Please be patient as the page loads
|
ETV4_HUMAN
|
||||||
| SwissProt Accessions | P43268, Q96AW9 | Gene names | ETV4, E1AF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 4 (Adenovirus E1A enhancer-binding protein) (E1A-F). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
ETV4_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 100 | |
| SwissProt Accessions | P43268, Q96AW9 | Gene names | ETV4, E1AF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 4 (Adenovirus E1A enhancer-binding protein) (E1A-F). | |||||
|
ETV4_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.993690 (rank : 2) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P28322 | Gene names | Etv4, Pea-3, Pea3 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 4 (Polyomavirus enhancer activator 3) (Protein PEA3). | |||||
|
ETV1_MOUSE
|
||||||
| θ value | 5.99046e-137 (rank : 3) | NC score | 0.976338 (rank : 3) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P41164 | Gene names | Etv1, Er81, Etsrp81 | |||
|
Domain Architecture |
|
|||||
| Description | ETS translocation variant 1 (ER81 protein). | |||||
|
ETV1_HUMAN
|
||||||
| θ value | 1.74296e-136 (rank : 4) | NC score | 0.975419 (rank : 4) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P50549, O75849, Q9UQ71, Q9Y636 | Gene names | ETV1, ER81 | |||
|
Domain Architecture |
|
|||||
| Description | ETS translocation variant 1 (ER81 protein). | |||||
|
ETV5_HUMAN
|
||||||
| θ value | 1.75107e-120 (rank : 5) | NC score | 0.968104 (rank : 5) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P41161 | Gene names | ETV5, ERM | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 5 (Ets-related protein ERM). | |||||
|
ETV5_MOUSE
|
||||||
| θ value | 1.13501e-119 (rank : 6) | NC score | 0.948396 (rank : 6) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9CXC9, Q3TG49, Q8C0F3, Q9JHB1 | Gene names | Etv5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 5. | |||||
|
ETS1_HUMAN
|
||||||
| θ value | 1.2049e-28 (rank : 7) | NC score | 0.869513 (rank : 14) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P14921, Q14278, Q16080 | Gene names | ETS1 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-1 protein (p54). | |||||
|
ETS1_MOUSE
|
||||||
| θ value | 1.2049e-28 (rank : 8) | NC score | 0.869230 (rank : 15) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P27577, Q61403 | Gene names | Ets1, Ets-1 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-1 protein (p54). | |||||
|
ELK4_MOUSE
|
||||||
| θ value | 2.68423e-28 (rank : 9) | NC score | 0.875401 (rank : 9) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P41158 | Gene names | Elk4, Sap1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-4 (Serum response factor accessory protein 1) (SAP-1). | |||||
|
ETS2_HUMAN
|
||||||
| θ value | 7.80994e-28 (rank : 10) | NC score | 0.866886 (rank : 16) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P15036 | Gene names | ETS2 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-2 protein. | |||||
|
ETS2_MOUSE
|
||||||
| θ value | 7.80994e-28 (rank : 11) | NC score | 0.866656 (rank : 17) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P15037 | Gene names | Ets2 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-2 protein. | |||||
|
ELK4_HUMAN
|
||||||
| θ value | 1.73987e-27 (rank : 12) | NC score | 0.860241 (rank : 18) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P28324, P28323 | Gene names | ELK4, SAP1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-4 (Serum response factor accessory protein 1) (SAP-1). | |||||
|
ELK1_HUMAN
|
||||||
| θ value | 4.28545e-26 (rank : 13) | NC score | 0.874555 (rank : 11) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P19419, O75606, O95058, Q969X8, Q9UJM4 | Gene names | ELK1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-1. | |||||
|
ELK1_MOUSE
|
||||||
| θ value | 4.28545e-26 (rank : 14) | NC score | 0.872708 (rank : 12) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P41969 | Gene names | Elk1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-1. | |||||
|
ELK3_HUMAN
|
||||||
| θ value | 1.62847e-25 (rank : 15) | NC score | 0.879472 (rank : 7) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P41970, Q6FG57, Q6GU29, Q9UD17 | Gene names | ELK3, NET, SAP2 | |||
|
Domain Architecture |
|
|||||
| Description | ETS domain-containing protein Elk-3 (ETS-related protein NET) (ETS- related protein ERP) (SRF accessory protein 2) (SAP-2). | |||||
|
ELK3_MOUSE
|
||||||
| θ value | 4.73814e-25 (rank : 16) | NC score | 0.876710 (rank : 8) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P41971, P97747, Q62346 | Gene names | Elk3, Erp, Net | |||
|
Domain Architecture |
|
|||||
| Description | ETS domain-containing protein Elk-3 (ETS-related protein NET) (ETS- related protein ERP). | |||||
|
FLI1_HUMAN
|
||||||
| θ value | 4.73814e-25 (rank : 17) | NC score | 0.843445 (rank : 22) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q01543, Q14319, Q92480, Q9UE07 | Gene names | FLI1 | |||
|
Domain Architecture |
|
|||||
| Description | Friend leukemia integration 1 transcription factor (Fli-1 proto- oncogene) (ERGB transcription factor). | |||||
|
ERG_MOUSE
|
||||||
| θ value | 6.18819e-25 (rank : 18) | NC score | 0.843226 (rank : 23) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P81270, Q8C5L4, Q920K7, Q920K8, Q920K9 | Gene names | Erg, Erg-3 | |||
|
Domain Architecture |
|
|||||
| Description | Transcriptional regulator ERG. | |||||
|
FLI1_MOUSE
|
||||||
| θ value | 6.18819e-25 (rank : 19) | NC score | 0.845451 (rank : 21) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P26323 | Gene names | Fli1, Fli-1 | |||
|
Domain Architecture |
|
|||||
| Description | Friend leukemia integration 1 transcription factor (Retroviral integration site protein Fli-1). | |||||
|
ERG_HUMAN
|
||||||
| θ value | 1.05554e-24 (rank : 20) | NC score | 0.843056 (rank : 24) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P11308, Q16113 | Gene names | ERG | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ERG (Transforming protein ERG). | |||||
|
ETV2_HUMAN
|
||||||
| θ value | 5.23862e-24 (rank : 21) | NC score | 0.875094 (rank : 10) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O00321, Q9UEA0 | Gene names | ETV2, ER71, ETSRP71 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 2 (Ets-related protein 71). | |||||
|
GABPA_MOUSE
|
||||||
| θ value | 3.39556e-23 (rank : 22) | NC score | 0.849737 (rank : 19) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q00422 | Gene names | Gabpa, E4tf1a | |||
|
Domain Architecture |
|
|||||
| Description | GA-binding protein alpha chain (GABP-subunit alpha). | |||||
|
GABPA_HUMAN
|
||||||
| θ value | 4.43474e-23 (rank : 23) | NC score | 0.848746 (rank : 20) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q06546, Q12939 | Gene names | GABPA, E4TF1A | |||
|
Domain Architecture |
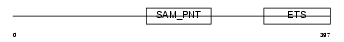
|
|||||
| Description | GA-binding protein alpha chain (GABP-subunit alpha) (Transcription factor E4TF1-60) (Nuclear respiratory factor 2 subunit alpha). | |||||
|
ETV2_MOUSE
|
||||||
| θ value | 1.29031e-22 (rank : 24) | NC score | 0.869927 (rank : 13) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P41163 | Gene names | Etv2, Er71, Etsrp71 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 2 (Ets-related protein 71). | |||||
|
ERF_HUMAN
|
||||||
| θ value | 1.02238e-19 (rank : 25) | NC score | 0.818512 (rank : 25) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P50548, Q9UPI7 | Gene names | ERF | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing transcription factor ERF (Ets2 repressor factor). | |||||
|
ERF_MOUSE
|
||||||
| θ value | 1.02238e-19 (rank : 26) | NC score | 0.817855 (rank : 26) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P70459 | Gene names | Erf | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing transcription factor ERF. | |||||
|
ETV3_HUMAN
|
||||||
| θ value | 2.97466e-19 (rank : 27) | NC score | 0.785781 (rank : 35) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P41162, Q8TAC8, Q9BX30 | Gene names | ETV3, METS, PE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
ETV3_MOUSE
|
||||||
| θ value | 2.97466e-19 (rank : 28) | NC score | 0.808652 (rank : 29) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8R4Z4, Q9QZW1 | Gene names | Etv3, Mets, Pe1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
ELF1_HUMAN
|
||||||
| θ value | 2.5182e-18 (rank : 29) | NC score | 0.787509 (rank : 32) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P32519, Q9UDE1 | Gene names | ELF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-1 (E74-like factor 1). | |||||
|
ELF4_HUMAN
|
||||||
| θ value | 3.28887e-18 (rank : 30) | NC score | 0.787482 (rank : 33) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q99607, O60435 | Gene names | ELF4, ELFR, MEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-4 (E74-like factor 4) (Myeloid Elf-1-like factor). | |||||
|
ELF1_MOUSE
|
||||||
| θ value | 4.29542e-18 (rank : 31) | NC score | 0.783031 (rank : 36) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q60775 | Gene names | Elf1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS-related transcription factor Elf-1 (E74-like factor 1). | |||||
|
ELF4_MOUSE
|
||||||
| θ value | 5.60996e-18 (rank : 32) | NC score | 0.768788 (rank : 39) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9Z2U4 | Gene names | Elf4, Mef | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-4 (E74-like factor 4) (Myeloid Elf-1-like factor). | |||||
|
SPDEF_HUMAN
|
||||||
| θ value | 1.38178e-16 (rank : 33) | NC score | 0.817412 (rank : 27) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O95238 | Gene names | SPDEF, PDEF, PSE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SAM pointed domain-containing Ets transcription factor (Prostate- derived Ets factor) (Prostate epithelium-specific Ets transcription factor) (Prostate-specific Ets). | |||||
|
ELF2_HUMAN
|
||||||
| θ value | 1.80466e-16 (rank : 34) | NC score | 0.787626 (rank : 31) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q15723, Q15724, Q15725, Q6P1K5 | Gene names | ELF2, NERF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-2 (E74-like factor 2) (New ETS- related factor). | |||||
|
ELF2_MOUSE
|
||||||
| θ value | 1.80466e-16 (rank : 35) | NC score | 0.787113 (rank : 34) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9JHC9, Q6NST2, Q8BTX8, Q9JHC7, Q9JHC8, Q9JHD0 | Gene names | Elf2, Nerf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-2 (E74-like factor 2) (New ETS- related factor). | |||||
|
SPDEF_MOUSE
|
||||||
| θ value | 1.80466e-16 (rank : 36) | NC score | 0.815278 (rank : 28) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9WTP3 | Gene names | Spdef, Pdef, Pse | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SAM pointed domain-containing Ets transcription factor (Prostate- derived Ets factor) (Prostate epithelium-specific Ets transcription factor) (Prostate-specific Ets). | |||||
|
ETV6_HUMAN
|
||||||
| θ value | 4.44505e-15 (rank : 37) | NC score | 0.780679 (rank : 38) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P41212, Q9UMF6 | Gene names | ETV6, TEL, TEL1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor ETV6 (ETS-related protein Tel1) (Tel) (ETS translocation variant 6). | |||||
|
ETV6_MOUSE
|
||||||
| θ value | 1.68911e-14 (rank : 38) | NC score | 0.781739 (rank : 37) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P97360 | Gene names | Etv6, Tel, Tel1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor ETV6 (ETS-related protein Tel1) (Tel) (ETS translocation variant 6). | |||||
|
ETV7_HUMAN
|
||||||
| θ value | 4.59992e-12 (rank : 39) | NC score | 0.802247 (rank : 30) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9Y603, Q9NZ65, Q9NZ66, Q9NZ68, Q9NZR8, Q9UNJ7, Q9Y5K4, Q9Y604 | Gene names | ETV7, TEL2, TELB, TREF | |||
|
Domain Architecture |
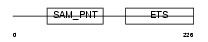
|
|||||
| Description | Transcription factor ETV7 (Transcription factor Tel-2) (ETS-related protein Tel2) (Tel-related Ets factor). | |||||
|
ELF5_HUMAN
|
||||||
| θ value | 1.133e-10 (rank : 40) | NC score | 0.754244 (rank : 40) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9UKW6, O95175, Q8N2K9, Q96QY3, Q9UKW5 | Gene names | ELF5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-5 (E74-like factor 5) (Epithelium-specific Ets transcription factor 2) (ESE-2) (Epithelium- restricted ESE-1-related Ets factor). | |||||
|
ELF5_MOUSE
|
||||||
| θ value | 1.133e-10 (rank : 41) | NC score | 0.753986 (rank : 41) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8VDK3, Q921H5, Q9Z2K6 | Gene names | Elf5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-5 (E74-like factor 5). | |||||
|
TOB1_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 42) | NC score | 0.041365 (rank : 55) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q61471 | Gene names | Tob1, Tob, Trob | |||
|
Domain Architecture |

