Please be patient as the page loads
|
TCFL5_HUMAN
|
||||||
| SwissProt Accessions | Q9UL49, O94771, Q9BYW0 | Gene names | TCFL5, CHA, E2BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor-like 5 protein (Cha transcription factor) (HPV-16 E2-binding protein 1) (E2BP-1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
TCFL5_HUMAN
|
||||||
| θ value | 3.96267e-181 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9UL49, O94771, Q9BYW0 | Gene names | TCFL5, CHA, E2BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor-like 5 protein (Cha transcription factor) (HPV-16 E2-binding protein 1) (E2BP-1). | |||||
|
BMAL1_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 2) | NC score | 0.205864 (rank : 10) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O00327, O00313, O00314, O00315, O00316, O00317, Q99631, Q99649 | Gene names | ARNTL, BMAL1, MOP3 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Member of PAS protein 3) (Basic-helix-loop- helix-PAS orphan MOP3) (bHLH-PAS protein JAP3). | |||||
|
BMAL1_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 3) | NC score | 0.205259 (rank : 11) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9WTL8, O88295, Q921S4, Q9R0U2, Q9WTL9 | Gene names | Arntl, Bmal1 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Arnt3). | |||||
|
MAD1_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 4) | NC score | 0.230905 (rank : 5) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P50538, Q60798, Q61825 | Gene names | Mxd1, Mad, Mad1 | |||
|
Domain Architecture |

|
|||||
| Description | MAD protein (MAX dimerizer) (MAX dimerization protein 1). | |||||
|
MLX_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 5) | NC score | 0.204771 (rank : 12) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O08609, Q9EQJ7, Q9EQJ8 | Gene names | Mlx, Tcfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
MXI1_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 6) | NC score | 0.225610 (rank : 6) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P50539, Q15887 | Gene names | MXI1 | |||
|
Domain Architecture |
|
|||||
| Description | MAX-interacting protein 1 (Protein MXI1). | |||||
|
MXI1_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 7) | NC score | 0.223373 (rank : 7) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P50540 | Gene names | Mxi1 | |||
|
Domain Architecture |
|
|||||
| Description | MAX-interacting protein 1 (Protein MXI1). | |||||
|
MNT_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 8) | NC score | 0.186407 (rank : 19) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 432 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O08789, P97349 | Gene names | Mnt, Rox | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
MAD1_HUMAN
|
||||||
| θ value | 0.125558 (rank : 9) | NC score | 0.216170 (rank : 9) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q05195 | Gene names | MXD1, MAD | |||
|
Domain Architecture |
|
|||||
| Description | MAD protein (MAX dimerizer) (MAX dimerization protein 1). | |||||
|
TFEB_HUMAN
|
||||||
| θ value | 0.125558 (rank : 10) | NC score | 0.195340 (rank : 15) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P19484, Q709B3, Q7Z6P9, Q9BRJ5, Q9UJD8 | Gene names | TFEB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
TFEB_MOUSE
|
||||||
| θ value | 0.125558 (rank : 11) | NC score | 0.196580 (rank : 13) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9R210 | Gene names | Tfeb, Tcfeb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
MLX_HUMAN
|
||||||
| θ value | 0.163984 (rank : 12) | NC score | 0.191355 (rank : 17) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9UH92, Q53XM6, Q96FL2, Q9H2V0, Q9H2V1, Q9H2V2, Q9NXN3 | Gene names | MLX, TCFL4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
ARNT_HUMAN
|
||||||
| θ value | 0.21417 (rank : 13) | NC score | 0.193817 (rank : 16) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P27540 | Gene names | ARNT | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
ARNT_MOUSE
|
||||||
| θ value | 0.21417 (rank : 14) | NC score | 0.190103 (rank : 18) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P53762, Q60661 | Gene names | Arnt | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
MNT_HUMAN
|
||||||
| θ value | 0.21417 (rank : 15) | NC score | 0.174868 (rank : 22) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 496 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q99583 | Gene names | MNT, ROX | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
USF2_HUMAN
|
||||||
| θ value | 0.21417 (rank : 16) | NC score | 0.232648 (rank : 3) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q15853, O00671, O00709, Q05750, Q07952, Q15851, Q15852 | Gene names | USF2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (FOS- interacting protein) (FIP) (Major late transcription factor 2). | |||||
|
USF1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 17) | NC score | 0.232757 (rank : 2) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P22415 | Gene names | USF1, USF | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF1_MOUSE
|
||||||
| θ value | 0.279714 (rank : 18) | NC score | 0.232540 (rank : 4) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q61069 | Gene names | Usf1, Usf | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
ARNT2_HUMAN
|
||||||
| θ value | 0.47712 (rank : 19) | NC score | 0.183829 (rank : 21) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9HBZ2, O15024 | Gene names | ARNT2, KIAA0307 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
ARNT2_MOUSE
|
||||||
| θ value | 0.47712 (rank : 20) | NC score | 0.184628 (rank : 20) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q61324 | Gene names | Arnt2 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
USF2_MOUSE
|
||||||
| θ value | 0.813845 (rank : 21) | NC score | 0.220047 (rank : 8) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q64705, Q3UIL2, Q8VE59 | Gene names | Usf2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (Major late transcription factor 2). | |||||
|
K0310_HUMAN
|
||||||
| θ value | 1.38821 (rank : 22) | NC score | 0.065014 (rank : 57) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O15027, Q5SXP0, Q5SXP1, Q8N347, Q96HP1 | Gene names | KIAA0310 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0310. | |||||
|
MAD4_HUMAN
|
||||||
| θ value | 2.36792 (rank : 23) | NC score | 0.195729 (rank : 14) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q14582, Q5TZX4 | Gene names | MXD4, MAD4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-interacting transcriptional repressor MAD4 (Max-associated protein 4) (MAX dimerization protein 4). | |||||
|
MAP1B_MOUSE
|
||||||
| θ value | 2.36792 (rank : 24) | NC score | 0.033651 (rank : 80) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 977 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P14873 | Gene names | Map1b, Mtap1b, Mtap5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) (MAP1.2) (MAP1(X)) [Contains: MAP1 light chain LC1]. | |||||
|
CLOCK_MOUSE
|
||||||
| θ value | 3.0926 (rank : 25) | NC score | 0.132758 (rank : 31) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O08785 | Gene names | Clock | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (mCLOCK). | |||||
|
CLOCK_HUMAN
|
||||||
| θ value | 4.03905 (rank : 26) | NC score | 0.130633 (rank : 32) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O15516, O14516, Q9UIT8 | Gene names | CLOCK, KIAA0334 | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (hCLOCK). | |||||
|
NALDL_HUMAN
|
||||||
| θ value | 4.03905 (rank : 27) | NC score | 0.022089 (rank : 87) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UQQ1, O43176 | Gene names | NAALADL1, NAALADASEL, NAALADL | |||
|
Domain Architecture |

