Please be patient as the page loads
|
MAD3_MOUSE
|
||||||
| SwissProt Accessions | Q80US8, Q60947 | Gene names | Mxd3, Mad3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Max-interacting transcriptional repressor MAD3 (Max-associated protein 3) (MAX dimerization protein 3). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
MAD3_MOUSE
|
||||||
| θ value | 9.0981e-77 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 94 | |
| SwissProt Accessions | Q80US8, Q60947 | Gene names | Mxd3, Mad3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Max-interacting transcriptional repressor MAD3 (Max-associated protein 3) (MAX dimerization protein 3). | |||||
|
MAD3_HUMAN
|
||||||
| θ value | 4.24587e-58 (rank : 2) | NC score | 0.972694 (rank : 2) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9BW11, Q53HK1, Q7Z4Y0, Q8NDJ7, Q96ME3 | Gene names | MXD3, MAD3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Max-interacting transcriptional repressor MAD3 (Max-associated protein 3) (MAX dimerization protein 3) (Myx). | |||||
|
MAD1_MOUSE
|
||||||
| θ value | 3.17079e-29 (rank : 3) | NC score | 0.893348 (rank : 5) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P50538, Q60798, Q61825 | Gene names | Mxd1, Mad, Mad1 | |||
|
Domain Architecture |

|
|||||
| Description | MAD protein (MAX dimerizer) (MAX dimerization protein 1). | |||||
|
MAD4_HUMAN
|
||||||
| θ value | 1.12775e-26 (rank : 4) | NC score | 0.889295 (rank : 8) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q14582, Q5TZX4 | Gene names | MXD4, MAD4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-interacting transcriptional repressor MAD4 (Max-associated protein 4) (MAX dimerization protein 4). | |||||
|
MAD1_HUMAN
|
||||||
| θ value | 3.28125e-26 (rank : 5) | NC score | 0.890323 (rank : 7) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q05195 | Gene names | MXD1, MAD | |||
|
Domain Architecture |
|
|||||
| Description | MAD protein (MAX dimerizer) (MAX dimerization protein 1). | |||||
|
MAD4_MOUSE
|
||||||
| θ value | 1.24688e-25 (rank : 6) | NC score | 0.892869 (rank : 6) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q60948 | Gene names | Mxd4, Mad4 | |||
|
Domain Architecture |
|
|||||
| Description | Max-interacting transcriptional repressor MAD4 (Max-associated protein 4) (MAX dimerization protein 4). | |||||
|
MXI1_MOUSE
|
||||||
| θ value | 3.07116e-24 (rank : 7) | NC score | 0.898064 (rank : 4) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P50540 | Gene names | Mxi1 | |||
|
Domain Architecture |
|
|||||
| Description | MAX-interacting protein 1 (Protein MXI1). | |||||
|
MXI1_HUMAN
|
||||||
| θ value | 6.84181e-24 (rank : 8) | NC score | 0.903462 (rank : 3) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P50539, Q15887 | Gene names | MXI1 | |||
|
Domain Architecture |
|
|||||
| Description | MAX-interacting protein 1 (Protein MXI1). | |||||
|
MNT_HUMAN
|
||||||
| θ value | 1.133e-10 (rank : 9) | NC score | 0.512521 (rank : 10) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 496 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q99583 | Gene names | MNT, ROX | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
MNT_MOUSE
|
||||||
| θ value | 4.30538e-10 (rank : 10) | NC score | 0.530833 (rank : 9) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 432 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O08789, P97349 | Gene names | Mnt, Rox | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
MYC_MOUSE
|
||||||
| θ value | 1.29631e-06 (rank : 11) | NC score | 0.379278 (rank : 11) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P01108, P70247, Q3UM70, Q61422 | Gene names | Myc | |||
|
Domain Architecture |
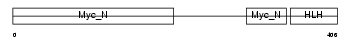
|
|||||
| Description | Myc proto-oncogene protein (c-Myc) (Transcription factor p64). | |||||
|
MYC_HUMAN
|
||||||
| θ value | 2.88788e-06 (rank : 12) | NC score | 0.376419 (rank : 12) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P01106, P01107, Q14026 | Gene names | MYC | |||
|
Domain Architecture |

|
|||||
| Description | Myc proto-oncogene protein (c-Myc) (Transcription factor p64). | |||||
|
MYCL2_HUMAN
|
||||||
| θ value | 0.000461057 (rank : 13) | NC score | 0.311234 (rank : 14) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P12525 | Gene names | MYCL2 | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-2 protein. | |||||
|
USF2_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 14) | NC score | 0.305938 (rank : 15) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q64705, Q3UIL2, Q8VE59 | Gene names | Usf2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (Major late transcription factor 2). | |||||
|
TFE3_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 15) | NC score | 0.252733 (rank : 22) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P19532, Q92757, Q92758, Q99964 | Gene names | TFE3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E3. | |||||
|
USF2_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 16) | NC score | 0.305783 (rank : 17) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q15853, O00671, O00709, Q05750, Q07952, Q15851, Q15852 | Gene names | USF2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (FOS- interacting protein) (FIP) (Major late transcription factor 2). | |||||
|
GOGA1_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 17) | NC score | 0.049651 (rank : 67) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1222 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q92805, Q8IYZ9 | Gene names | GOLGA1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 1 (Golgin-97). | |||||
|
AZI1_HUMAN
|
||||||
| θ value | 0.125558 (rank : 18) | NC score | 0.062893 (rank : 51) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1000 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9UPN4, Q96F50 | Gene names | AZI1, CEP131, KIAA1118 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 5-azacytidine-induced protein 1 (Pre-acrosome localization protein 1) (Centrosomal protein of 131 kDa) (Cep131 protein). | |||||
|
CCD96_HUMAN
|
||||||
| θ value | 0.125558 (rank : 19) | NC score | 0.075360 (rank : 42) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 595 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q2M329, Q8N2I7 | Gene names | CCDC96 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 96. | |||||
|
MITF_HUMAN
|
||||||
| θ value | 0.125558 (rank : 20) | NC score | 0.234482 (rank : 25) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O75030, Q14841, Q9P2V0, Q9P2V1, Q9P2V2, Q9P2Y8 | Gene names | MITF | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
MAX_HUMAN
|
||||||
| θ value | 0.21417 (rank : 21) | NC score | 0.312590 (rank : 13) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P61244, P25912, P52163 | Gene names | MAX | |||
|
Domain Architecture |

|
|||||
| Description | Protein max (Myc-associated factor X). | |||||
|
MAX_MOUSE
|
||||||
| θ value | 0.21417 (rank : 22) | NC score | 0.305815 (rank : 16) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P28574 | Gene names | Max, Myn | |||
|
Domain Architecture |

