Please be patient as the page loads
|
GPTC3_MOUSE
|
||||||
| SwissProt Accessions | Q8BIY1 | Gene names | Gpatc3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 3. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
GPTC3_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.986618 (rank : 2) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q96I76, Q5JYH2, Q8NDJ2, Q9H9Z3 | Gene names | GPATC3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 3. | |||||
|
GPTC3_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8BIY1 | Gene names | Gpatc3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 3. | |||||
|
TFP11_MOUSE
|
||||||
| θ value | 2.44474e-05 (rank : 3) | NC score | 0.288813 (rank : 3) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9ERA6, Q8VD06 | Gene names | Tfip11, Tip39 | |||
|
Domain Architecture |

|
|||||
| Description | Tuftelin-interacting protein 11 (Tuftelin-interacting protein 39). | |||||
|
TFP11_HUMAN
|
||||||
| θ value | 0.000158464 (rank : 4) | NC score | 0.274558 (rank : 4) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UBB9, O95908, Q5H8V8, Q9UGV7, Q9Y2Q8 | Gene names | TFIP11 | |||
|
Domain Architecture |

|
|||||
| Description | Tuftelin-interacting protein 11. | |||||
|
ZGPAT_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 5) | NC score | 0.175257 (rank : 5) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8VDM1, Q8BWW2 | Gene names | Zgpat | |||
|
Domain Architecture |
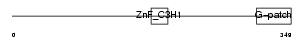
|
|||||
| Description | Zinc finger CCCH-type with G patch domain protein. | |||||
|
INADL_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 6) | NC score | 0.039929 (rank : 32) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q63ZW7, O70471, Q5PRG3, Q6P6J1, Q80YR8, Q8BPB9 | Gene names | Inadl, Cipp, Patj | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | InaD-like protein (Inadl protein) (Pals1-associated tight junction protein) (Protein associated to tight junctions) (Channel-interacting PDZ domain-containing protein). | |||||
|
RBM5_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 7) | NC score | 0.149154 (rank : 6) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P52756, Q93021, Q9BU14, Q9UKY8, Q9UL24 | Gene names | RBM5 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 5 (RNA-binding motif protein 5) (Tumor suppressor LUCA15) (Protein G15) (NY-REN-9 antigen). | |||||
|
RBM5_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 8) | NC score | 0.149126 (rank : 7) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q91YE7, Q3UZ19, Q99KV9 | Gene names | Rbm5, Luca15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein 5 (RNA-binding motif protein 5) (Tumor suppressor LUCA15). | |||||
|
SON_HUMAN
|
||||||
| θ value | 0.279714 (rank : 9) | NC score | 0.059098 (rank : 27) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
SON_MOUSE
|
||||||
| θ value | 0.279714 (rank : 10) | NC score | 0.086239 (rank : 24) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 957 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9QX47, Q9CQ12, Q9CQK6, Q9QXP5 | Gene names | Son | |||
|
Domain Architecture |

|
|||||
| Description | SON protein. | |||||
|
CLD19_HUMAN
|
||||||
| θ value | 0.47712 (rank : 11) | NC score | 0.021017 (rank : 41) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N6F1, Q8N8X0 | Gene names | CLDN19 | |||
|
Domain Architecture |

|
|||||
| Description | Claudin-19. | |||||
|
LEO1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 12) | NC score | 0.042310 (rank : 31) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
SF04_HUMAN
|
||||||
| θ value | 0.47712 (rank : 13) | NC score | 0.131720 (rank : 10) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8IWZ8, O60378, Q6P3X9, Q8TCQ4, Q8WWT4, Q8WWT5, Q9NTG3 | Gene names | SF4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor 4 (RNA-binding protein RBP). | |||||
|
CK5P2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 14) | NC score | 0.022095 (rank : 39) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1151 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8K389, Q6PCN1, Q6ZPL0 | Gene names | Cdk5rap2, Kiaa1633 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CDK5 regulatory subunit-associated protein 2 (CDK5 activator-binding protein C48). | |||||
|
ERIC1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 15) | NC score | 0.054284 (rank : 28) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q86X53, Q9P063 | Gene names | ERICH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate-rich protein 1. | |||||
|
SFR14_HUMAN
|
||||||
| θ value | 0.62314 (rank : 16) | NC score | 0.130742 (rank : 11) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8IX01, O15071, O60369, Q5JPH7, Q8WUF7 | Gene names | SFRS14, KIAA0365 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative splicing factor, arginine/serine-rich 14 (Arginine/serine- rich-splicing factor 14). | |||||
|
GPTC2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 17) | NC score | 0.109005 (rank : 15) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9NW75, Q86YE7 | Gene names | GPATC2 | |||
|
Domain Architecture |

