : 20) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6ZPK0, Q6XVG0, Q80Z33, Q8CAZ4, Q8VEC8 | Gene names | Phf21a, Bhc80, Kiaa1696, Pftf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (mBHC80) (BHC80a). | |||||
|
PF21B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 133) | NC score | 0.088615 (rank : 29) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q96EK2, Q5TFL2, Q6ICC0 | Gene names | PHF21B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
PF21B_MOUSE
|
||||||
| θ value | θ > 10 (rank : 134) | NC score | 0.094534 (rank : 25) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8C966 | Gene names | Phf21b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
PHF10_HUMAN
|
||||||
| θ value | θ > 10 (rank : 135) | NC score | 0.219201 (rank : 11) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8WUB8, Q2YDA3, Q9BXD2, Q9NV26 | Gene names | PHF10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 10 (XAP135). | |||||
|
PHF10_MOUSE
|
||||||
| θ value | θ > 10 (rank : 136) | NC score | 0.218346 (rank : 12) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9D8M7, Q99LV5 | Gene names | Phf10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 10. | |||||
|
PHF12_HUMAN
|
||||||
| θ value | θ > 10 (rank : 137) | NC score | 0.102934 (rank : 22) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96QT6, Q6ZML2, Q9BV34, Q9H7U9, Q9P205 | Gene names | PHF12, KIAA1523 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 12 (PHD factor 1) (Pf1). | |||||
|
PHF12_MOUSE
|
||||||
| θ value | θ > 10 (rank : 138) | NC score | 0.103312 (rank : 21) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q5SPL2, Q5SPL3, Q6ZPN8, Q80W50 | Gene names | Phf12, Kiaa1523 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 12 (PHD factor 1) (Pf1). | |||||
|
PHF14_HUMAN
|
||||||
| θ value | θ > 10 (rank : 139) | NC score | 0.062115 (rank : 57) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O94880 | Gene names | PHF14, KIAA0783 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 14. | |||||
|
PHF14_MOUSE
|
||||||
| θ value | θ > 10 (rank : 140) | NC score | 0.064449 (rank : 52) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9D4H9, Q810Z6, Q8C284, Q8CGF8, Q9D1Q8 | Gene names | Phf14 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 14. | |||||
|
PHF7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 141) | NC score | 0.062184 (rank : 56) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BWX1 | Gene names | PHF7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 7 (Testis development protein NYD-SP6). | |||||
|
REQU_HUMAN
|
||||||
| θ value | θ > 10 (rank : 142) | NC score | 0.175339 (rank : 15) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 523 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q92785 | Gene names | DPF2, REQ, UBID4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein ubi-d4 (Requiem) (Apoptosis response zinc finger protein) (D4, zinc and double PHD fingers family 2). | |||||
|
REQU_MOUSE
|
||||||
| θ value | θ > 10 (rank : 143) | NC score | 0.175340 (rank : 14) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q61103, Q3UNP5, Q60663, Q9QYA3 | Gene names | Dpf2, Req, Ubid4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein ubi-d4 (Requiem) (Apoptosis response zinc finger protein) (D4, zinc and double PHD fingers family 2). | |||||
|
RSF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 144) | NC score | 0.083669 (rank : 33) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 539 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96T23, Q86X86, Q9H3L8, Q9NVZ8, Q9NYU0 | Gene names | RSF1, HBXAP, XAP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Remodeling and spacing factor 1 (Rsf-1) (Hepatitis B virus X- associated protein) (HBV pX-associated protein 8) (p325 subunit of RSF chromatin remodelling complex). | |||||
|
TIF1A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 145) | NC score | 0.051838 (rank : 83) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O15164, O95854 | Gene names | TRIM24, RNF82, TIF1, TIF1A | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-alpha (TIF1-alpha) (Tripartite motif-containing protein 24) (RING finger protein 82). | |||||
|
TIF1A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 146) | NC score | 0.053229 (rank : 78) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q64127, Q64126 | Gene names | Trim24, Tif1, Tif1a | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-alpha (TIF1-alpha) (Tripartite motif-containing protein 24). | |||||
|
TP53B_MOUSE
|
||||||
| θ value | θ > 10 (rank : 147) | NC score | 0.071488 (rank : 44) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 644 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P70399, Q68FD0, Q8CI97, Q91YC9 | Gene names | Tp53bp1, Trp53bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor suppressor p53-binding protein 1 (p53-binding protein 1) (p53BP1) (53BP1). | |||||
|
TRI66_HUMAN
|
||||||
| θ value | θ > 10 (rank : 148) | NC score | 0.056837 (rank : 70) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O15016, Q9BQQ4 | Gene names | TRIM66, KIAA0298 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
TRI66_MOUSE
|
||||||
| θ value | θ > 10 (rank : 149) | NC score | 0.050999 (rank : 85) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q924W6 | Gene names | Trim66 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
TXND2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 150) | NC score | 0.058276 (rank : 67) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
UHRF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 151) | NC score | 0.075288 (rank : 40) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 272 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96T88, Q8J022, Q9H6S6, Q9P115, Q9P1U7 | Gene names | UHRF1, ICBP90, NP95, RNF106 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 1 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 1) (Inverted CCAAT box-binding protein of 90 kDa) (Transcription factor ICBP90) (Nuclear zinc finger protein Np95) (Nuclear protein 95) (HuNp95) (RING finger protein 106). | |||||
|
UHRF1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 152) | NC score | 0.067501 (rank : 48) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8VDF2, Q8C6F1, Q8VIA1, Q9Z1H6 | Gene names | Uhrf1, Np95 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 1 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 1) (Nuclear zinc finger protein Np95) (Nuclear protein 95). | |||||
|
UHRF2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 153) | NC score | 0.059982 (rank : 60) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96PU4, Q5VYR1, Q5VYR3, Q659C8, Q8TAG7 | Gene names | UHRF2, NIRF, RNF107 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 2 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 2) (Np95/ICBP90-like RING finger protein) (Np95-like RING finger protein) (Nuclear zinc finger protein Np97) (RING finger protein 107). | |||||
|
UHRF2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 154) | NC score | 0.055175 (rank : 74) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q7TMI3, Q8BG56, Q8BJP6, Q8BY30, Q8K1I5 | Gene names | Uhrf2, Nirf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 2 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 2) (Np95-like ring finger protein) (Nuclear zinc finger protein Np97) (NIRF). | |||||
|
TIP60_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 90 | |
| SwissProt Accessions | Q92993, O95624, Q13430, Q6GSE8, Q9BWK7 | Gene names | HTATIP, TIP60 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase HTATIP (EC 2.3.1.48) (EC 2.3.1.-) (60 kDa Tat interactive protein) (Tip60) (HIV-1 Tat interactive protein) (cPLA(2)-interacting protein). | |||||
|
TIP60_MOUSE
|
||||||
| NC score | 0.999444 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 88 | |
