Please be patient as the page loads
|
MAD3_HUMAN
|
||||||
| SwissProt Accessions | Q9BW11, Q53HK1, Q7Z4Y0, Q8NDJ7, Q96ME3 | Gene names | MXD3, MAD3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Max-interacting transcriptional repressor MAD3 (Max-associated protein 3) (MAX dimerization protein 3) (Myx). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
MAD3_HUMAN
|
||||||
| θ value | 8.81223e-72 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 92 | |
| SwissProt Accessions | Q9BW11, Q53HK1, Q7Z4Y0, Q8NDJ7, Q96ME3 | Gene names | MXD3, MAD3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Max-interacting transcriptional repressor MAD3 (Max-associated protein 3) (MAX dimerization protein 3) (Myx). | |||||
|
MAD3_MOUSE
|
||||||
| θ value | 4.24587e-58 (rank : 2) | NC score | 0.972694 (rank : 2) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q80US8, Q60947 | Gene names | Mxd3, Mad3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Max-interacting transcriptional repressor MAD3 (Max-associated protein 3) (MAX dimerization protein 3). | |||||
|
MAD1_MOUSE
|
||||||
| θ value | 3.3877e-31 (rank : 3) | NC score | 0.892487 (rank : 6) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P50538, Q60798, Q61825 | Gene names | Mxd1, Mad, Mad1 | |||
|
Domain Architecture |

|
|||||
| Description | MAD protein (MAX dimerizer) (MAX dimerization protein 1). | |||||
|
MAD1_HUMAN
|
||||||
| θ value | 1.02001e-27 (rank : 4) | NC score | 0.890053 (rank : 8) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q05195 | Gene names | MXD1, MAD | |||
|
Domain Architecture |
|
|||||
| Description | MAD protein (MAX dimerizer) (MAX dimerization protein 1). | |||||
|
MAD4_HUMAN
|
||||||
| θ value | 5.06226e-27 (rank : 5) | NC score | 0.895203 (rank : 5) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q14582, Q5TZX4 | Gene names | MXD4, MAD4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-interacting transcriptional repressor MAD4 (Max-associated protein 4) (MAX dimerization protein 4). | |||||
|
MAD4_MOUSE
|
||||||
| θ value | 5.06226e-27 (rank : 6) | NC score | 0.891616 (rank : 7) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q60948 | Gene names | Mxd4, Mad4 | |||
|
Domain Architecture |
|
|||||
| Description | Max-interacting transcriptional repressor MAD4 (Max-associated protein 4) (MAX dimerization protein 4). | |||||
|
MXI1_MOUSE
|
||||||
| θ value | 2.35151e-24 (rank : 7) | NC score | 0.897169 (rank : 4) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P50540 | Gene names | Mxi1 | |||
|
Domain Architecture |
|
|||||
| Description | MAX-interacting protein 1 (Protein MXI1). | |||||
|
MXI1_HUMAN
|
||||||
| θ value | 1.99067e-23 (rank : 8) | NC score | 0.902806 (rank : 3) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P50539, Q15887 | Gene names | MXI1 | |||
|
Domain Architecture |
|
|||||
| Description | MAX-interacting protein 1 (Protein MXI1). | |||||
|
MNT_MOUSE
|
||||||
| θ value | 9.59137e-10 (rank : 9) | NC score | 0.519868 (rank : 9) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 432 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O08789, P97349 | Gene names | Mnt, Rox | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
MNT_HUMAN
|
||||||
| θ value | 1.63604e-09 (rank : 10) | NC score | 0.499335 (rank : 10) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 496 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q99583 | Gene names | MNT, ROX | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
MYC_MOUSE
|
||||||
| θ value | 5.81887e-07 (rank : 11) | NC score | 0.386049 (rank : 11) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P01108, P70247, Q3UM70, Q61422 | Gene names | Myc | |||
|
Domain Architecture |
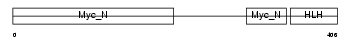
|
|||||
| Description | Myc proto-oncogene protein (c-Myc) (Transcription factor p64). | |||||
|
MYC_HUMAN
|
||||||
| θ value | 1.29631e-06 (rank : 12) | NC score | 0.384205 (rank : 12) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P01106, P01107, Q14026 | Gene names | MYC | |||
|
Domain Architecture |

|
|||||
| Description | Myc proto-oncogene protein (c-Myc) (Transcription factor p64). | |||||
|
MYCL2_HUMAN
|
||||||
| θ value | 0.00020696 (rank : 13) | NC score | 0.322557 (rank : 13) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P12525 | Gene names | MYCL2 | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-2 protein. | |||||
|
TFE3_HUMAN
|
||||||
| θ value | 0.000786445 (rank : 14) | NC score | 0.273559 (rank : 22) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P19532, Q92757, Q92758, Q99964 | Gene names | TFE3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E3. | |||||
|
USF2_MOUSE
|
||||||
| θ value | 0.00509761 (rank : 15) | NC score | 0.314071 (rank : 14) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q64705, Q3UIL2, Q8VE59 | Gene names | Usf2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (Major late transcription factor 2). | |||||
|
SRBP1_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 16) | NC score | 0.171002 (rank : 31) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9WTN3 | Gene names | Srebf1, Srebp1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
USF2_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 17) | NC score | 0.313247 (rank : 15) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q15853, O00671, O00709, Q05750, Q07952, Q15851, Q15852 | Gene names | USF2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (FOS- interacting protein) (FIP) (Major late transcription factor 2). | |||||
|
MITF_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 18) | NC score | 0.255856 (rank : 25) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O75030, Q14841, Q9P2V0, Q9P2V1, Q9P2V2, Q9P2Y8 | Gene names | MITF | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
MITF_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 19) | NC score | 0.246156 (rank : 26) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q08874, O08885, O88203, Q08843, Q60781, Q60782, Q9JIJ0, Q9JIJ1, Q9JIJ2, Q9JIJ3, Q9JIJ4, Q9JIJ5, Q9JIJ6, Q9JKX9 | Gene names | Mitf, Bw, Mi, Vit | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
MYCL1_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 20) | NC score | 0.295018 (rank : 18) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P12524, Q14897, Q5QPL0, Q5QPL1, Q9NUE9 | Gene names | MYCL1, LMYC, MYCL | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
MD1L1_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 21) | NC score | 0.055999 (rank : 59) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 893 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9Y6D9, Q13312, Q75MI0, Q86UM4, Q9UNH0 | Gene names | MAD1L1, MAD1, TXBP181 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitotic spindle assembly checkpoint protein MAD1 (Mitotic arrest deficient-like protein 1) (MAD1-like 1) (Mitotic checkpoint MAD1 protein-homolog) (HsMAD1) (hMAD1) (Tax-binding protein 181). | |||||
|
RABE2_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 22) | NC score | 0.063879 (rank : 53) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 598 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q91WG2, Q99KN3 | Gene names | Rabep2, Rabpt5b | |||
|
Domain Architecture |