|
|||||
| Description | N-acetylated-alpha-linked acidic dipeptidase-like protein (EC 3.4.17.21) (NAALADase L) (Ileal dipeptidylpeptidase) (100 kDa ileum brush border membrane protein) (I100). | |||||
|
PHAR1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 28) | NC score | 0.038505 (rank : 79) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q2M3X8, Q80VL9, Q8C873 | Gene names | Phactr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatase and actin regulator 1. | |||||
|
HAIR_HUMAN
|
||||||
| θ value | 5.27518 (rank : 29) | NC score | 0.022914 (rank : 85) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O43593, Q96H33, Q9NPE1 | Gene names | HR | |||
|
Domain Architecture |

|
|||||
| Description | Protein hairless. | |||||
|
NPAS2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 30) | NC score | 0.129367 (rank : 33) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q99743, Q99629 | Gene names | NPAS2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2) (Member of PAS protein 4) (MOP4). | |||||
|
NPAS2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 31) | NC score | 0.127927 (rank : 34) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P97460 | Gene names | Npas2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2). | |||||
|
TTC16_HUMAN
|
||||||
| θ value | 5.27518 (rank : 32) | NC score | 0.022821 (rank : 86) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NEE8, Q5JU66, Q96M72 | Gene names | TTC16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tetratricopeptide repeat protein 16 (TPR repeat protein 16). | |||||
|
DCP1A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 33) | NC score | 0.027546 (rank : 82) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NPI6 | Gene names | DCP1A, SMIF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | mRNA decapping enzyme 1A (EC 3.-.-.-) (Transcription factor SMIF) (Smad4-interacting transcriptional co-activator). | |||||
|
MAP1A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 34) | NC score | 0.020804 (rank : 88) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P78559, O95643, Q12973, Q15882, Q9UJT4 | Gene names | MAP1A | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 1A (MAP 1A) (Proliferation-related protein p80) [Contains: MAP1 light chain LC2]. | |||||
|
PDXL2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 35) | NC score | 0.025818 (rank : 83) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CAE9, Q8CFW3 | Gene names | Podxl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Podocalyxin-like protein 2 precursor (Endoglycan). | |||||
|
PLRG1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 36) | NC score | 0.007951 (rank : 89) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O43660, Q8WUD8 | Gene names | PLRG1 | |||
|
Domain Architecture |

|
|||||
| Description | Pleiotropic regulator 1. | |||||
|
WBS14_HUMAN
|
||||||
| θ value | 6.88961 (rank : 37) | NC score | 0.068592 (rank : 46) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9NP71, Q96E48, Q9BY03, Q9BY04, Q9BY05, Q9BY06, Q9Y2P3 | Gene names | MLXIPL, MIO, WBSCR14 | |||
|
Domain Architecture |
|
|||||
| Description | Williams-Beuren syndrome chromosome region 14 protein (WS basic-helix- loop-helix leucine zipper protein) (WS-bHLH) (Mlx interactor) (MLX- interacting protein-like). | |||||
|
IER5_MOUSE
|
||||||
| θ value | 8.99809 (rank : 38) | NC score | 0.023890 (rank : 84) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O89113, Q8CGI0 | Gene names | Ier5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Immediate early response gene 5 protein. | |||||
|
MAX_MOUSE
|
||||||
| θ value | 8.99809 (rank : 39) | NC score | 0.145998 (rank : 29) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P28574 | Gene names | Max, Myn | |||
|
Domain Architecture |