| SwissProt Accessions | Q8CHK4, Q8CGZ3, Q8CGZ4, Q8VIH0 | Gene names | Htatip, Tip60 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase HTATIP (EC 2.3.1.48) (EC 2.3.1.-) (60 kDa Tat interactive protein) (Tip60). | |||||
|
MYST1_HUMAN
|
||||||
| NC score | 0.952888 (rank : 3) | θ value | 5.14231e-88 (rank : 3) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9H7Z6, Q659G0, Q7LC17, Q8IY59, Q8WYB4, Q8WZ14, Q9HAC5, Q9NR35 | Gene names | MYST1, MOF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable histone acetyltransferase MYST1 (EC 2.3.1.48) (MYST protein 1) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 1) (hMOF). | |||||
|
MYST1_MOUSE
|
||||||
| NC score | 0.952429 (rank : 4) | θ value | 5.14231e-88 (rank : 4) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9D1P2, Q3UIY0, Q8BJ69, Q8BJ76, Q8CI73 | Gene names | Myst1, Mof | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable histone acetyltransferase MYST1 (EC 2.3.1.48) (MYST protein 1) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 1). | |||||
|
MYST2_HUMAN
|
||||||
| NC score | 0.927248 (rank : 5) | θ value | 1.95407e-87 (rank : 5) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O95251 | Gene names | MYST2, HBO1, HBOa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST2 (EC 2.3.1.48) (MYST protein 2) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 2) (Histone acetyltransferase binding to ORC1). | |||||
|
MYST2_MOUSE
|
||||||
| NC score | 0.927023 (rank : 6) | θ value | 1.95407e-87 (rank : 6) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q5SVQ0, Q5SVQ1, Q5SVQ2, Q5SVQ3, Q5SVQ7, Q6PGC6, Q80Y65 | Gene names | Myst2, Hbo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST2 (EC 2.3.1.48) (MYST protein 2) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 2) (Histone acetyltransferase binding to ORC1). | |||||
|
MYST4_MOUSE
|
||||||
| NC score | 0.766123 (rank : 7) | θ value | 2.02684e-76 (rank : 9) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8BRB7, Q7TNW5, Q8BG35, Q8C441, Q9JKX5 | Gene names | Myst4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Querkopf protein). | |||||
|
MYST3_MOUSE
|
||||||
| NC score | 0.733006 (rank : 8) | θ value | 3.22841e-82 (rank : 8) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 688 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8BZ21, Q8BW52, Q8BYH1, Q8C1F3 | Gene names | Myst3, Moz | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Monocytic leukemia zinc finger protein) (Monocytic leukemia zinc finger homolog). | |||||
|
MYST4_HUMAN
|
||||||
| NC score | 0.727395 (rank : 9) | θ value | 4.99229e-75 (rank : 10) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8WYB5, O15087, Q86Y05, Q8WU81, Q9UKW2, Q9UKW3, Q9UKX0 | Gene names | MYST4, KIAA0383, MORF, MOZ2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Histone acetyltransferase MOZ2) (Monocytic leukemia zinc finger protein- related factor) (Histone acetyltransferase MORF). | |||||
|
MYST3_HUMAN
|
||||||
| NC score | 0.719642 (rank : 10) | θ value | 2.47191e-82 (rank : 7) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q92794 | Gene names | MYST3, MOZ, RUNXBP2, ZNF220 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Runt-related transcription factor-binding protein 2) (Monocytic leukemia zinc finger protein) (Zinc finger protein 220). | |||||
|
PHF10_HUMAN
|
||||||
| NC score | 0.219201 (rank : 11) | θ value | θ > 10 (rank : 135) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8WUB8, Q2YDA3, Q9BXD2, Q9NV26 | Gene names | PHF10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 10 (XAP135). | |||||
|
PHF10_MOUSE
|
||||||
| NC score | 0.218346 (rank : 12) | θ value | θ > 10 (rank : 136) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9D8M7, Q99LV5 | Gene names | Phf10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 10. | |||||
|
DPF1_HUMAN
|
||||||
| NC score | 0.194398 (rank : 13) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q92782 | Gene names | DPF1, NEUD4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein neuro-d4 (D4, zinc and double PHD fingers family 1). | |||||
|
REQU_MOUSE
|
||||||
| NC score | 0.175340 (rank : 14) | θ value | θ > 10 (rank : 143) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q61103, Q3UNP5, Q60663, Q9QYA3 | Gene names | Dpf2, Req, Ubid4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein ubi-d4 (Requiem) (Apoptosis response zinc finger protein) (D4, zinc and double PHD fingers family 2). | |||||
|
REQU_HUMAN
|
||||||
| NC score | 0.175339 (rank : 15) | θ value | θ > 10 (rank : 142) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 523 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q92785 | Gene names | DPF2, REQ, UBID4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein ubi-d4 (Requiem) (Apoptosis response zinc finger protein) (D4, zinc and double PHD fingers family 2). | |||||
|
DPF1_MOUSE
|
||||||
| NC score | 0.169903 (rank : 16) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9QX66, Q9QX65, Q9QYA4 | Gene names | Dpf1, Neud4 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc-finger protein neuro-d4 (D4, zinc and double PHD fingers family 1). | |||||
|
BAZ1A_HUMAN
|
||||||
| NC score | 0.111998 (rank : 17) | θ value | θ > 10 (rank : 95) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 693 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9NRL2, Q9NZ15, Q9P065, Q9UIG1, Q9Y3V3 | Gene names | BAZ1A, ACF1, WCRF180 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1A (ATP-utilizing chromatin assembly and remodeling factor 1) (hACF1) (ATP-dependent chromatin remodelling protein) (Williams syndrome transcription factor-related chromatin remodeling factor 180) (WCRF180) (hWALp1) (CHRAC subunit ACF1). | |||||
|
MLL2_HUMAN
|
||||||
| NC score | 0.110088 (rank : 18) | θ value | 3.0926 (rank : 53) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
PF21A_HUMAN
|
||||||
| NC score | 0.108109 (rank : 19) | θ value | θ > 10 (rank : 131) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q96BD5, Q6AWA2, Q9C0G7, Q9H8V9, Q9HAK6, Q9NZE9 | Gene names | PHF21A, BHC80, KIAA1696 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (BHC80a). | |||||
|
PF21A_MOUSE
|
||||||
| NC score | 0.106067 (rank : 20) | θ value | θ > 10 (rank : 132) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6ZPK0, Q6XVG0, Q80Z33, Q8CAZ4, Q8VEC8 | Gene names | Phf21a, Bhc80, Kiaa1696, Pftf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21A (BRAF35-HDAC complex protein BHC80) (mBHC80) (BHC80a). | |||||
|
PHF12_MOUSE
|
||||||
| NC score | 0.103312 (rank : 21) | θ value | θ > 10 (rank : 138) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q5SPL2, Q5SPL3, Q6ZPN8, Q80W50 | Gene names | Phf12, Kiaa1523 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 12 (PHD factor 1) (Pf1). | |||||
|
PHF12_HUMAN
|
||||||
| NC score | 0.102934 (rank : 22) | θ value | θ > 10 (rank : 137) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96QT6, Q6ZML2, Q9BV34, Q9H7U9, Q9P205 | Gene names | PHF12, KIAA1523 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 12 (PHD factor 1) (Pf1). | |||||
|
MLL3_MOUSE
|
||||||
| NC score | 0.097073 (rank : 23) | θ value | θ > 10 (rank : 121) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
MLL3_HUMAN
|
||||||
| NC score | 0.096971 (rank : 24) | θ value | θ > 10 (rank : 120) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
PF21B_MOUSE
|
||||||
| NC score | 0.094534 (rank : 25) | θ value | θ > 10 (rank : 134) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8C966 | Gene names | Phf21b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
AIRE_HUMAN
|
||||||
| NC score | 0.092211 (rank : 26) | θ value | θ > 10 (rank : 91) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O43918, O43922, O43932, O75745 | Gene names | AIRE, APECED | |||
|
Domain Architecture |

