Please be patient as the page loads
|
IER5_MOUSE
|
||||||
| SwissProt Accessions | O89113, Q8CGI0 | Gene names | Ier5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Immediate early response gene 5 protein. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
IER5_MOUSE
|
||||||
| θ value | 7.12574e-114 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 110 | |
| SwissProt Accessions | O89113, Q8CGI0 | Gene names | Ier5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Immediate early response gene 5 protein. | |||||
|
IER5_HUMAN
|
||||||
| θ value | 8.51556e-75 (rank : 2) | NC score | 0.930859 (rank : 2) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q5VY09, Q8WY68, Q9NY49, Q9NZP9 | Gene names | IER5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Immediate early response gene 5 protein. | |||||
|
IER2_HUMAN
|
||||||
| θ value | 2.0648e-12 (rank : 3) | NC score | 0.611754 (rank : 3) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BTL4, Q03827, Q2TAZ2 | Gene names | IER2, ETR101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Immediate early response gene 2 protein (Protein ETR101). | |||||
|
IER2_MOUSE
|
||||||
| θ value | 5.62301e-10 (rank : 4) | NC score | 0.577807 (rank : 4) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P17950, Q64251, Q8BME8, Q99M25 | Gene names | Ier2 | |||
|
Domain Architecture |

|
|||||
| Description | Immediate early response gene 2 protein (T-lymphocyte-activated protein) (Cycloheximide-induced gene protein) (CHX1) (Growth factor- inducible immediate early protein) (Proline-rich-induced protein 92) (Pip92). | |||||
|
PRRT3_HUMAN
|
||||||
| θ value | 0.000786445 (rank : 5) | NC score | 0.123134 (rank : 5) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q5FWE3, Q49AD0, Q6UXY6, Q8NBC9 | Gene names | PRRT3, UNQ5823/PRO19642 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
MUC1_HUMAN
|
||||||
| θ value | 0.00665767 (rank : 6) | NC score | 0.087546 (rank : 19) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
MAGE1_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 7) | NC score | 0.069072 (rank : 34) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
GNAS1_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 8) | NC score | 0.066924 (rank : 41) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 651 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q6R0H7, Q6R0H4, Q6R0H5, Q6R2J5, Q9JJX0, Q9Z1N8 | Gene names | Gnas, Gnas1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Guanine nucleotide-binding protein G(s) subunit alpha isoforms XLas (Adenylate cyclase-stimulating G alpha protein) (Extra large alphas protein) (XLalphas). | |||||
|
ROBO3_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 9) | NC score | 0.044145 (rank : 88) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 570 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q96MS0 | Gene names | ROBO3 | |||
|
Domain Architecture |

|
|||||
| Description | Roundabout homolog 3 precursor (Roundabout-like protein 3). | |||||
|
TM16H_HUMAN
|
||||||
| θ value | 0.163984 (rank : 10) | NC score | 0.071330 (rank : 30) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9HCE9 | Gene names | TMEM16H, KIAA1623 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 16H. | |||||
|
NOTC3_HUMAN
|
||||||
| θ value | 0.21417 (rank : 11) | NC score | 0.024503 (rank : 120) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1062 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9UM47, Q9UEB3, Q9UPL3, Q9Y6L8 | Gene names | NOTCH3 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 3 precursor (Notch 3) [Contains: Notch 3 extracellular truncation; Notch 3 intracellular domain]. | |||||
|
ROBO3_MOUSE
|
||||||
| θ value | 0.21417 (rank : 12) | NC score | 0.041404 (rank : 94) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 524 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9Z2I4 | Gene names | Robo3, Rbig1 | |||
|
Domain Architecture |

|
|||||
| Description | Roundabout homolog 3 precursor (Retinoblastoma-inhibiting gene 1) (Rig-1). | |||||
|
FXL19_MOUSE
|
||||||
| θ value | 0.279714 (rank : 13) | NC score | 0.066455 (rank : 42) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q6PB97, Q7TSH0, Q8BIB4 | Gene names | Fbxl19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 19 (F-box and leucine-rich repeat protein 19). | |||||
|
IASPP_MOUSE
|
||||||
| θ value | 0.279714 (rank : 14) | NC score | 0.058762 (rank : 57) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1024 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q5I1X5 | Gene names | Ppp1r13l, Nkip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RelA-associated inhibitor (Inhibitor of ASPP protein) (Protein iASPP) (PPP1R13B-like protein) (NFkB-interacting protein 1). | |||||
|
IQEC3_MOUSE
|
||||||
| θ value | 0.279714 (rank : 15) | NC score | 0.045090 (rank : 86) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q3TES0, Q3UHP8, Q80TJ8 | Gene names | Iqsec3, Kiaa1110 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 3. | |||||
|
MARCS_MOUSE
|
||||||
| θ value | 0.279714 (rank : 16) | NC score | 0.098326 (rank : 12) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P26645 | Gene names | Marcks, Macs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS). | |||||
|
CO7A1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 17) | NC score | 0.041884 (rank : 93) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 807 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q02388, Q14054, Q16507 | Gene names | COL7A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(VII) chain precursor (Long-chain collagen) (LC collagen). | |||||
|
IP3KC_MOUSE
|
||||||
| θ value | 0.365318 (rank : 18) | NC score | 0.047209 (rank : 83) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7TS72, Q3U384 | Gene names | Itpkc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol-trisphosphate 3-kinase C (EC 2.7.1.127) (Inositol 1,4,5- trisphosphate 3-kinase C) (IP3K-C). | |||||
|
RIMB1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 19) | NC score | 0.056294 (rank : 61) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1479 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O95153, O75111, Q8N5W3 | Gene names | BZRAP1, KIAA0612, RBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Peripheral-type benzodiazepine receptor-associated protein 1 (PRAX-1) (Peripheral benzodiazepine receptor-interacting protein) (PBR-IP) (RIM-binding protein 1) (RIM-BP1). | |||||
|
WNK2_HUMAN
|
||||||
| θ value | 0.365318 (rank : 20) | NC score | 0.031422 (rank : 107) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1407 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9Y3S1, Q8IY36, Q9C0A3, Q9H3P4 | Gene names | WNK2, KIAA1760, PRKWNK2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK2 (EC 2.7.11.1) (Protein kinase with no lysine 2) (Protein kinase, lysine-deficient 2). | |||||
|
YLPM1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 21) | NC score | 0.089704 (rank : 17) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P49750, P49752, Q96I64, Q9P1V7 | Gene names | YLPM1, C14orf170, ZAP3 | |||
|
Domain Architecture |

|
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3) (ZAP113). | |||||
|
ZN385_HUMAN
|
||||||
| θ value | 0.47712 (rank : 22) | NC score | 0.060390 (rank : 53) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 614 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q96PM9, Q5VH53, Q9H7R6, Q9UFU3 | Gene names | ZNF385, HZF, RZF | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 385 (Hematopoietic zinc finger protein) (Retinal zinc finger protein). | |||||
|
HXA13_HUMAN
|
||||||
| θ value | 0.813845 (rank : 23) | NC score | 0.025683 (rank : 116) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 353 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P31271, O43371 | Gene names | HOXA13, HOX1J | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-A13 (Hox-1J). | |||||
|
OGFR_HUMAN
|
||||||
| θ value | 0.813845 (rank : 24) | NC score | 0.097047 (rank : 13) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
RGS3_MOUSE
|
||||||
| θ value | 0.813845 (rank : 25) | NC score | 0.071721 (rank : 28) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
SF3B1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 26) | NC score | 0.107371 (rank : 7) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O75533 | Gene names | SF3B1, SAP155 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3B subunit 1 (Spliceosome-associated protein 155) (SAP 155) (SF3b155) (Pre-mRNA-splicing factor SF3b 155 kDa subunit). | |||||
|
SF3B1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 27) | NC score | 0.107710 (rank : 6) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q99NB9, Q9CSK5 | Gene names | Sf3b1, Sap155 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3B subunit 1 (Spliceosome-associated protein 155) (SAP 155) (SF3b155) (Pre-mRNA-splicing factor SF3b 155 kDa subunit). | |||||
|
TCGAP_HUMAN
|
||||||
| θ value | 0.813845 (rank : 28) | NC score | 0.046295 (rank : 84) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 872 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O14559, O14552, O14560, Q6ZSP6, Q96CP3, Q9NT23 | Gene names | SNX26, TCGAP | |||
|
Domain Architecture |

|
|||||
| Description | TC10/CDC42 GTPase-activating protein (Sorting nexin-26). | |||||
|
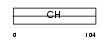
MILK1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 29) | NC score | 0.052038 (rank : 76) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8N3F8, Q5TI16, Q7RTP5, Q8N3N8, Q9BVL9, Q9BY92, Q9UH43, Q9UH44, Q9UH45 | Gene names | MIRAB13, KIAA1668 | |||
|
Domain Architecture |
|
|||||
| Description | Molecule interacting with Rab13 (MIRab13) (MICAL-like protein 1). | |||||
|
MYO9B_HUMAN
|
||||||
| θ value | 1.06291 (rank : 30) | NC score | 0.020435 (rank : 129) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 641 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q13459, O75314, Q9NUJ2, Q9UHN0 | Gene names | MYO9B, MYR5 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9B (Myosin IXb) (Unconventional myosin-9b). | |||||
|
NOTC4_HUMAN
|
||||||
| θ value | 1.06291 (rank : 31) | NC score | 0.019656 (rank : 131) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 905 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q99466, O00306, Q99458, Q99940, Q9H3S8, Q9UII9, Q9UIJ0 | Gene names | NOTCH4 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 4 precursor (Notch 4) (hNotch4) [Contains: Notch 4 extracellular truncation; Notch 4 intracellular domain]. | |||||
|
RECQ4_HUMAN
|
||||||
| θ value | 1.06291 (rank : 32) | NC score | 0.037939 (rank : 103) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O94761, Q3Y424, Q96DW2, Q96F55 | Gene names | RECQL4, RECQ4 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent DNA helicase Q4 (EC 3.6.1.-) (RecQ protein-like 4) (RecQ4) (RTS). | |||||
|
RTEL1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 33) | NC score | 0.042783 (rank : 91) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NZ71, Q5JTV4, Q9BW37, Q9H402, Q9H4X6, Q9NX25, Q9NZ73, Q9UPR4, Q9Y4R6 | Gene names | RTEL1, C20orf41, KIAA1088, NHL | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of telomere elongation helicase 1 (EC 3.6.1.-) (Helicase- like protein NHL). | |||||
|
BASP_MOUSE
|
||||||
| θ value | 1.38821 (rank : 34) | NC score | 0.100363 (rank : 11) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
CEL_HUMAN
|
||||||
| θ value | 1.38821 (rank : 35) | NC score | 0.050278 (rank : 81) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P19835, Q16398 | Gene names | CEL, BAL | |||
|
Domain Architecture |

