Please be patient as the page loads
|
ELK3_MOUSE
|
||||||
| SwissProt Accessions | P41971, P97747, Q62346 | Gene names | Elk3, Erp, Net | |||
|
Domain Architecture |
|
|||||
| Description | ETS domain-containing protein Elk-3 (ETS-related protein NET) (ETS- related protein ERP). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
ELK3_MOUSE
|
||||||
| θ value | 1.97579e-164 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | P41971, P97747, Q62346 | Gene names | Elk3, Erp, Net | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-3 (ETS-related protein NET) (ETS- related protein ERP). | |||||
|
ELK3_HUMAN
|
||||||
| θ value | 3.38586e-148 (rank : 2) | NC score | 0.997678 (rank : 2) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P41970, Q6FG57, Q6GU29, Q9UD17 | Gene names | ELK3, NET, SAP2 | |||
|
Domain Architecture |
|
|||||
| Description | ETS domain-containing protein Elk-3 (ETS-related protein NET) (ETS- related protein ERP) (SRF accessory protein 2) (SAP-2). | |||||
|
ELK4_HUMAN
|
||||||
| θ value | 8.79181e-80 (rank : 3) | NC score | 0.969171 (rank : 4) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P28324, P28323 | Gene names | ELK4, SAP1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-4 (Serum response factor accessory protein 1) (SAP-1). | |||||
|
ELK4_MOUSE
|
||||||
| θ value | 3.70236e-70 (rank : 4) | NC score | 0.977133 (rank : 3) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P41158 | Gene names | Elk4, Sap1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-4 (Serum response factor accessory protein 1) (SAP-1). | |||||
|
ELK1_MOUSE
|
||||||
| θ value | 1.2411e-41 (rank : 5) | NC score | 0.953667 (rank : 5) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P41969 | Gene names | Elk1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-1. | |||||
|
ELK1_HUMAN
|
||||||
| θ value | 1.79215e-40 (rank : 6) | NC score | 0.953017 (rank : 6) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P19419, O75606, O95058, Q969X8, Q9UJM4 | Gene names | ELK1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-1. | |||||
|
ETV4_HUMAN
|
||||||
| θ value | 4.73814e-25 (rank : 7) | NC score | 0.876710 (rank : 19) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P43268, Q96AW9 | Gene names | ETV4, E1AF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 4 (Adenovirus E1A enhancer-binding protein) (E1A-F). | |||||
|
ETV4_MOUSE
|
||||||
| θ value | 4.73814e-25 (rank : 8) | NC score | 0.877248 (rank : 18) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P28322 | Gene names | Etv4, Pea-3, Pea3 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 4 (Polyomavirus enhancer activator 3) (Protein PEA3). | |||||
|
ETV5_HUMAN
|
||||||
| θ value | 8.08199e-25 (rank : 9) | NC score | 0.869422 (rank : 24) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P41161 | Gene names | ETV5, ERM | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 5 (Ets-related protein ERM). | |||||
|
ETV5_MOUSE
|
||||||
| θ value | 1.80048e-24 (rank : 10) | NC score | 0.850077 (rank : 30) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9CXC9, Q3TG49, Q8C0F3, Q9JHB1 | Gene names | Etv5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 5. | |||||
|
ETV1_HUMAN
|
||||||
| θ value | 3.07116e-24 (rank : 11) | NC score | 0.874800 (rank : 21) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P50549, O75849, Q9UQ71, Q9Y636 | Gene names | ETV1, ER81 | |||
|
Domain Architecture |
|
|||||
| Description | ETS translocation variant 1 (ER81 protein). | |||||
|
ETV1_MOUSE
|
||||||
| θ value | 3.07116e-24 (rank : 12) | NC score | 0.875721 (rank : 20) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P41164 | Gene names | Etv1, Er81, Etsrp81 | |||
|
Domain Architecture |
|
|||||
| Description | ETS translocation variant 1 (ER81 protein). | |||||
|
FLI1_MOUSE
|
||||||
| θ value | 6.84181e-24 (rank : 13) | NC score | 0.882889 (rank : 15) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P26323 | Gene names | Fli1, Fli-1 | |||
|
Domain Architecture |
|
|||||
| Description | Friend leukemia integration 1 transcription factor (Retroviral integration site protein Fli-1). | |||||
|
FLI1_HUMAN
|
||||||
| θ value | 8.93572e-24 (rank : 14) | NC score | 0.881176 (rank : 16) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q01543, Q14319, Q92480, Q9UE07 | Gene names | FLI1 | |||
|
Domain Architecture |
|
|||||
| Description | Friend leukemia integration 1 transcription factor (Fli-1 proto- oncogene) (ERGB transcription factor). | |||||
|
ETS1_HUMAN
|
||||||
| θ value | 9.87957e-23 (rank : 15) | NC score | 0.890757 (rank : 11) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P14921, Q14278, Q16080 | Gene names | ETS1 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-1 protein (p54). | |||||
|
ETS1_MOUSE
|
||||||
| θ value | 9.87957e-23 (rank : 16) | NC score | 0.890637 (rank : 13) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P27577, Q61403 | Gene names | Ets1, Ets-1 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-1 protein (p54). | |||||
|
ETS2_HUMAN
|
||||||
| θ value | 1.29031e-22 (rank : 17) | NC score | 0.890863 (rank : 10) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P15036 | Gene names | ETS2 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-2 protein. | |||||
|
ETS2_MOUSE
|
||||||
| θ value | 1.29031e-22 (rank : 18) | NC score | 0.889583 (rank : 14) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P15037 | Gene names | Ets2 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-2 protein. | |||||
|
GABPA_HUMAN
|
||||||