|
|||||
| Description | Autoimmune regulator (Autoimmune polyendocrinopathy candidiasis ectodermal dystrophy protein) (APECED protein). | |||||
|
AIRE_MOUSE
|
||||||
| NC score | 0.091553 (rank : 27) | θ value | θ > 10 (rank : 92) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9Z0E3, Q9JLW0, Q9JLW1, Q9JLW2, Q9JLW3, Q9JLW4, Q9JLW5, Q9JLW6, Q9JLW7, Q9JLW8, Q9JLW9, Q9JLX0 | Gene names | Aire | |||
|
Domain Architecture |

|
|||||
| Description | Autoimmune regulator (Autoimmune polyendocrinopathy candidiasis ectodermal dystrophy protein homolog) (APECED protein homolog). | |||||
|
TXND2_HUMAN
|
||||||
| NC score | 0.090829 (rank : 28) | θ value | 3.0926 (rank : 57) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
PF21B_HUMAN
|
||||||
| NC score | 0.088615 (rank : 29) | θ value | θ > 10 (rank : 133) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q96EK2, Q5TFL2, Q6ICC0 | Gene names | PHF21B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
ARI4A_HUMAN
|
||||||
| NC score | 0.086835 (rank : 30) | θ value | 0.0563607 (rank : 12) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 434 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P29374, Q15991, Q15992, Q15993 | Gene names | ARID4A, RBBP1, RBP1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 4A (ARID domain- containing protein 4A) (Retinoblastoma-binding protein 1) (RBBP-1). | |||||
|
BAZ2A_MOUSE
|
||||||
| NC score | 0.085524 (rank : 31) | θ value | θ > 10 (rank : 99) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q91YE5 | Gene names | Baz2a, Tip5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5). | |||||
|
BAZ2A_HUMAN
|
||||||
| NC score | 0.084517 (rank : 32) | θ value | θ > 10 (rank : 98) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 780 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9UIF9, O00536, O15030, Q96H26 | Gene names | BAZ2A, KIAA0314, TIP5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5) (hWALp3). | |||||
|
RSF1_HUMAN
|
||||||
| NC score | 0.083669 (rank : 33) | θ value | θ > 10 (rank : 144) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 539 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96T23, Q86X86, Q9H3L8, Q9NVZ8, Q9NYU0 | Gene names | RSF1, HBXAP, XAP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Remodeling and spacing factor 1 (Rsf-1) (Hepatitis B virus X- associated protein) (HBV pX-associated protein 8) (p325 subunit of RSF chromatin remodelling complex). | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.082946 (rank : 34) | θ value | θ > 10 (rank : 109) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
BAZ1B_MOUSE
|
||||||
| NC score | 0.081527 (rank : 35) | θ value | θ > 10 (rank : 97) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9Z277, Q9CU68 | Gene names | Baz1b, Wbscr9 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein homolog) (WBRS9). | |||||
|
MLL4_HUMAN
|
||||||
| NC score | 0.079713 (rank : 36) | θ value | θ > 10 (rank : 122) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9UMN6, O15022, O95836, Q96GP2, Q96IP3, Q9UK25, Q9Y668, Q9Y669 | Gene names | MLL4, HRX2, KIAA0304, MLL2, TRX2 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 4 (Trithorax homolog 2). | |||||
|
BAZ2B_HUMAN
|
||||||
| NC score | 0.078116 (rank : 37) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 803 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9UIF8, Q96EA1, Q96SQ8, Q9P252, Q9Y4N8 | Gene names | BAZ2B, KIAA1476 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2B (hWALp4). | |||||
|
DPF3_MOUSE
|
||||||
| NC score | 0.076857 (rank : 38) | θ value | θ > 10 (rank : 106) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P58269 | Gene names | Dpf3, Cerd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc-finger protein Dpf3 (cer-d4). | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.075868 (rank : 39) | θ value | θ > 10 (rank : 110) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
UHRF1_HUMAN
|
||||||
| NC score | 0.075288 (rank : 40) | θ value | θ > 10 (rank : 151) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 272 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96T88, Q8J022, Q9H6S6, Q9P115, Q9P1U7 | Gene names | UHRF1, ICBP90, NP95, RNF106 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 1 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 1) (Inverted CCAAT box-binding protein of 90 kDa) (Transcription factor ICBP90) (Nuclear zinc finger protein Np95) (Nuclear protein 95) (HuNp95) (RING finger protein 106). | |||||
|
BAZ1B_HUMAN
|
||||||
| NC score | 0.074296 (rank : 41) | θ value | θ > 10 (rank : 96) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 701 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9UIG0, O95039, O95247, O95277 | Gene names | BAZ1B, WBSC10, WBSCR9, WSTF | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein) (Williams syndrome transcription factor) (hWALP2). | |||||
|
HRX_HUMAN
|
||||||
| NC score | 0.073579 (rank : 42) | θ value | θ > 10 (rank : 108) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
MDC1_HUMAN
|
||||||
| NC score | 0.072083 (rank : 43) | θ value | 0.21417 (rank : 15) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
TP53B_MOUSE
|
||||||
| NC score | 0.071488 (rank : 44) | θ value | θ > 10 (rank : 147) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 644 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P70399, Q68FD0, Q8CI97, Q91YC9 | Gene names | Tp53bp1, Trp53bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor suppressor p53-binding protein 1 (p53-binding protein 1) (p53BP1) (53BP1). | |||||
|
ERIC1_HUMAN
|
||||||
| NC score | 0.068930 (rank : 45) | θ value | θ > 10 (rank : 107) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q86X53, Q9P063 | Gene names | ERICH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate-rich protein 1. | |||||
|
JAD1C_HUMAN
|
||||||
| NC score | 0.067836 (rank : 46) | θ value | θ > 10 (rank : 112) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P41229 | Gene names | SMCX, DXS1272E, JARID1C, XE169 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1C (Protein SmcX) (Protein Xe169). | |||||
|
JAD1C_MOUSE
|
||||||
| NC score | 0.067551 (rank : 47) | θ value | θ > 10 (rank : 113) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P41230, O54995, Q9CVI4, Q9D0C3, Q9QVR8, Q9R039 | Gene names | Smcx, Jarid1c, Xe169 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1C (SmcX protein) (Xe169 protein). | |||||
|
UHRF1_MOUSE
|
||||||
| NC score | 0.067501 (rank : 48) | θ value | θ > 10 (rank : 152) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8VDF2, Q8C6F1, Q8VIA1, Q9Z1H6 | Gene names | Uhrf1, Np95 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 1 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 1) (Nuclear zinc finger protein Np95) (Nuclear protein 95). | |||||
|
AKA12_HUMAN
|
||||||
| NC score | 0.066672 (rank : 49) | θ value | θ > 10 (rank : 93) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q02952, O00310, O00498, Q99970 | Gene names | AKAP12, AKAP250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 12 (A-kinase anchor protein 250 kDa) (AKAP 250) (Myasthenia gravis autoantigen gravin). | |||||