|
|||||
| Description | Tob1 protein (Transducer of erbB-2 1). | |||||
|
WASIP_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 43) | NC score | 0.041430 (rank : 54) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O43516, Q15220, Q6MZU9, Q9BU37, Q9UNP1 | Gene names | WASPIP, WIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein) (PRPL-2 protein). | |||||
|
ARI1A_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 44) | NC score | 0.037845 (rank : 57) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 787 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O14497, Q53FK9, Q5T0W1, Q5T0W2, Q5T0W3, Q8NFD6, Q96T89, Q9BY33, Q9HBJ5, Q9UPZ1 | Gene names | ARID1A, C1orf4, OSA1, SMARCF1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 1A (ARID domain- containing protein 1A) (SWI/SNF-related, matrix-associated, actin- dependent regulator of chromatin subfamily F member 1) (SWI-SNF complex protein p270) (B120) (SWI-like protein) (Osa homolog 1) (hOSA1) (hELD) (BRG1-associated factor 250) (BAF250) (BRG1-associated factor 250a) (BAF250A). | |||||
|
NCOA6_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 45) | NC score | 0.051725 (rank : 50) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 1076 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9JL19, Q9JLT9 | Gene names | Ncoa6, Aib3, Prip, Rap250, Trbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC) (Thyroid hormone receptor-binding protein). | |||||
|
PTN23_MOUSE
|
||||||
| θ value | 0.125558 (rank : 46) | NC score | 0.032029 (rank : 61) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 726 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6PB44, Q69ZJ0, Q8R1Z5, Q923E6 | Gene names | Ptpn23, Kiaa1471 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 23 (EC 3.1.3.48). | |||||
|
SSXT_HUMAN
|
||||||
| θ value | 0.125558 (rank : 47) | NC score | 0.052621 (rank : 49) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q15532, Q16404, Q9BXC6 | Gene names | SS18, SSXT, SYT | |||
|
Domain Architecture |
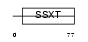
|
|||||
| Description | SSXT protein (Synovial sarcoma, translocated to X chromosome) (SYT protein). | |||||
|
TOB1_HUMAN
|
||||||
| θ value | 0.125558 (rank : 48) | NC score | 0.034870 (rank : 58) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P50616, Q4KMQ0 | Gene names | TOB1, TOB | |||
|
Domain Architecture |

