Please be patient as the page loads
|
MYCN_HUMAN
|
||||||
| SwissProt Accessions | P04198, Q6LDT9 | Gene names | MYCN, NMYC | |||
|
Domain Architecture |

|
|||||
| Description | N-myc proto-oncogene protein. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
MYCN_HUMAN
|
||||||
| θ value | 2.33393e-165 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 127 | |
| SwissProt Accessions | P04198, Q6LDT9 | Gene names | MYCN, NMYC | |||
|
Domain Architecture |

|
|||||
| Description | N-myc proto-oncogene protein. | |||||
|
MYCN_MOUSE
|
||||||
| θ value | 4.72459e-150 (rank : 2) | NC score | 0.981433 (rank : 2) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 86 | |
| SwissProt Accessions | P03966, Q61978 | Gene names | Mycn, Nmyc, Nmyc1 | |||
|
Domain Architecture |

|
|||||
| Description | N-myc proto-oncogene protein. | |||||
|
MYCS_MOUSE
|
||||||
| θ value | 4.77967e-110 (rank : 3) | NC score | 0.950047 (rank : 3) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9Z304 | Gene names | Mycs | |||
|
Domain Architecture |
|
|||||
| Description | Protein S-myc. | |||||
|
MYC_MOUSE
|
||||||
| θ value | 8.06329e-33 (rank : 4) | NC score | 0.827060 (rank : 4) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P01108, P70247, Q3UM70, Q61422 | Gene names | Myc | |||
|
Domain Architecture |
|
|||||
| Description | Myc proto-oncogene protein (c-Myc) (Transcription factor p64). | |||||
|
MYC_HUMAN
|
||||||
| θ value | 1.37539e-32 (rank : 5) | NC score | 0.820426 (rank : 5) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P01106, P01107, Q14026 | Gene names | MYC | |||
|
Domain Architecture |

|
|||||
| Description | Myc proto-oncogene protein (c-Myc) (Transcription factor p64). | |||||
|
MYCL1_HUMAN
|
||||||
| θ value | 1.24688e-25 (rank : 6) | NC score | 0.778661 (rank : 6) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P12524, Q14897, Q5QPL0, Q5QPL1, Q9NUE9 | Gene names | MYCL1, LMYC, MYCL | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
MYCL2_HUMAN
|
||||||
| θ value | 7.56453e-23 (rank : 7) | NC score | 0.743515 (rank : 7) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P12525 | Gene names | MYCL2 | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-2 protein. | |||||
|
MYCL1_MOUSE
|
||||||
| θ value | 1.33526e-19 (rank : 8) | NC score | 0.742050 (rank : 8) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P10166, Q5FWI7 | Gene names | Mycl1, Lmyc1, Mycl | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
MYCB_MOUSE
|
||||||
| θ value | 2.43908e-13 (rank : 9) | NC score | 0.689254 (rank : 9) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6P8Z1, Q9D6E3 | Gene names | Mycb, Bmyc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein B-Myc. | |||||
|
MAX_HUMAN
|
||||||
| θ value | 2.79066e-09 (rank : 10) | NC score | 0.621615 (rank : 10) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P61244, P25912, P52163 | Gene names | MAX | |||
|
Domain Architecture |

|
|||||
| Description | Protein max (Myc-associated factor X). | |||||
|
MAX_MOUSE
|
||||||
| θ value | 8.11959e-09 (rank : 11) | NC score | 0.613570 (rank : 11) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P28574 | Gene names | Max, Myn | |||
|
Domain Architecture |

|
|||||
| Description | Protein max (Myc-associated factor X) (Protein myn) (Myc-binding novel HLH/LZ protein). | |||||
|
FIGLA_MOUSE
|
||||||
| θ value | 6.43352e-06 (rank : 12) | NC score | 0.338721 (rank : 16) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O55208 | Gene names | Figla | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Factor in the germline alpha (Transcription factor FIGa) (FIGalpha). | |||||
|
NFH_MOUSE
|
||||||
| θ value | 2.44474e-05 (rank : 13) | NC score | 0.038134 (rank : 138) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
MNT_HUMAN
|
||||||
| θ value | 7.1131e-05 (rank : 14) | NC score | 0.304686 (rank : 20) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 496 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q99583 | Gene names | MNT, ROX | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
MNT_MOUSE
|
||||||
| θ value | 7.1131e-05 (rank : 15) | NC score | 0.312423 (rank : 19) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 432 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O08789, P97349 | Gene names | Mnt, Rox | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
SRBP2_HUMAN
|
||||||
| θ value | 7.1131e-05 (rank : 16) | NC score | 0.290005 (rank : 23) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q12772, Q9UH04 | Gene names | SREBF2, SREBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 2 (SREBP-2) (Sterol regulatory element-binding transcription factor 2). | |||||
|
FIGLA_HUMAN
|
||||||
| θ value | 0.000602161 (rank : 17) | NC score | 0.320982 (rank : 17) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q6QHK4 | Gene names | FIGLA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Factor in the germline alpha (Transcription factor FIGa) (FIGalpha). | |||||
|
MITF_HUMAN
|
||||||
| θ value | 0.000602161 (rank : 18) | NC score | 0.288869 (rank : 25) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O75030, Q14841, Q9P2V0, Q9P2V1, Q9P2V2, Q9P2Y8 | Gene names | MITF | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
USF2_HUMAN
|
||||||
| θ value | 0.000602161 (rank : 19) | NC score | 0.361386 (rank : 12) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q15853, O00671, O00709, Q05750, Q07952, Q15851, Q15852 | Gene names | USF2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (FOS- interacting protein) (FIP) (Major late transcription factor 2). | |||||
|
USF2_MOUSE
|
||||||
| θ value | 0.000602161 (rank : 20) | NC score | 0.359233 (rank : 13) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q64705, Q3UIL2, Q8VE59 | Gene names | Usf2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (Major late transcription factor 2). | |||||
|
MITF_MOUSE
|
||||||
| θ value | 0.00102713 (rank : 21) | NC score | 0.286236 (rank : 27) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q08874, O08885, O88203, Q08843, Q60781, Q60782, Q9JIJ0, Q9JIJ1, Q9JIJ2, Q9JIJ3, Q9JIJ4, Q9JIJ5, Q9JIJ6, Q9JKX9 | Gene names | Mitf, Bw, Mi, Vit | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
USF1_HUMAN
|
||||||
| θ value | 0.00134147 (rank : 22) | NC score | 0.356055 (rank : 15) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P22415 | Gene names | USF1, USF | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF1_MOUSE
|
||||||
| θ value | 0.00134147 (rank : 23) | NC score | 0.358516 (rank : 14) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q61069 | Gene names | Usf1, Usf | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
TFE3_HUMAN
|
||||||
| θ value | 0.00298849 (rank : 24) | NC score | 0.293634 (rank : 22) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P19532, Q92757, Q92758, Q99964 | Gene names | TFE3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E3. | |||||
|
WBS14_HUMAN
|
||||||
| θ value | 0.00298849 (rank : 25) | NC score | 0.175194 (rank : 48) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9NP71, Q96E48, Q9BY03, Q9BY04, Q9BY05, Q9BY06, Q9Y2P3 | Gene names | MLXIPL, MIO, WBSCR14 | |||
|
Domain Architecture |
|
|||||
| Description | Williams-Beuren syndrome chromosome region 14 protein (WS basic-helix- loop-helix leucine zipper protein) (WS-bHLH) (Mlx interactor) (MLX- interacting protein-like). | |||||
|
GRIN1_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 26) | NC score | 0.058631 (rank : 119) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q7Z2K8, Q8ND74, Q96PZ4 | Gene names | GPRIN1, KIAA1893 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
ASCL1_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 27) | NC score | 0.250606 (rank : 32) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P50553, Q9BQ30 | Gene names | ASCL1, ASH1 | |||
|
Domain Architecture |

|
|||||
| Description | Achaete-scute homolog 1 (HASH1). | |||||
|
MAD4_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 28) | NC score | 0.289872 (rank : 24) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q14582, Q5TZX4 | Gene names | MXD4, MAD4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-interacting transcriptional repressor MAD4 (Max-associated protein 4) (MAX dimerization protein 4). | |||||
|
TFAP4_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 29) | NC score | 0.314233 (rank : 18) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q01664, O60409 | Gene names | TFAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor AP-4 (Activating enhancer-binding protein 4). | |||||
|
ASCL1_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 30) | NC score | 0.238249 (rank : 37) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q02067 | Gene names | Ascl1, Ash1, Mash-1, Mash1 | |||
|
Domain Architecture |

|
|||||
| Description | Achaete-scute homolog 1 (Mash-1). | |||||
|
BHLH8_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 31) | NC score | 0.241005 (rank : 35) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q7RTS1 | Gene names | BHLHB8, MIST1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Class B basic helix-loop-helix protein 8 (bHLHB8) (Muscle, intestine and stomach expression 1) (MIST-1). | |||||
|
BHLH8_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 32) | NC score | 0.247501 (rank : 34) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9QYC3, Q9QYE4 | Gene names | Bhlhb8, Mist1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Class B basic helix-loop-helix protein 8 (bHLHB8) (Muscle, intestine and stomach expression 1) (MIST-1). | |||||
|
PTF1A_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 33) | NC score | 0.187526 (rank : 45) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9QX98, Q9QYF5, Q9QYF6 | Gene names | Ptf1a, Ptf1p48 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pancreas transcription factor 1 subunit alpha (Pancreas-specific transcription factor 1a) (bHLH transcription factor p48) (p48 DNA- binding subunit of transcription factor PTF1) (PTF1-p48). | |||||
|
SRBP1_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 34) | NC score | 0.250968 (rank : 31) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P36956, Q16062 | Gene names | SREBF1, SREBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
MAD4_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 35) | NC score | 0.238531 (rank : 36) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q60948 | Gene names | Mxd4, Mad4 | |||
|
Domain Architecture |
|
|||||
| Description | Max-interacting transcriptional repressor MAD4 (Max-associated protein 4) (MAX dimerization protein 4). | |||||
|
MXI1_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 36) | NC score | 0.300630 (rank : 21) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P50540 | Gene names | Mxi1 | |||
|
Domain Architecture |
|
|||||
| Description | MAX-interacting protein 1 (Protein MXI1). | |||||
|
OGFR_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 37) | NC score | 0.038704 (rank : 137) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
SRBP1_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 38) | NC score | 0.237097 (rank : 38) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9WTN3 | Gene names | Srebf1, Srebp1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
ARNT_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 39) | NC score | 0.140270 (rank : 54) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P27540 | Gene names | ARNT | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
ARNT_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 40) | NC score | 0.135805 (rank : 59) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P53762, Q60661 | Gene names | Arnt | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
ASCL3_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 41) | NC score | 0.173750 (rank : 49) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9NQ33, Q8WYQ6 | Gene names | ASCL3, HASH3, SGN1 | |||
|
Domain Architecture |
|
|||||
| Description | Achaete-scute homolog 3 (bHLH transcriptional regulator Sgn-1). | |||||
|
MLX_HUMAN
|
||||||
| θ value | 0.125558 (rank : 42) | NC score | 0.268579 (rank : 29) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UH92, Q53XM6, Q96FL2, Q9H2V0, Q9H2V1, Q9H2V2, Q9NXN3 | Gene names | MLX, TCFL4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
MLX_MOUSE
|
||||||
| θ value | 0.125558 (rank : 43) | NC score | 0.270075 (rank : 28) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O08609, Q9EQJ7, Q9EQJ8 | Gene names | Mlx, Tcfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
MXI1_HUMAN
|
||||||
| θ value | 0.125558 (rank : 44) | NC score | 0.288526 (rank : 26) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P50539, Q15887 | Gene names | MXI1 | |||
|
Domain Architecture |
|
|||||
| Description | MAX-interacting protein 1 (Protein MXI1). | |||||
|
NDF2_HUMAN
|
||||||
| θ value | 0.125558 (rank : 45) | NC score | 0.120114 (rank : 66) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q15784, Q8TBI7, Q9UQC6 | Gene names | NEUROD2, NDRF | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic differentiation factor 2 (NeuroD2) (NeuroD-related factor) (NDRF). | |||||
|
NDF2_MOUSE
|
||||||
| θ value | 0.125558 (rank : 46) | NC score | 0.119961 (rank : 67) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q62414, Q61952, Q925V5 | Gene names | Neurod2, Ndrf | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic differentiation factor 2 (NeuroD2) (NeuroD-related factor) (NDRF). | |||||
|
PTF1A_HUMAN
|
||||||
| θ value | 0.163984 (rank : 47) | NC score | 0.184899 (rank : 47) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q7RTS3, Q9HC25 | Gene names | PTF1A, PTF1P48 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pancreas transcription factor 1 subunit alpha (Pancreas-specific transcription factor 1a) (bHLH transcription factor p48) (p48 DNA- binding subunit of transcription factor PTF1) (PTF1-p48). | |||||
|
ARNT2_HUMAN
|
||||||
| θ value | 0.21417 (rank : 48) | NC score | 0.127967 (rank : 63) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9HBZ2, O15024 | Gene names | ARNT2, KIAA0307 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
ARNT2_MOUSE
|
||||||
| θ value | 0.21417 (rank : 49) | NC score | 0.128427 (rank : 61) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q61324 | Gene names | Arnt2 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
ATS13_HUMAN
|
||||||
| θ value | 0.21417 (rank : 50) | NC score | 0.014809 (rank : 161) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q76LX8, Q6UY16, Q710F6, Q711T8, Q96L37, Q9H0G3 | Gene names | ADAMTS13, C9orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAMTS-13 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase with thrombospondin motifs 13) (ADAM-TS 13) (ADAM-TS13) (von Willebrand factor-cleaving protease) (vWF-cleaving protease) (vWF-CP). | |||||
|
DEND_MOUSE
|
||||||
| θ value | 0.21417 (rank : 51) | NC score | 0.046402 (rank : 134) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80TS7 | Gene names | Ddn, Kiaa0749 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dendrin. | |||||
|
ASCL3_MOUSE
|
||||||
| θ value | 0.279714 (rank : 52) | NC score | 0.160347 (rank : 52) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9JJR7 | Gene names | Ascl3, Mash3, Sgn1 | |||
|
Domain Architecture |

