Please be patient as the page loads
|
AHR_HUMAN
|
||||||
| SwissProt Accessions | P35869, Q13728, Q13803, Q13804 | Gene names | AHR | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
AHR_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P35869, Q13728, Q13803, Q13804 | Gene names | AHR | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
AHR_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.993935 (rank : 2) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P30561, Q8QZX6, Q8R4S3, Q99P79, Q9QVY1 | Gene names | Ahr | |||
|
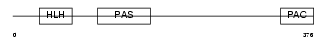
Domain Architecture |
|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
SIM2_HUMAN
|
||||||
| θ value | 3.17079e-29 (rank : 3) | NC score | 0.842738 (rank : 3) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q14190, O60766, Q15470, Q15471, Q15472, Q15473, Q16532 | Gene names | SIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2. | |||||
|
SIM2_MOUSE
|
||||||
| θ value | 4.57862e-28 (rank : 4) | NC score | 0.838702 (rank : 4) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q61079, O35391, Q61046, Q61904 | Gene names | Sim2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2 (SIM transcription factor) (mSIM). | |||||
|
SIM1_MOUSE
|
||||||
| θ value | 5.97985e-28 (rank : 5) | NC score | 0.834601 (rank : 6) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q61045, O70284, P70183 | Gene names | Sim1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1 (mSIM1). | |||||
|
HIF1A_MOUSE
|
||||||
| θ value | 7.80994e-28 (rank : 6) | NC score | 0.830453 (rank : 7) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q61221, O08741, O08993, Q61664, Q61665, Q8C681, Q8CC19, Q8CCB6, Q8R385, Q9CYA8 | Gene names | Hif1a | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein). | |||||
|
SIM1_HUMAN
|
||||||
| θ value | 1.33217e-27 (rank : 7) | NC score | 0.835341 (rank : 5) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P81133, Q5TDP7 | Gene names | SIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1. | |||||
|
HIF1A_HUMAN
|
||||||
| θ value | 8.63488e-27 (rank : 8) | NC score | 0.826852 (rank : 8) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q16665, Q96PT9, Q9UPB1 | Gene names | HIF1A | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein) (Member of PAS protein 1) (MOP1). | |||||
|
NPAS3_MOUSE
|
||||||
| θ value | 2.51237e-26 (rank : 9) | NC score | 0.817248 (rank : 9) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9QZQ0, Q9EQP4 | Gene names | Npas3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
NPAS3_HUMAN
|
||||||
| θ value | 2.77775e-25 (rank : 10) | NC score | 0.815377 (rank : 12) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8IXF0, Q86US6, Q86US7, Q8IXF2, Q9BY81, Q9H323, Q9Y4L8 | Gene names | NPAS3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
EPAS1_MOUSE
|
||||||
| θ value | 9.87957e-23 (rank : 11) | NC score | 0.815426 (rank : 11) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P97481, O08787, O55046 | Gene names | Epas1, Hif2a | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Hypoxia- inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (MHLF) (HIF-related factor) (HRF). | |||||
|
EPAS1_HUMAN
|
||||||
| θ value | 1.6852e-22 (rank : 12) | NC score | 0.816161 (rank : 10) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q99814, Q86VA2, Q99630 | Gene names | EPAS1, HIF2A | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Member of PAS protein 2) (MOP2) (Hypoxia-inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (HLF). | |||||
|
CLOCK_HUMAN
|
||||||
| θ value | 1.5773e-20 (rank : 13) | NC score | 0.753697 (rank : 15) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O15516, O14516, Q9UIT8 | Gene names | CLOCK, KIAA0334 | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (hCLOCK). | |||||
|
CLOCK_MOUSE
|
||||||
| θ value | 4.58923e-20 (rank : 14) | NC score | 0.750577 (rank : 16) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O08785 | Gene names | Clock | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (mCLOCK). | |||||
|
BMAL1_MOUSE
|
||||||
| θ value | 1.33526e-19 (rank : 15) | NC score | 0.740411 (rank : 18) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9WTL8, O88295, Q921S4, Q9R0U2, Q9WTL9 | Gene names | Arntl, Bmal1 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Arnt3). | |||||
|
NPAS1_HUMAN
|
||||||
| θ value | 2.27762e-19 (rank : 16) | NC score | 0.805712 (rank : 13) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q99742, Q99632, Q9BY83 | Gene names | NPAS1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1) (Member of PAS protein 5) (MOP5). | |||||
|
BMAL1_HUMAN
|
||||||
| θ value | 6.62687e-19 (rank : 17) | NC score | 0.740904 (rank : 17) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O00327, O00313, O00314, O00315, O00316, O00317, Q99631, Q99649 | Gene names | ARNTL, BMAL1, MOP3 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Member of PAS protein 3) (Basic-helix-loop- helix-PAS orphan MOP3) (bHLH-PAS protein JAP3). | |||||
