Please be patient as the page loads
|
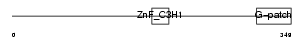
ZGPAT_HUMAN
|
||||||
| SwissProt Accessions | Q8N5A5, Q5JWI9, Q8NC55, Q96JI0, Q96JU4, Q9H401 | Gene names | ZGPAT, KIAA1847, ZC3H9, ZC3HDC9 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger CCCH-type with G patch domain protein (Zinc finger CCCH domain-containing protein 9). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
ZGPAT_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 97 | |
| SwissProt Accessions | Q8N5A5, Q5JWI9, Q8NC55, Q96JI0, Q96JU4, Q9H401 | Gene names | ZGPAT, KIAA1847, ZC3H9, ZC3HDC9 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger CCCH-type with G patch domain protein (Zinc finger CCCH domain-containing protein 9). | |||||
|
ZGPAT_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.975006 (rank : 2) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q8VDM1, Q8BWW2 | Gene names | Zgpat | |||
|
Domain Architecture |
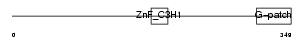
|
|||||
| Description | Zinc finger CCCH-type with G patch domain protein. | |||||
|
TFP11_HUMAN
|
||||||
| θ value | 2.61198e-07 (rank : 3) | NC score | 0.381002 (rank : 4) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9UBB9, O95908, Q5H8V8, Q9UGV7, Q9Y2Q8 | Gene names | TFIP11 | |||
|
Domain Architecture |

|
|||||
| Description | Tuftelin-interacting protein 11. | |||||
|
TFP11_MOUSE
|
||||||
| θ value | 2.61198e-07 (rank : 4) | NC score | 0.381523 (rank : 3) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9ERA6, Q8VD06 | Gene names | Tfip11, Tip39 | |||
|
Domain Architecture |

|
|||||
| Description | Tuftelin-interacting protein 11 (Tuftelin-interacting protein 39). | |||||
|
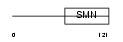
SPF30_HUMAN
|
||||||
| θ value | 4.1701e-05 (rank : 5) | NC score | 0.275203 (rank : 6) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O75940 | Gene names | SMNDC1, SMNR, SPF30 | |||
|
Domain Architecture |
|
|||||
| Description | Survival of motor neuron-related-splicing factor 30 (SMN-related protein) (30 kDa splicing factor SMNrp) (Survival motor neuron domain- containing protein 1). | |||||
|
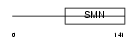
SPF30_MOUSE
|
||||||
| θ value | 4.1701e-05 (rank : 6) | NC score | 0.277805 (rank : 5) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BGT7, Q4QQL0 | Gene names | Smndc1, Smnr, Spf30 | |||
|
Domain Architecture |
|
|||||
| Description | Survival of motor neuron-related-splicing factor 30 (SMN-related protein) (30 kDa splicing factor SMNrp) (Survival motor neuron domain- containing protein 1). | |||||
|
RBM10_HUMAN
|
||||||
| θ value | 0.00298849 (rank : 7) | NC score | 0.225506 (rank : 9) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P98175, Q14136 | Gene names | RBM10, DXS8237E, KIAA0122 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 10 (RNA-binding motif protein 10). | |||||
|
GPTC2_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 8) | NC score | 0.242651 (rank : 7) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q7TQC7, Q8BNJ9, Q8BPM1, Q8CDH9 | Gene names | Gpatc2 | |||
|
Domain Architecture |

|
|||||
| Description | G patch domain-containing protein 2. | |||||
|
GPTC2_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 9) | NC score | 0.238104 (rank : 8) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9NW75, Q86YE7 | Gene names | GPATC2 | |||
|
Domain Architecture |

|
|||||
| Description | G patch domain-containing protein 2. | |||||
|
CCD13_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 10) | NC score | 0.058816 (rank : 54) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 721 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8IYE1 | Gene names | CCDC13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 13. | |||||
|
CGRE1_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 11) | NC score | 0.066211 (rank : 53) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q99674 | Gene names | CGREF1, CGR11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell growth regulator with EF hand domain 1 (Cell growth regulatory gene 11 protein). | |||||
|
SON_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 12) | NC score | 0.084671 (rank : 49) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
SON_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 13) | NC score | 0.123854 (rank : 38) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 957 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9QX47, Q9CQ12, Q9CQK6, Q9QXP5 | Gene names | Son | |||
|
Domain Architecture |