|
|||||
| Description | G patch domain-containing protein 2. | |||||
|
K0082_HUMAN
|
||||||
| θ value | 1.06291 (rank : 18) | NC score | 0.097458 (rank : 20) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8N1G2, Q14670, Q96FJ9 | Gene names | KIAA0082 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0082. | |||||
|
SF04_MOUSE
|
||||||
| θ value | 1.06291 (rank : 19) | NC score | 0.124687 (rank : 12) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8CH02, Q8R094 | Gene names | Sf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor 4. | |||||
|
SFR14_MOUSE
|
||||||
| θ value | 1.06291 (rank : 20) | NC score | 0.124006 (rank : 13) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8CH09, Q6PG19, Q80UY8, Q8BY32, Q8CFM0 | Gene names | Sfrs14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative splicing factor, arginine/serine-rich 14 (Arginine/serine- rich-splicing factor 14). | |||||
|
TCAL5_HUMAN
|
||||||
| θ value | 1.06291 (rank : 21) | NC score | 0.027986 (rank : 36) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 806 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q5H9L2 | Gene names | TCEAL5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 5 (TCEA-like protein 5) (Transcription elongation factor S-II protein-like 5). | |||||
|
ZGPAT_HUMAN
|
||||||
| θ value | 1.06291 (rank : 22) | NC score | 0.142913 (rank : 8) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8N5A5, Q5JWI9, Q8NC55, Q96JI0, Q96JU4, Q9H401 | Gene names | ZGPAT, KIAA1847, ZC3H9, ZC3HDC9 | |||
|
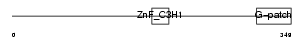
Domain Architecture |
|
|||||
| Description | Zinc finger CCCH-type with G patch domain protein (Zinc finger CCCH domain-containing protein 9). | |||||
|
PINX1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 23) | NC score | 0.133760 (rank : 9) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9CZX5, Q91WZ9, Q9D0C2 | Gene names | Pinx1, Lpts | |||
|
Domain Architecture |

|
|||||
| Description | Pin2-interacting protein X1 (TRF1-interacting protein 1) (Liver- related putative tumor suppressor) (LPTS1) (67-11-3 protein). | |||||
|
AGGF1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 24) | NC score | 0.106345 (rank : 17) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8N302, O00581, Q53YS3, Q9BU84, Q9NW66 | Gene names | AGGF1, VG5Q | |||
|
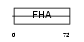
Domain Architecture |
|
|||||
| Description | Angiogenic factor with G patch and FHA domains 1 (Angiogenic factor VG5Q) (Vasculogenesis gene on 5q protein) (hVG5Q). | |||||
|
BASI_HUMAN
|
||||||
| θ value | 1.81305 (rank : 25) | NC score | 0.033529 (rank : 35) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P35613, Q7Z796, Q8IZL7 | Gene names | BSG | |||
|
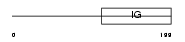
Domain Architecture |
|
|||||
| Description | Basigin precursor (Leukocyte activation antigen M6) (Collagenase stimulatory factor) (Extracellular matrix metalloproteinase inducer) (EMMPRIN) (5F7) (Tumor cell-derived collagenase stimulatory factor) (TCSF) (OK blood group antigen) (CD147 antigen). | |||||
|
AGGF1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 26) | NC score | 0.107123 (rank : 16) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q7TN31, Q8R2S6, Q9CQR9, Q9CU87, Q9D768 | Gene names | Aggf1, Vg5q | |||
|
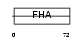
Domain Architecture |
|
|||||
| Description | Angiogenic factor with G patch and FHA domains 1 (Angiogenic factor VG5Q) (mVG5Q). | |||||
|
GANAB_HUMAN
|
||||||
| θ value | 2.36792 (rank : 27) | NC score | 0.019836 (rank : 44) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q14697, Q8WTS9, Q9P0X0 | Gene names | GANAB, G2AN, KIAA0088 | |||
|
Domain Architecture |