|
|||||
| Description | Bile salt-activated lipase precursor (EC 3.1.1.3) (EC 3.1.1.13) (BAL) (Bile salt-stimulated lipase) (BSSL) (Carboxyl ester lipase) (Sterol esterase) (Cholesterol esterase) (Pancreatic lysophospholipase). | |||||
|
HCN2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 36) | NC score | 0.042152 (rank : 92) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 369 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9UL51, O60742, O60743, O75267, Q9UBS2 | Gene names | HCN2, BCNG2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 2 (Brain cyclic nucleotide-gated channel 2) (BCNG-2). | |||||
|
INVS_MOUSE
|
||||||
| θ value | 1.38821 (rank : 37) | NC score | 0.026102 (rank : 115) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 609 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O89019, O88849 | Gene names | Invs, Inv, Nphp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inversin (Inversion of embryo turning protein) (Nephrocystin-2). | |||||
|
SALL3_HUMAN
|
||||||
| θ value | 1.38821 (rank : 38) | NC score | 0.018237 (rank : 132) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1066 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9BXA9, Q9UGH1 | Gene names | SALL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sal-like protein 3 (Zinc finger protein SALL3) (hSALL3). | |||||
|
GP135_HUMAN
|
||||||
| θ value | 1.81305 (rank : 39) | NC score | 0.010737 (rank : 144) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 669 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8IZ08, Q7Z604, Q86SM3, Q8NH39 | Gene names | GPR135 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 135. | |||||
|
HXA13_MOUSE
|
||||||
| θ value | 1.81305 (rank : 40) | NC score | 0.023466 (rank : 127) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q62424 | Gene names | Hoxa13, Hox-1.10 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-A13 (Hox-1.10). | |||||
|
AGRIN_HUMAN
|
||||||
| θ value | 2.36792 (rank : 41) | NC score | 0.023415 (rank : 128) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 582 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O00468, Q5SVA2, Q60FE1, Q7KYS8, Q8N4J5, Q96IC1, Q9BTD4 | Gene names | AGRIN, AGRN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Agrin precursor. | |||||
|
CSKI1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 42) | NC score | 0.039462 (rank : 97) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1031 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q6P9K8, Q6ZPU2, Q8BWU2, Q8BX99, Q9CXH0 | Gene names | Caskin1, Kiaa1306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-1 (CASK-interacting protein 1). | |||||
|
FXL19_HUMAN
|
||||||
| θ value | 2.36792 (rank : 43) | NC score | 0.060474 (rank : 52) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6PCT2, Q8N789, Q9NT14 | Gene names | FBXL19, FBL19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 19 (F-box and leucine-rich repeat protein 19). | |||||
|
GP108_HUMAN
|
||||||
| θ value | 2.36792 (rank : 44) | NC score | 0.028896 (rank : 108) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NPR9 | Gene names | GPR108 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein GPR108 precursor. | |||||
|
HRX_HUMAN
|
||||||
| θ value | 2.36792 (rank : 45) | NC score | 0.038302 (rank : 100) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
KKCC1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 46) | NC score | 0.007535 (rank : 154) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 866 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8VBY2, Q9R054 | Gene names | Camkk1, Camkk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium/calmodulin-dependent protein kinase kinase 1 (EC 2.7.11.17) (Calcium/calmodulin-dependent protein kinase kinase alpha) (CaM-kinase kinase alpha) (CaM-KK alpha) (CaMKK alpha) (CaMKK 1) (CaM-kinase IV kinase). | |||||
|
MRCKB_MOUSE
|
||||||
| θ value | 2.36792 (rank : 47) | NC score | 0.006173 (rank : 156) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 2180 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q7TT50, Q69ZR5, Q80W33 | Gene names | Cdc42bpb, Kiaa1124 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK beta (EC 2.7.11.1) (CDC42-binding protein kinase beta) (Myotonic dystrophy kinase-related CDC42-binding kinase beta) (Myotonic dystrophy protein kinase-like beta) (MRCK beta) (DMPK-like beta). | |||||
|
NRCAM_HUMAN
|
||||||
| θ value | 2.36792 (rank : 48) | NC score | 0.026735 (rank : 111) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 439 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q92823, O15051, O15179, Q9UHI3, Q9UHI4 | Gene names | NRCAM, KIAA0343 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal cell adhesion molecule precursor (Nr-CAM) (NgCAM-related cell adhesion molecule) (Ng-CAM-related) (hBravo). | |||||
|
NRCAM_MOUSE
|
||||||
| θ value | 2.36792 (rank : 49) | NC score | 0.026222 (rank : 114) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 432 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q810U4, Q80U33, Q8BLG8, Q8BX92, Q8BYJ8 | Gene names | Nrcam, Kiaa0343 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal cell adhesion molecule precursor (Nr-CAM) (NgCAM-related cell adhesion molecule) (Ng-CAM-related) (mBravo). | |||||
|
OBF1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 50) | NC score | 0.067069 (rank : 40) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q16633, Q14983 | Gene names | POU2AF1, OBF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | POU domain class 2-associating factor 1 (B-cell-specific coactivator OBF-1) (OCT-binding factor 1) (BOB-1) (OCA-B). | |||||
|
OBF1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 51) | NC score | 0.058887 (rank : 56) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q64693 | Gene names | Pou2af1, Obf-1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | POU domain class 2-associating factor 1 (B-cell-specific coactivator OBF-1) (OCT-binding factor 1) (BOB-1) (BOB1) (OCA-B). | |||||
|
BASP_HUMAN
|
||||||
| θ value | 3.0926 (rank : 52) | NC score | 0.089157 (rank : 18) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
CSPG5_HUMAN
|
||||||
| θ value | 3.0926 (rank : 53) | NC score | 0.043824 (rank : 90) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O95196, Q71M39, Q71M40 | Gene names | CSPG5, CALEB, NGC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate proteoglycan 5 precursor (Neuroglycan C) (Acidic leucine-rich EGF-like domain-containing brain protein). | |||||
|
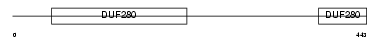
DRD4_HUMAN
|
||||||
| θ value | 3.0926 (rank : 54) | NC score | 0.011234 (rank : 141) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 886 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P21917 | Gene names | DRD4 | |||
|
Domain Architecture |
|
|||||
| Description | D(4) dopamine receptor (Dopamine D4 receptor) (D(2C) dopamine receptor). | |||||
|
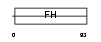
FOXC1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 55) | NC score | 0.023889 (rank : 123) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q12948, Q86UP7, Q9BYM1, Q9NUE5, Q9UDD0, Q9UP06 | Gene names | FOXC1, FKHL7, FREAC3 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein C1 (Forkhead-related protein FKHL7) (Forkhead- related transcription factor 3) (FREAC-3). | |||||
|
JPH2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 56) | NC score | 0.037997 (rank : 102) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 639 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9BR39, O95913, Q5JY74, Q9UJN4 | Gene names | JPH2, JP2 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-2 (Junctophilin type 2) (JP-2). | |||||
|
KLF14_HUMAN
|
||||||
| θ value | 3.0926 (rank : 57) | NC score | 0.005643 (rank : 157) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 725 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8TD94 | Gene names | KLF14, BTEB5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Krueppel-like factor 14 (Transcription factor BTEB5) (Basic transcription element-binding protein 5) (BTE-binding protein 5). | |||||
|
KLF2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 58) | NC score | 0.009092 (rank : 149) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 755 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Y5W3, Q9UJS5, Q9UKR6 | Gene names | KLF2, LKLF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Krueppel-like factor 2 (Lung krueppel-like factor). | |||||
|
PO4F2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 59) | NC score | 0.013433 (rank : 136) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q63934, Q63954 | Gene names | Pou4f2, Brn3b | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 4, transcription factor 2 (Brain-specific homeobox/POU domain protein 3B) (Brn-3B) (Brn-3.2). | |||||
|
SRTD2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 60) | NC score | 0.035514 (rank : 105) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q14140 | Gene names | SERTAD2, KIAA0127 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SERTA domain-containing protein 2 (Transcriptional regulator interacting with the PHD-bromodomain 2) (TRIP-Br2). | |||||
|
ZN447_HUMAN
|
||||||
| θ value | 3.0926 (rank : 61) | NC score | 0.008781 (rank : 150) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 899 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8TBC5, Q9BRK7, Q9H9A0 | Gene names | ZNF447 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 447. | |||||
|
BORG1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 62) | NC score | 0.025287 (rank : 118) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O14613, Q9UNS0 | Gene names | CDC42EP2, BORG1, CEP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cdc42 effector protein 2 (Binder of Rho GTPases 1). | |||||
|
CTR9_MOUSE
|
||||||
| θ value | 4.03905 (rank : 63) | NC score | 0.023856 (rank : 124) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q62018, Q3UFF5, Q3UY40, Q66JX4, Q7TPS6, Q8BND9, Q8BRD1, Q8C9W7, Q8C9Y3, Q8CHI1 | Gene names | Ctr9, Kiaa0155, Sh2bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein CTR9 homolog (SH2 domain-binding protein 1) (Tetratricopeptide repeat-containing, SH2-binding phosphoprotein of 150 kDa) (TPR-containing, SH2-binding phosphoprotein of 150 kDa) (p150TSP). | |||||
|
M3K6_HUMAN
|
||||||
| θ value | 4.03905 (rank : 64) | NC score | 0.008159 (rank : 153) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 889 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O95382 | Gene names | MAP3K6, MAPKKK6 | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase 6 (EC 2.7.11.25). | |||||
|
MINK1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 65) | NC score | 0.010407 (rank : 146) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1451 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9JM52, Q921M6, Q9JM92 | Gene names | Mink1, Map4k6, Mink | |||
|
Domain Architecture |

|
|||||
| Description | Misshapen-like kinase 1 (EC 2.7.11.1) (Mitogen-activated protein kinase kinase kinase kinase 6) (MAPK/ERK kinase kinase kinase 6) (MEK kinase kinase 6) (MEKKK 6) (Misshapen/NIK-related kinase) (GCK family kinase MiNK). | |||||
|
MKL1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 66) | NC score | 0.036918 (rank : 104) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 667 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8K4J6 | Gene names | Mkl1, Bsac | |||
|
Domain Architecture |

|
|||||
| Description | MKL/myocardin-like protein 1 (Myocardin-related transcription factor A) (MRTF-A) (Megakaryoblastic leukemia 1 protein homolog) (Basic SAP coiled-coil transcription activator). | |||||
|
MYCL1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 67) | NC score | 0.023764 (rank : 125) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P12524, Q14897, Q5QPL0, Q5QPL1, Q9NUE9 | Gene names | MYCL1, LMYC, MYCL | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
NRG3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 68) | NC score | 0.039082 (rank : 98) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O35181 | Gene names | Nrg3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pro-neuregulin-3, membrane-bound isoform precursor (Pro-NRG3) [Contains: Neuregulin-3 (NRG-3)]. | |||||
|
PELP1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 69) | NC score | 0.068063 (rank : 38) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9DBD5, Q5F2E2, Q6PEM0, Q91YM9 | Gene names | Pelp1, Mnar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor). | |||||
|
R3HD2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 70) | NC score | 0.045265 (rank : 85) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q80TM6, Q80YB1, Q8BLS5, Q9CW50 | Gene names | R3hdm2, Kiaa1002 | |||
|
Domain Architecture |