|
|||||
| Description | Rab GTPase-binding effector protein 2 (Rabaptin-5beta). | |||||
|
BPAEA_MOUSE
|
||||||
| θ value | 0.125558 (rank : 23) | NC score | 0.049777 (rank : 65) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1480 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q91ZU8, Q8K5D4 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoform 5 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
NEK11_HUMAN
|
||||||
| θ value | 0.125558 (rank : 24) | NC score | 0.007302 (rank : 104) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1043 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8NG66, Q5JPC0, Q8NG65, Q8TBY1, Q9H5F4 | Gene names | NEK11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase Nek11 (EC 2.7.11.1) (NimA-related protein kinase 11) (Never in mitosis A-related kinase 11). | |||||
|
ANKR6_HUMAN
|
||||||
| θ value | 0.163984 (rank : 25) | NC score | 0.015527 (rank : 101) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 439 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Y2G4, Q9NU24, Q9UFQ9 | Gene names | ANKRD6, KIAA0957 | |||
|
Domain Architecture |
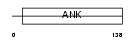
|
|||||
| Description | Ankyrin repeat domain-containing protein 6. | |||||
|
KIF5C_MOUSE
|
||||||
| θ value | 0.163984 (rank : 26) | NC score | 0.038530 (rank : 82) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1040 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P28738, Q9Z2F8 | Gene names | Kif5c, Nkhc2 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin heavy chain isoform 5C (Kinesin heavy chain neuron-specific 2). | |||||
|
MYCL1_MOUSE
|
||||||
| θ value | 0.163984 (rank : 27) | NC score | 0.292473 (rank : 19) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P10166, Q5FWI7 | Gene names | Mycl1, Lmyc1, Mycl | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
CP135_MOUSE
|
||||||
| θ value | 0.279714 (rank : 28) | NC score | 0.041583 (rank : 75) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1278 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q6P5D4, Q6A030 | Gene names | Cep135, Cep4, Kiaa0635 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 135 kDa (Cep135 protein) (Centrosomal protein 4). | |||||
|
TACC2_HUMAN
|
||||||
| θ value | 0.365318 (rank : 29) | NC score | 0.057029 (rank : 58) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 856 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O95359, Q9NZ41, Q9NZR5 | Gene names | TACC2 | |||
|
Domain Architecture |

|
|||||
| Description | Transforming acidic coiled-coil-containing protein 2 (Anti Zuai-1) (AZU-1). | |||||
|
CT2NL_MOUSE
|
||||||
| θ value | 0.47712 (rank : 30) | NC score | 0.043133 (rank : 71) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 902 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q99LJ0, Q6ZPR0, Q8BSV1 | Gene names | Cttnbp2nl, Kiaa1433 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CTTNBP2 N-terminal-like protein. | |||||
|
GOGA5_MOUSE
|
||||||
| θ value | 0.47712 (rank : 31) | NC score | 0.038832 (rank : 81) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1434 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9QYE6, O88317, Q3TGE7, Q3U6S5, Q3UUF9 | Gene names | Golga5, Retii | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 5 (Golgin-84) (Sumiko protein) (Ret-II protein). | |||||
|
SRBP1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 32) | NC score | 0.150475 (rank : 34) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P36956, Q16062 | Gene names | SREBF1, SREBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
CCD96_MOUSE
|
||||||
| θ value | 0.62314 (rank : 33) | NC score | 0.068215 (rank : 52) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9CR92 | Gene names | Ccdc96 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 96. | |||||
|
CCD96_HUMAN
|
||||||
| θ value | 0.813845 (rank : 34) | NC score | 0.077033 (rank : 48) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 595 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q2M329, Q8N2I7 | Gene names | CCDC96 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 96. | |||||
|
TACC2_MOUSE
|
||||||
| θ value | 0.813845 (rank : 35) | NC score | 0.055503 (rank : 60) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 774 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9JJG0 | Gene names | Tacc2 | |||
|
Domain Architecture |

|
|||||
| Description | Transforming acidic coiled-coil-containing protein 2. | |||||
|
ZWINT_HUMAN
|
||||||
| θ value | 0.813845 (rank : 36) | NC score | 0.063631 (rank : 54) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O95229, Q9BWD0 | Gene names | ZWINT | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ZW10 interactor (ZW10-interacting protein 1) (Zwint-1). | |||||
|
AZI1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 37) | NC score | 0.052878 (rank : 62) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 990 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q62036 | Gene names | Azi1, Az1, Azi | |||
|
Domain Architecture |

|
|||||
| Description | 5-azacytidine-induced protein 1 (Pre-acrosome localization protein 1). | |||||
|
CROCC_HUMAN
|
||||||
| θ value | 1.06291 (rank : 38) | NC score | 0.048497 (rank : 66) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1409 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q5TZA2, Q2VHY3, Q66GT7, Q7Z2L4, Q7Z5D7 | Gene names | CROCC, KIAA0445 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rootletin (Ciliary rootlet coiled-coil protein). | |||||
|
CYLN2_MOUSE
|
||||||
| θ value | 1.06291 (rank : 39) | NC score | 0.043426 (rank : 70) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1002 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9Z0H8, Q7TSI9, Q8CHU1, Q9EP81 | Gene names | Cyln2, Kiaa0291 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoplasmic linker protein 2 (Cytoplasmic linker protein 115) (CLIP- 115). | |||||
|
MAX_HUMAN
|
||||||
| θ value | 1.06291 (rank : 40) | NC score | 0.310358 (rank : 16) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P61244, P25912, P52163 | Gene names | MAX | |||
|
Domain Architecture |

|
|||||
| Description | Protein max (Myc-associated factor X). | |||||
|
RCOR3_HUMAN
|
||||||
| θ value | 1.06291 (rank : 41) | NC score | 0.039831 (rank : 80) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9P2K3, Q5VT47, Q7L9I5, Q8N5U3, Q9NV83 | Gene names | RCOR3, KIAA1343 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 3. | |||||
|
RCOR3_MOUSE
|
||||||
| θ value | 1.06291 (rank : 42) | NC score | 0.042309 (rank : 72) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q6PGA0 | Gene names | Rcor3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 3. | |||||
|
TFEB_HUMAN
|
||||||
| θ value | 1.06291 (rank : 43) | NC score | 0.211619 (rank : 27) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P19484, Q709B3, Q7Z6P9, Q9BRJ5, Q9UJD8 | Gene names | TFEB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
USF1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 44) | NC score | 0.283053 (rank : 21) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P22415 | Gene names | USF1, USF | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 45) | NC score | 0.284224 (rank : 20) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q61069 | Gene names | Usf1, Usf | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
BHLH2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 46) | NC score | 0.139083 (rank : 37) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O35185, P97289 | Gene names | Bhlhb2, Clast5, Stra13 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 2 (bHLHB2) (Stimulated with retinoic acid 13) (E47 interaction protein 1) (eipl). | |||||
|
BHLH3_HUMAN
|
||||||
| θ value | 1.38821 (rank : 47) | NC score | 0.151486 (rank : 33) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9C0J9 | Gene names | BHLHB3, DEC2, SHARP1 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (hDEC2) (Enhancer-of-split and hairy-related protein 1) (SHARP-1). | |||||
|
ARHGC_MOUSE
|
||||||
| θ value | 1.81305 (rank : 48) | NC score | 0.018228 (rank : 98) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8R4H2, Q80U18 | Gene names | Arhgef12, Kiaa0382, Larg | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 12 (Leukemia-associated RhoGEF). | |||||
|
BHLH3_MOUSE
|
||||||
| θ value | 1.81305 (rank : 49) | NC score | 0.143988 (rank : 36) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q99PV5 | Gene names | Bhlhb3, Dec2 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (mDEC2). | |||||
|
MAX_MOUSE
|
||||||
| θ value | 1.81305 (rank : 50) | NC score | 0.303550 (rank : 17) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P28574 | Gene names | Max, Myn | |||
|
Domain Architecture |

