Please be patient as the page loads
|
HES5_MOUSE
|
||||||
| SwissProt Accessions | P70120, Q8BVI1 | Gene names | Hes5, Hes-5 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-5 (Hairy and enhancer of split 5). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
HES5_MOUSE
|
||||||
| θ value | 6.11685e-65 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P70120, Q8BVI1 | Gene names | Hes5, Hes-5 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-5 (Hairy and enhancer of split 5). | |||||
|
HES1_HUMAN
|
||||||
| θ value | 1.74796e-11 (rank : 2) | NC score | 0.821421 (rank : 6) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q14469 | Gene names | HES1, HL, HRY | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor HES-1 (Hairy and enhancer of split 1) (Hairy- like) (HHL) (Hairy homolog). | |||||
|
HES1_MOUSE
|
||||||
| θ value | 1.74796e-11 (rank : 3) | NC score | 0.821829 (rank : 5) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P35428 | Gene names | Hes1, Hes-1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-1 (Hairy and enhancer of split 1). | |||||
|
HES7_MOUSE
|
||||||
| θ value | 8.67504e-11 (rank : 4) | NC score | 0.772129 (rank : 8) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8BKT2, Q99JA6 | Gene names | Hes7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor HES-7 (Hairy and enhancer of split 7) (bHLH factor Hes7). | |||||
|
HES7_HUMAN
|
||||||
| θ value | 3.29651e-10 (rank : 5) | NC score | 0.772983 (rank : 7) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9BYE0 | Gene names | HES7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor HES-7 (Hairy and enhancer of split 7) (bHLH factor Hes7). | |||||
|
HES4_HUMAN
|
||||||
| θ value | 4.30538e-10 (rank : 6) | NC score | 0.849466 (rank : 2) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9HCC6 | Gene names | HES4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-4 (Hairy and enhancer of split 4) (bHLH factor Hes4). | |||||
|
HEY1_HUMAN
|
||||||
| θ value | 1.25267e-09 (rank : 7) | NC score | 0.750373 (rank : 10) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9Y5J3, Q5TZS3, Q9NYP4 | Gene names | HEY1, CHF2, HESR1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1) (Cardiovascular helix-loop-helix factor 2) (HES-related repressor protein 2) (HERP2). | |||||
|
HEY1_MOUSE
|
||||||
| θ value | 1.25267e-09 (rank : 8) | NC score | 0.753503 (rank : 9) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9WV93 | Gene names | Hey1, Hesr1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1). | |||||
|
HES2_HUMAN
|
||||||
| θ value | 2.79066e-09 (rank : 9) | NC score | 0.826413 (rank : 4) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9Y543, Q96EN4, Q9Y542 | Gene names | HES2 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor HES-2 (Hairy and enhancer of split 2). | |||||
|
HES2_MOUSE
|
||||||
| θ value | 6.21693e-09 (rank : 10) | NC score | 0.831350 (rank : 3) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O54792 | Gene names | Hes2 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor HES-2 (Hairy and enhancer of split 2). | |||||
|
HEY2_HUMAN
|
||||||
| θ value | 2.36244e-08 (rank : 11) | NC score | 0.745967 (rank : 12) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9UBP5 | Gene names | HEY2, CHF1, GRL, HERP, HERP1, HRT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Cardiovascular helix-loop-helix factor 1) (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES- related repressor protein 2) (Protein gridlock homolog). | |||||
|
HEY2_MOUSE
|
||||||
| θ value | 5.26297e-08 (rank : 12) | NC score | 0.744202 (rank : 13) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9QUS4, Q3TZ99, Q8CD44 | Gene names | Hey2, Chf1, Herp, Herp1, Hesr2, Hrt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES-related repressor protein 2). | |||||
|
HES3_MOUSE
|
||||||
| θ value | 8.97725e-08 (rank : 13) | NC score | 0.749049 (rank : 11) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q61657, O09083, O09150 | Gene names | Hes3, Hes-3 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor HES-3 (Hairy and enhancer of split 3). | |||||
|
HES6_HUMAN
|
||||||
| θ value | 3.77169e-06 (rank : 14) | NC score | 0.722615 (rank : 14) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96HZ4, Q8N2J2, Q96T93, Q9P2S3 | Gene names | HES6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription cofactor HES-6 (Hairy and enhancer of split 6) (C- HAIRY1). | |||||
|
HES6_MOUSE
|
||||||
| θ value | 3.77169e-06 (rank : 15) | NC score | 0.714157 (rank : 15) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9JHE6, Q3U2D7 | Gene names | Hes6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription cofactor HES-6 (Hairy and enhancer of split 6). | |||||