|
|||||
| Description | R3H domain-containing protein 2. | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 71) | NC score | 0.065484 (rank : 44) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
TIF1B_MOUSE
|
||||||
| θ value | 4.03905 (rank : 72) | NC score | 0.016308 (rank : 134) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q62318, P70391, Q8C283, Q99PN4 | Gene names | Trim28, Krip1, Tif1b | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-beta (TIF1-beta) (Tripartite motif-containing protein 28) (KRAB-A-interacting protein) (KRIP-1). | |||||
|
UBP31_HUMAN
|
||||||
| θ value | 4.03905 (rank : 73) | NC score | 0.023944 (rank : 121) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 293 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q70CQ4, Q6AW97, Q6ZTC0, Q6ZTN2, Q9ULL7 | Gene names | USP31, KIAA1203 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 31 (EC 3.1.2.15) (Ubiquitin thioesterase 31) (Ubiquitin-specific-processing protease 31) (Deubiquitinating enzyme 31). | |||||
|
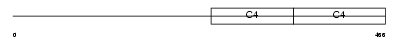
CO4A4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 74) | NC score | 0.040373 (rank : 95) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 440 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P53420 | Gene names | COL4A4 | |||
|
Domain Architecture |
|
|||||
| Description | Collagen alpha-4(IV) chain precursor. | |||||
|
FOXD1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 75) | NC score | 0.010157 (rank : 147) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61345 | Gene names | Foxd1, Fkhl8, Freac4, Hfhbf2 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein D1 (Forkhead-related protein FKHL8) (Forkhead- related transcription factor 4) (FREAC-4) (Brain factor 2) (HFH-BF-2). | |||||
|
HGS_MOUSE
|
||||||
| θ value | 5.27518 (rank : 76) | NC score | 0.026321 (rank : 113) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q99LI8, Q61691, Q8BQW3 | Gene names | Hgs, Hrs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate. | |||||
|
IGHG3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 77) | NC score | 0.009623 (rank : 148) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P01860 | Gene names | IGHG3 | |||
|
Domain Architecture |
|
|||||
| Description | Ig gamma-3 chain C region (Heavy chain disease protein) (HDC). | |||||
|
KCNA5_MOUSE
|
||||||
| θ value | 5.27518 (rank : 78) | NC score | 0.007213 (rank : 155) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61762 | Gene names | Kcna5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 5 (Voltage-gated potassium channel subunit Kv1.5) (KV1-5). | |||||
|
LBXCO_HUMAN
|
||||||
| θ value | 5.27518 (rank : 79) | NC score | 0.043942 (rank : 89) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P84550 | Gene names | LBXCOR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ladybird homeobox corepressor 1. | |||||
|
PODO_HUMAN
|
||||||
| θ value | 5.27518 (rank : 80) | NC score | 0.017811 (rank : 133) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NP85, Q8N6Q5 | Gene names | NPHS2 | |||
|
Domain Architecture |
|
|||||
| Description | Podocin. | |||||
|
PRR13_MOUSE
|
||||||
| θ value | 5.27518 (rank : 81) | NC score | 0.107209 (rank : 8) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9CQJ5, Q3U3U4 | Gene names | Prr13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 13. | |||||
|
PYR1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 82) | NC score | 0.019817 (rank : 130) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P27708, Q6P0Q0 | Gene names | CAD | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CAD protein [Includes: Glutamine-dependent carbamoyl-phosphate synthase (EC 6.3.5.5); Aspartate carbamoyltransferase (EC 2.1.3.2); Dihydroorotase (EC 3.5.2.3)]. | |||||
|
TAF4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 83) | NC score | 0.092641 (rank : 16) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 687 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O00268, Q5TBP6, Q99721, Q9BR40, Q9BX42 | Gene names | TAF4, TAF2C, TAF2C1, TAF4A, TAFII130, TAFII135 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 4 (TBP-associated factor 4) (Transcription initiation factor TFIID 135 kDa subunit) (TAF(II)135) (TAFII-135) (TAFII135) (TAFII-130) (TAFII130). | |||||
|
TGON2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 84) | NC score | 0.038241 (rank : 101) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
UBIP1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 85) | NC score | 0.014315 (rank : 135) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q811S7, Q3UPR3, Q3US11, Q60786, Q8C514 | Gene names | Ubp1, Cp2b, Nf2d9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Upstream-binding protein 1 (Nuclear factor 2d9) (NF2d9). | |||||
|
ARP5_MOUSE
|
||||||
| θ value | 6.88961 (rank : 86) | NC score | 0.013048 (rank : 137) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q80US4, Q8BL26 | Gene names | Actr5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Actin-related protein 5. | |||||
|
CABIN_HUMAN
|
||||||
| θ value | 6.88961 (rank : 87) | NC score | 0.055162 (rank : 66) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 490 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9Y6J0, Q9Y460 | Gene names | CABIN1, KIAA0330 | |||
|
Domain Architecture |

|
|||||
| Description | Calcineurin-binding protein Cabin 1 (Calcineurin inhibitor) (CAIN). | |||||
|
DAB2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 88) | NC score | 0.024922 (rank : 119) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P98082, Q13598, Q9BTY0, Q9UK04 | Gene names | DAB2, DOC2 | |||
|
Domain Architecture |

|
|||||
| Description | Disabled homolog 2 (Differentially expressed protein 2) (DOC-2). | |||||
|
GAK_HUMAN
|
||||||
| θ value | 6.88961 (rank : 89) | NC score | 0.012600 (rank : 138) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1028 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O14976, Q9BVY6 | Gene names | GAK | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin G-associated kinase (EC 2.7.11.1). | |||||
|
HXB1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 90) | NC score | 0.008188 (rank : 152) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P14653 | Gene names | HOXB1, HOX2I | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-B1 (Hox-2I). | |||||
|
LEUK_HUMAN
|
||||||
| θ value | 6.88961 (rank : 91) | NC score | 0.039832 (rank : 96) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P16150 | Gene names | SPN, CD43 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leukosialin precursor (Leukocyte sialoglycoprotein) (Sialophorin) (Galactoglycoprotein) (GALGP) (CD43 antigen). | |||||
|
LSD1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 92) | NC score | 0.026992 (rank : 110) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O60341, Q5TH94, Q5TH95, Q86VT7, Q8IXK4, Q8NDP6, Q8TAZ3, Q96AW4 | Gene names | AOF2, KIAA0601, LSD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lysine-specific histone demethylase 1 (EC 1.-.-.-) (Flavin-containing amine oxidase domain-containing protein 2) (BRAF35-HDAC complex protein BHC110). | |||||
|
MARCS_HUMAN
|
||||||
| θ value | 6.88961 (rank : 93) | NC score | 0.079849 (rank : 22) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P29966, Q2LA83, Q5TDB7 | Gene names | MARCKS, MACS, PRKCSL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS) (Protein kinase C substrate, 80 kDa protein, light chain) (PKCSL) (80K-L protein). | |||||
|
MINK1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 94) | NC score | 0.012020 (rank : 139) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8N4C8, Q9P1X1, Q9P2R8 | Gene names | MINK1, MAP4K6, MINK | |||
|
Domain Architecture |

|
|||||
| Description | Misshapen-like kinase 1 (EC 2.7.11.1) (Mitogen-activated protein kinase kinase kinase kinase 6) (MAPK/ERK kinase kinase kinase 6) (MEK kinase kinase 6) (MEKKK 6) (Misshapen/NIK-related kinase) (GCK family kinase MiNK). | |||||
|
PCLO_MOUSE
|
||||||
| θ value | 6.88961 (rank : 95) | NC score | 0.055545 (rank : 63) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
PHLPP_HUMAN
|
||||||
| θ value | 6.88961 (rank : 96) | NC score | 0.023656 (rank : 126) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 604 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O60346, Q641Q7, Q6P4C4, Q6PJI6, Q86TN6, Q96FK2, Q9NUY1 | Gene names | PHLPP, KIAA0606, PLEKHE1, SCOP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH domain leucine-rich repeat-containing protein phosphatase (EC 3.1.3.16) (PH domain leucine-rich repeat protein phosphatase) (Pleckstrin homology domain-containing family E protein 1) (Suprachiasmatic nucleus circadian oscillatory protein) (hSCOP). | |||||
|
PSL1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 97) | NC score | 0.044614 (rank : 87) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8TCT7, O60365, Q567S3, Q8IUH9, Q9BUY6, Q9H3M4, Q9NPN2, Q9P1Z6 | Gene names | SPPL2B, KIAA1532, PSL1 | |||
|
Domain Architecture |

|
|||||
| Description | Signal peptide peptidase-like 2B (EC 3.4.23.-) (Protein SPP-like 2B) (Protein SPPL2b) (Intramembrane protease 4) (IMP4) (Presenilin-like protein 1). | |||||
|
SYN1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 98) | NC score | 0.061356 (rank : 49) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P17600, O75825, Q5H9A9 | Gene names | SYN1 | |||
|
Domain Architecture |

|
|||||
| Description | Synapsin-1 (Synapsin I) (Brain protein 4.1). | |||||
|
UB7I1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 99) | NC score | 0.025530 (rank : 117) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P58283, Q3U493, Q3UE56, Q3UGM3, Q68FN0, Q6P1H8, Q6PWY5, Q8BN27, Q8C1U3 | Gene names | Ubce7ip1, Triad3, Uip83, Zin | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin ligase TRIAD3 (EC 6.3.2.-) (Ubiquitin-conjugating enzyme 7-interacting protein 1) (UbcM4-interacting protein 83) (Triad domain- containing protein 3). | |||||
|
CF155_HUMAN
|
||||||
| θ value | 8.99809 (rank : 100) | NC score | 0.038447 (rank : 99) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H8W2, Q5TG13 | Gene names | C6orf155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf155. | |||||
|
CHIT1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 101) | NC score | 0.008611 (rank : 151) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q13231, Q3ZAR1, Q6ISC2, Q9H3V8 | Gene names | CHIT1 | |||
|
Domain Architecture |

|
|||||
| Description | Chitotriosidase-1 precursor (EC 3.2.1.14) (Chitinase-1). | |||||
|
CO2A1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 102) | NC score | 0.026599 (rank : 112) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 483 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P28481 | Gene names | Col2a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(II) chain precursor [Contains: Chondrocalcin]. | |||||
|
CRSP7_HUMAN
|
||||||
| θ value | 8.99809 (rank : 103) | NC score | 0.031479 (rank : 106) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O95402 | Gene names | CRSP7, ARC70 | |||
|
Domain Architecture |

|
|||||
| Description | CRSP complex subunit 7 (Cofactor required for Sp1 transcriptional activation subunit 7) (Transcriptional coactivator CRSP70) (Activator- recruited cofactor 70 kDa component) (ARC70). | |||||
|
DTX1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 104) | NC score | 0.027213 (rank : 109) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q61010, Q3TER5, Q8C2H2, Q9ER09 | Gene names | Dtx1 | |||
|
Domain Architecture |
|
|||||
| Description | Protein deltex-1 (Deltex-1) (Deltex1) (mDTX1) (FXI-T1). | |||||
|
E2AK1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 105) | NC score | 0.010918 (rank : 143) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 833 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9BQI3, Q8NBW3, Q9HC02, Q9NYE0, Q9P0V6, Q9P1J5, Q9P2H8, Q9UHG4 | Gene names | EIF2AK1, HRI, KIAA1369 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2-alpha kinase 1 (EC 2.7.11.1) (Heme-regulated eukaryotic initiation factor eIF-2-alpha kinase) (Heme-regulated inhibitor) (Heme-controlled repressor) (HCR) (Hemin-sensitive initiation factor 2-alpha kinase). | |||||
|
E2AK1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 106) | NC score | 0.010611 (rank : 145) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Z2R9, Q8C024, Q8K123, Q9CTP5, Q9D601 | Gene names | Eif2ak1, Hri | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2-alpha kinase 1 (EC 2.7.11.1) (Heme-regulated eukaryotic initiation factor eIF-2-alpha kinase) (Heme-regulated inhibitor) (Heme-controlled repressor) (HCR) (Hemin-sensitive initiation factor 2-alpha kinase). | |||||
|
GAK5_HUMAN
|
||||||
| θ value | 8.99809 (rank : 107) | NC score | 0.011232 (rank : 142) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 429 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P62684 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19p13.11 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K113 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
LTK_MOUSE
|
||||||
| θ value | 8.99809 (rank : 108) | NC score | 0.003040 (rank : 158) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 770 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P08923 | Gene names | Ltk | |||
|
Domain Architecture |

|
|||||
| Description | Leukocyte tyrosine kinase receptor precursor (EC 2.7.10.1). | |||||
|
MAGI2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 109) | NC score | 0.011373 (rank : 140) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 373 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q86UL8, O60434, O60510, Q86UI7, Q9NP44, Q9UDQ5, Q9UDU1 | Gene names | MAGI2, ACVRINP1, AIP1, KIAA0705 | |||
|
Domain Architecture |