|
|||||
| Description | Protein max (Myc-associated factor X) (Protein myn) (Myc-binding novel HLH/LZ protein). | |||||
|
MITF_MOUSE
|
||||||
| θ value | 0.279714 (rank : 23) | NC score | 0.225053 (rank : 26) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q08874, O08885, O88203, Q08843, Q60781, Q60782, Q9JIJ0, Q9JIJ1, Q9JIJ2, Q9JIJ3, Q9JIJ4, Q9JIJ5, Q9JIJ6, Q9JKX9 | Gene names | Mitf, Bw, Mi, Vit | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
BPAEA_MOUSE
|
||||||
| θ value | 0.365318 (rank : 24) | NC score | 0.053198 (rank : 62) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1480 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q91ZU8, Q8K5D4 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoform 5 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
MYCL1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 25) | NC score | 0.277945 (rank : 18) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P12524, Q14897, Q5QPL0, Q5QPL1, Q9NUE9 | Gene names | MYCL1, LMYC, MYCL | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
MYCL1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 26) | NC score | 0.274319 (rank : 19) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P10166, Q5FWI7 | Gene names | Mycl1, Lmyc1, Mycl | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
CCD21_HUMAN
|
||||||
| θ value | 0.62314 (rank : 27) | NC score | 0.046689 (rank : 73) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1228 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q6P2H3, Q5VY68, Q5VY70, Q9H6Q1, Q9H828, Q9UF52 | Gene names | CCDC21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 21. | |||||
|
CCD96_MOUSE
|
||||||
| θ value | 0.62314 (rank : 28) | NC score | 0.074697 (rank : 44) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9CR92 | Gene names | Ccdc96 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 96. | |||||
|
GO45_HUMAN
|
||||||
| θ value | 0.62314 (rank : 29) | NC score | 0.061828 (rank : 53) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9H2G9, O15298, Q5T531, Q9GZX4 | Gene names | BLZF1, JEM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgin 45 (Basic leucine zipper nuclear factor 1) (JEM-1) (p45 basic leucine-zipper nuclear factor). | |||||
|
GOGA3_HUMAN
|
||||||
| θ value | 0.62314 (rank : 30) | NC score | 0.049750 (rank : 66) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1317 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q08378, O43241, Q6P9C7, Q86XW3, Q8TDA9, Q8WZA3 | Gene names | GOLGA3 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 3 (Golgin-160) (Golgi complex-associated protein of 170 kDa) (GCP170). | |||||
|
NEK11_HUMAN
|
||||||
| θ value | 0.62314 (rank : 31) | NC score | 0.006915 (rank : 112) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1043 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8NG66, Q5JPC0, Q8NG65, Q8TBY1, Q9H5F4 | Gene names | NEK11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase Nek11 (EC 2.7.11.1) (NimA-related protein kinase 11) (Never in mitosis A-related kinase 11). | |||||
|
SYCP1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 32) | NC score | 0.046872 (rank : 72) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1313 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q15431, O14963 | Gene names | SYCP1, SCP1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptonemal complex protein 1 (SCP-1). | |||||
|
ZWINT_HUMAN
|
||||||
| θ value | 0.62314 (rank : 33) | NC score | 0.068265 (rank : 48) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O95229, Q9BWD0 | Gene names | ZWINT | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ZW10 interactor (ZW10-interacting protein 1) (Zwint-1). | |||||
|
USF1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 34) | NC score | 0.272051 (rank : 21) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P22415 | Gene names | USF1, USF | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 35) | NC score | 0.273335 (rank : 20) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q61069 | Gene names | Usf1, Usf | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
TFEB_HUMAN
|
||||||
| θ value | 1.06291 (rank : 36) | NC score | 0.194375 (rank : 29) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P19484, Q709B3, Q7Z6P9, Q9BRJ5, Q9UJD8 | Gene names | TFEB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
TFEB_MOUSE
|
||||||
| θ value | 1.06291 (rank : 37) | NC score | 0.194557 (rank : 28) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9R210 | Gene names | Tfeb, Tcfeb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
MYCN_HUMAN
|
||||||
| θ value | 1.38821 (rank : 38) | NC score | 0.248938 (rank : 24) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P04198, Q6LDT9 | Gene names | MYCN, NMYC | |||
|
Domain Architecture |

|
|||||
| Description | N-myc proto-oncogene protein. | |||||
|
RCOR3_HUMAN
|
||||||
| θ value | 1.38821 (rank : 39) | NC score | 0.037369 (rank : 88) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9P2K3, Q5VT47, Q7L9I5, Q8N5U3, Q9NV83 | Gene names | RCOR3, KIAA1343 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 3. | |||||
|
RCOR3_MOUSE
|
||||||
| θ value | 1.38821 (rank : 40) | NC score | 0.039673 (rank : 80) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6PGA0 | Gene names | Rcor3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 3. | |||||
|
BHLH2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 41) | NC score | 0.109046 (rank : 37) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O14503, Q96TD3 | Gene names | BHLHB2, DEC1, SHARP2, STRA13 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 2 (bHLHB2) (Differentially expressed in chondrocytes protein 1) (DEC1) (Enhancer-of-split and hairy-related protein 2) (SHARP-2) (Stimulated with retinoic acid 13). | |||||
|
BMAL1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 42) | NC score | 0.075101 (rank : 43) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O00327, O00313, O00314, O00315, O00316, O00317, Q99631, Q99649 | Gene names | ARNTL, BMAL1, MOP3 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Member of PAS protein 3) (Basic-helix-loop- helix-PAS orphan MOP3) (bHLH-PAS protein JAP3). | |||||
|
BMAL1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 43) | NC score | 0.074061 (rank : 46) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9WTL8, O88295, Q921S4, Q9R0U2, Q9WTL9 | Gene names | Arntl, Bmal1 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Arnt3). | |||||
|
CENPF_HUMAN
|
||||||
| θ value | 1.81305 (rank : 44) | NC score | 0.046015 (rank : 75) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1932 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P49454, Q13171, Q13246 | Gene names | CENPF | |||
|
Domain Architecture |

|
|||||
| Description | Centromere protein F (Kinetochore protein CENP-F) (Mitosin) (AH antigen). | |||||
|
PCNT_MOUSE
|
||||||
| θ value | 1.81305 (rank : 45) | NC score | 0.045229 (rank : 77) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1207 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P48725 | Gene names | Pcnt, Pcnt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pericentrin. | |||||
|
RABE2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 46) | NC score | 0.058226 (rank : 57) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 598 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q91WG2, Q99KN3 | Gene names | Rabep2, Rabpt5b | |||
|
Domain Architecture |

|
|||||
| Description | Rab GTPase-binding effector protein 2 (Rabaptin-5beta). | |||||
|
BHLH2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 47) | NC score | 0.119760 (rank : 35) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O35185, P97289 | Gene names | Bhlhb2, Clast5, Stra13 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 2 (bHLHB2) (Stimulated with retinoic acid 13) (E47 interaction protein 1) (eipl). | |||||
|
BICD2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 48) | NC score | 0.049786 (rank : 65) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 856 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8TD16, O75181, Q5TBQ2, Q5TBQ3, Q96LH2, Q9BT84, Q9H561 | Gene names | BICD2, KIAA0699 | |||
|
Domain Architecture |
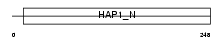
|
|||||
| Description | Cytoskeleton-like bicaudal D protein homolog 2. | |||||
|
BICD2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 49) | NC score | 0.049607 (rank : 68) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 864 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q921C5, Q80TU1, Q8BTE3, Q9DCL3 | Gene names | Bicd2, Kiaa0699 | |||
|
Domain Architecture |
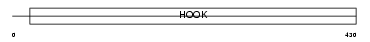
|
|||||
| Description | Cytoskeleton-like bicaudal D protein homolog 2. | |||||
|
CCD62_HUMAN
|
||||||
| θ value | 2.36792 (rank : 50) | NC score | 0.048059 (rank : 70) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 631 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q6P9F0, Q6ZVF2, Q86VJ0, Q9BYZ5 | Gene names | CCDC62 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 60 (Aaa-protein) (Protein TSP- NY). | |||||
|
HIP1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 51) | NC score | 0.046233 (rank : 74) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1016 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | O00291, O00328, Q8TDL4 | Gene names | HIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin-interacting protein 1 (HIP-I). | |||||
|
NGN1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 52) | NC score | 0.023147 (rank : 105) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P70660 | Gene names | Neurog1, Ath4c, Neurod3, Ngn, Ngn1 | |||
|
Domain Architecture |
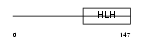
|
|||||
| Description | Neurogenin 1 (Neurogenic differentiation factor 3) (NeuroD3) (Neurogenic basic-helix-loop-helix protein) (Helix-loop-helix protein mATH-4C) (MATH4C). | |||||
|
CCDC6_HUMAN
|
||||||
| θ value | 3.0926 (rank : 53) | NC score | 0.057834 (rank : 58) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 723 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q16204, Q15250 | Gene names | CCDC6, D10S170 | |||
|
Domain Architecture |
|
|||||
| Description | Coiled-coil domain-containing protein 6 (H4 protein) (Papillary thyroid carcinoma-encoded protein). | |||||
|
DZIP1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 54) | NC score | 0.042552 (rank : 78) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 530 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8BMD2, Q6ZQ10, Q9CRK5 | Gene names | Dzip1, Kiaa0996 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein DZIP1 (DAZ-interacting protein 1 homolog). | |||||
|
MYCN_MOUSE
|
||||||
| θ value | 3.0926 (rank : 55) | NC score | 0.250521 (rank : 23) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P03966, Q61978 | Gene names | Mycn, Nmyc, Nmyc1 | |||
|
Domain Architecture |

|
|||||
| Description | N-myc proto-oncogene protein. | |||||
|
STX1A_HUMAN
|
||||||
| θ value | 3.0926 (rank : 56) | NC score | 0.035376 (rank : 96) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q16623, O15447, O15448, Q12936, Q9BPZ6 | Gene names | STX1A, STX1 | |||
|
Domain Architecture |