|
NSD1_MOUSE
|
||||||
| NC score | 0.066068 (rank : 50) | θ value | θ > 10 (rank : 129) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O88491, Q8C480, Q9CT70 | Gene names | Nsd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein). | |||||
|
NASP_HUMAN
|
||||||
| NC score | 0.066054 (rank : 51) | θ value | θ > 10 (rank : 124) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P49321, Q96A69, Q9BTW2 | Gene names | NASP | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear autoantigenic sperm protein (NASP). | |||||
|
PHF14_MOUSE
|
||||||
| NC score | 0.064449 (rank : 52) | θ value | θ > 10 (rank : 140) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9D4H9, Q810Z6, Q8C284, Q8CGF8, Q9D1Q8 | Gene names | Phf14 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 14. | |||||
|
BPTF_HUMAN
|
||||||
| NC score | 0.064109 (rank : 53) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 786 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q12830, Q6NX67, Q7Z7D6, Q9UIG2 | Gene names | FALZ, BPTF, FAC1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleosome remodeling factor subunit BPTF (Bromodomain and PHD finger- containing transcription factor) (Fetal Alzheimer antigen) (Fetal Alz- 50 clone 1 protein). | |||||
|
MUC7_HUMAN
|
||||||
| NC score | 0.062704 (rank : 54) | θ value | 0.0736092 (rank : 13) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 538 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8TAX7 | Gene names | MUC7, MG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-7 precursor (MUC-7) (Salivary mucin-7) (Apo-MG2). | |||||
|
JAD1D_HUMAN
|
||||||
| NC score | 0.062320 (rank : 55) | θ value | θ > 10 (rank : 114) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BY66, Q92509, Q92809, Q9HCU1 | Gene names | SMCY, HY, HYA, JARID1D, KIAA0234 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1D (SmcY protein) (Histocompatibility Y antigen) (H-Y). | |||||
|
PHF7_HUMAN
|
||||||
| NC score | 0.062184 (rank : 56) | θ value | θ > 10 (rank : 141) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BWX1 | Gene names | PHF7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 7 (Testis development protein NYD-SP6). | |||||
|
PHF14_HUMAN
|
||||||
| NC score | 0.062115 (rank : 57) | θ value | θ > 10 (rank : 139) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O94880 | Gene names | PHF14, KIAA0783 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 14. | |||||
|
CJ012_HUMAN
|
||||||
| NC score | 0.061351 (rank : 58) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8N655, Q9H945, Q9Y457 | Gene names | C10orf12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf12. | |||||
|
LEO1_MOUSE
|
||||||
| NC score | 0.060385 (rank : 59) | θ value | θ > 10 (rank : 119) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
UHRF2_HUMAN
|
||||||
| NC score | 0.059982 (rank : 60) | θ value | θ > 10 (rank : 153) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96PU4, Q5VYR1, Q5VYR3, Q659C8, Q8TAG7 | Gene names | UHRF2, NIRF, RNF107 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 2 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 2) (Np95/ICBP90-like RING finger protein) (Np95-like RING finger protein) (Nuclear zinc finger protein Np97) (RING finger protein 107). | |||||
|
CBP_MOUSE
|
||||||
| NC score | 0.059707 (rank : 61) | θ value | 0.47712 (rank : 20) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
JAD1D_MOUSE
|
||||||
| NC score | 0.059568 (rank : 62) | θ value | θ > 10 (rank : 115) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q62240, Q9QVR9, Q9R040 | Gene names | Smcy, Hya, Jarid1d | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1D (SmcY protein) (Histocompatibility Y antigen) (H-Y). | |||||
|
NSD1_HUMAN
|
||||||
| NC score | 0.059418 (rank : 63) | θ value | θ > 10 (rank : 128) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q96L73, Q96PD8, Q96RN7 | Gene names | NSD1, ARA267 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein) (Androgen receptor-associated coregulator 267). | |||||
|
NCOA6_HUMAN
|
||||||
| NC score | 0.059350 (rank : 64) | θ value | θ > 10 (rank : 125) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1202 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q14686, Q9NTZ9, Q9UH74, Q9UK86 | Gene names | NCOA6, AIB3, KIAA0181, RAP250, TRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (PRIP) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC RAP250) (Thyroid hormone receptor-binding protein). | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.059157 (rank : 65) | θ value | 0.365318 (rank : 18) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
JAD1A_HUMAN
|
||||||
| NC score | 0.058697 (rank : 66) | θ value | θ > 10 (rank : 111) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P29375 | Gene names | JARID1A, RBBP2, RBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji/ARID domain-containing protein 1A (Retinoblastoma-binding protein 2) (RBBP-2). | |||||
|
TXND2_MOUSE
|
||||||
| NC score | 0.058276 (rank : 67) | θ value | θ > 10 (rank : 150) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
LEO1_HUMAN
|
||||||
| NC score | 0.057753 (rank : 68) | θ value | θ > 10 (rank : 118) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
NSBP1_MOUSE
|
||||||
| NC score | 0.057247 (rank : 69) | θ value | θ > 10 (rank : 127) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
TRI66_HUMAN
|
||||||
| NC score | 0.056837 (rank : 70) | θ value | θ > 10 (rank : 148) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O15016, Q9BQQ4 | Gene names | TRIM66, KIAA0298 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
NIPBL_HUMAN
|
||||||
| NC score | 0.056600 (rank : 71) | θ value | θ > 10 (rank : 126) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.055985 (rank : 72) | θ value | 0.62314 (rank : 24) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
MYT1L_HUMAN
|
||||||
| NC score | 0.055643 (rank : 73) | θ value | θ > 10 (rank : 123) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UL68, Q6IQ17, Q9UPP6 | Gene names | MYT1L, KIAA1106 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1-like protein (MyT1L protein) (MyT1-L). | |||||
|
UHRF2_MOUSE
|
||||||
| NC score | 0.055175 (rank : 74) | θ value | θ > 10 (rank : 154) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q7TMI3, Q8BG56, Q8BJP6, Q8BY30, Q8K1I5 | Gene names | Uhrf2, Nirf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-like PHD and RING finger domain-containing protein 2 (EC 6.3.2.-) (Ubiquitin-like-containing PHD and RING finger domains protein 2) (Np95-like ring finger protein) (Nuclear zinc finger protein Np97) (NIRF). | |||||
|
PCLO_MOUSE
|
||||||
| NC score | 0.054137 (rank : 75) | θ value | 0.0431538 (rank : 11) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
KI67_HUMAN
|
||||||
| NC score | 0.053319 (rank : 76) | θ value | θ > 10 (rank : 117) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P46013 | Gene names | MKI67 | |||
|
Domain Architecture |