|
|||||
| Description | Protein max (Myc-associated factor X) (Protein myn) (Myc-binding novel HLH/LZ protein). | |||||
|
BHLH2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 51) | NC score | 0.127778 (rank : 38) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O14503, Q96TD3 | Gene names | BHLHB2, DEC1, SHARP2, STRA13 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 2 (bHLHB2) (Differentially expressed in chondrocytes protein 1) (DEC1) (Enhancer-of-split and hairy-related protein 2) (SHARP-2) (Stimulated with retinoic acid 13). | |||||
|
BHLH8_HUMAN
|
||||||
| θ value | 2.36792 (rank : 52) | NC score | 0.057991 (rank : 57) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q7RTS1 | Gene names | BHLHB8, MIST1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Class B basic helix-loop-helix protein 8 (bHLHB8) (Muscle, intestine and stomach expression 1) (MIST-1). | |||||
|
KIF5C_HUMAN
|
||||||
| θ value | 2.36792 (rank : 53) | NC score | 0.036760 (rank : 83) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 974 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O60282, O95079 | Gene names | KIF5C, KIAA0531, NKHC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin heavy chain isoform 5C (Kinesin heavy chain neuron-specific 2). | |||||
|
ZGPAT_MOUSE
|
||||||
| θ value | 2.36792 (rank : 54) | NC score | 0.028752 (rank : 93) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8VDM1, Q8BWW2 | Gene names | Zgpat | |||
|
Domain Architecture |
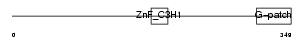
|
|||||
| Description | Zinc finger CCCH-type with G patch domain protein. | |||||
|
BMAL1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 55) | NC score | 0.089841 (rank : 42) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O00327, O00313, O00314, O00315, O00316, O00317, Q99631, Q99649 | Gene names | ARNTL, BMAL1, MOP3 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Member of PAS protein 3) (Basic-helix-loop- helix-PAS orphan MOP3) (bHLH-PAS protein JAP3). | |||||
|
BMAL1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 56) | NC score | 0.088454 (rank : 43) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9WTL8, O88295, Q921S4, Q9R0U2, Q9WTL9 | Gene names | Arntl, Bmal1 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Arnt3). | |||||
|
MYCN_HUMAN
|
||||||
| θ value | 3.0926 (rank : 57) | NC score | 0.256197 (rank : 24) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P04198, Q6LDT9 | Gene names | MYCN, NMYC | |||
|
Domain Architecture |

|
|||||
| Description | N-myc proto-oncogene protein. | |||||
|
MYCN_MOUSE
|
||||||
| θ value | 3.0926 (rank : 58) | NC score | 0.257458 (rank : 23) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P03966, Q61978 | Gene names | Mycn, Nmyc, Nmyc1 | |||
|
Domain Architecture |

|
|||||
| Description | N-myc proto-oncogene protein. | |||||
|
PLEC1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 59) | NC score | 0.040932 (rank : 77) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1326 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q15149, Q15148, Q16640 | Gene names | PLEC1 | |||
|
Domain Architecture |

|
|||||
| Description | Plectin-1 (PLTN) (PCN) (Hemidesmosomal protein 1) (HD1). | |||||
|
TFEB_MOUSE
|
||||||
| θ value | 3.0926 (rank : 60) | NC score | 0.211359 (rank : 28) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9R210 | Gene names | Tfeb, Tcfeb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
ARNT2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 61) | NC score | 0.072784 (rank : 50) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9HBZ2, O15024 | Gene names | ARNT2, KIAA0307 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
ARNT2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 62) | NC score | 0.073191 (rank : 49) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q61324 | Gene names | Arnt2 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
CCD21_HUMAN
|
||||||
| θ value | 4.03905 (rank : 63) | NC score | 0.041241 (rank : 76) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1228 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q6P2H3, Q5VY68, Q5VY70, Q9H6Q1, Q9H828, Q9UF52 | Gene names | CCDC21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 21. | |||||
|
CYLN2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 64) | NC score | 0.041908 (rank : 74) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1004 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9UDT6, O14527, O43611 | Gene names | CYLN2, KIAA0291, WBSCR4, WSCR4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoplasmic linker protein 2 (Cytoplasmic linker protein 115) (CLIP- 115) (Williams-Beuren syndrome chromosome region 4 protein). | |||||
|
HES2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 65) | NC score | 0.044445 (rank : 69) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O54792 | Gene names | Hes2 | |||
|
Domain Architecture |
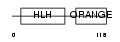
|
|||||
| Description | Transcription factor HES-2 (Hairy and enhancer of split 2). | |||||
|
HIP1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 66) | NC score | 0.042177 (rank : 73) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1016 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O00291, O00328, Q8TDL4 | Gene names | HIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin-interacting protein 1 (HIP-I). | |||||
|
MACF4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 67) | NC score | 0.031820 (rank : 88) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1321 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q96PK2, Q8WXY1 | Gene names | MACF1, ABP620, ACF7, KIAA0465, KIAA1251 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1, isoform 4. | |||||
|
SYNE1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 68) | NC score | 0.031367 (rank : 91) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1484 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8NF91, O94890, Q5JV19, Q5JV22, Q8N9P7, Q8TCP1, Q8WWW6, Q8WWW7, Q8WXF6, Q96N17, Q9C0A7, Q9H525, Q9H526, Q9NS36, Q9NU50, Q9UJ06, Q9UJ07, Q9ULF8 | Gene names | SYNE1, KIAA0796, KIAA1262, KIAA1756, MYNE1 | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-1 (Nuclear envelope spectrin repeat protein 1) (Synaptic nuclear envelope protein 1) (Syne-1) (Myocyte nuclear envelope protein 1) (Myne-1) (Enaptin). | |||||
|
CENPF_HUMAN
|
||||||
| θ value | 5.27518 (rank : 69) | NC score | 0.040604 (rank : 78) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1932 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P49454, Q13171, Q13246 | Gene names | CENPF | |||
|
Domain Architecture |

|
|||||
| Description | Centromere protein F (Kinetochore protein CENP-F) (Mitosin) (AH antigen). | |||||
|
CING_MOUSE
|
||||||
| θ value | 5.27518 (rank : 70) | NC score | 0.045607 (rank : 67) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1432 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P59242 | Gene names | Cgn | |||
|
Domain Architecture |

|
|||||
| Description | Cingulin. | |||||
|
CROCC_MOUSE
|
||||||
| θ value | 5.27518 (rank : 71) | NC score | 0.045570 (rank : 68) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1460 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8CJ40, Q7TQL2, Q80U01, Q8R0B9 | Gene names | Crocc, Kiaa0445 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rootletin (Ciliary rootlet coiled-coil protein). | |||||
|
IFT81_MOUSE
|
||||||
| θ value | 5.27518 (rank : 72) | NC score | 0.034083 (rank : 85) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 959 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O35594, Q8R4W2, Q9CSA1, Q9EQM1 | Gene names | Ift81, Cdv-1, Cdv1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intraflagellar transport 81 (Carnitine deficiency-associated protein expressed in ventricle 1) (CDV-1 protein). | |||||
|
LIPB2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 73) | NC score | 0.031992 (rank : 87) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 570 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O35711, Q7TMG4, Q8CBS6, Q99KX6 | Gene names | Ppfibp2, Cclp1 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-beta-2 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein-binding protein 2) (PTPRF-interacting protein-binding protein 2) (Coiled-coil-like protein 1). | |||||
|
MY18A_HUMAN
|
||||||
| θ value | 5.27518 (rank : 74) | NC score | 0.031499 (rank : 90) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1929 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q92614, Q8IXP8 | Gene names | MYO18A, KIAA0216, MYSPDZ | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
RIMB1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 75) | NC score | 0.031649 (rank : 89) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q7TNF8, Q5NCP7, Q80TV9, Q8BIH5 | Gene names | Bzrap1, Kiaa0612, Rbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Peripheral-type benzodiazepine receptor-associated protein 1 (PRAX-1) (Peripheral benzodiazepine receptor-interacting protein) (PBR-IP) (RIM-binding protein 1) (RIM-BP1). | |||||
|
TPR_HUMAN
|
||||||
| θ value | 5.27518 (rank : 76) | NC score | 0.033890 (rank : 86) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
YAP1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 77) | NC score | 0.011746 (rank : 102) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P46938, Q52KJ5 | Gene names | Yap1, Yap, Yap65 | |||
|
Domain Architecture |

|
|||||
| Description | 65 kDa Yes-associated protein (YAP65). | |||||
|
AMOL2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 78) | NC score | 0.035288 (rank : 84) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 591 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8K371, Q3TPM1, Q7TPE4, Q8BP84, Q8BS08, Q9QUS0 | Gene names | Amotl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin-like protein 2. | |||||
|
ARHGC_HUMAN
|
||||||
| θ value | 6.88961 (rank : 79) | NC score | 0.016162 (rank : 100) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NZN5, O15086 | Gene names | ARHGEF12, KIAA0382, LARG | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 12 (Leukemia-associated RhoGEF). | |||||
|
ARNT_HUMAN
|
||||||
| θ value | 6.88961 (rank : 80) | NC score | 0.085911 (rank : 44) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P27540 | Gene names | ARNT | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
ARNT_MOUSE
|
||||||
| θ value | 6.88961 (rank : 81) | NC score | 0.078608 (rank : 46) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P53762, Q60661 | Gene names | Arnt | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
BRPF1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 82) | NC score | 0.011473 (rank : 103) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P55201, Q9UHI0 | Gene names | BRPF1, BR140 | |||
|
Domain Architecture |