|
|||||
| Description | Syntaxin-1A (Neuron-specific antigen HPC-1). | |||||
|
STX1A_MOUSE
|
||||||
| θ value | 3.0926 (rank : 57) | NC score | 0.035579 (rank : 94) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O35526 | Gene names | Stx1a | |||
|
Domain Architecture |

|
|||||
| Description | Syntaxin-1A (Neuron-specific antigen HPC-1). | |||||
|
ATOH1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 58) | NC score | 0.038301 (rank : 86) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q92858 | Gene names | ATOH1, ATH1 | |||
|
Domain Architecture |

|
|||||
| Description | Atonal protein homolog 1 (Helix-loop-helix protein hATH-1). | |||||
|
ATOH1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 59) | NC score | 0.038342 (rank : 85) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P48985 | Gene names | Atoh1, Ath1 | |||
|
Domain Architecture |

|
|||||
| Description | Atonal protein homolog 1 (Helix-loop-helix protein mATH-1) (MATH1). | |||||
|
CCD16_HUMAN
|
||||||
| θ value | 4.03905 (rank : 60) | NC score | 0.021043 (rank : 106) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96NB3, Q96F60, Q96GZ5, Q9BU38 | Gene names | CCDC16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 16. | |||||
|
CING_MOUSE
|
||||||
| θ value | 4.03905 (rank : 61) | NC score | 0.049070 (rank : 69) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1432 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | P59242 | Gene names | Cgn | |||
|
Domain Architecture |

|
|||||
| Description | Cingulin. | |||||
|
DESP_HUMAN
|
||||||
| θ value | 4.03905 (rank : 62) | NC score | 0.047607 (rank : 71) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P15924, O75993, Q14189, Q9UHN4 | Gene names | DSP | |||
|
Domain Architecture |

|
|||||
| Description | Desmoplakin (DP) (250/210 kDa paraneoplastic pemphigus antigen). | |||||
|
GO45_MOUSE
|
||||||
| θ value | 4.03905 (rank : 63) | NC score | 0.055737 (rank : 59) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8R2X8, Q3TJC4, Q8C0U6, Q9DAV7 | Gene names | Blzf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgin 45 (Basic leucine zipper nuclear factor 1). | |||||
|
LIPB2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 64) | NC score | 0.038693 (rank : 82) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 570 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O35711, Q7TMG4, Q8CBS6, Q99KX6 | Gene names | Ppfibp2, Cclp1 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-beta-2 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein-binding protein 2) (PTPRF-interacting protein-binding protein 2) (Coiled-coil-like protein 1). | |||||
|
ANR26_MOUSE
|
||||||
| θ value | 5.27518 (rank : 65) | NC score | 0.031831 (rank : 103) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1554 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q811D2, Q69ZS2, Q9CS61 | Gene names | Ankrd26, Kiaa1074 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 26. | |||||
|
BRPF1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 66) | NC score | 0.013385 (rank : 111) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P55201, Q9UHI0 | Gene names | BRPF1, BR140 | |||
|
Domain Architecture |

|
|||||
| Description | Peregrin (Bromodomain and PHD finger-containing protein 1) (BR140 protein). | |||||
|
CTRO_MOUSE
|
||||||
| θ value | 5.27518 (rank : 67) | NC score | 0.019476 (rank : 108) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 2344 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P49025, O88528, O88937, O88938, Q3UM99, Q8CIJ1 | Gene names | Cit | |||
|
Domain Architecture |

|
|||||
| Description | Citron Rho-interacting kinase (EC 2.7.11.1) (CRIK) (Rho-interacting, serine/threonine-protein kinase 21). | |||||
|
EP15R_HUMAN
|
||||||
| θ value | 5.27518 (rank : 68) | NC score | 0.037661 (rank : 87) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1083 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9UBC2 | Gene names | EPS15L1, EPS15R | |||
|
Domain Architecture |

|
|||||
| Description | Epidermal growth factor receptor substrate 15-like 1 (Eps15-related protein) (Eps15R). | |||||
|
GOGA4_MOUSE
|
||||||
| θ value | 5.27518 (rank : 69) | NC score | 0.038464 (rank : 84) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1939 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q91VW5, O70365, Q8CGH6 | Gene names | Golga4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (tGolgin-1). | |||||
|
OPTN_HUMAN
|
||||||
| θ value | 5.27518 (rank : 70) | NC score | 0.038684 (rank : 83) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 827 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q96CV9, Q5T672, Q5T673, Q5T674, Q5T675, Q7LDL9, Q8N562, Q9UET9, Q9UEV4, Q9Y218 | Gene names | OPTN, FIP2, GLC1E, NRP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Optineurin (Optic neuropathy-inducing protein) (E3-14.7K-interacting protein) (FIP-2) (Huntingtin-interacting protein HYPL) (NEMO-related protein) (Transcription factor IIIA-interacting protein) (TFIIIA- IntP). | |||||
|
SRBP1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 71) | NC score | 0.116597 (rank : 36) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9WTN3 | Gene names | Srebf1, Srebp1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
BHLH3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 72) | NC score | 0.128418 (rank : 32) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9C0J9 | Gene names | BHLHB3, DEC2, SHARP1 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (hDEC2) (Enhancer-of-split and hairy-related protein 1) (SHARP-1). | |||||
|
BICD1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 73) | NC score | 0.045487 (rank : 76) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 844 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q96G01, O43892, O43893 | Gene names | BICD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-like bicaudal D protein homolog 1. | |||||
|
BPA1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 74) | NC score | 0.033976 (rank : 101) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q91ZU6, Q91ZU7 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 1/2/3/4 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
DTBP1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 75) | NC score | 0.019571 (rank : 107) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q96EV8, Q96NV2, Q9H0U2, Q9H3J5 | Gene names | DTNBP1 | |||
|
Domain Architecture |
|
|||||
| Description | Dystrobrevin-binding protein 1 (Dysbindin) (Hermansky-Pudlak syndrome 7 protein). | |||||
|
EP15R_MOUSE
|
||||||
| θ value | 6.88961 (rank : 76) | NC score | 0.036027 (rank : 91) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q60902, Q8CB60, Q8CB70, Q91WH8 | Gene names | Eps15l1, Eps15-rs, Eps15R | |||
|
Domain Architecture |

|
|||||
| Description | Epidermal growth factor receptor substrate 15-like 1 (Eps15-related protein) (Eps15R) (Epidermal growth factor receptor pathway substrate 15-related sequence) (Eps15-rs). | |||||
|
MY18A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 77) | NC score | 0.034392 (rank : 98) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1929 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q92614, Q8IXP8 | Gene names | MYO18A, KIAA0216, MYSPDZ | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
MY18A_MOUSE
|
||||||
| θ value | 6.88961 (rank : 78) | NC score | 0.036029 (rank : 90) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1989 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9JMH9, Q811D7 | Gene names | Myo18a, Myspdz | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
ROCK2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 79) | NC score | 0.017644 (rank : 110) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 2073 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O75116, Q9UQN5 | Gene names | ROCK2, KIAA0619 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 2 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 2) (p164 ROCK-2) (Rho kinase 2). | |||||
|
ROCK2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 80) | NC score | 0.018035 (rank : 109) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 2072 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P70336, Q8BR64, Q8CBR0 | Gene names | Rock2 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 2 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 2) (p164 ROCK-2). | |||||
|
STX1B_HUMAN
|
||||||
| θ value | 6.88961 (rank : 81) | NC score | 0.034049 (rank : 99) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 252 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q15531 | Gene names | STX1B1, STX1B | |||
|
Domain Architecture |

|
|||||
| Description | Syntaxin-1B1 (Syntaxin 1B). | |||||
|
STX1C_HUMAN
|
||||||
| θ value | 6.88961 (rank : 82) | NC score | 0.035633 (rank : 92) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 270 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P61266 | Gene names | STX1B2 | |||
|
Domain Architecture |

|
|||||
| Description | Syntaxin-1B2. | |||||
|
STX1C_MOUSE
|
||||||
| θ value | 6.88961 (rank : 83) | NC score | 0.035633 (rank : 93) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 270 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P61264, P32853 | Gene names | Stx1b2, Stx1b | |||
|
Domain Architecture |