|
|||||
| Description | Tob1 protein (Transducer of erbB-2 1). | |||||
|
MYST3_MOUSE
|
||||||
| θ value | 0.21417 (rank : 49) | NC score | 0.025220 (rank : 69) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 688 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8BZ21, Q8BW52, Q8BYH1, Q8C1F3 | Gene names | Myst3, Moz | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Monocytic leukemia zinc finger protein) (Monocytic leukemia zinc finger homolog). | |||||
|
NCOA6_HUMAN
|
||||||
| θ value | 0.21417 (rank : 50) | NC score | 0.049289 (rank : 51) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 1202 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q14686, Q9NTZ9, Q9UH74, Q9UK86 | Gene names | NCOA6, AIB3, KIAA0181, RAP250, TRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (PRIP) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC RAP250) (Thyroid hormone receptor-binding protein). | |||||
|
PTN23_HUMAN
|
||||||
| θ value | 0.21417 (rank : 51) | NC score | 0.034700 (rank : 59) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 646 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9H3S7, Q7KZF8, Q8N6Z5, Q9BSR5, Q9P257, Q9UG03, Q9UMZ4 | Gene names | PTPN23, KIAA1471 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 23 (EC 3.1.3.48) (His- domain-containing protein tyrosine phosphatase) (HD-PTP) (Protein tyrosine phosphatase TD14) (PTP-TD14). | |||||
|
MAML3_HUMAN
|
||||||
| θ value | 0.365318 (rank : 52) | NC score | 0.034229 (rank : 60) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q96JK9 | Gene names | MAML3, KIAA1816 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 3 (Mam-3). | |||||
|
DPOLM_MOUSE
|
||||||
| θ value | 0.47712 (rank : 53) | NC score | 0.030725 (rank : 63) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9JIW4, Q9JJW9 | Gene names | Polm | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase mu (EC 2.7.7.7) (Pol Mu). | |||||
|
LMO6_MOUSE
|
||||||
| θ value | 0.62314 (rank : 54) | NC score | 0.008456 (rank : 88) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80VL3, Q8CAY9 | Gene names | Lmo6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | LIM domain only protein 6 (Triple LIM domain protein 6). | |||||
|
BOC_HUMAN
|
||||||
| θ value | 0.813845 (rank : 55) | NC score | 0.005314 (rank : 93) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BWV1, Q6UXJ5, Q8N2P7, Q8NF26 | Gene names | BOC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brother of CDO precursor (Protein BOC). | |||||
|
IQEC2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 56) | NC score | 0.026610 (rank : 66) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q5JU85, O60275 | Gene names | IQSEC2, KIAA0522 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 2. | |||||
|
IQEC2_MOUSE
|
||||||
| θ value | 0.813845 (rank : 57) | NC score | 0.026605 (rank : 67) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q5DU25 | Gene names | Iqsec2, Kiaa0522 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 2. | |||||
|
SSXT_MOUSE
|
||||||
| θ value | 0.813845 (rank : 58) | NC score | 0.048331 (rank : 52) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q62280 | Gene names | Ss18, Ssxt, Syt | |||
|
Domain Architecture |
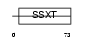
|
|||||
| Description | SSXT protein (SYT protein) (Synovial sarcoma-associated Ss18-alpha). | |||||
|
VAX2_MOUSE
|
||||||
| θ value | 0.813845 (rank : 59) | NC score | 0.009288 (rank : 85) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 338 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9WTP9 | Gene names | Vax2, Vex | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ventral anterior homeobox 2 (Ventral retina homeodomain protein). | |||||
|
BRD4_HUMAN
|
||||||
| θ value | 1.38821 (rank : 60) | NC score | 0.031561 (rank : 62) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O60885, O60433, Q96PD3 | Gene names | BRD4, HUNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 4 (HUNK1 protein). | |||||
|
NELF_HUMAN
|
||||||
| θ value | 1.38821 (rank : 61) | NC score | 0.018734 (rank : 72) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6X4W1, Q6X4V7, Q6X4V8, Q6X4V9, Q8N2M2, Q96SY1, Q9NPM4, Q9NPP3, Q9NPS3 | Gene names | NELF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nasal embryonic luteinizing hormone-releasing hormone factor (Nasal embryonic LHRH factor). | |||||
|
GSCR1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 62) | NC score | 0.054368 (rank : 48) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9NZM4 | Gene names | GLTSCR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glioma tumor suppressor candidate region gene 1 protein. | |||||
|
INCE_HUMAN
|
||||||
| θ value | 1.81305 (rank : 63) | NC score | 0.001629 (rank : 100) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 978 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NQS7, Q5Y192 | Gene names | INCENP | |||
|
Domain Architecture |

|
|||||
| Description | Inner centromere protein. | |||||
|
NFAT5_HUMAN
|
||||||
| θ value | 1.81305 (rank : 64) | NC score | 0.017816 (rank : 73) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O94916, O95693, Q9UN18 | Gene names | NFAT5, KIAA0827, TONEBP | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor of activated T-cells 5 (T-cell transcription factor NFAT5) (NF-AT5) (Tonicity-responsive enhancer-binding protein) (TonE- binding protein) (TonEBP). | |||||
|
PDC6I_HUMAN
|
||||||
| θ value | 1.81305 (rank : 65) | NC score | 0.045971 (rank : 53) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8WUM4, Q9BX86, Q9NUN0, Q9P2H2, Q9UKL5 | Gene names | PDCD6IP, AIP1, ALIX, KIAA1375 | |||
|
Domain Architecture |
|
|||||
| Description | Programmed cell death 6-interacting protein (PDCD6-interacting protein) (ALG-2-interacting protein 1) (Hp95). | |||||
|
SETB1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 66) | NC score | 0.011065 (rank : 82) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q15047, Q96GM9 | Gene names | SETDB1, KIAA0067 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 4 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 4) (H3-K9-HMTase 4) (SET domain bifurcated 1) (ERG-associated protein with SET domain) (ESET). | |||||
|
SPRR3_MOUSE
|
||||||
| θ value | 1.81305 (rank : 67) | NC score | 0.040090 (rank : 56) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O09116 | Gene names | Sprr3 | |||
|
Domain Architecture |

|
|||||
| Description | Small proline-rich protein 3 (Cornifin beta). | |||||
|
FOXJ2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 68) | NC score | 0.013924 (rank : 79) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9P0K8, Q96PS9, Q9NSN5 | Gene names | FOXJ2, FHX | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein J2 (Fork head homologous X). | |||||
|
BSN_MOUSE
|
||||||
| θ value | 3.0926 (rank : 69) | NC score | 0.019946 (rank : 71) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
GSLG1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 70) | NC score | 0.015109 (rank : 76) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61543, Q9QZ40 | Gene names | Glg1, Esl1, Mg160, Selel | |||
|
Domain Architecture |
|
|||||
| Description | Golgi apparatus protein 1 precursor (Golgi sialoglycoprotein MG-160) (E-selectin ligand 1) (ESL-1) (Selel). | |||||
|
KIF1C_MOUSE
|
||||||
| θ value | 3.0926 (rank : 71) | NC score | 0.002342 (rank : 98) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O35071, Q5SX62 | Gene names | Kif1c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin-like protein KIF1C. | |||||
|
SNTB2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 72) | NC score | 0.007676 (rank : 91) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61235 | Gene names | Sntb2, Snt2b2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-2-syntrophin (59 kDa dystrophin-associated protein A1 basic component 2) (Syntrophin 3) (SNT3) (Syntrophin-like) (SNTL). | |||||
|
DGKK_HUMAN
|
||||||
| θ value | 4.03905 (rank : 73) | NC score | 0.015350 (rank : 75) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 378 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q5KSL6 | Gene names | DGKK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Diacylglycerol kinase kappa (EC 2.7.1.107) (Diglyceride kinase kappa) (DGK-kappa) (DAG kinase kappa) (142 kDa diacylglycerol kinase). | |||||
|
PTPRN_MOUSE
|
||||||
| θ value | 4.03905 (rank : 74) | NC score | 0.010164 (rank : 84) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q60673, Q62129 | Gene names | Ptprn, Ptp35 | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase-like N precursor (R-PTP-N) (PTP IA-2). | |||||
|
RREB1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 75) | NC score | -0.002816 (rank : 105) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 879 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q92766, O75567 | Gene names | RREB1 | |||
|
Domain Architecture |