|
NPAS2_MOUSE
|
||||||
| θ value | 4.74913e-17 (rank : 18) | NC score | 0.728155 (rank : 19) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P97460 | Gene names | Npas2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2). | |||||
|
NPAS1_MOUSE
|
||||||
| θ value | 1.058e-16 (rank : 19) | NC score | 0.801844 (rank : 14) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P97459 | Gene names | Npas1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1). | |||||
|
NPAS2_HUMAN
|
||||||
| θ value | 1.38178e-16 (rank : 20) | NC score | 0.721429 (rank : 20) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q99743, Q99629 | Gene names | NPAS2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2) (Member of PAS protein 4) (MOP4). | |||||
|
NPAS4_HUMAN
|
||||||
| θ value | 7.58209e-15 (rank : 21) | NC score | 0.660700 (rank : 23) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8IUM7, Q8N8S5, Q8N9Q9 | Gene names | NPAS4, NXF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF). | |||||
|
NPAS4_MOUSE
|
||||||
| θ value | 2.20605e-14 (rank : 22) | NC score | 0.649385 (rank : 25) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8BGD7, Q3V3U3 | Gene names | Npas4, Nxf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF) (Limbic-enhanced PAS protein) (LE-PAS). | |||||
|
NCOA3_HUMAN
|
||||||
| θ value | 3.52202e-12 (rank : 23) | NC score | 0.589777 (rank : 29) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9Y6Q9, Q5JYD9, Q5JYE0, Q9BR49, Q9UPC9, Q9UPG4, Q9UPG7 | Gene names | NCOA3, AIB1, RAC3, TRAM1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP-interacting protein) (pCIP). | |||||
|
NCOA1_HUMAN
|
||||||
| θ value | 7.84624e-12 (rank : 24) | NC score | 0.632270 (rank : 28) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q15788, O00150, O43792, O43793, Q13071, Q13420, Q6GVI5, Q7KYV3 | Gene names | NCOA1, SRC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (RIP160) (Protein Hin-2) (NY-REN-52 antigen). | |||||
|
NCOA1_MOUSE
|
||||||
| θ value | 7.84624e-12 (rank : 25) | NC score | 0.634284 (rank : 27) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P70365, P70366, Q61202, Q8CBI9 | Gene names | Ncoa1, Src1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (Nuclear receptor coactivator protein 1) (mNRC-1). | |||||
|
NCOA2_HUMAN
|
||||||
| θ value | 8.67504e-11 (rank : 26) | NC score | 0.576231 (rank : 30) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q15596 | Gene names | NCOA2, TIF2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 2 (NCoA-2) (Transcriptional intermediary factor 2). | |||||
|
ARNT2_HUMAN
|
||||||
| θ value | 2.52405e-10 (rank : 27) | NC score | 0.665594 (rank : 21) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9HBZ2, O15024 | Gene names | ARNT2, KIAA0307 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
NCOA2_MOUSE
|
||||||
| θ value | 4.30538e-10 (rank : 28) | NC score | 0.573492 (rank : 31) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q61026, O09001, P97759 | Gene names | Ncoa2, Grip1, Tif2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 2 (NCoA-2) (Transcriptional intermediary factor 2) (Glucocorticoid receptor-interacting protein 1) (GRIP-1). | |||||
|
ARNT2_MOUSE
|
||||||
| θ value | 5.62301e-10 (rank : 29) | NC score | 0.663710 (rank : 22) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q61324 | Gene names | Arnt2 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
ARNT_HUMAN
|
||||||
| θ value | 9.59137e-10 (rank : 30) | NC score | 0.660566 (rank : 24) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P27540 | Gene names | ARNT | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
ARNT_MOUSE
|
||||||
| θ value | 1.63604e-09 (rank : 31) | NC score | 0.648789 (rank : 26) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P53762, Q60661 | Gene names | Arnt | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
NCOA3_MOUSE
|
||||||
| θ value | 4.76016e-09 (rank : 32) | NC score | 0.551777 (rank : 32) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O09000, Q9CRD5 | Gene names | Ncoa3, Aib1, Pcip, Rac3, Tram1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein homolog) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP- interacting protein) (p/CIP) (pCIP). | |||||
|
GCC2_MOUSE
|
||||||
| θ value | 0.47712 (rank : 33) | NC score | 0.002886 (rank : 64) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 1619 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CHG3, Q8BR44, Q8R2Q5, Q9CT45 | Gene names | Gcc2, Kiaa0336 | |||
|
Domain Architecture |

|
|||||
| Description | GRIP and coiled-coil domain-containing protein 2 (Golgi coiled coil protein GCC185). | |||||
|
HES5_MOUSE
|
||||||
| θ value | 1.06291 (rank : 34) | NC score | 0.105869 (rank : 39) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P70120, Q8BVI1 | Gene names | Hes5, Hes-5 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-5 (Hairy and enhancer of split 5). | |||||
|
PTN22_HUMAN
|
||||||