|
|||||
| Description | SON protein. | |||||
|
MKRN4_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 14) | NC score | 0.125482 (rank : 37) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q13434 | Gene names | MKRN4, RNF64, ZNF127L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Makorin-4 (Zinc finger protein 127-Xp) (ZNF127-Xp) (RING finger protein 64). | |||||
|
DHX57_HUMAN
|
||||||
| θ value | 0.125558 (rank : 15) | NC score | 0.055286 (rank : 55) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6P158, Q53SI4, Q6P9G1, Q7Z6H3, Q8NG17, Q96M33 | Gene names | DHX57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative ATP-dependent RNA helicase DHX57 (EC 3.6.1.-) (DEAH box protein 57). | |||||
|
RBM5_HUMAN
|
||||||
| θ value | 0.163984 (rank : 16) | NC score | 0.190585 (rank : 10) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P52756, Q93021, Q9BU14, Q9UKY8, Q9UL24 | Gene names | RBM5 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 5 (RNA-binding motif protein 5) (Tumor suppressor LUCA15) (Protein G15) (NY-REN-9 antigen). | |||||
|
RBM5_MOUSE
|
||||||
| θ value | 0.163984 (rank : 17) | NC score | 0.190332 (rank : 11) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q91YE7, Q3UZ19, Q99KV9 | Gene names | Rbm5, Luca15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein 5 (RNA-binding motif protein 5) (Tumor suppressor LUCA15). | |||||
|
MKRN2_HUMAN
|
||||||
| θ value | 0.47712 (rank : 18) | NC score | 0.113664 (rank : 40) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9H000, Q8N391, Q96BD4, Q9BUY2, Q9NRY1 | Gene names | MKRN2, RNF62 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-2 (RING finger protein 62). | |||||
|
RIMB1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 19) | NC score | 0.034491 (rank : 68) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q7TNF8, Q5NCP7, Q80TV9, Q8BIH5 | Gene names | Bzrap1, Kiaa0612, Rbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Peripheral-type benzodiazepine receptor-associated protein 1 (PRAX-1) (Peripheral benzodiazepine receptor-interacting protein) (PBR-IP) (RIM-binding protein 1) (RIM-BP1). | |||||
|
SFR14_MOUSE
|
||||||
| θ value | 0.47712 (rank : 20) | NC score | 0.147945 (rank : 14) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8CH09, Q6PG19, Q80UY8, Q8BY32, Q8CFM0 | Gene names | Sfrs14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative splicing factor, arginine/serine-rich 14 (Arginine/serine- rich-splicing factor 14). | |||||
|
BAT4_MOUSE
|
||||||
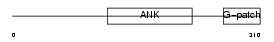
| θ value | 0.62314 (rank : 21) | NC score | 0.138118 (rank : 27) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q61858, Q9R065 | Gene names | Bat4, Bat-4, G5 | |||
|
Domain Architecture |
|
|||||
| Description | Protein BAT4 (HLA-B-associated transcript 4) (G5 protein). | |||||
|
K0082_MOUSE
|
||||||
| θ value | 0.62314 (rank : 22) | NC score | 0.130157 (rank : 34) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9DBC3, Q3U3G5, Q3U7Y9, Q6A0D5, Q8C7V0 | Gene names | Kiaa0082 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0082. | |||||
|
K1C14_HUMAN
|
||||||
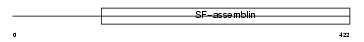
| θ value | 0.62314 (rank : 23) | NC score | 0.034008 (rank : 69) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P02533, Q14715, Q53XY3, Q9BUE3, Q9UBN2, Q9UBN3, Q9UCY4 | Gene names | KRT14 | |||
|
Domain Architecture |
|
|||||
| Description | Keratin, type I cytoskeletal 14 (Cytokeratin-14) (CK-14) (Keratin-14) (K14). | |||||
|
VPK3_HUMAN
|
||||||
| θ value | 0.62314 (rank : 24) | NC score | 0.138654 (rank : 26) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P63120 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19q12 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K(C19) Pro protein) (Protease) (Proteinase) (PR). | |||||
|
LIPA3_HUMAN
|
||||||
| θ value | 0.813845 (rank : 25) | NC score | 0.032805 (rank : 71) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O75145, Q9H8B5, Q9UEW4 | Gene names | PPFIA3, KIAA0654 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-3 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-3) (PTPRF-interacting protein alpha-3). | |||||
|
MKRN1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 26) | NC score | 0.108518 (rank : 43) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9UHC7, Q6GSF1, Q9H0G0, Q9UEZ7 | Gene names | MKRN1, RNF61 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-1 (RING finger protein 61). | |||||
|
VPK7_HUMAN
|
||||||
| θ value | 0.813845 (rank : 27) | NC score | 0.141916 (rank : 19) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P63123 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_1q23.3 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K18 Pro protein) (HERV-K110 Pro protein) (HERV-K(C1a) Pro protein) (Protease) (Proteinase) (PR). | |||||
|
CKAP4_HUMAN
|
||||||
| θ value | 1.06291 (rank : 28) | NC score | 0.042202 (rank : 60) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 592 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q07065, Q504S5, Q53ES6 | Gene names | CKAP4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-associated protein 4 (63 kDa membrane protein) (p63). | |||||
|
GPTC3_MOUSE
|
||||||
| θ value | 1.06291 (rank : 29) | NC score | 0.142913 (rank : 18) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8BIY1 | Gene names | Gpatc3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 3. | |||||
|
MKRN1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 30) | NC score | 0.106732 (rank : 44) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9QXP6 | Gene names | Mkrn1 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-1. | |||||
|
ZC3H6_MOUSE
|
||||||
| θ value | 1.06291 (rank : 31) | NC score | 0.085500 (rank : 48) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8BYK8, Q80UK3, Q8C1H1, Q9D604 | Gene names | Zc3h6, Zc3hdc6 | |||
|
Domain Architecture |
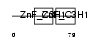
|
|||||
| Description | Zinc finger CCCH domain-containing protein 6. | |||||
|
K0082_HUMAN
|
||||||
| θ value | 1.38821 (rank : 32) | NC score | 0.134847 (rank : 30) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8N1G2, Q14670, Q96FJ9 | Gene names | KIAA0082 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0082. | |||||
|
K1C10_MOUSE
|
||||||
| θ value | 1.38821 (rank : 33) | NC score | 0.028315 (rank : 80) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 466 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P02535, P08731, Q8BUX3, Q8BV09, Q8BVU3, Q9CXH6, Q9JKB4 | Gene names | Krt10, Krt1-10 | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type I cytoskeletal 10 (Cytokeratin-10) (CK-10) (Keratin-10) (K10) (56 kDa cytokeratin) (Keratin, type I cytoskeletal 59 kDa). | |||||
|
K1C14_MOUSE
|
||||||
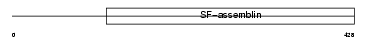
| θ value | 1.38821 (rank : 34) | NC score | 0.032076 (rank : 73) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 501 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q61781, Q91VQ4 | Gene names | Krt14, Krt1-14 | |||
|
Domain Architecture |
|
|||||
| Description | Keratin, type I cytoskeletal 14 (Cytokeratin-14) (CK-14) (Keratin-14) (K14). | |||||
|
MYH14_HUMAN
|
||||||
| θ value | 1.38821 (rank : 35) | NC score | 0.029590 (rank : 79) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 995 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q7Z406, Q5CZ75, Q6XYE4, Q76B62, Q8WV23, Q96I22, Q9BT27, Q9BW35, Q9H882 | Gene names | MYH14, KIAA2034 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-14 (Myosin heavy chain, nonmuscle IIc) (Nonmuscle myosin heavy chain IIc) (NMHC II-C). | |||||
|
VPK10_HUMAN
|
||||||
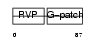
| θ value | 1.38821 (rank : 36) | NC score | 0.139237 (rank : 24) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P10265, Q9UKH6 | Gene names | ||||
|
Domain Architecture |
|
|||||
| Description | HERV-K_5q33.3 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K10 Pro protein) (HERV-K107 Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK12_HUMAN
|
||||||
| θ value | 1.38821 (rank : 37) | NC score | 0.139238 (rank : 23) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P63131, Q9UKI0 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_1q22 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K102 Pro protein) (HERV-K(III) Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 38) | NC score | 0.139232 (rank : 25) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P63119 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_12q14.1 provirus ancestral Pro protein (EC 3.4.23.-) (Protease) (Proteinase) (PR). | |||||
|
VPK5_HUMAN
|
||||||
| θ value | 1.38821 (rank : 39) | NC score | 0.139480 (rank : 22) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P63121 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19p13.11 provirus ancestral Pro protein (EC 3.4.23.-) (HERV- K113 envelope protein) (Protease) (Proteinase) (PR). | |||||
|
VPK9_HUMAN
|
||||||
| θ value | 1.38821 (rank : 40) | NC score | 0.139973 (rank : 21) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P63124 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_5q13.3 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K104 Pro protein) (Protease) (Proteinase) (PR). | |||||
|
ANR26_MOUSE
|
||||||
| θ value | 1.81305 (rank : 41) | NC score | 0.026547 (rank : 85) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 1554 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q811D2, Q69ZS2, Q9CS61 | Gene names | Ankrd26, Kiaa1074 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 26. | |||||
|
K1C10_HUMAN
|
||||||
| θ value | 1.81305 (rank : 42) | NC score | 0.028252 (rank : 82) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 526 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P13645, Q14664, Q8N175 | Gene names | KRT10 | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type I cytoskeletal 10 (Cytokeratin-10) (CK-10) (Keratin-10) (K10). | |||||
|
NKRF_HUMAN
|
||||||
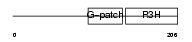
| θ value | 1.81305 (rank : 43) | NC score | 0.146565 (rank : 16) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O15226, Q9UJ91 | Gene names | NKRF, ITBA4, NRF | |||
|
Domain Architecture |
|
|||||
| Description | NF-kappa-B-repressing factor (NFKB-repressing factor) (Transcription factor NRF) (ITBA4 protein). | |||||
|
NKRF_MOUSE
|
||||||
| θ value | 1.81305 (rank : 44) | NC score | 0.147013 (rank : 15) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8BY02, Q8BJU5, Q9CRL2 | Gene names | Nkrf, Nrf | |||
|
Domain Architecture |

|
|||||
| Description | NF-kappa-B-repressing factor (NFKB-repressing factor) (Transcription factor NRF). | |||||
|
PINX1_HUMAN
|
||||||
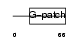
| θ value | 1.81305 (rank : 45) | NC score | 0.154820 (rank : 13) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q96BK5, Q7Z7J8, Q96QD7, Q9HBU7, Q9NWW2 | Gene names | PINX1, LPTS | |||
|
Domain Architecture |
|
|||||
| Description | Pin2-interacting protein X1 (TRF1-interacting protein 1) (Liver- related putative tumor suppressor) (67-11-3 protein). | |||||
|
PINX1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 46) | NC score | 0.170419 (rank : 12) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9CZX5, Q91WZ9, Q9D0C2 | Gene names | Pinx1, Lpts | |||
|
Domain Architecture |