|
|||||
| Description | Achaete-scute homolog 3 (bHLH transcriptional regulator Sgn-1) (Mash- 3). | |||||
|
ID1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 53) | NC score | 0.130465 (rank : 60) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P41134, O00651, O00652, Q16371, Q16377, Q5TE66, Q5TE67, Q969Z7, Q9H0Z5, Q9H109 | Gene names | ID1, ID | |||
|
Domain Architecture |
|
|||||
| Description | DNA-binding protein inhibitor ID-1 (Inhibitor of DNA binding 1). | |||||
|
ID1_MOUSE
|
||||||
| θ value | 0.279714 (rank : 54) | NC score | 0.128009 (rank : 62) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P20067, Q61101, Q9D897 | Gene names | Id1, Id, Id-1, Idb1 | |||
|
Domain Architecture |
|
|||||
| Description | DNA-binding protein inhibitor ID-1 (Inhibitor of DNA binding 1). | |||||
|
ITF2_HUMAN
|
||||||
| θ value | 0.279714 (rank : 55) | NC score | 0.085149 (rank : 89) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P15884, Q15439, Q15440 | Gene names | TCF4, ITF2, SEF2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 4 (Immunoglobulin transcription factor 2) (ITF-2) (SL3-3 enhancer factor 2) (SEF-2). | |||||
|
ITF2_MOUSE
|
||||||
| θ value | 0.279714 (rank : 56) | NC score | 0.086380 (rank : 88) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q60722, Q60721, Q62211, Q64072, Q80UE8 | Gene names | Tcf4, Itf2, Sef2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 4 (Immunoglobulin transcription factor 2) (ITF-2) (MITF-2) (SL3-3 enhancer factor 2) (SEF-2) (Class A helix-loop-helix transcription factor ME2). | |||||
|
PTN13_MOUSE
|
||||||
| θ value | 0.279714 (rank : 57) | NC score | 0.010609 (rank : 173) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 426 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q64512, Q61494, Q62135, Q64499 | Gene names | Ptpn13, Ptp14 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 13 (EC 3.1.3.48) (Protein tyrosine phosphatase PTP-BL) (Protein-tyrosine phosphatase RIP) (protein tyrosine phosphatase DPZPTP) (PTP36). | |||||
|
ASCL2_MOUSE
|
||||||
| θ value | 0.365318 (rank : 58) | NC score | 0.219993 (rank : 41) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O35885, Q9WUJ7 | Gene names | Ascl2, Mash2 | |||
|
Domain Architecture |

|
|||||
| Description | Achaete-scute homolog 2 (Mash-2). | |||||
|
GDIA_HUMAN
|
||||||
| θ value | 0.47712 (rank : 59) | NC score | 0.063529 (rank : 113) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P31150, P50394, Q7Z2G6, Q7Z2G9, Q7Z2H5, Q7Z2I6 | Gene names | GDI1, OPHN2, RABGDIA, XAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Rab GDP dissociation inhibitor alpha (Rab GDI alpha) (Guanosine diphosphate dissociation inhibitor 1) (GDI-1) (XAP-4) (Oligophrenin- 2). | |||||
|
HAND2_HUMAN
|
||||||
| θ value | 0.47712 (rank : 60) | NC score | 0.194363 (rank : 43) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P61296, O95300, O95301, P97833 | Gene names | HAND2, DHAND | |||
|
Domain Architecture |
|
|||||
| Description | Heart- and neural crest derivatives-expressed protein 2 (Deciduum, heart, autonomic nervous system and neural crest derivatives-expressed protein 2) (dHAND). | |||||
|
HAND2_MOUSE
|
||||||
| θ value | 0.47712 (rank : 61) | NC score | 0.194363 (rank : 44) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q61039, Q61100 | Gene names | Hand2, Dhand, Hed, Thing2 | |||
|
Domain Architecture |

|
|||||
| Description | Heart- and neural crest derivatives-expressed protein 2 (Deciduum, heart, autonomic nervous system and neural crest derivatives-expressed protein 2) (dHAND) (Helix-loop-helix transcription factor expressed in embryo and deciduum) (Thing-2). | |||||
|
HTF4_HUMAN
|
||||||
| θ value | 0.47712 (rank : 62) | NC score | 0.078793 (rank : 97) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q99081 | Gene names | TCF12, HEB, HTF4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 12 (Transcription factor HTF-4) (E-box-binding protein) (DNA-binding protein HTF4). | |||||
|
HAND1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 63) | NC score | 0.166016 (rank : 51) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O96004 | Gene names | HAND1, EHAND | |||
|
Domain Architecture |
|
|||||
| Description | Heart- and neural crest derivatives-expressed protein 1 (Extraembryonic tissues, heart, autonomic nervous system and neural crest derivatives-expressed protein 1) (eHAND). | |||||
|
HAND1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 64) | NC score | 0.171338 (rank : 50) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q64279, Q61099 | Gene names | Hand1, Ehand, Hxt, Thing1 | |||
|
Domain Architecture |
|
|||||
| Description | Heart- and neural crest derivatives-expressed protein 1 (Extraembryonic tissues, heart, autonomic nervous system and neural crest derivatives-expressed protein 1) (eHAND) (Helix-loop-helix transcription factor expressed in extraembryonic mesoderm and trophoblast) (Thing-1) (Th1). | |||||
|
HTF4_MOUSE
|
||||||
| θ value | 0.62314 (rank : 65) | NC score | 0.077577 (rank : 99) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q61286 | Gene names | Tcf12, Alf1, Meb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 12 (Transcription factor HTF-4) (E-box-binding protein) (DNA-binding protein HTF4) (Class A helix-loop-helix transcription factor ME1). | |||||
|
NOL6_MOUSE
|
||||||
| θ value | 0.62314 (rank : 66) | NC score | 0.032562 (rank : 140) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R5K4, Q6PAN8, Q8C6V4, Q8R5K3, Q8VCG0, Q8WTY7 | Gene names | Nol6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleolar protein 6 (Nucleolar RNA-associated protein) (Nrap). | |||||
|
EPN3_MOUSE
|
||||||
| θ value | 0.813845 (rank : 67) | NC score | 0.027184 (rank : 145) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q91W69, Q9CV55 | Gene names | Epn3 | |||
|
Domain Architecture |
|
|||||
| Description | Epsin-3 (EPS-15-interacting protein 3). | |||||
|
GDIA_MOUSE
|
||||||
| θ value | 0.813845 (rank : 68) | NC score | 0.060480 (rank : 117) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P50396, Q8VHM3, Q91Y71, Q96CX5 | Gene names | Gdi1, Rabgdia | |||
|
Domain Architecture |
|
|||||
| Description | Rab GDP dissociation inhibitor alpha (Rab GDI alpha) (Guanosine diphosphate dissociation inhibitor 1) (GDI-1). | |||||
|
GDIB_MOUSE
|
||||||
| θ value | 0.813845 (rank : 69) | NC score | 0.060205 (rank : 118) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61598, Q541Z9, Q8C530, Q9D8M9 | Gene names | Gdi2, Gdi3 | |||
|
Domain Architecture |
|
|||||
| Description | Rab GDP dissociation inhibitor beta (Rab GDI beta) (Guanosine diphosphate dissociation inhibitor 2) (GDI-2) (GDI-3). | |||||
|
MAD1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 70) | NC score | 0.208077 (rank : 42) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P50538, Q60798, Q61825 | Gene names | Mxd1, Mad, Mad1 | |||
|
Domain Architecture |

|
|||||
| Description | MAD protein (MAX dimerizer) (MAX dimerization protein 1). | |||||
|
TFEB_HUMAN
|
||||||
| θ value | 0.813845 (rank : 71) | NC score | 0.231732 (rank : 40) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P19484, Q709B3, Q7Z6P9, Q9BRJ5, Q9UJD8 | Gene names | TFEB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
TFEB_MOUSE
|
||||||
| θ value | 0.813845 (rank : 72) | NC score | 0.232756 (rank : 39) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9R210 | Gene names | Tfeb, Tcfeb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
ZFP28_HUMAN
|
||||||
| θ value | 0.813845 (rank : 73) | NC score | -0.000950 (rank : 186) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 843 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8NHY6, Q9BY30, Q9P2B6 | Gene names | ZFP28, KIAA1431 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger protein 28 homolog (Zfp-28) (Krueppel-like zinc finger factor X6). | |||||
|
CS007_MOUSE
|
||||||
| θ value | 1.06291 (rank : 74) | NC score | 0.031409 (rank : 141) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 464 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6ZPZ3 | Gene names | Kiaa1064 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein C19orf7 homolog. | |||||
|
NGN3_HUMAN
|
||||||
| θ value | 1.06291 (rank : 75) | NC score | 0.115057 (rank : 68) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9Y4Z2, Q9BY24 | Gene names | NEUROG3, NGN3 | |||
|
Domain Architecture |
|
|||||
| Description | Neurogenin 3. | |||||
|
ACINU_HUMAN
|
||||||
| θ value | 1.38821 (rank : 76) | NC score | 0.042919 (rank : 135) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9UKV3, O75158, Q9UG91, Q9UKV1, Q9UKV2 | Gene names | ACIN1, ACINUS, KIAA0670 | |||
|
Domain Architecture |

|
|||||
| Description | Apoptotic chromatin condensation inducer in the nucleus (Acinus). | |||||
|
ASCL2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 77) | NC score | 0.185394 (rank : 46) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q99929, Q9UM68 | Gene names | ASCL2 | |||
|
Domain Architecture |
|
|||||
| Description | Achaete-scute homolog 2 (Mash2) (HASH2). | |||||
|
FGFP3_HUMAN
|
||||||
| θ value | 1.38821 (rank : 78) | NC score | 0.039515 (rank : 136) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8TAT2, Q8NBN0 | Gene names | FGFBP3, C10orf13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fibroblast growth factor-binding protein 3 precursor (FGF-binding protein 3) (FGF-BP3) (FGFBP-3). | |||||
|
MAD3_MOUSE
|
||||||
| θ value | 1.38821 (rank : 79) | NC score | 0.248938 (rank : 33) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q80US8, Q60947 | Gene names | Mxd3, Mad3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Max-interacting transcriptional repressor MAD3 (Max-associated protein 3) (MAX dimerization protein 3). | |||||
|
MYH6_HUMAN
|
||||||
| θ value | 1.38821 (rank : 80) | NC score | 0.013277 (rank : 165) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1426 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P13533, Q13943, Q14906, Q14907 | Gene names | MYH6, MYHCA | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
PHLPP_HUMAN
|
||||||
| θ value | 1.81305 (rank : 81) | NC score | 0.014047 (rank : 163) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 604 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O60346, Q641Q7, Q6P4C4, Q6PJI6, Q86TN6, Q96FK2, Q9NUY1 | Gene names | PHLPP, KIAA0606, PLEKHE1, SCOP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH domain leucine-rich repeat-containing protein phosphatase (EC 3.1.3.16) (PH domain leucine-rich repeat protein phosphatase) (Pleckstrin homology domain-containing family E protein 1) (Suprachiasmatic nucleus circadian oscillatory protein) (hSCOP). | |||||
|
CASP_MOUSE
|
||||||
| θ value | 2.36792 (rank : 82) | NC score | 0.020284 (rank : 152) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1152 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P70403, Q7TQJ4, Q91WN6 | Gene names | Cutl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein CASP. | |||||
|
CUTL1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 83) | NC score | 0.022399 (rank : 150) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P53564, O08994, P70301, Q571L6, Q91ZD2 | Gene names | Cutl1, Cux, Cux1, Kiaa4047 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP) (Homeobox protein Cux). | |||||
|
GDIB_HUMAN
|
||||||
| θ value | 2.36792 (rank : 84) | NC score | 0.051675 (rank : 129) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P50395, O43928, Q9UQM6 | Gene names | GDI2, RABGDIB | |||
|
Domain Architecture |
|
|||||
| Description | Rab GDP dissociation inhibitor beta (Rab GDI beta) (Guanosine diphosphate dissociation inhibitor 2) (GDI-2). | |||||
|
MYH7_HUMAN
|
||||||
| θ value | 2.36792 (rank : 85) | NC score | 0.013213 (rank : 166) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P12883, Q14836, Q14837, Q14904, Q16579, Q92679, Q9H1D5, Q9UMM8 | Gene names | MYH7, MYHCB | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle beta isoform (MyHC-beta). | |||||
|
PEPL_MOUSE
|
||||||
| θ value | 2.36792 (rank : 86) | NC score | 0.012843 (rank : 167) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1105 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9R269, O70231, Q9CUT1, Q9JLZ7 | Gene names | Ppl | |||
|
Domain Architecture |