|
BHLH3_HUMAN
|
||||||
| θ value | 8.40245e-06 (rank : 16) | NC score | 0.575593 (rank : 16) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9C0J9 | Gene names | BHLHB3, DEC2, SHARP1 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (hDEC2) (Enhancer-of-split and hairy-related protein 1) (SHARP-1). | |||||
|
BHLH3_MOUSE
|
||||||
| θ value | 1.09739e-05 (rank : 17) | NC score | 0.574536 (rank : 17) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q99PV5 | Gene names | Bhlhb3, Dec2 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (mDEC2). | |||||
|
BHLH2_MOUSE
|
||||||
| θ value | 0.00035302 (rank : 18) | NC score | 0.548708 (rank : 19) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O35185, P97289 | Gene names | Bhlhb2, Clast5, Stra13 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 2 (bHLHB2) (Stimulated with retinoic acid 13) (E47 interaction protein 1) (eipl). | |||||
|
BHLH2_HUMAN
|
||||||
| θ value | 0.000461057 (rank : 19) | NC score | 0.560765 (rank : 18) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O14503, Q96TD3 | Gene names | BHLHB2, DEC1, SHARP2, STRA13 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 2 (bHLHB2) (Differentially expressed in chondrocytes protein 1) (DEC1) (Enhancer-of-split and hairy-related protein 2) (SHARP-2) (Stimulated with retinoic acid 13). | |||||
|
AHR_MOUSE
|
||||||
| θ value | 0.21417 (rank : 20) | NC score | 0.107679 (rank : 31) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P30561, Q8QZX6, Q8R4S3, Q99P79, Q9QVY1 | Gene names | Ahr | |||
|
Domain Architecture |
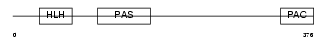
|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
BMAL1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 21) | NC score | 0.169443 (rank : 21) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O00327, O00313, O00314, O00315, O00316, O00317, Q99631, Q99649 | Gene names | ARNTL, BMAL1, MOP3 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Member of PAS protein 3) (Basic-helix-loop- helix-PAS orphan MOP3) (bHLH-PAS protein JAP3). | |||||
|
BMAL1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 22) | NC score | 0.172604 (rank : 20) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9WTL8, O88295, Q921S4, Q9R0U2, Q9WTL9 | Gene names | Arntl, Bmal1 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Arnt3). | |||||
|
CLOCK_HUMAN
|
||||||
| θ value | 0.62314 (rank : 23) | NC score | 0.121847 (rank : 27) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O15516, O14516, Q9UIT8 | Gene names | CLOCK, KIAA0334 | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (hCLOCK). | |||||
|
AHR_HUMAN
|
||||||
| θ value | 1.06291 (rank : 24) | NC score | 0.105869 (rank : 35) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P35869, Q13728, Q13803, Q13804 | Gene names | AHR | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
CLOCK_MOUSE
|
||||||
| θ value | 1.06291 (rank : 25) | NC score | 0.117486 (rank : 29) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O08785 | Gene names | Clock | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (mCLOCK). | |||||
|
USF2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 26) | NC score | 0.129244 (rank : 24) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q64705, Q3UIL2, Q8VE59 | Gene names | Usf2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (Major late transcription factor 2). | |||||
|
USF1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 27) | NC score | 0.140174 (rank : 22) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P22415 | Gene names | USF1, USF | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 28) | NC score | 0.140169 (rank : 23) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q61069 | Gene names | Usf1, Usf | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
EPAS1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 29) | NC score | 0.065676 (rank : 53) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P97481, O08787, O55046 | Gene names | Epas1, Hif2a | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Hypoxia- inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (MHLF) (HIF-related factor) (HRF). | |||||
|
SRBP2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 30) | NC score | 0.114189 (rank : 30) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q12772, Q9UH04 | Gene names | SREBF2, SREBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 2 (SREBP-2) (Sterol regulatory element-binding transcription factor 2). | |||||
|