|
|||||
| Description | Pin2-interacting protein X1 (TRF1-interacting protein 1) (Liver- related putative tumor suppressor) (LPTS1) (67-11-3 protein). | |||||
|
SFR14_HUMAN
|
||||||
| θ value | 1.81305 (rank : 47) | NC score | 0.143724 (rank : 17) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8IX01, O15071, O60369, Q5JPH7, Q8WUF7 | Gene names | SFRS14, KIAA0365 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative splicing factor, arginine/serine-rich 14 (Arginine/serine- rich-splicing factor 14). | |||||
|
VPK17_HUMAN
|
||||||
| θ value | 1.81305 (rank : 48) | NC score | 0.136948 (rank : 28) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P63125 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_11q22.1 provirus ancestral Pro protein (EC 3.4.23.-) (Protease) (Proteinase) (PR). | |||||
|
VPK2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 49) | NC score | 0.134668 (rank : 31) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9Y6I0, Q9UKH5 | Gene names | ERVK6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_7p22.1 provirus ancestral Pro protein (EC 3.4.23.-) (HERV- K(HML-2.HOM) Pro protein) (HERV-K108 Pro protein) (HERV-K(C7) Pro protein) (Protease) (Proteinase) (PR). | |||||
|
ZC3H6_HUMAN
|
||||||
| θ value | 1.81305 (rank : 50) | NC score | 0.080313 (rank : 50) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P61129 | Gene names | ZC3H6, KIAA2035, ZC3HDC6 | |||
|
Domain Architecture |
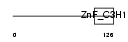
|
|||||
| Description | Zinc finger CCCH domain-containing protein 6. | |||||
|
BAT4_HUMAN
|
||||||
| θ value | 2.36792 (rank : 51) | NC score | 0.127393 (rank : 36) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O95872 | Gene names | BAT4, G5 | |||
|
Domain Architecture |
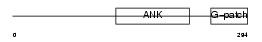
|
|||||
| Description | Protein BAT4 (HLA-B-associated transcript 4) (G5 protein). | |||||
|
GPTC3_HUMAN
|
||||||
| θ value | 2.36792 (rank : 52) | NC score | 0.121204 (rank : 39) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96I76, Q5JYH2, Q8NDJ2, Q9H9Z3 | Gene names | GPATC3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 3. | |||||
|
SF04_HUMAN
|
||||||
| θ value | 2.36792 (rank : 53) | NC score | 0.140578 (rank : 20) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8IWZ8, O60378, Q6P3X9, Q8TCQ4, Q8WWT4, Q8WWT5, Q9NTG3 | Gene names | SF4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor 4 (RNA-binding protein RBP). | |||||
|
SF04_MOUSE
|
||||||
| θ value | 2.36792 (rank : 54) | NC score | 0.135939 (rank : 29) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8CH02, Q8R094 | Gene names | Sf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor 4. | |||||
|
CD72_HUMAN
|
||||||
| θ value | 3.0926 (rank : 55) | NC score | 0.028310 (rank : 81) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P21854 | Gene names | CD72 | |||
|
Domain Architecture |

|
|||||
| Description | B-cell differentiation antigen CD72 (Lyb-2). | |||||
|
CP250_HUMAN
|
||||||
| θ value | 3.0926 (rank : 56) | NC score | 0.031154 (rank : 74) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 1805 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9BV73, O14812, O60588, Q9H450 | Gene names | CEP250, CEP2, CNAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Centrosome-associated protein CEP250 (Centrosomal protein 2) (Centrosomal Nek2-associated protein 1) (C-Nap1). | |||||
|
CRF_MOUSE
|
||||||
| θ value | 3.0926 (rank : 57) | NC score | 0.037265 (rank : 64) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CIT0 | Gene names | Crh | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Corticoliberin precursor (Corticotropin-releasing factor) (CRF) (Corticotropin-releasing hormone). | |||||
|
DHX57_MOUSE
|
||||||
| θ value | 3.0926 (rank : 58) | NC score | 0.040792 (rank : 61) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6P5D3, Q3TS93, Q6NZK4, Q6P1B4, Q8BI63, Q8BIA2, Q8R360 | Gene names | Dhx57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative ATP-dependent RNA helicase DHX57 (EC 3.6.1.-) (DEAH box protein 57). | |||||
|
K1C16_MOUSE
|
||||||
| θ value | 3.0926 (rank : 59) | NC score | 0.028224 (rank : 83) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 444 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9Z2K1 | Gene names | Krt16, Krt1-16 | |||
|
Domain Architecture |
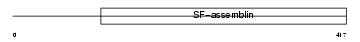
|
|||||
| Description | Keratin, type I cytoskeletal 16 (Cytokeratin-16) (CK-16) (Keratin-16) (K16). | |||||
|
MYH13_HUMAN
|
||||||
| θ value | 3.0926 (rank : 60) | NC score | 0.024894 (rank : 88) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 1504 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UKX3, O95252 | Gene names | MYH13 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-13 (Myosin heavy chain, skeletal muscle, extraocular) (MyHC- eo). | |||||
|
RUFY1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 61) | NC score | 0.026499 (rank : 86) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 765 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8BIJ7, Q8BKQ4, Q8BL21, Q9EPM6 | Gene names | Rufy1, Rabip4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and FYVE domain-containing protein 1 (Rab4-interacting protein). | |||||
|
SYTL2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 62) | NC score | 0.013005 (rank : 99) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 253 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99N50, Q8BT37, Q99J89, Q99J90, Q99N51, Q99N52, Q99N55, Q99N56 | Gene names | Sytl2, Slp2 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 2 (Exophilin-4). | |||||
|
ZO1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 63) | NC score | 0.011799 (rank : 101) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 453 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P39447 | Gene names | Tjp1, Zo1 | |||
|
Domain Architecture |

|
|||||
| Description | Tight junction protein ZO-1 (Zonula occludens 1 protein) (Zona occludens 1 protein) (Tight junction protein 1). | |||||
|
AGGF1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 64) | NC score | 0.132831 (rank : 33) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8N302, O00581, Q53YS3, Q9BU84, Q9NW66 | Gene names | AGGF1, VG5Q | |||
|
Domain Architecture |
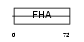
|
|||||
| Description | Angiogenic factor with G patch and FHA domains 1 (Angiogenic factor VG5Q) (Vasculogenesis gene on 5q protein) (hVG5Q). | |||||
|
AGGF1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 65) | NC score | 0.133956 (rank : 32) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q7TN31, Q8R2S6, Q9CQR9, Q9CU87, Q9D768 | Gene names | Aggf1, Vg5q | |||
|
Domain Architecture |
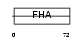
|
|||||
| Description | Angiogenic factor with G patch and FHA domains 1 (Angiogenic factor VG5Q) (mVG5Q). | |||||
|
CCD21_MOUSE
|
||||||
| θ value | 4.03905 (rank : 66) | NC score | 0.033863 (rank : 70) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8BMK0, Q8BUF1, Q8K0E6 | Gene names | Ccdc21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 21. | |||||
|
CS007_HUMAN
|
||||||
| θ value | 4.03905 (rank : 67) | NC score | 0.071379 (rank : 51) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UPT8, Q9Y420 | Gene names | C19orf7, KIAA1064 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein C19orf7. | |||||
|
CS007_MOUSE
|
||||||
| θ value | 4.03905 (rank : 68) | NC score | 0.068905 (rank : 52) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 464 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q6ZPZ3 | Gene names | Kiaa1064 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein C19orf7 homolog. | |||||
|
EVPL_MOUSE
|
||||||
| θ value | 4.03905 (rank : 69) | NC score | 0.024726 (rank : 90) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 952 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9D952 | Gene names | Evpl | |||
|
Domain Architecture |