| θ value | 1.38821 (rank : 35) | NC score | 0.007401 (rank : 63) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y2R2, O95063, O95064 | Gene names | PTPN22, PTPN8 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 22 (EC 3.1.3.48) (Hematopoietic cell protein-tyrosine phosphatase 70Z-PEP) (Lymphoid phosphatase) (LyP). | |||||
|
HES3_MOUSE
|
||||||
| θ value | 1.81305 (rank : 36) | NC score | 0.081527 (rank : 43) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q61657, O09083, O09150 | Gene names | Hes3, Hes-3 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor HES-3 (Hairy and enhancer of split 3). | |||||
|
R3HD2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 37) | NC score | 0.016127 (rank : 60) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80TM6, Q80YB1, Q8BLS5, Q9CW50 | Gene names | R3hdm2, Kiaa1002 | |||
|
Domain Architecture |

|
|||||
| Description | R3H domain-containing protein 2. | |||||
|
ZBT40_HUMAN
|
||||||
| θ value | 4.03905 (rank : 38) | NC score | -0.002082 (rank : 67) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 949 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NUA8, O75066, Q5TFU5, Q8N1R1 | Gene names | ZBTB40, KIAA0478 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger and BTB domain-containing protein 40. | |||||
|
MITF_HUMAN
|
||||||
| θ value | 5.27518 (rank : 39) | NC score | 0.080537 (rank : 44) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O75030, Q14841, Q9P2V0, Q9P2V1, Q9P2V2, Q9P2Y8 | Gene names | MITF | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
HES7_HUMAN
|
||||||
| θ value | 6.88961 (rank : 40) | NC score | 0.054063 (rank : 56) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9BYE0 | Gene names | HES7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor HES-7 (Hairy and enhancer of split 7) (bHLH factor Hes7). | |||||
|
HES7_MOUSE
|
||||||
| θ value | 6.88961 (rank : 41) | NC score | 0.052024 (rank : 58) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8BKT2, Q99JA6 | Gene names | Hes7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor HES-7 (Hairy and enhancer of split 7) (bHLH factor Hes7). | |||||
|
ITSN2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 42) | NC score | -0.000676 (rank : 66) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 1371 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Z0R6, Q9Z0R5 | Gene names | Itsn2, Ese2, Sh3d1B | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-2 (SH3 domain-containing protein 1B) (EH and SH3 domains protein 2) (EH domain and SH3 domain regulator of endocytosis 2). | |||||
|
MITF_MOUSE
|
||||||
| θ value | 8.99809 (rank : 43) | NC score | 0.080130 (rank : 45) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q08874, O08885, O88203, Q08843, Q60781, Q60782, Q9JIJ0, Q9JIJ1, Q9JIJ2, Q9JIJ3, Q9JIJ4, Q9JIJ5, Q9JIJ6, Q9JKX9 | Gene names | Mitf, Bw, Mi, Vit | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
NAF1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 44) | NC score | 0.001728 (rank : 65) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 852 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9WUU8, Q922A9, Q922F7, Q9EPP8, Q9R0X3 | Gene names | Tnip1, Abin, Naf1 | |||
|
Domain Architecture |

|
|||||
| Description | Nef-associated factor 1 (Naf1) (TNFAIP3-interacting protein 1) (A20- binding inhibitor of NF-kappa-B activation) (ABIN) (Virion-associated nuclear shuttling protein) (VAN) (mVAN). | |||||
|
OXA1L_MOUSE
|
||||||
| θ value | 8.99809 (rank : 45) | NC score | 0.009738 (rank : 61) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BGA9, Q8BK01, Q8R091, Q9D8X7 | Gene names | Oxa1l | |||
|
Domain Architecture |

|
|||||
| Description | Inner membrane protein OXA1L, mitochondrial precursor (Oxidase assembly 1-like protein) (OXA1-like protein). | |||||
|
RPA1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 46) | NC score | 0.022382 (rank : 59) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O95602, Q9UEH0, Q9UFT9 | Gene names | POLR1A | |||
|
Domain Architecture |

|
|||||
| Description | DNA-directed RNA polymerase I largest subunit (EC 2.7.7.6) (RNA polymerase I 194 kDa subunit) (RPA194). | |||||
|
TJAP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 47) | NC score | 0.007477 (rank : 62) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 295 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9DCD5, Q8CFL7, Q8R5I2 | Gene names | Tjap1, Pilt, Tjp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tight junction-associated protein 1 (Tight junction protein 4) (Protein incorporated later into tight junctions). | |||||
|
HEY1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 48) | NC score | 0.079452 (rank : 46) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9Y5J3, Q5TZS3, Q9NYP4 | Gene names | HEY1, CHF2, HESR1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1) (Cardiovascular helix-loop-helix factor 2) (HES-related repressor protein 2) (HERP2). | |||||
|