MAP1A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 31) | NC score | 0.008666 (rank : 62) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P78559, O95643, Q12973, Q15882, Q9UJT4 | Gene names | MAP1A | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 1A (MAP 1A) (Proliferation-related protein p80) [Contains: MAP1 light chain LC2]. | |||||
|
NPAS2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 32) | NC score | 0.106422 (rank : 33) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q99743, Q99629 | Gene names | NPAS2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2) (Member of PAS protein 4) (MOP4). | |||||
|
NPAS2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 33) | NC score | 0.106550 (rank : 32) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P97460 | Gene names | Npas2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2). | |||||
|
SIM2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 34) | NC score | 0.073616 (rank : 46) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q14190, O60766, Q15470, Q15471, Q15472, Q15473, Q16532 | Gene names | SIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2. | |||||
|
LYST_HUMAN
|
||||||
| θ value | 8.99809 (rank : 35) | NC score | 0.008449 (rank : 63) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 324 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99698, O43274, Q5T2U9, Q96TD7, Q96TD8, Q99709, Q9H133 | Gene names | LYST, CHS, CHS1 | |||
|
Domain Architecture |

|
|||||
| Description | Lysosomal-trafficking regulator (Beige homolog). | |||||
|
MILK1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 36) | NC score | 0.005798 (rank : 64) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 788 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BGT6, Q8BJ60 | Gene names | Mirab13, D15Mit260, Kiaa1668 | |||
|
Domain Architecture |

|
|||||
| Description | Molecule interacting with Rab13 (MIRab13) (MICAL-like protein 1). | |||||
|
SIM1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 37) | NC score | 0.070755 (rank : 48) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P81133, Q5TDP7 | Gene names | SIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1. | |||||
|
SIM1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 38) | NC score | 0.070877 (rank : 47) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q61045, O70284, P70183 | Gene names | Sim1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1 (mSIM1). | |||||
|
ARNT2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 39) | NC score | 0.093902 (rank : 40) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9HBZ2, O15024 | Gene names | ARNT2, KIAA0307 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
ARNT2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 40) | NC score | 0.095276 (rank : 39) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q61324 | Gene names | Arnt2 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
ARNT_HUMAN
|
||||||
| θ value | θ > 10 (rank : 41) | NC score | 0.092928 (rank : 41) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P27540 | Gene names | ARNT | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
ARNT_MOUSE
|
||||||
| θ value | θ > 10 (rank : 42) | NC score | 0.092254 (rank : 42) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P53762, Q60661 | Gene names | Arnt | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
EPAS1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 43) | NC score | 0.061536 (rank : 56) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q99814, Q86VA2, Q99630 | Gene names | EPAS1, HIF2A | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Member of PAS protein 2) (MOP2) (Hypoxia-inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (HLF). | |||||
|
HIF1A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 44) | NC score | 0.064582 (rank : 54) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q16665, Q96PT9, Q9UPB1 | Gene names | HIF1A | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein) (Member of PAS protein 1) (MOP1). | |||||
|
HIF1A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 45) | NC score | 0.062725 (rank : 55) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q61221, O08741, O08993, Q61664, Q61665, Q8C681, Q8CC19, Q8CCB6, Q8R385, Q9CYA8 | Gene names | Hif1a | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein). | |||||
|
MAD1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 46) | NC score | 0.051111 (rank : 61) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q05195 | Gene names | MXD1, MAD | |||
|
Domain Architecture |
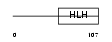
|
|||||
| Description | MAD protein (MAX dimerizer) (MAX dimerization protein 1). | |||||
|