|
|||||
| Description | Antigen KI-67. | |||||
|
TCOF_MOUSE
|
||||||
| NC score | 0.053273 (rank : 77) | θ value | 0.365318 (rank : 19) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O08784, O08857 | Gene names | Tcof1 | |||
|
Domain Architecture |

|
|||||
| Description | Treacle protein (Treacher Collins syndrome protein homolog). | |||||
|
TIF1A_MOUSE
|
||||||
| NC score | 0.053229 (rank : 78) | θ value | θ > 10 (rank : 146) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q64127, Q64126 | Gene names | Trim24, Tif1, Tif1a | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-alpha (TIF1-alpha) (Tripartite motif-containing protein 24). | |||||
|
OGFR_MOUSE
|
||||||
| NC score | 0.053041 (rank : 79) | θ value | θ > 10 (rank : 130) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 584 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q99PG2, Q99L33 | Gene names | Ogfr | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor). | |||||
|
TAF4_HUMAN
|
||||||
| NC score | 0.052599 (rank : 80) | θ value | 0.279714 (rank : 16) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 687 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O00268, Q5TBP6, Q99721, Q9BR40, Q9BX42 | Gene names | TAF4, TAF2C, TAF2C1, TAF4A, TAFII130, TAFII135 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 4 (TBP-associated factor 4) (Transcription initiation factor TFIID 135 kDa subunit) (TAF(II)135) (TAFII-135) (TAFII135) (TAFII-130) (TAFII130). | |||||
|
ASPH_HUMAN
|
||||||
| NC score | 0.052382 (rank : 81) | θ value | θ > 10 (rank : 94) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q12797 | Gene names | ASPH | |||
|
Domain Architecture |

|
|||||
| Description | Aspartyl/asparaginyl beta-hydroxylase (EC 1.14.11.16) (Aspartate beta- hydroxylase) (ASP beta-hydroxylase) (Peptide-aspartate beta- dioxygenase). | |||||
|
FYB_HUMAN
|
||||||
| NC score | 0.052095 (rank : 82) | θ value | 0.813845 (rank : 29) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 379 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O15117, O00359 | Gene names | FYB, SLAP130 | |||
|
Domain Architecture |

|
|||||
| Description | FYN-binding protein (FYN-T-binding protein) (FYB-120/130) (p120/p130) (SLP-76-associated phosphoprotein) (SLAP-130). | |||||
|
TIF1A_HUMAN
|
||||||
| NC score | 0.051838 (rank : 83) | θ value | θ > 10 (rank : 145) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O15164, O95854 | Gene names | TRIM24, RNF82, TIF1, TIF1A | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-alpha (TIF1-alpha) (Tripartite motif-containing protein 24) (RING finger protein 82). | |||||
|
SET1A_HUMAN
|
||||||
| NC score | 0.051061 (rank : 84) | θ value | 0.47712 (rank : 22) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O15047, Q6PIF3, Q8TAJ6 | Gene names | SETD1A, KIAA0339, SET1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET1 (EC 2.1.1.43) (Set1/Ash2 histone methyltransferase complex subunit SET1) (SET-domain-containing protein 1A). | |||||
|
TRI66_MOUSE
|
||||||
| NC score | 0.050999 (rank : 85) | θ value | θ > 10 (rank : 149) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q924W6 | Gene names | Trim66 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
ADDB_MOUSE
|
||||||
| NC score | 0.050694 (rank : 86) | θ value | 6.88961 (rank : 71) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 574 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9QYB8, Q3U0E1, Q80VH9, Q8C1C4, Q9CXE3, Q9JLE5, Q9QYB7 | Gene names | Add2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-adducin (Erythrocyte adducin subunit beta) (Add97). | |||||
|
CHD5_HUMAN
|
||||||
| NC score | 0.050682 (rank : 87) | θ value | 6.88961 (rank : 74) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8TDI0, O75032, Q5TG89, Q9UFR9 | Gene names | CHD5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain helicase-DNA-binding protein 5 (EC 3.6.1.-) (ATP- dependent helicase CHD5) (CHD-5). | |||||
|
JADE2_MOUSE
|
||||||
| NC score | 0.050163 (rank : 88) | θ value | θ > 10 (rank : 116) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q6ZQF7, Q3UHD5, Q6IE83 | Gene names | Phf15, Jade2, Kiaa0239 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-2 (PHD finger protein 15). | |||||
|
CD2_HUMAN
|
||||||
| NC score | 0.050131 (rank : 89) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P06729, Q96TE5 | Gene names | CD2 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell surface antigen CD2 precursor (T-cell surface antigen T11/Leu- 5) (LFA-2) (LFA-3 receptor) (Erythrocyte receptor) (Rosette receptor). | |||||
|
AF17_HUMAN
|
||||||
| NC score | 0.049488 (rank : 90) | θ value | 0.21417 (rank : 14) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P55198, Q9UF49 | Gene names | MLLT6, AF17 | |||
|
Domain Architecture |

