Please be patient as the page loads
|
ERG_HUMAN
|
||||||
| SwissProt Accessions | P11308, Q16113 | Gene names | ERG | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ERG (Transforming protein ERG). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
ERG_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 86 | |
| SwissProt Accessions | P11308, Q16113 | Gene names | ERG | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ERG (Transforming protein ERG). | |||||
|
ERG_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.997316 (rank : 2) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | P81270, Q8C5L4, Q920K7, Q920K8, Q920K9 | Gene names | Erg, Erg-3 | |||
|
Domain Architecture |
|
|||||
| Description | Transcriptional regulator ERG. | |||||
|
FLI1_HUMAN
|
||||||
| θ value | 5.54228e-175 (rank : 3) | NC score | 0.991702 (rank : 4) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q01543, Q14319, Q92480, Q9UE07 | Gene names | FLI1 | |||
|
Domain Architecture |
|
|||||
| Description | Friend leukemia integration 1 transcription factor (Fli-1 proto- oncogene) (ERGB transcription factor). | |||||
|
FLI1_MOUSE
|
||||||
| θ value | 5.55514e-167 (rank : 4) | NC score | 0.991845 (rank : 3) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P26323 | Gene names | Fli1, Fli-1 | |||
|
Domain Architecture |
|
|||||
| Description | Friend leukemia integration 1 transcription factor (Retroviral integration site protein Fli-1). | |||||
|
GABPA_MOUSE
|
||||||
| θ value | 1.62093e-41 (rank : 5) | NC score | 0.924419 (rank : 5) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q00422 | Gene names | Gabpa, E4tf1a | |||
|
Domain Architecture |
|
|||||
| Description | GA-binding protein alpha chain (GABP-subunit alpha). | |||||
|
GABPA_HUMAN
|
||||||
| θ value | 2.11701e-41 (rank : 6) | NC score | 0.922535 (rank : 6) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q06546, Q12939 | Gene names | GABPA, E4TF1A | |||
|
Domain Architecture |
|
|||||
| Description | GA-binding protein alpha chain (GABP-subunit alpha) (Transcription factor E4TF1-60) (Nuclear respiratory factor 2 subunit alpha). | |||||
|
ETS1_HUMAN
|
||||||
| θ value | 7.54701e-31 (rank : 7) | NC score | 0.895053 (rank : 9) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P14921, Q14278, Q16080 | Gene names | ETS1 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-1 protein (p54). | |||||
|
ETS1_MOUSE
|
||||||
| θ value | 7.54701e-31 (rank : 8) | NC score | 0.894970 (rank : 10) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P27577, Q61403 | Gene names | Ets1, Ets-1 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-1 protein (p54). | |||||
|
ETS2_HUMAN
|
||||||
| θ value | 7.54701e-31 (rank : 9) | NC score | 0.894663 (rank : 11) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P15036 | Gene names | ETS2 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-2 protein. | |||||
|
ETS2_MOUSE
|
||||||
| θ value | 7.54701e-31 (rank : 10) | NC score | 0.893488 (rank : 12) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P15037 | Gene names | Ets2 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-2 protein. | |||||
|
ERF_HUMAN
|
||||||
| θ value | 1.2049e-28 (rank : 11) | NC score | 0.876605 (rank : 16) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P50548, Q9UPI7 | Gene names | ERF | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing transcription factor ERF (Ets2 repressor factor). | |||||
|
ERF_MOUSE
|
||||||
| θ value | 1.2049e-28 (rank : 12) | NC score | 0.876063 (rank : 18) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P70459 | Gene names | Erf | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing transcription factor ERF. | |||||
|
ETV3_MOUSE
|
||||||
| θ value | 1.02001e-27 (rank : 13) | NC score | 0.869177 (rank : 20) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q8R4Z4, Q9QZW1 | Gene names | Etv3, Mets, Pe1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
ETV3_HUMAN
|
||||||
| θ value | 5.06226e-27 (rank : 14) | NC score | 0.842441 (rank : 28) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P41162, Q8TAC8, Q9BX30 | Gene names | ETV3, METS, PE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