|
|||||
| Description | Neutral alpha-glucosidase AB precursor (EC 3.2.1.84) (Glucosidase II subunit alpha). | |||||
|
NSBP1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 28) | NC score | 0.037826 (rank : 33) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
GPTC2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 29) | NC score | 0.101097 (rank : 19) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q7TQC7, Q8BNJ9, Q8BPM1, Q8CDH9 | Gene names | Gpatc2 | |||
|
Domain Architecture |

|
|||||
| Description | G patch domain-containing protein 2. | |||||
|
LEO1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 30) | NC score | 0.035326 (rank : 34) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
MAP1A_HUMAN
|
||||||
| θ value | 3.0926 (rank : 31) | NC score | 0.021327 (rank : 40) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P78559, O95643, Q12973, Q15882, Q9UJT4 | Gene names | MAP1A | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 1A (MAP 1A) (Proliferation-related protein p80) [Contains: MAP1 light chain LC2]. | |||||
|
NKRF_MOUSE
|
||||||
| θ value | 3.0926 (rank : 32) | NC score | 0.089505 (rank : 22) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8BY02, Q8BJU5, Q9CRL2 | Gene names | Nkrf, Nrf | |||
|
Domain Architecture |

|
|||||
| Description | NF-kappa-B-repressing factor (NFKB-repressing factor) (Transcription factor NRF). | |||||
|
ACINU_HUMAN
|
||||||
| θ value | 4.03905 (rank : 33) | NC score | 0.026155 (rank : 38) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UKV3, O75158, Q9UG91, Q9UKV1, Q9UKV2 | Gene names | ACIN1, ACINUS, KIAA0670 | |||
|
Domain Architecture |

|
|||||
| Description | Apoptotic chromatin condensation inducer in the nucleus (Acinus). | |||||
|
ANK1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 34) | NC score | 0.008393 (rank : 49) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P16157, O43400, Q13768, Q59FP2, Q8N604, Q99407 | Gene names | ANK1, ANK | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-1 (Erythrocyte ankyrin) (Ankyrin-R). | |||||
|
K0082_MOUSE
|
||||||
| θ value | 4.03905 (rank : 35) | NC score | 0.092002 (rank : 21) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9DBC3, Q3U3G5, Q3U7Y9, Q6A0D5, Q8C7V0 | Gene names | Kiaa0082 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0082. | |||||
|
IWS1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 36) | NC score | 0.020206 (rank : 43) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
NKRF_HUMAN
|
||||||
| θ value | 5.27518 (rank : 37) | NC score | 0.087083 (rank : 23) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O15226, Q9UJ91 | Gene names | NKRF, ITBA4, NRF | |||
|
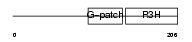
Domain Architecture |
|
|||||
| Description | NF-kappa-B-repressing factor (NFKB-repressing factor) (Transcription factor NRF) (ITBA4 protein). | |||||
|
SAS10_HUMAN
|
||||||
| θ value | 5.27518 (rank : 38) | NC score | 0.067975 (rank : 25) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NQZ2, Q6FI82 | Gene names | CRLZ1, SAS10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Something about silencing protein 10 (Disrupter of silencing SAS10) (Charged amino acid-rich leucine zipper 1). | |||||
|
CK5P2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 39) | NC score | 0.020360 (rank : 42) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1433 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q96SN8, Q7Z3L4, Q7Z3U1, Q7Z7I6, Q9BSW0, Q9H6J6, Q9HCD9, Q9NV90, Q9UIW9 | Gene names | CDK5RAP2, CEP215, KIAA1633 | |||
|
Domain Architecture |

|
|||||
| Description | CDK5 regulatory subunit-associated protein 2 (CDK5 activator-binding protein C48) (Centrosome-associated protein 215). | |||||
|
PACN2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 40) | NC score | 0.012779 (rank : 47) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 382 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9WVE8 | Gene names | Pacsin2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C and casein kinase substrate in neurons protein 2. | |||||
|
PINX1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 41) | NC score | 0.105920 (rank : 18) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q96BK5, Q7Z7J8, Q96QD7, Q9HBU7, Q9NWW2 | Gene names | PINX1, LPTS | |||
|
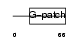
Domain Architecture |
|
|||||
| Description | Pin2-interacting protein X1 (TRF1-interacting protein 1) (Liver- related putative tumor suppressor) (67-11-3 protein). | |||||
|
RAD_MOUSE
|
||||||
| θ value | 6.88961 (rank : 42) | NC score | 0.007962 (rank : 50) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O88667 | Gene names | Rrad | |||
|
Domain Architecture |