|
|||||
| Description | Periplakin. | |||||
|
SMC4_HUMAN
|
||||||
| θ value | 2.36792 (rank : 87) | NC score | 0.014867 (rank : 160) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 937 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9NTJ3, O95752, Q8NDL4, Q9UNT9 | Gene names | SMC4L1, CAPC, SMC4 | |||
|
Domain Architecture |
|
|||||
| Description | Structural maintenance of chromosomes 4-like 1 protein (Chromosome- associated polypeptide C) (hCAP-C) (XCAP-C homolog). | |||||
|
CASP_HUMAN
|
||||||
| θ value | 3.0926 (rank : 88) | NC score | 0.019595 (rank : 153) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1131 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q13948, Q53GU9, Q8TBS3 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein CASP. | |||||
|
CDS1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 89) | NC score | 0.015286 (rank : 158) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q92903, O00163 | Gene names | CDS1, CDS | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidate cytidylyltransferase 1 (EC 2.7.7.41) (CDP-diglyceride synthetase 1) (CDP-diglyceride pyrophosphorylase 1) (CDP- diacylglycerol synthase 1) (CDS 1) (CTP:phosphatidate cytidylyltransferase 1) (CDP-DAG synthase 1) (CDP-DG synthetase 1). | |||||
|
CUTL1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 90) | NC score | 0.023569 (rank : 149) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1574 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P39880, Q6NYH4, Q9UEV5 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP). | |||||
|
FOG1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 91) | NC score | 0.008176 (rank : 176) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1237 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O35615 | Gene names | Zfpm1, Fog, Fog1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein ZFPM1 (Zinc finger protein multitype 1) (Friend of GATA protein 1) (Friend of GATA-1) (FOG-1). | |||||
|
MAD3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 92) | NC score | 0.256197 (rank : 30) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9BW11, Q53HK1, Q7Z4Y0, Q8NDJ7, Q96ME3 | Gene names | MXD3, MAD3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Max-interacting transcriptional repressor MAD3 (Max-associated protein 3) (MAX dimerization protein 3) (Myx). | |||||
|
P80C_HUMAN
|
||||||
| θ value | 3.0926 (rank : 93) | NC score | 0.032952 (rank : 139) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P38432 | Gene names | COIL, CLN80 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coilin (p80). | |||||
|
SPIB_MOUSE
|
||||||
| θ value | 3.0926 (rank : 94) | NC score | 0.027163 (rank : 146) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O35906, O35907, O35909, O55199 | Gene names | Spib | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor Spi-B. | |||||
|
CAMKV_HUMAN
|
||||||
| θ value | 4.03905 (rank : 95) | NC score | -0.000340 (rank : 185) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1084 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8NCB2, Q6FIB8, Q8NBS8, Q8NC85, Q8NDU4, Q8WTT8, Q9BQC9, Q9H0Q5 | Gene names | CAMKV | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CaM kinase-like vesicle-associated protein. | |||||
|
K0310_HUMAN
|
||||||
| θ value | 4.03905 (rank : 96) | NC score | 0.029809 (rank : 143) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O15027, Q5SXP0, Q5SXP1, Q8N347, Q96HP1 | Gene names | KIAA0310 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0310. | |||||
|
MYH6_MOUSE
|
||||||
| θ value | 4.03905 (rank : 97) | NC score | 0.012532 (rank : 168) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1501 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q02566, Q64258, Q64738 | Gene names | Myh6, Myhca | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
ZN142_HUMAN
|
||||||
| θ value | 4.03905 (rank : 98) | NC score | 0.000066 (rank : 184) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1007 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P52746, Q92510 | Gene names | ZNF142, KIAA0236 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 142 (HA4654). | |||||
|
HXB3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 99) | NC score | 0.001649 (rank : 182) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 493 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P09026, P10285, Q4PJ06, Q61680 | Gene names | Hoxb3, Hox-2.7, Hoxb-3 | |||
|
Domain Architecture |
|
|||||
| Description | Homeobox protein Hox-B3 (Hox-2.7) (Homeobox protein MH-23). | |||||
|
RHG05_HUMAN
|
||||||
| θ value | 5.27518 (rank : 100) | NC score | 0.015626 (rank : 157) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q13017 | Gene names | ARHGAP5, RHOGAP5 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 5 (p190-B). | |||||
|
RHG05_MOUSE
|
||||||
| θ value | 5.27518 (rank : 101) | NC score | 0.015180 (rank : 159) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P97393 | Gene names | Arhgap5, Rhogap5 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 5 (p190-B). | |||||
|
SALL1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 102) | NC score | 0.001315 (rank : 183) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 826 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9ER74, Q920R5 | Gene names | Sall1, Sal3 | |||
|
Domain Architecture |

|
|||||
| Description | Sal-like protein 1 (Zinc finger protein Spalt-3) (Sal-3) (MSal-3). | |||||
|
SALL4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 103) | NC score | 0.001791 (rank : 181) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 799 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UJQ4 | Gene names | SALL4 | |||
|
Domain Architecture |

|
|||||
| Description | Sal-like protein 4 (Zinc finger protein SALL4). | |||||
|
SDPR_MOUSE
|
||||||
| θ value | 5.27518 (rank : 104) | NC score | 0.025945 (rank : 148) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 485 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q63918, Q3V1P6, Q78EC3, Q8CBT4 | Gene names | Sdpr, Sdr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serum deprivation-response protein (Phosphatidylserine-binding protein). | |||||
|
TB182_HUMAN
|
||||||
| θ value | 5.27518 (rank : 105) | NC score | 0.030419 (rank : 142) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9C0C2 | Gene names | TNKS1BP1, KIAA1741, TAB182 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 182 kDa tankyrase 1-binding protein. | |||||
|
TRIPB_HUMAN
|
||||||
| θ value | 5.27518 (rank : 106) | NC score | 0.013853 (rank : 164) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1587 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q15643, O14689, O15154, O95949 | Gene names | TRIP11, CEV14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid receptor-interacting protein 11 (TRIP-11) (Golgi-associated microtubule-binding protein 210) (GMAP-210) (Trip230) (Clonal evolution-related gene on chromosome 14). | |||||
|
TTBK2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 107) | NC score | 0.003709 (rank : 179) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 811 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q3UVR3, Q3TSR6, Q3UFW0, Q571D1, Q8BKA4, Q924U8 | Gene names | Ttbk2, Kiaa0847, Ttbk1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tau-tubulin kinase 2. | |||||
|
WASF2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 108) | NC score | 0.014698 (rank : 162) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y6W5, O60794, Q9UDY7 | Gene names | WASF2, WAVE2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein family member 2 (WASP-family protein member 2) (Protein WAVE-2) (Verprolin homology domain-containing protein 2). | |||||
|
CEL_HUMAN
|
||||||
| θ value | 6.88961 (rank : 109) | NC score | 0.009512 (rank : 174) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P19835, Q16398 | Gene names | CEL, BAL | |||
|
Domain Architecture |

|
|||||
| Description | Bile salt-activated lipase precursor (EC 3.1.1.3) (EC 3.1.1.13) (BAL) (Bile salt-stimulated lipase) (BSSL) (Carboxyl ester lipase) (Sterol esterase) (Cholesterol esterase) (Pancreatic lysophospholipase). | |||||
|
CO002_HUMAN
|
||||||
| θ value | 6.88961 (rank : 110) | NC score | 0.012487 (rank : 169) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NZP6 | Gene names | C15orf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C15orf2. | |||||
|
CSKI1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 111) | NC score | 0.008247 (rank : 175) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1031 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6P9K8, Q6ZPU2, Q8BWU2, Q8BX99, Q9CXH0 | Gene names | Caskin1, Kiaa1306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-1 (CASK-interacting protein 1). | |||||
|
EPN1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 112) | NC score | 0.028102 (rank : 144) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y6I3, Q86ST3, Q9HA18 | Gene names | EPN1 | |||
|
Domain Architecture |
|
|||||
| Description | Epsin-1 (EPS-15-interacting protein 1) (EH domain-binding mitotic phosphoprotein). | |||||
|
IL16_HUMAN
|
||||||
| θ value | 6.88961 (rank : 113) | NC score | 0.011426 (rank : 172) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q14005, Q16435, Q9UP18 | Gene names | IL16 | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin-16 precursor (IL-16) (Lymphocyte chemoattractant factor) (LCF). | |||||
|
IRS2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 114) | NC score | 0.018518 (rank : 154) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P81122 | Gene names | Irs2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Insulin receptor substrate 2 (IRS-2) (4PS). | |||||
|
OSBP2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 115) | NC score | 0.015762 (rank : 156) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q969R2, O60396, Q9BY96, Q9BZF0 | Gene names | OSBP2, KIAA1664, ORP4, OSBPL4 | |||
|
Domain Architecture |

|
|||||
| Description | Oxysterol-binding protein 2 (Oxysterol-binding protein-related protein 4) (OSBP-related protein 4) (ORP-4). | |||||
|
POT15_HUMAN
|
||||||
| θ value | 6.88961 (rank : 116) | NC score | 0.003741 (rank : 178) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 893 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6S5H4, Q6NXN7, Q6S5H7 | Gene names | POTE15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prostate, ovary, testis-expressed protein on chromosome 15. | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 117) | NC score | 0.021151 (rank : 151) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
ETV3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 118) | NC score | 0.011677 (rank : 171) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P41162, Q8TAC8, Q9BX30 | Gene names | ETV3, METS, PE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
ID4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 119) | NC score | 0.073985 (rank : 102) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P47928, Q13005 | Gene names | ID4 | |||
|
Domain Architecture |
|
|||||
| Description | DNA-binding protein inhibitor ID-4 (Inhibitor of DNA binding 4). | |||||
|
ID4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 120) | NC score | 0.072338 (rank : 106) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P41139 | Gene names | Id4, Id-4, Idb4 | |||
|
Domain Architecture |
|
|||||
| Description | DNA-binding protein inhibitor ID-4 (Inhibitor of DNA binding 4). | |||||
|
IWS1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 121) | NC score | 0.026337 (rank : 147) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
MAGD4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 122) | NC score | 0.006184 (rank : 177) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 338 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q96JG8, Q8WXW4, Q9BQ84, Q9BYH3, Q9BYH4 | Gene names | MAGED4, KIAA1859, MAGEE1 | |||
|
Domain Architecture |

|
|||||
| Description | Melanoma-associated antigen D4 (MAGE-D4 antigen) (MAGE-E1 antigen). | |||||
|
NPM2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 123) | NC score | 0.012090 (rank : 170) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q80W85, Q8BW23 | Gene names | Npm2 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoplasmin-2. | |||||
|
PCLO_MOUSE
|
||||||
| θ value | 8.99809 (rank : 124) | NC score | 0.016560 (rank : 155) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
TTBK2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 125) | NC score | 0.003334 (rank : 180) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 822 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6IQ55, O94932, Q6ZN52, Q8IVV1 | Gene names | TTBK2, KIAA0847 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tau-tubulin kinase 2 (EC 2.7.11.1). | |||||
|
WBS14_MOUSE
|
||||||
| θ value | 8.99809 (rank : 126) | NC score | 0.124875 (rank : 65) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q99MZ3, Q99MY9, Q99MZ0, Q99MZ1, Q99MZ2, Q9JLM5 | Gene names | Mlxipl, Mio, Wbscr14 | |||
|
Domain Architecture |

|
|||||
| Description | Williams-Beuren syndrome chromosome region 14 protein homolog (Mlx interactor) (MLX-interacting protein-like). | |||||
|
ZN275_HUMAN
|
||||||
| θ value | 8.99809 (rank : 127) | NC score | -0.002311 (rank : 187) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 761 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9NSD4 | Gene names | ZNF275 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 275. | |||||
|
ATOH1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 128) | NC score | 0.109769 (rank : 70) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q92858 | Gene names | ATOH1, ATH1 | |||
|
Domain Architecture |

|
|||||
| Description | Atonal protein homolog 1 (Helix-loop-helix protein hATH-1). | |||||
|
ATOH1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 129) | NC score | 0.110254 (rank : 69) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P48985 | Gene names | Atoh1, Ath1 | |||
|
Domain Architecture |

|
|||||
| Description | Atonal protein homolog 1 (Helix-loop-helix protein mATH-1) (MATH1). | |||||
|
BHLH3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 130) | NC score | 0.051119 (rank : 131) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9C0J9 | Gene names | BHLHB3, DEC2, SHARP1 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (hDEC2) (Enhancer-of-split and hairy-related protein 1) (SHARP-1). | |||||
|
BHLH4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 131) | NC score | 0.054284 (rank : 125) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8NDY6 | Gene names | BHLHB4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Class B basic helix-loop-helix protein 4 (bHLHB4). | |||||
|
BHLH4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 132) | NC score | 0.054077 (rank : 126) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8BGW3, Q71MB7 | Gene names | Bhlhb4, H20 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Class B basic helix-loop-helix protein 4 (bHLHB4). | |||||
|
BMAL1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 133) | NC score | 0.079257 (rank : 93) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O00327, O00313, O00314, O00315, O00316, O00317, Q99631, Q99649 | Gene names | ARNTL, BMAL1, MOP3 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Member of PAS protein 3) (Basic-helix-loop- helix-PAS orphan MOP3) (bHLH-PAS protein JAP3). | |||||
|
BMAL1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 134) | NC score | 0.078797 (rank : 96) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9WTL8, O88295, Q921S4, Q9R0U2, Q9WTL9 | Gene names | Arntl, Bmal1 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Arnt3). | |||||
|
HEN1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 135) | NC score | 0.092814 (rank : 77) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q02575 | Gene names | NHLH1, HEN1 | |||
|
Domain Architecture |
|
|||||
| Description | Helix-loop-helix protein 1 (HEN1) (Nescient helix loop helix 1) (NSCL- 1). | |||||
|
HEN1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 136) | NC score | 0.094808 (rank : 73) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q02576 | Gene names | Nhlh1, Hen1 | |||
|
Domain Architecture |
|
|||||
| Description | Helix-loop-helix protein 1 (HEN1) (Nescient helix loop helix 1) (NSCL- 1). | |||||
|
HEN2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 137) | NC score | 0.092616 (rank : 78) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q02577 | Gene names | NHLH2, HEN2 | |||
|
Domain Architecture |
|
|||||
| Description | Helix-loop-helix protein 2 (HEN2) (Nescient helix loop helix 2) (NSCL- 2). | |||||
|
HEN2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 138) | NC score | 0.091896 (rank : 79) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q64221 | Gene names | Nhlh2, Hen2 | |||
|
Domain Architecture |
|
|||||
| Description | Helix-loop-helix protein 2 (HEN2) (Nescient helix loop helix 2) (NSCL- 2). | |||||
|
HEY1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 139) | NC score | 0.050366 (rank : 132) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9Y5J3, Q5TZS3, Q9NYP4 | Gene names | HEY1, CHF2, HESR1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1) (Cardiovascular helix-loop-helix factor 2) (HES-related repressor protein 2) (HERP2). | |||||
|
HEY1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 140) | NC score | 0.051601 (rank : 130) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9WV93 | Gene names | Hey1, Hesr1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1). | |||||
|
HEY2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 141) | NC score | 0.055388 (rank : 123) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9UBP5 | Gene names | HEY2, CHF1, GRL, HERP, HERP1, HRT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Cardiovascular helix-loop-helix factor 1) (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES- related repressor protein 2) (Protein gridlock homolog). | |||||
|
HEY2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 142) | NC score | 0.055631 (rank : 122) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9QUS4, Q3TZ99, Q8CD44 | Gene names | Hey2, Chf1, Herp, Herp1, Hesr2, Hrt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES-related repressor protein 2). | |||||
|
ID2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 143) | NC score | 0.063326 (rank : 114) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q02363 | Gene names | ID2 | |||
|
Domain Architecture |
|
|||||
| Description | DNA-binding protein inhibitor ID-2 (Inhibitor of DNA binding 2). | |||||
|
ID2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 144) | NC score | 0.058505 (rank : 120) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P41136, O88604 | Gene names | Id2, Id-2, Idb2 | |||
|
Domain Architecture |
|
|||||
| Description | DNA-binding protein inhibitor ID-2 (Inhibitor of DNA binding 2). | |||||
|
LYL1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 145) | NC score | 0.093806 (rank : 75) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P12980, O76102 | Gene names | LYL1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein lyl-1 (Lymphoblastic leukemia-derived sequence 1). | |||||
|
LYL1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 146) | NC score | 0.094170 (rank : 74) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P27792 | Gene names | Lyl1, Lyl-1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein lyl-1 (Lymphoblastic leukemia-derived sequence 1). | |||||
|
MAD1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 147) | NC score | 0.143353 (rank : 53) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q05195 | Gene names | MXD1, MAD | |||
|
Domain Architecture |
|
|||||
| Description | MAD protein (MAX dimerizer) (MAX dimerization protein 1). | |||||
|
MUSC_HUMAN
|
||||||
| θ value | θ > 10 (rank : 148) | NC score | 0.072930 (rank : 104) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O60682, O75946, Q9BRE7 | Gene names | MSC, ABF1 | |||
|
Domain Architecture |