|
|||||
| Description | Syntaxin-1B2 (Syntaxin 1B). | |||||
|
TAXB1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 84) | NC score | 0.040451 (rank : 79) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 886 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q3UKC1, Q91YT6, Q9CVF0, Q9DC45 | Gene names | Tax1bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tax1-binding protein 1 homolog. | |||||
|
ASCL2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 85) | NC score | 0.035505 (rank : 95) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q99929, Q9UM68 | Gene names | ASCL2 | |||
|
Domain Architecture |
|
|||||
| Description | Achaete-scute homolog 2 (Mash2) (HASH2). | |||||
|
AZI1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 86) | NC score | 0.062187 (rank : 52) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 990 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q62036 | Gene names | Azi1, Az1, Azi | |||
|
Domain Architecture |

|
|||||
| Description | 5-azacytidine-induced protein 1 (Pre-acrosome localization protein 1). | |||||
|
BHLH3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 87) | NC score | 0.120488 (rank : 34) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q99PV5 | Gene names | Bhlhb3, Dec2 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (mDEC2). | |||||
|
CASP_HUMAN
|
||||||
| θ value | 8.99809 (rank : 88) | NC score | 0.038697 (rank : 81) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1131 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q13948, Q53GU9, Q8TBS3 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein CASP. | |||||
|
CCD46_MOUSE
|
||||||
| θ value | 8.99809 (rank : 89) | NC score | 0.037068 (rank : 89) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q5PR68, Q80ZN6, Q99JS4, Q9D9T1 | Gene names | Ccdc46 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled coil domain-containing protein 46. | |||||
|
CENPE_HUMAN
|
||||||
| θ value | 8.99809 (rank : 90) | NC score | 0.034474 (rank : 97) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 2060 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q02224 | Gene names | CENPE | |||
|
Domain Architecture |

|
|||||
| Description | Centromeric protein E (CENP-E). | |||||
|
CUTL1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 91) | NC score | 0.033762 (rank : 102) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1574 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P39880, Q6NYH4, Q9UEV5 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP). | |||||
|
K1802_MOUSE
|
||||||
| θ value | 8.99809 (rank : 92) | NC score | 0.006361 (rank : 113) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
MYH13_HUMAN
|
||||||
| θ value | 8.99809 (rank : 93) | NC score | 0.030582 (rank : 104) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1504 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9UKX3, O95252 | Gene names | MYH13 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-13 (Myosin heavy chain, skeletal muscle, extraocular) (MyHC- eo). | |||||
|
MYH9_MOUSE
|
||||||
| θ value | 8.99809 (rank : 94) | NC score | 0.033984 (rank : 100) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1829 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q8VDD5 | Gene names | Myh9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
ARNT2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 95) | NC score | 0.050258 (rank : 64) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q61324 | Gene names | Arnt2 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
ARNT_HUMAN
|
||||||
| θ value | θ > 10 (rank : 96) | NC score | 0.063429 (rank : 49) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P27540 | Gene names | ARNT | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
ARNT_MOUSE
|
||||||
| θ value | θ > 10 (rank : 97) | NC score | 0.055509 (rank : 60) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P53762, Q60661 | Gene names | Arnt | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
BHLH8_HUMAN
|
||||||
| θ value | θ > 10 (rank : 98) | NC score | 0.058843 (rank : 55) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q7RTS1 | Gene names | BHLHB8, MIST1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Class B basic helix-loop-helix protein 8 (bHLHB8) (Muscle, intestine and stomach expression 1) (MIST-1). | |||||
|
BHLH8_MOUSE
|
||||||
| θ value | θ > 10 (rank : 99) | NC score | 0.063400 (rank : 50) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9QYC3, Q9QYE4 | Gene names | Bhlhb8, Mist1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Class B basic helix-loop-helix protein 8 (bHLHB8) (Muscle, intestine and stomach expression 1) (MIST-1). | |||||
|
FIGLA_HUMAN
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.053664 (rank : 61) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6QHK4 | Gene names | FIGLA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Factor in the germline alpha (Transcription factor FIGa) (FIGalpha). | |||||
|
HES1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.081368 (rank : 41) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q14469 | Gene names | HES1, HL, HRY | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor HES-1 (Hairy and enhancer of split 1) (Hairy- like) (HHL) (Hairy homolog). | |||||
|
HES1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.081705 (rank : 40) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P35428 | Gene names | Hes1, Hes-1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-1 (Hairy and enhancer of split 1). | |||||
|
HES4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.058355 (rank : 56) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9HCC6 | Gene names | HES4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-4 (Hairy and enhancer of split 4) (bHLH factor Hes4). | |||||
|
MLX_HUMAN
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.071306 (rank : 47) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UH92, Q53XM6, Q96FL2, Q9H2V0, Q9H2V1, Q9H2V2, Q9NXN3 | Gene names | MLX, TCFL4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
MLX_MOUSE
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.074460 (rank : 45) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O08609, Q9EQJ7, Q9EQJ8 | Gene names | Mlx, Tcfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
MYCB_MOUSE
|
||||||
| θ value | θ > 10 (rank : 106) | NC score | 0.140713 (rank : 31) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6P8Z1, Q9D6E3 | Gene names | Mycb, Bmyc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein B-Myc. | |||||
|
MYCS_MOUSE
|
||||||
| θ value | θ > 10 (rank : 107) | NC score | 0.197739 (rank : 27) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9Z304 | Gene names | Mycs | |||
|
Domain Architecture |
|
|||||
| Description | Protein S-myc. | |||||
|
SRBP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 108) | NC score | 0.094297 (rank : 39) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P36956, Q16062 | Gene names | SREBF1, SREBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
SRBP2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 109) | NC score | 0.126033 (rank : 33) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q12772, Q9UH04 | Gene names | SREBF2, SREBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 2 (SREBP-2) (Sterol regulatory element-binding transcription factor 2). | |||||
|
TCFL5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 110) | NC score | 0.152605 (rank : 30) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9UL49, O94771, Q9BYW0 | Gene names | TCFL5, CHA, E2BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor-like 5 protein (Cha transcription factor) (HPV-16 E2-binding protein 1) (E2BP-1). | |||||
|
TFAP4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 111) | NC score | 0.098781 (rank : 38) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q01664, O60409 | Gene names | TFAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor AP-4 (Activating enhancer-binding protein 4). | |||||
|
TRHY_HUMAN
|
||||||
| θ value | θ > 10 (rank : 112) | NC score | 0.051682 (rank : 63) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1385 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q07283 | Gene names | THH, THL, TRHY | |||
|
Domain Architecture |

|
|||||
| Description | Trichohyalin. | |||||
|
WBS14_HUMAN
|
||||||
| θ value | θ > 10 (rank : 113) | NC score | 0.060392 (rank : 54) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9NP71, Q96E48, Q9BY03, Q9BY04, Q9BY05, Q9BY06, Q9Y2P3 | Gene names | MLXIPL, MIO, WBSCR14 | |||
|
Domain Architecture |
|
|||||
| Description | Williams-Beuren syndrome chromosome region 14 protein (WS basic-helix- loop-helix leucine zipper protein) (WS-bHLH) (Mlx interactor) (MLX- interacting protein-like). | |||||
|
MAD3_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 9.0981e-77 (rank : 1) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 94 | |
| SwissProt Accessions | Q80US8, Q60947 | Gene names | Mxd3, Mad3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Max-interacting transcriptional repressor MAD3 (Max-associated protein 3) (MAX dimerization protein 3). | |||||
|
MAD3_HUMAN
|
||||||
| NC score | 0.972694 (rank : 2) | θ value | 4.24587e-58 (rank : 2) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9BW11, Q53HK1, Q7Z4Y0, Q8NDJ7, Q96ME3 | Gene names | MXD3, MAD3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Max-interacting transcriptional repressor MAD3 (Max-associated protein 3) (MAX dimerization protein 3) (Myx). | |||||
|
MXI1_HUMAN
|
||||||
| NC score | 0.903462 (rank : 3) | θ value | 6.84181e-24 (rank : 8) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P50539, Q15887 | Gene names | MXI1 | |||
|
Domain Architecture |

|
|||||
| Description | MAX-interacting protein 1 (Protein MXI1). | |||||
|
MXI1_MOUSE
|
||||||
| NC score | 0.898064 (rank : 4) | θ value | 3.07116e-24 (rank : 7) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P50540 | Gene names | Mxi1 | |||
|
Domain Architecture |