|
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 2 (Membrane-associated guanylate kinase inverted 2) (MAGI-2) (Atrophin-1-interacting protein 1) (Atrophin-1-interacting protein A). | |||||
|
TCFL5_HUMAN
|
||||||
| θ value | 8.99809 (rank : 110) | NC score | 0.023890 (rank : 122) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UL49, O94771, Q9BYW0 | Gene names | TCFL5, CHA, E2BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor-like 5 protein (Cha transcription factor) (HPV-16 E2-binding protein 1) (E2BP-1). | |||||
|
ATN1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 111) | NC score | 0.060556 (rank : 51) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
BAT2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 112) | NC score | 0.065016 (rank : 45) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
BAT3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 113) | NC score | 0.051247 (rank : 80) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 518 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9Z1R2, Q8SNA3 | Gene names | Bat3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT3 (HLA-B-associated transcript 3). | |||||
|
BSN_MOUSE
|
||||||
| θ value | θ > 10 (rank : 114) | NC score | 0.054864 (rank : 67) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
CIC_HUMAN
|
||||||
| θ value | θ > 10 (rank : 115) | NC score | 0.054206 (rank : 70) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 992 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q96RK0, Q7LGI1, Q9UEG5, Q9Y6T1 | Gene names | CIC, KIAA0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
CN032_HUMAN
|
||||||
| θ value | θ > 10 (rank : 116) | NC score | 0.068498 (rank : 36) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8NDC0, Q96BG5 | Gene names | C14orf32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf32. | |||||
|
CN032_MOUSE
|
||||||
| θ value | θ > 10 (rank : 117) | NC score | 0.080939 (rank : 21) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8BH93, Q3TF12, Q3TGL0, Q8CC90 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf32 homolog. | |||||
|
CN155_HUMAN
|
||||||
| θ value | θ > 10 (rank : 118) | NC score | 0.056506 (rank : 59) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q5H9T9, Q5H9U7, Q86YI2, Q9H0J3 | Gene names | C14orf155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf155. | |||||
|
CPSF6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 119) | NC score | 0.065005 (rank : 46) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q16630, Q53ES1, Q9BSJ7, Q9BW18 | Gene names | CPSF6, CFIM68 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage and polyadenylation specificity factor 6 (Cleavage and polyadenylation specificity factor 68 kDa subunit) (CPSF 68 kDa subunit) (Pre-mRNA cleavage factor Im 68 kDa subunit) (Protein HPBRII- 4/7). | |||||
|
CPSF6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 120) | NC score | 0.066289 (rank : 43) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q6NVF9, Q8BX86, Q8BXI8 | Gene names | Cpsf6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage and polyadenylation specificity factor 6. | |||||
|
CS016_HUMAN
|
||||||
| θ value | θ > 10 (rank : 121) | NC score | 0.052061 (rank : 75) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8ND99 | Gene names | C19orf16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C19orf16. | |||||
|
ENAH_MOUSE
|
||||||
| θ value | θ > 10 (rank : 122) | NC score | 0.054212 (rank : 69) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q03173, P70430, P70431, P70432, P70433, Q5D053 | Gene names | Enah, Mena, Ndpp1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein enabled homolog (NPC-derived proline-rich protein 1) (NDPP-1). | |||||
|
FMN2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 123) | NC score | 0.051833 (rank : 78) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 484 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9NZ56, Q59GF6, Q5VU37, Q9NZ55 | Gene names | FMN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Formin-2. | |||||
|
GGN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 124) | NC score | 0.070931 (rank : 31) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q86UU5, Q7RTU6, Q86UU4, Q8NAA1 | Gene names | GGN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin. | |||||
|
GGN_MOUSE
|
||||||
| θ value | θ > 10 (rank : 125) | NC score | 0.054439 (rank : 68) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q80WJ1, Q5EBP4, Q80WI9, Q80WJ0 | Gene names | Ggn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin. | |||||
|
GSCR1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 126) | NC score | 0.067300 (rank : 39) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9NZM4 | Gene names | GLTSCR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glioma tumor suppressor candidate region gene 1 protein. | |||||
|
MBD6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 127) | NC score | 0.068404 (rank : 37) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q96DN6, Q8N3M0, Q8NA81, Q96Q00 | Gene names | MBD6, KIAA1887 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 6 (Methyl-CpG-binding protein MBD6). | |||||
|
MILK1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 128) | NC score | 0.051838 (rank : 77) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 788 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8BGT6, Q8BJ60 | Gene names | Mirab13, D15Mit260, Kiaa1668 | |||
|
Domain Architecture |

|
|||||
| Description | Molecule interacting with Rab13 (MIRab13) (MICAL-like protein 1). | |||||
|
MUC1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 129) | NC score | 0.061199 (rank : 50) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q02496 | Gene names | Muc1, Muc-1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (Polymorphic epithelial mucin) (PEMT) (Episialin). | |||||
|
MUC7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 130) | NC score | 0.071563 (rank : 29) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 538 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8TAX7 | Gene names | MUC7, MG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-7 precursor (MUC-7) (Salivary mucin-7) (Apo-MG2). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | θ > 10 (rank : 131) | NC score | 0.054192 (rank : 71) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
PELP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 132) | NC score | 0.054122 (rank : 72) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 596 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8IZL8, O15450, Q5EGN3, Q6NTE6, Q96FT1, Q9BU60 | Gene names | PELP1, HMX3, MNAR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor) (Transcription factor HMX3). | |||||
|
PRB3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 133) | NC score | 0.105187 (rank : 9) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q04118, Q15188, Q7M4M9 | Gene names | PRB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 3 precursor (Parotid salivary glycoprotein G1) (Proline-rich protein G1). | |||||
|
PRB4S_HUMAN
|
||||||
| θ value | θ > 10 (rank : 134) | NC score | 0.096888 (rank : 14) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P10163, P02813 | Gene names | PRB4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 4 allele S precursor (Salivary proline-rich protein Po) (Parotid o protein) [Contains: Protein N1; Glycosylated protein A]. | |||||
|
PRP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 135) | NC score | 0.100724 (rank : 10) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P04280, Q08805, Q15186, Q15187, Q15214, Q15215, Q16038 | Gene names | PRB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 1 precursor (Salivary proline-rich protein) [Contains: Basic peptide IB-6; Peptide P-H]. | |||||
|
PRP2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 136) | NC score | 0.092706 (rank : 15) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P05142 | Gene names | Prh1, Prp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein MP-2 precursor. | |||||
|
PRP5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 137) | NC score | 0.053016 (rank : 73) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P04281 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic proline-rich peptide IB-1. | |||||
|
PRPE_HUMAN
|
||||||
| θ value | θ > 10 (rank : 138) | NC score | 0.062741 (rank : 48) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P02811 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic proline-rich peptide P-E (IB-9). | |||||
|
PRR12_HUMAN
|
||||||
| θ value | θ > 10 (rank : 139) | NC score | 0.072141 (rank : 27) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9ULL5, Q8N4J6 | Gene names | PRR12, KIAA1205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 12. | |||||
|
PRR13_HUMAN
|
||||||
| θ value | θ > 10 (rank : 140) | NC score | 0.078889 (rank : 23) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9NZ81, Q6FIG7, Q6MZP8, Q6NXQ6, Q6PKF9 | Gene names | PRR13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 13. | |||||
|
PRRT3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 141) | NC score | 0.081655 (rank : 20) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q6PE13 | Gene names | Prrt3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
SF01_HUMAN
|
||||||
| θ value | θ > 10 (rank : 142) | NC score | 0.055205 (rank : 65) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q15637, Q14818, Q14819, Q15913, Q8IY00, Q92744, Q92745, Q969H7, Q9BW01, Q9UEI0 | Gene names | SF1, ZFM1, ZNF162 | |||
|
Domain Architecture |
|
|||||
| Description | Splicing factor 1 (Zinc finger protein 162) (Transcription factor ZFM1) (Zinc finger gene in MEN1 locus) (Mammalian branch point-binding protein mBBP) (BBP). | |||||
|
SF3A2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 143) | NC score | 0.070671 (rank : 32) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q15428, O75245 | Gene names | SF3A2, SAP62 | |||
|
Domain Architecture |
|
|||||
| Description | Splicing factor 3A subunit 2 (Spliceosome-associated protein 62) (SAP 62) (SF3a66). | |||||
|
SF3A2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 144) | NC score | 0.068965 (rank : 35) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q62203 | Gene names | Sf3a2, Sap62 | |||
|
Domain Architecture |
|
|||||
| Description | Splicing factor 3A subunit 2 (Spliceosome-associated protein 62) (SAP 62) (SF3a66). | |||||
|
SFPQ_HUMAN
|
||||||
| θ value | θ > 10 (rank : 145) | NC score | 0.052338 (rank : 74) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 986 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P23246, P30808 | Gene names | SFPQ, PSF | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, proline- and glutamine-rich (Polypyrimidine tract- binding protein-associated-splicing factor) (PTB-associated-splicing factor) (PSF) (DNA-binding p52/p100 complex, 100 kDa subunit) (100 kDa DNA-pairing protein) (hPOMp100). | |||||
|
SMR3A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 146) | NC score | 0.056365 (rank : 60) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q99954 | Gene names | SMR3A, PBI, PROL5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Submaxillary gland androgen-regulated protein 3 homolog A precursor (Proline-rich protein 5) (Proline-rich protein PBI). | |||||
|
SRRM1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 147) | NC score | 0.050259 (rank : 82) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
SRRM1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 148) | NC score | 0.055333 (rank : 64) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 149) | NC score | 0.056161 (rank : 62) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
WASIP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 150) | NC score | 0.072954 (rank : 26) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O43516, Q15220, Q6MZU9, Q9BU37, Q9UNP1 | Gene names | WASPIP, WIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein) (PRPL-2 protein). | |||||
|
WASIP_MOUSE
|
||||||
| θ value | θ > 10 (rank : 151) | NC score | 0.073296 (rank : 25) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8K1I7, Q3U0U8 | Gene names | Waspip, Wip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein). | |||||
|
WASL_MOUSE
|
||||||
| θ value | θ > 10 (rank : 152) | NC score | 0.051675 (rank : 79) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 339 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q91YD9 | Gene names | Wasl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neural Wiskott-Aldrich syndrome protein (N-WASP). | |||||
|
WBP11_HUMAN
|
||||||
| θ value | θ > 10 (rank : 153) | NC score | 0.069771 (rank : 33) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9Y2W2, Q96AY8 | Gene names | WBP11, NPWBP, SNP70 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 11 (WBP-11) (SH3 domain-binding protein SNP70) (Npw38-binding protein) (NpwBP). | |||||
|
WBP11_MOUSE
|
||||||
| θ value | θ > 10 (rank : 154) | NC score | 0.064661 (rank : 47) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q923D5, O88539, Q3UMD8, Q8VDI0 | Gene names | Wbp11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 11 (WBP-11). | |||||
|
WIRE_HUMAN
|
||||||
| θ value | θ > 10 (rank : 155) | NC score | 0.059458 (rank : 54) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8TF74, Q658J8, Q71RE1, Q8TE44 | Gene names | WIRE, WICH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WIP-related protein (WASP-interacting protein-related protein) (WIP- and CR16-homologous protein). | |||||
|
WIRE_MOUSE
|
||||||
| θ value | θ > 10 (rank : 156) | NC score | 0.059125 (rank : 55) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q6PEV3 | Gene names | Wire | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WIP-related protein (WASP-interacting protein-related protein). | |||||
|
YLPM1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 157) | NC score | 0.078786 (rank : 24) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
ZN207_HUMAN
|
||||||
| θ value | θ > 10 (rank : 158) | NC score | 0.058404 (rank : 58) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 326 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O43670, Q96HW5, Q9BUQ7 | Gene names | ZNF207 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 207. | |||||
|
IER5_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 7.12574e-114 (rank : 1) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 110 | |
| SwissProt Accessions | O89113, Q8CGI0 | Gene names | Ier5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Immediate early response gene 5 protein. | |||||
|
IER5_HUMAN
|
||||||
| NC score | 0.930859 (rank : 2) | θ value | 8.51556e-75 (rank : 2) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q5VY09, Q8WY68, Q9NY49, Q9NZP9 | Gene names | IER5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Immediate early response gene 5 protein. | |||||
|
IER2_HUMAN
|
||||||
| NC score | 0.611754 (rank : 3) | θ value | 2.0648e-12 (rank : 3) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BTL4, Q03827, Q2TAZ2 | Gene names | IER2, ETR101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Immediate early response gene 2 protein (Protein ETR101). | |||||
|
IER2_MOUSE
|
||||||
| NC score | 0.577807 (rank : 4) | θ value | 5.62301e-10 (rank : 4) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P17950, Q64251, Q8BME8, Q99M25 | Gene names | Ier2 | |||
|
Domain Architecture |