|
|||||
| Description | Musculin (Activated B-cell factor 1) (ABF-1). | |||||
|
MUSC_MOUSE
|
||||||
| θ value | θ > 10 (rank : 149) | NC score | 0.073664 (rank : 103) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O88940 | Gene names | Msc, Myor | |||
|
Domain Architecture |

|
|||||
| Description | Musculin (Myogenic repressor). | |||||
|
MYF5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 150) | NC score | 0.078929 (rank : 94) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P13349 | Gene names | MYF5 | |||
|
Domain Architecture |
|
|||||
| Description | Myogenic factor 5 (Myf-5). | |||||
|
MYF5_MOUSE
|
||||||
| θ value | θ > 10 (rank : 151) | NC score | 0.080936 (rank : 92) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P24699, Q543W7 | Gene names | Myf5, Myf-5 | |||
|
Domain Architecture |
|
|||||
| Description | Myogenic factor 5 (Myf-5). | |||||
|
MYF6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 152) | NC score | 0.090744 (rank : 83) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P23409, Q53X80, Q6FHI9 | Gene names | MYF6 | |||
|
Domain Architecture |
|
|||||
| Description | Myogenic factor 6 (Myf-6). | |||||
|
MYF6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 153) | NC score | 0.086669 (rank : 87) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P15375 | Gene names | Myf6, Myf-6 | |||
|
Domain Architecture |
|
|||||
| Description | Myogenic factor 6 (Myf-6) (Herculin). | |||||
|
MYOD1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 154) | NC score | 0.076824 (rank : 101) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P15172 | Gene names | MYOD1, MYF3, MYOD | |||
|
Domain Architecture |
|
|||||
| Description | Myoblast determination protein 1 (Myogenic factor 3) (Myf-3). | |||||
|
MYOD1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 155) | NC score | 0.078046 (rank : 98) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P10085 | Gene names | Myod1, Myod | |||
|
Domain Architecture |
|
|||||
| Description | Myoblast determination protein 1. | |||||
|
MYOG_HUMAN
|
||||||
| θ value | θ > 10 (rank : 156) | NC score | 0.087388 (rank : 85) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P15173 | Gene names | MYOG, MYF4 | |||
|
Domain Architecture |
|
|||||
| Description | Myogenin (Myogenic factor 4) (Myf-4). | |||||
|
MYOG_MOUSE
|
||||||
| θ value | θ > 10 (rank : 157) | NC score | 0.087196 (rank : 86) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P12979 | Gene names | Myog | |||
|
Domain Architecture |
|
|||||
| Description | Myogenin (MYOD1-related protein). | |||||
|
NDF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 158) | NC score | 0.077163 (rank : 100) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q13562, O00343, Q13340, Q5U095, Q96TH0, Q99455, Q9UEC8 | Gene names | NEUROD1, NEUROD | |||
|
Domain Architecture |
|
|||||
| Description | Neurogenic differentiation factor 1 (NeuroD1) (NeuroD). | |||||
|
NDF1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 159) | NC score | 0.078871 (rank : 95) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q60867, Q545N9, Q60897 | Gene names | Neurod1, Neurod | |||
|
Domain Architecture |
|
|||||
| Description | Neurogenic differentiation factor 1 (NeuroD1). | |||||
|
NDF4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 160) | NC score | 0.062880 (rank : 115) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9HD90 | Gene names | NEUROD4 | |||
|
Domain Architecture |
|
|||||
| Description | Neurogenic differentiation factor 4 (NeuroD4). | |||||
|
NDF4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 161) | NC score | 0.063733 (rank : 112) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O09105 | Gene names | Neurod4, Ath3, Atoh3 | |||
|
Domain Architecture |
|
|||||
| Description | Neurogenic differentiation factor 4 (NeuroD4) (Atonal protein homolog 3) (Helix-loop-helix protein mATH-3) (MATH3). | |||||
|
NDF6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 162) | NC score | 0.065808 (rank : 110) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q96NK8, Q9H3H6 | Gene names | NEUROD6 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic differentiation factor 6 (NeuroD6). | |||||
|
NDF6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 163) | NC score | 0.065618 (rank : 111) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P48986 | Gene names | Neurod6, Ath2, Atoh2, Nex1 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic differentiation factor 6 (NeuroD6) (Atonal protein homolog 2) (Helix-loop-helix protein mATH-2) (MATH2) (NEX-1 protein). | |||||
|
NGN1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 164) | NC score | 0.071207 (rank : 107) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q92886, Q96HE1 | Gene names | NEUROG1, NEUROD3, NGN, NGN1 | |||
|
Domain Architecture |
|
|||||
| Description | Neurogenin 1 (Neurogenic differentiation factor 3) (NeuroD3) (Neurogenic basic-helix-loop-helix protein). | |||||
|
NGN1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 165) | NC score | 0.072773 (rank : 105) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P70660 | Gene names | Neurog1, Ath4c, Neurod3, Ngn, Ngn1 | |||
|
Domain Architecture |
|
|||||
| Description | Neurogenin 1 (Neurogenic differentiation factor 3) (NeuroD3) (Neurogenic basic-helix-loop-helix protein) (Helix-loop-helix protein mATH-4C) (MATH4C). | |||||
|
NGN2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 166) | NC score | 0.091715 (rank : 80) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9H2A3, Q8N416 | Gene names | NEUROG2, NGN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neurogenin 2. | |||||
|
NGN2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 167) | NC score | 0.091248 (rank : 82) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P70447, P70237 | Gene names | Neurog2, Ath4a, Atoh4, Ngn2 | |||
|
Domain Architecture |
|
|||||
| Description | Neurogenin 2 (Atonal protein homolog 4) (Helix-loop-helix protein mATH-4A) (MATH4A). | |||||
|
NGN3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 168) | NC score | 0.091705 (rank : 81) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P70661 | Gene names | Neurog3, Ath4b, Atoh5, Ngn3 | |||
|
Domain Architecture |
|
|||||
| Description | Neurogenin 3 (Atonal protein homolog 5) (Helix-loop-helix protein mATH-4B) (MATH4B). | |||||
|
OLIG1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 169) | NC score | 0.050117 (rank : 133) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8TAK6, Q7RTS0 | Gene names | OLIG1 | |||
|
Domain Architecture |
|
|||||
| Description | Oligodendrocyte transcription factor 1 (Oligo1). | |||||
|
OLIG2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 170) | NC score | 0.051870 (rank : 128) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q13516, Q86X04, Q9NZ14 | Gene names | OLIG2, BHLHB1, PRKCBP2, RACK17 | |||
|
Domain Architecture |

|
|||||
| Description | Oligodendrocyte transcription factor 2 (Oligo2) (Basic helix-loop- helix protein class B 1) (Protein kinase C-binding protein RACK17) (Protein kinase C-binding protein 2). | |||||
|
OLIG2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 171) | NC score | 0.052336 (rank : 127) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9EQW6, Q9JKN4 | Gene names | Olig2 | |||
|
Domain Architecture |

|
|||||
| Description | Oligodendrocyte transcription factor 2 (Oligo2). | |||||
|
SCX_MOUSE
|
||||||
| θ value | θ > 10 (rank : 172) | NC score | 0.124919 (rank : 64) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q64124 | Gene names | Scx | |||
|
Domain Architecture |
|
|||||
| Description | Basic helix-loop-helix transcription factor scleraxis. | |||||
|
TAL1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 173) | NC score | 0.102063 (rank : 72) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P17542 | Gene names | TAL1, SCL, TCL5 | |||
|
Domain Architecture |
|
|||||
| Description | T-cell acute lymphocytic leukemia-1 protein (TAL-1 protein) (Stem cell protein) (T-cell leukemia/lymphoma-5 protein). | |||||
|
TAL1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 174) | NC score | 0.105168 (rank : 71) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P22091 | Gene names | Tal1, Scl, Tal-1 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell acute lymphocytic leukemia-1 protein homolog (TAL-1 protein) (Stem cell protein). | |||||
|
TAL2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 175) | NC score | 0.084565 (rank : 90) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q16559 | Gene names | TAL2 | |||
|
Domain Architecture |
|
|||||
| Description | T-cell acute lymphocytic leukemia-2 protein (TAL-2 protein). | |||||
|
TAL2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 176) | NC score | 0.084288 (rank : 91) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q62282 | Gene names | Tal2 | |||
|
Domain Architecture |
|
|||||
| Description | T-cell acute lymphocytic leukemia-2 protein homolog (TAL-2 protein). | |||||
|
TCF15_HUMAN
|
||||||
| θ value | θ > 10 (rank : 177) | NC score | 0.089765 (rank : 84) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q12870, Q9NQQ1 | Gene names | TCF15, BHLHEC2 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor 15 (bHLH-EC2 protein) (Paraxis). | |||||
|
TCF15_MOUSE
|
||||||
| θ value | θ > 10 (rank : 178) | NC score | 0.093438 (rank : 76) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q60756, Q60788 | Gene names | Tcf15, Bhlhec2 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor 15 (bHLH-EC2 protein) (Paraxis) (Meso1). | |||||
|
TCF21_HUMAN
|
||||||
| θ value | θ > 10 (rank : 179) | NC score | 0.069471 (rank : 108) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O43680, O43545, Q9BZ14 | Gene names | TCF21, POD1 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor 21 (Podocyte-expressed 1) (Pod-1) (Epicardin) (Capsulin). | |||||
|
TCF21_MOUSE
|
||||||
| θ value | θ > 10 (rank : 180) | NC score | 0.068632 (rank : 109) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O35437, Q3U023 | Gene names | Tcf21, Pod1 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor 21 (Podocyte-expressed 1) (Pod-1) (Epicardin) (Capsulin). | |||||
|
TCFL5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 181) | NC score | 0.062617 (rank : 116) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9UL49, O94771, Q9BYW0 | Gene names | TCFL5, CHA, E2BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor-like 5 protein (Cha transcription factor) (HPV-16 E2-binding protein 1) (E2BP-1). | |||||
|
TFE2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 182) | NC score | 0.056963 (rank : 121) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P15923, P15883, Q14208, Q14635, Q14636, Q9UPI9 | Gene names | TCF3, E2A, ITF1 | |||
|
Domain Architecture |
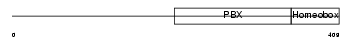
|
|||||
| Description | Transcription factor E2-alpha (Immunoglobulin enhancer-binding factor E12/E47) (Transcription factor 3) (TCF-3) (Immunoglobulin transcription factor 1) (Transcription factor ITF-1) (Kappa-E2-binding factor). | |||||
|
TFE2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 183) | NC score | 0.054679 (rank : 124) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P15806, Q3U153, Q8CAH9, Q8VCY4, Q922S2, Q99MB8, Q9CYJ4 | Gene names | Tcf3, Alf2, Me2, Tcfe2a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor E2-alpha (Immunoglobulin enhancer-binding factor E12/E47) (Transcription factor 3) (TCF-3) (Transcription factor A1). | |||||
|
TWST1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 184) | NC score | 0.136354 (rank : 57) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q15672, Q92487, Q99804 | Gene names | TWIST1, TWIST | |||
|
Domain Architecture |

|
|||||
| Description | Twist-related protein 1 (H-twist). | |||||
|
TWST1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 185) | NC score | 0.136185 (rank : 58) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P26687 | Gene names | Twist1, Twist | |||
|
Domain Architecture |

|
|||||
| Description | Twist-related protein 1 (M-twist). | |||||
|
TWST2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 186) | NC score | 0.137238 (rank : 55) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8WVJ9 | Gene names | TWIST2, DERMO1 | |||
|
Domain Architecture |
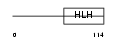
|
|||||
| Description | Twist-related protein 2 (Dermis expressed protein 1) (Dermo-1). | |||||
|
TWST2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 187) | NC score | 0.137212 (rank : 56) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9D030 | Gene names | Twist2, Dermo1 | |||
|
Domain Architecture |
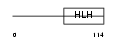
|
|||||
| Description | Twist-related protein 2 (Dermis expressed protein 1) (Dermo-1). | |||||
|
MYCN_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 2.33393e-165 (rank : 1) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 127 | |
| SwissProt Accessions | P04198, Q6LDT9 | Gene names | MYCN, NMYC | |||
|
Domain Architecture |

|
|||||
| Description | N-myc proto-oncogene protein. | |||||
|
MYCN_MOUSE
|
||||||
| NC score | 0.981433 (rank : 2) | θ value | 4.72459e-150 (rank : 2) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 86 | |
| SwissProt Accessions | P03966, Q61978 | Gene names | Mycn, Nmyc, Nmyc1 | |||
|
Domain Architecture |

|
|||||
| Description | N-myc proto-oncogene protein. | |||||
|
MYCS_MOUSE
|
||||||
| NC score | 0.950047 (rank : 3) | θ value | 4.77967e-110 (rank : 3) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9Z304 | Gene names | Mycs | |||
|
Domain Architecture |

|
|||||
| Description | Protein S-myc. | |||||
|
MYC_MOUSE
|
||||||
| NC score | 0.827060 (rank : 4) | θ value | 8.06329e-33 (rank : 4) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P01108, P70247, Q3UM70, Q61422 | Gene names | Myc | |||
|
Domain Architecture |

|
|||||
| Description | Myc proto-oncogene protein (c-Myc) (Transcription factor p64). | |||||
|
MYC_HUMAN
|
||||||
| NC score | 0.820426 (rank : 5) | θ value | 1.37539e-32 (rank : 5) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P01106, P01107, Q14026 | Gene names | MYC | |||
|
Domain Architecture |

|
|||||
| Description | Myc proto-oncogene protein (c-Myc) (Transcription factor p64). | |||||
|
MYCL1_HUMAN
|
||||||
| NC score | 0.778661 (rank : 6) | θ value | 1.24688e-25 (rank : 6) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P12524, Q14897, Q5QPL0, Q5QPL1, Q9NUE9 | Gene names | MYCL1, LMYC, MYCL | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
MYCL2_HUMAN
|
||||||
| NC score | 0.743515 (rank : 7) | θ value | 7.56453e-23 (rank : 7) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P12525 | Gene names | MYCL2 | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-2 protein. | |||||
|
MYCL1_MOUSE
|
||||||
| NC score | 0.742050 (rank : 8) | θ value | 1.33526e-19 (rank : 8) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P10166, Q5FWI7 | Gene names | Mycl1, Lmyc1, Mycl | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
MYCB_MOUSE
|
||||||
| NC score | 0.689254 (rank : 9) | θ value | 2.43908e-13 (rank : 9) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6P8Z1, Q9D6E3 | Gene names | Mycb, Bmyc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein B-Myc. | |||||
|
MAX_HUMAN
|
||||||
| NC score | 0.621615 (rank : 10) | θ value | 2.79066e-09 (rank : 10) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P61244, P25912, P52163 | Gene names | MAX | |||
|
Domain Architecture |