|
|||||
| Description | MAX-interacting protein 1 (Protein MXI1). | |||||
|
MAD1_MOUSE
|
||||||
| NC score | 0.893348 (rank : 5) | θ value | 3.17079e-29 (rank : 3) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P50538, Q60798, Q61825 | Gene names | Mxd1, Mad, Mad1 | |||
|
Domain Architecture |

|
|||||
| Description | MAD protein (MAX dimerizer) (MAX dimerization protein 1). | |||||
|
MAD4_MOUSE
|
||||||
| NC score | 0.892869 (rank : 6) | θ value | 1.24688e-25 (rank : 6) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q60948 | Gene names | Mxd4, Mad4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-interacting transcriptional repressor MAD4 (Max-associated protein 4) (MAX dimerization protein 4). | |||||
|
MAD1_HUMAN
|
||||||
| NC score | 0.890323 (rank : 7) | θ value | 3.28125e-26 (rank : 5) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q05195 | Gene names | MXD1, MAD | |||
|
Domain Architecture |

|
|||||
| Description | MAD protein (MAX dimerizer) (MAX dimerization protein 1). | |||||
|
MAD4_HUMAN
|
||||||
| NC score | 0.889295 (rank : 8) | θ value | 1.12775e-26 (rank : 4) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q14582, Q5TZX4 | Gene names | MXD4, MAD4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-interacting transcriptional repressor MAD4 (Max-associated protein 4) (MAX dimerization protein 4). | |||||
|
MNT_MOUSE
|
||||||
| NC score | 0.530833 (rank : 9) | θ value | 4.30538e-10 (rank : 10) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 432 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O08789, P97349 | Gene names | Mnt, Rox | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
MNT_HUMAN
|
||||||
| NC score | 0.512521 (rank : 10) | θ value | 1.133e-10 (rank : 9) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 496 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q99583 | Gene names | MNT, ROX | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
MYC_MOUSE
|
||||||
| NC score | 0.379278 (rank : 11) | θ value | 1.29631e-06 (rank : 11) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P01108, P70247, Q3UM70, Q61422 | Gene names | Myc | |||
|
Domain Architecture |

|
|||||
| Description | Myc proto-oncogene protein (c-Myc) (Transcription factor p64). | |||||
|
MYC_HUMAN
|
||||||
| NC score | 0.376419 (rank : 12) | θ value | 2.88788e-06 (rank : 12) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P01106, P01107, Q14026 | Gene names | MYC | |||
|
Domain Architecture |

|
|||||
| Description | Myc proto-oncogene protein (c-Myc) (Transcription factor p64). | |||||
|
MAX_HUMAN
|
||||||
| NC score | 0.312590 (rank : 13) | θ value | 0.21417 (rank : 21) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P61244, P25912, P52163 | Gene names | MAX | |||
|
Domain Architecture |

|
|||||
| Description | Protein max (Myc-associated factor X). | |||||
|
MYCL2_HUMAN
|
||||||
| NC score | 0.311234 (rank : 14) | θ value | 0.000461057 (rank : 13) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P12525 | Gene names | MYCL2 | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-2 protein. | |||||
|
USF2_MOUSE
|
||||||
| NC score | 0.305938 (rank : 15) | θ value | 0.00390308 (rank : 14) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q64705, Q3UIL2, Q8VE59 | Gene names | Usf2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (Major late transcription factor 2). | |||||
|
MAX_MOUSE
|
||||||
| NC score | 0.305815 (rank : 16) | θ value | 0.21417 (rank : 22) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P28574 | Gene names | Max, Myn | |||
|
Domain Architecture |

|
|||||
| Description | Protein max (Myc-associated factor X) (Protein myn) (Myc-binding novel HLH/LZ protein). | |||||
|
USF2_HUMAN
|
||||||
| NC score | 0.305783 (rank : 17) | θ value | 0.00869519 (rank : 16) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q15853, O00671, O00709, Q05750, Q07952, Q15851, Q15852 | Gene names | USF2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (FOS- interacting protein) (FIP) (Major late transcription factor 2). | |||||
|
MYCL1_HUMAN
|
||||||
| NC score | 0.277945 (rank : 18) | θ value | 0.365318 (rank : 25) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P12524, Q14897, Q5QPL0, Q5QPL1, Q9NUE9 | Gene names | MYCL1, LMYC, MYCL | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
MYCL1_MOUSE
|
||||||
| NC score | 0.274319 (rank : 19) | θ value | 0.365318 (rank : 26) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P10166, Q5FWI7 | Gene names | Mycl1, Lmyc1, Mycl | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
USF1_MOUSE
|
||||||
| NC score | 0.273335 (rank : 20) | θ value | 0.813845 (rank : 35) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q61069 | Gene names | Usf1, Usf | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF1_HUMAN
|
||||||
| NC score | 0.272051 (rank : 21) | θ value | 0.813845 (rank : 34) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P22415 | Gene names | USF1, USF | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
TFE3_HUMAN
|
||||||
| NC score | 0.252733 (rank : 22) | θ value | 0.00509761 (rank : 15) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P19532, Q92757, Q92758, Q99964 | Gene names | TFE3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E3. | |||||
|
MYCN_MOUSE
|
||||||
| NC score | 0.250521 (rank : 23) | θ value | 3.0926 (rank : 55) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P03966, Q61978 | Gene names | Mycn, Nmyc, Nmyc1 | |||
|
Domain Architecture |

|
|||||
| Description | N-myc proto-oncogene protein. | |||||
|
MYCN_HUMAN
|
||||||
| NC score | 0.248938 (rank : 24) | θ value | 1.38821 (rank : 38) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P04198, Q6LDT9 | Gene names | MYCN, NMYC | |||
|
Domain Architecture |

|
|||||
| Description | N-myc proto-oncogene protein. | |||||
|
MITF_HUMAN
|
||||||
| NC score | 0.234482 (rank : 25) | θ value | 0.125558 (rank : 20) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O75030, Q14841, Q9P2V0, Q9P2V1, Q9P2V2, Q9P2Y8 | Gene names | MITF | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
MITF_MOUSE
|
||||||
| NC score | 0.225053 (rank : 26) | θ value | 0.279714 (rank : 23) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q08874, O08885, O88203, Q08843, Q60781, Q60782, Q9JIJ0, Q9JIJ1, Q9JIJ2, Q9JIJ3, Q9JIJ4, Q9JIJ5, Q9JIJ6, Q9JKX9 | Gene names | Mitf, Bw, Mi, Vit | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
MYCS_MOUSE
|
||||||
| NC score | 0.197739 (rank : 27) | θ value | θ > 10 (rank : 107) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9Z304 | Gene names | Mycs | |||
|
Domain Architecture |