|
|||||
| Description | GTP-binding protein RAD. | |||||
|
SPF45_HUMAN
|
||||||
| θ value | 6.88961 (rank : 43) | NC score | 0.052779 (rank : 30) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q96I25, Q96GY6 | Gene names | RBM17, SPF45 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 45 (45 kDa-splicing factor) (RNA-binding motif protein 17). | |||||
|
SPF45_MOUSE
|
||||||
| θ value | 6.88961 (rank : 44) | NC score | 0.053825 (rank : 29) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8JZX4, O75939 | Gene names | Rbm17, Spf45 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 45 (45 kDa-splicing factor) (RNA-binding motif protein 17). | |||||
|
ADPPT_MOUSE
|
||||||
| θ value | 8.99809 (rank : 45) | NC score | 0.027806 (rank : 37) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9CQF6, Q5U5W8, Q9CU40, Q9D827 | Gene names | Aasdhppt | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | L-aminoadipate-semialdehyde dehydrogenase-phosphopantetheinyl transferase (EC 2.7.8.-) (4'-phosphopantetheinyl transferase) (Alpha- aminoadipic semialdehyde dehydrogenase-phosphopantetheinyl transferase) (AASD-PPT). | |||||
|
LAG3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 46) | NC score | 0.014239 (rank : 46) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P18627 | Gene names | LAG3, FDC | |||
|
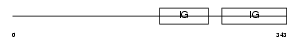
Domain Architecture |
|
|||||
| Description | Lymphocyte activation gene 3 protein precursor (LAG-3) (FDC protein) (CD223 antigen). | |||||
|
MLL4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 47) | NC score | 0.012087 (rank : 48) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UMN6, O15022, O95836, Q96GP2, Q96IP3, Q9UK25, Q9Y668, Q9Y669 | Gene names | MLL4, HRX2, KIAA0304, MLL2, TRX2 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 4 (Trithorax homolog 2). | |||||
|
RAD_HUMAN
|
||||||
| θ value | 8.99809 (rank : 48) | NC score | 0.006920 (rank : 51) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 224 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P55042, Q96F39 | Gene names | RRAD, RAD | |||
|
Domain Architecture |

|
|||||
| Description | GTP-binding protein RAD (RAS associated with diabetes) (RAD1). | |||||
|
RBM10_HUMAN
|
||||||
| θ value | 8.99809 (rank : 49) | NC score | 0.111727 (rank : 14) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P98175, Q14136 | Gene names | RBM10, DXS8237E, KIAA0122 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 10 (RNA-binding motif protein 10). | |||||
|
SAS10_MOUSE
|
||||||
| θ value | 8.99809 (rank : 50) | NC score | 0.060051 (rank : 26) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9JI13, Q3UL95, Q8R5C6, Q9JJ12 | Gene names | Crlz1, Crl1, Sas10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Something about silencing protein 10 (Disrupter of silencing SAS10) (Charged amino acid-rich leucine zipper 1) (Crl-1). | |||||
|
SIM2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 51) | NC score | 0.006805 (rank : 52) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q14190, O60766, Q15470, Q15471, Q15472, Q15473, Q16532 | Gene names | SIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2. | |||||
|
TGON2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 52) | NC score | 0.016257 (rank : 45) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q62314 | Gene names | Tgoln2, Ttgn2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (TGN38B). | |||||
|
GPTC3_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8BIY1 | Gene names | Gpatc3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 3. | |||||
|
GPTC3_HUMAN
|
||||||
| NC score | 0.986618 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q96I76, Q5JYH2, Q8NDJ2, Q9H9Z3 | Gene names | GPATC3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 3. | |||||
|
TFP11_MOUSE
|
||||||
| NC score | 0.288813 (rank : 3) | θ value | 2.44474e-05 (rank : 3) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9ERA6, Q8VD06 | Gene names | Tfip11, Tip39 | |||
|
Domain Architecture |