| θ value | 6.40375e-22 (rank : 19) | NC score | 0.890735 (rank : 12) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q06546, Q12939 | Gene names | GABPA, E4TF1A | |||
|
Domain Architecture |
|
|||||
| Description | GA-binding protein alpha chain (GABP-subunit alpha) (Transcription factor E4TF1-60) (Nuclear respiratory factor 2 subunit alpha). | |||||
|
GABPA_MOUSE
|
||||||
| θ value | 6.40375e-22 (rank : 20) | NC score | 0.892166 (rank : 9) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q00422 | Gene names | Gabpa, E4tf1a | |||
|
Domain Architecture |
|
|||||
| Description | GA-binding protein alpha chain (GABP-subunit alpha). | |||||
|
ERG_HUMAN
|
||||||
| θ value | 4.58923e-20 (rank : 21) | NC score | 0.873928 (rank : 22) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P11308, Q16113 | Gene names | ERG | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ERG (Transforming protein ERG). | |||||
|
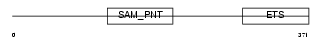
ERG_MOUSE
|
||||||
| θ value | 4.58923e-20 (rank : 22) | NC score | 0.873287 (rank : 23) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P81270, Q8C5L4, Q920K7, Q920K8, Q920K9 | Gene names | Erg, Erg-3 | |||
|
Domain Architecture |
|
|||||
| Description | Transcriptional regulator ERG. | |||||
|
ERF_HUMAN
|
||||||
| θ value | 7.82807e-20 (rank : 23) | NC score | 0.866106 (rank : 25) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P50548, Q9UPI7 | Gene names | ERF | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing transcription factor ERF (Ets2 repressor factor). | |||||
|
ERF_MOUSE
|
||||||
| θ value | 7.82807e-20 (rank : 24) | NC score | 0.865573 (rank : 26) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P70459 | Gene names | Erf | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing transcription factor ERF. | |||||
|
ETV2_HUMAN
|
||||||
| θ value | 2.97466e-19 (rank : 25) | NC score | 0.903376 (rank : 7) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O00321, Q9UEA0 | Gene names | ETV2, ER71, ETSRP71 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 2 (Ets-related protein 71). | |||||
|
ETV3_HUMAN
|
||||||
| θ value | 3.88503e-19 (rank : 26) | NC score | 0.834713 (rank : 36) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P41162, Q8TAC8, Q9BX30 | Gene names | ETV3, METS, PE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
ETV3_MOUSE
|
||||||
| θ value | 3.88503e-19 (rank : 27) | NC score | 0.860194 (rank : 29) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8R4Z4, Q9QZW1 | Gene names | Etv3, Mets, Pe1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
ETV2_MOUSE
|
||||||
| θ value | 8.65492e-19 (rank : 28) | NC score | 0.900661 (rank : 8) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P41163 | Gene names | Etv2, Er71, Etsrp71 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 2 (Ets-related protein 71). | |||||
|
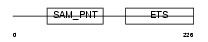
ETV7_HUMAN
|
||||||
| θ value | 2.7842e-17 (rank : 29) | NC score | 0.879345 (rank : 17) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9Y603, Q9NZ65, Q9NZ66, Q9NZ68, Q9NZR8, Q9UNJ7, Q9Y5K4, Q9Y604 | Gene names | ETV7, TEL2, TELB, TREF | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor ETV7 (Transcription factor Tel-2) (ETS-related protein Tel2) (Tel-related Ets factor). | |||||
|
ELF2_HUMAN
|
||||||
| θ value | 6.20254e-17 (rank : 30) | NC score | 0.839517 (rank : 33) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q15723, Q15724, Q15725, Q6P1K5 | Gene names | ELF2, NERF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-2 (E74-like factor 2) (New ETS- related factor). | |||||
|
ELF4_HUMAN
|
||||||
| θ value | 8.10077e-17 (rank : 31) | NC score | 0.835601 (rank : 35) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q99607, O60435 | Gene names | ELF4, ELFR, MEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-4 (E74-like factor 4) (Myeloid Elf-1-like factor). | |||||
|
ELF4_MOUSE
|
||||||
| θ value | 8.10077e-17 (rank : 32) | NC score | 0.813684 (rank : 41) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9Z2U4 | Gene names | Elf4, Mef | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-4 (E74-like factor 4) (Myeloid Elf-1-like factor). | |||||
|
ELF2_MOUSE
|
||||||
| θ value | 1.058e-16 (rank : 33) | NC score | 0.840427 (rank : 32) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9JHC9, Q6NST2, Q8BTX8, Q9JHC7, Q9JHC8, Q9JHD0 | Gene names | Elf2, Nerf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-2 (E74-like factor 2) (New ETS- related factor). | |||||
|
ETV6_HUMAN
|
||||||
| θ value | 3.07829e-16 (rank : 34) | NC score | 0.836908 (rank : 34) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P41212, Q9UMF6 | Gene names | ETV6, TEL, TEL1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor ETV6 (ETS-related protein Tel1) (Tel) (ETS translocation variant 6). | |||||
|
ETV6_MOUSE
|
||||||
| θ value | 3.07829e-16 (rank : 35) | NC score | 0.840648 (rank : 31) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P97360 | Gene names | Etv6, Tel, Tel1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor ETV6 (ETS-related protein Tel1) (Tel) (ETS translocation variant 6). | |||||
|
ELF1_HUMAN
|
||||||
| θ value | 4.02038e-16 (rank : 36) | NC score | 0.833822 (rank : 37) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P32519, Q9UDE1 | Gene names | ELF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-1 (E74-like factor 1). | |||||
|
ELF1_MOUSE
|
||||||