|
|||||
| Description | Protein S-myc. | |||||
|
TFEB_MOUSE
|
||||||
| NC score | 0.194557 (rank : 28) | θ value | 1.06291 (rank : 37) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9R210 | Gene names | Tfeb, Tcfeb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
TFEB_HUMAN
|
||||||
| NC score | 0.194375 (rank : 29) | θ value | 1.06291 (rank : 36) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P19484, Q709B3, Q7Z6P9, Q9BRJ5, Q9UJD8 | Gene names | TFEB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
TCFL5_HUMAN
|
||||||
| NC score | 0.152605 (rank : 30) | θ value | θ > 10 (rank : 110) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9UL49, O94771, Q9BYW0 | Gene names | TCFL5, CHA, E2BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor-like 5 protein (Cha transcription factor) (HPV-16 E2-binding protein 1) (E2BP-1). | |||||
|
MYCB_MOUSE
|
||||||
| NC score | 0.140713 (rank : 31) | θ value | θ > 10 (rank : 106) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6P8Z1, Q9D6E3 | Gene names | Mycb, Bmyc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein B-Myc. | |||||
|
BHLH3_HUMAN
|
||||||
| NC score | 0.128418 (rank : 32) | θ value | 6.88961 (rank : 72) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9C0J9 | Gene names | BHLHB3, DEC2, SHARP1 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (hDEC2) (Enhancer-of-split and hairy-related protein 1) (SHARP-1). | |||||
|
SRBP2_HUMAN
|
||||||
| NC score | 0.126033 (rank : 33) | θ value | θ > 10 (rank : 109) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q12772, Q9UH04 | Gene names | SREBF2, SREBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 2 (SREBP-2) (Sterol regulatory element-binding transcription factor 2). | |||||
|
BHLH3_MOUSE
|
||||||
| NC score | 0.120488 (rank : 34) | θ value | 8.99809 (rank : 87) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q99PV5 | Gene names | Bhlhb3, Dec2 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (mDEC2). | |||||
|
BHLH2_MOUSE
|
||||||
| NC score | 0.119760 (rank : 35) | θ value | 2.36792 (rank : 47) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O35185, P97289 | Gene names | Bhlhb2, Clast5, Stra13 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 2 (bHLHB2) (Stimulated with retinoic acid 13) (E47 interaction protein 1) (eipl). | |||||
|
SRBP1_MOUSE
|
||||||
| NC score | 0.116597 (rank : 36) | θ value | 5.27518 (rank : 71) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9WTN3 | Gene names | Srebf1, Srebp1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
BHLH2_HUMAN
|
||||||
| NC score | 0.109046 (rank : 37) | θ value | 1.81305 (rank : 41) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O14503, Q96TD3 | Gene names | BHLHB2, DEC1, SHARP2, STRA13 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 2 (bHLHB2) (Differentially expressed in chondrocytes protein 1) (DEC1) (Enhancer-of-split and hairy-related protein 2) (SHARP-2) (Stimulated with retinoic acid 13). | |||||
|
TFAP4_HUMAN
|
||||||
| NC score | 0.098781 (rank : 38) | θ value | θ > 10 (rank : 111) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q01664, O60409 | Gene names | TFAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor AP-4 (Activating enhancer-binding protein 4). | |||||
|
SRBP1_HUMAN
|
||||||
| NC score | 0.094297 (rank : 39) | θ value | θ > 10 (rank : 108) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P36956, Q16062 | Gene names | SREBF1, SREBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
HES1_MOUSE
|
||||||
| NC score | 0.081705 (rank : 40) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P35428 | Gene names | Hes1, Hes-1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-1 (Hairy and enhancer of split 1). | |||||
|
HES1_HUMAN
|
||||||
| NC score | 0.081368 (rank : 41) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q14469 | Gene names | HES1, HL, HRY | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-1 (Hairy and enhancer of split 1) (Hairy- like) (HHL) (Hairy homolog). | |||||
|
CCD96_HUMAN
|
||||||
| NC score | 0.075360 (rank : 42) | θ value | 0.125558 (rank : 19) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 595 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q2M329, Q8N2I7 | Gene names | CCDC96 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 96. | |||||
|
BMAL1_HUMAN
|
||||||
| NC score | 0.075101 (rank : 43) | θ value | 1.81305 (rank : 42) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O00327, O00313, O00314, O00315, O00316, O00317, Q99631, Q99649 | Gene names | ARNTL, BMAL1, MOP3 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Member of PAS protein 3) (Basic-helix-loop- helix-PAS orphan MOP3) (bHLH-PAS protein JAP3). | |||||
|
CCD96_MOUSE
|
||||||
| NC score | 0.074697 (rank : 44) | θ value | 0.62314 (rank : 28) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9CR92 | Gene names | Ccdc96 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 96. | |||||
|
MLX_MOUSE
|
||||||
| NC score | 0.074460 (rank : 45) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O08609, Q9EQJ7, Q9EQJ8 | Gene names | Mlx, Tcfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
BMAL1_MOUSE
|
||||||
| NC score | 0.074061 (rank : 46) | θ value | 1.81305 (rank : 43) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9WTL8, O88295, Q921S4, Q9R0U2, Q9WTL9 | Gene names | Arntl, Bmal1 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Arnt3). | |||||
|
MLX_HUMAN
|
||||||
| NC score | 0.071306 (rank : 47) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UH92, Q53XM6, Q96FL2, Q9H2V0, Q9H2V1, Q9H2V2, Q9NXN3 | Gene names | MLX, TCFL4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
ZWINT_HUMAN
|
||||||
| NC score | 0.068265 (rank : 48) | θ value | 0.62314 (rank : 33) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O95229, Q9BWD0 | Gene names | ZWINT | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ZW10 interactor (ZW10-interacting protein 1) (Zwint-1). | |||||
|
ARNT_HUMAN
|
||||||
| NC score | 0.063429 (rank : 49) | θ value | θ > 10 (rank : 96) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P27540 | Gene names | ARNT | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
BHLH8_MOUSE
|
||||||
| NC score | 0.063400 (rank : 50) | θ value | θ > 10 (rank : 99) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9QYC3, Q9QYE4 | Gene names | Bhlhb8, Mist1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Class B basic helix-loop-helix protein 8 (bHLHB8) (Muscle, intestine and stomach expression 1) (MIST-1). | |||||
|
AZI1_HUMAN
|
||||||
| NC score | 0.062893 (rank : 51) | θ value | 0.125558 (rank : 18) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1000 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9UPN4, Q96F50 | Gene names | AZI1, CEP131, KIAA1118 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 5-azacytidine-induced protein 1 (Pre-acrosome localization protein 1) (Centrosomal protein of 131 kDa) (Cep131 protein). | |||||
|
AZI1_MOUSE
|
||||||
| NC score | 0.062187 (rank : 52) | θ value | 8.99809 (rank : 86) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 990 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q62036 | Gene names | Azi1, Az1, Azi | |||
|
Domain Architecture |

|
|||||
| Description | 5-azacytidine-induced protein 1 (Pre-acrosome localization protein 1). | |||||
|
GO45_HUMAN
|
||||||
| NC score | 0.061828 (rank : 53) | θ value | 0.62314 (rank : 29) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9H2G9, O15298, Q5T531, Q9GZX4 | Gene names | BLZF1, JEM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgin 45 (Basic leucine zipper nuclear factor 1) (JEM-1) (p45 basic leucine-zipper nuclear factor). | |||||
|
WBS14_HUMAN
|
||||||
| NC score | 0.060392 (rank : 54) | θ value | θ > 10 (rank : 113) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9NP71, Q96E48, Q9BY03, Q9BY04, Q9BY05, Q9BY06, Q9Y2P3 | Gene names | MLXIPL, MIO, WBSCR14 | |||
|
Domain Architecture |

|
|||||
| Description | Williams-Beuren syndrome chromosome region 14 protein (WS basic-helix- loop-helix leucine zipper protein) (WS-bHLH) (Mlx interactor) (MLX- interacting protein-like). | |||||
|
BHLH8_HUMAN
|
||||||
| NC score | 0.058843 (rank : 55) | θ value | θ > 10 (rank : 98) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q7RTS1 | Gene names | BHLHB8, MIST1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Class B basic helix-loop-helix protein 8 (bHLHB8) (Muscle, intestine and stomach expression 1) (MIST-1). | |||||
|
HES4_HUMAN
|
||||||
| NC score | 0.058355 (rank : 56) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9HCC6 | Gene names | HES4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-4 (Hairy and enhancer of split 4) (bHLH factor Hes4). | |||||
|
RABE2_MOUSE
|
||||||
| NC score | 0.058226 (rank : 57) | θ value | 1.81305 (rank : 46) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 598 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q91WG2, Q99KN3 | Gene names | Rabep2, Rabpt5b | |||
|
Domain Architecture |

|
|||||
| Description | Rab GTPase-binding effector protein 2 (Rabaptin-5beta). | |||||
|
CCDC6_HUMAN
|
||||||
| NC score | 0.057834 (rank : 58) | θ value | 3.0926 (rank : 53) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 723 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q16204, Q15250 | Gene names | CCDC6, D10S170 | |||
|
Domain Architecture |