HEY1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 49) | NC score | 0.082927 (rank : 42) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9WV93 | Gene names | Hey1, Hesr1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1). | |||||
|
HEY2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 50) | NC score | 0.086502 (rank : 40) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9UBP5 | Gene names | HEY2, CHF1, GRL, HERP, HERP1, HRT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Cardiovascular helix-loop-helix factor 1) (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES- related repressor protein 2) (Protein gridlock homolog). | |||||
|
HEY2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 51) | NC score | 0.084388 (rank : 41) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9QUS4, Q3TZ99, Q8CD44 | Gene names | Hey2, Chf1, Herp, Herp1, Hesr2, Hrt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES-related repressor protein 2). | |||||
|
MLX_HUMAN
|
||||||
| θ value | θ > 10 (rank : 52) | NC score | 0.067215 (rank : 51) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UH92, Q53XM6, Q96FL2, Q9H2V0, Q9H2V1, Q9H2V2, Q9NXN3 | Gene names | MLX, TCFL4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
MLX_MOUSE
|
||||||
| θ value | θ > 10 (rank : 53) | NC score | 0.066840 (rank : 52) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O08609, Q9EQJ7, Q9EQJ8 | Gene names | Mlx, Tcfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
PER1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 54) | NC score | 0.168498 (rank : 36) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O15534 | Gene names | PER1, KIAA0482, PER, RIGUI | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (hPER). | |||||
|
PER1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 55) | NC score | 0.164582 (rank : 37) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O35973 | Gene names | Per1, Per, Rigui | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (mPER) (M-Rigui). | |||||
|
PER2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 56) | NC score | 0.171213 (rank : 34) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O15055 | Gene names | PER2, KIAA0347 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2. | |||||
|
PER2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 57) | NC score | 0.156428 (rank : 38) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O54943, O54954 | Gene names | Per2 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2 (mPER2). | |||||
|
PER3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 58) | NC score | 0.169382 (rank : 35) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P56645, Q5H8X4, Q969K6, Q96S77, Q96S78, Q9C0J3, Q9NSP9, Q9UGU8 | Gene names | PER3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (hPER3). | |||||
|
PER3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 59) | NC score | 0.187356 (rank : 33) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O70361 | Gene names | Per3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (mPER3). | |||||
|
TCFL5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 60) | NC score | 0.068199 (rank : 48) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UL49, O94771, Q9BYW0 | Gene names | TCFL5, CHA, E2BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor-like 5 protein (Cha transcription factor) (HPV-16 E2-binding protein 1) (E2BP-1). | |||||
|
TFE3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 61) | NC score | 0.073574 (rank : 47) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P19532, Q92757, Q92758, Q99964 | Gene names | TFE3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E3. | |||||
|
TFEB_HUMAN
|
||||||
| θ value | θ > 10 (rank : 62) | NC score | 0.067353 (rank : 49) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P19484, Q709B3, Q7Z6P9, Q9BRJ5, Q9UJD8 | Gene names | TFEB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
TFEB_MOUSE
|
||||||
| θ value | θ > 10 (rank : 63) | NC score | 0.067217 (rank : 50) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9R210 | Gene names | Tfeb, Tcfeb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
USF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 64) | NC score | 0.059898 (rank : 53) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P22415 | Gene names | USF1, USF | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 65) | NC score | 0.059871 (rank : 54) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q61069 | Gene names | Usf1, Usf | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 66) | NC score | 0.057900 (rank : 55) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q15853, O00671, O00709, Q05750, Q07952, Q15851, Q15852 | Gene names | USF2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (FOS- interacting protein) (FIP) (Major late transcription factor 2). | |||||
|