| θ value | 4.02038e-16 (rank : 37) | NC score | 0.826465 (rank : 38) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q60775 | Gene names | Elf1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS-related transcription factor Elf-1 (E74-like factor 1). | |||||
|
SPDEF_HUMAN
|
||||||
| θ value | 9.90251e-15 (rank : 38) | NC score | 0.864217 (rank : 27) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O95238 | Gene names | SPDEF, PDEF, PSE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SAM pointed domain-containing Ets transcription factor (Prostate- derived Ets factor) (Prostate epithelium-specific Ets transcription factor) (Prostate-specific Ets). | |||||
|
SPDEF_MOUSE
|
||||||
| θ value | 1.29331e-14 (rank : 39) | NC score | 0.862075 (rank : 28) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9WTP3 | Gene names | Spdef, Pdef, Pse | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SAM pointed domain-containing Ets transcription factor (Prostate- derived Ets factor) (Prostate epithelium-specific Ets transcription factor) (Prostate-specific Ets). | |||||
|
ELF5_HUMAN
|
||||||
| θ value | 1.33837e-11 (rank : 40) | NC score | 0.819778 (rank : 40) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9UKW6, O95175, Q8N2K9, Q96QY3, Q9UKW5 | Gene names | ELF5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-5 (E74-like factor 5) (Epithelium-specific Ets transcription factor 2) (ESE-2) (Epithelium- restricted ESE-1-related Ets factor). | |||||
|
ELF5_MOUSE
|
||||||
| θ value | 1.33837e-11 (rank : 41) | NC score | 0.820710 (rank : 39) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8VDK3, Q921H5, Q9Z2K6 | Gene names | Elf5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-5 (E74-like factor 5). | |||||
|
SPIB_HUMAN
|
||||||
| θ value | 0.00035302 (rank : 42) | NC score | 0.609557 (rank : 44) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q01892, Q15359 | Gene names | SPIB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor Spi-B. | |||||
|
SPI1_HUMAN
|
||||||
| θ value | 0.000786445 (rank : 43) | NC score | 0.618334 (rank : 43) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P17947 | Gene names | SPI1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor PU.1 (31 kDa transforming protein). | |||||
|
SPI1_MOUSE
|
||||||
| θ value | 0.000786445 (rank : 44) | NC score | 0.619619 (rank : 42) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P17433, Q99L57 | Gene names | Spi1, Sfpi-1, Sfpi1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor PU.1 (31 kDa transforming protein) (SFFV proviral integration 1 protein). | |||||
|
SPIC_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 45) | NC score | 0.569431 (rank : 46) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8N5J4 | Gene names | SPIC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor Spi-C. | |||||
|
SPIB_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 46) | NC score | 0.589283 (rank : 45) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O35906, O35907, O35909, O55199 | Gene names | Spib | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor Spi-B. | |||||
|
SPIC_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 47) | NC score | 0.535219 (rank : 47) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q6P3D7, Q9Z0Y0 | Gene names | Spic | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor Spi-C (Pu.1-related factor) (Prf). | |||||
|
BCAS1_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 48) | NC score | 0.039643 (rank : 48) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80YN3, Q9CVA1 | Gene names | Bcas1, Nabc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Breast carcinoma amplified sequence 1 homolog (Novel amplified in breast cancer 1 homolog). | |||||
|
MAP9_HUMAN
|
||||||
| θ value | 0.279714 (rank : 49) | NC score | 0.017020 (rank : 56) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 518 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q49MG5, Q4W5I7, Q68DU1, Q9H781, Q9H7B6 | Gene names | MAP9, ASAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 9 (Aster-associated protein). | |||||
|
MYCB2_MOUSE
|
||||||
| θ value | 0.279714 (rank : 50) | NC score | 0.019875 (rank : 54) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7TPH6, Q6PCM8 | Gene names | Mycbp2, Pam, Phr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ubiquitin ligase protein MYCBP2 (EC 6.3.2.-) (Myc-binding protein 2) (Protein associated with Myc) (Pam/highwire/rpm-1 protein). | |||||
|
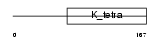
ZBTB3_HUMAN
|
||||||
| θ value | 0.279714 (rank : 51) | NC score | 0.002501 (rank : 70) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 910 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9H5J0 | Gene names | ZBTB3 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger and BTB domain-containing protein 3. | |||||
|
BCL9_HUMAN
|
||||||
| θ value | 0.62314 (rank : 52) | NC score | 0.024293 (rank : 52) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 1008 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O00512 | Gene names | BCL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9) (Legless homolog). | |||||
|
ALEX_HUMAN
|
||||||
| θ value | 0.813845 (rank : 53) | NC score | 0.030033 (rank : 50) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P84996 | Gene names | GNAS, GNAS1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ALEX (Alternative gene product encoded by XL-exon). | |||||
|
ALMS1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 54) | NC score | 0.017051 (rank : 55) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8K4E0, Q6A084, Q8C9N9 | Gene names | Alms1, Kiaa0328 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alstrom syndrome protein 1 homolog. | |||||