ETV1_HUMAN
|
||||||
| θ value | 8.63488e-27 (rank : 15) | NC score | 0.848320 (rank : 24) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P50549, O75849, Q9UQ71, Q9Y636 | Gene names | ETV1, ER81 | |||
|
Domain Architecture |
|
|||||
| Description | ETS translocation variant 1 (ER81 protein). | |||||
|
ETV1_MOUSE
|
||||||
| θ value | 8.63488e-27 (rank : 16) | NC score | 0.849210 (rank : 23) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P41164 | Gene names | Etv1, Er81, Etsrp81 | |||
|
Domain Architecture |
|
|||||
| Description | ETS translocation variant 1 (ER81 protein). | |||||
|
ETV2_HUMAN
|
||||||
| θ value | 2.51237e-26 (rank : 17) | NC score | 0.905631 (rank : 7) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O00321, Q9UEA0 | Gene names | ETV2, ER71, ETSRP71 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 2 (Ets-related protein 71). | |||||
|
ETV2_MOUSE
|
||||||
| θ value | 3.28125e-26 (rank : 18) | NC score | 0.903625 (rank : 8) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P41163 | Gene names | Etv2, Er71, Etsrp71 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 2 (Ets-related protein 71). | |||||
|
ETV5_HUMAN
|
||||||
| θ value | 1.24688e-25 (rank : 19) | NC score | 0.841862 (rank : 29) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P41161 | Gene names | ETV5, ERM | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 5 (Ets-related protein ERM). | |||||
|
ETV5_MOUSE
|
||||||
| θ value | 1.24688e-25 (rank : 20) | NC score | 0.823837 (rank : 31) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9CXC9, Q3TG49, Q8C0F3, Q9JHB1 | Gene names | Etv5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 5. | |||||
|
ELK4_MOUSE
|
||||||
| θ value | 6.18819e-25 (rank : 21) | NC score | 0.876739 (rank : 15) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P41158 | Gene names | Elk4, Sap1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-4 (Serum response factor accessory protein 1) (SAP-1). | |||||
|
ETV4_HUMAN
|
||||||
| θ value | 1.05554e-24 (rank : 22) | NC score | 0.843056 (rank : 27) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P43268, Q96AW9 | Gene names | ETV4, E1AF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 4 (Adenovirus E1A enhancer-binding protein) (E1A-F). | |||||
|
ETV4_MOUSE
|
||||||
| θ value | 1.05554e-24 (rank : 23) | NC score | 0.843414 (rank : 26) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P28322 | Gene names | Etv4, Pea-3, Pea3 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 4 (Polyomavirus enhancer activator 3) (Protein PEA3). | |||||
|
ELK1_MOUSE
|
||||||
| θ value | 8.93572e-24 (rank : 24) | NC score | 0.879676 (rank : 14) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P41969 | Gene names | Elk1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-1. | |||||
|
ELK1_HUMAN
|
||||||
| θ value | 1.16704e-23 (rank : 25) | NC score | 0.882059 (rank : 13) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P19419, O75606, O95058, Q969X8, Q9UJM4 | Gene names | ELK1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-1. | |||||
|
ELK4_HUMAN
|
||||||
| θ value | 1.16704e-23 (rank : 26) | NC score | 0.860759 (rank : 22) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P28324, P28323 | Gene names | ELK4, SAP1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-4 (Serum response factor accessory protein 1) (SAP-1). | |||||
|
ETV7_HUMAN
|
||||||
| θ value | 9.87957e-23 (rank : 27) | NC score | 0.867063 (rank : 21) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9Y603, Q9NZ65, Q9NZ66, Q9NZ68, Q9NZR8, Q9UNJ7, Q9Y5K4, Q9Y604 | Gene names | ETV7, TEL2, TELB, TREF | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor ETV7 (Transcription factor Tel-2) (ETS-related protein Tel2) (Tel-related Ets factor). | |||||
|
SPDEF_MOUSE
|
||||||
| θ value | 2.69047e-20 (rank : 28) | NC score | 0.844817 (rank : 25) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9WTP3 | Gene names | Spdef, Pdef, Pse | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SAM pointed domain-containing Ets transcription factor (Prostate- derived Ets factor) (Prostate epithelium-specific Ets transcription factor) (Prostate-specific Ets). | |||||