|
|||||
| Description | Envoplakin (p210) (210 kDa cornified envelope precursor protein). | |||||
|
HIP1R_MOUSE
|
||||||
| θ value | 4.03905 (rank : 70) | NC score | 0.035984 (rank : 65) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 959 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9JKY5 | Gene names | Hip1r | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin-interacting protein 1-related protein (Hip1-related). | |||||
|
MKRN3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 71) | NC score | 0.094311 (rank : 45) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q13064 | Gene names | MKRN3, RNF63, ZNF127 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-3 (Zinc finger protein 127) (RING finger protein 63). | |||||
|
VPK11_HUMAN
|
||||||
| θ value | 4.03905 (rank : 72) | NC score | 0.127687 (rank : 35) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P63129, Q9UKI1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_22q11.21 provirus ancestral Pro protein (EC 3.4.23.-) (HERV- K101 envelope protein) (Protease) (Proteinase) (PR). | |||||
|
AMOL2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 73) | NC score | 0.024764 (rank : 89) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 591 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8K371, Q3TPM1, Q7TPE4, Q8BP84, Q8BS08, Q9QUS0 | Gene names | Amotl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin-like protein 2. | |||||
|
CROCC_HUMAN
|
||||||
| θ value | 5.27518 (rank : 74) | NC score | 0.032265 (rank : 72) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 1409 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q5TZA2, Q2VHY3, Q66GT7, Q7Z2L4, Q7Z5D7 | Gene names | CROCC, KIAA0445 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rootletin (Ciliary rootlet coiled-coil protein). | |||||
|
GOG8A_HUMAN
|
||||||
| θ value | 5.27518 (rank : 75) | NC score | 0.029652 (rank : 78) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 635 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9NZW0, O94937, Q9NZG8, Q9NZW3 | Gene names | GOLGA8A, KIAA0855 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgin subfamily A member 8A/B (Golgi autoantigen golgin-67) (88 kDa Golgi protein) (Gm88 autoantigen). | |||||
|
MKRN2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 76) | NC score | 0.090283 (rank : 47) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9ERV1, Q9D0L9 | Gene names | Mkrn2 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-2. | |||||
|
MKRN3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 77) | NC score | 0.092480 (rank : 46) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q60764, Q7TQE4 | Gene names | Mkrn3, Zfp127, Znf127 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-3 (Zinc finger protein 127). | |||||
|
RFX5_HUMAN
|
||||||
| θ value | 5.27518 (rank : 78) | NC score | 0.030122 (rank : 76) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P48382 | Gene names | RFX5 | |||
|
Domain Architecture |
|
|||||
| Description | DNA-binding protein RFX5 (Regulatory factor X subunit 5). | |||||
|
TDRD4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 79) | NC score | 0.039346 (rank : 63) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NUY9 | Gene names | TDRD4 | |||
|
Domain Architecture |

|
|||||
| Description | Tudor domain-containing protein 4. | |||||
|
CENPU_MOUSE
|
||||||
| θ value | 6.88961 (rank : 80) | NC score | 0.023082 (rank : 95) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8C4M7, Q6UNA2, Q9D9U1 | Gene names | Mlf1ip, Cenpu | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein U (CENP-U) (MLF1-interacting protein). | |||||
|
FYCO1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 81) | NC score | 0.030079 (rank : 77) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 1059 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8VDC1, Q3V385, Q7TMT0, Q8BJN2 | Gene names | Fyco1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE and coiled-coil domain-containing protein 1. | |||||
|
GOGA5_MOUSE
|
||||||
| θ value | 6.88961 (rank : 82) | NC score | 0.024527 (rank : 91) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 1434 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9QYE6, O88317, Q3TGE7, Q3U6S5, Q3UUF9 | Gene names | Golga5, Retii | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 5 (Golgin-84) (Sumiko protein) (Ret-II protein). | |||||
|
MYH11_HUMAN
|
||||||
| θ value | 6.88961 (rank : 83) | NC score | 0.023788 (rank : 93) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 1806 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P35749, O00396, O94944, P78422 | Gene names | MYH11, KIAA0866 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
NUCB1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 84) | NC score | 0.024300 (rank : 92) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q02818, Q15838, Q7Z4J7, Q9BUR1 | Gene names | NUCB1, NUC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleobindin-1 precursor (CALNUC). | |||||
|
PACN3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 85) | NC score | 0.012839 (rank : 100) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UKS6, Q9H331, Q9NWV9 | Gene names | PACSIN3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C and casein kinase substrate in neurons protein 3 (SH3 domain-containing protein 6511) (Endophilin I). | |||||
|
PAR12_MOUSE
|
||||||
| θ value | 6.88961 (rank : 86) | NC score | 0.035225 (rank : 66) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8BZ20, Q80VL6, Q8K333, Q8R3U2 | Gene names | Parp12, Zc3hdc1 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 12 (EC 2.4.2.30) (PARP-12) (Zinc finger CCCH domain-containing protein 1). | |||||
|
TPR_HUMAN
|
||||||
| θ value | 6.88961 (rank : 87) | NC score | 0.027851 (rank : 84) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
CP250_MOUSE
|
||||||
| θ value | 8.99809 (rank : 88) | NC score | 0.030967 (rank : 75) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 1583 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q60952, Q2I8G3, Q3UTR4, Q6PFF6, Q8BLC6 | Gene names | Cep250, Cep2, Inmp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome-associated protein CEP250 (Centrosomal protein 2) (Centrosomal Nek2-associated protein 1) (C-Nap1) (Intranuclear matrix protein). | |||||
|
FOSL2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 89) | NC score | 0.016056 (rank : 98) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P15408 | Gene names | FOSL2, FRA2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fos-related antigen 2. | |||||
|
FOSL2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 90) | NC score | 0.016802 (rank : 97) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P47930 | Gene names | Fosl2, Fra-2, Fra2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fos-related antigen 2. | |||||
|
KPCD_MOUSE
|
||||||
| θ value | 8.99809 (rank : 91) | NC score | -0.002050 (rank : 102) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 888 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P28867, Q91V85, Q9Z333 | Gene names | Prkcd, Pkcd | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C delta type (EC 2.7.11.13) (nPKC-delta). | |||||
|
MYH11_MOUSE
|
||||||
| θ value | 8.99809 (rank : 92) | NC score | 0.026244 (rank : 87) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 1777 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O08638, O08639, Q62462, Q64195 | Gene names | Myh11 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
NUPL2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 93) | NC score | 0.045349 (rank : 59) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O15504, Q49AE7, Q9BS49 | Gene names | NUPL2, CG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoporin-like 2 (NLP-1) (hCG1) (NUP42 homolog). | |||||
|
NUPL2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 94) | NC score | 0.040379 (rank : 62) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8CIC2, Q8BJX3, Q8BVP7, Q924S0 | Gene names | Nupl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoporin-like 2 (NLP-1). | |||||
|
ORC3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 95) | NC score | 0.022378 (rank : 96) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UBD5, Q13565, Q6IUY7, Q9UG44, Q9UNT6 | Gene names | ORC3L, LATHEO | |||
|
Domain Architecture |
|
|||||
| Description | Origin recognition complex subunit 3 (Origin recognition complex subunit Latheo). | |||||
|
ORC3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 96) | NC score | 0.023230 (rank : 94) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9JK30 | Gene names | Orc3l, Orc3 | |||
|
Domain Architecture |
|
|||||
| Description | Origin recognition complex subunit 3 (Origin recognition complex subunit Latheo). | |||||
|
PAR12_HUMAN
|
||||||
| θ value | 8.99809 (rank : 97) | NC score | 0.034617 (rank : 67) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9H0J9, Q9H610, Q9NP36, Q9NTI3 | Gene names | PARP12, ZC3HDC1 | |||
|
Domain Architecture |
|
|||||
| Description | Poly [ADP-ribose] polymerase 12 (EC 2.4.2.30) (PARP-12) (Zinc finger CCCH domain-containing protein 1). | |||||
|
RBM6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 98) | NC score | 0.055272 (rank : 56) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P78332, O60549, O75524 | Gene names | RBM6, DEF3 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 6 (RNA-binding motif protein 6) (RNA-binding protein DEF-3) (Lung cancer antigen NY-LU-12) (Protein G16). | |||||
|
VPK4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 99) | NC score | 0.108876 (rank : 42) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P63127, Q9UKH4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_6q14.1 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K109 Pro protein) (HERV-K(C6) Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.108900 (rank : 41) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P63122 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_8p23.1 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K115 Pro protein) (Protease) (Proteinase) (PR). | |||||
|
ZC3H8_HUMAN
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.050296 (rank : 58) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8N5P1, Q9BZ75 | Gene names | ZC3H8, ZC3HDC8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 8. | |||||
|
ZC3H8_MOUSE
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.051740 (rank : 57) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9JJ48, Q80X92 | Gene names | Zc3h8, Fliz1, Zc3hdc8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 8 (Fetal liver zinc finger protein 1). | |||||
|
ZGPAT_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 97 | |
| SwissProt Accessions | Q8N5A5, Q5JWI9, Q8NC55, Q96JI0, Q96JU4, Q9H401 | Gene names | ZGPAT, KIAA1847, ZC3H9, ZC3HDC9 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger CCCH-type with G patch domain protein (Zinc finger CCCH domain-containing protein 9). | |||||
|
ZGPAT_MOUSE
|
||||||
| NC score | 0.975006 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q8VDM1, Q8BWW2 | Gene names | Zgpat | |||
|
Domain Architecture |
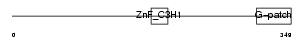
|
|||||
| Description | Zinc finger CCCH-type with G patch domain protein. | |||||
|
TFP11_MOUSE
|
||||||
| NC score | 0.381523 (rank : 3) | θ value | 2.61198e-07 (rank : 4) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9ERA6, Q8VD06 | Gene names | Tfip11, Tip39 | |||
|
Domain Architecture |