USF2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 67) | NC score | 0.053961 (rank : 57) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q64705, Q3UIL2, Q8VE59 | Gene names | Usf2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (Major late transcription factor 2). | |||||
|
AHR_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P35869, Q13728, Q13803, Q13804 | Gene names | AHR | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
AHR_MOUSE
|
||||||
| NC score | 0.993935 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P30561, Q8QZX6, Q8R4S3, Q99P79, Q9QVY1 | Gene names | Ahr | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
SIM2_HUMAN
|
||||||
| NC score | 0.842738 (rank : 3) | θ value | 3.17079e-29 (rank : 3) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q14190, O60766, Q15470, Q15471, Q15472, Q15473, Q16532 | Gene names | SIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2. | |||||
|
SIM2_MOUSE
|
||||||
| NC score | 0.838702 (rank : 4) | θ value | 4.57862e-28 (rank : 4) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q61079, O35391, Q61046, Q61904 | Gene names | Sim2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2 (SIM transcription factor) (mSIM). | |||||
|
SIM1_HUMAN
|
||||||
| NC score | 0.835341 (rank : 5) | θ value | 1.33217e-27 (rank : 7) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P81133, Q5TDP7 | Gene names | SIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1. | |||||
|
SIM1_MOUSE
|
||||||
| NC score | 0.834601 (rank : 6) | θ value | 5.97985e-28 (rank : 5) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q61045, O70284, P70183 | Gene names | Sim1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1 (mSIM1). | |||||
|
HIF1A_MOUSE
|
||||||
| NC score | 0.830453 (rank : 7) | θ value | 7.80994e-28 (rank : 6) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q61221, O08741, O08993, Q61664, Q61665, Q8C681, Q8CC19, Q8CCB6, Q8R385, Q9CYA8 | Gene names | Hif1a | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein). | |||||
|
HIF1A_HUMAN
|
||||||
| NC score | 0.826852 (rank : 8) | θ value | 8.63488e-27 (rank : 8) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q16665, Q96PT9, Q9UPB1 | Gene names | HIF1A | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein) (Member of PAS protein 1) (MOP1). | |||||
|
NPAS3_MOUSE
|
||||||
| NC score | 0.817248 (rank : 9) | θ value | 2.51237e-26 (rank : 9) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9QZQ0, Q9EQP4 | Gene names | Npas3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
EPAS1_HUMAN
|
||||||
| NC score | 0.816161 (rank : 10) | θ value | 1.6852e-22 (rank : 12) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q99814, Q86VA2, Q99630 | Gene names | EPAS1, HIF2A | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Member of PAS protein 2) (MOP2) (Hypoxia-inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (HLF). | |||||
|
EPAS1_MOUSE
|
||||||
| NC score | 0.815426 (rank : 11) | θ value | 9.87957e-23 (rank : 11) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P97481, O08787, O55046 | Gene names | Epas1, Hif2a | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Hypoxia- inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (MHLF) (HIF-related factor) (HRF). | |||||
|
NPAS3_HUMAN
|
||||||
| NC score | 0.815377 (rank : 12) | θ value | 2.77775e-25 (rank : 10) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8IXF0, Q86US6, Q86US7, Q8IXF2, Q9BY81, Q9H323, Q9Y4L8 | Gene names | NPAS3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
NPAS1_HUMAN
|
||||||
| NC score | 0.805712 (rank : 13) | θ value | 2.27762e-19 (rank : 16) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q99742, Q99632, Q9BY83 | Gene names | NPAS1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1) (Member of PAS protein 5) (MOP5). | |||||
|
NPAS1_MOUSE
|
||||||
| NC score | 0.801844 (rank : 14) | θ value | 1.058e-16 (rank : 19) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P97459 | Gene names | Npas1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1). | |||||
|
CLOCK_HUMAN
|
||||||
| NC score | 0.753697 (rank : 15) | θ value | 1.5773e-20 (rank : 13) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O15516, O14516, Q9UIT8 | Gene names | CLOCK, KIAA0334 | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (hCLOCK). | |||||
|
CLOCK_MOUSE
|
||||||
| NC score | 0.750577 (rank : 16) | θ value | 4.58923e-20 (rank : 14) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O08785 | Gene names | Clock | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (mCLOCK). | |||||
|
BMAL1_HUMAN
|
||||||
| NC score | 0.740904 (rank : 17) | θ value | 6.62687e-19 (rank : 17) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O00327, O00313, O00314, O00315, O00316, O00317, Q99631, Q99649 | Gene names | ARNTL, BMAL1, MOP3 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Member of PAS protein 3) (Basic-helix-loop- helix-PAS orphan MOP3) (bHLH-PAS protein JAP3). | |||||
|
BMAL1_MOUSE
|
||||||