|
|||||
| Description | RAS-responsive element-binding protein 1 (RREB-1) (Raf-responsive zinc finger protein LZ321). | |||||
|
AMOL1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 76) | NC score | 0.005016 (rank : 96) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8IY63, Q63HK7, Q8NDN0, Q8WXD1, Q96CM5 | Gene names | AMOTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin-like protein 1. | |||||
|
BARH2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 77) | NC score | 0.001256 (rank : 102) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8VIB5, Q66L43 | Gene names | Barhl2, Barh1, Mbh1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BarH-like 2 homeobox protein (Bar-class homeodomain protein MBH1) (Homeobox protein B-H1). | |||||
|
BRD4_MOUSE
|
||||||
| θ value | 5.27518 (rank : 78) | NC score | 0.026953 (rank : 65) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 845 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9ESU6, Q8VHF7, Q8VHF8 | Gene names | Brd4, Mcap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 4 (Mitotic chromosome-associated protein) (MCAP). | |||||
|
IWS1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 79) | NC score | 0.008456 (rank : 87) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
MINT_MOUSE
|
||||||
| θ value | 5.27518 (rank : 80) | NC score | 0.011452 (rank : 81) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
PDC6I_MOUSE
|
||||||
| θ value | 5.27518 (rank : 81) | NC score | 0.030658 (rank : 64) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9WU78, O88695, O89014, Q8BSL8, Q8R0H5, Q99LR3, Q9QZN8 | Gene names | Pdcd6ip, Aip1, Alix | |||
|
Domain Architecture |
|
|||||
| Description | Programmed cell death 6-interacting protein (ALG-2-interacting protein X) (ALG-2-interacting protein 1) (E2F1-inducible protein) (Eig2). | |||||
|
PYGO2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 82) | NC score | 0.012491 (rank : 80) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9BRQ0, Q8WYZ4, Q96CY2 | Gene names | PYGO2 | |||
|
Domain Architecture |

|
|||||
| Description | Pygopus homolog 2. | |||||
|
SPR1A_HUMAN
|
||||||
| θ value | 5.27518 (rank : 83) | NC score | 0.026380 (rank : 68) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P35321, Q9UDG4 | Gene names | SPRR1A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cornifin A (Small proline-rich protein IA) (SPR-IA) (SPRK) (19 kDa pancornulin). | |||||
|
UBP31_HUMAN
|
||||||
| θ value | 5.27518 (rank : 84) | NC score | 0.003826 (rank : 97) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 293 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q70CQ4, Q6AW97, Q6ZTC0, Q6ZTN2, Q9ULL7 | Gene names | USP31, KIAA1203 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 31 (EC 3.1.2.15) (Ubiquitin thioesterase 31) (Ubiquitin-specific-processing protease 31) (Deubiquitinating enzyme 31). | |||||
|
AUTS2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 85) | NC score | 0.024456 (rank : 70) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8WXX7, Q9Y4F2 | Gene names | AUTS2, KIAA0442 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Autism susceptibility gene 2 protein. | |||||
|
HNRL1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 86) | NC score | 0.009016 (rank : 86) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BUJ2, O76022, Q6ZSZ0, Q7L8P4, Q8N6Z4, Q96G37, Q9HAL3, Q9UG75 | Gene names | HNRPUL1, E1BAP5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heterogeneous nuclear ribonucleoprotein U-like protein 1 (Adenovirus early region 1B-associated protein 5) (E1B-55 kDa-associated protein 5) (E1B-AP5). | |||||
|
IF4G3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 87) | NC score | 0.005062 (rank : 94) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O43432, Q15597, Q8NEN1 | Gene names | EIF4G3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 3 (eIF-4-gamma 3) (eIF-4G 3) (eIF4G 3) (eIF-4-gamma II) (eIF4GII). | |||||
|
LTBP2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 88) | NC score | -0.000984 (rank : 104) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 617 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O08999, Q8C6W9 | Gene names | Ltbp2 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein 2 precursor (LTBP-2). | |||||
|
PI5PA_HUMAN
|
||||||
| θ value | 6.88961 (rank : 89) | NC score | 0.014630 (rank : 77) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 749 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q15735, Q32M61, Q6ZTH6, Q8N902, Q9UDT9 | Gene names | PIB5PA, PIPP | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 4,5-bisphosphate 5-phosphatase A (EC 3.1.3.56). | |||||
|
RUSC2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 90) | NC score | 0.007873 (rank : 89) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q80U22, Q8R1D6 | Gene names | Rusc2, Kiaa0375 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and SH3 domain-containing protein 2. | |||||
|
SYN1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 91) | NC score | 0.010845 (rank : 83) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P17600, O75825, Q5H9A9 | Gene names | SYN1 | |||
|
Domain Architecture |

|
|||||
| Description | Synapsin-1 (Synapsin I) (Brain protein 4.1). | |||||
|
VAX2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 92) | NC score | 0.005019 (rank : 95) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UIW0 | Gene names | VAX2 | |||
|
Domain Architecture |
|
|||||
| Description | Ventral anterior homeobox 2. | |||||
|
WNK2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 93) | NC score | 0.001632 (rank : 99) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 1407 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9Y3S1, Q8IY36, Q9C0A3, Q9H3P4 | Gene names | WNK2, KIAA1760, PRKWNK2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK2 (EC 2.7.11.1) (Protein kinase with no lysine 2) (Protein kinase, lysine-deficient 2). | |||||
|
CRSP7_HUMAN
|
||||||
| θ value | 8.99809 (rank : 94) | NC score | 0.006750 (rank : 92) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O95402 | Gene names | CRSP7, ARC70 | |||
|
Domain Architecture |

|
|||||
| Description | CRSP complex subunit 7 (Cofactor required for Sp1 transcriptional activation subunit 7) (Transcriptional coactivator CRSP70) (Activator- recruited cofactor 70 kDa component) (ARC70). | |||||
|
DDEF2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 95) | NC score | 0.000835 (rank : 103) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 599 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O43150 | Gene names | DDEF2, KIAA0400 | |||
|
Domain Architecture |

|
|||||
| Description | Development and differentiation-enhancing factor 2 (Pyk2 C-terminus- associated protein) (PAP) (Paxillin-associated protein with ARFGAP activity 3) (PAG3). | |||||
|
K1802_MOUSE
|
||||||
| θ value | 8.99809 (rank : 96) | NC score | 0.014206 (rank : 78) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
SIX2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 97) | NC score | 0.001540 (rank : 101) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NPC8, Q9BXH7 | Gene names | SIX2 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein SIX2 (Sine oculis homeobox homolog 2). | |||||
|
TRI66_HUMAN
|
||||||
| θ value | 8.99809 (rank : 98) | NC score | 0.007868 (rank : 90) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O15016, Q9BQQ4 | Gene names | TRIM66, KIAA0298 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
YLPM1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 99) | NC score | 0.017028 (rank : 74) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P49750, P49752, Q96I64, Q9P1V7 | Gene names | YLPM1, C14orf170, ZAP3 | |||
|
Domain Architecture |