|
BCL9_MOUSE
|
||||||
| θ value | 1.81305 (rank : 55) | NC score | 0.020899 (rank : 53) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 770 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9D219, Q67FX9, Q8BUJ8, Q8VE74 | Gene names | Bcl9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9). | |||||
|
UROL1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 56) | NC score | 0.015867 (rank : 57) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5DID3, Q5DID1, Q5DID2 | Gene names | Umodl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uromodulin-like 1 precursor (Olfactorin). | |||||
|
ZN613_HUMAN
|
||||||
| θ value | 1.81305 (rank : 57) | NC score | -0.004245 (rank : 72) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 746 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6PF04, Q96SS9 | Gene names | ZNF613 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 613. | |||||
|
HEY2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 58) | NC score | 0.015843 (rank : 58) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9QUS4, Q3TZ99, Q8CD44 | Gene names | Hey2, Chf1, Herp, Herp1, Hesr2, Hrt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES-related repressor protein 2). | |||||
|
PTN22_MOUSE
|
||||||
| θ value | 2.36792 (rank : 59) | NC score | 0.009082 (rank : 62) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 179 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P29352, Q7TMP9 | Gene names | Ptpn22, Ptpn8 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 22 (EC 3.1.3.48) (Hematopoietic cell protein-tyrosine phosphatase 70Z-PEP). | |||||
|
ACHA4_MOUSE
|
||||||
| θ value | 3.0926 (rank : 60) | NC score | 0.003076 (rank : 69) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O70174, Q8BHE9, Q8VI10, Q9ET51 | Gene names | Chrna4, Acra4 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal acetylcholine receptor protein subunit alpha-4 precursor. | |||||
|
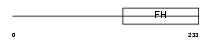
FOXK2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 61) | NC score | 0.005735 (rank : 65) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q01167, Q13622, Q13623, Q13624 | Gene names | FOXK2, ILF, ILF1 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein K2 (Interleukin enhancer-binding factor 1) (Cellular transcription factor ILF-1). | |||||
|
NEIL3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 62) | NC score | 0.013290 (rank : 59) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8TAT5, Q2PPJ3, Q8NG51, Q9NV95 | Gene names | NEIL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endonuclease VIII-like 3 (Nei-like 3) (DNA glycosylase FPG2). | |||||
|
ACD_HUMAN
|
||||||
| θ value | 4.03905 (rank : 63) | NC score | 0.030248 (rank : 49) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96AP0, Q562H5, Q9H8F9 | Gene names | ACD, PIP1, PTOP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Adrenocortical dysplasia protein homolog (POT1 and TIN2-interacting protein). | |||||
|
WDHD1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 64) | NC score | 0.004178 (rank : 67) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P59328, Q6P408 | Gene names | Wdhd1, And1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat and HMG-box DNA-binding protein 1 (Acidic nucleoplasmic DNA- binding protein 1) (And-1). | |||||
|
KCNQ5_HUMAN
|
||||||
| θ value | 5.27518 (rank : 65) | NC score | 0.003709 (rank : 68) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NR82, Q9NRN0, Q9NYA6 | Gene names | KCNQ5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 5 (Voltage-gated potassium channel subunit Kv7.5) (Potassium channel subunit alpha KvLQT5) (KQT-like 5). | |||||
|
RTN4_MOUSE
|
||||||
| θ value | 5.27518 (rank : 66) | NC score | 0.007109 (rank : 64) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99P72, Q5DTK9, Q7TNB7, Q80W95, Q8BGK7, Q8BGM9, Q8K290, Q8K3G8, Q9CTE3 | Gene names | Rtn4, Kiaa0886, Nogo | |||
|
Domain Architecture |

|
|||||
| Description | Reticulon-4 (Neurite outgrowth inhibitor) (Nogo protein). | |||||
|
CSMD3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 67) | NC score | 0.001178 (rank : 71) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7Z407, Q96PZ3 | Gene names | CSMD3, KIAA1894 | |||
|
Domain Architecture |

|
|||||
| Description | CUB and sushi domain-containing protein 3 precursor (CUB and sushi multiple domains protein 3). | |||||
|
MLL2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 68) | NC score | 0.013023 (rank : 60) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | 6.88961 (rank : 69) | NC score | 0.025566 (rank : 51) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
YETS2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 70) | NC score | 0.010455 (rank : 61) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9ULM3, Q641P6, Q9NW96 | Gene names | YEATS2, KIAA1197 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YEATS domain-containing protein 2. | |||||
|
HNF1B_HUMAN
|
||||||
| θ value | 8.99809 (rank : 71) | NC score | 0.005141 (rank : 66) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P35680 | Gene names | TCF2, HNF1B | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 1-beta (HNF-1beta) (HNF-1B) (Variant hepatic nuclear factor 1) (VHNF1) (Homeoprotein LFB3) (Transcription factor 2) (TCF-2). | |||||
|
TPP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 72) | NC score | 0.007332 (rank : 63) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O89023, Q543Q8, Q9QUS7 | Gene names | Tpp1, Cln2 | |||