| NC score | 0.740411 (rank : 18) | θ value | 1.33526e-19 (rank : 15) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9WTL8, O88295, Q921S4, Q9R0U2, Q9WTL9 | Gene names | Arntl, Bmal1 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Arnt3). | |||||
|
NPAS2_MOUSE
|
||||||
| NC score | 0.728155 (rank : 19) | θ value | 4.74913e-17 (rank : 18) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P97460 | Gene names | Npas2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2). | |||||
|
NPAS2_HUMAN
|
||||||
| NC score | 0.721429 (rank : 20) | θ value | 1.38178e-16 (rank : 20) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q99743, Q99629 | Gene names | NPAS2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2) (Member of PAS protein 4) (MOP4). | |||||
|
ARNT2_HUMAN
|
||||||
| NC score | 0.665594 (rank : 21) | θ value | 2.52405e-10 (rank : 27) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9HBZ2, O15024 | Gene names | ARNT2, KIAA0307 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
ARNT2_MOUSE
|
||||||
| NC score | 0.663710 (rank : 22) | θ value | 5.62301e-10 (rank : 29) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q61324 | Gene names | Arnt2 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
NPAS4_HUMAN
|
||||||
| NC score | 0.660700 (rank : 23) | θ value | 7.58209e-15 (rank : 21) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8IUM7, Q8N8S5, Q8N9Q9 | Gene names | NPAS4, NXF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF). | |||||
|
ARNT_HUMAN
|
||||||
| NC score | 0.660566 (rank : 24) | θ value | 9.59137e-10 (rank : 30) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P27540 | Gene names | ARNT | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
NPAS4_MOUSE
|
||||||
| NC score | 0.649385 (rank : 25) | θ value | 2.20605e-14 (rank : 22) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8BGD7, Q3V3U3 | Gene names | Npas4, Nxf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuronal PAS domain-containing protein 4 (Neuronal PAS4) (HLH-PAS transcription factor NXF) (Limbic-enhanced PAS protein) (LE-PAS). | |||||
|
ARNT_MOUSE
|
||||||
| NC score | 0.648789 (rank : 26) | θ value | 1.63604e-09 (rank : 31) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P53762, Q60661 | Gene names | Arnt | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
NCOA1_MOUSE
|
||||||
| NC score | 0.634284 (rank : 27) | θ value | 7.84624e-12 (rank : 25) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P70365, P70366, Q61202, Q8CBI9 | Gene names | Ncoa1, Src1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (Nuclear receptor coactivator protein 1) (mNRC-1). | |||||
|
NCOA1_HUMAN
|
||||||
| NC score | 0.632270 (rank : 28) | θ value | 7.84624e-12 (rank : 24) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q15788, O00150, O43792, O43793, Q13071, Q13420, Q6GVI5, Q7KYV3 | Gene names | NCOA1, SRC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (RIP160) (Protein Hin-2) (NY-REN-52 antigen). | |||||
|
NCOA3_HUMAN
|
||||||
| NC score | 0.589777 (rank : 29) | θ value | 3.52202e-12 (rank : 23) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9Y6Q9, Q5JYD9, Q5JYE0, Q9BR49, Q9UPC9, Q9UPG4, Q9UPG7 | Gene names | NCOA3, AIB1, RAC3, TRAM1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP-interacting protein) (pCIP). | |||||
|
NCOA2_HUMAN
|
||||||
| NC score | 0.576231 (rank : 30) | θ value | 8.67504e-11 (rank : 26) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q15596 | Gene names | NCOA2, TIF2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 2 (NCoA-2) (Transcriptional intermediary factor 2). | |||||
|
NCOA2_MOUSE
|
||||||
| NC score | 0.573492 (rank : 31) | θ value | 4.30538e-10 (rank : 28) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q61026, O09001, P97759 | Gene names | Ncoa2, Grip1, Tif2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 2 (NCoA-2) (Transcriptional intermediary factor 2) (Glucocorticoid receptor-interacting protein 1) (GRIP-1). | |||||
|
NCOA3_MOUSE
|
||||||
| NC score | 0.551777 (rank : 32) | θ value | 4.76016e-09 (rank : 32) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O09000, Q9CRD5 | Gene names | Ncoa3, Aib1, Pcip, Rac3, Tram1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein homolog) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP- interacting protein) (p/CIP) (pCIP). | |||||
|
PER3_MOUSE
|
||||||
| NC score | 0.187356 (rank : 33) | θ value | θ > 10 (rank : 59) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O70361 | Gene names | Per3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (mPER3). | |||||
|
PER2_HUMAN
|
||||||
| NC score | 0.171213 (rank : 34) | θ value | θ > 10 (rank : 56) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O15055 | Gene names | PER2, KIAA0347 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2. | |||||
|
PER3_HUMAN
|
||||||
| NC score | 0.169382 (rank : 35) | θ value | θ > 10 (rank : 58) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P56645, Q5H8X4, Q969K6, Q96S77, Q96S78, Q9C0J3, Q9NSP9, Q9UGU8 | Gene names | PER3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (hPER3). | |||||