|
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3) (ZAP113). | |||||
|
ZN646_HUMAN
|
||||||
| θ value | 8.99809 (rank : 100) | NC score | -0.004546 (rank : 106) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 813 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O15015 | Gene names | ZNF646, KIAA0296 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 646. | |||||
|
SPI1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.493164 (rank : 44) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P17947 | Gene names | SPI1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor PU.1 (31 kDa transforming protein). | |||||
|
SPI1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.494091 (rank : 43) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P17433, Q99L57 | Gene names | Spi1, Sfpi-1, Sfpi1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor PU.1 (31 kDa transforming protein) (SFFV proviral integration 1 protein). | |||||
|
SPIB_HUMAN
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.496549 (rank : 42) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q01892, Q15359 | Gene names | SPIB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor Spi-B. | |||||
|
SPIB_MOUSE
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.482396 (rank : 45) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O35906, O35907, O35909, O55199 | Gene names | Spib | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor Spi-B. | |||||
|
SPIC_HUMAN
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.413516 (rank : 46) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8N5J4 | Gene names | SPIC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor Spi-C. | |||||
|
SPIC_MOUSE
|
||||||
| θ value | θ > 10 (rank : 106) | NC score | 0.387929 (rank : 47) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q6P3D7, Q9Z0Y0 | Gene names | Spic | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor Spi-C (Pu.1-related factor) (Prf). | |||||
|
ETV4_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 100 | |
| SwissProt Accessions | P43268, Q96AW9 | Gene names | ETV4, E1AF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 4 (Adenovirus E1A enhancer-binding protein) (E1A-F). | |||||
|
ETV4_MOUSE
|
||||||
| NC score | 0.993690 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P28322 | Gene names | Etv4, Pea-3, Pea3 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 4 (Polyomavirus enhancer activator 3) (Protein PEA3). | |||||
|
ETV1_MOUSE
|
||||||
| NC score | 0.976338 (rank : 3) | θ value | 5.99046e-137 (rank : 3) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P41164 | Gene names | Etv1, Er81, Etsrp81 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 1 (ER81 protein). | |||||
|
ETV1_HUMAN
|
||||||
| NC score | 0.975419 (rank : 4) | θ value | 1.74296e-136 (rank : 4) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P50549, O75849, Q9UQ71, Q9Y636 | Gene names | ETV1, ER81 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 1 (ER81 protein). | |||||
|
ETV5_HUMAN
|
||||||
| NC score | 0.968104 (rank : 5) | θ value | 1.75107e-120 (rank : 5) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P41161 | Gene names | ETV5, ERM | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 5 (Ets-related protein ERM). | |||||
|
ETV5_MOUSE
|
||||||
| NC score | 0.948396 (rank : 6) | θ value | 1.13501e-119 (rank : 6) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9CXC9, Q3TG49, Q8C0F3, Q9JHB1 | Gene names | Etv5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 5. | |||||
|
ELK3_HUMAN
|
||||||
| NC score | 0.879472 (rank : 7) | θ value | 1.62847e-25 (rank : 15) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P41970, Q6FG57, Q6GU29, Q9UD17 | Gene names | ELK3, NET, SAP2 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-3 (ETS-related protein NET) (ETS- related protein ERP) (SRF accessory protein 2) (SAP-2). | |||||
|
ELK3_MOUSE
|
||||||
| NC score | 0.876710 (rank : 8) | θ value | 4.73814e-25 (rank : 16) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P41971, P97747, Q62346 | Gene names | Elk3, Erp, Net | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-3 (ETS-related protein NET) (ETS- related protein ERP). | |||||
|
ELK4_MOUSE
|
||||||
| NC score | 0.875401 (rank : 9) | θ value | 2.68423e-28 (rank : 9) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P41158 | Gene names | Elk4, Sap1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-4 (Serum response factor accessory protein 1) (SAP-1). | |||||
|
ETV2_HUMAN
|
||||||
| NC score | 0.875094 (rank : 10) | θ value | 5.23862e-24 (rank : 21) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O00321, Q9UEA0 | Gene names | ETV2, ER71, ETSRP71 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 2 (Ets-related protein 71). | |||||
|
ELK1_HUMAN
|
||||||
| NC score | 0.874555 (rank : 11) | θ value | 4.28545e-26 (rank : 13) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P19419, O75606, O95058, Q969X8, Q9UJM4 | Gene names | ELK1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-1. | |||||
|
ELK1_MOUSE
|
||||||
| NC score | 0.872708 (rank : 12) | θ value | 4.28545e-26 (rank : 14) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P41969 | Gene names | Elk1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-1. | |||||
|
ETV2_MOUSE
|
||||||
| NC score | 0.869927 (rank : 13) | θ value | 1.29031e-22 (rank : 24) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P41163 | Gene names | Etv2, Er71, Etsrp71 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 2 (Ets-related protein 71). | |||||
|
ETS1_HUMAN
|
||||||
| NC score | 0.869513 (rank : 14) | θ value | 1.2049e-28 (rank : 7) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P14921, Q14278, Q16080 | Gene names | ETS1 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-1 protein (p54). | |||||
|
ETS1_MOUSE
|
||||||
| NC score | 0.869230 (rank : 15) | θ value | 1.2049e-28 (rank : 8) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P27577, Q61403 | Gene names | Ets1, Ets-1 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-1 protein (p54). | |||||
|
ETS2_HUMAN
|
||||||
| NC score | 0.866886 (rank : 16) | θ value | 7.80994e-28 (rank : 10) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P15036 | Gene names | ETS2 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-2 protein. | |||||
|
ETS2_MOUSE
|
||||||
| NC score | 0.866656 (rank : 17) | θ value | 7.80994e-28 (rank : 11) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P15037 | Gene names | Ets2 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-2 protein. | |||||
|
ELK4_HUMAN
|
||||||
| NC score | 0.860241 (rank : 18) | θ value | 1.73987e-27 (rank : 12) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P28324, P28323 | Gene names | ELK4, SAP1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-4 (Serum response factor accessory protein 1) (SAP-1). | |||||
|
GABPA_MOUSE
|
||||||
| NC score | 0.849737 (rank : 19) | θ value | 3.39556e-23 (rank : 22) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q00422 | Gene names | Gabpa, E4tf1a | |||
|
Domain Architecture |

|
|||||
| Description | GA-binding protein alpha chain (GABP-subunit alpha). | |||||
|
GABPA_HUMAN
|
||||||
| NC score | 0.848746 (rank : 20) | θ value | 4.43474e-23 (rank : 23) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q06546, Q12939 | Gene names | GABPA, E4TF1A | |||
|
Domain Architecture |

|
|||||
| Description | GA-binding protein alpha chain (GABP-subunit alpha) (Transcription factor E4TF1-60) (Nuclear respiratory factor 2 subunit alpha). | |||||
|
FLI1_MOUSE
|
||||||
| NC score | 0.845451 (rank : 21) | θ value | 6.18819e-25 (rank : 19) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P26323 | Gene names | Fli1, Fli-1 | |||
|
Domain Architecture |

|
|||||
| Description | Friend leukemia integration 1 transcription factor (Retroviral integration site protein Fli-1). | |||||
|
FLI1_HUMAN
|
||||||
| NC score | 0.843445 (rank : 22) | θ value | 4.73814e-25 (rank : 17) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q01543, Q14319, Q92480, Q9UE07 | Gene names | FLI1 | |||
|
Domain Architecture |