MAD1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 47) | NC score | 0.066073 (rank : 52) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P50538, Q60798, Q61825 | Gene names | Mxd1, Mad, Mad1 | |||
|
Domain Architecture |

|
|||||
| Description | MAD protein (MAX dimerizer) (MAX dimerization protein 1). | |||||
|
MITF_HUMAN
|
||||||
| θ value | θ > 10 (rank : 48) | NC score | 0.103555 (rank : 36) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O75030, Q14841, Q9P2V0, Q9P2V1, Q9P2V2, Q9P2Y8 | Gene names | MITF | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
MITF_MOUSE
|
||||||
| θ value | θ > 10 (rank : 49) | NC score | 0.099599 (rank : 38) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q08874, O08885, O88203, Q08843, Q60781, Q60782, Q9JIJ0, Q9JIJ1, Q9JIJ2, Q9JIJ3, Q9JIJ4, Q9JIJ5, Q9JIJ6, Q9JKX9 | Gene names | Mitf, Bw, Mi, Vit | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
MLX_HUMAN
|
||||||
| θ value | θ > 10 (rank : 50) | NC score | 0.123943 (rank : 25) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9UH92, Q53XM6, Q96FL2, Q9H2V0, Q9H2V1, Q9H2V2, Q9NXN3 | Gene names | MLX, TCFL4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
MLX_MOUSE
|
||||||
| θ value | θ > 10 (rank : 51) | NC score | 0.123772 (rank : 26) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O08609, Q9EQJ7, Q9EQJ8 | Gene names | Mlx, Tcfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
NCOA1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 52) | NC score | 0.068503 (rank : 51) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q15788, O00150, O43792, O43793, Q13071, Q13420, Q6GVI5, Q7KYV3 | Gene names | NCOA1, SRC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (RIP160) (Protein Hin-2) (NY-REN-52 antigen). | |||||
|
NCOA1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 53) | NC score | 0.068851 (rank : 50) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P70365, P70366, Q61202, Q8CBI9 | Gene names | Ncoa1, Src1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (Nuclear receptor coactivator protein 1) (mNRC-1). | |||||
|
NPAS1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 54) | NC score | 0.052870 (rank : 59) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q99742, Q99632, Q9BY83 | Gene names | NPAS1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1) (Member of PAS protein 5) (MOP5). | |||||
|
NPAS1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 55) | NC score | 0.052823 (rank : 60) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P97459 | Gene names | Npas1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1). | |||||
|
NPAS3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 56) | NC score | 0.061048 (rank : 58) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8IXF0, Q86US6, Q86US7, Q8IXF2, Q9BY81, Q9H323, Q9Y4L8 | Gene names | NPAS3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
NPAS3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 57) | NC score | 0.061246 (rank : 57) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9QZQ0, Q9EQP4 | Gene names | Npas3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
SIM2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 58) | NC score | 0.069140 (rank : 49) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q61079, O35391, Q61046, Q61904 | Gene names | Sim2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2 (SIM transcription factor) (mSIM). | |||||
|
SRBP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 59) | NC score | 0.100162 (rank : 37) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P36956, Q16062 | Gene names | SREBF1, SREBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
SRBP1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 60) | NC score | 0.090193 (rank : 43) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9WTN3 | Gene names | Srebf1, Srebp1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
TFE3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 61) | NC score | 0.105941 (rank : 34) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P19532, Q92757, Q92758, Q99964 | Gene names | TFE3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E3. | |||||
|
TFEB_HUMAN
|
||||||
| θ value | θ > 10 (rank : 62) | NC score | 0.083392 (rank : 45) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P19484, Q709B3, Q7Z6P9, Q9BRJ5, Q9UJD8 | Gene names | TFEB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
TFEB_MOUSE
|
||||||
| θ value | θ > 10 (rank : 63) | NC score | 0.084680 (rank : 44) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9R210 | Gene names | Tfeb, Tcfeb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