|
PER1_HUMAN
|
||||||
| NC score | 0.168498 (rank : 36) | θ value | θ > 10 (rank : 54) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O15534 | Gene names | PER1, KIAA0482, PER, RIGUI | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (hPER). | |||||
|
PER1_MOUSE
|
||||||
| NC score | 0.164582 (rank : 37) | θ value | θ > 10 (rank : 55) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O35973 | Gene names | Per1, Per, Rigui | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (mPER) (M-Rigui). | |||||
|
PER2_MOUSE
|
||||||
| NC score | 0.156428 (rank : 38) | θ value | θ > 10 (rank : 57) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O54943, O54954 | Gene names | Per2 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2 (mPER2). | |||||
|
HES5_MOUSE
|
||||||
| NC score | 0.105869 (rank : 39) | θ value | 1.06291 (rank : 34) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P70120, Q8BVI1 | Gene names | Hes5, Hes-5 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-5 (Hairy and enhancer of split 5). | |||||
|
HEY2_HUMAN
|
||||||
| NC score | 0.086502 (rank : 40) | θ value | θ > 10 (rank : 50) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9UBP5 | Gene names | HEY2, CHF1, GRL, HERP, HERP1, HRT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Cardiovascular helix-loop-helix factor 1) (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES- related repressor protein 2) (Protein gridlock homolog). | |||||
|
HEY2_MOUSE
|
||||||
| NC score | 0.084388 (rank : 41) | θ value | θ > 10 (rank : 51) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9QUS4, Q3TZ99, Q8CD44 | Gene names | Hey2, Chf1, Herp, Herp1, Hesr2, Hrt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES-related repressor protein 2). | |||||
|
HEY1_MOUSE
|
||||||
| NC score | 0.082927 (rank : 42) | θ value | θ > 10 (rank : 49) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9WV93 | Gene names | Hey1, Hesr1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1). | |||||
|
HES3_MOUSE
|
||||||
| NC score | 0.081527 (rank : 43) | θ value | 1.81305 (rank : 36) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q61657, O09083, O09150 | Gene names | Hes3, Hes-3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-3 (Hairy and enhancer of split 3). | |||||
|
MITF_HUMAN
|
||||||
| NC score | 0.080537 (rank : 44) | θ value | 5.27518 (rank : 39) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O75030, Q14841, Q9P2V0, Q9P2V1, Q9P2V2, Q9P2Y8 | Gene names | MITF | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
MITF_MOUSE
|
||||||
| NC score | 0.080130 (rank : 45) | θ value | 8.99809 (rank : 43) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q08874, O08885, O88203, Q08843, Q60781, Q60782, Q9JIJ0, Q9JIJ1, Q9JIJ2, Q9JIJ3, Q9JIJ4, Q9JIJ5, Q9JIJ6, Q9JKX9 | Gene names | Mitf, Bw, Mi, Vit | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
HEY1_HUMAN
|
||||||
| NC score | 0.079452 (rank : 46) | θ value | θ > 10 (rank : 48) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9Y5J3, Q5TZS3, Q9NYP4 | Gene names | HEY1, CHF2, HESR1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1) (Cardiovascular helix-loop-helix factor 2) (HES-related repressor protein 2) (HERP2). | |||||
|
TFE3_HUMAN
|
||||||
| NC score | 0.073574 (rank : 47) | θ value | θ > 10 (rank : 61) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P19532, Q92757, Q92758, Q99964 | Gene names | TFE3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E3. | |||||
|
TCFL5_HUMAN
|
||||||
| NC score | 0.068199 (rank : 48) | θ value | θ > 10 (rank : 60) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UL49, O94771, Q9BYW0 | Gene names | TCFL5, CHA, E2BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor-like 5 protein (Cha transcription factor) (HPV-16 E2-binding protein 1) (E2BP-1). | |||||
|
TFEB_HUMAN
|
||||||
| NC score | 0.067353 (rank : 49) | θ value | θ > 10 (rank : 62) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P19484, Q709B3, Q7Z6P9, Q9BRJ5, Q9UJD8 | Gene names | TFEB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
TFEB_MOUSE
|
||||||
| NC score | 0.067217 (rank : 50) | θ value | θ > 10 (rank : 63) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9R210 | Gene names | Tfeb, Tcfeb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
MLX_HUMAN
|
||||||
| NC score | 0.067215 (rank : 51) | θ value | θ > 10 (rank : 52) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UH92, Q53XM6, Q96FL2, Q9H2V0, Q9H2V1, Q9H2V2, Q9NXN3 | Gene names | MLX, TCFL4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
MLX_MOUSE
|
||||||
| NC score | 0.066840 (rank : 52) | θ value | θ > 10 (rank : 53) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O08609, Q9EQJ7, Q9EQJ8 | Gene names | Mlx, Tcfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
USF1_HUMAN
|
||||||
| NC score | 0.059898 (rank : 53) | θ value | θ > 10 (rank : 64) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P22415 | Gene names | USF1, USF | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF1_MOUSE