|
|||||
| Description | Protein max (Myc-associated factor X). | |||||
|
MAX_MOUSE
|
||||||
| NC score | 0.613570 (rank : 11) | θ value | 8.11959e-09 (rank : 11) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P28574 | Gene names | Max, Myn | |||
|
Domain Architecture |

|
|||||
| Description | Protein max (Myc-associated factor X) (Protein myn) (Myc-binding novel HLH/LZ protein). | |||||
|
USF2_HUMAN
|
||||||
| NC score | 0.361386 (rank : 12) | θ value | 0.000602161 (rank : 19) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q15853, O00671, O00709, Q05750, Q07952, Q15851, Q15852 | Gene names | USF2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (FOS- interacting protein) (FIP) (Major late transcription factor 2). | |||||
|
USF2_MOUSE
|
||||||
| NC score | 0.359233 (rank : 13) | θ value | 0.000602161 (rank : 20) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q64705, Q3UIL2, Q8VE59 | Gene names | Usf2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (Major late transcription factor 2). | |||||
|
USF1_MOUSE
|
||||||
| NC score | 0.358516 (rank : 14) | θ value | 0.00134147 (rank : 23) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q61069 | Gene names | Usf1, Usf | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF1_HUMAN
|
||||||
| NC score | 0.356055 (rank : 15) | θ value | 0.00134147 (rank : 22) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P22415 | Gene names | USF1, USF | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
FIGLA_MOUSE
|
||||||
| NC score | 0.338721 (rank : 16) | θ value | 6.43352e-06 (rank : 12) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O55208 | Gene names | Figla | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Factor in the germline alpha (Transcription factor FIGa) (FIGalpha). | |||||
|
FIGLA_HUMAN
|
||||||
| NC score | 0.320982 (rank : 17) | θ value | 0.000602161 (rank : 17) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q6QHK4 | Gene names | FIGLA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Factor in the germline alpha (Transcription factor FIGa) (FIGalpha). | |||||
|
TFAP4_HUMAN
|
||||||
| NC score | 0.314233 (rank : 18) | θ value | 0.00509761 (rank : 29) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q01664, O60409 | Gene names | TFAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor AP-4 (Activating enhancer-binding protein 4). | |||||
|
MNT_MOUSE
|
||||||
| NC score | 0.312423 (rank : 19) | θ value | 7.1131e-05 (rank : 15) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 432 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O08789, P97349 | Gene names | Mnt, Rox | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
MNT_HUMAN
|
||||||
| NC score | 0.304686 (rank : 20) | θ value | 7.1131e-05 (rank : 14) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 496 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q99583 | Gene names | MNT, ROX | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
MXI1_MOUSE
|
||||||
| NC score | 0.300630 (rank : 21) | θ value | 0.0330416 (rank : 36) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P50540 | Gene names | Mxi1 | |||
|
Domain Architecture |

|
|||||
| Description | MAX-interacting protein 1 (Protein MXI1). | |||||
|
TFE3_HUMAN
|
||||||
| NC score | 0.293634 (rank : 22) | θ value | 0.00298849 (rank : 24) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P19532, Q92757, Q92758, Q99964 | Gene names | TFE3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E3. | |||||
|
SRBP2_HUMAN
|
||||||
| NC score | 0.290005 (rank : 23) | θ value | 7.1131e-05 (rank : 16) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q12772, Q9UH04 | Gene names | SREBF2, SREBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 2 (SREBP-2) (Sterol regulatory element-binding transcription factor 2). | |||||
|
MAD4_HUMAN
|
||||||
| NC score | 0.289872 (rank : 24) | θ value | 0.00509761 (rank : 28) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q14582, Q5TZX4 | Gene names | MXD4, MAD4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-interacting transcriptional repressor MAD4 (Max-associated protein 4) (MAX dimerization protein 4). | |||||
|
MITF_HUMAN
|
||||||
| NC score | 0.288869 (rank : 25) | θ value | 0.000602161 (rank : 18) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O75030, Q14841, Q9P2V0, Q9P2V1, Q9P2V2, Q9P2Y8 | Gene names | MITF | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
MXI1_HUMAN
|
||||||
| NC score | 0.288526 (rank : 26) | θ value | 0.125558 (rank : 44) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P50539, Q15887 | Gene names | MXI1 | |||
|
Domain Architecture |

|
|||||
| Description | MAX-interacting protein 1 (Protein MXI1). | |||||
|
MITF_MOUSE
|
||||||
| NC score | 0.286236 (rank : 27) | θ value | 0.00102713 (rank : 21) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q08874, O08885, O88203, Q08843, Q60781, Q60782, Q9JIJ0, Q9JIJ1, Q9JIJ2, Q9JIJ3, Q9JIJ4, Q9JIJ5, Q9JIJ6, Q9JKX9 | Gene names | Mitf, Bw, Mi, Vit | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
MLX_MOUSE
|
||||||
| NC score | 0.270075 (rank : 28) | θ value | 0.125558 (rank : 43) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O08609, Q9EQJ7, Q9EQJ8 | Gene names | Mlx, Tcfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
MLX_HUMAN
|
||||||
| NC score | 0.268579 (rank : 29) | θ value | 0.125558 (rank : 42) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UH92, Q53XM6, Q96FL2, Q9H2V0, Q9H2V1, Q9H2V2, Q9NXN3 | Gene names | MLX, TCFL4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
MAD3_HUMAN
|
||||||
| NC score | 0.256197 (rank : 30) | θ value | 3.0926 (rank : 92) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9BW11, Q53HK1, Q7Z4Y0, Q8NDJ7, Q96ME3 | Gene names | MXD3, MAD3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Max-interacting transcriptional repressor MAD3 (Max-associated protein 3) (MAX dimerization protein 3) (Myx). | |||||
|
SRBP1_HUMAN
|
||||||
| NC score | 0.250968 (rank : 31) | θ value | 0.0252991 (rank : 34) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P36956, Q16062 | Gene names | SREBF1, SREBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
ASCL1_HUMAN
|
||||||
| NC score | 0.250606 (rank : 32) | θ value | 0.00509761 (rank : 27) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P50553, Q9BQ30 | Gene names | ASCL1, ASH1 | |||
|
Domain Architecture |

|
|||||
| Description | Achaete-scute homolog 1 (HASH1). | |||||
|
MAD3_MOUSE
|
||||||
| NC score | 0.248938 (rank : 33) | θ value | 1.38821 (rank : 79) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q80US8, Q60947 | Gene names | Mxd3, Mad3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Max-interacting transcriptional repressor MAD3 (Max-associated protein 3) (MAX dimerization protein 3). | |||||
|
BHLH8_MOUSE
|
||||||
| NC score | 0.247501 (rank : 34) | θ value | 0.0113563 (rank : 32) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9QYC3, Q9QYE4 | Gene names | Bhlhb8, Mist1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Class B basic helix-loop-helix protein 8 (bHLHB8) (Muscle, intestine and stomach expression 1) (MIST-1). | |||||
|
BHLH8_HUMAN
|
||||||
| NC score | 0.241005 (rank : 35) | θ value | 0.0113563 (rank : 31) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q7RTS1 | Gene names | BHLHB8, MIST1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Class B basic helix-loop-helix protein 8 (bHLHB8) (Muscle, intestine and stomach expression 1) (MIST-1). | |||||
|
MAD4_MOUSE
|
||||||
| NC score | 0.238531 (rank : 36) | θ value | 0.0330416 (rank : 35) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q60948 | Gene names | Mxd4, Mad4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-interacting transcriptional repressor MAD4 (Max-associated protein 4) (MAX dimerization protein 4). | |||||
|
ASCL1_MOUSE
|
||||||
| NC score | 0.238249 (rank : 37) | θ value | 0.0113563 (rank : 30) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q02067 | Gene names | Ascl1, Ash1, Mash-1, Mash1 | |||
|
Domain Architecture |

|
|||||
| Description | Achaete-scute homolog 1 (Mash-1). | |||||
|
SRBP1_MOUSE
|
||||||
| NC score | 0.237097 (rank : 38) | θ value | 0.0736092 (rank : 38) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9WTN3 | Gene names | Srebf1, Srebp1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
TFEB_MOUSE
|
||||||
| NC score | 0.232756 (rank : 39) | θ value | 0.813845 (rank : 72) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9R210 | Gene names | Tfeb, Tcfeb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
TFEB_HUMAN
|
||||||
| NC score | 0.231732 (rank : 40) | θ value | 0.813845 (rank : 71) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P19484, Q709B3, Q7Z6P9, Q9BRJ5, Q9UJD8 | Gene names | TFEB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
ASCL2_MOUSE
|
||||||
| NC score | 0.219993 (rank : 41) | θ value | 0.365318 (rank : 58) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O35885, Q9WUJ7 | Gene names | Ascl2, Mash2 | |||
|
Domain Architecture |

|
|||||
| Description | Achaete-scute homolog 2 (Mash-2). | |||||
|
MAD1_MOUSE
|
||||||
| NC score | 0.208077 (rank : 42) | θ value | 0.813845 (rank : 70) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P50538, Q60798, Q61825 | Gene names | Mxd1, Mad, Mad1 | |||
|
Domain Architecture |

|
|||||
| Description | MAD protein (MAX dimerizer) (MAX dimerization protein 1). | |||||
|
HAND2_HUMAN
|
||||||
| NC score | 0.194363 (rank : 43) | θ value | 0.47712 (rank : 60) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P61296, O95300, O95301, P97833 | Gene names | HAND2, DHAND | |||
|
Domain Architecture |

|
|||||
| Description | Heart- and neural crest derivatives-expressed protein 2 (Deciduum, heart, autonomic nervous system and neural crest derivatives-expressed protein 2) (dHAND). | |||||
|
HAND2_MOUSE
|
||||||
| NC score | 0.194363 (rank : 44) | θ value | 0.47712 (rank : 61) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q61039, Q61100 | Gene names | Hand2, Dhand, Hed, Thing2 | |||
|
Domain Architecture |

|
|||||
| Description | Heart- and neural crest derivatives-expressed protein 2 (Deciduum, heart, autonomic nervous system and neural crest derivatives-expressed protein 2) (dHAND) (Helix-loop-helix transcription factor expressed in embryo and deciduum) (Thing-2). | |||||
|
PTF1A_MOUSE
|
||||||
| NC score | 0.187526 (rank : 45) | θ value | 0.0193708 (rank : 33) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9QX98, Q9QYF5, Q9QYF6 | Gene names | Ptf1a, Ptf1p48 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pancreas transcription factor 1 subunit alpha (Pancreas-specific transcription factor 1a) (bHLH transcription factor p48) (p48 DNA- binding subunit of transcription factor PTF1) (PTF1-p48). | |||||
|
ASCL2_HUMAN
|
||||||
| NC score | 0.185394 (rank : 46) | θ value | 1.38821 (rank : 77) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q99929, Q9UM68 | Gene names | ASCL2 | |||
|
Domain Architecture |

|
|||||
| Description | Achaete-scute homolog 2 (Mash2) (HASH2). | |||||
|
PTF1A_HUMAN
|
||||||
| NC score | 0.184899 (rank : 47) | θ value | 0.163984 (rank : 47) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q7RTS3, Q9HC25 | Gene names | PTF1A, PTF1P48 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pancreas transcription factor 1 subunit alpha (Pancreas-specific transcription factor 1a) (bHLH transcription factor p48) (p48 DNA- binding subunit of transcription factor PTF1) (PTF1-p48). | |||||
|
WBS14_HUMAN
|
||||||
| NC score | 0.175194 (rank : 48) | θ value | 0.00298849 (rank : 25) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9NP71, Q96E48, Q9BY03, Q9BY04, Q9BY05, Q9BY06, Q9Y2P3 | Gene names | MLXIPL, MIO, WBSCR14 | |||
|
Domain Architecture |

|
|||||
| Description | Williams-Beuren syndrome chromosome region 14 protein (WS basic-helix- loop-helix leucine zipper protein) (WS-bHLH) (Mlx interactor) (MLX- interacting protein-like). | |||||
|
ASCL3_HUMAN
|
||||||
| NC score | 0.173750 (rank : 49) | θ value | 0.0961366 (rank : 41) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9NQ33, Q8WYQ6 | Gene names | ASCL3, HASH3, SGN1 | |||
|
Domain Architecture |

|
|||||
| Description | Achaete-scute homolog 3 (bHLH transcriptional regulator Sgn-1). | |||||
|
HAND1_MOUSE
|
||||||
| NC score | 0.171338 (rank : 50) | θ value | 0.62314 (rank : 64) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q64279, Q61099 | Gene names | Hand1, Ehand, Hxt, Thing1 | |||
|
Domain Architecture |

|
|||||
| Description | Heart- and neural crest derivatives-expressed protein 1 (Extraembryonic tissues, heart, autonomic nervous system and neural crest derivatives-expressed protein 1) (eHAND) (Helix-loop-helix transcription factor expressed in extraembryonic mesoderm and trophoblast) (Thing-1) (Th1). | |||||
|
HAND1_HUMAN
|
||||||
| NC score | 0.166016 (rank : 51) | θ value | 0.62314 (rank : 63) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O96004 | Gene names | HAND1, EHAND | |||
|
Domain Architecture |

|
|||||
| Description | Heart- and neural crest derivatives-expressed protein 1 (Extraembryonic tissues, heart, autonomic nervous system and neural crest derivatives-expressed protein 1) (eHAND). | |||||
|
ASCL3_MOUSE
|
||||||
| NC score | 0.160347 (rank : 52) | θ value | 0.279714 (rank : 52) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9JJR7 | Gene names | Ascl3, Mash3, Sgn1 | |||
|
Domain Architecture |

|
|||||
| Description | Achaete-scute homolog 3 (bHLH transcriptional regulator Sgn-1) (Mash- 3). | |||||
|
MAD1_HUMAN
|
||||||
| NC score | 0.143353 (rank : 53) | θ value | θ > 10 (rank : 147) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q05195 | Gene names | MXD1, MAD | |||
|
Domain Architecture |