|
|||||
| Description | Friend leukemia integration 1 transcription factor (Fli-1 proto- oncogene) (ERGB transcription factor). | |||||
|
ERG_MOUSE
|
||||||
| NC score | 0.843226 (rank : 23) | θ value | 6.18819e-25 (rank : 18) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P81270, Q8C5L4, Q920K7, Q920K8, Q920K9 | Gene names | Erg, Erg-3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ERG. | |||||
|
ERG_HUMAN
|
||||||
| NC score | 0.843056 (rank : 24) | θ value | 1.05554e-24 (rank : 20) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P11308, Q16113 | Gene names | ERG | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ERG (Transforming protein ERG). | |||||
|
ERF_HUMAN
|
||||||
| NC score | 0.818512 (rank : 25) | θ value | 1.02238e-19 (rank : 25) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P50548, Q9UPI7 | Gene names | ERF | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing transcription factor ERF (Ets2 repressor factor). | |||||
|
ERF_MOUSE
|
||||||
| NC score | 0.817855 (rank : 26) | θ value | 1.02238e-19 (rank : 26) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P70459 | Gene names | Erf | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing transcription factor ERF. | |||||
|
SPDEF_HUMAN
|
||||||
| NC score | 0.817412 (rank : 27) | θ value | 1.38178e-16 (rank : 33) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O95238 | Gene names | SPDEF, PDEF, PSE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SAM pointed domain-containing Ets transcription factor (Prostate- derived Ets factor) (Prostate epithelium-specific Ets transcription factor) (Prostate-specific Ets). | |||||
|
SPDEF_MOUSE
|
||||||
| NC score | 0.815278 (rank : 28) | θ value | 1.80466e-16 (rank : 36) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9WTP3 | Gene names | Spdef, Pdef, Pse | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SAM pointed domain-containing Ets transcription factor (Prostate- derived Ets factor) (Prostate epithelium-specific Ets transcription factor) (Prostate-specific Ets). | |||||
|
ETV3_MOUSE
|
||||||
| NC score | 0.808652 (rank : 29) | θ value | 2.97466e-19 (rank : 28) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8R4Z4, Q9QZW1 | Gene names | Etv3, Mets, Pe1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
ETV7_HUMAN
|
||||||
| NC score | 0.802247 (rank : 30) | θ value | 4.59992e-12 (rank : 39) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9Y603, Q9NZ65, Q9NZ66, Q9NZ68, Q9NZR8, Q9UNJ7, Q9Y5K4, Q9Y604 | Gene names | ETV7, TEL2, TELB, TREF | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor ETV7 (Transcription factor Tel-2) (ETS-related protein Tel2) (Tel-related Ets factor). | |||||
|
ELF2_HUMAN
|
||||||
| NC score | 0.787626 (rank : 31) | θ value | 1.80466e-16 (rank : 34) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q15723, Q15724, Q15725, Q6P1K5 | Gene names | ELF2, NERF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-2 (E74-like factor 2) (New ETS- related factor). | |||||
|
ELF1_HUMAN
|
||||||
| NC score | 0.787509 (rank : 32) | θ value | 2.5182e-18 (rank : 29) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P32519, Q9UDE1 | Gene names | ELF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-1 (E74-like factor 1). | |||||
|
ELF4_HUMAN
|
||||||
| NC score | 0.787482 (rank : 33) | θ value | 3.28887e-18 (rank : 30) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q99607, O60435 | Gene names | ELF4, ELFR, MEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-4 (E74-like factor 4) (Myeloid Elf-1-like factor). | |||||
|
ELF2_MOUSE
|
||||||
| NC score | 0.787113 (rank : 34) | θ value | 1.80466e-16 (rank : 35) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9JHC9, Q6NST2, Q8BTX8, Q9JHC7, Q9JHC8, Q9JHD0 | Gene names | Elf2, Nerf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-2 (E74-like factor 2) (New ETS- related factor). | |||||
|
ETV3_HUMAN
|
||||||
| NC score | 0.785781 (rank : 35) | θ value | 2.97466e-19 (rank : 27) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P41162, Q8TAC8, Q9BX30 | Gene names | ETV3, METS, PE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
ELF1_MOUSE
|
||||||
| NC score | 0.783031 (rank : 36) | θ value | 4.29542e-18 (rank : 31) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q60775 | Gene names | Elf1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS-related transcription factor Elf-1 (E74-like factor 1). | |||||
|
ETV6_MOUSE
|
||||||
| NC score | 0.781739 (rank : 37) | θ value | 1.68911e-14 (rank : 38) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P97360 | Gene names | Etv6, Tel, Tel1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor ETV6 (ETS-related protein Tel1) (Tel) (ETS translocation variant 6). | |||||
|
ETV6_HUMAN
|
||||||
| NC score | 0.780679 (rank : 38) | θ value | 4.44505e-15 (rank : 37) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P41212, Q9UMF6 | Gene names | ETV6, TEL, TEL1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor ETV6 (ETS-related protein Tel1) (Tel) (ETS translocation variant 6). | |||||
|
ELF4_MOUSE
|
||||||
| NC score | 0.768788 (rank : 39) | θ value | 5.60996e-18 (rank : 32) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9Z2U4 | Gene names | Elf4, Mef | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-4 (E74-like factor 4) (Myeloid Elf-1-like factor). | |||||
|
ELF5_HUMAN
|
||||||
| NC score | 0.754244 (rank : 40) | θ value | 1.133e-10 (rank : 40) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9UKW6, O95175, Q8N2K9, Q96QY3, Q9UKW5 | Gene names | ELF5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-5 (E74-like factor 5) (Epithelium-specific Ets transcription factor 2) (ESE-2) (Epithelium- restricted ESE-1-related Ets factor). | |||||
|
ELF5_MOUSE
|
||||||
| NC score | 0.753986 (rank : 41) | θ value | 1.133e-10 (rank : 41) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8VDK3, Q921H5, Q9Z2K6 | Gene names | Elf5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-5 (E74-like factor 5). | |||||
|
SPIB_HUMAN
|
||||||
| NC score | 0.496549 (rank : 42) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q01892, Q15359 | Gene names | SPIB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor Spi-B. | |||||
|
SPI1_MOUSE
|
||||||
| NC score | 0.494091 (rank : 43) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P17433, Q99L57 | Gene names | Spi1, Sfpi-1, Sfpi1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor PU.1 (31 kDa transforming protein) (SFFV proviral integration 1 protein). | |||||
|
SPI1_HUMAN
|
||||||
| NC score | 0.493164 (rank : 44) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P17947 | Gene names | SPI1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor PU.1 (31 kDa transforming protein). | |||||
|
SPIB_MOUSE
|
||||||
| NC score | 0.482396 (rank : 45) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O35906, O35907, O35909, O55199 | Gene names | Spib | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor Spi-B. | |||||
|
SPIC_HUMAN
|
||||||
| NC score | 0.413516 (rank : 46) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8N5J4 | Gene names | SPIC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor Spi-C. | |||||
|
SPIC_MOUSE
|
||||||
| NC score | 0.387929 (rank : 47) | θ value | θ > 10 (rank : 106) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q6P3D7, Q9Z0Y0 | Gene names | Spic | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor Spi-C (Pu.1-related factor) (Prf). | |||||
|
GSCR1_HUMAN
|
||||||
| NC score | 0.054368 (rank : 48) | θ value | 1.81305 (rank : 62) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9NZM4 | Gene names | GLTSCR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glioma tumor suppressor candidate region gene 1 protein. | |||||
|
SSXT_HUMAN
|
||||||
| NC score | 0.052621 (rank : 49) | θ value | 0.125558 (rank : 47) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q15532, Q16404, Q9BXC6 | Gene names | SS18, SSXT, SYT | |||
|
Domain Architecture |

|
|||||
| Description | SSXT protein (Synovial sarcoma, translocated to X chromosome) (SYT protein). | |||||
|
NCOA6_MOUSE
|
||||||
| NC score | 0.051725 (rank : 50) | θ value | 0.0961366 (rank : 45) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 1076 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9JL19, Q9JLT9 | Gene names | Ncoa6, Aib3, Prip, Rap250, Trbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC) (Thyroid hormone receptor-binding protein). | |||||
|
NCOA6_HUMAN
|
||||||
| NC score | 0.049289 (rank : 51) | θ value | 0.21417 (rank : 50) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 1202 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q14686, Q9NTZ9, Q9UH74, Q9UK86 | Gene names | NCOA6, AIB3, KIAA0181, RAP250, TRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (PRIP) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC RAP250) (Thyroid hormone receptor-binding protein). | |||||
|
SSXT_MOUSE
|
||||||
| NC score | 0.048331 (rank : 52) | θ value | 0.813845 (rank : 58) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q62280 | Gene names | Ss18, Ssxt, Syt | |||
|
Domain Architecture |

|
|||||
| Description | SSXT protein (SYT protein) (Synovial sarcoma-associated Ss18-alpha). | |||||
|
PDC6I_HUMAN
|
||||||
| NC score | 0.045971 (rank : 53) | θ value | 1.81305 (rank : 65) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8WUM4, Q9BX86, Q9NUN0, Q9P2H2, Q9UKL5 | Gene names | PDCD6IP, AIP1, ALIX, KIAA1375 | |||
|
Domain Architecture |

|
|||||
| Description | Programmed cell death 6-interacting protein (PDCD6-interacting protein) (ALG-2-interacting protein 1) (Hp95). | |||||
|
WASIP_HUMAN
|
||||||
| NC score | 0.041430 (rank : 54) | θ value | 0.0736092 (rank : 43) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O43516, Q15220, Q6MZU9, Q9BU37, Q9UNP1 | Gene names | WASPIP, WIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein) (PRPL-2 protein). | |||||
|
TOB1_MOUSE
|
||||||
| NC score | 0.041365 (rank : 55) | θ value | 0.0563607 (rank : 42) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q61471 | Gene names | Tob1, Tob, Trob | |||
|
Domain Architecture |

|
|||||
| Description | Tob1 protein (Transducer of erbB-2 1). | |||||
|
SPRR3_MOUSE
|
||||||
| NC score | 0.040090 (rank : 56) | θ value | 1.81305 (rank : 67) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O09116 | Gene names | Sprr3 | |||
|
Domain Architecture |