|
|||||
| Description | Protein AF-17. | |||||
|
MAP1B_HUMAN
|
||||||
| NC score | 0.048402 (rank : 91) | θ value | 5.27518 (rank : 67) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P46821 | Gene names | MAP1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) [Contains: MAP1 light chain LC1]. | |||||
|
MINT_MOUSE
|
||||||
| NC score | 0.048181 (rank : 92) | θ value | 0.365318 (rank : 17) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
PCLO_HUMAN
|
||||||
| NC score | 0.047925 (rank : 93) | θ value | 1.81305 (rank : 40) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
TAF6_HUMAN
|
||||||
| NC score | 0.047313 (rank : 94) | θ value | 1.06291 (rank : 35) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P49848 | Gene names | TAF6, TAF2E, TAFII70 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 6 (Transcription initiation factor TFIID 70 kDa subunit) (TAF(II)70) (TAFII-70) (TAFII- 80) (TAFII80). | |||||
|
RGS3_MOUSE
|
||||||
| NC score | 0.046227 (rank : 95) | θ value | 0.813845 (rank : 30) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.045408 (rank : 96) | θ value | 2.36792 (rank : 47) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
SRRM1_MOUSE
|
||||||
| NC score | 0.044773 (rank : 97) | θ value | 8.99809 (rank : 88) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
ATX2_MOUSE
|
||||||
| NC score | 0.043611 (rank : 98) | θ value | 1.38821 (rank : 36) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O70305, P97421 | Gene names | Atxn2, Atx2, Sca2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2 (Spinocerebellar ataxia type 2 protein homolog). | |||||
|
CBP_HUMAN
|
||||||
| NC score | 0.042720 (rank : 99) | θ value | 0.813845 (rank : 28) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
TAF6_MOUSE
|
||||||
| NC score | 0.042328 (rank : 100) | θ value | 1.81305 (rank : 43) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q62311 | Gene names | Taf6, Taf2e | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 6 (Transcription initiation factor TFIID 70 kDa subunit) (TAF(II)70) (TAFII-70) (TAFII- 80) (TAFII80) (p80). | |||||
|
RKHD1_HUMAN
|
||||||
| NC score | 0.040267 (rank : 101) | θ value | 1.38821 (rank : 38) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q86XN8 | Gene names | RKHD1, KIAA2031 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger and KH domain-containing protein 1. | |||||
|
SYNPO_HUMAN
|
||||||
| NC score | 0.040126 (rank : 102) | θ value | 1.06291 (rank : 34) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8N3V7, O15271, Q9UPX1 | Gene names | SYNPO, KIAA1029 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptopodin. | |||||
|
CS016_HUMAN
|
||||||
| NC score | 0.038980 (rank : 103) | θ value | 2.36792 (rank : 44) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8ND99 | Gene names | C19orf16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C19orf16. | |||||
|
CTDP1_HUMAN
|
||||||
| NC score | 0.037898 (rank : 104) | θ value | 4.03905 (rank : 60) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9Y5B0, Q7Z644, Q96BZ1, Q9Y6F5 | Gene names | CTDP1, FCP1 | |||
|
Domain Architecture |

|
|||||
| Description | RNA polymerase II subunit A C-terminal domain phosphatase (EC 3.1.3.16) (TFIIF-associating CTD phosphatase). | |||||
|
TCOF_HUMAN
|
||||||
| NC score | 0.037895 (rank : 105) | θ value | 3.0926 (rank : 56) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q13428, Q99408, Q99860 | Gene names | TCOF1 | |||
|
Domain Architecture |

|
|||||
| Description | Treacle protein (Treacher Collins syndrome protein). | |||||
|
DB125_HUMAN
|
||||||
| NC score | 0.036452 (rank : 106) | θ value | 5.27518 (rank : 65) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8N687, Q7Z7B9 | Gene names | DEFB125, DEFB25 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-defensin 125 precursor (Defensin, beta 125) (Beta-defensin 25) (DEFB-25). | |||||
|
SYNJ1_MOUSE
|
||||||
| NC score | 0.033958 (rank : 107) | θ value | 0.813845 (rank : 31) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8CHC4 | Gene names | Synj1, Kiaa0910 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptojanin-1 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 1). | |||||
|
K1802_MOUSE
|
||||||
| NC score | 0.032981 (rank : 108) | θ value | 4.03905 (rank : 64) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
ZMYM3_HUMAN
|
||||||
| NC score | 0.032233 (rank : 109) | θ value | 0.62314 (rank : 27) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q14202, O15089 | Gene names | ZMYM3, DXS6673E, KIAA0385, ZNF261 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger MYM-type protein 3 (Zinc finger protein 261). | |||||
|
ADDB_HUMAN
|
||||||
| NC score | 0.027975 (rank : 110) | θ value | 3.0926 (rank : 50) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P35612, Q13482, Q16412, Q59G82, Q5U5P4, Q6P0P2, Q6PGQ4, Q7Z688, Q7Z689, Q7Z690, Q7Z691 | Gene names | ADD2, ADDB | |||
|
Domain Architecture |

|
|||||
| Description | Beta-adducin (Erythrocyte adducin subunit beta). | |||||
|
K1802_HUMAN
|
||||||
| NC score | 0.026864 (rank : 111) | θ value | 3.0926 (rank : 52) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1378 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q96JM3, Q6P181, Q8NC88, Q9BST0 | Gene names | KIAA1802, C13orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
UBP31_HUMAN
|
||||||
| NC score | 0.025841 (rank : 112) | θ value | 0.62314 (rank : 26) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 293 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q70CQ4, Q6AW97, Q6ZTC0, Q6ZTN2, Q9ULL7 | Gene names | USP31, KIAA1203 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 31 (EC 3.1.2.15) (Ubiquitin thioesterase 31) (Ubiquitin-specific-processing protease 31) (Deubiquitinating enzyme 31). | |||||
|
ST5_HUMAN
|
||||||
| NC score | 0.025785 (rank : 113) | θ value | 0.62314 (rank : 25) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P78524, P78523, Q16492, Q7KYY2, Q7KZ12, Q8NE12, Q9BQQ6 | Gene names | ST5, DENND2B, HTS1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppression of tumorigenicity 5 (HeLa tumor suppression 1) (DENN domain-containing protein 2B). | |||||
|
ZAN_HUMAN
|
||||||
| NC score | 0.024013 (rank : 114) | θ value | 5.27518 (rank : 69) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 679 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9Y493, O00218, Q96L85, Q96L86, Q96L87, Q96L88, Q96L89, Q96L90, Q9BXN9, Q9BZ83, Q9BZ84, Q9BZ85, Q9BZ86, Q9BZ87, Q9BZ88 | Gene names | ZAN | |||
|
Domain Architecture |

|
|||||
| Description | Zonadhesin precursor. | |||||
|
PININ_MOUSE
|
||||||
| NC score | 0.023346 (rank : 115) | θ value | 3.0926 (rank : 54) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 777 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O35691, Q8CD89, Q8CGU3 | Gene names | Pnn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pinin. | |||||
|
SORC1_HUMAN
|
||||||
| NC score | 0.022768 (rank : 116) | θ value | 0.47712 (rank : 23) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8WY21, Q59GG7, Q5JVT7, Q5JVT8, Q5VY14, Q86WQ1, Q86WQ2, Q9H1Y1, Q9H1Y2 | Gene names | SORCS1, SORCS | |||
|
Domain Architecture |