|
|||||
| Description | MAD protein (MAX dimerizer) (MAX dimerization protein 1). | |||||
|
ARNT_HUMAN
|
||||||
| NC score | 0.140270 (rank : 54) | θ value | 0.0961366 (rank : 39) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P27540 | Gene names | ARNT | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
TWST2_HUMAN
|
||||||
| NC score | 0.137238 (rank : 55) | θ value | θ > 10 (rank : 186) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8WVJ9 | Gene names | TWIST2, DERMO1 | |||
|
Domain Architecture |

|
|||||
| Description | Twist-related protein 2 (Dermis expressed protein 1) (Dermo-1). | |||||
|
TWST2_MOUSE
|
||||||
| NC score | 0.137212 (rank : 56) | θ value | θ > 10 (rank : 187) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9D030 | Gene names | Twist2, Dermo1 | |||
|
Domain Architecture |

|
|||||
| Description | Twist-related protein 2 (Dermis expressed protein 1) (Dermo-1). | |||||
|
TWST1_HUMAN
|
||||||
| NC score | 0.136354 (rank : 57) | θ value | θ > 10 (rank : 184) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q15672, Q92487, Q99804 | Gene names | TWIST1, TWIST | |||
|
Domain Architecture |

|
|||||
| Description | Twist-related protein 1 (H-twist). | |||||
|
TWST1_MOUSE
|
||||||
| NC score | 0.136185 (rank : 58) | θ value | θ > 10 (rank : 185) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P26687 | Gene names | Twist1, Twist | |||
|
Domain Architecture |

|
|||||
| Description | Twist-related protein 1 (M-twist). | |||||
|
ARNT_MOUSE
|
||||||
| NC score | 0.135805 (rank : 59) | θ value | 0.0961366 (rank : 40) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P53762, Q60661 | Gene names | Arnt | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
ID1_HUMAN
|
||||||
| NC score | 0.130465 (rank : 60) | θ value | 0.279714 (rank : 53) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P41134, O00651, O00652, Q16371, Q16377, Q5TE66, Q5TE67, Q969Z7, Q9H0Z5, Q9H109 | Gene names | ID1, ID | |||
|
Domain Architecture |

|
|||||
| Description | DNA-binding protein inhibitor ID-1 (Inhibitor of DNA binding 1). | |||||
|
ARNT2_MOUSE
|
||||||
| NC score | 0.128427 (rank : 61) | θ value | 0.21417 (rank : 49) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q61324 | Gene names | Arnt2 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
ID1_MOUSE
|
||||||
| NC score | 0.128009 (rank : 62) | θ value | 0.279714 (rank : 54) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P20067, Q61101, Q9D897 | Gene names | Id1, Id, Id-1, Idb1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-binding protein inhibitor ID-1 (Inhibitor of DNA binding 1). | |||||
|
ARNT2_HUMAN
|
||||||
| NC score | 0.127967 (rank : 63) | θ value | 0.21417 (rank : 48) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9HBZ2, O15024 | Gene names | ARNT2, KIAA0307 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
SCX_MOUSE
|
||||||
| NC score | 0.124919 (rank : 64) | θ value | θ > 10 (rank : 172) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q64124 | Gene names | Scx | |||
|
Domain Architecture |

|
|||||
| Description | Basic helix-loop-helix transcription factor scleraxis. | |||||
|
WBS14_MOUSE
|
||||||
| NC score | 0.124875 (rank : 65) | θ value | 8.99809 (rank : 126) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q99MZ3, Q99MY9, Q99MZ0, Q99MZ1, Q99MZ2, Q9JLM5 | Gene names | Mlxipl, Mio, Wbscr14 | |||
|
Domain Architecture |

|
|||||
| Description | Williams-Beuren syndrome chromosome region 14 protein homolog (Mlx interactor) (MLX-interacting protein-like). | |||||
|
NDF2_HUMAN
|
||||||
| NC score | 0.120114 (rank : 66) | θ value | 0.125558 (rank : 45) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q15784, Q8TBI7, Q9UQC6 | Gene names | NEUROD2, NDRF | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic differentiation factor 2 (NeuroD2) (NeuroD-related factor) (NDRF). | |||||
|
NDF2_MOUSE
|
||||||
| NC score | 0.119961 (rank : 67) | θ value | 0.125558 (rank : 46) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q62414, Q61952, Q925V5 | Gene names | Neurod2, Ndrf | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic differentiation factor 2 (NeuroD2) (NeuroD-related factor) (NDRF). | |||||
|
NGN3_HUMAN
|
||||||
| NC score | 0.115057 (rank : 68) | θ value | 1.06291 (rank : 75) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9Y4Z2, Q9BY24 | Gene names | NEUROG3, NGN3 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenin 3. | |||||
|
ATOH1_MOUSE
|
||||||
| NC score | 0.110254 (rank : 69) | θ value | θ > 10 (rank : 129) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P48985 | Gene names | Atoh1, Ath1 | |||
|
Domain Architecture |

|
|||||
| Description | Atonal protein homolog 1 (Helix-loop-helix protein mATH-1) (MATH1). | |||||
|
ATOH1_HUMAN
|
||||||
| NC score | 0.109769 (rank : 70) | θ value | θ > 10 (rank : 128) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q92858 | Gene names | ATOH1, ATH1 | |||
|
Domain Architecture |

|
|||||
| Description | Atonal protein homolog 1 (Helix-loop-helix protein hATH-1). | |||||
|
TAL1_MOUSE
|
||||||
| NC score | 0.105168 (rank : 71) | θ value | θ > 10 (rank : 174) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P22091 | Gene names | Tal1, Scl, Tal-1 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell acute lymphocytic leukemia-1 protein homolog (TAL-1 protein) (Stem cell protein). | |||||
|
TAL1_HUMAN
|
||||||
| NC score | 0.102063 (rank : 72) | θ value | θ > 10 (rank : 173) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P17542 | Gene names | TAL1, SCL, TCL5 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell acute lymphocytic leukemia-1 protein (TAL-1 protein) (Stem cell protein) (T-cell leukemia/lymphoma-5 protein). | |||||
|
HEN1_MOUSE
|
||||||
| NC score | 0.094808 (rank : 73) | θ value | θ > 10 (rank : 136) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q02576 | Gene names | Nhlh1, Hen1 | |||
|
Domain Architecture |

|
|||||
| Description | Helix-loop-helix protein 1 (HEN1) (Nescient helix loop helix 1) (NSCL- 1). | |||||
|
LYL1_MOUSE
|
||||||
| NC score | 0.094170 (rank : 74) | θ value | θ > 10 (rank : 146) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P27792 | Gene names | Lyl1, Lyl-1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein lyl-1 (Lymphoblastic leukemia-derived sequence 1). | |||||
|
LYL1_HUMAN
|
||||||
| NC score | 0.093806 (rank : 75) | θ value | θ > 10 (rank : 145) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P12980, O76102 | Gene names | LYL1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein lyl-1 (Lymphoblastic leukemia-derived sequence 1). | |||||
|
TCF15_MOUSE
|
||||||
| NC score | 0.093438 (rank : 76) | θ value | θ > 10 (rank : 178) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q60756, Q60788 | Gene names | Tcf15, Bhlhec2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 15 (bHLH-EC2 protein) (Paraxis) (Meso1). | |||||
|
HEN1_HUMAN
|
||||||
| NC score | 0.092814 (rank : 77) | θ value | θ > 10 (rank : 135) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q02575 | Gene names | NHLH1, HEN1 | |||
|
Domain Architecture |

|
|||||
| Description | Helix-loop-helix protein 1 (HEN1) (Nescient helix loop helix 1) (NSCL- 1). | |||||
|
HEN2_HUMAN
|
||||||
| NC score | 0.092616 (rank : 78) | θ value | θ > 10 (rank : 137) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q02577 | Gene names | NHLH2, HEN2 | |||
|
Domain Architecture |

|
|||||
| Description | Helix-loop-helix protein 2 (HEN2) (Nescient helix loop helix 2) (NSCL- 2). | |||||
|
HEN2_MOUSE
|
||||||
| NC score | 0.091896 (rank : 79) | θ value | θ > 10 (rank : 138) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q64221 | Gene names | Nhlh2, Hen2 | |||
|
Domain Architecture |

|
|||||
| Description | Helix-loop-helix protein 2 (HEN2) (Nescient helix loop helix 2) (NSCL- 2). | |||||
|
NGN2_HUMAN
|
||||||
| NC score | 0.091715 (rank : 80) | θ value | θ > 10 (rank : 166) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9H2A3, Q8N416 | Gene names | NEUROG2, NGN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neurogenin 2. | |||||
|
NGN3_MOUSE
|
||||||
| NC score | 0.091705 (rank : 81) | θ value | θ > 10 (rank : 168) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P70661 | Gene names | Neurog3, Ath4b, Atoh5, Ngn3 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenin 3 (Atonal protein homolog 5) (Helix-loop-helix protein mATH-4B) (MATH4B). | |||||
|
NGN2_MOUSE
|
||||||
| NC score | 0.091248 (rank : 82) | θ value | θ > 10 (rank : 167) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P70447, P70237 | Gene names | Neurog2, Ath4a, Atoh4, Ngn2 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenin 2 (Atonal protein homolog 4) (Helix-loop-helix protein mATH-4A) (MATH4A). | |||||
|
MYF6_HUMAN
|
||||||
| NC score | 0.090744 (rank : 83) | θ value | θ > 10 (rank : 152) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P23409, Q53X80, Q6FHI9 | Gene names | MYF6 | |||
|
Domain Architecture |

|
|||||
| Description | Myogenic factor 6 (Myf-6). | |||||
|
TCF15_HUMAN
|
||||||
| NC score | 0.089765 (rank : 84) | θ value | θ > 10 (rank : 177) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q12870, Q9NQQ1 | Gene names | TCF15, BHLHEC2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 15 (bHLH-EC2 protein) (Paraxis). | |||||
|
MYOG_HUMAN
|
||||||
| NC score | 0.087388 (rank : 85) | θ value | θ > 10 (rank : 156) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P15173 | Gene names | MYOG, MYF4 | |||
|
Domain Architecture |

|
|||||
| Description | Myogenin (Myogenic factor 4) (Myf-4). | |||||
|
MYOG_MOUSE
|
||||||
| NC score | 0.087196 (rank : 86) | θ value | θ > 10 (rank : 157) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P12979 | Gene names | Myog | |||
|
Domain Architecture |

|
|||||
| Description | Myogenin (MYOD1-related protein). | |||||
|
MYF6_MOUSE
|
||||||
| NC score | 0.086669 (rank : 87) | θ value | θ > 10 (rank : 153) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P15375 | Gene names | Myf6, Myf-6 | |||
|
Domain Architecture |

|
|||||
| Description | Myogenic factor 6 (Myf-6) (Herculin). | |||||
|
ITF2_MOUSE
|
||||||
| NC score | 0.086380 (rank : 88) | θ value | 0.279714 (rank : 56) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q60722, Q60721, Q62211, Q64072, Q80UE8 | Gene names | Tcf4, Itf2, Sef2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 4 (Immunoglobulin transcription factor 2) (ITF-2) (MITF-2) (SL3-3 enhancer factor 2) (SEF-2) (Class A helix-loop-helix transcription factor ME2). | |||||
|
ITF2_HUMAN
|
||||||
| NC score | 0.085149 (rank : 89) | θ value | 0.279714 (rank : 55) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P15884, Q15439, Q15440 | Gene names | TCF4, ITF2, SEF2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 4 (Immunoglobulin transcription factor 2) (ITF-2) (SL3-3 enhancer factor 2) (SEF-2). | |||||
|
TAL2_HUMAN
|
||||||
| NC score | 0.084565 (rank : 90) | θ value | θ > 10 (rank : 175) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q16559 | Gene names | TAL2 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell acute lymphocytic leukemia-2 protein (TAL-2 protein). | |||||
|
TAL2_MOUSE
|
||||||
| NC score | 0.084288 (rank : 91) | θ value | θ > 10 (rank : 176) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q62282 | Gene names | Tal2 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell acute lymphocytic leukemia-2 protein homolog (TAL-2 protein). | |||||
|
MYF5_MOUSE
|
||||||
| NC score | 0.080936 (rank : 92) | θ value | θ > 10 (rank : 151) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P24699, Q543W7 | Gene names | Myf5, Myf-5 | |||
|
Domain Architecture |

|
|||||
| Description | Myogenic factor 5 (Myf-5). | |||||
|
BMAL1_HUMAN
|
||||||
| NC score | 0.079257 (rank : 93) | θ value | θ > 10 (rank : 133) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O00327, O00313, O00314, O00315, O00316, O00317, Q99631, Q99649 | Gene names | ARNTL, BMAL1, MOP3 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Member of PAS protein 3) (Basic-helix-loop- helix-PAS orphan MOP3) (bHLH-PAS protein JAP3). | |||||
|
MYF5_HUMAN
|
||||||
| NC score | 0.078929 (rank : 94) | θ value | θ > 10 (rank : 150) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P13349 | Gene names | MYF5 | |||
|
Domain Architecture |

|
|||||
| Description | Myogenic factor 5 (Myf-5). | |||||
|
NDF1_MOUSE
|
||||||
| NC score | 0.078871 (rank : 95) | θ value | θ > 10 (rank : 159) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q60867, Q545N9, Q60897 | Gene names | Neurod1, Neurod | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic differentiation factor 1 (NeuroD1). | |||||
|
BMAL1_MOUSE
|
||||||
| NC score | 0.078797 (rank : 96) | θ value | θ > 10 (rank : 134) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9WTL8, O88295, Q921S4, Q9R0U2, Q9WTL9 | Gene names | Arntl, Bmal1 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Arnt3). | |||||
|
HTF4_HUMAN
|
||||||
| NC score | 0.078793 (rank : 97) | θ value | 0.47712 (rank : 62) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q99081 | Gene names | TCF12, HEB, HTF4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 12 (Transcription factor HTF-4) (E-box-binding protein) (DNA-binding protein HTF4). | |||||
|
MYOD1_MOUSE
|
||||||
| NC score | 0.078046 (rank : 98) | θ value | θ > 10 (rank : 155) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P10085 | Gene names | Myod1, Myod | |||
|
Domain Architecture |