USF2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 64) | NC score | 0.117727 (rank : 28) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q15853, O00671, O00709, Q05750, Q07952, Q15851, Q15852 | Gene names | USF2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (FOS- interacting protein) (FIP) (Major late transcription factor 2). | |||||
|
HES5_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 6.11685e-65 (rank : 1) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P70120, Q8BVI1 | Gene names | Hes5, Hes-5 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-5 (Hairy and enhancer of split 5). | |||||
|
HES4_HUMAN
|
||||||
| NC score | 0.849466 (rank : 2) | θ value | 4.30538e-10 (rank : 6) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9HCC6 | Gene names | HES4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-4 (Hairy and enhancer of split 4) (bHLH factor Hes4). | |||||
|
HES2_MOUSE
|
||||||
| NC score | 0.831350 (rank : 3) | θ value | 6.21693e-09 (rank : 10) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O54792 | Gene names | Hes2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-2 (Hairy and enhancer of split 2). | |||||
|
HES2_HUMAN
|
||||||
| NC score | 0.826413 (rank : 4) | θ value | 2.79066e-09 (rank : 9) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9Y543, Q96EN4, Q9Y542 | Gene names | HES2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-2 (Hairy and enhancer of split 2). | |||||
|
HES1_MOUSE
|
||||||
| NC score | 0.821829 (rank : 5) | θ value | 1.74796e-11 (rank : 3) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P35428 | Gene names | Hes1, Hes-1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-1 (Hairy and enhancer of split 1). | |||||
|
HES1_HUMAN
|
||||||
| NC score | 0.821421 (rank : 6) | θ value | 1.74796e-11 (rank : 2) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q14469 | Gene names | HES1, HL, HRY | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-1 (Hairy and enhancer of split 1) (Hairy- like) (HHL) (Hairy homolog). | |||||
|
HES7_HUMAN
|
||||||
| NC score | 0.772983 (rank : 7) | θ value | 3.29651e-10 (rank : 5) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9BYE0 | Gene names | HES7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor HES-7 (Hairy and enhancer of split 7) (bHLH factor Hes7). | |||||
|
HES7_MOUSE
|
||||||
| NC score | 0.772129 (rank : 8) | θ value | 8.67504e-11 (rank : 4) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8BKT2, Q99JA6 | Gene names | Hes7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor HES-7 (Hairy and enhancer of split 7) (bHLH factor Hes7). | |||||
|
HEY1_MOUSE
|
||||||
| NC score | 0.753503 (rank : 9) | θ value | 1.25267e-09 (rank : 8) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9WV93 | Gene names | Hey1, Hesr1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1). | |||||
|
HEY1_HUMAN
|
||||||
| NC score | 0.750373 (rank : 10) | θ value | 1.25267e-09 (rank : 7) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9Y5J3, Q5TZS3, Q9NYP4 | Gene names | HEY1, CHF2, HESR1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1) (Cardiovascular helix-loop-helix factor 2) (HES-related repressor protein 2) (HERP2). | |||||
|
HES3_MOUSE
|
||||||
| NC score | 0.749049 (rank : 11) | θ value | 8.97725e-08 (rank : 13) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q61657, O09083, O09150 | Gene names | Hes3, Hes-3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-3 (Hairy and enhancer of split 3). | |||||
|
HEY2_HUMAN
|
||||||
| NC score | 0.745967 (rank : 12) | θ value | 2.36244e-08 (rank : 11) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9UBP5 | Gene names | HEY2, CHF1, GRL, HERP, HERP1, HRT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Cardiovascular helix-loop-helix factor 1) (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES- related repressor protein 2) (Protein gridlock homolog). | |||||
|
HEY2_MOUSE
|
||||||
| NC score | 0.744202 (rank : 13) | θ value | 5.26297e-08 (rank : 12) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9QUS4, Q3TZ99, Q8CD44 | Gene names | Hey2, Chf1, Herp, Herp1, Hesr2, Hrt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES-related repressor protein 2). | |||||
|
HES6_HUMAN
|
||||||
| NC score | 0.722615 (rank : 14) | θ value | 3.77169e-06 (rank : 14) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96HZ4, Q8N2J2, Q96T93, Q9P2S3 | Gene names | HES6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription cofactor HES-6 (Hairy and enhancer of split 6) (C- HAIRY1). | |||||
|
HES6_MOUSE
|
||||||