|
|||||
| Description | Protein max (Myc-associated factor X) (Protein myn) (Myc-binding novel HLH/LZ protein). | |||||
|
MITF_HUMAN
|
||||||
| θ value | 8.99809 (rank : 40) | NC score | 0.158218 (rank : 24) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O75030, Q14841, Q9P2V0, Q9P2V1, Q9P2V2, Q9P2Y8 | Gene names | MITF | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
MITF_MOUSE
|
||||||
| θ value | 8.99809 (rank : 41) | NC score | 0.155810 (rank : 25) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q08874, O08885, O88203, Q08843, Q60781, Q60782, Q9JIJ0, Q9JIJ1, Q9JIJ2, Q9JIJ3, Q9JIJ4, Q9JIJ5, Q9JIJ6, Q9JKX9 | Gene names | Mitf, Bw, Mi, Vit | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
PHAR1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 42) | NC score | 0.032707 (rank : 81) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9C0D0, Q3MJ93, Q5JSJ2 | Gene names | PHACTR1, KIAA1733, RPEL1 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatase and actin regulator 1. | |||||
|
TFE3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 43) | NC score | 0.160439 (rank : 23) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P19532, Q92757, Q92758, Q99964 | Gene names | TFE3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E3. | |||||
|
AHR_HUMAN
|
||||||
| θ value | θ > 10 (rank : 44) | NC score | 0.068199 (rank : 48) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P35869, Q13728, Q13803, Q13804 | Gene names | AHR | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
AHR_MOUSE
|
||||||
| θ value | θ > 10 (rank : 45) | NC score | 0.065126 (rank : 56) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P30561, Q8QZX6, Q8R4S3, Q99P79, Q9QVY1 | Gene names | Ahr | |||
|
Domain Architecture |
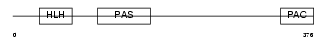
|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
BHLH3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 46) | NC score | 0.056871 (rank : 70) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9C0J9 | Gene names | BHLHB3, DEC2, SHARP1 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (hDEC2) (Enhancer-of-split and hairy-related protein 1) (SHARP-1). | |||||
|
BHLH3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 47) | NC score | 0.054346 (rank : 77) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q99PV5 | Gene names | Bhlhb3, Dec2 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (mDEC2). | |||||
|
EPAS1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 48) | NC score | 0.078977 (rank : 35) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q99814, Q86VA2, Q99630 | Gene names | EPAS1, HIF2A | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Member of PAS protein 2) (MOP2) (Hypoxia-inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (HLF). | |||||
|
EPAS1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 49) | NC score | 0.078190 (rank : 36) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P97481, O08787, O55046 | Gene names | Epas1, Hif2a | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Hypoxia- inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (MHLF) (HIF-related factor) (HRF). | |||||
|
HEY1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 50) | NC score | 0.065846 (rank : 53) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9Y5J3, Q5TZS3, Q9NYP4 | Gene names | HEY1, CHF2, HESR1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1) (Cardiovascular helix-loop-helix factor 2) (HES-related repressor protein 2) (HERP2). | |||||
|
HEY1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 51) | NC score | 0.068125 (rank : 49) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9WV93 | Gene names | Hey1, Hesr1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1). | |||||
|
HEY2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 52) | NC score | 0.070499 (rank : 39) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9UBP5 | Gene names | HEY2, CHF1, GRL, HERP, HERP1, HRT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Cardiovascular helix-loop-helix factor 1) (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES- related repressor protein 2) (Protein gridlock homolog). | |||||
|
HEY2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 53) | NC score | 0.068575 (rank : 47) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9QUS4, Q3TZ99, Q8CD44 | Gene names | Hey2, Chf1, Herp, Herp1, Hesr2, Hrt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES-related repressor protein 2). | |||||
|
HIF1A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 54) | NC score | 0.075848 (rank : 37) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q16665, Q96PT9, Q9UPB1 | Gene names | HIF1A | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein) (Member of PAS protein 1) (MOP1). | |||||
|
HIF1A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 55) | NC score | 0.075700 (rank : 38) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q61221, O08741, O08993, Q61664, Q61665, Q8C681, Q8CC19, Q8CCB6, Q8R385, Q9CYA8 | Gene names | Hif1a | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein). | |||||
|
MAD3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 56) | NC score | 0.153789 (rank : 26) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9BW11, Q53HK1, Q7Z4Y0, Q8NDJ7, Q96ME3 | Gene names | MXD3, MAD3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Max-interacting transcriptional repressor MAD3 (Max-associated protein 3) (MAX dimerization protein 3) (Myx). | |||||
|
MAD3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 57) | NC score | 0.152605 (rank : 27) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q80US8, Q60947 | Gene names | Mxd3, Mad3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Max-interacting transcriptional repressor MAD3 (Max-associated protein 3) (MAX dimerization protein 3). | |||||
|
MAD4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 58) | NC score | 0.152343 (rank : 28) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q60948 | Gene names | Mxd4, Mad4 | |||
|
Domain Architecture |
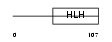
|
|||||
| Description | Max-interacting transcriptional repressor MAD4 (Max-associated protein 4) (MAX dimerization protein 4). | |||||
|
MAX_HUMAN
|
||||||
| θ value | θ > 10 (rank : 59) | NC score | 0.134367 (rank : 30) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P61244, P25912, P52163 | Gene names | MAX | |||
|
Domain Architecture |

|
|||||
| Description | Protein max (Myc-associated factor X). | |||||
|
MYCL1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 60) | NC score | 0.065700 (rank : 54) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P12524, Q14897, Q5QPL0, Q5QPL1, Q9NUE9 | Gene names | MYCL1, LMYC, MYCL | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
MYCL1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 61) | NC score | 0.070082 (rank : 42) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P10166, Q5FWI7 | Gene names | Mycl1, Lmyc1, Mycl | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
MYCN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 62) | NC score | 0.062617 (rank : 59) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P04198, Q6LDT9 | Gene names | MYCN, NMYC | |||
|
Domain Architecture |

|
|||||
| Description | N-myc proto-oncogene protein. | |||||
|
MYCN_MOUSE
|
||||||
| θ value | θ > 10 (rank : 63) | NC score | 0.062015 (rank : 61) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P03966, Q61978 | Gene names | Mycn, Nmyc, Nmyc1 | |||
|
Domain Architecture |

|
|||||
| Description | N-myc proto-oncogene protein. | |||||
|
MYCS_MOUSE
|
||||||
| θ value | θ > 10 (rank : 64) | NC score | 0.054725 (rank : 76) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9Z304 | Gene names | Mycs | |||
|
Domain Architecture |
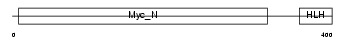
|
|||||
| Description | Protein S-myc. | |||||
|
MYC_HUMAN
|
||||||
| θ value | θ > 10 (rank : 65) | NC score | 0.067639 (rank : 51) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P01106, P01107, Q14026 | Gene names | MYC | |||
|
Domain Architecture |