|
|||||
| Description | Coiled-coil domain-containing protein 6 (H4 protein) (Papillary thyroid carcinoma-encoded protein). | |||||
|
GO45_MOUSE
|
||||||
| NC score | 0.055737 (rank : 59) | θ value | 4.03905 (rank : 63) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8R2X8, Q3TJC4, Q8C0U6, Q9DAV7 | Gene names | Blzf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgin 45 (Basic leucine zipper nuclear factor 1). | |||||
|
ARNT_MOUSE
|
||||||
| NC score | 0.055509 (rank : 60) | θ value | θ > 10 (rank : 97) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P53762, Q60661 | Gene names | Arnt | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
FIGLA_HUMAN
|
||||||
| NC score | 0.053664 (rank : 61) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6QHK4 | Gene names | FIGLA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Factor in the germline alpha (Transcription factor FIGa) (FIGalpha). | |||||
|
BPAEA_MOUSE
|
||||||
| NC score | 0.053198 (rank : 62) | θ value | 0.365318 (rank : 24) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1480 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q91ZU8, Q8K5D4 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoform 5 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
TRHY_HUMAN
|
||||||
| NC score | 0.051682 (rank : 63) | θ value | θ > 10 (rank : 112) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1385 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q07283 | Gene names | THH, THL, TRHY | |||
|
Domain Architecture |

|
|||||
| Description | Trichohyalin. | |||||
|
ARNT2_MOUSE
|
||||||
| NC score | 0.050258 (rank : 64) | θ value | θ > 10 (rank : 95) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q61324 | Gene names | Arnt2 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
BICD2_HUMAN
|
||||||
| NC score | 0.049786 (rank : 65) | θ value | 2.36792 (rank : 48) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 856 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8TD16, O75181, Q5TBQ2, Q5TBQ3, Q96LH2, Q9BT84, Q9H561 | Gene names | BICD2, KIAA0699 | |||
|
Domain Architecture |

|
|||||
| Description | Cytoskeleton-like bicaudal D protein homolog 2. | |||||
|
GOGA3_HUMAN
|
||||||
| NC score | 0.049750 (rank : 66) | θ value | 0.62314 (rank : 30) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1317 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q08378, O43241, Q6P9C7, Q86XW3, Q8TDA9, Q8WZA3 | Gene names | GOLGA3 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 3 (Golgin-160) (Golgi complex-associated protein of 170 kDa) (GCP170). | |||||
|
GOGA1_HUMAN
|
||||||
| NC score | 0.049651 (rank : 67) | θ value | 0.0563607 (rank : 17) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1222 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q92805, Q8IYZ9 | Gene names | GOLGA1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 1 (Golgin-97). | |||||
|
BICD2_MOUSE
|
||||||
| NC score | 0.049607 (rank : 68) | θ value | 2.36792 (rank : 49) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 864 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q921C5, Q80TU1, Q8BTE3, Q9DCL3 | Gene names | Bicd2, Kiaa0699 | |||
|
Domain Architecture |

|
|||||
| Description | Cytoskeleton-like bicaudal D protein homolog 2. | |||||
|
CING_MOUSE
|
||||||
| NC score | 0.049070 (rank : 69) | θ value | 4.03905 (rank : 61) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1432 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | P59242 | Gene names | Cgn | |||
|
Domain Architecture |

|
|||||
| Description | Cingulin. | |||||
|
CCD62_HUMAN
|
||||||
| NC score | 0.048059 (rank : 70) | θ value | 2.36792 (rank : 50) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 631 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q6P9F0, Q6ZVF2, Q86VJ0, Q9BYZ5 | Gene names | CCDC62 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 60 (Aaa-protein) (Protein TSP- NY). | |||||
|
DESP_HUMAN
|
||||||
| NC score | 0.047607 (rank : 71) | θ value | 4.03905 (rank : 62) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P15924, O75993, Q14189, Q9UHN4 | Gene names | DSP | |||
|
Domain Architecture |

|
|||||
| Description | Desmoplakin (DP) (250/210 kDa paraneoplastic pemphigus antigen). | |||||
|
SYCP1_HUMAN
|
||||||
| NC score | 0.046872 (rank : 72) | θ value | 0.62314 (rank : 32) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1313 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q15431, O14963 | Gene names | SYCP1, SCP1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptonemal complex protein 1 (SCP-1). | |||||
|
CCD21_HUMAN
|
||||||
| NC score | 0.046689 (rank : 73) | θ value | 0.62314 (rank : 27) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1228 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q6P2H3, Q5VY68, Q5VY70, Q9H6Q1, Q9H828, Q9UF52 | Gene names | CCDC21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 21. | |||||
|
HIP1_HUMAN
|
||||||
| NC score | 0.046233 (rank : 74) | θ value | 2.36792 (rank : 51) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1016 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | O00291, O00328, Q8TDL4 | Gene names | HIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin-interacting protein 1 (HIP-I). | |||||
|
CENPF_HUMAN
|
||||||
| NC score | 0.046015 (rank : 75) | θ value | 1.81305 (rank : 44) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1932 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P49454, Q13171, Q13246 | Gene names | CENPF | |||
|
Domain Architecture |

|
|||||
| Description | Centromere protein F (Kinetochore protein CENP-F) (Mitosin) (AH antigen). | |||||
|
BICD1_HUMAN
|
||||||
| NC score | 0.045487 (rank : 76) | θ value | 6.88961 (rank : 73) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 844 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q96G01, O43892, O43893 | Gene names | BICD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-like bicaudal D protein homolog 1. | |||||
|
PCNT_MOUSE
|
||||||
| NC score | 0.045229 (rank : 77) | θ value | 1.81305 (rank : 45) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1207 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P48725 | Gene names | Pcnt, Pcnt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pericentrin. | |||||
|
DZIP1_MOUSE
|
||||||
| NC score | 0.042552 (rank : 78) | θ value | 3.0926 (rank : 54) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 530 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8BMD2, Q6ZQ10, Q9CRK5 | Gene names | Dzip1, Kiaa0996 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein DZIP1 (DAZ-interacting protein 1 homolog). | |||||
|
TAXB1_MOUSE
|
||||||
| NC score | 0.040451 (rank : 79) | θ value | 6.88961 (rank : 84) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 886 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q3UKC1, Q91YT6, Q9CVF0, Q9DC45 | Gene names | Tax1bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tax1-binding protein 1 homolog. | |||||
|
RCOR3_MOUSE
|
||||||
| NC score | 0.039673 (rank : 80) | θ value | 1.38821 (rank : 40) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6PGA0 | Gene names | Rcor3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 3. | |||||
|
CASP_HUMAN
|
||||||
| NC score | 0.038697 (rank : 81) | θ value | 8.99809 (rank : 88) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1131 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q13948, Q53GU9, Q8TBS3 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein CASP. | |||||
|
LIPB2_MOUSE
|
||||||
| NC score | 0.038693 (rank : 82) | θ value | 4.03905 (rank : 64) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 570 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O35711, Q7TMG4, Q8CBS6, Q99KX6 | Gene names | Ppfibp2, Cclp1 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-beta-2 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein-binding protein 2) (PTPRF-interacting protein-binding protein 2) (Coiled-coil-like protein 1). | |||||
|
OPTN_HUMAN
|
||||||
| NC score | 0.038684 (rank : 83) | θ value | 5.27518 (rank : 70) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 827 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q96CV9, Q5T672, Q5T673, Q5T674, Q5T675, Q7LDL9, Q8N562, Q9UET9, Q9UEV4, Q9Y218 | Gene names | OPTN, FIP2, GLC1E, NRP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Optineurin (Optic neuropathy-inducing protein) (E3-14.7K-interacting protein) (FIP-2) (Huntingtin-interacting protein HYPL) (NEMO-related protein) (Transcription factor IIIA-interacting protein) (TFIIIA- IntP). | |||||
|
GOGA4_MOUSE
|
||||||
| NC score | 0.038464 (rank : 84) | θ value | 5.27518 (rank : 69) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1939 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q91VW5, O70365, Q8CGH6 | Gene names | Golga4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (tGolgin-1). | |||||
|
ATOH1_MOUSE
|
||||||
| NC score | 0.038342 (rank : 85) | θ value | 4.03905 (rank : 59) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P48985 | Gene names | Atoh1, Ath1 | |||
|
Domain Architecture |

|
|||||
| Description | Atonal protein homolog 1 (Helix-loop-helix protein mATH-1) (MATH1). | |||||
|
ATOH1_HUMAN
|
||||||
| NC score | 0.038301 (rank : 86) | θ value | 4.03905 (rank : 58) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q92858 | Gene names | ATOH1, ATH1 | |||
|
Domain Architecture |

|
|||||
| Description | Atonal protein homolog 1 (Helix-loop-helix protein hATH-1). | |||||
|
EP15R_HUMAN
|
||||||
| NC score | 0.037661 (rank : 87) | θ value | 5.27518 (rank : 68) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1083 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9UBC2 | Gene names | EPS15L1, EPS15R | |||
|
Domain Architecture |