|
|||||
| Description | Immediate early response gene 2 protein (T-lymphocyte-activated protein) (Cycloheximide-induced gene protein) (CHX1) (Growth factor- inducible immediate early protein) (Proline-rich-induced protein 92) (Pip92). | |||||
|
PRRT3_HUMAN
|
||||||
| NC score | 0.123134 (rank : 5) | θ value | 0.000786445 (rank : 5) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q5FWE3, Q49AD0, Q6UXY6, Q8NBC9 | Gene names | PRRT3, UNQ5823/PRO19642 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
SF3B1_MOUSE
|
||||||
| NC score | 0.107710 (rank : 6) | θ value | 0.813845 (rank : 27) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q99NB9, Q9CSK5 | Gene names | Sf3b1, Sap155 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3B subunit 1 (Spliceosome-associated protein 155) (SAP 155) (SF3b155) (Pre-mRNA-splicing factor SF3b 155 kDa subunit). | |||||
|
SF3B1_HUMAN
|
||||||
| NC score | 0.107371 (rank : 7) | θ value | 0.813845 (rank : 26) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O75533 | Gene names | SF3B1, SAP155 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3B subunit 1 (Spliceosome-associated protein 155) (SAP 155) (SF3b155) (Pre-mRNA-splicing factor SF3b 155 kDa subunit). | |||||
|
PRR13_MOUSE
|
||||||
| NC score | 0.107209 (rank : 8) | θ value | 5.27518 (rank : 81) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9CQJ5, Q3U3U4 | Gene names | Prr13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 13. | |||||
|
PRB3_HUMAN
|
||||||
| NC score | 0.105187 (rank : 9) | θ value | θ > 10 (rank : 133) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q04118, Q15188, Q7M4M9 | Gene names | PRB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 3 precursor (Parotid salivary glycoprotein G1) (Proline-rich protein G1). | |||||
|
PRP1_HUMAN
|
||||||
| NC score | 0.100724 (rank : 10) | θ value | θ > 10 (rank : 135) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P04280, Q08805, Q15186, Q15187, Q15214, Q15215, Q16038 | Gene names | PRB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 1 precursor (Salivary proline-rich protein) [Contains: Basic peptide IB-6; Peptide P-H]. | |||||
|
BASP_MOUSE
|
||||||
| NC score | 0.100363 (rank : 11) | θ value | 1.38821 (rank : 34) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
MARCS_MOUSE
|
||||||
| NC score | 0.098326 (rank : 12) | θ value | 0.279714 (rank : 16) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P26645 | Gene names | Marcks, Macs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS). | |||||
|
OGFR_HUMAN
|
||||||
| NC score | 0.097047 (rank : 13) | θ value | 0.813845 (rank : 24) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
PRB4S_HUMAN
|
||||||
| NC score | 0.096888 (rank : 14) | θ value | θ > 10 (rank : 134) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P10163, P02813 | Gene names | PRB4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 4 allele S precursor (Salivary proline-rich protein Po) (Parotid o protein) [Contains: Protein N1; Glycosylated protein A]. | |||||
|
PRP2_MOUSE
|
||||||
| NC score | 0.092706 (rank : 15) | θ value | θ > 10 (rank : 136) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P05142 | Gene names | Prh1, Prp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein MP-2 precursor. | |||||
|
TAF4_HUMAN
|
||||||
| NC score | 0.092641 (rank : 16) | θ value | 5.27518 (rank : 83) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 687 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O00268, Q5TBP6, Q99721, Q9BR40, Q9BX42 | Gene names | TAF4, TAF2C, TAF2C1, TAF4A, TAFII130, TAFII135 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 4 (TBP-associated factor 4) (Transcription initiation factor TFIID 135 kDa subunit) (TAF(II)135) (TAFII-135) (TAFII135) (TAFII-130) (TAFII130). | |||||
|
YLPM1_HUMAN
|
||||||
| NC score | 0.089704 (rank : 17) | θ value | 0.365318 (rank : 21) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P49750, P49752, Q96I64, Q9P1V7 | Gene names | YLPM1, C14orf170, ZAP3 | |||
|
Domain Architecture |

|
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3) (ZAP113). | |||||
|
BASP_HUMAN
|
||||||
| NC score | 0.089157 (rank : 18) | θ value | 3.0926 (rank : 52) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
MUC1_HUMAN
|
||||||
| NC score | 0.087546 (rank : 19) | θ value | 0.00665767 (rank : 6) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
PRRT3_MOUSE
|
||||||
| NC score | 0.081655 (rank : 20) | θ value | θ > 10 (rank : 141) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q6PE13 | Gene names | Prrt3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
CN032_MOUSE
|
||||||
| NC score | 0.080939 (rank : 21) | θ value | θ > 10 (rank : 117) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8BH93, Q3TF12, Q3TGL0, Q8CC90 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf32 homolog. | |||||
|
MARCS_HUMAN
|
||||||
| NC score | 0.079849 (rank : 22) | θ value | 6.88961 (rank : 93) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P29966, Q2LA83, Q5TDB7 | Gene names | MARCKS, MACS, PRKCSL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS) (Protein kinase C substrate, 80 kDa protein, light chain) (PKCSL) (80K-L protein). | |||||
|
PRR13_HUMAN
|
||||||
| NC score | 0.078889 (rank : 23) | θ value | θ > 10 (rank : 140) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9NZ81, Q6FIG7, Q6MZP8, Q6NXQ6, Q6PKF9 | Gene names | PRR13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 13. | |||||
|
YLPM1_MOUSE
|
||||||
| NC score | 0.078786 (rank : 24) | θ value | θ > 10 (rank : 157) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
WASIP_MOUSE
|
||||||
| NC score | 0.073296 (rank : 25) | θ value | θ > 10 (rank : 151) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8K1I7, Q3U0U8 | Gene names | Waspip, Wip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein). | |||||
|
WASIP_HUMAN
|
||||||
| NC score | 0.072954 (rank : 26) | θ value | θ > 10 (rank : 150) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O43516, Q15220, Q6MZU9, Q9BU37, Q9UNP1 | Gene names | WASPIP, WIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein) (PRPL-2 protein). | |||||
|
PRR12_HUMAN
|
||||||
| NC score | 0.072141 (rank : 27) | θ value | θ > 10 (rank : 139) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9ULL5, Q8N4J6 | Gene names | PRR12, KIAA1205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 12. | |||||
|
RGS3_MOUSE
|
||||||
| NC score | 0.071721 (rank : 28) | θ value | 0.813845 (rank : 25) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
MUC7_HUMAN
|
||||||
| NC score | 0.071563 (rank : 29) | θ value | θ > 10 (rank : 130) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 538 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8TAX7 | Gene names | MUC7, MG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-7 precursor (MUC-7) (Salivary mucin-7) (Apo-MG2). | |||||
|
TM16H_HUMAN
|
||||||
| NC score | 0.071330 (rank : 30) | θ value | 0.163984 (rank : 10) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9HCE9 | Gene names | TMEM16H, KIAA1623 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 16H. | |||||
|
GGN_HUMAN
|
||||||
| NC score | 0.070931 (rank : 31) | θ value | θ > 10 (rank : 124) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q86UU5, Q7RTU6, Q86UU4, Q8NAA1 | Gene names | GGN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin. | |||||
|
SF3A2_HUMAN
|
||||||
| NC score | 0.070671 (rank : 32) | θ value | θ > 10 (rank : 143) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q15428, O75245 | Gene names | SF3A2, SAP62 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3A subunit 2 (Spliceosome-associated protein 62) (SAP 62) (SF3a66). | |||||
|
WBP11_HUMAN
|
||||||
| NC score | 0.069771 (rank : 33) | θ value | θ > 10 (rank : 153) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9Y2W2, Q96AY8 | Gene names | WBP11, NPWBP, SNP70 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 11 (WBP-11) (SH3 domain-binding protein SNP70) (Npw38-binding protein) (NpwBP). | |||||
|
MAGE1_HUMAN
|
||||||
| NC score | 0.069072 (rank : 34) | θ value | 0.0330416 (rank : 7) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
SF3A2_MOUSE
|
||||||
| NC score | 0.068965 (rank : 35) | θ value | θ > 10 (rank : 144) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q62203 | Gene names | Sf3a2, Sap62 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3A subunit 2 (Spliceosome-associated protein 62) (SAP 62) (SF3a66). | |||||
|
CN032_HUMAN
|
||||||
| NC score | 0.068498 (rank : 36) | θ value | θ > 10 (rank : 116) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8NDC0, Q96BG5 | Gene names | C14orf32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf32. | |||||
|
MBD6_HUMAN
|
||||||
| NC score | 0.068404 (rank : 37) | θ value | θ > 10 (rank : 127) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q96DN6, Q8N3M0, Q8NA81, Q96Q00 | Gene names | MBD6, KIAA1887 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 6 (Methyl-CpG-binding protein MBD6). | |||||
|
PELP1_MOUSE
|
||||||
| NC score | 0.068063 (rank : 38) | θ value | 4.03905 (rank : 69) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9DBD5, Q5F2E2, Q6PEM0, Q91YM9 | Gene names | Pelp1, Mnar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor). | |||||
|
GSCR1_HUMAN
|
||||||
| NC score | 0.067300 (rank : 39) | θ value | θ > 10 (rank : 126) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9NZM4 | Gene names | GLTSCR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glioma tumor suppressor candidate region gene 1 protein. | |||||
|
OBF1_HUMAN
|
||||||
| NC score | 0.067069 (rank : 40) | θ value | 2.36792 (rank : 50) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q16633, Q14983 | Gene names | POU2AF1, OBF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | POU domain class 2-associating factor 1 (B-cell-specific coactivator OBF-1) (OCT-binding factor 1) (BOB-1) (OCA-B). | |||||
|
GNAS1_MOUSE
|
||||||
| NC score | 0.066924 (rank : 41) | θ value | 0.0736092 (rank : 8) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 651 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q6R0H7, Q6R0H4, Q6R0H5, Q6R2J5, Q9JJX0, Q9Z1N8 | Gene names | Gnas, Gnas1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Guanine nucleotide-binding protein G(s) subunit alpha isoforms XLas (Adenylate cyclase-stimulating G alpha protein) (Extra large alphas protein) (XLalphas). | |||||
|
FXL19_MOUSE
|
||||||
| NC score | 0.066455 (rank : 42) | θ value | 0.279714 (rank : 13) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q6PB97, Q7TSH0, Q8BIB4 | Gene names | Fbxl19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 19 (F-box and leucine-rich repeat protein 19). | |||||
|
CPSF6_MOUSE
|
||||||
| NC score | 0.066289 (rank : 43) | θ value | θ > 10 (rank : 120) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q6NVF9, Q8BX86, Q8BXI8 | Gene names | Cpsf6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage and polyadenylation specificity factor 6. | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.065484 (rank : 44) | θ value | 4.03905 (rank : 71) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
BAT2_HUMAN
|
||||||
| NC score | 0.065016 (rank : 45) | θ value | θ > 10 (rank : 112) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
CPSF6_HUMAN
|
||||||
| NC score | 0.065005 (rank : 46) | θ value | θ > 10 (rank : 119) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q16630, Q53ES1, Q9BSJ7, Q9BW18 | Gene names | CPSF6, CFIM68 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage and polyadenylation specificity factor 6 (Cleavage and polyadenylation specificity factor 68 kDa subunit) (CPSF 68 kDa subunit) (Pre-mRNA cleavage factor Im 68 kDa subunit) (Protein HPBRII- 4/7). | |||||
|
WBP11_MOUSE
|
||||||
| NC score | 0.064661 (rank : 47) | θ value | θ > 10 (rank : 154) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q923D5, O88539, Q3UMD8, Q8VDI0 | Gene names | Wbp11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-binding protein 11 (WBP-11). | |||||
|
PRPE_HUMAN
|
||||||
| NC score | 0.062741 (rank : 48) | θ value | θ > 10 (rank : 138) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P02811 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic proline-rich peptide P-E (IB-9). | |||||
|
SYN1_HUMAN
|
||||||
| NC score | 0.061356 (rank : 49) | θ value | 6.88961 (rank : 98) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P17600, O75825, Q5H9A9 | Gene names | SYN1 | |||
|
Domain Architecture |