|
|||||
| Description | Tuftelin-interacting protein 11 (Tuftelin-interacting protein 39). | |||||
|
TFP11_HUMAN
|
||||||
| NC score | 0.381002 (rank : 4) | θ value | 2.61198e-07 (rank : 3) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9UBB9, O95908, Q5H8V8, Q9UGV7, Q9Y2Q8 | Gene names | TFIP11 | |||
|
Domain Architecture |

|
|||||
| Description | Tuftelin-interacting protein 11. | |||||
|
SPF30_MOUSE
|
||||||
| NC score | 0.277805 (rank : 5) | θ value | 4.1701e-05 (rank : 6) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BGT7, Q4QQL0 | Gene names | Smndc1, Smnr, Spf30 | |||
|
Domain Architecture |

|
|||||
| Description | Survival of motor neuron-related-splicing factor 30 (SMN-related protein) (30 kDa splicing factor SMNrp) (Survival motor neuron domain- containing protein 1). | |||||
|
SPF30_HUMAN
|
||||||
| NC score | 0.275203 (rank : 6) | θ value | 4.1701e-05 (rank : 5) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O75940 | Gene names | SMNDC1, SMNR, SPF30 | |||
|
Domain Architecture |

|
|||||
| Description | Survival of motor neuron-related-splicing factor 30 (SMN-related protein) (30 kDa splicing factor SMNrp) (Survival motor neuron domain- containing protein 1). | |||||
|
GPTC2_MOUSE
|
||||||
| NC score | 0.242651 (rank : 7) | θ value | 0.00390308 (rank : 8) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q7TQC7, Q8BNJ9, Q8BPM1, Q8CDH9 | Gene names | Gpatc2 | |||
|
Domain Architecture |

|
|||||
| Description | G patch domain-containing protein 2. | |||||
|
GPTC2_HUMAN
|
||||||
| NC score | 0.238104 (rank : 8) | θ value | 0.00869519 (rank : 9) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9NW75, Q86YE7 | Gene names | GPATC2 | |||
|
Domain Architecture |

|
|||||
| Description | G patch domain-containing protein 2. | |||||
|
RBM10_HUMAN
|
||||||
| NC score | 0.225506 (rank : 9) | θ value | 0.00298849 (rank : 7) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P98175, Q14136 | Gene names | RBM10, DXS8237E, KIAA0122 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 10 (RNA-binding motif protein 10). | |||||
|
RBM5_HUMAN
|
||||||
| NC score | 0.190585 (rank : 10) | θ value | 0.163984 (rank : 16) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P52756, Q93021, Q9BU14, Q9UKY8, Q9UL24 | Gene names | RBM5 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 5 (RNA-binding motif protein 5) (Tumor suppressor LUCA15) (Protein G15) (NY-REN-9 antigen). | |||||
|
RBM5_MOUSE
|
||||||
| NC score | 0.190332 (rank : 11) | θ value | 0.163984 (rank : 17) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q91YE7, Q3UZ19, Q99KV9 | Gene names | Rbm5, Luca15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein 5 (RNA-binding motif protein 5) (Tumor suppressor LUCA15). | |||||
|
PINX1_MOUSE
|
||||||
| NC score | 0.170419 (rank : 12) | θ value | 1.81305 (rank : 46) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9CZX5, Q91WZ9, Q9D0C2 | Gene names | Pinx1, Lpts | |||
|
Domain Architecture |

|
|||||
| Description | Pin2-interacting protein X1 (TRF1-interacting protein 1) (Liver- related putative tumor suppressor) (LPTS1) (67-11-3 protein). | |||||
|
PINX1_HUMAN
|
||||||
| NC score | 0.154820 (rank : 13) | θ value | 1.81305 (rank : 45) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q96BK5, Q7Z7J8, Q96QD7, Q9HBU7, Q9NWW2 | Gene names | PINX1, LPTS | |||
|
Domain Architecture |

|
|||||
| Description | Pin2-interacting protein X1 (TRF1-interacting protein 1) (Liver- related putative tumor suppressor) (67-11-3 protein). | |||||
|
SFR14_MOUSE
|
||||||
| NC score | 0.147945 (rank : 14) | θ value | 0.47712 (rank : 20) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8CH09, Q6PG19, Q80UY8, Q8BY32, Q8CFM0 | Gene names | Sfrs14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative splicing factor, arginine/serine-rich 14 (Arginine/serine- rich-splicing factor 14). | |||||
|
NKRF_MOUSE
|
||||||
| NC score | 0.147013 (rank : 15) | θ value | 1.81305 (rank : 44) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8BY02, Q8BJU5, Q9CRL2 | Gene names | Nkrf, Nrf | |||
|
Domain Architecture |

|
|||||
| Description | NF-kappa-B-repressing factor (NFKB-repressing factor) (Transcription factor NRF). | |||||
|
NKRF_HUMAN
|
||||||
| NC score | 0.146565 (rank : 16) | θ value | 1.81305 (rank : 43) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O15226, Q9UJ91 | Gene names | NKRF, ITBA4, NRF | |||
|
Domain Architecture |

|
|||||
| Description | NF-kappa-B-repressing factor (NFKB-repressing factor) (Transcription factor NRF) (ITBA4 protein). | |||||
|
SFR14_HUMAN
|
||||||
| NC score | 0.143724 (rank : 17) | θ value | 1.81305 (rank : 47) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8IX01, O15071, O60369, Q5JPH7, Q8WUF7 | Gene names | SFRS14, KIAA0365 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative splicing factor, arginine/serine-rich 14 (Arginine/serine- rich-splicing factor 14). | |||||
|
GPTC3_MOUSE
|
||||||
| NC score | 0.142913 (rank : 18) | θ value | 1.06291 (rank : 29) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8BIY1 | Gene names | Gpatc3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 3. | |||||
|
VPK7_HUMAN
|
||||||
| NC score | 0.141916 (rank : 19) | θ value | 0.813845 (rank : 27) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P63123 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_1q23.3 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K18 Pro protein) (HERV-K110 Pro protein) (HERV-K(C1a) Pro protein) (Protease) (Proteinase) (PR). | |||||
|
SF04_HUMAN
|
||||||
| NC score | 0.140578 (rank : 20) | θ value | 2.36792 (rank : 53) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8IWZ8, O60378, Q6P3X9, Q8TCQ4, Q8WWT4, Q8WWT5, Q9NTG3 | Gene names | SF4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor 4 (RNA-binding protein RBP). | |||||
|
VPK9_HUMAN
|
||||||
| NC score | 0.139973 (rank : 21) | θ value | 1.38821 (rank : 40) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P63124 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_5q13.3 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K104 Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK5_HUMAN
|
||||||
| NC score | 0.139480 (rank : 22) | θ value | 1.38821 (rank : 39) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P63121 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19p13.11 provirus ancestral Pro protein (EC 3.4.23.-) (HERV- K113 envelope protein) (Protease) (Proteinase) (PR). | |||||
|
VPK12_HUMAN
|
||||||
| NC score | 0.139238 (rank : 23) | θ value | 1.38821 (rank : 37) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P63131, Q9UKI0 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_1q22 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K102 Pro protein) (HERV-K(III) Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK10_HUMAN
|
||||||
| NC score | 0.139237 (rank : 24) | θ value | 1.38821 (rank : 36) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P10265, Q9UKH6 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | HERV-K_5q33.3 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K10 Pro protein) (HERV-K107 Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK1_HUMAN
|
||||||
| NC score | 0.139232 (rank : 25) | θ value | 1.38821 (rank : 38) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P63119 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_12q14.1 provirus ancestral Pro protein (EC 3.4.23.-) (Protease) (Proteinase) (PR). | |||||
|
VPK3_HUMAN
|
||||||
| NC score | 0.138654 (rank : 26) | θ value | 0.62314 (rank : 24) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P63120 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19q12 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K(C19) Pro protein) (Protease) (Proteinase) (PR). | |||||
|
BAT4_MOUSE
|
||||||
| NC score | 0.138118 (rank : 27) | θ value | 0.62314 (rank : 21) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q61858, Q9R065 | Gene names | Bat4, Bat-4, G5 | |||
|
Domain Architecture |