|
|||||
| Description | Myc proto-oncogene protein (c-Myc) (Transcription factor p64). | |||||
|
MYC_MOUSE
|
||||||
| θ value | θ > 10 (rank : 66) | NC score | 0.070245 (rank : 40) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P01108, P70247, Q3UM70, Q61422 | Gene names | Myc | |||
|
Domain Architecture |
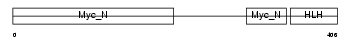
|
|||||
| Description | Myc proto-oncogene protein (c-Myc) (Transcription factor p64). | |||||
|
NCOA1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 67) | NC score | 0.062012 (rank : 62) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q15788, O00150, O43792, O43793, Q13071, Q13420, Q6GVI5, Q7KYV3 | Gene names | NCOA1, SRC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (RIP160) (Protein Hin-2) (NY-REN-52 antigen). | |||||
|
NCOA1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 68) | NC score | 0.062117 (rank : 60) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P70365, P70366, Q61202, Q8CBI9 | Gene names | Ncoa1, Src1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (Nuclear receptor coactivator protein 1) (mNRC-1). | |||||
|
NCOA3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 69) | NC score | 0.059109 (rank : 67) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y6Q9, Q5JYD9, Q5JYE0, Q9BR49, Q9UPC9, Q9UPG4, Q9UPG7 | Gene names | NCOA3, AIB1, RAC3, TRAM1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP-interacting protein) (pCIP). | |||||
|
NCOA3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 70) | NC score | 0.051627 (rank : 78) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O09000, Q9CRD5 | Gene names | Ncoa3, Aib1, Pcip, Rac3, Tram1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein homolog) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP- interacting protein) (p/CIP) (pCIP). | |||||
|
NPAS1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 71) | NC score | 0.060381 (rank : 65) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q99742, Q99632, Q9BY83 | Gene names | NPAS1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1) (Member of PAS protein 5) (MOP5). | |||||
|
NPAS1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 72) | NC score | 0.060532 (rank : 64) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P97459 | Gene names | Npas1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1). | |||||
|
NPAS3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 73) | NC score | 0.063824 (rank : 58) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8IXF0, Q86US6, Q86US7, Q8IXF2, Q9BY81, Q9H323, Q9Y4L8 | Gene names | NPAS3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
NPAS3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 74) | NC score | 0.065597 (rank : 55) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9QZQ0, Q9EQP4 | Gene names | Npas3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
PER1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 75) | NC score | 0.055475 (rank : 73) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O15534 | Gene names | PER1, KIAA0482, PER, RIGUI | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (hPER). | |||||
|
PER1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 76) | NC score | 0.056786 (rank : 71) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O35973 | Gene names | Per1, Per, Rigui | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (mPER) (M-Rigui). | |||||
|
PER2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 77) | NC score | 0.059229 (rank : 66) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O15055 | Gene names | PER2, KIAA0347 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2. | |||||
|
PER2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 78) | NC score | 0.058812 (rank : 69) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O54943, O54954 | Gene names | Per2 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2 (mPER2). | |||||
|
PER3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 79) | NC score | 0.055197 (rank : 74) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P56645, Q5H8X4, Q969K6, Q96S77, Q96S78, Q9C0J3, Q9NSP9, Q9UGU8 | Gene names | PER3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (hPER3). | |||||
|
PER3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 80) | NC score | 0.059061 (rank : 68) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O70361 | Gene names | Per3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (mPER3). | |||||
|
SIM1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 81) | NC score | 0.069678 (rank : 44) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P81133, Q5TDP7 | Gene names | SIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1. | |||||
|
SIM1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 82) | NC score | 0.069764 (rank : 43) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q61045, O70284, P70183 | Gene names | Sim1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1 (mSIM1). | |||||
|
SIM2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 83) | NC score | 0.070143 (rank : 41) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q14190, O60766, Q15470, Q15471, Q15472, Q15473, Q16532 | Gene names | SIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2. | |||||
|
SIM2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 84) | NC score | 0.068799 (rank : 45) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q61079, O35391, Q61046, Q61904 | Gene names | Sim2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2 (SIM transcription factor) (mSIM). | |||||
|
SRBP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 85) | NC score | 0.067982 (rank : 50) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P36956, Q16062 | Gene names | SREBF1, SREBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
SRBP1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 86) | NC score | 0.067114 (rank : 52) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9WTN3 | Gene names | Srebf1, Srebp1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
SRBP2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 87) | NC score | 0.060946 (rank : 63) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q12772, Q9UH04 | Gene names | SREBF2, SREBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 2 (SREBP-2) (Sterol regulatory element-binding transcription factor 2). | |||||
|
TFAP4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 88) | NC score | 0.054856 (rank : 75) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q01664, O60409 | Gene names | TFAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor AP-4 (Activating enhancer-binding protein 4). | |||||
|
WBS14_MOUSE
|
||||||
| θ value | θ > 10 (rank : 89) | NC score | 0.056333 (rank : 72) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q99MZ3, Q99MY9, Q99MZ0, Q99MZ1, Q99MZ2, Q9JLM5 | Gene names | Mlxipl, Mio, Wbscr14 | |||
|
Domain Architecture |

|
|||||
| Description | Williams-Beuren syndrome chromosome region 14 protein homolog (Mlx interactor) (MLX-interacting protein-like). | |||||
|
TCFL5_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 3.96267e-181 (rank : 1) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9UL49, O94771, Q9BYW0 | Gene names | TCFL5, CHA, E2BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor-like 5 protein (Cha transcription factor) (HPV-16 E2-binding protein 1) (E2BP-1). | |||||
|
USF1_HUMAN
|
||||||
| NC score | 0.232757 (rank : 2) | θ value | 0.279714 (rank : 17) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P22415 | Gene names | USF1, USF | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF2_HUMAN
|
||||||
| NC score | 0.232648 (rank : 3) | θ value | 0.21417 (rank : 16) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q15853, O00671, O00709, Q05750, Q07952, Q15851, Q15852 | Gene names | USF2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (FOS- interacting protein) (FIP) (Major late transcription factor 2). | |||||
|
USF1_MOUSE
|
||||||
| NC score | 0.232540 (rank : 4) | θ value | 0.279714 (rank : 18) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q61069 | Gene names | Usf1, Usf | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
MAD1_MOUSE
|
||||||
| NC score | 0.230905 (rank : 5) | θ value | 0.0113563 (rank : 4) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P50538, Q60798, Q61825 | Gene names | Mxd1, Mad, Mad1 | |||
|
Domain Architecture |

|
|||||
| Description | MAD protein (MAX dimerizer) (MAX dimerization protein 1). | |||||
|
MXI1_HUMAN
|
||||||
| NC score | 0.225610 (rank : 6) | θ value | 0.0736092 (rank : 6) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P50539, Q15887 | Gene names | MXI1 | |||
|
Domain Architecture |

|
|||||
| Description | MAX-interacting protein 1 (Protein MXI1). | |||||
|
MXI1_MOUSE
|
||||||
| NC score | 0.223373 (rank : 7) | θ value | 0.0736092 (rank : 7) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P50540 | Gene names | Mxi1 | |||
|
Domain Architecture |