|
Domain Architecture |

|
|||||
| Description | Tripeptidyl-peptidase 1 precursor (EC 3.4.14.9) (Tripeptidyl-peptidase I) (TPP-I) (Tripeptidyl aminopeptidase) (Lysosomal pepstatin insensitive protease) (LPIC). | |||||
|
ELK3_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 1.97579e-164 (rank : 1) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | P41971, P97747, Q62346 | Gene names | Elk3, Erp, Net | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-3 (ETS-related protein NET) (ETS- related protein ERP). | |||||
|
ELK3_HUMAN
|
||||||
| NC score | 0.997678 (rank : 2) | θ value | 3.38586e-148 (rank : 2) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P41970, Q6FG57, Q6GU29, Q9UD17 | Gene names | ELK3, NET, SAP2 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-3 (ETS-related protein NET) (ETS- related protein ERP) (SRF accessory protein 2) (SAP-2). | |||||
|
ELK4_MOUSE
|
||||||
| NC score | 0.977133 (rank : 3) | θ value | 3.70236e-70 (rank : 4) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P41158 | Gene names | Elk4, Sap1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-4 (Serum response factor accessory protein 1) (SAP-1). | |||||
|
ELK4_HUMAN
|
||||||
| NC score | 0.969171 (rank : 4) | θ value | 8.79181e-80 (rank : 3) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P28324, P28323 | Gene names | ELK4, SAP1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-4 (Serum response factor accessory protein 1) (SAP-1). | |||||
|
ELK1_MOUSE
|
||||||
| NC score | 0.953667 (rank : 5) | θ value | 1.2411e-41 (rank : 5) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P41969 | Gene names | Elk1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-1. | |||||
|
ELK1_HUMAN
|
||||||
| NC score | 0.953017 (rank : 6) | θ value | 1.79215e-40 (rank : 6) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P19419, O75606, O95058, Q969X8, Q9UJM4 | Gene names | ELK1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-1. | |||||
|
ETV2_HUMAN
|
||||||
| NC score | 0.903376 (rank : 7) | θ value | 2.97466e-19 (rank : 25) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O00321, Q9UEA0 | Gene names | ETV2, ER71, ETSRP71 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 2 (Ets-related protein 71). | |||||
|
ETV2_MOUSE
|
||||||
| NC score | 0.900661 (rank : 8) | θ value | 8.65492e-19 (rank : 28) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P41163 | Gene names | Etv2, Er71, Etsrp71 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 2 (Ets-related protein 71). | |||||
|
GABPA_MOUSE
|
||||||
| NC score | 0.892166 (rank : 9) | θ value | 6.40375e-22 (rank : 20) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q00422 | Gene names | Gabpa, E4tf1a | |||
|
Domain Architecture |

|
|||||
| Description | GA-binding protein alpha chain (GABP-subunit alpha). | |||||
|
ETS2_HUMAN
|
||||||
| NC score | 0.890863 (rank : 10) | θ value | 1.29031e-22 (rank : 17) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P15036 | Gene names | ETS2 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-2 protein. | |||||
|
ETS1_HUMAN
|
||||||
| NC score | 0.890757 (rank : 11) | θ value | 9.87957e-23 (rank : 15) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P14921, Q14278, Q16080 | Gene names | ETS1 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-1 protein (p54). | |||||
|
GABPA_HUMAN
|
||||||
| NC score | 0.890735 (rank : 12) | θ value | 6.40375e-22 (rank : 19) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q06546, Q12939 | Gene names | GABPA, E4TF1A | |||
|
Domain Architecture |

|
|||||
| Description | GA-binding protein alpha chain (GABP-subunit alpha) (Transcription factor E4TF1-60) (Nuclear respiratory factor 2 subunit alpha). | |||||
|
ETS1_MOUSE
|
||||||
| NC score | 0.890637 (rank : 13) | θ value | 9.87957e-23 (rank : 16) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P27577, Q61403 | Gene names | Ets1, Ets-1 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-1 protein (p54). | |||||
|
ETS2_MOUSE
|
||||||
| NC score | 0.889583 (rank : 14) | θ value | 1.29031e-22 (rank : 18) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P15037 | Gene names | Ets2 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-2 protein. | |||||
|
FLI1_MOUSE
|
||||||
| NC score | 0.882889 (rank : 15) | θ value | 6.84181e-24 (rank : 13) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P26323 | Gene names | Fli1, Fli-1 | |||
|
Domain Architecture |

|
|||||
| Description | Friend leukemia integration 1 transcription factor (Retroviral integration site protein Fli-1). | |||||
|
FLI1_HUMAN
|
||||||
| NC score | 0.881176 (rank : 16) | θ value | 8.93572e-24 (rank : 14) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q01543, Q14319, Q92480, Q9UE07 | Gene names | FLI1 | |||
|
Domain Architecture |

|
|||||
| Description | Friend leukemia integration 1 transcription factor (Fli-1 proto- oncogene) (ERGB transcription factor). | |||||
|
ETV7_HUMAN
|
||||||
| NC score | 0.879345 (rank : 17) | θ value | 2.7842e-17 (rank : 29) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9Y603, Q9NZ65, Q9NZ66, Q9NZ68, Q9NZR8, Q9UNJ7, Q9Y5K4, Q9Y604 | Gene names | ETV7, TEL2, TELB, TREF | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor ETV7 (Transcription factor Tel-2) (ETS-related protein Tel2) (Tel-related Ets factor). | |||||
|
ETV4_MOUSE
|
||||||
| NC score | 0.877248 (rank : 18) | θ value | 4.73814e-25 (rank : 8) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P28322 | Gene names | Etv4, Pea-3, Pea3 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 4 (Polyomavirus enhancer activator 3) (Protein PEA3). | |||||