|
|||||
| Description | Synapsin-1 (Synapsin I) (Brain protein 4.1). | |||||
|
MUC1_MOUSE
|
||||||
| NC score | 0.061199 (rank : 50) | θ value | θ > 10 (rank : 129) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q02496 | Gene names | Muc1, Muc-1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (Polymorphic epithelial mucin) (PEMT) (Episialin). | |||||
|
ATN1_HUMAN
|
||||||
| NC score | 0.060556 (rank : 51) | θ value | θ > 10 (rank : 111) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
FXL19_HUMAN
|
||||||
| NC score | 0.060474 (rank : 52) | θ value | 2.36792 (rank : 43) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6PCT2, Q8N789, Q9NT14 | Gene names | FBXL19, FBL19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box/LRR-repeat protein 19 (F-box and leucine-rich repeat protein 19). | |||||
|
ZN385_HUMAN
|
||||||
| NC score | 0.060390 (rank : 53) | θ value | 0.47712 (rank : 22) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 614 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q96PM9, Q5VH53, Q9H7R6, Q9UFU3 | Gene names | ZNF385, HZF, RZF | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 385 (Hematopoietic zinc finger protein) (Retinal zinc finger protein). | |||||
|
WIRE_HUMAN
|
||||||
| NC score | 0.059458 (rank : 54) | θ value | θ > 10 (rank : 155) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8TF74, Q658J8, Q71RE1, Q8TE44 | Gene names | WIRE, WICH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WIP-related protein (WASP-interacting protein-related protein) (WIP- and CR16-homologous protein). | |||||
|
WIRE_MOUSE
|
||||||
| NC score | 0.059125 (rank : 55) | θ value | θ > 10 (rank : 156) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q6PEV3 | Gene names | Wire | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WIP-related protein (WASP-interacting protein-related protein). | |||||
|
OBF1_MOUSE
|
||||||
| NC score | 0.058887 (rank : 56) | θ value | 2.36792 (rank : 51) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q64693 | Gene names | Pou2af1, Obf-1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | POU domain class 2-associating factor 1 (B-cell-specific coactivator OBF-1) (OCT-binding factor 1) (BOB-1) (BOB1) (OCA-B). | |||||
|
IASPP_MOUSE
|
||||||
| NC score | 0.058762 (rank : 57) | θ value | 0.279714 (rank : 14) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1024 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q5I1X5 | Gene names | Ppp1r13l, Nkip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RelA-associated inhibitor (Inhibitor of ASPP protein) (Protein iASPP) (PPP1R13B-like protein) (NFkB-interacting protein 1). | |||||
|
ZN207_HUMAN
|
||||||
| NC score | 0.058404 (rank : 58) | θ value | θ > 10 (rank : 158) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 326 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O43670, Q96HW5, Q9BUQ7 | Gene names | ZNF207 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 207. | |||||
|
CN155_HUMAN
|
||||||
| NC score | 0.056506 (rank : 59) | θ value | θ > 10 (rank : 118) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q5H9T9, Q5H9U7, Q86YI2, Q9H0J3 | Gene names | C14orf155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf155. | |||||
|
SMR3A_HUMAN
|
||||||
| NC score | 0.056365 (rank : 60) | θ value | θ > 10 (rank : 146) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q99954 | Gene names | SMR3A, PBI, PROL5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Submaxillary gland androgen-regulated protein 3 homolog A precursor (Proline-rich protein 5) (Proline-rich protein PBI). | |||||
|
RIMB1_HUMAN
|
||||||
| NC score | 0.056294 (rank : 61) | θ value | 0.365318 (rank : 19) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1479 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | O95153, O75111, Q8N5W3 | Gene names | BZRAP1, KIAA0612, RBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Peripheral-type benzodiazepine receptor-associated protein 1 (PRAX-1) (Peripheral benzodiazepine receptor-interacting protein) (PBR-IP) (RIM-binding protein 1) (RIM-BP1). | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.056161 (rank : 62) | θ value | θ > 10 (rank : 149) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
PCLO_MOUSE
|
||||||
| NC score | 0.055545 (rank : 63) | θ value | 6.88961 (rank : 95) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
SRRM1_MOUSE
|
||||||
| NC score | 0.055333 (rank : 64) | θ value | θ > 10 (rank : 148) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
SF01_HUMAN
|
||||||
| NC score | 0.055205 (rank : 65) | θ value | θ > 10 (rank : 142) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q15637, Q14818, Q14819, Q15913, Q8IY00, Q92744, Q92745, Q969H7, Q9BW01, Q9UEI0 | Gene names | SF1, ZFM1, ZNF162 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 1 (Zinc finger protein 162) (Transcription factor ZFM1) (Zinc finger gene in MEN1 locus) (Mammalian branch point-binding protein mBBP) (BBP). | |||||
|
CABIN_HUMAN
|
||||||
| NC score | 0.055162 (rank : 66) | θ value | 6.88961 (rank : 87) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 490 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9Y6J0, Q9Y460 | Gene names | CABIN1, KIAA0330 | |||
|
Domain Architecture |

|
|||||
| Description | Calcineurin-binding protein Cabin 1 (Calcineurin inhibitor) (CAIN). | |||||
|
BSN_MOUSE
|
||||||
| NC score | 0.054864 (rank : 67) | θ value | θ > 10 (rank : 114) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
GGN_MOUSE
|
||||||
| NC score | 0.054439 (rank : 68) | θ value | θ > 10 (rank : 125) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q80WJ1, Q5EBP4, Q80WI9, Q80WJ0 | Gene names | Ggn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin. | |||||
|
ENAH_MOUSE
|
||||||
| NC score | 0.054212 (rank : 69) | θ value | θ > 10 (rank : 122) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q03173, P70430, P70431, P70432, P70433, Q5D053 | Gene names | Enah, Mena, Ndpp1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein enabled homolog (NPC-derived proline-rich protein 1) (NDPP-1). | |||||
|
CIC_HUMAN
|
||||||
| NC score | 0.054206 (rank : 70) | θ value | θ > 10 (rank : 115) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 992 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q96RK0, Q7LGI1, Q9UEG5, Q9Y6T1 | Gene names | CIC, KIAA0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.054192 (rank : 71) | θ value | θ > 10 (rank : 131) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
PELP1_HUMAN
|
||||||
| NC score | 0.054122 (rank : 72) | θ value | θ > 10 (rank : 132) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 596 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8IZL8, O15450, Q5EGN3, Q6NTE6, Q96FT1, Q9BU60 | Gene names | PELP1, HMX3, MNAR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor) (Transcription factor HMX3). | |||||
|
PRP5_HUMAN
|
||||||
| NC score | 0.053016 (rank : 73) | θ value | θ > 10 (rank : 137) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P04281 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic proline-rich peptide IB-1. | |||||
|
SFPQ_HUMAN
|
||||||
| NC score | 0.052338 (rank : 74) | θ value | θ > 10 (rank : 145) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 986 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P23246, P30808 | Gene names | SFPQ, PSF | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, proline- and glutamine-rich (Polypyrimidine tract- binding protein-associated-splicing factor) (PTB-associated-splicing factor) (PSF) (DNA-binding p52/p100 complex, 100 kDa subunit) (100 kDa DNA-pairing protein) (hPOMp100). | |||||
|
CS016_HUMAN
|
||||||
| NC score | 0.052061 (rank : 75) | θ value | θ > 10 (rank : 121) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8ND99 | Gene names | C19orf16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C19orf16. | |||||
|
MILK1_HUMAN
|
||||||
| NC score | 0.052038 (rank : 76) | θ value | 1.06291 (rank : 29) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8N3F8, Q5TI16, Q7RTP5, Q8N3N8, Q9BVL9, Q9BY92, Q9UH43, Q9UH44, Q9UH45 | Gene names | MIRAB13, KIAA1668 | |||
|
Domain Architecture |

|
|||||
| Description | Molecule interacting with Rab13 (MIRab13) (MICAL-like protein 1). | |||||
|
MILK1_MOUSE
|
||||||
| NC score | 0.051838 (rank : 77) | θ value | θ > 10 (rank : 128) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 788 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8BGT6, Q8BJ60 | Gene names | Mirab13, D15Mit260, Kiaa1668 | |||
|
Domain Architecture |

|
|||||
| Description | Molecule interacting with Rab13 (MIRab13) (MICAL-like protein 1). | |||||
|
FMN2_HUMAN
|
||||||
| NC score | 0.051833 (rank : 78) | θ value | θ > 10 (rank : 123) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 484 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9NZ56, Q59GF6, Q5VU37, Q9NZ55 | Gene names | FMN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Formin-2. | |||||
|
WASL_MOUSE
|
||||||
| NC score | 0.051675 (rank : 79) | θ value | θ > 10 (rank : 152) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 339 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q91YD9 | Gene names | Wasl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neural Wiskott-Aldrich syndrome protein (N-WASP). | |||||
|
BAT3_MOUSE
|
||||||
| NC score | 0.051247 (rank : 80) | θ value | θ > 10 (rank : 113) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 518 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9Z1R2, Q8SNA3 | Gene names | Bat3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Large proline-rich protein BAT3 (HLA-B-associated transcript 3). | |||||
|
CEL_HUMAN
|
||||||
| NC score | 0.050278 (rank : 81) | θ value | 1.38821 (rank : 35) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P19835, Q16398 | Gene names | CEL, BAL | |||
|
Domain Architecture |

|
|||||
| Description | Bile salt-activated lipase precursor (EC 3.1.1.3) (EC 3.1.1.13) (BAL) (Bile salt-stimulated lipase) (BSSL) (Carboxyl ester lipase) (Sterol esterase) (Cholesterol esterase) (Pancreatic lysophospholipase). | |||||
|
SRRM1_HUMAN
|
||||||
| NC score | 0.050259 (rank : 82) | θ value | θ > 10 (rank : 147) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
IP3KC_MOUSE
|
||||||
| NC score | 0.047209 (rank : 83) | θ value | 0.365318 (rank : 18) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7TS72, Q3U384 | Gene names | Itpkc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol-trisphosphate 3-kinase C (EC 2.7.1.127) (Inositol 1,4,5- trisphosphate 3-kinase C) (IP3K-C). | |||||
|
TCGAP_HUMAN
|
||||||
| NC score | 0.046295 (rank : 84) | θ value | 0.813845 (rank : 28) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 872 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O14559, O14552, O14560, Q6ZSP6, Q96CP3, Q9NT23 | Gene names | SNX26, TCGAP | |||
|
Domain Architecture |

|
|||||
| Description | TC10/CDC42 GTPase-activating protein (Sorting nexin-26). | |||||
|
R3HD2_MOUSE
|
||||||
| NC score | 0.045265 (rank : 85) | θ value | 4.03905 (rank : 70) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q80TM6, Q80YB1, Q8BLS5, Q9CW50 | Gene names | R3hdm2, Kiaa1002 | |||
|
Domain Architecture |

|
|||||
| Description | R3H domain-containing protein 2. | |||||
|
IQEC3_MOUSE
|
||||||
| NC score | 0.045090 (rank : 86) | θ value | 0.279714 (rank : 15) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q3TES0, Q3UHP8, Q80TJ8 | Gene names | Iqsec3, Kiaa1110 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 3. | |||||
|
PSL1_HUMAN
|
||||||
| NC score | 0.044614 (rank : 87) | θ value | 6.88961 (rank : 97) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8TCT7, O60365, Q567S3, Q8IUH9, Q9BUY6, Q9H3M4, Q9NPN2, Q9P1Z6 | Gene names | SPPL2B, KIAA1532, PSL1 | |||
|
Domain Architecture |

|
|||||
| Description | Signal peptide peptidase-like 2B (EC 3.4.23.-) (Protein SPP-like 2B) (Protein SPPL2b) (Intramembrane protease 4) (IMP4) (Presenilin-like protein 1). | |||||
|
ROBO3_HUMAN
|
||||||
| NC score | 0.044145 (rank : 88) | θ value | 0.0736092 (rank : 9) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 570 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q96MS0 | Gene names | ROBO3 | |||
|
Domain Architecture |

|
|||||
| Description | Roundabout homolog 3 precursor (Roundabout-like protein 3). | |||||
|
LBXCO_HUMAN
|
||||||
| NC score | 0.043942 (rank : 89) | θ value | 5.27518 (rank : 79) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P84550 | Gene names | LBXCOR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ladybird homeobox corepressor 1. | |||||
|
CSPG5_HUMAN
|
||||||
| NC score | 0.043824 (rank : 90) | θ value | 3.0926 (rank : 53) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O95196, Q71M39, Q71M40 | Gene names | CSPG5, CALEB, NGC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate proteoglycan 5 precursor (Neuroglycan C) (Acidic leucine-rich EGF-like domain-containing brain protein). | |||||
|
RTEL1_HUMAN
|
||||||
| NC score | 0.042783 (rank : 91) | θ value | 1.06291 (rank : 33) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NZ71, Q5JTV4, Q9BW37, Q9H402, Q9H4X6, Q9NX25, Q9NZ73, Q9UPR4, Q9Y4R6 | Gene names | RTEL1, C20orf41, KIAA1088, NHL | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of telomere elongation helicase 1 (EC 3.6.1.-) (Helicase- like protein NHL). | |||||
|
HCN2_HUMAN
|
||||||
| NC score | 0.042152 (rank : 92) | θ value | 1.38821 (rank : 36) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 369 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9UL51, O60742, O60743, O75267, Q9UBS2 | Gene names | HCN2, BCNG2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 2 (Brain cyclic nucleotide-gated channel 2) (BCNG-2). | |||||
|
CO7A1_HUMAN
|
||||||
| NC score | 0.041884 (rank : 93) | θ value | 0.365318 (rank : 17) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 807 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q02388, Q14054, Q16507 | Gene names | COL7A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(VII) chain precursor (Long-chain collagen) (LC collagen). | |||||
|
ROBO3_MOUSE
|
||||||
| NC score | 0.041404 (rank : 94) | θ value | 0.21417 (rank : 12) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 524 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9Z2I4 | Gene names | Robo3, Rbig1 | |||
|
Domain Architecture |