|
|||||
| Description | MAX-interacting protein 1 (Protein MXI1). | |||||
|
USF2_MOUSE
|
||||||
| NC score | 0.220047 (rank : 8) | θ value | 0.813845 (rank : 21) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q64705, Q3UIL2, Q8VE59 | Gene names | Usf2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (Major late transcription factor 2). | |||||
|
MAD1_HUMAN
|
||||||
| NC score | 0.216170 (rank : 9) | θ value | 0.125558 (rank : 9) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q05195 | Gene names | MXD1, MAD | |||
|
Domain Architecture |

|
|||||
| Description | MAD protein (MAX dimerizer) (MAX dimerization protein 1). | |||||
|
BMAL1_HUMAN
|
||||||
| NC score | 0.205864 (rank : 10) | θ value | 0.0113563 (rank : 2) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O00327, O00313, O00314, O00315, O00316, O00317, Q99631, Q99649 | Gene names | ARNTL, BMAL1, MOP3 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Member of PAS protein 3) (Basic-helix-loop- helix-PAS orphan MOP3) (bHLH-PAS protein JAP3). | |||||
|
BMAL1_MOUSE
|
||||||
| NC score | 0.205259 (rank : 11) | θ value | 0.0113563 (rank : 3) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9WTL8, O88295, Q921S4, Q9R0U2, Q9WTL9 | Gene names | Arntl, Bmal1 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Arnt3). | |||||
|
MLX_MOUSE
|
||||||
| NC score | 0.204771 (rank : 12) | θ value | 0.0563607 (rank : 5) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O08609, Q9EQJ7, Q9EQJ8 | Gene names | Mlx, Tcfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
TFEB_MOUSE
|
||||||
| NC score | 0.196580 (rank : 13) | θ value | 0.125558 (rank : 11) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9R210 | Gene names | Tfeb, Tcfeb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
MAD4_HUMAN
|
||||||
| NC score | 0.195729 (rank : 14) | θ value | 2.36792 (rank : 23) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q14582, Q5TZX4 | Gene names | MXD4, MAD4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-interacting transcriptional repressor MAD4 (Max-associated protein 4) (MAX dimerization protein 4). | |||||
|
TFEB_HUMAN
|
||||||
| NC score | 0.195340 (rank : 15) | θ value | 0.125558 (rank : 10) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P19484, Q709B3, Q7Z6P9, Q9BRJ5, Q9UJD8 | Gene names | TFEB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
ARNT_HUMAN
|
||||||
| NC score | 0.193817 (rank : 16) | θ value | 0.21417 (rank : 13) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P27540 | Gene names | ARNT | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
MLX_HUMAN
|
||||||
| NC score | 0.191355 (rank : 17) | θ value | 0.163984 (rank : 12) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9UH92, Q53XM6, Q96FL2, Q9H2V0, Q9H2V1, Q9H2V2, Q9NXN3 | Gene names | MLX, TCFL4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
ARNT_MOUSE
|
||||||
| NC score | 0.190103 (rank : 18) | θ value | 0.21417 (rank : 14) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P53762, Q60661 | Gene names | Arnt | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
MNT_MOUSE
|
||||||
| NC score | 0.186407 (rank : 19) | θ value | 0.0961366 (rank : 8) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 432 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O08789, P97349 | Gene names | Mnt, Rox | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
ARNT2_MOUSE
|
||||||
| NC score | 0.184628 (rank : 20) | θ value | 0.47712 (rank : 20) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q61324 | Gene names | Arnt2 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
ARNT2_HUMAN
|
||||||
| NC score | 0.183829 (rank : 21) | θ value | 0.47712 (rank : 19) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9HBZ2, O15024 | Gene names | ARNT2, KIAA0307 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
MNT_HUMAN
|
||||||
| NC score | 0.174868 (rank : 22) | θ value | 0.21417 (rank : 15) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 496 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q99583 | Gene names | MNT, ROX | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
TFE3_HUMAN
|
||||||
| NC score | 0.160439 (rank : 23) | θ value | 8.99809 (rank : 43) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P19532, Q92757, Q92758, Q99964 | Gene names | TFE3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E3. | |||||
|
MITF_HUMAN
|
||||||
| NC score | 0.158218 (rank : 24) | θ value | 8.99809 (rank : 40) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O75030, Q14841, Q9P2V0, Q9P2V1, Q9P2V2, Q9P2Y8 | Gene names | MITF | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
MITF_MOUSE
|
||||||
| NC score | 0.155810 (rank : 25) | θ value | 8.99809 (rank : 41) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q08874, O08885, O88203, Q08843, Q60781, Q60782, Q9JIJ0, Q9JIJ1, Q9JIJ2, Q9JIJ3, Q9JIJ4, Q9JIJ5, Q9JIJ6, Q9JKX9 | Gene names | Mitf, Bw, Mi, Vit | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
MAD3_HUMAN
|
||||||
| NC score | 0.153789 (rank : 26) | θ value | θ > 10 (rank : 56) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9BW11, Q53HK1, Q7Z4Y0, Q8NDJ7, Q96ME3 | Gene names | MXD3, MAD3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Max-interacting transcriptional repressor MAD3 (Max-associated protein 3) (MAX dimerization protein 3) (Myx). | |||||
|
MAD3_MOUSE
|
||||||
| NC score | 0.152605 (rank : 27) | θ value | θ > 10 (rank : 57) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q80US8, Q60947 | Gene names | Mxd3, Mad3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Max-interacting transcriptional repressor MAD3 (Max-associated protein 3) (MAX dimerization protein 3). | |||||
|
MAD4_MOUSE
|
||||||
| NC score | 0.152343 (rank : 28) | θ value | θ > 10 (rank : 58) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q60948 | Gene names | Mxd4, Mad4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-interacting transcriptional repressor MAD4 (Max-associated protein 4) (MAX dimerization protein 4). | |||||
|
MAX_MOUSE
|
||||||
| NC score | 0.145998 (rank : 29) | θ value | 8.99809 (rank : 39) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P28574 | Gene names | Max, Myn | |||
|
Domain Architecture |