|
|||||
| Description | Tuftelin-interacting protein 11 (Tuftelin-interacting protein 39). | |||||
|
TFP11_HUMAN
|
||||||
| NC score | 0.274558 (rank : 4) | θ value | 0.000158464 (rank : 4) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UBB9, O95908, Q5H8V8, Q9UGV7, Q9Y2Q8 | Gene names | TFIP11 | |||
|
Domain Architecture |

|
|||||
| Description | Tuftelin-interacting protein 11. | |||||
|
ZGPAT_MOUSE
|
||||||
| NC score | 0.175257 (rank : 5) | θ value | 0.0431538 (rank : 5) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8VDM1, Q8BWW2 | Gene names | Zgpat | |||
|
Domain Architecture |
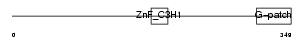
|
|||||
| Description | Zinc finger CCCH-type with G patch domain protein. | |||||
|
RBM5_HUMAN
|
||||||
| NC score | 0.149154 (rank : 6) | θ value | 0.0961366 (rank : 7) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P52756, Q93021, Q9BU14, Q9UKY8, Q9UL24 | Gene names | RBM5 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 5 (RNA-binding motif protein 5) (Tumor suppressor LUCA15) (Protein G15) (NY-REN-9 antigen). | |||||
|
RBM5_MOUSE
|
||||||
| NC score | 0.149126 (rank : 7) | θ value | 0.0961366 (rank : 8) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q91YE7, Q3UZ19, Q99KV9 | Gene names | Rbm5, Luca15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein 5 (RNA-binding motif protein 5) (Tumor suppressor LUCA15). | |||||
|
ZGPAT_HUMAN
|
||||||
| NC score | 0.142913 (rank : 8) | θ value | 1.06291 (rank : 22) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8N5A5, Q5JWI9, Q8NC55, Q96JI0, Q96JU4, Q9H401 | Gene names | ZGPAT, KIAA1847, ZC3H9, ZC3HDC9 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger CCCH-type with G patch domain protein (Zinc finger CCCH domain-containing protein 9). | |||||
|
PINX1_MOUSE
|
||||||
| NC score | 0.133760 (rank : 9) | θ value | 1.38821 (rank : 23) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9CZX5, Q91WZ9, Q9D0C2 | Gene names | Pinx1, Lpts | |||
|
Domain Architecture |

|
|||||
| Description | Pin2-interacting protein X1 (TRF1-interacting protein 1) (Liver- related putative tumor suppressor) (LPTS1) (67-11-3 protein). | |||||
|
SF04_HUMAN
|
||||||
| NC score | 0.131720 (rank : 10) | θ value | 0.47712 (rank : 13) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8IWZ8, O60378, Q6P3X9, Q8TCQ4, Q8WWT4, Q8WWT5, Q9NTG3 | Gene names | SF4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor 4 (RNA-binding protein RBP). | |||||
|
SFR14_HUMAN
|
||||||
| NC score | 0.130742 (rank : 11) | θ value | 0.62314 (rank : 16) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8IX01, O15071, O60369, Q5JPH7, Q8WUF7 | Gene names | SFRS14, KIAA0365 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative splicing factor, arginine/serine-rich 14 (Arginine/serine- rich-splicing factor 14). | |||||
|
SF04_MOUSE
|
||||||
| NC score | 0.124687 (rank : 12) | θ value | 1.06291 (rank : 19) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8CH02, Q8R094 | Gene names | Sf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor 4. | |||||
|
SFR14_MOUSE
|
||||||
| NC score | 0.124006 (rank : 13) | θ value | 1.06291 (rank : 20) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8CH09, Q6PG19, Q80UY8, Q8BY32, Q8CFM0 | Gene names | Sfrs14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative splicing factor, arginine/serine-rich 14 (Arginine/serine- rich-splicing factor 14). | |||||
|
RBM10_HUMAN
|
||||||
| NC score | 0.111727 (rank : 14) | θ value | 8.99809 (rank : 49) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P98175, Q14136 | Gene names | RBM10, DXS8237E, KIAA0122 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 10 (RNA-binding motif protein 10). | |||||
|
GPTC2_HUMAN
|
||||||
| NC score | 0.109005 (rank : 15) | θ value | 1.06291 (rank : 17) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9NW75, Q86YE7 | Gene names | GPATC2 | |||
|
Domain Architecture |