|
|||||
| Description | Peregrin (Bromodomain and PHD finger-containing protein 1) (BR140 protein). | |||||
|
FIGLA_HUMAN
|
||||||
| θ value | 6.88961 (rank : 83) | NC score | 0.054390 (rank : 61) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q6QHK4 | Gene names | FIGLA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Factor in the germline alpha (Transcription factor FIGa) (FIGalpha). | |||||
|
SRBP2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 84) | NC score | 0.177631 (rank : 30) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q12772, Q9UH04 | Gene names | SREBF2, SREBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 2 (SREBP-2) (Sterol regulatory element-binding transcription factor 2). | |||||
|
STIM1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 85) | NC score | 0.026225 (rank : 96) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 665 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q13586 | Gene names | STIM1, GOK | |||
|
Domain Architecture |
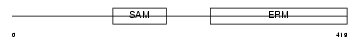
|
|||||
| Description | Stromal interaction molecule 1 precursor. | |||||
|
CCDC3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 86) | NC score | 0.017022 (rank : 99) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9BQI4 | Gene names | CCDC3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 3 precursor. | |||||
|
HIP1R_HUMAN
|
||||||
| θ value | 8.99809 (rank : 87) | NC score | 0.040427 (rank : 79) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 809 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O75146, Q9UED9 | Gene names | HIP1R, HIP12, KIAA0655 | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin-interacting protein 1-related protein (Hip1-related) (Hip 12). | |||||
|
K1802_MOUSE
|
||||||
| θ value | 8.99809 (rank : 88) | NC score | 0.007157 (rank : 105) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
MY18B_HUMAN
|
||||||
| θ value | 8.99809 (rank : 89) | NC score | 0.029088 (rank : 92) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1299 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8IUG5, Q8NDI8, Q8TE65, Q8WWS0, Q96KH2, Q96KR8, Q96KR9 | Gene names | MYO18B | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18B (Myosin XVIIIb). | |||||
|
MYH6_MOUSE
|
||||||
| θ value | 8.99809 (rank : 90) | NC score | 0.026793 (rank : 94) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1501 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q02566, Q64258, Q64738 | Gene names | Myh6, Myhca | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
SNW1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 91) | NC score | 0.026437 (rank : 95) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q13573, Q13483, Q32N03, Q5D0D6 | Gene names | SNW1, SKIIP, SKIP | |||
|
Domain Architecture |
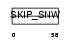
|
|||||
| Description | SNW domain-containing protein 1 (Nuclear protein SkiP) (Ski- interacting protein) (Nuclear receptor coactivator NCoA-62). | |||||
|
SNW1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 92) | NC score | 0.026225 (rank : 97) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9CSN1 | Gene names | Snw1, Skiip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SNW domain-containing protein 1 (Nuclear protein SkiP) (Ski- interacting protein). | |||||
|
BHLH8_MOUSE
|
||||||
| θ value | θ > 10 (rank : 93) | NC score | 0.061209 (rank : 56) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9QYC3, Q9QYE4 | Gene names | Bhlhb8, Mist1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Class B basic helix-loop-helix protein 8 (bHLHB8) (Muscle, intestine and stomach expression 1) (MIST-1). | |||||
|
HES1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 94) | NC score | 0.093307 (rank : 41) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q14469 | Gene names | HES1, HL, HRY | |||
|
Domain Architecture |
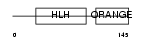
|
|||||
| Description | Transcription factor HES-1 (Hairy and enhancer of split 1) (Hairy- like) (HHL) (Hairy homolog). | |||||
|
HES1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 95) | NC score | 0.093715 (rank : 40) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P35428 | Gene names | Hes1, Hes-1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-1 (Hairy and enhancer of split 1). | |||||
|
HES4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 96) | NC score | 0.072754 (rank : 51) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9HCC6 | Gene names | HES4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-4 (Hairy and enhancer of split 4) (bHLH factor Hes4). | |||||
|
HEY2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 97) | NC score | 0.050810 (rank : 64) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9UBP5 | Gene names | HEY2, CHF1, GRL, HERP, HERP1, HRT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Cardiovascular helix-loop-helix factor 1) (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES- related repressor protein 2) (Protein gridlock homolog). | |||||
|
HEY2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 98) | NC score | 0.050924 (rank : 63) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9QUS4, Q3TZ99, Q8CD44 | Gene names | Hey2, Chf1, Herp, Herp1, Hesr2, Hrt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES-related repressor protein 2). | |||||
|
MLX_HUMAN
|
||||||
| θ value | θ > 10 (rank : 99) | NC score | 0.077650 (rank : 47) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9UH92, Q53XM6, Q96FL2, Q9H2V0, Q9H2V1, Q9H2V2, Q9NXN3 | Gene names | MLX, TCFL4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
MLX_MOUSE
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.080152 (rank : 45) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O08609, Q9EQJ7, Q9EQJ8 | Gene names | Mlx, Tcfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
MYCB_MOUSE
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.145098 (rank : 35) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6P8Z1, Q9D6E3 | Gene names | Mycb, Bmyc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein B-Myc. | |||||
|
MYCS_MOUSE
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.207112 (rank : 29) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9Z304 | Gene names | Mycs | |||
|
Domain Architecture |
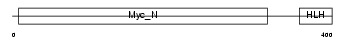
|
|||||
| Description | Protein S-myc. | |||||
|
TCFL5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.153789 (rank : 32) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UL49, O94771, Q9BYW0 | Gene names | TCFL5, CHA, E2BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor-like 5 protein (Cha transcription factor) (HPV-16 E2-binding protein 1) (E2BP-1). | |||||
|
TFAP4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.112162 (rank : 39) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q01664, O60409 | Gene names | TFAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor AP-4 (Activating enhancer-binding protein 4). | |||||
|
WBS14_HUMAN
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.062311 (rank : 55) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NP71, Q96E48, Q9BY03, Q9BY04, Q9BY05, Q9BY06, Q9Y2P3 | Gene names | MLXIPL, MIO, WBSCR14 | |||
|
Domain Architecture |
|
|||||
| Description | Williams-Beuren syndrome chromosome region 14 protein (WS basic-helix- loop-helix leucine zipper protein) (WS-bHLH) (Mlx interactor) (MLX- interacting protein-like). | |||||
|
MAD3_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 8.81223e-72 (rank : 1) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 92 | |
| SwissProt Accessions | Q9BW11, Q53HK1, Q7Z4Y0, Q8NDJ7, Q96ME3 | Gene names | MXD3, MAD3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Max-interacting transcriptional repressor MAD3 (Max-associated protein 3) (MAX dimerization protein 3) (Myx). | |||||
|
MAD3_MOUSE
|
||||||
| NC score | 0.972694 (rank : 2) | θ value | 4.24587e-58 (rank : 2) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q80US8, Q60947 | Gene names | Mxd3, Mad3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Max-interacting transcriptional repressor MAD3 (Max-associated protein 3) (MAX dimerization protein 3). | |||||
|
MXI1_HUMAN
|
||||||
| NC score | 0.902806 (rank : 3) | θ value | 1.99067e-23 (rank : 8) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P50539, Q15887 | Gene names | MXI1 | |||
|
Domain Architecture |

|
|||||
| Description | MAX-interacting protein 1 (Protein MXI1). | |||||
|
MXI1_MOUSE
|
||||||
| NC score | 0.897169 (rank : 4) | θ value | 2.35151e-24 (rank : 7) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P50540 | Gene names | Mxi1 | |||
|
Domain Architecture |