|
|||||
| Description | Protein BAT4 (HLA-B-associated transcript 4) (G5 protein). | |||||
|
VPK17_HUMAN
|
||||||
| NC score | 0.136948 (rank : 28) | θ value | 1.81305 (rank : 48) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P63125 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_11q22.1 provirus ancestral Pro protein (EC 3.4.23.-) (Protease) (Proteinase) (PR). | |||||
|
SF04_MOUSE
|
||||||
| NC score | 0.135939 (rank : 29) | θ value | 2.36792 (rank : 54) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8CH02, Q8R094 | Gene names | Sf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor 4. | |||||
|
K0082_HUMAN
|
||||||
| NC score | 0.134847 (rank : 30) | θ value | 1.38821 (rank : 32) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8N1G2, Q14670, Q96FJ9 | Gene names | KIAA0082 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0082. | |||||
|
VPK2_HUMAN
|
||||||
| NC score | 0.134668 (rank : 31) | θ value | 1.81305 (rank : 49) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9Y6I0, Q9UKH5 | Gene names | ERVK6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_7p22.1 provirus ancestral Pro protein (EC 3.4.23.-) (HERV- K(HML-2.HOM) Pro protein) (HERV-K108 Pro protein) (HERV-K(C7) Pro protein) (Protease) (Proteinase) (PR). | |||||
|
AGGF1_MOUSE
|
||||||
| NC score | 0.133956 (rank : 32) | θ value | 4.03905 (rank : 65) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q7TN31, Q8R2S6, Q9CQR9, Q9CU87, Q9D768 | Gene names | Aggf1, Vg5q | |||
|
Domain Architecture |

|
|||||
| Description | Angiogenic factor with G patch and FHA domains 1 (Angiogenic factor VG5Q) (mVG5Q). | |||||
|
AGGF1_HUMAN
|
||||||
| NC score | 0.132831 (rank : 33) | θ value | 4.03905 (rank : 64) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8N302, O00581, Q53YS3, Q9BU84, Q9NW66 | Gene names | AGGF1, VG5Q | |||
|
Domain Architecture |

|
|||||
| Description | Angiogenic factor with G patch and FHA domains 1 (Angiogenic factor VG5Q) (Vasculogenesis gene on 5q protein) (hVG5Q). | |||||
|
K0082_MOUSE
|
||||||
| NC score | 0.130157 (rank : 34) | θ value | 0.62314 (rank : 22) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9DBC3, Q3U3G5, Q3U7Y9, Q6A0D5, Q8C7V0 | Gene names | Kiaa0082 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0082. | |||||
|
VPK11_HUMAN
|
||||||
| NC score | 0.127687 (rank : 35) | θ value | 4.03905 (rank : 72) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P63129, Q9UKI1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_22q11.21 provirus ancestral Pro protein (EC 3.4.23.-) (HERV- K101 envelope protein) (Protease) (Proteinase) (PR). | |||||
|
BAT4_HUMAN
|
||||||
| NC score | 0.127393 (rank : 36) | θ value | 2.36792 (rank : 51) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O95872 | Gene names | BAT4, G5 | |||
|
Domain Architecture |

|
|||||
| Description | Protein BAT4 (HLA-B-associated transcript 4) (G5 protein). | |||||
|
MKRN4_HUMAN
|
||||||
| NC score | 0.125482 (rank : 37) | θ value | 0.0736092 (rank : 14) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q13434 | Gene names | MKRN4, RNF64, ZNF127L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Makorin-4 (Zinc finger protein 127-Xp) (ZNF127-Xp) (RING finger protein 64). | |||||
|
SON_MOUSE
|
||||||
| NC score | 0.123854 (rank : 38) | θ value | 0.0330416 (rank : 13) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 957 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9QX47, Q9CQ12, Q9CQK6, Q9QXP5 | Gene names | Son | |||
|
Domain Architecture |

|
|||||
| Description | SON protein. | |||||
|
GPTC3_HUMAN
|
||||||
| NC score | 0.121204 (rank : 39) | θ value | 2.36792 (rank : 52) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96I76, Q5JYH2, Q8NDJ2, Q9H9Z3 | Gene names | GPATC3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 3. | |||||
|
MKRN2_HUMAN
|
||||||
| NC score | 0.113664 (rank : 40) | θ value | 0.47712 (rank : 18) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9H000, Q8N391, Q96BD4, Q9BUY2, Q9NRY1 | Gene names | MKRN2, RNF62 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-2 (RING finger protein 62). | |||||
|
VPK6_HUMAN
|
||||||
| NC score | 0.108900 (rank : 41) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P63122 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_8p23.1 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K115 Pro protein) (Protease) (Proteinase) (PR). | |||||
|
VPK4_HUMAN
|
||||||
| NC score | 0.108876 (rank : 42) | θ value | θ > 10 (rank : 99) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P63127, Q9UKH4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_6q14.1 provirus ancestral Pro protein (EC 3.4.23.-) (HERV-K109 Pro protein) (HERV-K(C6) Pro protein) (Protease) (Proteinase) (PR). | |||||
|
MKRN1_HUMAN
|
||||||
| NC score | 0.108518 (rank : 43) | θ value | 0.813845 (rank : 26) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9UHC7, Q6GSF1, Q9H0G0, Q9UEZ7 | Gene names | MKRN1, RNF61 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-1 (RING finger protein 61). | |||||
|
MKRN1_MOUSE
|
||||||
| NC score | 0.106732 (rank : 44) | θ value | 1.06291 (rank : 30) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9QXP6 | Gene names | Mkrn1 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-1. | |||||
|
MKRN3_HUMAN
|
||||||
| NC score | 0.094311 (rank : 45) | θ value | 4.03905 (rank : 71) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q13064 | Gene names | MKRN3, RNF63, ZNF127 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-3 (Zinc finger protein 127) (RING finger protein 63). | |||||
|
MKRN3_MOUSE
|
||||||
| NC score | 0.092480 (rank : 46) | θ value | 5.27518 (rank : 77) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q60764, Q7TQE4 | Gene names | Mkrn3, Zfp127, Znf127 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-3 (Zinc finger protein 127). | |||||
|
MKRN2_MOUSE
|
||||||
| NC score | 0.090283 (rank : 47) | θ value | 5.27518 (rank : 76) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9ERV1, Q9D0L9 | Gene names | Mkrn2 | |||
|
Domain Architecture |