| NC score | 0.714157 (rank : 15) | θ value | 3.77169e-06 (rank : 15) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9JHE6, Q3U2D7 | Gene names | Hes6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription cofactor HES-6 (Hairy and enhancer of split 6). | |||||
|
BHLH3_HUMAN
|
||||||
| NC score | 0.575593 (rank : 16) | θ value | 8.40245e-06 (rank : 16) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9C0J9 | Gene names | BHLHB3, DEC2, SHARP1 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (hDEC2) (Enhancer-of-split and hairy-related protein 1) (SHARP-1). | |||||
|
BHLH3_MOUSE
|
||||||
| NC score | 0.574536 (rank : 17) | θ value | 1.09739e-05 (rank : 17) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q99PV5 | Gene names | Bhlhb3, Dec2 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (mDEC2). | |||||
|
BHLH2_HUMAN
|
||||||
| NC score | 0.560765 (rank : 18) | θ value | 0.000461057 (rank : 19) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O14503, Q96TD3 | Gene names | BHLHB2, DEC1, SHARP2, STRA13 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 2 (bHLHB2) (Differentially expressed in chondrocytes protein 1) (DEC1) (Enhancer-of-split and hairy-related protein 2) (SHARP-2) (Stimulated with retinoic acid 13). | |||||
|
BHLH2_MOUSE
|
||||||
| NC score | 0.548708 (rank : 19) | θ value | 0.00035302 (rank : 18) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O35185, P97289 | Gene names | Bhlhb2, Clast5, Stra13 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 2 (bHLHB2) (Stimulated with retinoic acid 13) (E47 interaction protein 1) (eipl). | |||||
|
BMAL1_MOUSE
|
||||||
| NC score | 0.172604 (rank : 20) | θ value | 0.47712 (rank : 22) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9WTL8, O88295, Q921S4, Q9R0U2, Q9WTL9 | Gene names | Arntl, Bmal1 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Arnt3). | |||||
|
BMAL1_HUMAN
|
||||||
| NC score | 0.169443 (rank : 21) | θ value | 0.47712 (rank : 21) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O00327, O00313, O00314, O00315, O00316, O00317, Q99631, Q99649 | Gene names | ARNTL, BMAL1, MOP3 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Member of PAS protein 3) (Basic-helix-loop- helix-PAS orphan MOP3) (bHLH-PAS protein JAP3). | |||||
|
USF1_HUMAN
|
||||||
| NC score | 0.140174 (rank : 22) | θ value | 3.0926 (rank : 27) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P22415 | Gene names | USF1, USF | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF1_MOUSE
|
||||||
| NC score | 0.140169 (rank : 23) | θ value | 3.0926 (rank : 28) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q61069 | Gene names | Usf1, Usf | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF2_MOUSE
|
||||||
| NC score | 0.129244 (rank : 24) | θ value | 2.36792 (rank : 26) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q64705, Q3UIL2, Q8VE59 | Gene names | Usf2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (Major late transcription factor 2). | |||||
|
MLX_HUMAN
|
||||||
| NC score | 0.123943 (rank : 25) | θ value | θ > 10 (rank : 50) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9UH92, Q53XM6, Q96FL2, Q9H2V0, Q9H2V1, Q9H2V2, Q9NXN3 | Gene names | MLX, TCFL4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
MLX_MOUSE
|
||||||
| NC score | 0.123772 (rank : 26) | θ value | θ > 10 (rank : 51) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O08609, Q9EQJ7, Q9EQJ8 | Gene names | Mlx, Tcfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
CLOCK_HUMAN
|
||||||
| NC score | 0.121847 (rank : 27) | θ value | 0.62314 (rank : 23) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O15516, O14516, Q9UIT8 | Gene names | CLOCK, KIAA0334 | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (hCLOCK). | |||||
|
USF2_HUMAN
|
||||||
| NC score | 0.117727 (rank : 28) | θ value | θ > 10 (rank : 64) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q15853, O00671, O00709, Q05750, Q07952, Q15851, Q15852 | Gene names | USF2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (FOS- interacting protein) (FIP) (Major late transcription factor 2). | |||||
|
CLOCK_MOUSE
|
||||||
| NC score | 0.117486 (rank : 29) | θ value | 1.06291 (rank : 25) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O08785 | Gene names | Clock | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (mCLOCK). | |||||
|
SRBP2_HUMAN
|
||||||
| NC score | 0.114189 (rank : 30) | θ value | 5.27518 (rank : 30) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q12772, Q9UH04 | Gene names | SREBF2, SREBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 2 (SREBP-2) (Sterol regulatory element-binding transcription factor 2). | |||||
|
AHR_MOUSE
|
||||||