|
|||||
| Description | Myoblast determination protein 1. | |||||
|
HTF4_MOUSE
|
||||||
| NC score | 0.077577 (rank : 99) | θ value | 0.62314 (rank : 65) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q61286 | Gene names | Tcf12, Alf1, Meb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 12 (Transcription factor HTF-4) (E-box-binding protein) (DNA-binding protein HTF4) (Class A helix-loop-helix transcription factor ME1). | |||||
|
NDF1_HUMAN
|
||||||
| NC score | 0.077163 (rank : 100) | θ value | θ > 10 (rank : 158) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q13562, O00343, Q13340, Q5U095, Q96TH0, Q99455, Q9UEC8 | Gene names | NEUROD1, NEUROD | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic differentiation factor 1 (NeuroD1) (NeuroD). | |||||
|
MYOD1_HUMAN
|
||||||
| NC score | 0.076824 (rank : 101) | θ value | θ > 10 (rank : 154) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P15172 | Gene names | MYOD1, MYF3, MYOD | |||
|
Domain Architecture |

|
|||||
| Description | Myoblast determination protein 1 (Myogenic factor 3) (Myf-3). | |||||
|
ID4_HUMAN
|
||||||
| NC score | 0.073985 (rank : 102) | θ value | 8.99809 (rank : 119) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P47928, Q13005 | Gene names | ID4 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-binding protein inhibitor ID-4 (Inhibitor of DNA binding 4). | |||||
|
MUSC_MOUSE
|
||||||
| NC score | 0.073664 (rank : 103) | θ value | θ > 10 (rank : 149) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O88940 | Gene names | Msc, Myor | |||
|
Domain Architecture |

|
|||||
| Description | Musculin (Myogenic repressor). | |||||
|
MUSC_HUMAN
|
||||||
| NC score | 0.072930 (rank : 104) | θ value | θ > 10 (rank : 148) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O60682, O75946, Q9BRE7 | Gene names | MSC, ABF1 | |||
|
Domain Architecture |

|
|||||
| Description | Musculin (Activated B-cell factor 1) (ABF-1). | |||||
|
NGN1_MOUSE
|
||||||
| NC score | 0.072773 (rank : 105) | θ value | θ > 10 (rank : 165) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P70660 | Gene names | Neurog1, Ath4c, Neurod3, Ngn, Ngn1 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenin 1 (Neurogenic differentiation factor 3) (NeuroD3) (Neurogenic basic-helix-loop-helix protein) (Helix-loop-helix protein mATH-4C) (MATH4C). | |||||
|
ID4_MOUSE
|
||||||
| NC score | 0.072338 (rank : 106) | θ value | 8.99809 (rank : 120) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P41139 | Gene names | Id4, Id-4, Idb4 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-binding protein inhibitor ID-4 (Inhibitor of DNA binding 4). | |||||
|
NGN1_HUMAN
|
||||||
| NC score | 0.071207 (rank : 107) | θ value | θ > 10 (rank : 164) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q92886, Q96HE1 | Gene names | NEUROG1, NEUROD3, NGN, NGN1 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenin 1 (Neurogenic differentiation factor 3) (NeuroD3) (Neurogenic basic-helix-loop-helix protein). | |||||
|
TCF21_HUMAN
|
||||||
| NC score | 0.069471 (rank : 108) | θ value | θ > 10 (rank : 179) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O43680, O43545, Q9BZ14 | Gene names | TCF21, POD1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 21 (Podocyte-expressed 1) (Pod-1) (Epicardin) (Capsulin). | |||||
|
TCF21_MOUSE
|
||||||
| NC score | 0.068632 (rank : 109) | θ value | θ > 10 (rank : 180) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O35437, Q3U023 | Gene names | Tcf21, Pod1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 21 (Podocyte-expressed 1) (Pod-1) (Epicardin) (Capsulin). | |||||
|
NDF6_HUMAN
|
||||||
| NC score | 0.065808 (rank : 110) | θ value | θ > 10 (rank : 162) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q96NK8, Q9H3H6 | Gene names | NEUROD6 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic differentiation factor 6 (NeuroD6). | |||||
|
NDF6_MOUSE
|
||||||
| NC score | 0.065618 (rank : 111) | θ value | θ > 10 (rank : 163) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P48986 | Gene names | Neurod6, Ath2, Atoh2, Nex1 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic differentiation factor 6 (NeuroD6) (Atonal protein homolog 2) (Helix-loop-helix protein mATH-2) (MATH2) (NEX-1 protein). | |||||
|
NDF4_MOUSE
|
||||||
| NC score | 0.063733 (rank : 112) | θ value | θ > 10 (rank : 161) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O09105 | Gene names | Neurod4, Ath3, Atoh3 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic differentiation factor 4 (NeuroD4) (Atonal protein homolog 3) (Helix-loop-helix protein mATH-3) (MATH3). | |||||
|
GDIA_HUMAN
|
||||||
| NC score | 0.063529 (rank : 113) | θ value | 0.47712 (rank : 59) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P31150, P50394, Q7Z2G6, Q7Z2G9, Q7Z2H5, Q7Z2I6 | Gene names | GDI1, OPHN2, RABGDIA, XAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Rab GDP dissociation inhibitor alpha (Rab GDI alpha) (Guanosine diphosphate dissociation inhibitor 1) (GDI-1) (XAP-4) (Oligophrenin- 2). | |||||
|
ID2_HUMAN
|
||||||
| NC score | 0.063326 (rank : 114) | θ value | θ > 10 (rank : 143) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q02363 | Gene names | ID2 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-binding protein inhibitor ID-2 (Inhibitor of DNA binding 2). | |||||
|
NDF4_HUMAN
|
||||||
| NC score | 0.062880 (rank : 115) | θ value | θ > 10 (rank : 160) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9HD90 | Gene names | NEUROD4 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic differentiation factor 4 (NeuroD4). | |||||
|
TCFL5_HUMAN
|
||||||
| NC score | 0.062617 (rank : 116) | θ value | θ > 10 (rank : 181) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9UL49, O94771, Q9BYW0 | Gene names | TCFL5, CHA, E2BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor-like 5 protein (Cha transcription factor) (HPV-16 E2-binding protein 1) (E2BP-1). | |||||
|
GDIA_MOUSE
|
||||||
| NC score | 0.060480 (rank : 117) | θ value | 0.813845 (rank : 68) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P50396, Q8VHM3, Q91Y71, Q96CX5 | Gene names | Gdi1, Rabgdia | |||
|
Domain Architecture |

|
|||||
| Description | Rab GDP dissociation inhibitor alpha (Rab GDI alpha) (Guanosine diphosphate dissociation inhibitor 1) (GDI-1). | |||||
|
GDIB_MOUSE
|
||||||
| NC score | 0.060205 (rank : 118) | θ value | 0.813845 (rank : 69) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61598, Q541Z9, Q8C530, Q9D8M9 | Gene names | Gdi2, Gdi3 | |||
|
Domain Architecture |

|
|||||
| Description | Rab GDP dissociation inhibitor beta (Rab GDI beta) (Guanosine diphosphate dissociation inhibitor 2) (GDI-2) (GDI-3). | |||||
|
GRIN1_HUMAN
|
||||||
| NC score | 0.058631 (rank : 119) | θ value | 0.00390308 (rank : 26) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q7Z2K8, Q8ND74, Q96PZ4 | Gene names | GPRIN1, KIAA1893 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
ID2_MOUSE
|
||||||
| NC score | 0.058505 (rank : 120) | θ value | θ > 10 (rank : 144) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P41136, O88604 | Gene names | Id2, Id-2, Idb2 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-binding protein inhibitor ID-2 (Inhibitor of DNA binding 2). | |||||
|
TFE2_HUMAN
|
||||||
| NC score | 0.056963 (rank : 121) | θ value | θ > 10 (rank : 182) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P15923, P15883, Q14208, Q14635, Q14636, Q9UPI9 | Gene names | TCF3, E2A, ITF1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E2-alpha (Immunoglobulin enhancer-binding factor E12/E47) (Transcription factor 3) (TCF-3) (Immunoglobulin transcription factor 1) (Transcription factor ITF-1) (Kappa-E2-binding factor). | |||||
|
HEY2_MOUSE
|
||||||
| NC score | 0.055631 (rank : 122) | θ value | θ > 10 (rank : 142) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9QUS4, Q3TZ99, Q8CD44 | Gene names | Hey2, Chf1, Herp, Herp1, Hesr2, Hrt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES-related repressor protein 2). | |||||
|
HEY2_HUMAN
|
||||||
| NC score | 0.055388 (rank : 123) | θ value | θ > 10 (rank : 141) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9UBP5 | Gene names | HEY2, CHF1, GRL, HERP, HERP1, HRT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Cardiovascular helix-loop-helix factor 1) (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES- related repressor protein 2) (Protein gridlock homolog). | |||||
|
TFE2_MOUSE
|
||||||
| NC score | 0.054679 (rank : 124) | θ value | θ > 10 (rank : 183) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P15806, Q3U153, Q8CAH9, Q8VCY4, Q922S2, Q99MB8, Q9CYJ4 | Gene names | Tcf3, Alf2, Me2, Tcfe2a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor E2-alpha (Immunoglobulin enhancer-binding factor E12/E47) (Transcription factor 3) (TCF-3) (Transcription factor A1). | |||||
|
BHLH4_HUMAN
|
||||||
| NC score | 0.054284 (rank : 125) | θ value | θ > 10 (rank : 131) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8NDY6 | Gene names | BHLHB4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Class B basic helix-loop-helix protein 4 (bHLHB4). | |||||
|
BHLH4_MOUSE
|
||||||
| NC score | 0.054077 (rank : 126) | θ value | θ > 10 (rank : 132) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8BGW3, Q71MB7 | Gene names | Bhlhb4, H20 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Class B basic helix-loop-helix protein 4 (bHLHB4). | |||||
|
OLIG2_MOUSE
|
||||||
| NC score | 0.052336 (rank : 127) | θ value | θ > 10 (rank : 171) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9EQW6, Q9JKN4 | Gene names | Olig2 | |||
|
Domain Architecture |

|
|||||
| Description | Oligodendrocyte transcription factor 2 (Oligo2). | |||||
|
OLIG2_HUMAN
|
||||||
| NC score | 0.051870 (rank : 128) | θ value | θ > 10 (rank : 170) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q13516, Q86X04, Q9NZ14 | Gene names | OLIG2, BHLHB1, PRKCBP2, RACK17 | |||
|
Domain Architecture |

|
|||||
| Description | Oligodendrocyte transcription factor 2 (Oligo2) (Basic helix-loop- helix protein class B 1) (Protein kinase C-binding protein RACK17) (Protein kinase C-binding protein 2). | |||||
|
GDIB_HUMAN
|
||||||
| NC score | 0.051675 (rank : 129) | θ value | 2.36792 (rank : 84) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P50395, O43928, Q9UQM6 | Gene names | GDI2, RABGDIB | |||
|
Domain Architecture |

|
|||||
| Description | Rab GDP dissociation inhibitor beta (Rab GDI beta) (Guanosine diphosphate dissociation inhibitor 2) (GDI-2). | |||||
|
HEY1_MOUSE
|
||||||
| NC score | 0.051601 (rank : 130) | θ value | θ > 10 (rank : 140) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9WV93 | Gene names | Hey1, Hesr1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1). | |||||
|
BHLH3_HUMAN
|
||||||
| NC score | 0.051119 (rank : 131) | θ value | θ > 10 (rank : 130) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9C0J9 | Gene names | BHLHB3, DEC2, SHARP1 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (hDEC2) (Enhancer-of-split and hairy-related protein 1) (SHARP-1). | |||||
|
HEY1_HUMAN
|
||||||
| NC score | 0.050366 (rank : 132) | θ value | θ > 10 (rank : 139) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9Y5J3, Q5TZS3, Q9NYP4 | Gene names | HEY1, CHF2, HESR1 | |||
|
Domain Architecture |

|
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 1 (Hairy and enhancer of split-related protein 1) (HESR-1) (Cardiovascular helix-loop-helix factor 2) (HES-related repressor protein 2) (HERP2). | |||||
|
OLIG1_HUMAN
|
||||||
| NC score | 0.050117 (rank : 133) | θ value | θ > 10 (rank : 169) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8TAK6, Q7RTS0 | Gene names | OLIG1 | |||
|
Domain Architecture |

|
|||||
| Description | Oligodendrocyte transcription factor 1 (Oligo1). | |||||
|
DEND_MOUSE
|
||||||
| NC score | 0.046402 (rank : 134) | θ value | 0.21417 (rank : 51) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80TS7 | Gene names | Ddn, Kiaa0749 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dendrin. | |||||
|
ACINU_HUMAN
|
||||||
| NC score | 0.042919 (rank : 135) | θ value | 1.38821 (rank : 76) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9UKV3, O75158, Q9UG91, Q9UKV1, Q9UKV2 | Gene names | ACIN1, ACINUS, KIAA0670 | |||
|
Domain Architecture |

|
|||||
| Description | Apoptotic chromatin condensation inducer in the nucleus (Acinus). | |||||
|
FGFP3_HUMAN
|
||||||
| NC score | 0.039515 (rank : 136) | θ value | 1.38821 (rank : 78) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8TAT2, Q8NBN0 | Gene names | FGFBP3, C10orf13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fibroblast growth factor-binding protein 3 precursor (FGF-binding protein 3) (FGF-BP3) (FGFBP-3). | |||||
|
OGFR_HUMAN
|
||||||
| NC score | 0.038704 (rank : 137) | θ value | 0.0563607 (rank : 37) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
NFH_MOUSE
|
||||||
| NC score | 0.038134 (rank : 138) | θ value | 2.44474e-05 (rank : 13) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
P80C_HUMAN
|
||||||
| NC score | 0.032952 (rank : 139) | θ value | 3.0926 (rank : 93) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P38432 | Gene names | COIL, CLN80 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coilin (p80). | |||||
|
NOL6_MOUSE
|
||||||
| NC score | 0.032562 (rank : 140) | θ value | 0.62314 (rank : 66) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8R5K4, Q6PAN8, Q8C6V4, Q8R5K3, Q8VCG0, Q8WTY7 | Gene names | Nol6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleolar protein 6 (Nucleolar RNA-associated protein) (Nrap). | |||||
|
CS007_MOUSE
|
||||||
| NC score | 0.031409 (rank : 141) | θ value | 1.06291 (rank : 74) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 464 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6ZPZ3 | Gene names | Kiaa1064 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein C19orf7 homolog. | |||||
|
TB182_HUMAN
|
||||||
| NC score | 0.030419 (rank : 142) | θ value | 5.27518 (rank : 105) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9C0C2 | Gene names | TNKS1BP1, KIAA1741, TAB182 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 182 kDa tankyrase 1-binding protein. | |||||
|
K0310_HUMAN
|
||||||
| NC score | 0.029809 (rank : 143) | θ value | 4.03905 (rank : 96) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O15027, Q5SXP0, Q5SXP1, Q8N347, Q96HP1 | Gene names | KIAA0310 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0310. | |||||
|
EPN1_HUMAN
|
||||||
| NC score | 0.028102 (rank : 144) | θ value | 6.88961 (rank : 112) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y6I3, Q86ST3, Q9HA18 | Gene names | EPN1 | |||
|
Domain Architecture |