|
|||||
| Description | MAX-interacting protein 1 (Protein MXI1). | |||||
|
MAD4_HUMAN
|
||||||
| NC score | 0.895203 (rank : 5) | θ value | 5.06226e-27 (rank : 5) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q14582, Q5TZX4 | Gene names | MXD4, MAD4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-interacting transcriptional repressor MAD4 (Max-associated protein 4) (MAX dimerization protein 4). | |||||
|
MAD1_MOUSE
|
||||||
| NC score | 0.892487 (rank : 6) | θ value | 3.3877e-31 (rank : 3) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P50538, Q60798, Q61825 | Gene names | Mxd1, Mad, Mad1 | |||
|
Domain Architecture |

|
|||||
| Description | MAD protein (MAX dimerizer) (MAX dimerization protein 1). | |||||
|
MAD4_MOUSE
|
||||||
| NC score | 0.891616 (rank : 7) | θ value | 5.06226e-27 (rank : 6) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q60948 | Gene names | Mxd4, Mad4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-interacting transcriptional repressor MAD4 (Max-associated protein 4) (MAX dimerization protein 4). | |||||
|
MAD1_HUMAN
|
||||||
| NC score | 0.890053 (rank : 8) | θ value | 1.02001e-27 (rank : 4) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q05195 | Gene names | MXD1, MAD | |||
|
Domain Architecture |

|
|||||
| Description | MAD protein (MAX dimerizer) (MAX dimerization protein 1). | |||||
|
MNT_MOUSE
|
||||||
| NC score | 0.519868 (rank : 9) | θ value | 9.59137e-10 (rank : 9) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 432 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O08789, P97349 | Gene names | Mnt, Rox | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
MNT_HUMAN
|
||||||
| NC score | 0.499335 (rank : 10) | θ value | 1.63604e-09 (rank : 10) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 496 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q99583 | Gene names | MNT, ROX | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
MYC_MOUSE
|
||||||
| NC score | 0.386049 (rank : 11) | θ value | 5.81887e-07 (rank : 11) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P01108, P70247, Q3UM70, Q61422 | Gene names | Myc | |||
|
Domain Architecture |

|
|||||
| Description | Myc proto-oncogene protein (c-Myc) (Transcription factor p64). | |||||
|
MYC_HUMAN
|
||||||
| NC score | 0.384205 (rank : 12) | θ value | 1.29631e-06 (rank : 12) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P01106, P01107, Q14026 | Gene names | MYC | |||
|
Domain Architecture |

|
|||||
| Description | Myc proto-oncogene protein (c-Myc) (Transcription factor p64). | |||||
|
MYCL2_HUMAN
|
||||||
| NC score | 0.322557 (rank : 13) | θ value | 0.00020696 (rank : 13) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P12525 | Gene names | MYCL2 | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-2 protein. | |||||
|
USF2_MOUSE
|
||||||
| NC score | 0.314071 (rank : 14) | θ value | 0.00509761 (rank : 15) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q64705, Q3UIL2, Q8VE59 | Gene names | Usf2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (Major late transcription factor 2). | |||||
|
USF2_HUMAN
|
||||||
| NC score | 0.313247 (rank : 15) | θ value | 0.0113563 (rank : 17) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q15853, O00671, O00709, Q05750, Q07952, Q15851, Q15852 | Gene names | USF2 | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 2 (Upstream transcription factor 2) (FOS- interacting protein) (FIP) (Major late transcription factor 2). | |||||
|
MAX_HUMAN
|
||||||
| NC score | 0.310358 (rank : 16) | θ value | 1.06291 (rank : 40) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P61244, P25912, P52163 | Gene names | MAX | |||
|
Domain Architecture |

|
|||||
| Description | Protein max (Myc-associated factor X). | |||||
|
MAX_MOUSE
|
||||||
| NC score | 0.303550 (rank : 17) | θ value | 1.81305 (rank : 50) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P28574 | Gene names | Max, Myn | |||
|
Domain Architecture |

|
|||||
| Description | Protein max (Myc-associated factor X) (Protein myn) (Myc-binding novel HLH/LZ protein). | |||||
|
MYCL1_HUMAN
|
||||||
| NC score | 0.295018 (rank : 18) | θ value | 0.0736092 (rank : 20) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P12524, Q14897, Q5QPL0, Q5QPL1, Q9NUE9 | Gene names | MYCL1, LMYC, MYCL | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
MYCL1_MOUSE
|
||||||
| NC score | 0.292473 (rank : 19) | θ value | 0.163984 (rank : 27) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P10166, Q5FWI7 | Gene names | Mycl1, Lmyc1, Mycl | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
USF1_MOUSE
|
||||||
| NC score | 0.284224 (rank : 20) | θ value | 1.06291 (rank : 45) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q61069 | Gene names | Usf1, Usf | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
USF1_HUMAN
|
||||||
| NC score | 0.283053 (rank : 21) | θ value | 1.06291 (rank : 44) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P22415 | Gene names | USF1, USF | |||
|
Domain Architecture |

|
|||||
| Description | Upstream stimulatory factor 1 (Major late transcription factor 1). | |||||
|
TFE3_HUMAN
|
||||||
| NC score | 0.273559 (rank : 22) | θ value | 0.000786445 (rank : 14) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P19532, Q92757, Q92758, Q99964 | Gene names | TFE3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E3. | |||||
|
MYCN_MOUSE
|
||||||
| NC score | 0.257458 (rank : 23) | θ value | 3.0926 (rank : 58) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P03966, Q61978 | Gene names | Mycn, Nmyc, Nmyc1 | |||
|
Domain Architecture |

|
|||||
| Description | N-myc proto-oncogene protein. | |||||
|
MYCN_HUMAN
|
||||||
| NC score | 0.256197 (rank : 24) | θ value | 3.0926 (rank : 57) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P04198, Q6LDT9 | Gene names | MYCN, NMYC | |||
|
Domain Architecture |

|
|||||
| Description | N-myc proto-oncogene protein. | |||||
|
MITF_HUMAN
|
||||||
| NC score | 0.255856 (rank : 25) | θ value | 0.0252991 (rank : 18) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O75030, Q14841, Q9P2V0, Q9P2V1, Q9P2V2, Q9P2Y8 | Gene names | MITF | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
MITF_MOUSE
|
||||||
| NC score | 0.246156 (rank : 26) | θ value | 0.0563607 (rank : 19) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q08874, O08885, O88203, Q08843, Q60781, Q60782, Q9JIJ0, Q9JIJ1, Q9JIJ2, Q9JIJ3, Q9JIJ4, Q9JIJ5, Q9JIJ6, Q9JKX9 | Gene names | Mitf, Bw, Mi, Vit | |||
|
Domain Architecture |

|
|||||
| Description | Microphthalmia-associated transcription factor. | |||||
|
TFEB_HUMAN
|
||||||
| NC score | 0.211619 (rank : 27) | θ value | 1.06291 (rank : 43) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P19484, Q709B3, Q7Z6P9, Q9BRJ5, Q9UJD8 | Gene names | TFEB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
TFEB_MOUSE
|
||||||
| NC score | 0.211359 (rank : 28) | θ value | 3.0926 (rank : 60) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9R210 | Gene names | Tfeb, Tcfeb | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor EB. | |||||
|
MYCS_MOUSE
|
||||||
| NC score | 0.207112 (rank : 29) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9Z304 | Gene names | Mycs | |||
|
Domain Architecture |