|
ELK3_HUMAN
|
||||||
| θ value | 4.58923e-20 (rank : 29) | NC score | 0.876407 (rank : 17) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P41970, Q6FG57, Q6GU29, Q9UD17 | Gene names | ELK3, NET, SAP2 | |||
|
Domain Architecture |
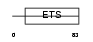
|
|||||
| Description | ETS domain-containing protein Elk-3 (ETS-related protein NET) (ETS- related protein ERP) (SRF accessory protein 2) (SAP-2). | |||||
|
ELK3_MOUSE
|
||||||
| θ value | 4.58923e-20 (rank : 30) | NC score | 0.873928 (rank : 19) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P41971, P97747, Q62346 | Gene names | Elk3, Erp, Net | |||
|
Domain Architecture |
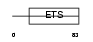
|
|||||
| Description | ETS domain-containing protein Elk-3 (ETS-related protein NET) (ETS- related protein ERP). | |||||
|
ELF1_HUMAN
|
||||||
| θ value | 1.99529e-15 (rank : 31) | NC score | 0.792988 (rank : 38) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P32519, Q9UDE1 | Gene names | ELF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-1 (E74-like factor 1). | |||||
|
ELF1_MOUSE
|
||||||
| θ value | 1.99529e-15 (rank : 32) | NC score | 0.786126 (rank : 40) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q60775 | Gene names | Elf1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS-related transcription factor Elf-1 (E74-like factor 1). | |||||
|
ELF4_MOUSE
|
||||||
| θ value | 3.40345e-15 (rank : 33) | NC score | 0.771411 (rank : 41) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9Z2U4 | Gene names | Elf4, Mef | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-4 (E74-like factor 4) (Myeloid Elf-1-like factor). | |||||
|
ELF2_MOUSE
|
||||||
| θ value | 4.44505e-15 (rank : 34) | NC score | 0.798358 (rank : 36) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9JHC9, Q6NST2, Q8BTX8, Q9JHC7, Q9JHC8, Q9JHD0 | Gene names | Elf2, Nerf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-2 (E74-like factor 2) (New ETS- related factor). | |||||
|
ELF4_HUMAN
|
||||||
| θ value | 4.44505e-15 (rank : 35) | NC score | 0.791369 (rank : 39) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q99607, O60435 | Gene names | ELF4, ELFR, MEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-4 (E74-like factor 4) (Myeloid Elf-1-like factor). | |||||
|
ELF2_HUMAN
|
||||||
| θ value | 5.8054e-15 (rank : 36) | NC score | 0.796180 (rank : 37) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q15723, Q15724, Q15725, Q6P1K5 | Gene names | ELF2, NERF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-2 (E74-like factor 2) (New ETS- related factor). | |||||
|
ETV6_HUMAN
|
||||||
| θ value | 2.20605e-14 (rank : 37) | NC score | 0.799950 (rank : 35) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P41212, Q9UMF6 | Gene names | ETV6, TEL, TEL1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor ETV6 (ETS-related protein Tel1) (Tel) (ETS translocation variant 6). | |||||
|
ETV6_MOUSE
|
||||||
| θ value | 2.20605e-14 (rank : 38) | NC score | 0.804678 (rank : 33) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P97360 | Gene names | Etv6, Tel, Tel1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor ETV6 (ETS-related protein Tel1) (Tel) (ETS translocation variant 6). | |||||
|
ELF5_MOUSE
|
||||||
| θ value | 1.42992e-13 (rank : 39) | NC score | 0.804917 (rank : 32) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8VDK3, Q921H5, Q9Z2K6 | Gene names | Elf5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-5 (E74-like factor 5). | |||||
|
SPDEF_HUMAN
|
||||||
| θ value | 5.43371e-13 (rank : 40) | NC score | 0.839816 (rank : 30) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O95238 | Gene names | SPDEF, PDEF, PSE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SAM pointed domain-containing Ets transcription factor (Prostate- derived Ets factor) (Prostate epithelium-specific Ets transcription factor) (Prostate-specific Ets). | |||||
|
ELF5_HUMAN
|
||||||