|
|||||
| Description | Protein max (Myc-associated factor X) (Protein myn) (Myc-binding novel HLH/LZ protein). | |||||
|
MAX_HUMAN
|
||||||
| NC score | 0.134367 (rank : 30) | θ value | θ > 10 (rank : 59) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P61244, P25912, P52163 | Gene names | MAX | |||
|
Domain Architecture |

|
|||||
| Description | Protein max (Myc-associated factor X). | |||||
|
CLOCK_MOUSE
|
||||||
| NC score | 0.132758 (rank : 31) | θ value | 3.0926 (rank : 25) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O08785 | Gene names | Clock | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (mCLOCK). | |||||
|
CLOCK_HUMAN
|
||||||
| NC score | 0.130633 (rank : 32) | θ value | 4.03905 (rank : 26) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O15516, O14516, Q9UIT8 | Gene names | CLOCK, KIAA0334 | |||
|
Domain Architecture |

|
|||||
| Description | Circadian locomoter output cycles protein kaput (hCLOCK). | |||||
|
NPAS2_HUMAN
|
||||||
| NC score | 0.129367 (rank : 33) | θ value | 5.27518 (rank : 30) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q99743, Q99629 | Gene names | NPAS2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2) (Member of PAS protein 4) (MOP4). | |||||
|
NPAS2_MOUSE
|
||||||
| NC score | 0.127927 (rank : 34) | θ value | 5.27518 (rank : 31) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P97460 | Gene names | Npas2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2). | |||||
|
EPAS1_HUMAN
|
||||||
| NC score | 0.078977 (rank : 35) | θ value | θ > 10 (rank : 48) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q99814, Q86VA2, Q99630 | Gene names | EPAS1, HIF2A | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Member of PAS protein 2) (MOP2) (Hypoxia-inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (HLF). | |||||
|
EPAS1_MOUSE
|
||||||
| NC score | 0.078190 (rank : 36) | θ value | θ > 10 (rank : 49) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P97481, O08787, O55046 | Gene names | Epas1, Hif2a | |||
|
Domain Architecture |

|
|||||
| Description | Endothelial PAS domain-containing protein 1 (EPAS-1) (Hypoxia- inducible factor 2 alpha) (HIF-2 alpha) (HIF2 alpha) (HIF-1 alpha-like factor) (MHLF) (HIF-related factor) (HRF). | |||||
|
HIF1A_HUMAN
|
||||||
| NC score | 0.075848 (rank : 37) | θ value | θ > 10 (rank : 54) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q16665, Q96PT9, Q9UPB1 | Gene names | HIF1A | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein) (Member of PAS protein 1) (MOP1). | |||||
|
HIF1A_MOUSE
|
||||||
| NC score | 0.075700 (rank : 38) | θ value | θ > 10 (rank : 55) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q61221, O08741, O08993, Q61664, Q61665, Q8C681, Q8CC19, Q8CCB6, Q8R385, Q9CYA8 | Gene names | Hif1a | |||
|
Domain Architecture |

|
|||||
| Description | Hypoxia-inducible factor 1 alpha (HIF-1 alpha) (HIF1 alpha) (ARNT- interacting protein). | |||||
|
HEY2_HUMAN
|
||||||
| NC score | 0.070499 (rank : 39) | θ value | θ > 10 (rank : 52) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9UBP5 | Gene names | HEY2, CHF1, GRL, HERP, HERP1, HRT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Cardiovascular helix-loop-helix factor 1) (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES- related repressor protein 2) (Protein gridlock homolog). | |||||
|
MYC_MOUSE
|
||||||
| NC score | 0.070245 (rank : 40) | θ value | θ > 10 (rank : 66) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P01108, P70247, Q3UM70, Q61422 | Gene names | Myc | |||
|
Domain Architecture |

|
|||||
| Description | Myc proto-oncogene protein (c-Myc) (Transcription factor p64). | |||||
|
SIM2_HUMAN
|
||||||
| NC score | 0.070143 (rank : 41) | θ value | θ > 10 (rank : 83) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q14190, O60766, Q15470, Q15471, Q15472, Q15473, Q16532 | Gene names | SIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2. | |||||
|
MYCL1_MOUSE
|
||||||
| NC score | 0.070082 (rank : 42) | θ value | θ > 10 (rank : 61) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P10166, Q5FWI7 | Gene names | Mycl1, Lmyc1, Mycl | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
SIM1_MOUSE
|
||||||
| NC score | 0.069764 (rank : 43) | θ value | θ > 10 (rank : 82) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q61045, O70284, P70183 | Gene names | Sim1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1 (mSIM1). | |||||
|
SIM1_HUMAN
|
||||||
| NC score | 0.069678 (rank : 44) | θ value | θ > 10 (rank : 81) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P81133, Q5TDP7 | Gene names | SIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 1. | |||||
|
SIM2_MOUSE
|
||||||
| NC score | 0.068799 (rank : 45) | θ value | θ > 10 (rank : 84) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q61079, O35391, Q61046, Q61904 | Gene names | Sim2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2 (SIM transcription factor) (mSIM). | |||||
|
WBS14_HUMAN
|
||||||
| NC score | 0.068592 (rank : 46) | θ value | 6.88961 (rank : 37) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9NP71, Q96E48, Q9BY03, Q9BY04, Q9BY05, Q9BY06, Q9Y2P3 | Gene names | MLXIPL, MIO, WBSCR14 | |||
|
Domain Architecture |

|
|||||
| Description | Williams-Beuren syndrome chromosome region 14 protein (WS basic-helix- loop-helix leucine zipper protein) (WS-bHLH) (Mlx interactor) (MLX- interacting protein-like). | |||||
|
HEY2_MOUSE
|
||||||
| NC score | 0.068575 (rank : 47) | θ value | θ > 10 (rank : 53) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9QUS4, Q3TZ99, Q8CD44 | Gene names | Hey2, Chf1, Herp, Herp1, Hesr2, Hrt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES-related repressor protein 2). | |||||
|
AHR_HUMAN
|
||||||
| NC score | 0.068199 (rank : 48) | θ value | θ > 10 (rank : 44) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P35869, Q13728, Q13803, Q13804 | Gene names | AHR | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
HEY1_MOUSE
|
||||||
| NC score | 0.068125 (rank : 49) | θ value | θ > 10 (rank : 51) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9WV93 | Gene names | Hey1, Hesr1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1). | |||||
|
SRBP1_HUMAN
|
||||||
| NC score | 0.067982 (rank : 50) | θ value | θ > 10 (rank : 85) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P36956, Q16062 | Gene names | SREBF1, SREBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
MYC_HUMAN
|
||||||
| NC score | 0.067639 (rank : 51) | θ value | θ > 10 (rank : 65) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P01106, P01107, Q14026 | Gene names | MYC | |||
|
Domain Architecture |