|
|||||
| Description | G patch domain-containing protein 2. | |||||
|
AGGF1_MOUSE
|
||||||
| NC score | 0.107123 (rank : 16) | θ value | 2.36792 (rank : 26) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q7TN31, Q8R2S6, Q9CQR9, Q9CU87, Q9D768 | Gene names | Aggf1, Vg5q | |||
|
Domain Architecture |

|
|||||
| Description | Angiogenic factor with G patch and FHA domains 1 (Angiogenic factor VG5Q) (mVG5Q). | |||||
|
AGGF1_HUMAN
|
||||||
| NC score | 0.106345 (rank : 17) | θ value | 1.81305 (rank : 24) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8N302, O00581, Q53YS3, Q9BU84, Q9NW66 | Gene names | AGGF1, VG5Q | |||
|
Domain Architecture |

|
|||||
| Description | Angiogenic factor with G patch and FHA domains 1 (Angiogenic factor VG5Q) (Vasculogenesis gene on 5q protein) (hVG5Q). | |||||
|
PINX1_HUMAN
|
||||||
| NC score | 0.105920 (rank : 18) | θ value | 6.88961 (rank : 41) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q96BK5, Q7Z7J8, Q96QD7, Q9HBU7, Q9NWW2 | Gene names | PINX1, LPTS | |||
|
Domain Architecture |

|
|||||
| Description | Pin2-interacting protein X1 (TRF1-interacting protein 1) (Liver- related putative tumor suppressor) (67-11-3 protein). | |||||
|
GPTC2_MOUSE
|
||||||
| NC score | 0.101097 (rank : 19) | θ value | 3.0926 (rank : 29) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q7TQC7, Q8BNJ9, Q8BPM1, Q8CDH9 | Gene names | Gpatc2 | |||
|
Domain Architecture |

|
|||||
| Description | G patch domain-containing protein 2. | |||||
|
K0082_HUMAN
|
||||||
| NC score | 0.097458 (rank : 20) | θ value | 1.06291 (rank : 18) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8N1G2, Q14670, Q96FJ9 | Gene names | KIAA0082 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0082. | |||||
|
K0082_MOUSE
|
||||||
| NC score | 0.092002 (rank : 21) | θ value | 4.03905 (rank : 35) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9DBC3, Q3U3G5, Q3U7Y9, Q6A0D5, Q8C7V0 | Gene names | Kiaa0082 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0082. | |||||
|
NKRF_MOUSE
|
||||||
| NC score | 0.089505 (rank : 22) | θ value | 3.0926 (rank : 32) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8BY02, Q8BJU5, Q9CRL2 | Gene names | Nkrf, Nrf | |||
|
Domain Architecture |

|
|||||
| Description | NF-kappa-B-repressing factor (NFKB-repressing factor) (Transcription factor NRF). | |||||
|
NKRF_HUMAN
|
||||||
| NC score | 0.087083 (rank : 23) | θ value | 5.27518 (rank : 37) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O15226, Q9UJ91 | Gene names | NKRF, ITBA4, NRF | |||
|
Domain Architecture |

|
|||||
| Description | NF-kappa-B-repressing factor (NFKB-repressing factor) (Transcription factor NRF) (ITBA4 protein). | |||||
|
SON_MOUSE
|
||||||
| NC score | 0.086239 (rank : 24) | θ value | 0.279714 (rank : 10) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 957 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9QX47, Q9CQ12, Q9CQK6, Q9QXP5 | Gene names | Son | |||
|
Domain Architecture |