|
|||||
| Description | Protein S-myc. | |||||
|
SRBP2_HUMAN
|
||||||
| NC score | 0.177631 (rank : 30) | θ value | 6.88961 (rank : 84) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q12772, Q9UH04 | Gene names | SREBF2, SREBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 2 (SREBP-2) (Sterol regulatory element-binding transcription factor 2). | |||||
|
SRBP1_MOUSE
|
||||||
| NC score | 0.171002 (rank : 31) | θ value | 0.0113563 (rank : 16) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9WTN3 | Gene names | Srebf1, Srebp1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
TCFL5_HUMAN
|
||||||
| NC score | 0.153789 (rank : 32) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UL49, O94771, Q9BYW0 | Gene names | TCFL5, CHA, E2BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor-like 5 protein (Cha transcription factor) (HPV-16 E2-binding protein 1) (E2BP-1). | |||||
|
BHLH3_HUMAN
|
||||||
| NC score | 0.151486 (rank : 33) | θ value | 1.38821 (rank : 47) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9C0J9 | Gene names | BHLHB3, DEC2, SHARP1 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (hDEC2) (Enhancer-of-split and hairy-related protein 1) (SHARP-1). | |||||
|
SRBP1_HUMAN
|
||||||
| NC score | 0.150475 (rank : 34) | θ value | 0.47712 (rank : 32) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P36956, Q16062 | Gene names | SREBF1, SREBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
MYCB_MOUSE
|
||||||
| NC score | 0.145098 (rank : 35) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6P8Z1, Q9D6E3 | Gene names | Mycb, Bmyc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein B-Myc. | |||||
|
BHLH3_MOUSE
|
||||||
| NC score | 0.143988 (rank : 36) | θ value | 1.81305 (rank : 49) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q99PV5 | Gene names | Bhlhb3, Dec2 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (mDEC2). | |||||
|
BHLH2_MOUSE
|
||||||
| NC score | 0.139083 (rank : 37) | θ value | 1.38821 (rank : 46) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O35185, P97289 | Gene names | Bhlhb2, Clast5, Stra13 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 2 (bHLHB2) (Stimulated with retinoic acid 13) (E47 interaction protein 1) (eipl). | |||||
|
BHLH2_HUMAN
|
||||||
| NC score | 0.127778 (rank : 38) | θ value | 2.36792 (rank : 51) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O14503, Q96TD3 | Gene names | BHLHB2, DEC1, SHARP2, STRA13 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 2 (bHLHB2) (Differentially expressed in chondrocytes protein 1) (DEC1) (Enhancer-of-split and hairy-related protein 2) (SHARP-2) (Stimulated with retinoic acid 13). | |||||
|
TFAP4_HUMAN
|
||||||
| NC score | 0.112162 (rank : 39) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q01664, O60409 | Gene names | TFAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor AP-4 (Activating enhancer-binding protein 4). | |||||
|
HES1_MOUSE
|
||||||
| NC score | 0.093715 (rank : 40) | θ value | θ > 10 (rank : 95) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P35428 | Gene names | Hes1, Hes-1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-1 (Hairy and enhancer of split 1). | |||||
|
HES1_HUMAN
|
||||||
| NC score | 0.093307 (rank : 41) | θ value | θ > 10 (rank : 94) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q14469 | Gene names | HES1, HL, HRY | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-1 (Hairy and enhancer of split 1) (Hairy- like) (HHL) (Hairy homolog). | |||||
|
BMAL1_HUMAN
|
||||||
| NC score | 0.089841 (rank : 42) | θ value | 3.0926 (rank : 55) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O00327, O00313, O00314, O00315, O00316, O00317, Q99631, Q99649 | Gene names | ARNTL, BMAL1, MOP3 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Member of PAS protein 3) (Basic-helix-loop- helix-PAS orphan MOP3) (bHLH-PAS protein JAP3). | |||||
|
BMAL1_MOUSE
|
||||||
| NC score | 0.088454 (rank : 43) | θ value | 3.0926 (rank : 56) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9WTL8, O88295, Q921S4, Q9R0U2, Q9WTL9 | Gene names | Arntl, Bmal1 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator-like protein 1 (Brain and muscle ARNT-like 1) (Arnt3). | |||||
|
ARNT_HUMAN
|
||||||
| NC score | 0.085911 (rank : 44) | θ value | 6.88961 (rank : 80) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P27540 | Gene names | ARNT | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
MLX_MOUSE
|
||||||
| NC score | 0.080152 (rank : 45) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O08609, Q9EQJ7, Q9EQJ8 | Gene names | Mlx, Tcfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
ARNT_MOUSE
|
||||||
| NC score | 0.078608 (rank : 46) | θ value | 6.88961 (rank : 81) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P53762, Q60661 | Gene names | Arnt | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator (ARNT protein) (Dioxin receptor, nuclear translocator) (Hypoxia-inducible factor 1 beta) (HIF-1 beta). | |||||
|
MLX_HUMAN
|
||||||
| NC score | 0.077650 (rank : 47) | θ value | θ > 10 (rank : 99) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9UH92, Q53XM6, Q96FL2, Q9H2V0, Q9H2V1, Q9H2V2, Q9NXN3 | Gene names | MLX, TCFL4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
CCD96_HUMAN
|
||||||
| NC score | 0.077033 (rank : 48) | θ value | 0.813845 (rank : 34) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 595 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q2M329, Q8N2I7 | Gene names | CCDC96 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 96. | |||||
|
ARNT2_MOUSE
|
||||||
| NC score | 0.073191 (rank : 49) | θ value | 4.03905 (rank : 62) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q61324 | Gene names | Arnt2 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
ARNT2_HUMAN
|
||||||
| NC score | 0.072784 (rank : 50) | θ value | 4.03905 (rank : 61) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9HBZ2, O15024 | Gene names | ARNT2, KIAA0307 | |||
|
Domain Architecture |

|
|||||
| Description | Aryl hydrocarbon receptor nuclear translocator 2 (ARNT protein 2). | |||||
|
HES4_HUMAN
|
||||||
| NC score | 0.072754 (rank : 51) | θ value | θ > 10 (rank : 96) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9HCC6 | Gene names | HES4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-4 (Hairy and enhancer of split 4) (bHLH factor Hes4). | |||||
|
CCD96_MOUSE
|
||||||
| NC score | 0.068215 (rank : 52) | θ value | 0.62314 (rank : 33) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9CR92 | Gene names | Ccdc96 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 96. | |||||
|
RABE2_MOUSE
|
||||||
| NC score | 0.063879 (rank : 53) | θ value | 0.0961366 (rank : 22) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 598 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q91WG2, Q99KN3 | Gene names | Rabep2, Rabpt5b | |||
|
Domain Architecture |

|
|||||
| Description | Rab GTPase-binding effector protein 2 (Rabaptin-5beta). | |||||
|
ZWINT_HUMAN
|
||||||
| NC score | 0.063631 (rank : 54) | θ value | 0.813845 (rank : 36) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O95229, Q9BWD0 | Gene names | ZWINT | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ZW10 interactor (ZW10-interacting protein 1) (Zwint-1). | |||||
|
WBS14_HUMAN
|
||||||
| NC score | 0.062311 (rank : 55) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NP71, Q96E48, Q9BY03, Q9BY04, Q9BY05, Q9BY06, Q9Y2P3 | Gene names | MLXIPL, MIO, WBSCR14 | |||
|
Domain Architecture |

|
|||||
| Description | Williams-Beuren syndrome chromosome region 14 protein (WS basic-helix- loop-helix leucine zipper protein) (WS-bHLH) (Mlx interactor) (MLX- interacting protein-like). | |||||
|
BHLH8_MOUSE
|
||||||
| NC score | 0.061209 (rank : 56) | θ value | θ > 10 (rank : 93) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9QYC3, Q9QYE4 | Gene names | Bhlhb8, Mist1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Class B basic helix-loop-helix protein 8 (bHLHB8) (Muscle, intestine and stomach expression 1) (MIST-1). | |||||
|
BHLH8_HUMAN
|
||||||
| NC score | 0.057991 (rank : 57) | θ value | 2.36792 (rank : 52) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q7RTS1 | Gene names | BHLHB8, MIST1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Class B basic helix-loop-helix protein 8 (bHLHB8) (Muscle, intestine and stomach expression 1) (MIST-1). | |||||
|
TACC2_HUMAN
|
||||||
| NC score | 0.057029 (rank : 58) | θ value | 0.365318 (rank : 29) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 856 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O95359, Q9NZ41, Q9NZR5 | Gene names | TACC2 | |||
|
Domain Architecture |