|
ETV4_HUMAN
|
||||||
| NC score | 0.876710 (rank : 19) | θ value | 4.73814e-25 (rank : 7) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P43268, Q96AW9 | Gene names | ETV4, E1AF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 4 (Adenovirus E1A enhancer-binding protein) (E1A-F). | |||||
|
ETV1_MOUSE
|
||||||
| NC score | 0.875721 (rank : 20) | θ value | 3.07116e-24 (rank : 12) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P41164 | Gene names | Etv1, Er81, Etsrp81 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 1 (ER81 protein). | |||||
|
ETV1_HUMAN
|
||||||
| NC score | 0.874800 (rank : 21) | θ value | 3.07116e-24 (rank : 11) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P50549, O75849, Q9UQ71, Q9Y636 | Gene names | ETV1, ER81 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 1 (ER81 protein). | |||||
|
ERG_HUMAN
|
||||||
| NC score | 0.873928 (rank : 22) | θ value | 4.58923e-20 (rank : 21) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P11308, Q16113 | Gene names | ERG | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ERG (Transforming protein ERG). | |||||
|
ERG_MOUSE
|
||||||
| NC score | 0.873287 (rank : 23) | θ value | 4.58923e-20 (rank : 22) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P81270, Q8C5L4, Q920K7, Q920K8, Q920K9 | Gene names | Erg, Erg-3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ERG. | |||||
|
ETV5_HUMAN
|
||||||
| NC score | 0.869422 (rank : 24) | θ value | 8.08199e-25 (rank : 9) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P41161 | Gene names | ETV5, ERM | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 5 (Ets-related protein ERM). | |||||
|
ERF_HUMAN
|
||||||
| NC score | 0.866106 (rank : 25) | θ value | 7.82807e-20 (rank : 23) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P50548, Q9UPI7 | Gene names | ERF | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing transcription factor ERF (Ets2 repressor factor). | |||||
|
ERF_MOUSE
|
||||||
| NC score | 0.865573 (rank : 26) | θ value | 7.82807e-20 (rank : 24) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P70459 | Gene names | Erf | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing transcription factor ERF. | |||||
|
SPDEF_HUMAN
|
||||||
| NC score | 0.864217 (rank : 27) | θ value | 9.90251e-15 (rank : 38) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O95238 | Gene names | SPDEF, PDEF, PSE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SAM pointed domain-containing Ets transcription factor (Prostate- derived Ets factor) (Prostate epithelium-specific Ets transcription factor) (Prostate-specific Ets). | |||||
|
SPDEF_MOUSE
|
||||||
| NC score | 0.862075 (rank : 28) | θ value | 1.29331e-14 (rank : 39) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9WTP3 | Gene names | Spdef, Pdef, Pse | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SAM pointed domain-containing Ets transcription factor (Prostate- derived Ets factor) (Prostate epithelium-specific Ets transcription factor) (Prostate-specific Ets). | |||||
|
ETV3_MOUSE
|
||||||
| NC score | 0.860194 (rank : 29) | θ value | 3.88503e-19 (rank : 27) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8R4Z4, Q9QZW1 | Gene names | Etv3, Mets, Pe1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
ETV5_MOUSE
|
||||||
| NC score | 0.850077 (rank : 30) | θ value | 1.80048e-24 (rank : 10) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9CXC9, Q3TG49, Q8C0F3, Q9JHB1 | Gene names | Etv5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 5. | |||||
|
ETV6_MOUSE
|
||||||
| NC score | 0.840648 (rank : 31) | θ value | 3.07829e-16 (rank : 35) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P97360 | Gene names | Etv6, Tel, Tel1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor ETV6 (ETS-related protein Tel1) (Tel) (ETS translocation variant 6). | |||||
|
ELF2_MOUSE
|
||||||
| NC score | 0.840427 (rank : 32) | θ value | 1.058e-16 (rank : 33) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9JHC9, Q6NST2, Q8BTX8, Q9JHC7, Q9JHC8, Q9JHD0 | Gene names | Elf2, Nerf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-2 (E74-like factor 2) (New ETS- related factor). | |||||
|
ELF2_HUMAN
|
||||||
| NC score | 0.839517 (rank : 33) | θ value | 6.20254e-17 (rank : 30) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q15723, Q15724, Q15725, Q6P1K5 | Gene names | ELF2, NERF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-2 (E74-like factor 2) (New ETS- related factor). | |||||
|
ETV6_HUMAN
|
||||||
| NC score | 0.836908 (rank : 34) | θ value | 3.07829e-16 (rank : 34) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P41212, Q9UMF6 | Gene names | ETV6, TEL, TEL1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor ETV6 (ETS-related protein Tel1) (Tel) (ETS translocation variant 6). | |||||
|
ELF4_HUMAN
|
||||||
| NC score | 0.835601 (rank : 35) | θ value | 8.10077e-17 (rank : 31) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q99607, O60435 | Gene names | ELF4, ELFR, MEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-4 (E74-like factor 4) (Myeloid Elf-1-like factor). | |||||
|
ETV3_HUMAN
|
||||||
| NC score | 0.834713 (rank : 36) | θ value | 3.88503e-19 (rank : 26) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P41162, Q8TAC8, Q9BX30 | Gene names | ETV3, METS, PE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
ELF1_HUMAN
|
||||||