| NC score | 0.107679 (rank : 31) | θ value | 0.21417 (rank : 20) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P30561, Q8QZX6, Q8R4S3, Q99P79, Q9QVY1 | Gene names | Ahr | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
NPAS2_MOUSE
|
||||||
| NC score | 0.106550 (rank : 32) | θ value | 6.88961 (rank : 33) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P97460 | Gene names | Npas2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2). | |||||
|
NPAS2_HUMAN
|
||||||
| NC score | 0.106422 (rank : 33) | θ value | 6.88961 (rank : 32) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q99743, Q99629 | Gene names | NPAS2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2) (Member of PAS protein 4) (MOP4). | |||||
|
TFE3_HUMAN
|
||||||
| NC score | 0.105941 (rank : 34) | θ value | θ > 10 (rank : 61) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P19532, Q92757, Q92758, Q99964 | Gene names | TFE3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E3. | |||||
|
AHR_HUMAN
|
||||||
| NC score | 0.105869 (rank : 35) | θ value | 1.06291 (rank : 24) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P35869, Q13728, Q13803, Q13804 | Gene names | AHR | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
MITF_HUMAN
|
||||||
| NC score | 0.103555 (rank : 36) | θ value | θ > 10 (rank : 48) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O75030, Q14841, Q9P2V0, Q9P2V1, Q9P2V2, Q9P2Y8 | Gene names | MITF | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
SRBP1_HUMAN
|
||||||
| NC score | 0.100162 (rank : 37) | θ value | θ > 10 (rank : 59) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P36956, Q16062 | Gene names | SREBF1, SREBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
MITF_MOUSE
|
||||||
| NC score | 0.099599 (rank : 38) | θ value | θ > 10 (rank : 49) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q08874, O08885, O88203, Q08843, Q60781, Q60782, Q9JIJ0, Q9JIJ1, Q9JIJ2, Q9JIJ3, Q9JIJ4, Q9JIJ5, Q9JIJ6, Q9JKX9 | Gene names | Mitf, Bw, Mi, Vit | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
ARNT2_MOUSE
|
||||||
| NC score | 0.095276 (rank : 39) | θ value | θ > 10 (rank : 40) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q61324 | Gene names | Arnt2 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
ARNT2_HUMAN
|
||||||
| NC score | 0.093902 (rank : 40) | θ value | θ > 10 (rank : 39) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9HBZ2, O15024 | Gene names | ARNT2, KIAA0307 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
ARNT_HUMAN
|
||||||
| NC score | 0.092928 (rank : 41) | θ value | θ > 10 (rank : 41) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P27540 | Gene names | ARNT | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
ARNT_MOUSE
|
||||||
| NC score | 0.092254 (rank : 42) | θ value | θ > 10 (rank : 42) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P53762, Q60661 | Gene names | Arnt | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
SRBP1_MOUSE
|
||||||
| NC score | 0.090193 (rank : 43) | θ value | θ > 10 (rank : 60) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9WTN3 | Gene names | Srebf1, Srebp1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
TFEB_MOUSE
|
||||||
| NC score | 0.084680 (rank : 44) | θ value | θ > 10 (rank : 63) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9R210 | Gene names | Tfeb, Tcfeb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
TFEB_HUMAN
|
||||||
| NC score | 0.083392 (rank : 45) | θ value | θ > 10 (rank : 62) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P19484, Q709B3, Q7Z6P9, Q9BRJ5, Q9UJD8 | Gene names | TFEB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
SIM2_HUMAN
|
||||||
| NC score | 0.073616 (rank : 46) | θ value | 6.88961 (rank : 34) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q14190, O60766, Q15470, Q15471, Q15472, Q15473, Q16532 | Gene names | SIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2. | |||||
|
SIM1_MOUSE
|
||||||
| NC score | 0.070877 (rank : 47) | θ value | 8.99809 (rank : 38) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q61045, O70284, P70183 | Gene names | Sim1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1 (mSIM1). | |||||
|
SIM1_HUMAN
|
||||||
| NC score | 0.070755 (rank : 48) | θ value | 8.99809 (rank : 37) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P81133, Q5TDP7 | Gene names | SIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1. | |||||
|
SIM2_MOUSE
|
||||||
| NC score | 0.069140 (rank : 49) | θ value | θ > 10 (rank : 58) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q61079, O35391, Q61046, Q61904 | Gene names | Sim2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2 (SIM transcription factor) (mSIM). | |||||
|
NCOA1_MOUSE
|
||||||