|
||||||
| NC score | 0.059871 (rank : 54) | θ value | θ > 10 (rank : 65) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q61069 | Gene names | Usf1, Usf | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF2_HUMAN
|
||||||
| NC score | 0.057900 (rank : 55) | θ value | θ > 10 (rank : 66) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q15853, O00671, O00709, Q05750, Q07952, Q15851, Q15852 | Gene names | USF2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (FOS- interacting protein) (FIP) (Major late transcription factor 2). | |||||
|
HES7_HUMAN
|
||||||
| NC score | 0.054063 (rank : 56) | θ value | 6.88961 (rank : 40) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9BYE0 | Gene names | HES7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor HES-7 (Hairy and enhancer of split 7) (bHLH factor Hes7). | |||||
|
USF2_MOUSE
|
||||||
| NC score | 0.053961 (rank : 57) | θ value | θ > 10 (rank : 67) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q64705, Q3UIL2, Q8VE59 | Gene names | Usf2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (Major late transcription factor 2). | |||||
|
HES7_MOUSE
|
||||||
| NC score | 0.052024 (rank : 58) | θ value | 6.88961 (rank : 41) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8BKT2, Q99JA6 | Gene names | Hes7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor HES-7 (Hairy and enhancer of split 7) (bHLH factor Hes7). | |||||
|
RPA1_HUMAN
|
||||||
| NC score | 0.022382 (rank : 59) | θ value | 8.99809 (rank : 46) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O95602, Q9UEH0, Q9UFT9 | Gene names | POLR1A | |||
|
Domain Architecture |

|
|||||
| Description | DNA-directed RNA polymerase I largest subunit (EC 2.7.7.6) (RNA polymerase I 194 kDa subunit) (RPA194). | |||||
|
R3HD2_MOUSE
|
||||||
| NC score | 0.016127 (rank : 60) | θ value | 4.03905 (rank : 37) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80TM6, Q80YB1, Q8BLS5, Q9CW50 | Gene names | R3hdm2, Kiaa1002 | |||
|
Domain Architecture |

|
|||||
| Description | R3H domain-containing protein 2. | |||||
|
OXA1L_MOUSE
|
||||||
| NC score | 0.009738 (rank : 61) | θ value | 8.99809 (rank : 45) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BGA9, Q8BK01, Q8R091, Q9D8X7 | Gene names | Oxa1l | |||
|
Domain Architecture |

|
|||||
| Description | Inner membrane protein OXA1L, mitochondrial precursor (Oxidase assembly 1-like protein) (OXA1-like protein). | |||||
|
TJAP1_MOUSE
|
||||||
| NC score | 0.007477 (rank : 62) | θ value | 8.99809 (rank : 47) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 295 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9DCD5, Q8CFL7, Q8R5I2 | Gene names | Tjap1, Pilt, Tjp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tight junction-associated protein 1 (Tight junction protein 4) (Protein incorporated later into tight junctions). | |||||
|
PTN22_HUMAN
|
||||||
| NC score | 0.007401 (rank : 63) | θ value | 1.38821 (rank : 35) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y2R2, O95063, O95064 | Gene names | PTPN22, PTPN8 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 22 (EC 3.1.3.48) (Hematopoietic cell protein-tyrosine phosphatase 70Z-PEP) (Lymphoid phosphatase) (LyP). | |||||
|
GCC2_MOUSE
|
||||||
| NC score | 0.002886 (rank : 64) | θ value | 0.47712 (rank : 33) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 1619 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CHG3, Q8BR44, Q8R2Q5, Q9CT45 | Gene names | Gcc2, Kiaa0336 | |||
|
Domain Architecture |

|
|||||
| Description | GRIP and coiled-coil domain-containing protein 2 (Golgi coiled coil protein GCC185). | |||||
|
NAF1_MOUSE
|
||||||
| NC score | 0.001728 (rank : 65) | θ value | 8.99809 (rank : 44) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 852 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9WUU8, Q922A9, Q922F7, Q9EPP8, Q9R0X3 | Gene names | Tnip1, Abin, Naf1 | |||
|
Domain Architecture |

|
|||||
| Description | Nef-associated factor 1 (Naf1) (TNFAIP3-interacting protein 1) (A20- binding inhibitor of NF-kappa-B activation) (ABIN) (Virion-associated nuclear shuttling protein) (VAN) (mVAN). | |||||
|
ITSN2_MOUSE
|
||||||
| NC score | -0.000676 (rank : 66) | θ value | 6.88961 (rank : 42) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 1371 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Z0R6, Q9Z0R5 | Gene names | Itsn2, Ese2, Sh3d1B | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-2 (SH3 domain-containing protein 1B) (EH and SH3 domains protein 2) (EH domain and SH3 domain regulator of endocytosis 2). | |||||
|
ZBT40_HUMAN
|
||||||
| NC score | -0.002082 (rank : 67) | θ value | 4.03905 (rank : 38) | |||
| Query Neighborhood Hits | 47 | Target Neighborhood Hits | 949 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NUA8, O75066, Q5TFU5, Q8N1R1 | Gene names | ZBTB40, KIAA0478 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 40. | |||||