| NC score | 0.833822 (rank : 37) | θ value | 4.02038e-16 (rank : 36) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P32519, Q9UDE1 | Gene names | ELF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-1 (E74-like factor 1). | |||||
|
ELF1_MOUSE
|
||||||
| NC score | 0.826465 (rank : 38) | θ value | 4.02038e-16 (rank : 37) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q60775 | Gene names | Elf1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS-related transcription factor Elf-1 (E74-like factor 1). | |||||
|
ELF5_MOUSE
|
||||||
| NC score | 0.820710 (rank : 39) | θ value | 1.33837e-11 (rank : 41) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8VDK3, Q921H5, Q9Z2K6 | Gene names | Elf5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-5 (E74-like factor 5). | |||||
|
ELF5_HUMAN
|
||||||
| NC score | 0.819778 (rank : 40) | θ value | 1.33837e-11 (rank : 40) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9UKW6, O95175, Q8N2K9, Q96QY3, Q9UKW5 | Gene names | ELF5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-5 (E74-like factor 5) (Epithelium-specific Ets transcription factor 2) (ESE-2) (Epithelium- restricted ESE-1-related Ets factor). | |||||
|
ELF4_MOUSE
|
||||||
| NC score | 0.813684 (rank : 41) | θ value | 8.10077e-17 (rank : 32) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9Z2U4 | Gene names | Elf4, Mef | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-4 (E74-like factor 4) (Myeloid Elf-1-like factor). | |||||
|
SPI1_MOUSE
|
||||||
| NC score | 0.619619 (rank : 42) | θ value | 0.000786445 (rank : 44) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P17433, Q99L57 | Gene names | Spi1, Sfpi-1, Sfpi1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor PU.1 (31 kDa transforming protein) (SFFV proviral integration 1 protein). | |||||
|
SPI1_HUMAN
|
||||||
| NC score | 0.618334 (rank : 43) | θ value | 0.000786445 (rank : 43) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P17947 | Gene names | SPI1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor PU.1 (31 kDa transforming protein). | |||||
|
SPIB_HUMAN
|
||||||
| NC score | 0.609557 (rank : 44) | θ value | 0.00035302 (rank : 42) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q01892, Q15359 | Gene names | SPIB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor Spi-B. | |||||
|
SPIB_MOUSE
|
||||||
| NC score | 0.589283 (rank : 45) | θ value | 0.0431538 (rank : 46) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O35906, O35907, O35909, O55199 | Gene names | Spib | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor Spi-B. | |||||
|
SPIC_HUMAN
|
||||||
| NC score | 0.569431 (rank : 46) | θ value | 0.00869519 (rank : 45) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8N5J4 | Gene names | SPIC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor Spi-C. | |||||
|
SPIC_MOUSE
|
||||||
| NC score | 0.535219 (rank : 47) | θ value | 0.0431538 (rank : 47) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q6P3D7, Q9Z0Y0 | Gene names | Spic | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor Spi-C (Pu.1-related factor) (Prf). | |||||
|
BCAS1_MOUSE
|
||||||
| NC score | 0.039643 (rank : 48) | θ value | 0.0961366 (rank : 48) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q80YN3, Q9CVA1 | Gene names | Bcas1, Nabc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Breast carcinoma amplified sequence 1 homolog (Novel amplified in breast cancer 1 homolog). | |||||
|
ACD_HUMAN
|
||||||
| NC score | 0.030248 (rank : 49) | θ value | 4.03905 (rank : 63) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96AP0, Q562H5, Q9H8F9 | Gene names | ACD, PIP1, PTOP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Adrenocortical dysplasia protein homolog (POT1 and TIN2-interacting protein). | |||||
|
ALEX_HUMAN
|
||||||
| NC score | 0.030033 (rank : 50) | θ value | 0.813845 (rank : 53) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P84996 | Gene names | GNAS, GNAS1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ALEX (Alternative gene product encoded by XL-exon). | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.025566 (rank : 51) | θ value | 6.88961 (rank : 69) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
BCL9_HUMAN
|
||||||
| NC score | 0.024293 (rank : 52) | θ value | 0.62314 (rank : 52) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 1008 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O00512 | Gene names | BCL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9) (Legless homolog). | |||||
|
BCL9_MOUSE
|
||||||
| NC score | 0.020899 (rank : 53) | θ value | 1.81305 (rank : 55) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 770 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9D219, Q67FX9, Q8BUJ8, Q8VE74 | Gene names | Bcl9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9). | |||||
|
MYCB2_MOUSE
|
||||||
| NC score | 0.019875 (rank : 54) | θ value | 0.279714 (rank : 50) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7TPH6, Q6PCM8 | Gene names | Mycbp2, Pam, Phr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ubiquitin ligase protein MYCBP2 (EC 6.3.2.-) (Myc-binding protein 2) (Protein associated with Myc) (Pam/highwire/rpm-1 protein). | |||||
|
ALMS1_MOUSE
|
||||||
| NC score | 0.017051 (rank : 55) | θ value | 1.38821 (rank : 54) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8K4E0, Q6A084, Q8C9N9 | Gene names | Alms1, Kiaa0328 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alstrom syndrome protein 1 homolog. | |||||
|
MAP9_HUMAN
|
||||||