|
|||||
| Description | VPS10 domain-containing receptor SorCS1 precursor (hSorCS). | |||||
|
CCNK_MOUSE
|
||||||
| NC score | 0.022462 (rank : 117) | θ value | 6.88961 (rank : 72) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O88874, Q8R068 | Gene names | Ccnk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-K. | |||||
|
CSKI1_MOUSE
|
||||||
| NC score | 0.022171 (rank : 118) | θ value | 0.47712 (rank : 21) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1031 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q6P9K8, Q6ZPU2, Q8BWU2, Q8BX99, Q9CXH0 | Gene names | Caskin1, Kiaa1306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-1 (CASK-interacting protein 1). | |||||
|
TREF1_MOUSE
|
||||||
| NC score | 0.021868 (rank : 119) | θ value | 2.36792 (rank : 49) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8BXJ2, Q6PCQ4, Q80X25, Q810H8, Q8BY31 | Gene names | Trerf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcriptional-regulating factor 1 (Transcriptional-regulating protein 132) (Zinc finger transcription factor TReP-132). | |||||
|
SYP2L_HUMAN
|
||||||
| NC score | 0.021696 (rank : 120) | θ value | 5.27518 (rank : 68) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 400 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9H987 | Gene names | SYNPO2L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptopodin 2-like protein. | |||||
|
PAPOA_MOUSE
|
||||||
| NC score | 0.021353 (rank : 121) | θ value | 1.81305 (rank : 39) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q61183, Q61208, Q61209, Q8K4X2 | Gene names | Papola, Pap, Plap | |||
|
Domain Architecture |

|
|||||
| Description | Poly(A) polymerase alpha (EC 2.7.7.19) (PAP) (Polynucleotide adenylyltransferase). | |||||
|
TACC1_MOUSE
|
||||||
| NC score | 0.020957 (rank : 122) | θ value | 1.81305 (rank : 42) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 667 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q6Y685, Q6Y686 | Gene names | Tacc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transforming acidic coiled-coil-containing protein 1. | |||||
|
SRBS1_MOUSE
|
||||||
| NC score | 0.019639 (rank : 123) | θ value | 3.0926 (rank : 55) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q62417, Q80TF8, Q8BZI3, Q8K3Y2, Q921F8, Q9Z0Z8, Q9Z0Z9 | Gene names | Sorbs1, Kiaa1296, Sh3d5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sorbin and SH3 domain-containing protein 1 (Ponsin) (c-Cbl-associated protein) (CAP) (SH3 domain protein 5) (SH3P12). | |||||
|
PO6F2_MOUSE
|
||||||
| NC score | 0.018141 (rank : 124) | θ value | 1.38821 (rank : 37) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 639 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8BJI4 | Gene names | Pou6f2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | POU domain, class 6, transcription factor 2. | |||||
|
CN092_MOUSE
|
||||||
| NC score | 0.017598 (rank : 125) | θ value | 6.88961 (rank : 75) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8BU11, Q80UI2, Q99PN9, Q9CS16 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | Epidermal Langerhans cell protein LCP1. | |||||
|
REPS2_HUMAN
|
||||||
| NC score | 0.016689 (rank : 126) | θ value | 6.88961 (rank : 79) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8NFH8, O43428, Q8NFI5 | Gene names | REPS2, POB1 | |||
|
Domain Architecture |

|
|||||
| Description | RalBP1-associated Eps domain-containing protein 2 (RalBP1-interacting protein 2) (Partner of RalBP1). | |||||
|
GDF5_HUMAN
|
||||||
| NC score | 0.016168 (rank : 127) | θ value | 1.06291 (rank : 33) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P43026, Q96SB1 | Gene names | GDF5, CDMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Growth/differentiation factor 5 precursor (GDF-5) (Cartilage-derived morphogenetic protein 1) (CDMP-1) (Radotermin). | |||||
|
CCNL2_MOUSE
|
||||||
| NC score | 0.015758 (rank : 128) | θ value | 6.88961 (rank : 73) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9JJA7, Q5XK66, Q60995, Q8C136, Q8CIJ8, Q99L73, Q9QXH5 | Gene names | Ccnl2, Ania6b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-L2 (Cyclin Ania-6b) (Paneth cell-enhanced expression protein) (PCEE). | |||||
|
DDEF2_MOUSE
|
||||||
| NC score | 0.015583 (rank : 129) | θ value | 2.36792 (rank : 45) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 625 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q7SIG6, Q501K1, Q66JN2 | Gene names | Ddef2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Development and differentiation-enhancing factor 2 (Pyk2 C-terminus- associated protein) (PAP) (Paxillin-associated protein with ARFGAP activity 3) (PAG3). | |||||
|
SHAN1_HUMAN
|
||||||
| NC score | 0.015581 (rank : 130) | θ value | 1.81305 (rank : 41) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9Y566, Q9NYW9 | Gene names | SHANK1 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 1 (Shank1) (Somatostatin receptor-interacting protein) (SSTR-interacting protein) (SSTRIP). | |||||
|
GP2_HUMAN
|
||||||
| NC score | 0.015245 (rank : 131) | θ value | 4.03905 (rank : 63) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P55259, Q13338, Q9UIF1 | Gene names | GP2 | |||
|
Domain Architecture |

|
|||||
| Description | Pancreatic secretory granule membrane major glycoprotein GP2 precursor (Pancreatic zymogen granule membrane protein GP-2) (ZAP75). | |||||
|
PALB2_HUMAN
|
||||||
| NC score | 0.015234 (rank : 132) | θ value | 6.88961 (rank : 78) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q86YC2, Q8N7Y6, Q8ND31, Q9H6W1 | Gene names | PALB2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Partner and localizer of BRCA2. | |||||
|
IF4G1_HUMAN
|
||||||
| NC score | 0.013459 (rank : 133) | θ value | 8.99809 (rank : 85) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q04637, O43177, O95066, Q5HYG0, Q6ZN21, Q8N102 | Gene names | EIF4G1, EIF4G, EIF4GI | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 1 (eIF-4-gamma 1) (eIF-4G1) (eIF-4G 1) (p220). | |||||
|
RHG26_HUMAN
|
||||||
| NC score | 0.013303 (rank : 134) | θ value | 2.36792 (rank : 48) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UNA1, O75117, Q9BYS6, Q9BYS7, Q9UJ00 | Gene names | ARHGAP26, GRAF, KIAA0621, OPHN1L | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 26 (Oligophrenin-1-like protein) (GTPase regulator associated with focal adhesion kinase). | |||||
|
ZAN_MOUSE
|
||||||
| NC score | 0.012978 (rank : 135) | θ value | 5.27518 (rank : 70) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 808 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O88799, O08647 | Gene names | Zan | |||
|
Domain Architecture |