|
|||||
| Description | Roundabout homolog 3 precursor (Retinoblastoma-inhibiting gene 1) (Rig-1). | |||||
|
CO4A4_HUMAN
|
||||||
| NC score | 0.040373 (rank : 95) | θ value | 5.27518 (rank : 74) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 440 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P53420 | Gene names | COL4A4 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-4(IV) chain precursor. | |||||
|
LEUK_HUMAN
|
||||||
| NC score | 0.039832 (rank : 96) | θ value | 6.88961 (rank : 91) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P16150 | Gene names | SPN, CD43 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leukosialin precursor (Leukocyte sialoglycoprotein) (Sialophorin) (Galactoglycoprotein) (GALGP) (CD43 antigen). | |||||
|
CSKI1_MOUSE
|
||||||
| NC score | 0.039462 (rank : 97) | θ value | 2.36792 (rank : 42) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1031 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q6P9K8, Q6ZPU2, Q8BWU2, Q8BX99, Q9CXH0 | Gene names | Caskin1, Kiaa1306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-1 (CASK-interacting protein 1). | |||||
|
NRG3_MOUSE
|
||||||
| NC score | 0.039082 (rank : 98) | θ value | 4.03905 (rank : 68) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O35181 | Gene names | Nrg3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pro-neuregulin-3, membrane-bound isoform precursor (Pro-NRG3) [Contains: Neuregulin-3 (NRG-3)]. | |||||
|
CF155_HUMAN
|
||||||
| NC score | 0.038447 (rank : 99) | θ value | 8.99809 (rank : 100) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H8W2, Q5TG13 | Gene names | C6orf155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf155. | |||||
|
HRX_HUMAN
|
||||||
| NC score | 0.038302 (rank : 100) | θ value | 2.36792 (rank : 45) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
TGON2_HUMAN
|
||||||
| NC score | 0.038241 (rank : 101) | θ value | 5.27518 (rank : 84) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
JPH2_HUMAN
|
||||||
| NC score | 0.037997 (rank : 102) | θ value | 3.0926 (rank : 56) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 639 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9BR39, O95913, Q5JY74, Q9UJN4 | Gene names | JPH2, JP2 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-2 (Junctophilin type 2) (JP-2). | |||||
|
RECQ4_HUMAN
|
||||||
| NC score | 0.037939 (rank : 103) | θ value | 1.06291 (rank : 32) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O94761, Q3Y424, Q96DW2, Q96F55 | Gene names | RECQL4, RECQ4 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-dependent DNA helicase Q4 (EC 3.6.1.-) (RecQ protein-like 4) (RecQ4) (RTS). | |||||
|
MKL1_MOUSE
|
||||||
| NC score | 0.036918 (rank : 104) | θ value | 4.03905 (rank : 66) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 667 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8K4J6 | Gene names | Mkl1, Bsac | |||
|
Domain Architecture |

|
|||||
| Description | MKL/myocardin-like protein 1 (Myocardin-related transcription factor A) (MRTF-A) (Megakaryoblastic leukemia 1 protein homolog) (Basic SAP coiled-coil transcription activator). | |||||
|
SRTD2_HUMAN
|
||||||
| NC score | 0.035514 (rank : 105) | θ value | 3.0926 (rank : 60) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q14140 | Gene names | SERTAD2, KIAA0127 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SERTA domain-containing protein 2 (Transcriptional regulator interacting with the PHD-bromodomain 2) (TRIP-Br2). | |||||
|
CRSP7_HUMAN
|
||||||
| NC score | 0.031479 (rank : 106) | θ value | 8.99809 (rank : 103) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O95402 | Gene names | CRSP7, ARC70 | |||
|
Domain Architecture |

|
|||||
| Description | CRSP complex subunit 7 (Cofactor required for Sp1 transcriptional activation subunit 7) (Transcriptional coactivator CRSP70) (Activator- recruited cofactor 70 kDa component) (ARC70). | |||||
|
WNK2_HUMAN
|
||||||
| NC score | 0.031422 (rank : 107) | θ value | 0.365318 (rank : 20) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1407 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9Y3S1, Q8IY36, Q9C0A3, Q9H3P4 | Gene names | WNK2, KIAA1760, PRKWNK2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK2 (EC 2.7.11.1) (Protein kinase with no lysine 2) (Protein kinase, lysine-deficient 2). | |||||
|
GP108_HUMAN
|
||||||
| NC score | 0.028896 (rank : 108) | θ value | 2.36792 (rank : 44) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NPR9 | Gene names | GPR108 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein GPR108 precursor. | |||||
|
DTX1_MOUSE
|
||||||
| NC score | 0.027213 (rank : 109) | θ value | 8.99809 (rank : 104) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q61010, Q3TER5, Q8C2H2, Q9ER09 | Gene names | Dtx1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein deltex-1 (Deltex-1) (Deltex1) (mDTX1) (FXI-T1). | |||||
|
LSD1_HUMAN
|
||||||
| NC score | 0.026992 (rank : 110) | θ value | 6.88961 (rank : 92) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O60341, Q5TH94, Q5TH95, Q86VT7, Q8IXK4, Q8NDP6, Q8TAZ3, Q96AW4 | Gene names | AOF2, KIAA0601, LSD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lysine-specific histone demethylase 1 (EC 1.-.-.-) (Flavin-containing amine oxidase domain-containing protein 2) (BRAF35-HDAC complex protein BHC110). | |||||
|
NRCAM_HUMAN
|
||||||
| NC score | 0.026735 (rank : 111) | θ value | 2.36792 (rank : 48) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 439 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q92823, O15051, O15179, Q9UHI3, Q9UHI4 | Gene names | NRCAM, KIAA0343 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal cell adhesion molecule precursor (Nr-CAM) (NgCAM-related cell adhesion molecule) (Ng-CAM-related) (hBravo). | |||||
|
CO2A1_MOUSE
|
||||||
| NC score | 0.026599 (rank : 112) | θ value | 8.99809 (rank : 102) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 483 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P28481 | Gene names | Col2a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(II) chain precursor [Contains: Chondrocalcin]. | |||||
|
HGS_MOUSE
|
||||||
| NC score | 0.026321 (rank : 113) | θ value | 5.27518 (rank : 76) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q99LI8, Q61691, Q8BQW3 | Gene names | Hgs, Hrs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate. | |||||
|
NRCAM_MOUSE
|
||||||
| NC score | 0.026222 (rank : 114) | θ value | 2.36792 (rank : 49) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 432 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q810U4, Q80U33, Q8BLG8, Q8BX92, Q8BYJ8 | Gene names | Nrcam, Kiaa0343 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal cell adhesion molecule precursor (Nr-CAM) (NgCAM-related cell adhesion molecule) (Ng-CAM-related) (mBravo). | |||||
|
INVS_MOUSE
|
||||||
| NC score | 0.026102 (rank : 115) | θ value | 1.38821 (rank : 37) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 609 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O89019, O88849 | Gene names | Invs, Inv, Nphp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inversin (Inversion of embryo turning protein) (Nephrocystin-2). | |||||
|
HXA13_HUMAN
|
||||||
| NC score | 0.025683 (rank : 116) | θ value | 0.813845 (rank : 23) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 353 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P31271, O43371 | Gene names | HOXA13, HOX1J | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-A13 (Hox-1J). | |||||
|
UB7I1_MOUSE
|
||||||
| NC score | 0.025530 (rank : 117) | θ value | 6.88961 (rank : 99) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P58283, Q3U493, Q3UE56, Q3UGM3, Q68FN0, Q6P1H8, Q6PWY5, Q8BN27, Q8C1U3 | Gene names | Ubce7ip1, Triad3, Uip83, Zin | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin ligase TRIAD3 (EC 6.3.2.-) (Ubiquitin-conjugating enzyme 7-interacting protein 1) (UbcM4-interacting protein 83) (Triad domain- containing protein 3). | |||||
|
BORG1_HUMAN
|
||||||
| NC score | 0.025287 (rank : 118) | θ value | 4.03905 (rank : 62) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O14613, Q9UNS0 | Gene names | CDC42EP2, BORG1, CEP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cdc42 effector protein 2 (Binder of Rho GTPases 1). | |||||
|
DAB2_HUMAN
|
||||||
| NC score | 0.024922 (rank : 119) | θ value | 6.88961 (rank : 88) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P98082, Q13598, Q9BTY0, Q9UK04 | Gene names | DAB2, DOC2 | |||
|
Domain Architecture |

|
|||||
| Description | Disabled homolog 2 (Differentially expressed protein 2) (DOC-2). | |||||
|
NOTC3_HUMAN
|
||||||
| NC score | 0.024503 (rank : 120) | θ value | 0.21417 (rank : 11) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1062 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9UM47, Q9UEB3, Q9UPL3, Q9Y6L8 | Gene names | NOTCH3 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 3 precursor (Notch 3) [Contains: Notch 3 extracellular truncation; Notch 3 intracellular domain]. | |||||
|
UBP31_HUMAN
|
||||||
| NC score | 0.023944 (rank : 121) | θ value | 4.03905 (rank : 73) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 293 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q70CQ4, Q6AW97, Q6ZTC0, Q6ZTN2, Q9ULL7 | Gene names | USP31, KIAA1203 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 31 (EC 3.1.2.15) (Ubiquitin thioesterase 31) (Ubiquitin-specific-processing protease 31) (Deubiquitinating enzyme 31). | |||||
|
TCFL5_HUMAN
|
||||||
| NC score | 0.023890 (rank : 122) | θ value | 8.99809 (rank : 110) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UL49, O94771, Q9BYW0 | Gene names | TCFL5, CHA, E2BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor-like 5 protein (Cha transcription factor) (HPV-16 E2-binding protein 1) (E2BP-1). | |||||
|
FOXC1_HUMAN
|
||||||
| NC score | 0.023889 (rank : 123) | θ value | 3.0926 (rank : 55) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q12948, Q86UP7, Q9BYM1, Q9NUE5, Q9UDD0, Q9UP06 | Gene names | FOXC1, FKHL7, FREAC3 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein C1 (Forkhead-related protein FKHL7) (Forkhead- related transcription factor 3) (FREAC-3). | |||||
|
CTR9_MOUSE
|
||||||
| NC score | 0.023856 (rank : 124) | θ value | 4.03905 (rank : 63) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q62018, Q3UFF5, Q3UY40, Q66JX4, Q7TPS6, Q8BND9, Q8BRD1, Q8C9W7, Q8C9Y3, Q8CHI1 | Gene names | Ctr9, Kiaa0155, Sh2bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein CTR9 homolog (SH2 domain-binding protein 1) (Tetratricopeptide repeat-containing, SH2-binding phosphoprotein of 150 kDa) (TPR-containing, SH2-binding phosphoprotein of 150 kDa) (p150TSP). | |||||
|
MYCL1_HUMAN
|
||||||
| NC score | 0.023764 (rank : 125) | θ value | 4.03905 (rank : 67) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P12524, Q14897, Q5QPL0, Q5QPL1, Q9NUE9 | Gene names | MYCL1, LMYC, MYCL | |||
|
Domain Architecture |