| NC score | 0.017020 (rank : 56) | θ value | 0.279714 (rank : 49) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 518 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q49MG5, Q4W5I7, Q68DU1, Q9H781, Q9H7B6 | Gene names | MAP9, ASAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 9 (Aster-associated protein). | |||||
|
UROL1_MOUSE
|
||||||
| NC score | 0.015867 (rank : 57) | θ value | 1.81305 (rank : 56) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5DID3, Q5DID1, Q5DID2 | Gene names | Umodl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uromodulin-like 1 precursor (Olfactorin). | |||||
|
HEY2_MOUSE
|
||||||
| NC score | 0.015843 (rank : 58) | θ value | 2.36792 (rank : 58) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9QUS4, Q3TZ99, Q8CD44 | Gene names | Hey2, Chf1, Herp, Herp1, Hesr2, Hrt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hairy/enhancer-of-split related with YRPW motif 2 (Hairy and enhancer of split-related protein 2) (HESR-2) (Hairy-related transcription factor 2) (HES-related repressor protein 2). | |||||
|
NEIL3_HUMAN
|
||||||
| NC score | 0.013290 (rank : 59) | θ value | 3.0926 (rank : 62) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8TAT5, Q2PPJ3, Q8NG51, Q9NV95 | Gene names | NEIL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endonuclease VIII-like 3 (Nei-like 3) (DNA glycosylase FPG2). | |||||
|
MLL2_HUMAN
|
||||||
| NC score | 0.013023 (rank : 60) | θ value | 6.88961 (rank : 68) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
YETS2_HUMAN
|
||||||
| NC score | 0.010455 (rank : 61) | θ value | 6.88961 (rank : 70) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9ULM3, Q641P6, Q9NW96 | Gene names | YEATS2, KIAA1197 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YEATS domain-containing protein 2. | |||||
|
PTN22_MOUSE
|
||||||
| NC score | 0.009082 (rank : 62) | θ value | 2.36792 (rank : 59) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 179 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P29352, Q7TMP9 | Gene names | Ptpn22, Ptpn8 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 22 (EC 3.1.3.48) (Hematopoietic cell protein-tyrosine phosphatase 70Z-PEP). | |||||
|
TPP1_MOUSE
|
||||||
| NC score | 0.007332 (rank : 63) | θ value | 8.99809 (rank : 72) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O89023, Q543Q8, Q9QUS7 | Gene names | Tpp1, Cln2 | |||
|
Domain Architecture |

|
|||||
| Description | Tripeptidyl-peptidase 1 precursor (EC 3.4.14.9) (Tripeptidyl-peptidase I) (TPP-I) (Tripeptidyl aminopeptidase) (Lysosomal pepstatin insensitive protease) (LPIC). | |||||
|
RTN4_MOUSE
|
||||||
| NC score | 0.007109 (rank : 64) | θ value | 5.27518 (rank : 66) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99P72, Q5DTK9, Q7TNB7, Q80W95, Q8BGK7, Q8BGM9, Q8K290, Q8K3G8, Q9CTE3 | Gene names | Rtn4, Kiaa0886, Nogo | |||
|
Domain Architecture |

|
|||||
| Description | Reticulon-4 (Neurite outgrowth inhibitor) (Nogo protein). | |||||
|
FOXK2_HUMAN
|
||||||
| NC score | 0.005735 (rank : 65) | θ value | 3.0926 (rank : 61) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q01167, Q13622, Q13623, Q13624 | Gene names | FOXK2, ILF, ILF1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein K2 (Interleukin enhancer-binding factor 1) (Cellular transcription factor ILF-1). | |||||
|
HNF1B_HUMAN
|
||||||
| NC score | 0.005141 (rank : 66) | θ value | 8.99809 (rank : 71) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P35680 | Gene names | TCF2, HNF1B | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 1-beta (HNF-1beta) (HNF-1B) (Variant hepatic nuclear factor 1) (VHNF1) (Homeoprotein LFB3) (Transcription factor 2) (TCF-2). | |||||
|
WDHD1_MOUSE
|
||||||
| NC score | 0.004178 (rank : 67) | θ value | 4.03905 (rank : 64) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P59328, Q6P408 | Gene names | Wdhd1, And1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat and HMG-box DNA-binding protein 1 (Acidic nucleoplasmic DNA- binding protein 1) (And-1). | |||||
|
KCNQ5_HUMAN
|
||||||
| NC score | 0.003709 (rank : 68) | θ value | 5.27518 (rank : 65) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NR82, Q9NRN0, Q9NYA6 | Gene names | KCNQ5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 5 (Voltage-gated potassium channel subunit Kv7.5) (Potassium channel subunit alpha KvLQT5) (KQT-like 5). | |||||
|
ACHA4_MOUSE
|
||||||
| NC score | 0.003076 (rank : 69) | θ value | 3.0926 (rank : 60) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O70174, Q8BHE9, Q8VI10, Q9ET51 | Gene names | Chrna4, Acra4 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal acetylcholine receptor protein subunit alpha-4 precursor. | |||||
|
ZBTB3_HUMAN
|
||||||
| NC score | 0.002501 (rank : 70) | θ value | 0.279714 (rank : 51) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 910 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9H5J0 | Gene names | ZBTB3 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 3. | |||||
|
CSMD3_HUMAN
|
||||||
| NC score | 0.001178 (rank : 71) | θ value | 6.88961 (rank : 67) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7Z407, Q96PZ3 | Gene names | CSMD3, KIAA1894 | |||
|
Domain Architecture |

|
|||||
| Description | CUB and sushi domain-containing protein 3 precursor (CUB and sushi multiple domains protein 3). | |||||
|
ZN613_HUMAN
|
||||||
| NC score | -0.004245 (rank : 72) | θ value | 1.81305 (rank : 57) | |||
| Query Neighborhood Hits | 72 | Target Neighborhood Hits | 746 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6PF04, Q96SS9 | Gene names | ZNF613 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 613. | |||||