|
|||||
| Description | Makorin-2. | |||||
|
ZC3H6_MOUSE
|
||||||
| NC score | 0.085500 (rank : 48) | θ value | 1.06291 (rank : 31) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8BYK8, Q80UK3, Q8C1H1, Q9D604 | Gene names | Zc3h6, Zc3hdc6 | |||
|
Domain Architecture |
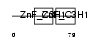
|
|||||
| Description | Zinc finger CCCH domain-containing protein 6. | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.084671 (rank : 49) | θ value | 0.0330416 (rank : 12) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
ZC3H6_HUMAN
|
||||||
| NC score | 0.080313 (rank : 50) | θ value | 1.81305 (rank : 50) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P61129 | Gene names | ZC3H6, KIAA2035, ZC3HDC6 | |||
|
Domain Architecture |
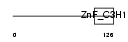
|
|||||
| Description | Zinc finger CCCH domain-containing protein 6. | |||||
|
CS007_HUMAN
|
||||||
| NC score | 0.071379 (rank : 51) | θ value | 4.03905 (rank : 67) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UPT8, Q9Y420 | Gene names | C19orf7, KIAA1064 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein C19orf7. | |||||
|
CS007_MOUSE
|
||||||
| NC score | 0.068905 (rank : 52) | θ value | 4.03905 (rank : 68) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 464 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q6ZPZ3 | Gene names | Kiaa1064 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein C19orf7 homolog. | |||||
|
CGRE1_HUMAN
|
||||||
| NC score | 0.066211 (rank : 53) | θ value | 0.0330416 (rank : 11) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q99674 | Gene names | CGREF1, CGR11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell growth regulator with EF hand domain 1 (Cell growth regulatory gene 11 protein). | |||||
|
CCD13_HUMAN
|
||||||
| NC score | 0.058816 (rank : 54) | θ value | 0.0330416 (rank : 10) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 721 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8IYE1 | Gene names | CCDC13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 13. | |||||
|
DHX57_HUMAN
|
||||||
| NC score | 0.055286 (rank : 55) | θ value | 0.125558 (rank : 15) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6P158, Q53SI4, Q6P9G1, Q7Z6H3, Q8NG17, Q96M33 | Gene names | DHX57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative ATP-dependent RNA helicase DHX57 (EC 3.6.1.-) (DEAH box protein 57). | |||||
|
RBM6_HUMAN
|
||||||
| NC score | 0.055272 (rank : 56) | θ value | θ > 10 (rank : 98) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P78332, O60549, O75524 | Gene names | RBM6, DEF3 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 6 (RNA-binding motif protein 6) (RNA-binding protein DEF-3) (Lung cancer antigen NY-LU-12) (Protein G16). | |||||
|
ZC3H8_MOUSE
|
||||||
| NC score | 0.051740 (rank : 57) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9JJ48, Q80X92 | Gene names | Zc3h8, Fliz1, Zc3hdc8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 8 (Fetal liver zinc finger protein 1). | |||||
|
ZC3H8_HUMAN
|
||||||
| NC score | 0.050296 (rank : 58) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8N5P1, Q9BZ75 | Gene names | ZC3H8, ZC3HDC8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 8. | |||||
|
NUPL2_HUMAN
|
||||||
| NC score | 0.045349 (rank : 59) | θ value | 8.99809 (rank : 93) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O15504, Q49AE7, Q9BS49 | Gene names | NUPL2, CG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoporin-like 2 (NLP-1) (hCG1) (NUP42 homolog). | |||||
|
CKAP4_HUMAN
|
||||||
| NC score | 0.042202 (rank : 60) | θ value | 1.06291 (rank : 28) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 592 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q07065, Q504S5, Q53ES6 | Gene names | CKAP4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-associated protein 4 (63 kDa membrane protein) (p63). | |||||
|
DHX57_MOUSE
|
||||||
| NC score | 0.040792 (rank : 61) | θ value | 3.0926 (rank : 58) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6P5D3, Q3TS93, Q6NZK4, Q6P1B4, Q8BI63, Q8BIA2, Q8R360 | Gene names | Dhx57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative ATP-dependent RNA helicase DHX57 (EC 3.6.1.-) (DEAH box protein 57). | |||||
|
NUPL2_MOUSE
|
||||||
| NC score | 0.040379 (rank : 62) | θ value | 8.99809 (rank : 94) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8CIC2, Q8BJX3, Q8BVP7, Q924S0 | Gene names | Nupl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleoporin-like 2 (NLP-1). | |||||
|
TDRD4_HUMAN
|
||||||
| NC score | 0.039346 (rank : 63) | θ value | 5.27518 (rank : 79) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NUY9 | Gene names | TDRD4 | |||
|
Domain Architecture |

|
|||||
| Description | Tudor domain-containing protein 4. | |||||
|
CRF_MOUSE
|
||||||
| NC score | 0.037265 (rank : 64) | θ value | 3.0926 (rank : 57) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CIT0 | Gene names | Crh | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Corticoliberin precursor (Corticotropin-releasing factor) (CRF) (Corticotropin-releasing hormone). | |||||
|
HIP1R_MOUSE
|
||||||
| NC score | 0.035984 (rank : 65) | θ value | 4.03905 (rank : 70) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 959 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9JKY5 | Gene names | Hip1r | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin-interacting protein 1-related protein (Hip1-related). | |||||
|
PAR12_MOUSE
|
||||||
| NC score | 0.035225 (rank : 66) | θ value | 6.88961 (rank : 86) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8BZ20, Q80VL6, Q8K333, Q8R3U2 | Gene names | Parp12, Zc3hdc1 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 12 (EC 2.4.2.30) (PARP-12) (Zinc finger CCCH domain-containing protein 1). | |||||
|
PAR12_HUMAN
|
||||||
| NC score | 0.034617 (rank : 67) | θ value | 8.99809 (rank : 97) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9H0J9, Q9H610, Q9NP36, Q9NTI3 | Gene names | PARP12, ZC3HDC1 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 12 (EC 2.4.2.30) (PARP-12) (Zinc finger CCCH domain-containing protein 1). | |||||
|
RIMB1_MOUSE
|
||||||
| NC score | 0.034491 (rank : 68) | θ value | 0.47712 (rank : 19) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q7TNF8, Q5NCP7, Q80TV9, Q8BIH5 | Gene names | Bzrap1, Kiaa0612, Rbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Peripheral-type benzodiazepine receptor-associated protein 1 (PRAX-1) (Peripheral benzodiazepine receptor-interacting protein) (PBR-IP) (RIM-binding protein 1) (RIM-BP1). | |||||
|
K1C14_HUMAN
|
||||||
| NC score | 0.034008 (rank : 69) | θ value | 0.62314 (rank : 23) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P02533, Q14715, Q53XY3, Q9BUE3, Q9UBN2, Q9UBN3, Q9UCY4 | Gene names | KRT14 | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type I cytoskeletal 14 (Cytokeratin-14) (CK-14) (Keratin-14) (K14). | |||||
|
CCD21_MOUSE
|
||||||
| NC score | 0.033863 (rank : 70) | θ value | 4.03905 (rank : 66) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8BMK0, Q8BUF1, Q8K0E6 | Gene names | Ccdc21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 21. | |||||
|
LIPA3_HUMAN
|
||||||
| NC score | 0.032805 (rank : 71) | θ value | 0.813845 (rank : 25) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O75145, Q9H8B5, Q9UEW4 | Gene names | PPFIA3, KIAA0654 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-3 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-3) (PTPRF-interacting protein alpha-3). | |||||
|
CROCC_HUMAN
|
||||||
| NC score | 0.032265 (rank : 72) | θ value | 5.27518 (rank : 74) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 1409 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q5TZA2, Q2VHY3, Q66GT7, Q7Z2L4, Q7Z5D7 | Gene names | CROCC, KIAA0445 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rootletin (Ciliary rootlet coiled-coil protein). | |||||
|
K1C14_MOUSE
|
||||||
| NC score | 0.032076 (rank : 73) | θ value | 1.38821 (rank : 34) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 501 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q61781, Q91VQ4 | Gene names | Krt14, Krt1-14 | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type I cytoskeletal 14 (Cytokeratin-14) (CK-14) (Keratin-14) (K14). | |||||
|
CP250_HUMAN
|
||||||
| NC score | 0.031154 (rank : 74) | θ value | 3.0926 (rank : 56) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 1805 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9BV73, O14812, O60588, Q9H450 | Gene names | CEP250, CEP2, CNAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Centrosome-associated protein CEP250 (Centrosomal protein 2) (Centrosomal Nek2-associated protein 1) (C-Nap1). | |||||
|
CP250_MOUSE
|
||||||
| NC score | 0.030967 (rank : 75) | θ value | 8.99809 (rank : 88) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 1583 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q60952, Q2I8G3, Q3UTR4, Q6PFF6, Q8BLC6 | Gene names | Cep250, Cep2, Inmp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome-associated protein CEP250 (Centrosomal protein 2) (Centrosomal Nek2-associated protein 1) (C-Nap1) (Intranuclear matrix protein). | |||||
|
RFX5_HUMAN
|
||||||
| NC score | 0.030122 (rank : 76) | θ value | 5.27518 (rank : 78) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P48382 | Gene names | RFX5 | |||
|
Domain Architecture |
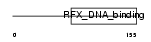
|
|||||
| Description | DNA-binding protein RFX5 (Regulatory factor X subunit 5). | |||||
|
FYCO1_MOUSE
|
||||||
| NC score | 0.030079 (rank : 77) | θ value | 6.88961 (rank : 81) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 1059 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8VDC1, Q3V385, Q7TMT0, Q8BJN2 | Gene names | Fyco1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE and coiled-coil domain-containing protein 1. | |||||
|
GOG8A_HUMAN
|
||||||
| NC score | 0.029652 (rank : 78) | θ value | 5.27518 (rank : 75) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 635 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9NZW0, O94937, Q9NZG8, Q9NZW3 | Gene names | GOLGA8A, KIAA0855 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgin subfamily A member 8A/B (Golgi autoantigen golgin-67) (88 kDa Golgi protein) (Gm88 autoantigen). | |||||
|
MYH14_HUMAN
|
||||||
| NC score | 0.029590 (rank : 79) | θ value | 1.38821 (rank : 35) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 995 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q7Z406, Q5CZ75, Q6XYE4, Q76B62, Q8WV23, Q96I22, Q9BT27, Q9BW35, Q9H882 | Gene names | MYH14, KIAA2034 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-14 (Myosin heavy chain, nonmuscle IIc) (Nonmuscle myosin heavy chain IIc) (NMHC II-C). | |||||
|
K1C10_MOUSE
|
||||||
| NC score | 0.028315 (rank : 80) | θ value | 1.38821 (rank : 33) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 466 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P02535, P08731, Q8BUX3, Q8BV09, Q8BVU3, Q9CXH6, Q9JKB4 | Gene names | Krt10, Krt1-10 | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type I cytoskeletal 10 (Cytokeratin-10) (CK-10) (Keratin-10) (K10) (56 kDa cytokeratin) (Keratin, type I cytoskeletal 59 kDa). | |||||
|
CD72_HUMAN
|
||||||
| NC score | 0.028310 (rank : 81) | θ value | 3.0926 (rank : 55) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P21854 | Gene names | CD72 | |||
|
Domain Architecture |