|
|||||
| Description | Epidermal growth factor receptor substrate 15-like 1 (Eps15-related protein) (Eps15R). | |||||
|
RCOR3_HUMAN
|
||||||
| NC score | 0.037369 (rank : 88) | θ value | 1.38821 (rank : 39) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9P2K3, Q5VT47, Q7L9I5, Q8N5U3, Q9NV83 | Gene names | RCOR3, KIAA1343 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 3. | |||||
|
CCD46_MOUSE
|
||||||
| NC score | 0.037068 (rank : 89) | θ value | 8.99809 (rank : 89) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q5PR68, Q80ZN6, Q99JS4, Q9D9T1 | Gene names | Ccdc46 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled coil domain-containing protein 46. | |||||
|
MY18A_MOUSE
|
||||||
| NC score | 0.036029 (rank : 90) | θ value | 6.88961 (rank : 78) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1989 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9JMH9, Q811D7 | Gene names | Myo18a, Myspdz | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
EP15R_MOUSE
|
||||||
| NC score | 0.036027 (rank : 91) | θ value | 6.88961 (rank : 76) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q60902, Q8CB60, Q8CB70, Q91WH8 | Gene names | Eps15l1, Eps15-rs, Eps15R | |||
|
Domain Architecture |

|
|||||
| Description | Epidermal growth factor receptor substrate 15-like 1 (Eps15-related protein) (Eps15R) (Epidermal growth factor receptor pathway substrate 15-related sequence) (Eps15-rs). | |||||
|
STX1C_HUMAN
|
||||||
| NC score | 0.035633 (rank : 92) | θ value | 6.88961 (rank : 82) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 270 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P61266 | Gene names | STX1B2 | |||
|
Domain Architecture |

|
|||||
| Description | Syntaxin-1B2. | |||||
|
STX1C_MOUSE
|
||||||
| NC score | 0.035633 (rank : 93) | θ value | 6.88961 (rank : 83) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 270 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P61264, P32853 | Gene names | Stx1b2, Stx1b | |||
|
Domain Architecture |

|
|||||
| Description | Syntaxin-1B2 (Syntaxin 1B). | |||||
|
STX1A_MOUSE
|
||||||
| NC score | 0.035579 (rank : 94) | θ value | 3.0926 (rank : 57) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O35526 | Gene names | Stx1a | |||
|
Domain Architecture |

|
|||||
| Description | Syntaxin-1A (Neuron-specific antigen HPC-1). | |||||
|
ASCL2_HUMAN
|
||||||
| NC score | 0.035505 (rank : 95) | θ value | 8.99809 (rank : 85) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q99929, Q9UM68 | Gene names | ASCL2 | |||
|
Domain Architecture |

|
|||||
| Description | Achaete-scute homolog 2 (Mash2) (HASH2). | |||||
|
STX1A_HUMAN
|
||||||
| NC score | 0.035376 (rank : 96) | θ value | 3.0926 (rank : 56) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q16623, O15447, O15448, Q12936, Q9BPZ6 | Gene names | STX1A, STX1 | |||
|
Domain Architecture |

|
|||||
| Description | Syntaxin-1A (Neuron-specific antigen HPC-1). | |||||
|
CENPE_HUMAN
|
||||||
| NC score | 0.034474 (rank : 97) | θ value | 8.99809 (rank : 90) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 2060 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q02224 | Gene names | CENPE | |||
|
Domain Architecture |

|
|||||
| Description | Centromeric protein E (CENP-E). | |||||
|
MY18A_HUMAN
|
||||||
| NC score | 0.034392 (rank : 98) | θ value | 6.88961 (rank : 77) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1929 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q92614, Q8IXP8 | Gene names | MYO18A, KIAA0216, MYSPDZ | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
STX1B_HUMAN
|
||||||
| NC score | 0.034049 (rank : 99) | θ value | 6.88961 (rank : 81) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 252 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q15531 | Gene names | STX1B1, STX1B | |||
|
Domain Architecture |

|
|||||
| Description | Syntaxin-1B1 (Syntaxin 1B). | |||||
|
MYH9_MOUSE
|
||||||
| NC score | 0.033984 (rank : 100) | θ value | 8.99809 (rank : 94) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1829 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q8VDD5 | Gene names | Myh9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
BPA1_MOUSE
|
||||||
| NC score | 0.033976 (rank : 101) | θ value | 6.88961 (rank : 74) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q91ZU6, Q91ZU7 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 1/2/3/4 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
CUTL1_HUMAN
|
||||||
| NC score | 0.033762 (rank : 102) | θ value | 8.99809 (rank : 91) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1574 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P39880, Q6NYH4, Q9UEV5 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP). | |||||
|
ANR26_MOUSE
|
||||||
| NC score | 0.031831 (rank : 103) | θ value | 5.27518 (rank : 65) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1554 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q811D2, Q69ZS2, Q9CS61 | Gene names | Ankrd26, Kiaa1074 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 26. | |||||
|
MYH13_HUMAN
|
||||||
| NC score | 0.030582 (rank : 104) | θ value | 8.99809 (rank : 93) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1504 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9UKX3, O95252 | Gene names | MYH13 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-13 (Myosin heavy chain, skeletal muscle, extraocular) (MyHC- eo). | |||||
|
NGN1_MOUSE
|
||||||
| NC score | 0.023147 (rank : 105) | θ value | 2.36792 (rank : 52) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P70660 | Gene names | Neurog1, Ath4c, Neurod3, Ngn, Ngn1 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenin 1 (Neurogenic differentiation factor 3) (NeuroD3) (Neurogenic basic-helix-loop-helix protein) (Helix-loop-helix protein mATH-4C) (MATH4C). | |||||
|
CCD16_HUMAN
|
||||||
| NC score | 0.021043 (rank : 106) | θ value | 4.03905 (rank : 60) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96NB3, Q96F60, Q96GZ5, Q9BU38 | Gene names | CCDC16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 16. | |||||
|
DTBP1_HUMAN
|
||||||
| NC score | 0.019571 (rank : 107) | θ value | 6.88961 (rank : 75) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q96EV8, Q96NV2, Q9H0U2, Q9H3J5 | Gene names | DTNBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Dystrobrevin-binding protein 1 (Dysbindin) (Hermansky-Pudlak syndrome 7 protein). | |||||
|
CTRO_MOUSE
|
||||||
| NC score | 0.019476 (rank : 108) | θ value | 5.27518 (rank : 67) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 2344 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P49025, O88528, O88937, O88938, Q3UM99, Q8CIJ1 | Gene names | Cit | |||
|
Domain Architecture |

|
|||||
| Description | Citron Rho-interacting kinase (EC 2.7.11.1) (CRIK) (Rho-interacting, serine/threonine-protein kinase 21). | |||||
|
ROCK2_MOUSE
|
||||||
| NC score | 0.018035 (rank : 109) | θ value | 6.88961 (rank : 80) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 2072 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P70336, Q8BR64, Q8CBR0 | Gene names | Rock2 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 2 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 2) (p164 ROCK-2). | |||||
|
ROCK2_HUMAN
|
||||||
| NC score | 0.017644 (rank : 110) | θ value | 6.88961 (rank : 79) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 2073 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | O75116, Q9UQN5 | Gene names | ROCK2, KIAA0619 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 2 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 2) (p164 ROCK-2) (Rho kinase 2). | |||||
|
BRPF1_HUMAN
|
||||||
| NC score | 0.013385 (rank : 111) | θ value | 5.27518 (rank : 66) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P55201, Q9UHI0 | Gene names | BRPF1, BR140 | |||
|
Domain Architecture |

|
|||||
| Description | Peregrin (Bromodomain and PHD finger-containing protein 1) (BR140 protein). | |||||
|
NEK11_HUMAN
|
||||||
| NC score | 0.006915 (rank : 112) | θ value | 0.62314 (rank : 31) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1043 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8NG66, Q5JPC0, Q8NG65, Q8TBY1, Q9H5F4 | Gene names | NEK11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase Nek11 (EC 2.7.11.1) (NimA-related protein kinase 11) (Never in mitosis A-related kinase 11). | |||||
|
K1802_MOUSE
|
||||||
| NC score | 0.006361 (rank : 113) | θ value | 8.99809 (rank : 92) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||