| NC score | 0.068851 (rank : 50) | θ value | θ > 10 (rank : 53) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P70365, P70366, Q61202, Q8CBI9 | Gene names | Ncoa1, Src1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (Nuclear receptor coactivator protein 1) (mNRC-1). | |||||
|
NCOA1_HUMAN
|
||||||
| NC score | 0.068503 (rank : 51) | θ value | θ > 10 (rank : 52) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q15788, O00150, O43792, O43793, Q13071, Q13420, Q6GVI5, Q7KYV3 | Gene names | NCOA1, SRC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (RIP160) (Protein Hin-2) (NY-REN-52 antigen). | |||||
|
MAD1_MOUSE
|
||||||
| NC score | 0.066073 (rank : 52) | θ value | θ > 10 (rank : 47) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P50538, Q60798, Q61825 | Gene names | Mxd1, Mad, Mad1 | |||
|
Domain Architecture |

|
|||||
| Description | MAD protein (MAX dimerizer) (MAX dimerization protein 1). | |||||
|
EPAS1_MOUSE
|
||||||
| NC score | 0.065676 (rank : 53) | θ value | 4.03905 (rank : 29) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P97481, O08787, O55046 | Gene names | Epas1, Hif2a | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Hypoxia- inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (MHLF) (HIF-related factor) (HRF). | |||||
|
HIF1A_HUMAN
|
||||||
| NC score | 0.064582 (rank : 54) | θ value | θ > 10 (rank : 44) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q16665, Q96PT9, Q9UPB1 | Gene names | HIF1A | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein) (Member of PAS protein 1) (MOP1). | |||||
|
HIF1A_MOUSE
|
||||||
| NC score | 0.062725 (rank : 55) | θ value | θ > 10 (rank : 45) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q61221, O08741, O08993, Q61664, Q61665, Q8C681, Q8CC19, Q8CCB6, Q8R385, Q9CYA8 | Gene names | Hif1a | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein). | |||||
|
EPAS1_HUMAN
|
||||||
| NC score | 0.061536 (rank : 56) | θ value | θ > 10 (rank : 43) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q99814, Q86VA2, Q99630 | Gene names | EPAS1, HIF2A | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Member of PAS protein 2) (MOP2) (Hypoxia-inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (HLF). | |||||
|
NPAS3_MOUSE
|
||||||
| NC score | 0.061246 (rank : 57) | θ value | θ > 10 (rank : 57) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9QZQ0, Q9EQP4 | Gene names | Npas3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
NPAS3_HUMAN
|
||||||
| NC score | 0.061048 (rank : 58) | θ value | θ > 10 (rank : 56) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8IXF0, Q86US6, Q86US7, Q8IXF2, Q9BY81, Q9H323, Q9Y4L8 | Gene names | NPAS3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
NPAS1_HUMAN
|
||||||
| NC score | 0.052870 (rank : 59) | θ value | θ > 10 (rank : 54) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q99742, Q99632, Q9BY83 | Gene names | NPAS1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1) (Member of PAS protein 5) (MOP5). | |||||
|
NPAS1_MOUSE
|
||||||
| NC score | 0.052823 (rank : 60) | θ value | θ > 10 (rank : 55) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P97459 | Gene names | Npas1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1). | |||||
|
MAD1_HUMAN
|
||||||
| NC score | 0.051111 (rank : 61) | θ value | θ > 10 (rank : 46) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q05195 | Gene names | MXD1, MAD | |||
|
Domain Architecture |

|
|||||
| Description | MAD protein (MAX dimerizer) (MAX dimerization protein 1). | |||||
|
MAP1A_HUMAN
|
||||||
| NC score | 0.008666 (rank : 62) | θ value | 6.88961 (rank : 31) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P78559, O95643, Q12973, Q15882, Q9UJT4 | Gene names | MAP1A | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 1A (MAP 1A) (Proliferation-related protein p80) [Contains: MAP1 light chain LC2]. | |||||
|
LYST_HUMAN
|
||||||
| NC score | 0.008449 (rank : 63) | θ value | 8.99809 (rank : 35) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 324 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99698, O43274, Q5T2U9, Q96TD7, Q96TD8, Q99709, Q9H133 | Gene names | LYST, CHS, CHS1 | |||
|
Domain Architecture |

|
|||||
| Description | Lysosomal-trafficking regulator (Beige homolog). | |||||
|
MILK1_MOUSE
|
||||||
| NC score | 0.005798 (rank : 64) | θ value | 8.99809 (rank : 36) | |||
| Query Neighborhood Hits | 38 | Target Neighborhood Hits | 788 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BGT6, Q8BJ60 | Gene names | Mirab13, D15Mit260, Kiaa1668 | |||
|
Domain Architecture |

|
|||||
| Description | Molecule interacting with Rab13 (MIRab13) (MICAL-like protein 1). | |||||