|
|||||
| Description | Small proline-rich protein 3 (Cornifin beta). | |||||
|
ARI1A_HUMAN
|
||||||
| NC score | 0.037845 (rank : 57) | θ value | 0.0961366 (rank : 44) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 787 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O14497, Q53FK9, Q5T0W1, Q5T0W2, Q5T0W3, Q8NFD6, Q96T89, Q9BY33, Q9HBJ5, Q9UPZ1 | Gene names | ARID1A, C1orf4, OSA1, SMARCF1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 1A (ARID domain- containing protein 1A) (SWI/SNF-related, matrix-associated, actin- dependent regulator of chromatin subfamily F member 1) (SWI-SNF complex protein p270) (B120) (SWI-like protein) (Osa homolog 1) (hOSA1) (hELD) (BRG1-associated factor 250) (BAF250) (BRG1-associated factor 250a) (BAF250A). | |||||
|
TOB1_HUMAN
|
||||||
| NC score | 0.034870 (rank : 58) | θ value | 0.125558 (rank : 48) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P50616, Q4KMQ0 | Gene names | TOB1, TOB | |||
|
Domain Architecture |

|
|||||
| Description | Tob1 protein (Transducer of erbB-2 1). | |||||
|
PTN23_HUMAN
|
||||||
| NC score | 0.034700 (rank : 59) | θ value | 0.21417 (rank : 51) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 646 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9H3S7, Q7KZF8, Q8N6Z5, Q9BSR5, Q9P257, Q9UG03, Q9UMZ4 | Gene names | PTPN23, KIAA1471 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 23 (EC 3.1.3.48) (His- domain-containing protein tyrosine phosphatase) (HD-PTP) (Protein tyrosine phosphatase TD14) (PTP-TD14). | |||||
|
MAML3_HUMAN
|
||||||
| NC score | 0.034229 (rank : 60) | θ value | 0.365318 (rank : 52) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q96JK9 | Gene names | MAML3, KIAA1816 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 3 (Mam-3). | |||||
|
PTN23_MOUSE
|
||||||
| NC score | 0.032029 (rank : 61) | θ value | 0.125558 (rank : 46) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 726 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q6PB44, Q69ZJ0, Q8R1Z5, Q923E6 | Gene names | Ptpn23, Kiaa1471 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 23 (EC 3.1.3.48). | |||||
|
BRD4_HUMAN
|
||||||
| NC score | 0.031561 (rank : 62) | θ value | 1.38821 (rank : 60) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O60885, O60433, Q96PD3 | Gene names | BRD4, HUNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 4 (HUNK1 protein). | |||||
|
DPOLM_MOUSE
|
||||||
| NC score | 0.030725 (rank : 63) | θ value | 0.47712 (rank : 53) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9JIW4, Q9JJW9 | Gene names | Polm | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase mu (EC 2.7.7.7) (Pol Mu). | |||||
|
PDC6I_MOUSE
|
||||||
| NC score | 0.030658 (rank : 64) | θ value | 5.27518 (rank : 81) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9WU78, O88695, O89014, Q8BSL8, Q8R0H5, Q99LR3, Q9QZN8 | Gene names | Pdcd6ip, Aip1, Alix | |||
|
Domain Architecture |

|
|||||
| Description | Programmed cell death 6-interacting protein (ALG-2-interacting protein X) (ALG-2-interacting protein 1) (E2F1-inducible protein) (Eig2). | |||||
|
BRD4_MOUSE
|
||||||
| NC score | 0.026953 (rank : 65) | θ value | 5.27518 (rank : 78) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 845 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9ESU6, Q8VHF7, Q8VHF8 | Gene names | Brd4, Mcap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 4 (Mitotic chromosome-associated protein) (MCAP). | |||||
|
IQEC2_HUMAN
|
||||||
| NC score | 0.026610 (rank : 66) | θ value | 0.813845 (rank : 56) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q5JU85, O60275 | Gene names | IQSEC2, KIAA0522 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 2. | |||||
|
IQEC2_MOUSE
|
||||||
| NC score | 0.026605 (rank : 67) | θ value | 0.813845 (rank : 57) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q5DU25 | Gene names | Iqsec2, Kiaa0522 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IQ motif and Sec7 domain-containing protein 2. | |||||
|
SPR1A_HUMAN
|
||||||
| NC score | 0.026380 (rank : 68) | θ value | 5.27518 (rank : 83) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P35321, Q9UDG4 | Gene names | SPRR1A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cornifin A (Small proline-rich protein IA) (SPR-IA) (SPRK) (19 kDa pancornulin). | |||||
|
MYST3_MOUSE
|
||||||
| NC score | 0.025220 (rank : 69) | θ value | 0.21417 (rank : 49) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 688 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8BZ21, Q8BW52, Q8BYH1, Q8C1F3 | Gene names | Myst3, Moz | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Monocytic leukemia zinc finger protein) (Monocytic leukemia zinc finger homolog). | |||||
|
AUTS2_HUMAN
|
||||||
| NC score | 0.024456 (rank : 70) | θ value | 6.88961 (rank : 85) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8WXX7, Q9Y4F2 | Gene names | AUTS2, KIAA0442 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Autism susceptibility gene 2 protein. | |||||
|
BSN_MOUSE
|
||||||
| NC score | 0.019946 (rank : 71) | θ value | 3.0926 (rank : 69) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
NELF_HUMAN
|
||||||
| NC score | 0.018734 (rank : 72) | θ value | 1.38821 (rank : 61) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6X4W1, Q6X4V7, Q6X4V8, Q6X4V9, Q8N2M2, Q96SY1, Q9NPM4, Q9NPP3, Q9NPS3 | Gene names | NELF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nasal embryonic luteinizing hormone-releasing hormone factor (Nasal embryonic LHRH factor). | |||||
|
NFAT5_HUMAN
|
||||||
| NC score | 0.017816 (rank : 73) | θ value | 1.81305 (rank : 64) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O94916, O95693, Q9UN18 | Gene names | NFAT5, KIAA0827, TONEBP | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor of activated T-cells 5 (T-cell transcription factor NFAT5) (NF-AT5) (Tonicity-responsive enhancer-binding protein) (TonE- binding protein) (TonEBP). | |||||
|
YLPM1_HUMAN
|
||||||
| NC score | 0.017028 (rank : 74) | θ value | 8.99809 (rank : 99) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P49750, P49752, Q96I64, Q9P1V7 | Gene names | YLPM1, C14orf170, ZAP3 | |||
|
Domain Architecture |

|
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3) (ZAP113). | |||||
|
DGKK_HUMAN
|
||||||
| NC score | 0.015350 (rank : 75) | θ value | 4.03905 (rank : 73) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 378 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q5KSL6 | Gene names | DGKK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Diacylglycerol kinase kappa (EC 2.7.1.107) (Diglyceride kinase kappa) (DGK-kappa) (DAG kinase kappa) (142 kDa diacylglycerol kinase). | |||||
|
GSLG1_MOUSE
|
||||||
| NC score | 0.015109 (rank : 76) | θ value | 3.0926 (rank : 70) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61543, Q9QZ40 | Gene names | Glg1, Esl1, Mg160, Selel | |||
|
Domain Architecture |
|
|||||
| Description | Golgi apparatus protein 1 precursor (Golgi sialoglycoprotein MG-160) (E-selectin ligand 1) (ESL-1) (Selel). | |||||
|
PI5PA_HUMAN
|
||||||
| NC score | 0.014630 (rank : 77) | θ value | 6.88961 (rank : 89) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 749 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q15735, Q32M61, Q6ZTH6, Q8N902, Q9UDT9 | Gene names | PIB5PA, PIPP | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 4,5-bisphosphate 5-phosphatase A (EC 3.1.3.56). | |||||
|
K1802_MOUSE
|
||||||
| NC score | 0.014206 (rank : 78) | θ value | 8.99809 (rank : 96) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
FOXJ2_HUMAN
|
||||||
| NC score | 0.013924 (rank : 79) | θ value | 2.36792 (rank : 68) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9P0K8, Q96PS9, Q9NSN5 | Gene names | FOXJ2, FHX | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein J2 (Fork head homologous X). | |||||
|
PYGO2_HUMAN
|
||||||
| NC score | 0.012491 (rank : 80) | θ value | 5.27518 (rank : 82) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9BRQ0, Q8WYZ4, Q96CY2 | Gene names | PYGO2 | |||
|
Domain Architecture |