|
|||||
| Description | Myc proto-oncogene protein (c-Myc) (Transcription factor p64). | |||||
|
SRBP1_MOUSE
|
||||||
| NC score | 0.067114 (rank : 52) | θ value | θ > 10 (rank : 86) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9WTN3 | Gene names | Srebf1, Srebp1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
HEY1_HUMAN
|
||||||
| NC score | 0.065846 (rank : 53) | θ value | θ > 10 (rank : 50) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9Y5J3, Q5TZS3, Q9NYP4 | Gene names | HEY1, CHF2, HESR1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1) (Cardiovascular helix-loop-helix factor 2) (HES-related repressor protein 2) (HERP2). | |||||
|
MYCL1_HUMAN
|
||||||
| NC score | 0.065700 (rank : 54) | θ value | θ > 10 (rank : 60) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P12524, Q14897, Q5QPL0, Q5QPL1, Q9NUE9 | Gene names | MYCL1, LMYC, MYCL | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
NPAS3_MOUSE
|
||||||
| NC score | 0.065597 (rank : 55) | θ value | θ > 10 (rank : 74) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9QZQ0, Q9EQP4 | Gene names | Npas3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
AHR_MOUSE
|
||||||
| NC score | 0.065126 (rank : 56) | θ value | θ > 10 (rank : 45) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P30561, Q8QZX6, Q8R4S3, Q99P79, Q9QVY1 | Gene names | Ahr | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor precursor (Ah receptor) (AhR). | |||||
|
K0310_HUMAN
|
||||||
| NC score | 0.065014 (rank : 57) | θ value | 1.38821 (rank : 22) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O15027, Q5SXP0, Q5SXP1, Q8N347, Q96HP1 | Gene names | KIAA0310 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0310. | |||||
|
NPAS3_HUMAN
|
||||||
| NC score | 0.063824 (rank : 58) | θ value | θ > 10 (rank : 73) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8IXF0, Q86US6, Q86US7, Q8IXF2, Q9BY81, Q9H323, Q9Y4L8 | Gene names | NPAS3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
MYCN_HUMAN
|
||||||
| NC score | 0.062617 (rank : 59) | θ value | θ > 10 (rank : 62) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P04198, Q6LDT9 | Gene names | MYCN, NMYC | |||
|
Domain Architecture |

|
|||||
| Description | N-myc proto-oncogene protein. | |||||
|
NCOA1_MOUSE
|
||||||
| NC score | 0.062117 (rank : 60) | θ value | θ > 10 (rank : 68) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P70365, P70366, Q61202, Q8CBI9 | Gene names | Ncoa1, Src1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (Nuclear receptor coactivator protein 1) (mNRC-1). | |||||
|
MYCN_MOUSE
|
||||||
| NC score | 0.062015 (rank : 61) | θ value | θ > 10 (rank : 63) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P03966, Q61978 | Gene names | Mycn, Nmyc, Nmyc1 | |||
|
Domain Architecture |

|
|||||
| Description | N-myc proto-oncogene protein. | |||||
|
NCOA1_HUMAN
|
||||||
| NC score | 0.062012 (rank : 62) | θ value | θ > 10 (rank : 67) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q15788, O00150, O43792, O43793, Q13071, Q13420, Q6GVI5, Q7KYV3 | Gene names | NCOA1, SRC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 1 (EC 2.3.1.48) (NCoA-1) (Steroid receptor coactivator 1) (SRC-1) (RIP160) (Protein Hin-2) (NY-REN-52 antigen). | |||||
|
SRBP2_HUMAN
|
||||||
| NC score | 0.060946 (rank : 63) | θ value | θ > 10 (rank : 87) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q12772, Q9UH04 | Gene names | SREBF2, SREBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 2 (SREBP-2) (Sterol regulatory element-binding transcription factor 2). | |||||
|
NPAS1_MOUSE
|
||||||
| NC score | 0.060532 (rank : 64) | θ value | θ > 10 (rank : 72) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P97459 | Gene names | Npas1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1). | |||||
|
NPAS1_HUMAN
|
||||||
| NC score | 0.060381 (rank : 65) | θ value | θ > 10 (rank : 71) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q99742, Q99632, Q9BY83 | Gene names | NPAS1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 1 (Neuronal PAS1) (Member of PAS protein 5) (MOP5). | |||||
|
PER2_HUMAN
|
||||||
| NC score | 0.059229 (rank : 66) | θ value | θ > 10 (rank : 77) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O15055 | Gene names | PER2, KIAA0347 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2. | |||||
|
NCOA3_HUMAN
|
||||||
| NC score | 0.059109 (rank : 67) | θ value | θ > 10 (rank : 69) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y6Q9, Q5JYD9, Q5JYE0, Q9BR49, Q9UPC9, Q9UPG4, Q9UPG7 | Gene names | NCOA3, AIB1, RAC3, TRAM1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP-interacting protein) (pCIP). | |||||
|
PER3_MOUSE
|
||||||
| NC score | 0.059061 (rank : 68) | θ value | θ > 10 (rank : 80) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O70361 | Gene names | Per3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (mPER3). | |||||
|
PER2_MOUSE
|
||||||
| NC score | 0.058812 (rank : 69) | θ value | θ > 10 (rank : 78) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O54943, O54954 | Gene names | Per2 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2 (mPER2). | |||||
|
BHLH3_HUMAN
|
||||||
| NC score | 0.056871 (rank : 70) | θ value | θ > 10 (rank : 46) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9C0J9 | Gene names | BHLHB3, DEC2, SHARP1 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (hDEC2) (Enhancer-of-split and hairy-related protein 1) (SHARP-1). | |||||
|
PER1_MOUSE
|
||||||
| NC score | 0.056786 (rank : 71) | θ value | θ > 10 (rank : 76) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O35973 | Gene names | Per1, Per, Rigui | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (mPER) (M-Rigui). | |||||
|
WBS14_MOUSE
|
||||||
| NC score | 0.056333 (rank : 72) | θ value | θ > 10 (rank : 89) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q99MZ3, Q99MY9, Q99MZ0, Q99MZ1, Q99MZ2, Q9JLM5 | Gene names | Mlxipl, Mio, Wbscr14 | |||
|
Domain Architecture |