|
|||||
| Description | B-cell differentiation antigen CD72 (Lyb-2). | |||||
|
K1C10_HUMAN
|
||||||
| NC score | 0.028252 (rank : 82) | θ value | 1.81305 (rank : 42) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 526 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P13645, Q14664, Q8N175 | Gene names | KRT10 | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type I cytoskeletal 10 (Cytokeratin-10) (CK-10) (Keratin-10) (K10). | |||||
|
K1C16_MOUSE
|
||||||
| NC score | 0.028224 (rank : 83) | θ value | 3.0926 (rank : 59) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 444 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9Z2K1 | Gene names | Krt16, Krt1-16 | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type I cytoskeletal 16 (Cytokeratin-16) (CK-16) (Keratin-16) (K16). | |||||
|
TPR_HUMAN
|
||||||
| NC score | 0.027851 (rank : 84) | θ value | 6.88961 (rank : 87) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
ANR26_MOUSE
|
||||||
| NC score | 0.026547 (rank : 85) | θ value | 1.81305 (rank : 41) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 1554 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q811D2, Q69ZS2, Q9CS61 | Gene names | Ankrd26, Kiaa1074 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 26. | |||||
|
RUFY1_MOUSE
|
||||||
| NC score | 0.026499 (rank : 86) | θ value | 3.0926 (rank : 61) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 765 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8BIJ7, Q8BKQ4, Q8BL21, Q9EPM6 | Gene names | Rufy1, Rabip4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and FYVE domain-containing protein 1 (Rab4-interacting protein). | |||||
|
MYH11_MOUSE
|
||||||
| NC score | 0.026244 (rank : 87) | θ value | 8.99809 (rank : 92) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 1777 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O08638, O08639, Q62462, Q64195 | Gene names | Myh11 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
MYH13_HUMAN
|
||||||
| NC score | 0.024894 (rank : 88) | θ value | 3.0926 (rank : 60) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 1504 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UKX3, O95252 | Gene names | MYH13 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-13 (Myosin heavy chain, skeletal muscle, extraocular) (MyHC- eo). | |||||
|
AMOL2_MOUSE
|
||||||
| NC score | 0.024764 (rank : 89) | θ value | 5.27518 (rank : 73) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 591 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8K371, Q3TPM1, Q7TPE4, Q8BP84, Q8BS08, Q9QUS0 | Gene names | Amotl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin-like protein 2. | |||||
|
EVPL_MOUSE
|
||||||
| NC score | 0.024726 (rank : 90) | θ value | 4.03905 (rank : 69) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 952 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9D952 | Gene names | Evpl | |||
|
Domain Architecture |

|
|||||
| Description | Envoplakin (p210) (210 kDa cornified envelope precursor protein). | |||||
|
GOGA5_MOUSE
|
||||||
| NC score | 0.024527 (rank : 91) | θ value | 6.88961 (rank : 82) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 1434 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9QYE6, O88317, Q3TGE7, Q3U6S5, Q3UUF9 | Gene names | Golga5, Retii | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 5 (Golgin-84) (Sumiko protein) (Ret-II protein). | |||||
|
NUCB1_HUMAN
|
||||||
| NC score | 0.024300 (rank : 92) | θ value | 6.88961 (rank : 84) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q02818, Q15838, Q7Z4J7, Q9BUR1 | Gene names | NUCB1, NUC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleobindin-1 precursor (CALNUC). | |||||
|
MYH11_HUMAN
|
||||||
| NC score | 0.023788 (rank : 93) | θ value | 6.88961 (rank : 83) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 1806 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P35749, O00396, O94944, P78422 | Gene names | MYH11, KIAA0866 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
ORC3_MOUSE
|
||||||
| NC score | 0.023230 (rank : 94) | θ value | 8.99809 (rank : 96) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9JK30 | Gene names | Orc3l, Orc3 | |||
|
Domain Architecture |

|
|||||
| Description | Origin recognition complex subunit 3 (Origin recognition complex subunit Latheo). | |||||
|
CENPU_MOUSE
|
||||||
| NC score | 0.023082 (rank : 95) | θ value | 6.88961 (rank : 80) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8C4M7, Q6UNA2, Q9D9U1 | Gene names | Mlf1ip, Cenpu | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein U (CENP-U) (MLF1-interacting protein). | |||||
|
ORC3_HUMAN
|
||||||
| NC score | 0.022378 (rank : 96) | θ value | 8.99809 (rank : 95) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UBD5, Q13565, Q6IUY7, Q9UG44, Q9UNT6 | Gene names | ORC3L, LATHEO | |||
|
Domain Architecture |

|
|||||
| Description | Origin recognition complex subunit 3 (Origin recognition complex subunit Latheo). | |||||
|
FOSL2_MOUSE
|
||||||
| NC score | 0.016802 (rank : 97) | θ value | 8.99809 (rank : 90) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P47930 | Gene names | Fosl2, Fra-2, Fra2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fos-related antigen 2. | |||||
|
FOSL2_HUMAN
|
||||||
| NC score | 0.016056 (rank : 98) | θ value | 8.99809 (rank : 89) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P15408 | Gene names | FOSL2, FRA2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fos-related antigen 2. | |||||
|
SYTL2_MOUSE
|
||||||
| NC score | 0.013005 (rank : 99) | θ value | 3.0926 (rank : 62) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 253 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99N50, Q8BT37, Q99J89, Q99J90, Q99N51, Q99N52, Q99N55, Q99N56 | Gene names | Sytl2, Slp2 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 2 (Exophilin-4). | |||||
|
PACN3_HUMAN
|
||||||
| NC score | 0.012839 (rank : 100) | θ value | 6.88961 (rank : 85) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UKS6, Q9H331, Q9NWV9 | Gene names | PACSIN3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C and casein kinase substrate in neurons protein 3 (SH3 domain-containing protein 6511) (Endophilin I). | |||||
|
ZO1_MOUSE
|
||||||
| NC score | 0.011799 (rank : 101) | θ value | 3.0926 (rank : 63) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 453 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P39447 | Gene names | Tjp1, Zo1 | |||
|
Domain Architecture |

|
|||||
| Description | Tight junction protein ZO-1 (Zonula occludens 1 protein) (Zona occludens 1 protein) (Tight junction protein 1). | |||||
|
KPCD_MOUSE
|
||||||
| NC score | -0.002050 (rank : 102) | θ value | 8.99809 (rank : 91) | |||
| Query Neighborhood Hits | 97 | Target Neighborhood Hits | 888 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P28867, Q91V85, Q9Z333 | Gene names | Prkcd, Pkcd | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C delta type (EC 2.7.11.13) (nPKC-delta). | |||||