|
|||||
| Description | Pygopus homolog 2. | |||||
|
MINT_MOUSE
|
||||||
| NC score | 0.011452 (rank : 81) | θ value | 5.27518 (rank : 80) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
SETB1_HUMAN
|
||||||
| NC score | 0.011065 (rank : 82) | θ value | 1.81305 (rank : 66) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q15047, Q96GM9 | Gene names | SETDB1, KIAA0067 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 4 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 4) (H3-K9-HMTase 4) (SET domain bifurcated 1) (ERG-associated protein with SET domain) (ESET). | |||||
|
SYN1_HUMAN
|
||||||
| NC score | 0.010845 (rank : 83) | θ value | 6.88961 (rank : 91) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P17600, O75825, Q5H9A9 | Gene names | SYN1 | |||
|
Domain Architecture |

|
|||||
| Description | Synapsin-1 (Synapsin I) (Brain protein 4.1). | |||||
|
PTPRN_MOUSE
|
||||||
| NC score | 0.010164 (rank : 84) | θ value | 4.03905 (rank : 74) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q60673, Q62129 | Gene names | Ptprn, Ptp35 | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase-like N precursor (R-PTP-N) (PTP IA-2). | |||||
|
VAX2_MOUSE
|
||||||
| NC score | 0.009288 (rank : 85) | θ value | 0.813845 (rank : 59) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 338 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9WTP9 | Gene names | Vax2, Vex | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ventral anterior homeobox 2 (Ventral retina homeodomain protein). | |||||
|
HNRL1_HUMAN
|
||||||
| NC score | 0.009016 (rank : 86) | θ value | 6.88961 (rank : 86) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BUJ2, O76022, Q6ZSZ0, Q7L8P4, Q8N6Z4, Q96G37, Q9HAL3, Q9UG75 | Gene names | HNRPUL1, E1BAP5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heterogeneous nuclear ribonucleoprotein U-like protein 1 (Adenovirus early region 1B-associated protein 5) (E1B-55 kDa-associated protein 5) (E1B-AP5). | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.008456 (rank : 87) | θ value | 5.27518 (rank : 79) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
LMO6_MOUSE
|
||||||
| NC score | 0.008456 (rank : 88) | θ value | 0.62314 (rank : 54) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80VL3, Q8CAY9 | Gene names | Lmo6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | LIM domain only protein 6 (Triple LIM domain protein 6). | |||||
|
RUSC2_MOUSE
|
||||||
| NC score | 0.007873 (rank : 89) | θ value | 6.88961 (rank : 90) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q80U22, Q8R1D6 | Gene names | Rusc2, Kiaa0375 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and SH3 domain-containing protein 2. | |||||
|
TRI66_HUMAN
|
||||||
| NC score | 0.007868 (rank : 90) | θ value | 8.99809 (rank : 98) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O15016, Q9BQQ4 | Gene names | TRIM66, KIAA0298 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 66. | |||||
|
SNTB2_MOUSE
|
||||||
| NC score | 0.007676 (rank : 91) | θ value | 3.0926 (rank : 72) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61235 | Gene names | Sntb2, Snt2b2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-2-syntrophin (59 kDa dystrophin-associated protein A1 basic component 2) (Syntrophin 3) (SNT3) (Syntrophin-like) (SNTL). | |||||
|
CRSP7_HUMAN
|
||||||
| NC score | 0.006750 (rank : 92) | θ value | 8.99809 (rank : 94) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O95402 | Gene names | CRSP7, ARC70 | |||
|
Domain Architecture |

|
|||||
| Description | CRSP complex subunit 7 (Cofactor required for Sp1 transcriptional activation subunit 7) (Transcriptional coactivator CRSP70) (Activator- recruited cofactor 70 kDa component) (ARC70). | |||||
|
BOC_HUMAN
|
||||||
| NC score | 0.005314 (rank : 93) | θ value | 0.813845 (rank : 55) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BWV1, Q6UXJ5, Q8N2P7, Q8NF26 | Gene names | BOC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brother of CDO precursor (Protein BOC). | |||||
|
IF4G3_HUMAN
|
||||||
| NC score | 0.005062 (rank : 94) | θ value | 6.88961 (rank : 87) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O43432, Q15597, Q8NEN1 | Gene names | EIF4G3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 3 (eIF-4-gamma 3) (eIF-4G 3) (eIF4G 3) (eIF-4-gamma II) (eIF4GII). | |||||
|
VAX2_HUMAN
|
||||||
| NC score | 0.005019 (rank : 95) | θ value | 6.88961 (rank : 92) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UIW0 | Gene names | VAX2 | |||
|
Domain Architecture |

|
|||||
| Description | Ventral anterior homeobox 2. | |||||
|
AMOL1_HUMAN
|
||||||
| NC score | 0.005016 (rank : 96) | θ value | 5.27518 (rank : 76) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8IY63, Q63HK7, Q8NDN0, Q8WXD1, Q96CM5 | Gene names | AMOTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin-like protein 1. | |||||
|
UBP31_HUMAN
|
||||||
| NC score | 0.003826 (rank : 97) | θ value | 5.27518 (rank : 84) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 293 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q70CQ4, Q6AW97, Q6ZTC0, Q6ZTN2, Q9ULL7 | Gene names | USP31, KIAA1203 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 31 (EC 3.1.2.15) (Ubiquitin thioesterase 31) (Ubiquitin-specific-processing protease 31) (Deubiquitinating enzyme 31). | |||||
|
KIF1C_MOUSE
|
||||||
| NC score | 0.002342 (rank : 98) | θ value | 3.0926 (rank : 71) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O35071, Q5SX62 | Gene names | Kif1c | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin-like protein KIF1C. | |||||
|
WNK2_HUMAN
|
||||||
| NC score | 0.001632 (rank : 99) | θ value | 6.88961 (rank : 93) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 1407 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9Y3S1, Q8IY36, Q9C0A3, Q9H3P4 | Gene names | WNK2, KIAA1760, PRKWNK2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK2 (EC 2.7.11.1) (Protein kinase with no lysine 2) (Protein kinase, lysine-deficient 2). | |||||
|
INCE_HUMAN
|
||||||
| NC score | 0.001629 (rank : 100) | θ value | 1.81305 (rank : 63) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 978 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NQS7, Q5Y192 | Gene names | INCENP | |||
|
Domain Architecture |

|
|||||
| Description | Inner centromere protein. | |||||
|
SIX2_HUMAN
|
||||||
| NC score | 0.001540 (rank : 101) | θ value | 8.99809 (rank : 97) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NPC8, Q9BXH7 | Gene names | SIX2 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein SIX2 (Sine oculis homeobox homolog 2). | |||||
|
BARH2_MOUSE
|
||||||
| NC score | 0.001256 (rank : 102) | θ value | 5.27518 (rank : 77) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8VIB5, Q66L43 | Gene names | Barhl2, Barh1, Mbh1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BarH-like 2 homeobox protein (Bar-class homeodomain protein MBH1) (Homeobox protein B-H1). | |||||
|
DDEF2_HUMAN
|
||||||
| NC score | 0.000835 (rank : 103) | θ value | 8.99809 (rank : 95) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 599 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O43150 | Gene names | DDEF2, KIAA0400 | |||
|
Domain Architecture |

|
|||||
| Description | Development and differentiation-enhancing factor 2 (Pyk2 C-terminus- associated protein) (PAP) (Paxillin-associated protein with ARFGAP activity 3) (PAG3). | |||||
|
LTBP2_MOUSE
|
||||||
| NC score | -0.000984 (rank : 104) | θ value | 6.88961 (rank : 88) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 617 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O08999, Q8C6W9 | Gene names | Ltbp2 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein 2 precursor (LTBP-2). | |||||
|
RREB1_HUMAN
|
||||||
| NC score | -0.002816 (rank : 105) | θ value | 4.03905 (rank : 75) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 879 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q92766, O75567 | Gene names | RREB1 | |||
|
Domain Architecture |

|
|||||
| Description | RAS-responsive element-binding protein 1 (RREB-1) (Raf-responsive zinc finger protein LZ321). | |||||
|
ZN646_HUMAN
|
||||||
| NC score | -0.004546 (rank : 106) | θ value | 8.99809 (rank : 100) | |||
| Query Neighborhood Hits | 100 | Target Neighborhood Hits | 813 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O15015 | Gene names | ZNF646, KIAA0296 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 646. | |||||