|
|||||
| Description | Epsin-1 (EPS-15-interacting protein 1) (EH domain-binding mitotic phosphoprotein). | |||||
|
EPN3_MOUSE
|
||||||
| NC score | 0.027184 (rank : 145) | θ value | 0.813845 (rank : 67) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q91W69, Q9CV55 | Gene names | Epn3 | |||
|
Domain Architecture |

|
|||||
| Description | Epsin-3 (EPS-15-interacting protein 3). | |||||
|
SPIB_MOUSE
|
||||||
| NC score | 0.027163 (rank : 146) | θ value | 3.0926 (rank : 94) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O35906, O35907, O35909, O55199 | Gene names | Spib | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor Spi-B. | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.026337 (rank : 147) | θ value | 8.99809 (rank : 121) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
SDPR_MOUSE
|
||||||
| NC score | 0.025945 (rank : 148) | θ value | 5.27518 (rank : 104) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 485 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q63918, Q3V1P6, Q78EC3, Q8CBT4 | Gene names | Sdpr, Sdr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serum deprivation-response protein (Phosphatidylserine-binding protein). | |||||
|
CUTL1_HUMAN
|
||||||
| NC score | 0.023569 (rank : 149) | θ value | 3.0926 (rank : 90) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1574 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P39880, Q6NYH4, Q9UEV5 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP). | |||||
|
CUTL1_MOUSE
|
||||||
| NC score | 0.022399 (rank : 150) | θ value | 2.36792 (rank : 83) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P53564, O08994, P70301, Q571L6, Q91ZD2 | Gene names | Cutl1, Cux, Cux1, Kiaa4047 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP) (Homeobox protein Cux). | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.021151 (rank : 151) | θ value | 6.88961 (rank : 117) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
CASP_MOUSE
|
||||||
| NC score | 0.020284 (rank : 152) | θ value | 2.36792 (rank : 82) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1152 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P70403, Q7TQJ4, Q91WN6 | Gene names | Cutl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein CASP. | |||||
|
CASP_HUMAN
|
||||||
| NC score | 0.019595 (rank : 153) | θ value | 3.0926 (rank : 88) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1131 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q13948, Q53GU9, Q8TBS3 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein CASP. | |||||
|
IRS2_MOUSE
|
||||||
| NC score | 0.018518 (rank : 154) | θ value | 6.88961 (rank : 114) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P81122 | Gene names | Irs2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Insulin receptor substrate 2 (IRS-2) (4PS). | |||||
|
PCLO_MOUSE
|
||||||
| NC score | 0.016560 (rank : 155) | θ value | 8.99809 (rank : 124) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
OSBP2_HUMAN
|
||||||
| NC score | 0.015762 (rank : 156) | θ value | 6.88961 (rank : 115) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q969R2, O60396, Q9BY96, Q9BZF0 | Gene names | OSBP2, KIAA1664, ORP4, OSBPL4 | |||
|
Domain Architecture |

|
|||||
| Description | Oxysterol-binding protein 2 (Oxysterol-binding protein-related protein 4) (OSBP-related protein 4) (ORP-4). | |||||
|
RHG05_HUMAN
|
||||||
| NC score | 0.015626 (rank : 157) | θ value | 5.27518 (rank : 100) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q13017 | Gene names | ARHGAP5, RHOGAP5 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 5 (p190-B). | |||||
|
CDS1_HUMAN
|
||||||
| NC score | 0.015286 (rank : 158) | θ value | 3.0926 (rank : 89) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q92903, O00163 | Gene names | CDS1, CDS | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidate cytidylyltransferase 1 (EC 2.7.7.41) (CDP-diglyceride synthetase 1) (CDP-diglyceride pyrophosphorylase 1) (CDP- diacylglycerol synthase 1) (CDS 1) (CTP:phosphatidate cytidylyltransferase 1) (CDP-DAG synthase 1) (CDP-DG synthetase 1). | |||||
|
RHG05_MOUSE
|
||||||
| NC score | 0.015180 (rank : 159) | θ value | 5.27518 (rank : 101) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P97393 | Gene names | Arhgap5, Rhogap5 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 5 (p190-B). | |||||
|
SMC4_HUMAN
|
||||||
| NC score | 0.014867 (rank : 160) | θ value | 2.36792 (rank : 87) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 937 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9NTJ3, O95752, Q8NDL4, Q9UNT9 | Gene names | SMC4L1, CAPC, SMC4 | |||
|
Domain Architecture |
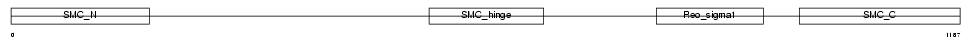
|
|||||
| Description | Structural maintenance of chromosomes 4-like 1 protein (Chromosome- associated polypeptide C) (hCAP-C) (XCAP-C homolog). | |||||
|
ATS13_HUMAN
|
||||||
| NC score | 0.014809 (rank : 161) | θ value | 0.21417 (rank : 50) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q76LX8, Q6UY16, Q710F6, Q711T8, Q96L37, Q9H0G3 | Gene names | ADAMTS13, C9orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAMTS-13 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase with thrombospondin motifs 13) (ADAM-TS 13) (ADAM-TS13) (von Willebrand factor-cleaving protease) (vWF-cleaving protease) (vWF-CP). | |||||
|
WASF2_HUMAN
|
||||||
| NC score | 0.014698 (rank : 162) | θ value | 5.27518 (rank : 108) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y6W5, O60794, Q9UDY7 | Gene names | WASF2, WAVE2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein family member 2 (WASP-family protein member 2) (Protein WAVE-2) (Verprolin homology domain-containing protein 2). | |||||
|
PHLPP_HUMAN
|
||||||
| NC score | 0.014047 (rank : 163) | θ value | 1.81305 (rank : 81) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 604 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O60346, Q641Q7, Q6P4C4, Q6PJI6, Q86TN6, Q96FK2, Q9NUY1 | Gene names | PHLPP, KIAA0606, PLEKHE1, SCOP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH domain leucine-rich repeat-containing protein phosphatase (EC 3.1.3.16) (PH domain leucine-rich repeat protein phosphatase) (Pleckstrin homology domain-containing family E protein 1) (Suprachiasmatic nucleus circadian oscillatory protein) (hSCOP). | |||||
|
TRIPB_HUMAN
|
||||||
| NC score | 0.013853 (rank : 164) | θ value | 5.27518 (rank : 106) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1587 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q15643, O14689, O15154, O95949 | Gene names | TRIP11, CEV14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid receptor-interacting protein 11 (TRIP-11) (Golgi-associated microtubule-binding protein 210) (GMAP-210) (Trip230) (Clonal evolution-related gene on chromosome 14). | |||||
|
MYH6_HUMAN
|
||||||
| NC score | 0.013277 (rank : 165) | θ value | 1.38821 (rank : 80) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1426 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P13533, Q13943, Q14906, Q14907 | Gene names | MYH6, MYHCA | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
MYH7_HUMAN
|
||||||
| NC score | 0.013213 (rank : 166) | θ value | 2.36792 (rank : 85) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P12883, Q14836, Q14837, Q14904, Q16579, Q92679, Q9H1D5, Q9UMM8 | Gene names | MYH7, MYHCB | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle beta isoform (MyHC-beta). | |||||
|
PEPL_MOUSE
|
||||||
| NC score | 0.012843 (rank : 167) | θ value | 2.36792 (rank : 86) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1105 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9R269, O70231, Q9CUT1, Q9JLZ7 | Gene names | Ppl | |||
|
Domain Architecture |

|
|||||
| Description | Periplakin. | |||||
|
MYH6_MOUSE
|
||||||
| NC score | 0.012532 (rank : 168) | θ value | 4.03905 (rank : 97) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1501 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q02566, Q64258, Q64738 | Gene names | Myh6, Myhca | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
CO002_HUMAN
|
||||||
| NC score | 0.012487 (rank : 169) | θ value | 6.88961 (rank : 110) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NZP6 | Gene names | C15orf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C15orf2. | |||||
|
NPM2_MOUSE
|
||||||
| NC score | 0.012090 (rank : 170) | θ value | 8.99809 (rank : 123) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q80W85, Q8BW23 | Gene names | Npm2 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoplasmin-2. | |||||
|
ETV3_HUMAN
|
||||||
| NC score | 0.011677 (rank : 171) | θ value | 8.99809 (rank : 118) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P41162, Q8TAC8, Q9BX30 | Gene names | ETV3, METS, PE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
IL16_HUMAN
|
||||||
| NC score | 0.011426 (rank : 172) | θ value | 6.88961 (rank : 113) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q14005, Q16435, Q9UP18 | Gene names | IL16 | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin-16 precursor (IL-16) (Lymphocyte chemoattractant factor) (LCF). | |||||
|
PTN13_MOUSE
|
||||||
| NC score | 0.010609 (rank : 173) | θ value | 0.279714 (rank : 57) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 426 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q64512, Q61494, Q62135, Q64499 | Gene names | Ptpn13, Ptp14 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 13 (EC 3.1.3.48) (Protein tyrosine phosphatase PTP-BL) (Protein-tyrosine phosphatase RIP) (protein tyrosine phosphatase DPZPTP) (PTP36). | |||||
|
CEL_HUMAN
|
||||||
| NC score | 0.009512 (rank : 174) | θ value | 6.88961 (rank : 109) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P19835, Q16398 | Gene names | CEL, BAL | |||
|
Domain Architecture |

|
|||||
| Description | Bile salt-activated lipase precursor (EC 3.1.1.3) (EC 3.1.1.13) (BAL) (Bile salt-stimulated lipase) (BSSL) (Carboxyl ester lipase) (Sterol esterase) (Cholesterol esterase) (Pancreatic lysophospholipase). | |||||
|
CSKI1_MOUSE
|
||||||
| NC score | 0.008247 (rank : 175) | θ value | 6.88961 (rank : 111) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1031 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6P9K8, Q6ZPU2, Q8BWU2, Q8BX99, Q9CXH0 | Gene names | Caskin1, Kiaa1306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-1 (CASK-interacting protein 1). | |||||
|
FOG1_MOUSE
|
||||||
| NC score | 0.008176 (rank : 176) | θ value | 3.0926 (rank : 91) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1237 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O35615 | Gene names | Zfpm1, Fog, Fog1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein ZFPM1 (Zinc finger protein multitype 1) (Friend of GATA protein 1) (Friend of GATA-1) (FOG-1). | |||||
|
MAGD4_HUMAN
|
||||||
| NC score | 0.006184 (rank : 177) | θ value | 8.99809 (rank : 122) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 338 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q96JG8, Q8WXW4, Q9BQ84, Q9BYH3, Q9BYH4 | Gene names | MAGED4, KIAA1859, MAGEE1 | |||
|
Domain Architecture |

|
|||||
| Description | Melanoma-associated antigen D4 (MAGE-D4 antigen) (MAGE-E1 antigen). | |||||
|
POT15_HUMAN
|
||||||
| NC score | 0.003741 (rank : 178) | θ value | 6.88961 (rank : 116) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 893 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6S5H4, Q6NXN7, Q6S5H7 | Gene names | POTE15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prostate, ovary, testis-expressed protein on chromosome 15. | |||||
|
TTBK2_MOUSE
|
||||||
| NC score | 0.003709 (rank : 179) | θ value | 5.27518 (rank : 107) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 811 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q3UVR3, Q3TSR6, Q3UFW0, Q571D1, Q8BKA4, Q924U8 | Gene names | Ttbk2, Kiaa0847, Ttbk1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tau-tubulin kinase 2. | |||||
|
TTBK2_HUMAN
|
||||||
| NC score | 0.003334 (rank : 180) | θ value | 8.99809 (rank : 125) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 822 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6IQ55, O94932, Q6ZN52, Q8IVV1 | Gene names | TTBK2, KIAA0847 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tau-tubulin kinase 2 (EC 2.7.11.1). | |||||
|
SALL4_HUMAN
|
||||||
| NC score | 0.001791 (rank : 181) | θ value | 5.27518 (rank : 103) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 799 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UJQ4 | Gene names | SALL4 | |||
|
Domain Architecture |

|
|||||
| Description | Sal-like protein 4 (Zinc finger protein SALL4). | |||||
|
HXB3_MOUSE
|
||||||
| NC score | 0.001649 (rank : 182) | θ value | 5.27518 (rank : 99) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 493 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P09026, P10285, Q4PJ06, Q61680 | Gene names | Hoxb3, Hox-2.7, Hoxb-3 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-B3 (Hox-2.7) (Homeobox protein MH-23). | |||||
|
SALL1_MOUSE
|
||||||
| NC score | 0.001315 (rank : 183) | θ value | 5.27518 (rank : 102) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 826 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9ER74, Q920R5 | Gene names | Sall1, Sal3 | |||
|
Domain Architecture |

|
|||||
| Description | Sal-like protein 1 (Zinc finger protein Spalt-3) (Sal-3) (MSal-3). | |||||
|
ZN142_HUMAN
|
||||||
| NC score | 0.000066 (rank : 184) | θ value | 4.03905 (rank : 98) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1007 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P52746, Q92510 | Gene names | ZNF142, KIAA0236 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 142 (HA4654). | |||||
|
CAMKV_HUMAN
|
||||||
| NC score | -0.000340 (rank : 185) | θ value | 4.03905 (rank : 95) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 1084 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8NCB2, Q6FIB8, Q8NBS8, Q8NC85, Q8NDU4, Q8WTT8, Q9BQC9, Q9H0Q5 | Gene names | CAMKV | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CaM kinase-like vesicle-associated protein. | |||||
|
ZFP28_HUMAN
|
||||||
| NC score | -0.000950 (rank : 186) | θ value | 0.813845 (rank : 73) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 843 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8NHY6, Q9BY30, Q9P2B6 | Gene names | ZFP28, KIAA1431 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 28 homolog (Zfp-28) (Krueppel-like zinc finger factor X6). | |||||
|
ZN275_HUMAN
|
||||||
| NC score | -0.002311 (rank : 187) | θ value | 8.99809 (rank : 127) | |||
| Query Neighborhood Hits | 127 | Target Neighborhood Hits | 761 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9NSD4 | Gene names | ZNF275 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 275. | |||||