|
|||||
| Description | Williams-Beuren syndrome chromosome region 14 protein homolog (Mlx interactor) (MLX-interacting protein-like). | |||||
|
PER1_HUMAN
|
||||||
| NC score | 0.055475 (rank : 73) | θ value | θ > 10 (rank : 75) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O15534 | Gene names | PER1, KIAA0482, PER, RIGUI | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (hPER). | |||||
|
PER3_HUMAN
|
||||||
| NC score | 0.055197 (rank : 74) | θ value | θ > 10 (rank : 79) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P56645, Q5H8X4, Q969K6, Q96S77, Q96S78, Q9C0J3, Q9NSP9, Q9UGU8 | Gene names | PER3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (hPER3). | |||||
|
TFAP4_HUMAN
|
||||||
| NC score | 0.054856 (rank : 75) | θ value | θ > 10 (rank : 88) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q01664, O60409 | Gene names | TFAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor AP-4 (Activating enhancer-binding protein 4). | |||||
|
MYCS_MOUSE
|
||||||
| NC score | 0.054725 (rank : 76) | θ value | θ > 10 (rank : 64) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9Z304 | Gene names | Mycs | |||
|
Domain Architecture |

|
|||||
| Description | Protein S-myc. | |||||
|
BHLH3_MOUSE
|
||||||
| NC score | 0.054346 (rank : 77) | θ value | θ > 10 (rank : 47) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q99PV5 | Gene names | Bhlhb3, Dec2 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (mDEC2). | |||||
|
NCOA3_MOUSE
|
||||||
| NC score | 0.051627 (rank : 78) | θ value | θ > 10 (rank : 70) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O09000, Q9CRD5 | Gene names | Ncoa3, Aib1, Pcip, Rac3, Tram1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein homolog) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP- interacting protein) (p/CIP) (pCIP). | |||||
|
PHAR1_MOUSE
|
||||||
| NC score | 0.038505 (rank : 79) | θ value | 4.03905 (rank : 28) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q2M3X8, Q80VL9, Q8C873 | Gene names | Phactr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatase and actin regulator 1. | |||||
|
MAP1B_MOUSE
|
||||||
| NC score | 0.033651 (rank : 80) | θ value | 2.36792 (rank : 24) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 977 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P14873 | Gene names | Map1b, Mtap1b, Mtap5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) (MAP1.2) (MAP1(X)) [Contains: MAP1 light chain LC1]. | |||||
|
PHAR1_HUMAN
|
||||||
| NC score | 0.032707 (rank : 81) | θ value | 8.99809 (rank : 42) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9C0D0, Q3MJ93, Q5JSJ2 | Gene names | PHACTR1, KIAA1733, RPEL1 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatase and actin regulator 1. | |||||
|
DCP1A_HUMAN
|
||||||
| NC score | 0.027546 (rank : 82) | θ value | 6.88961 (rank : 33) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NPI6 | Gene names | DCP1A, SMIF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | mRNA decapping enzyme 1A (EC 3.-.-.-) (Transcription factor SMIF) (Smad4-interacting transcriptional co-activator). | |||||
|
PDXL2_MOUSE
|
||||||
| NC score | 0.025818 (rank : 83) | θ value | 6.88961 (rank : 35) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CAE9, Q8CFW3 | Gene names | Podxl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Podocalyxin-like protein 2 precursor (Endoglycan). | |||||
|
IER5_MOUSE
|
||||||
| NC score | 0.023890 (rank : 84) | θ value | 8.99809 (rank : 38) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O89113, Q8CGI0 | Gene names | Ier5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Immediate early response gene 5 protein. | |||||
|
HAIR_HUMAN
|
||||||
| NC score | 0.022914 (rank : 85) | θ value | 5.27518 (rank : 29) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O43593, Q96H33, Q9NPE1 | Gene names | HR | |||
|
Domain Architecture |

|
|||||
| Description | Protein hairless. | |||||
|
TTC16_HUMAN
|
||||||
| NC score | 0.022821 (rank : 86) | θ value | 5.27518 (rank : 32) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NEE8, Q5JU66, Q96M72 | Gene names | TTC16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tetratricopeptide repeat protein 16 (TPR repeat protein 16). | |||||
|
NALDL_HUMAN
|
||||||
| NC score | 0.022089 (rank : 87) | θ value | 4.03905 (rank : 27) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UQQ1, O43176 | Gene names | NAALADL1, NAALADASEL, NAALADL | |||
|
Domain Architecture |

|
|||||
| Description | N-acetylated-alpha-linked acidic dipeptidase-like protein (EC 3.4.17.21) (NAALADase L) (Ileal dipeptidylpeptidase) (100 kDa ileum brush border membrane protein) (I100). | |||||
|
MAP1A_HUMAN
|
||||||
| NC score | 0.020804 (rank : 88) | θ value | 6.88961 (rank : 34) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P78559, O95643, Q12973, Q15882, Q9UJT4 | Gene names | MAP1A | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 1A (MAP 1A) (Proliferation-related protein p80) [Contains: MAP1 light chain LC2]. | |||||
|
PLRG1_HUMAN
|
||||||
| NC score | 0.007951 (rank : 89) | θ value | 6.88961 (rank : 36) | |||
| Query Neighborhood Hits | 43 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O43660, Q8WUD8 | Gene names | PLRG1 | |||
|
Domain Architecture |

|
|||||
| Description | Pleiotropic regulator 1. | |||||