|
|||||
| Description | SON protein. | |||||
|
SAS10_HUMAN
|
||||||
| NC score | 0.067975 (rank : 25) | θ value | 5.27518 (rank : 38) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NQZ2, Q6FI82 | Gene names | CRLZ1, SAS10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Something about silencing protein 10 (Disrupter of silencing SAS10) (Charged amino acid-rich leucine zipper 1). | |||||
|
SAS10_MOUSE
|
||||||
| NC score | 0.060051 (rank : 26) | θ value | 8.99809 (rank : 50) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9JI13, Q3UL95, Q8R5C6, Q9JJ12 | Gene names | Crlz1, Crl1, Sas10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Something about silencing protein 10 (Disrupter of silencing SAS10) (Charged amino acid-rich leucine zipper 1) (Crl-1). | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.059098 (rank : 27) | θ value | 0.279714 (rank : 9) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
ERIC1_HUMAN
|
||||||
| NC score | 0.054284 (rank : 28) | θ value | 0.62314 (rank : 15) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q86X53, Q9P063 | Gene names | ERICH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate-rich protein 1. | |||||
|
SPF45_MOUSE
|
||||||
| NC score | 0.053825 (rank : 29) | θ value | 6.88961 (rank : 44) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8JZX4, O75939 | Gene names | Rbm17, Spf45 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 45 (45 kDa-splicing factor) (RNA-binding motif protein 17). | |||||
|
SPF45_HUMAN
|
||||||
| NC score | 0.052779 (rank : 30) | θ value | 6.88961 (rank : 43) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q96I25, Q96GY6 | Gene names | RBM17, SPF45 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 45 (45 kDa-splicing factor) (RNA-binding motif protein 17). | |||||
|
LEO1_HUMAN
|
||||||
| NC score | 0.042310 (rank : 31) | θ value | 0.47712 (rank : 12) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
INADL_MOUSE
|
||||||
| NC score | 0.039929 (rank : 32) | θ value | 0.0961366 (rank : 6) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q63ZW7, O70471, Q5PRG3, Q6P6J1, Q80YR8, Q8BPB9 | Gene names | Inadl, Cipp, Patj | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | InaD-like protein (Inadl protein) (Pals1-associated tight junction protein) (Protein associated to tight junctions) (Channel-interacting PDZ domain-containing protein). | |||||
|
NSBP1_MOUSE
|
||||||
| NC score | 0.037826 (rank : 33) | θ value | 2.36792 (rank : 28) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
LEO1_MOUSE
|
||||||
| NC score | 0.035326 (rank : 34) | θ value | 3.0926 (rank : 30) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
BASI_HUMAN
|
||||||
| NC score | 0.033529 (rank : 35) | θ value | 1.81305 (rank : 25) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P35613, Q7Z796, Q8IZL7 | Gene names | BSG | |||
|
Domain Architecture |

|
|||||
| Description | Basigin precursor (Leukocyte activation antigen M6) (Collagenase stimulatory factor) (Extracellular matrix metalloproteinase inducer) (EMMPRIN) (5F7) (Tumor cell-derived collagenase stimulatory factor) (TCSF) (OK blood group antigen) (CD147 antigen). | |||||
|
TCAL5_HUMAN
|
||||||
| NC score | 0.027986 (rank : 36) | θ value | 1.06291 (rank : 21) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 806 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q5H9L2 | Gene names | TCEAL5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 5 (TCEA-like protein 5) (Transcription elongation factor S-II protein-like 5). | |||||
|
ADPPT_MOUSE
|
||||||
| NC score | 0.027806 (rank : 37) | θ value | 8.99809 (rank : 45) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9CQF6, Q5U5W8, Q9CU40, Q9D827 | Gene names | Aasdhppt | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | L-aminoadipate-semialdehyde dehydrogenase-phosphopantetheinyl transferase (EC 2.7.8.-) (4'-phosphopantetheinyl transferase) (Alpha- aminoadipic semialdehyde dehydrogenase-phosphopantetheinyl transferase) (AASD-PPT). | |||||
|
ACINU_HUMAN
|
||||||
| NC score | 0.026155 (rank : 38) | θ value | 4.03905 (rank : 33) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UKV3, O75158, Q9UG91, Q9UKV1, Q9UKV2 | Gene names | ACIN1, ACINUS, KIAA0670 | |||
|
Domain Architecture |