|
|||||
| Description | Transforming acidic coiled-coil-containing protein 2 (Anti Zuai-1) (AZU-1). | |||||
|
MD1L1_HUMAN
|
||||||
| NC score | 0.055999 (rank : 59) | θ value | 0.0961366 (rank : 21) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 893 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9Y6D9, Q13312, Q75MI0, Q86UM4, Q9UNH0 | Gene names | MAD1L1, MAD1, TXBP181 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitotic spindle assembly checkpoint protein MAD1 (Mitotic arrest deficient-like protein 1) (MAD1-like 1) (Mitotic checkpoint MAD1 protein-homolog) (HsMAD1) (hMAD1) (Tax-binding protein 181). | |||||
|
TACC2_MOUSE
|
||||||
| NC score | 0.055503 (rank : 60) | θ value | 0.813845 (rank : 35) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 774 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9JJG0 | Gene names | Tacc2 | |||
|
Domain Architecture |

|
|||||
| Description | Transforming acidic coiled-coil-containing protein 2. | |||||
|
FIGLA_HUMAN
|
||||||
| NC score | 0.054390 (rank : 61) | θ value | 6.88961 (rank : 83) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q6QHK4 | Gene names | FIGLA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Factor in the germline alpha (Transcription factor FIGa) (FIGalpha). | |||||
|
AZI1_MOUSE
|
||||||
| NC score | 0.052878 (rank : 62) | θ value | 1.06291 (rank : 37) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 990 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q62036 | Gene names | Azi1, Az1, Azi | |||
|
Domain Architecture |

|
|||||
| Description | 5-azacytidine-induced protein 1 (Pre-acrosome localization protein 1). | |||||
|
HEY2_MOUSE
|
||||||
| NC score | 0.050924 (rank : 63) | θ value | θ > 10 (rank : 98) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9QUS4, Q3TZ99, Q8CD44 | Gene names | Hey2, Chf1, Herp, Herp1, Hesr2, Hrt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES-related repressor protein 2). | |||||
|
HEY2_HUMAN
|
||||||
| NC score | 0.050810 (rank : 64) | θ value | θ > 10 (rank : 97) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9UBP5 | Gene names | HEY2, CHF1, GRL, HERP, HERP1, HRT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Cardiovascular helix-loop-helix factor 1) (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES- related repressor protein 2) (Protein gridlock homolog). | |||||
|
BPAEA_MOUSE
|
||||||
| NC score | 0.049777 (rank : 65) | θ value | 0.125558 (rank : 23) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1480 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q91ZU8, Q8K5D4 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoform 5 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
CROCC_HUMAN
|
||||||
| NC score | 0.048497 (rank : 66) | θ value | 1.06291 (rank : 38) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1409 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q5TZA2, Q2VHY3, Q66GT7, Q7Z2L4, Q7Z5D7 | Gene names | CROCC, KIAA0445 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rootletin (Ciliary rootlet coiled-coil protein). | |||||
|
CING_MOUSE
|
||||||
| NC score | 0.045607 (rank : 67) | θ value | 5.27518 (rank : 70) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1432 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P59242 | Gene names | Cgn | |||
|
Domain Architecture |

|
|||||
| Description | Cingulin. | |||||
|
CROCC_MOUSE
|
||||||
| NC score | 0.045570 (rank : 68) | θ value | 5.27518 (rank : 71) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1460 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8CJ40, Q7TQL2, Q80U01, Q8R0B9 | Gene names | Crocc, Kiaa0445 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rootletin (Ciliary rootlet coiled-coil protein). | |||||
|
HES2_MOUSE
|
||||||
| NC score | 0.044445 (rank : 69) | θ value | 4.03905 (rank : 65) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O54792 | Gene names | Hes2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor HES-2 (Hairy and enhancer of split 2). | |||||
|
CYLN2_MOUSE
|
||||||
| NC score | 0.043426 (rank : 70) | θ value | 1.06291 (rank : 39) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1002 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9Z0H8, Q7TSI9, Q8CHU1, Q9EP81 | Gene names | Cyln2, Kiaa0291 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoplasmic linker protein 2 (Cytoplasmic linker protein 115) (CLIP- 115). | |||||
|
CT2NL_MOUSE
|
||||||
| NC score | 0.043133 (rank : 71) | θ value | 0.47712 (rank : 30) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 902 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q99LJ0, Q6ZPR0, Q8BSV1 | Gene names | Cttnbp2nl, Kiaa1433 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CTTNBP2 N-terminal-like protein. | |||||
|
RCOR3_MOUSE
|
||||||
| NC score | 0.042309 (rank : 72) | θ value | 1.06291 (rank : 42) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q6PGA0 | Gene names | Rcor3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 3. | |||||
|
HIP1_HUMAN
|
||||||
| NC score | 0.042177 (rank : 73) | θ value | 4.03905 (rank : 66) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1016 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O00291, O00328, Q8TDL4 | Gene names | HIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin-interacting protein 1 (HIP-I). | |||||
|
CYLN2_HUMAN
|
||||||
| NC score | 0.041908 (rank : 74) | θ value | 4.03905 (rank : 64) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1004 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9UDT6, O14527, O43611 | Gene names | CYLN2, KIAA0291, WBSCR4, WSCR4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoplasmic linker protein 2 (Cytoplasmic linker protein 115) (CLIP- 115) (Williams-Beuren syndrome chromosome region 4 protein). | |||||
|
CP135_MOUSE
|
||||||
| NC score | 0.041583 (rank : 75) | θ value | 0.279714 (rank : 28) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1278 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q6P5D4, Q6A030 | Gene names | Cep135, Cep4, Kiaa0635 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 135 kDa (Cep135 protein) (Centrosomal protein 4). | |||||
|
CCD21_HUMAN
|
||||||
| NC score | 0.041241 (rank : 76) | θ value | 4.03905 (rank : 63) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1228 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q6P2H3, Q5VY68, Q5VY70, Q9H6Q1, Q9H828, Q9UF52 | Gene names | CCDC21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 21. | |||||
|
PLEC1_HUMAN
|
||||||
| NC score | 0.040932 (rank : 77) | θ value | 3.0926 (rank : 59) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1326 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q15149, Q15148, Q16640 | Gene names | PLEC1 | |||
|
Domain Architecture |

|
|||||
| Description | Plectin-1 (PLTN) (PCN) (Hemidesmosomal protein 1) (HD1). | |||||
|
CENPF_HUMAN
|
||||||
| NC score | 0.040604 (rank : 78) | θ value | 5.27518 (rank : 69) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1932 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P49454, Q13171, Q13246 | Gene names | CENPF | |||
|
Domain Architecture |