|
|||||
| Description | Zonadhesin precursor. | |||||
|
NGAP_HUMAN
|
||||||
| NC score | 0.012585 (rank : 136) | θ value | 6.88961 (rank : 77) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 517 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UJF2, O95174, Q5TFU9 | Gene names | RASAL2, NGAP | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein nGAP (RAS protein activator-like 1). | |||||
|
GGA2_HUMAN
|
||||||
| NC score | 0.011362 (rank : 137) | θ value | 6.88961 (rank : 76) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UJY4, O14564, Q9NYN2, Q9UPS2 | Gene names | GGA2, KIAA1080 | |||
|
Domain Architecture |

|
|||||
| Description | ADP-ribosylation factor-binding protein GGA2 (Golgi-localized, gamma ear-containing, ARF-binding protein 2) (Gamma-adaptin-related protein 2) (VHS domain and ear domain of gamma-adaptin) (Vear). | |||||
|
US6NL_MOUSE
|
||||||
| NC score | 0.009718 (rank : 138) | θ value | 8.99809 (rank : 89) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q80XC3, Q6ZQK8 | Gene names | Usp6nl, Kiaa0019 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USP6 N-terminal-like protein. | |||||
|
GNAS1_HUMAN
|
||||||
| NC score | 0.009599 (rank : 139) | θ value | 4.03905 (rank : 62) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q5JWF2, O75684, O75685, Q5JW67, Q5JWF1, Q9NY42 | Gene names | GNAS, GNAS1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Guanine nucleotide-binding protein G(s) subunit alpha isoforms XLas (Adenylate cyclase-stimulating G alpha protein) (Extra large alphas protein) (XLalphas). | |||||
|
CTGE2_HUMAN
|
||||||
| NC score | 0.009168 (rank : 140) | θ value | 4.03905 (rank : 61) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 594 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96RT6 | Gene names | CTAGE1, CTAGE2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein cTAGE-2. | |||||
|
FOG1_MOUSE
|
||||||
| NC score | 0.008905 (rank : 141) | θ value | 8.99809 (rank : 82) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1237 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O35615 | Gene names | Zfpm1, Fog, Fog1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein ZFPM1 (Zinc finger protein multitype 1) (Friend of GATA protein 1) (Friend of GATA-1) (FOG-1). | |||||
|
FOXQ1_HUMAN
|
||||||
| NC score | 0.008520 (rank : 142) | θ value | 5.27518 (rank : 66) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9C009, Q9NS06 | Gene names | FOXQ1, HFH1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein Q1 (Hepatocyte nuclear factor 3 forkhead homolog 1) (HNF-3/forkhead-like protein 1) (HFH-1). | |||||
|
FOXB1_MOUSE
|
||||||
| NC score | 0.006418 (rank : 143) | θ value | 8.99809 (rank : 84) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q64732, O08753 | Gene names | Foxb1, Fkh5, Foxb1a, Foxb1b, Mf3 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein B1 (Transcription factor FKH-5). | |||||
|
FOXB1_HUMAN
|
||||||
| NC score | 0.006336 (rank : 144) | θ value | 8.99809 (rank : 83) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q99853, O60652, O75917 | Gene names | FOXB1, FKH5 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein B1 (Transcription factor FKH-5). | |||||
|
RBP2_HUMAN
|
||||||
| NC score | 0.005912 (rank : 145) | θ value | 8.99809 (rank : 87) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P49792, Q13074, Q15280, Q53TE2, Q59FH7 | Gene names | RANBP2, NUP358 | |||
|
Domain Architecture |

|
|||||
| Description | Ran-binding protein 2 (RanBP2) (Nuclear pore complex protein Nup358) (Nucleoporin Nup358) (358 kDa nucleoporin) (P270). | |||||
|
ZBT38_HUMAN
|
||||||
| NC score | 0.005741 (rank : 146) | θ value | 0.813845 (rank : 32) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 885 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8NAP3 | Gene names | ZBTB38 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger and BTB domain-containing protein 38. | |||||
|
CROCC_HUMAN
|
||||||
| NC score | 0.005261 (rank : 147) | θ value | 3.0926 (rank : 51) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1409 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q5TZA2, Q2VHY3, Q66GT7, Q7Z2L4, Q7Z5D7 | Gene names | CROCC, KIAA0445 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rootletin (Ciliary rootlet coiled-coil protein). | |||||
|
TITIN_HUMAN
|
||||||
| NC score | 0.004767 (rank : 148) | θ value | 6.88961 (rank : 80) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 2923 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q8WZ42, Q10465, Q10466, Q15598, Q2XUS3, Q32Q60, Q4U1Z6, Q6NSG0, Q6PDB1, Q6PJP0, Q7KYM2, Q7KYN4, Q7KYN5, Q7LDM3, Q7Z2X3, Q8TCG8, Q8WZ51, Q8WZ52, Q8WZ53, Q8WZB3, Q92761, Q92762, Q9UD97, Q9UP84, Q9Y6L9 | Gene names | TTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Titin (EC 2.7.11.1) (Connectin) (Rhabdomyosarcoma antigen MU-RMS- 40.14). | |||||
|
LRIG2_HUMAN
|
||||||
| NC score | 0.004519 (rank : 149) | θ value | 2.36792 (rank : 46) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 595 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O94898, Q9NSN2 | Gene names | LRIG2, KIAA0806, LIG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeats and immunoglobulin-like domains protein 2 precursor (LIG-2). | |||||
|
ZBT26_HUMAN
|
||||||
| NC score | 0.002957 (rank : 150) | θ value | 3.0926 (rank : 58) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 784 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9HCK0, Q8WTR1 | Gene names | ZBTB26, KIAA1572, ZNF481 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 26 (Zinc finger protein 481) (Zinc finger protein Bioref). | |||||
|
ZN580_MOUSE
|
||||||
| NC score | 0.001642 (rank : 151) | θ value | 6.88961 (rank : 81) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 711 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9DB38 | Gene names | Znf580, Zfp580 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 580. | |||||
|
LATS1_MOUSE
|
||||||
| NC score | 0.001282 (rank : 152) | θ value | 8.99809 (rank : 86) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1027 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8BYR2, Q9Z0W4 | Gene names | Lats1, Warts | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase LATS1 (EC 2.7.11.1) (Large tumor suppressor homolog 1) (WARTS protein kinase). | |||||
|
ZN555_HUMAN
|
||||||
| NC score | 0.001004 (rank : 153) | θ value | 3.0926 (rank : 59) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 741 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8NEP9, Q8NA46, Q96MP1 | Gene names | ZNF555 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 555. | |||||
|
ZN324_HUMAN
|
||||||
| NC score | -0.000452 (rank : 154) | θ value | 8.99809 (rank : 90) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 765 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O75467 | Gene names | ZNF324 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 324 (Zinc finger protein ZF5128). | |||||