|
|||||
| Description | Apoptotic chromatin condensation inducer in the nucleus (Acinus). | |||||
|
CK5P2_MOUSE
|
||||||
| NC score | 0.022095 (rank : 39) | θ value | 0.62314 (rank : 14) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1151 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8K389, Q6PCN1, Q6ZPL0 | Gene names | Cdk5rap2, Kiaa1633 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CDK5 regulatory subunit-associated protein 2 (CDK5 activator-binding protein C48). | |||||
|
MAP1A_HUMAN
|
||||||
| NC score | 0.021327 (rank : 40) | θ value | 3.0926 (rank : 31) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P78559, O95643, Q12973, Q15882, Q9UJT4 | Gene names | MAP1A | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 1A (MAP 1A) (Proliferation-related protein p80) [Contains: MAP1 light chain LC2]. | |||||
|
CLD19_HUMAN
|
||||||
| NC score | 0.021017 (rank : 41) | θ value | 0.47712 (rank : 11) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N6F1, Q8N8X0 | Gene names | CLDN19 | |||
|
Domain Architecture |

|
|||||
| Description | Claudin-19. | |||||
|
CK5P2_HUMAN
|
||||||
| NC score | 0.020360 (rank : 42) | θ value | 6.88961 (rank : 39) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1433 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q96SN8, Q7Z3L4, Q7Z3U1, Q7Z7I6, Q9BSW0, Q9H6J6, Q9HCD9, Q9NV90, Q9UIW9 | Gene names | CDK5RAP2, CEP215, KIAA1633 | |||
|
Domain Architecture |

|
|||||
| Description | CDK5 regulatory subunit-associated protein 2 (CDK5 activator-binding protein C48) (Centrosome-associated protein 215). | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.020206 (rank : 43) | θ value | 5.27518 (rank : 36) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
GANAB_HUMAN
|
||||||
| NC score | 0.019836 (rank : 44) | θ value | 2.36792 (rank : 27) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q14697, Q8WTS9, Q9P0X0 | Gene names | GANAB, G2AN, KIAA0088 | |||
|
Domain Architecture |

|
|||||
| Description | Neutral alpha-glucosidase AB precursor (EC 3.2.1.84) (Glucosidase II subunit alpha). | |||||
|
TGON2_MOUSE
|
||||||
| NC score | 0.016257 (rank : 45) | θ value | 8.99809 (rank : 52) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q62314 | Gene names | Tgoln2, Ttgn2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (TGN38B). | |||||
|
LAG3_HUMAN
|
||||||
| NC score | 0.014239 (rank : 46) | θ value | 8.99809 (rank : 46) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P18627 | Gene names | LAG3, FDC | |||
|
Domain Architecture |

|
|||||
| Description | Lymphocyte activation gene 3 protein precursor (LAG-3) (FDC protein) (CD223 antigen). | |||||
|
PACN2_MOUSE
|
||||||
| NC score | 0.012779 (rank : 47) | θ value | 6.88961 (rank : 40) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 382 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9WVE8 | Gene names | Pacsin2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C and casein kinase substrate in neurons protein 2. | |||||
|
MLL4_HUMAN
|
||||||
| NC score | 0.012087 (rank : 48) | θ value | 8.99809 (rank : 47) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UMN6, O15022, O95836, Q96GP2, Q96IP3, Q9UK25, Q9Y668, Q9Y669 | Gene names | MLL4, HRX2, KIAA0304, MLL2, TRX2 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 4 (Trithorax homolog 2). | |||||
|
ANK1_HUMAN
|
||||||
| NC score | 0.008393 (rank : 49) | θ value | 4.03905 (rank : 34) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P16157, O43400, Q13768, Q59FP2, Q8N604, Q99407 | Gene names | ANK1, ANK | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-1 (Erythrocyte ankyrin) (Ankyrin-R). | |||||
|
RAD_MOUSE
|
||||||
| NC score | 0.007962 (rank : 50) | θ value | 6.88961 (rank : 42) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O88667 | Gene names | Rrad | |||
|
Domain Architecture |

|
|||||
| Description | GTP-binding protein RAD. | |||||
|
RAD_HUMAN
|
||||||
| NC score | 0.006920 (rank : 51) | θ value | 8.99809 (rank : 48) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 224 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P55042, Q96F39 | Gene names | RRAD, RAD | |||
|
Domain Architecture |

|
|||||
| Description | GTP-binding protein RAD (RAS associated with diabetes) (RAD1). | |||||
|
SIM2_HUMAN
|
||||||
| NC score | 0.006805 (rank : 52) | θ value | 8.99809 (rank : 51) | |||
| Query Neighborhood Hits | 52 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q14190, O60766, Q15470, Q15471, Q15472, Q15473, Q16532 | Gene names | SIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Single-minded homolog 2. | |||||