|
|||||
| Description | Centromere protein F (Kinetochore protein CENP-F) (Mitosin) (AH antigen). | |||||
|
HIP1R_HUMAN
|
||||||
| NC score | 0.040427 (rank : 79) | θ value | 8.99809 (rank : 87) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 809 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O75146, Q9UED9 | Gene names | HIP1R, HIP12, KIAA0655 | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin-interacting protein 1-related protein (Hip1-related) (Hip 12). | |||||
|
RCOR3_HUMAN
|
||||||
| NC score | 0.039831 (rank : 80) | θ value | 1.06291 (rank : 41) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9P2K3, Q5VT47, Q7L9I5, Q8N5U3, Q9NV83 | Gene names | RCOR3, KIAA1343 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 3. | |||||
|
GOGA5_MOUSE
|
||||||
| NC score | 0.038832 (rank : 81) | θ value | 0.47712 (rank : 31) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1434 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9QYE6, O88317, Q3TGE7, Q3U6S5, Q3UUF9 | Gene names | Golga5, Retii | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 5 (Golgin-84) (Sumiko protein) (Ret-II protein). | |||||
|
KIF5C_MOUSE
|
||||||
| NC score | 0.038530 (rank : 82) | θ value | 0.163984 (rank : 26) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1040 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P28738, Q9Z2F8 | Gene names | Kif5c, Nkhc2 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin heavy chain isoform 5C (Kinesin heavy chain neuron-specific 2). | |||||
|
KIF5C_HUMAN
|
||||||
| NC score | 0.036760 (rank : 83) | θ value | 2.36792 (rank : 53) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 974 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O60282, O95079 | Gene names | KIF5C, KIAA0531, NKHC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin heavy chain isoform 5C (Kinesin heavy chain neuron-specific 2). | |||||
|
AMOL2_MOUSE
|
||||||
| NC score | 0.035288 (rank : 84) | θ value | 6.88961 (rank : 78) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 591 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8K371, Q3TPM1, Q7TPE4, Q8BP84, Q8BS08, Q9QUS0 | Gene names | Amotl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin-like protein 2. | |||||
|
IFT81_MOUSE
|
||||||
| NC score | 0.034083 (rank : 85) | θ value | 5.27518 (rank : 72) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 959 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O35594, Q8R4W2, Q9CSA1, Q9EQM1 | Gene names | Ift81, Cdv-1, Cdv1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intraflagellar transport 81 (Carnitine deficiency-associated protein expressed in ventricle 1) (CDV-1 protein). | |||||
|
TPR_HUMAN
|
||||||
| NC score | 0.033890 (rank : 86) | θ value | 5.27518 (rank : 76) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
LIPB2_MOUSE
|
||||||
| NC score | 0.031992 (rank : 87) | θ value | 5.27518 (rank : 73) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 570 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O35711, Q7TMG4, Q8CBS6, Q99KX6 | Gene names | Ppfibp2, Cclp1 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-beta-2 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein-binding protein 2) (PTPRF-interacting protein-binding protein 2) (Coiled-coil-like protein 1). | |||||
|
MACF4_HUMAN
|
||||||
| NC score | 0.031820 (rank : 88) | θ value | 4.03905 (rank : 67) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1321 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q96PK2, Q8WXY1 | Gene names | MACF1, ABP620, ACF7, KIAA0465, KIAA1251 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1, isoform 4. | |||||
|
RIMB1_MOUSE
|
||||||
| NC score | 0.031649 (rank : 89) | θ value | 5.27518 (rank : 75) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q7TNF8, Q5NCP7, Q80TV9, Q8BIH5 | Gene names | Bzrap1, Kiaa0612, Rbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Peripheral-type benzodiazepine receptor-associated protein 1 (PRAX-1) (Peripheral benzodiazepine receptor-interacting protein) (PBR-IP) (RIM-binding protein 1) (RIM-BP1). | |||||
|
MY18A_HUMAN
|
||||||
| NC score | 0.031499 (rank : 90) | θ value | 5.27518 (rank : 74) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1929 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q92614, Q8IXP8 | Gene names | MYO18A, KIAA0216, MYSPDZ | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
SYNE1_HUMAN
|
||||||
| NC score | 0.031367 (rank : 91) | θ value | 4.03905 (rank : 68) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1484 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8NF91, O94890, Q5JV19, Q5JV22, Q8N9P7, Q8TCP1, Q8WWW6, Q8WWW7, Q8WXF6, Q96N17, Q9C0A7, Q9H525, Q9H526, Q9NS36, Q9NU50, Q9UJ06, Q9UJ07, Q9ULF8 | Gene names | SYNE1, KIAA0796, KIAA1262, KIAA1756, MYNE1 | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-1 (Nuclear envelope spectrin repeat protein 1) (Synaptic nuclear envelope protein 1) (Syne-1) (Myocyte nuclear envelope protein 1) (Myne-1) (Enaptin). | |||||
|
MY18B_HUMAN
|
||||||
| NC score | 0.029088 (rank : 92) | θ value | 8.99809 (rank : 89) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1299 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8IUG5, Q8NDI8, Q8TE65, Q8WWS0, Q96KH2, Q96KR8, Q96KR9 | Gene names | MYO18B | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18B (Myosin XVIIIb). | |||||
|
ZGPAT_MOUSE
|
||||||
| NC score | 0.028752 (rank : 93) | θ value | 2.36792 (rank : 54) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8VDM1, Q8BWW2 | Gene names | Zgpat | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger CCCH-type with G patch domain protein. | |||||
|
MYH6_MOUSE
|
||||||
| NC score | 0.026793 (rank : 94) | θ value | 8.99809 (rank : 90) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1501 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q02566, Q64258, Q64738 | Gene names | Myh6, Myhca | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle alpha isoform (MyHC-alpha). | |||||
|
SNW1_HUMAN
|
||||||
| NC score | 0.026437 (rank : 95) | θ value | 8.99809 (rank : 91) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q13573, Q13483, Q32N03, Q5D0D6 | Gene names | SNW1, SKIIP, SKIP | |||
|
Domain Architecture |

|
|||||
| Description | SNW domain-containing protein 1 (Nuclear protein SkiP) (Ski- interacting protein) (Nuclear receptor coactivator NCoA-62). | |||||
|
STIM1_HUMAN
|
||||||
| NC score | 0.026225 (rank : 96) | θ value | 6.88961 (rank : 85) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 665 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q13586 | Gene names | STIM1, GOK | |||
|
Domain Architecture |

|
|||||
| Description | Stromal interaction molecule 1 precursor. | |||||
|
SNW1_MOUSE
|
||||||
| NC score | 0.026225 (rank : 97) | θ value | 8.99809 (rank : 92) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9CSN1 | Gene names | Snw1, Skiip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SNW domain-containing protein 1 (Nuclear protein SkiP) (Ski- interacting protein). | |||||
|
ARHGC_MOUSE
|
||||||
| NC score | 0.018228 (rank : 98) | θ value | 1.81305 (rank : 48) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8R4H2, Q80U18 | Gene names | Arhgef12, Kiaa0382, Larg | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 12 (Leukemia-associated RhoGEF). | |||||
|
CCDC3_HUMAN
|
||||||
| NC score | 0.017022 (rank : 99) | θ value | 8.99809 (rank : 86) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9BQI4 | Gene names | CCDC3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 3 precursor. | |||||
|
ARHGC_HUMAN
|
||||||
| NC score | 0.016162 (rank : 100) | θ value | 6.88961 (rank : 79) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NZN5, O15086 | Gene names | ARHGEF12, KIAA0382, LARG | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 12 (Leukemia-associated RhoGEF). | |||||
|
ANKR6_HUMAN
|
||||||
| NC score | 0.015527 (rank : 101) | θ value | 0.163984 (rank : 25) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 439 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Y2G4, Q9NU24, Q9UFQ9 | Gene names | ANKRD6, KIAA0957 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin repeat domain-containing protein 6. | |||||
|
YAP1_MOUSE
|
||||||
| NC score | 0.011746 (rank : 102) | θ value | 5.27518 (rank : 77) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P46938, Q52KJ5 | Gene names | Yap1, Yap, Yap65 | |||
|
Domain Architecture |

|
|||||
| Description | 65 kDa Yes-associated protein (YAP65). | |||||
|
BRPF1_HUMAN
|
||||||
| NC score | 0.011473 (rank : 103) | θ value | 6.88961 (rank : 82) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P55201, Q9UHI0 | Gene names | BRPF1, BR140 | |||
|
Domain Architecture |

|
|||||
| Description | Peregrin (Bromodomain and PHD finger-containing protein 1) (BR140 protein). | |||||
|
NEK11_HUMAN
|
||||||
| NC score | 0.007302 (rank : 104) | θ value | 0.125558 (rank : 24) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1043 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8NG66, Q5JPC0, Q8NG65, Q8TBY1, Q9H5F4 | Gene names | NEK11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase Nek11 (EC 2.7.11.1) (NimA-related protein kinase 11) (Never in mitosis A-related kinase 11). | |||||
|
K1802_MOUSE
|
||||||
| NC score | 0.007157 (rank : 105) | θ value | 8.99809 (rank : 88) | |||
| Query Neighborhood Hits | 92 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||