|
|||||
| Description | L-myc-1 proto-oncogene protein. | |||||
|
PHLPP_HUMAN
|
||||||
| NC score | 0.023656 (rank : 126) | θ value | 6.88961 (rank : 96) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 604 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O60346, Q641Q7, Q6P4C4, Q6PJI6, Q86TN6, Q96FK2, Q9NUY1 | Gene names | PHLPP, KIAA0606, PLEKHE1, SCOP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH domain leucine-rich repeat-containing protein phosphatase (EC 3.1.3.16) (PH domain leucine-rich repeat protein phosphatase) (Pleckstrin homology domain-containing family E protein 1) (Suprachiasmatic nucleus circadian oscillatory protein) (hSCOP). | |||||
|
HXA13_MOUSE
|
||||||
| NC score | 0.023466 (rank : 127) | θ value | 1.81305 (rank : 40) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q62424 | Gene names | Hoxa13, Hox-1.10 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-A13 (Hox-1.10). | |||||
|
AGRIN_HUMAN
|
||||||
| NC score | 0.023415 (rank : 128) | θ value | 2.36792 (rank : 41) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 582 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O00468, Q5SVA2, Q60FE1, Q7KYS8, Q8N4J5, Q96IC1, Q9BTD4 | Gene names | AGRIN, AGRN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Agrin precursor. | |||||
|
MYO9B_HUMAN
|
||||||
| NC score | 0.020435 (rank : 129) | θ value | 1.06291 (rank : 30) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 641 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q13459, O75314, Q9NUJ2, Q9UHN0 | Gene names | MYO9B, MYR5 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9B (Myosin IXb) (Unconventional myosin-9b). | |||||
|
PYR1_HUMAN
|
||||||
| NC score | 0.019817 (rank : 130) | θ value | 5.27518 (rank : 82) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P27708, Q6P0Q0 | Gene names | CAD | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CAD protein [Includes: Glutamine-dependent carbamoyl-phosphate synthase (EC 6.3.5.5); Aspartate carbamoyltransferase (EC 2.1.3.2); Dihydroorotase (EC 3.5.2.3)]. | |||||
|
NOTC4_HUMAN
|
||||||
| NC score | 0.019656 (rank : 131) | θ value | 1.06291 (rank : 31) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 905 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q99466, O00306, Q99458, Q99940, Q9H3S8, Q9UII9, Q9UIJ0 | Gene names | NOTCH4 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 4 precursor (Notch 4) (hNotch4) [Contains: Notch 4 extracellular truncation; Notch 4 intracellular domain]. | |||||
|
SALL3_HUMAN
|
||||||
| NC score | 0.018237 (rank : 132) | θ value | 1.38821 (rank : 38) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1066 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9BXA9, Q9UGH1 | Gene names | SALL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sal-like protein 3 (Zinc finger protein SALL3) (hSALL3). | |||||
|
PODO_HUMAN
|
||||||
| NC score | 0.017811 (rank : 133) | θ value | 5.27518 (rank : 80) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NP85, Q8N6Q5 | Gene names | NPHS2 | |||
|
Domain Architecture |

|
|||||
| Description | Podocin. | |||||
|
TIF1B_MOUSE
|
||||||
| NC score | 0.016308 (rank : 134) | θ value | 4.03905 (rank : 72) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q62318, P70391, Q8C283, Q99PN4 | Gene names | Trim28, Krip1, Tif1b | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-beta (TIF1-beta) (Tripartite motif-containing protein 28) (KRAB-A-interacting protein) (KRIP-1). | |||||
|
UBIP1_MOUSE
|
||||||
| NC score | 0.014315 (rank : 135) | θ value | 5.27518 (rank : 85) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q811S7, Q3UPR3, Q3US11, Q60786, Q8C514 | Gene names | Ubp1, Cp2b, Nf2d9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Upstream-binding protein 1 (Nuclear factor 2d9) (NF2d9). | |||||
|
PO4F2_MOUSE
|
||||||
| NC score | 0.013433 (rank : 136) | θ value | 3.0926 (rank : 59) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q63934, Q63954 | Gene names | Pou4f2, Brn3b | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 4, transcription factor 2 (Brain-specific homeobox/POU domain protein 3B) (Brn-3B) (Brn-3.2). | |||||
|
ARP5_MOUSE
|
||||||
| NC score | 0.013048 (rank : 137) | θ value | 6.88961 (rank : 86) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q80US4, Q8BL26 | Gene names | Actr5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Actin-related protein 5. | |||||
|
GAK_HUMAN
|
||||||
| NC score | 0.012600 (rank : 138) | θ value | 6.88961 (rank : 89) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1028 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O14976, Q9BVY6 | Gene names | GAK | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin G-associated kinase (EC 2.7.11.1). | |||||
|
MINK1_HUMAN
|
||||||
| NC score | 0.012020 (rank : 139) | θ value | 6.88961 (rank : 94) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8N4C8, Q9P1X1, Q9P2R8 | Gene names | MINK1, MAP4K6, MINK | |||
|
Domain Architecture |

|
|||||
| Description | Misshapen-like kinase 1 (EC 2.7.11.1) (Mitogen-activated protein kinase kinase kinase kinase 6) (MAPK/ERK kinase kinase kinase 6) (MEK kinase kinase 6) (MEKKK 6) (Misshapen/NIK-related kinase) (GCK family kinase MiNK). | |||||
|
MAGI2_HUMAN
|
||||||
| NC score | 0.011373 (rank : 140) | θ value | 8.99809 (rank : 109) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 373 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q86UL8, O60434, O60510, Q86UI7, Q9NP44, Q9UDQ5, Q9UDU1 | Gene names | MAGI2, ACVRINP1, AIP1, KIAA0705 | |||
|
Domain Architecture |

|
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 2 (Membrane-associated guanylate kinase inverted 2) (MAGI-2) (Atrophin-1-interacting protein 1) (Atrophin-1-interacting protein A). | |||||
|
DRD4_HUMAN
|
||||||
| NC score | 0.011234 (rank : 141) | θ value | 3.0926 (rank : 54) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 886 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P21917 | Gene names | DRD4 | |||
|
Domain Architecture |

|
|||||
| Description | D(4) dopamine receptor (Dopamine D4 receptor) (D(2C) dopamine receptor). | |||||
|
GAK5_HUMAN
|
||||||
| NC score | 0.011232 (rank : 142) | θ value | 8.99809 (rank : 107) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 429 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P62684 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19p13.11 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K113 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
E2AK1_HUMAN
|
||||||
| NC score | 0.010918 (rank : 143) | θ value | 8.99809 (rank : 105) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 833 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9BQI3, Q8NBW3, Q9HC02, Q9NYE0, Q9P0V6, Q9P1J5, Q9P2H8, Q9UHG4 | Gene names | EIF2AK1, HRI, KIAA1369 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2-alpha kinase 1 (EC 2.7.11.1) (Heme-regulated eukaryotic initiation factor eIF-2-alpha kinase) (Heme-regulated inhibitor) (Heme-controlled repressor) (HCR) (Hemin-sensitive initiation factor 2-alpha kinase). | |||||
|
GP135_HUMAN
|
||||||
| NC score | 0.010737 (rank : 144) | θ value | 1.81305 (rank : 39) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 669 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8IZ08, Q7Z604, Q86SM3, Q8NH39 | Gene names | GPR135 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 135. | |||||
|
E2AK1_MOUSE
|
||||||
| NC score | 0.010611 (rank : 145) | θ value | 8.99809 (rank : 106) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Z2R9, Q8C024, Q8K123, Q9CTP5, Q9D601 | Gene names | Eif2ak1, Hri | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 2-alpha kinase 1 (EC 2.7.11.1) (Heme-regulated eukaryotic initiation factor eIF-2-alpha kinase) (Heme-regulated inhibitor) (Heme-controlled repressor) (HCR) (Hemin-sensitive initiation factor 2-alpha kinase). | |||||
|
MINK1_MOUSE
|
||||||
| NC score | 0.010407 (rank : 146) | θ value | 4.03905 (rank : 65) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1451 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9JM52, Q921M6, Q9JM92 | Gene names | Mink1, Map4k6, Mink | |||
|
Domain Architecture |

|
|||||
| Description | Misshapen-like kinase 1 (EC 2.7.11.1) (Mitogen-activated protein kinase kinase kinase kinase 6) (MAPK/ERK kinase kinase kinase 6) (MEK kinase kinase 6) (MEKKK 6) (Misshapen/NIK-related kinase) (GCK family kinase MiNK). | |||||
|
FOXD1_MOUSE
|
||||||
| NC score | 0.010157 (rank : 147) | θ value | 5.27518 (rank : 75) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61345 | Gene names | Foxd1, Fkhl8, Freac4, Hfhbf2 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein D1 (Forkhead-related protein FKHL8) (Forkhead- related transcription factor 4) (FREAC-4) (Brain factor 2) (HFH-BF-2). | |||||
|
IGHG3_HUMAN
|
||||||
| NC score | 0.009623 (rank : 148) | θ value | 5.27518 (rank : 77) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P01860 | Gene names | IGHG3 | |||
|
Domain Architecture |

|
|||||
| Description | Ig gamma-3 chain C region (Heavy chain disease protein) (HDC). | |||||
|
KLF2_HUMAN
|
||||||
| NC score | 0.009092 (rank : 149) | θ value | 3.0926 (rank : 58) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 755 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Y5W3, Q9UJS5, Q9UKR6 | Gene names | KLF2, LKLF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Krueppel-like factor 2 (Lung krueppel-like factor). | |||||
|
ZN447_HUMAN
|
||||||
| NC score | 0.008781 (rank : 150) | θ value | 3.0926 (rank : 61) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 899 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8TBC5, Q9BRK7, Q9H9A0 | Gene names | ZNF447 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 447. | |||||
|
CHIT1_HUMAN
|
||||||
| NC score | 0.008611 (rank : 151) | θ value | 8.99809 (rank : 101) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q13231, Q3ZAR1, Q6ISC2, Q9H3V8 | Gene names | CHIT1 | |||
|
Domain Architecture |

|
|||||
| Description | Chitotriosidase-1 precursor (EC 3.2.1.14) (Chitinase-1). | |||||
|
HXB1_HUMAN
|
||||||
| NC score | 0.008188 (rank : 152) | θ value | 6.88961 (rank : 90) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P14653 | Gene names | HOXB1, HOX2I | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-B1 (Hox-2I). | |||||
|
M3K6_HUMAN
|
||||||
| NC score | 0.008159 (rank : 153) | θ value | 4.03905 (rank : 64) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 889 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O95382 | Gene names | MAP3K6, MAPKKK6 | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase 6 (EC 2.7.11.25). | |||||
|
KKCC1_MOUSE
|
||||||
| NC score | 0.007535 (rank : 154) | θ value | 2.36792 (rank : 46) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 866 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8VBY2, Q9R054 | Gene names | Camkk1, Camkk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium/calmodulin-dependent protein kinase kinase 1 (EC 2.7.11.17) (Calcium/calmodulin-dependent protein kinase kinase alpha) (CaM-kinase kinase alpha) (CaM-KK alpha) (CaMKK alpha) (CaMKK 1) (CaM-kinase IV kinase). | |||||
|
KCNA5_MOUSE
|
||||||
| NC score | 0.007213 (rank : 155) | θ value | 5.27518 (rank : 78) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61762 | Gene names | Kcna5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 5 (Voltage-gated potassium channel subunit Kv1.5) (KV1-5). | |||||
|
MRCKB_MOUSE
|
||||||
| NC score | 0.006173 (rank : 156) | θ value | 2.36792 (rank : 47) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 2180 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q7TT50, Q69ZR5, Q80W33 | Gene names | Cdc42bpb, Kiaa1124 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK beta (EC 2.7.11.1) (CDC42-binding protein kinase beta) (Myotonic dystrophy kinase-related CDC42-binding kinase beta) (Myotonic dystrophy protein kinase-like beta) (MRCK beta) (DMPK-like beta). | |||||
|
KLF14_HUMAN
|
||||||
| NC score | 0.005643 (rank : 157) | θ value | 3.0926 (rank : 57) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 725 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8TD94 | Gene names | KLF14, BTEB5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Krueppel-like factor 14 (Transcription factor BTEB5) (Basic transcription element-binding protein 5) (BTE-binding protein 5). | |||||
|
LTK_MOUSE
|
||||||
| NC score | 0.003040 (rank : 158) | θ value | 8.99809 (rank : 108) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 770 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P08923 | Gene names | Ltk | |||
|
Domain Architecture |

|
|||||
| Description | Leukocyte tyrosine kinase receptor precursor (EC 2.7.10.1). | |||||