| θ value | 6.64225e-11 (rank : 41) | NC score | 0.801301 (rank : 34) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9UKW6, O95175, Q8N2K9, Q96QY3, Q9UKW5 | Gene names | ELF5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-5 (E74-like factor 5) (Epithelium-specific Ets transcription factor 2) (ESE-2) (Epithelium- restricted ESE-1-related Ets factor). | |||||
|
SPIB_HUMAN
|
||||||
| θ value | 1.99992e-07 (rank : 42) | NC score | 0.631467 (rank : 42) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q01892, Q15359 | Gene names | SPIB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor Spi-B. | |||||
|
SPIC_HUMAN
|
||||||
| θ value | 3.19293e-05 (rank : 43) | NC score | 0.590881 (rank : 46) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8N5J4 | Gene names | SPIC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor Spi-C. | |||||
|
SPIB_MOUSE
|
||||||
| θ value | 0.000461057 (rank : 44) | NC score | 0.608833 (rank : 45) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O35906, O35907, O35909, O55199 | Gene names | Spib | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor Spi-B. | |||||
|
SPI1_HUMAN
|
||||||
| θ value | 0.000786445 (rank : 45) | NC score | 0.612181 (rank : 43) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P17947 | Gene names | SPI1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor PU.1 (31 kDa transforming protein). | |||||
|
SPI1_MOUSE
|
||||||
| θ value | 0.000786445 (rank : 46) | NC score | 0.611772 (rank : 44) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P17433, Q99L57 | Gene names | Spi1, Sfpi-1, Sfpi1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor PU.1 (31 kDa transforming protein) (SFFV proviral integration 1 protein). | |||||
|
SPIC_MOUSE
|
||||||
| θ value | 0.000786445 (rank : 47) | NC score | 0.560360 (rank : 47) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q6P3D7, Q9Z0Y0 | Gene names | Spic | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor Spi-C (Pu.1-related factor) (Prf). | |||||
|
TFCP2_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 48) | NC score | 0.052777 (rank : 49) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q12800, Q12801, Q9UD77 | Gene names | TFCP2, LSF, SEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alpha-globin transcription factor CP2 (Transcription factor LSF) (SAA3 enhancer factor). | |||||
|
TFCP2_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 49) | NC score | 0.052792 (rank : 48) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9ERA0, Q3UGJ4, Q8VC13 | Gene names | Tcfcp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alpha-globin transcription factor CP2. | |||||
|
AHI1_HUMAN
|
||||||
| θ value | 0.125558 (rank : 50) | NC score | 0.015716 (rank : 58) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 537 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8N157, Q504T3, Q5TCP9, Q6P098, Q6PIT6, Q8NDX0, Q9H0H2 | Gene names | AHI1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Jouberin (Abelson helper integration site 1 protein homolog) (AHI-1). | |||||
|
NDST4_MOUSE
|
||||||
| θ value | 0.62314 (rank : 51) | NC score | 0.016239 (rank : 56) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9EQW8 | Gene names | Ndst4, Hsst4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bifunctional heparan sulfate N-deacetylase/N-sulfotransferase 4 (EC 2.8.2.8) (Glucosaminyl N-deacetylase/N-sulfotransferase 4) (NDST- 4) (N-heparan sulfate sulfotransferase 4) (N-HSST 4) [Includes: Heparan sulfate N-deacetylase 4 (EC 3.-.-.-); Heparan sulfate N- sulfotransferase 4 (EC 2.8.2.-)]. | |||||
|
WIRE_HUMAN
|
||||||
| θ value | 0.813845 (rank : 52) | NC score | 0.021357 (rank : 51) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8TF74, Q658J8, Q71RE1, Q8TE44 | Gene names | WIRE, WICH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WIP-related protein (WASP-interacting protein-related protein) (WIP- and CR16-homologous protein). | |||||
|
GLUD1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 53) | NC score | 0.020717 (rank : 52) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CIX8 | Gene names | Gluld1, Lgs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate--ammonia ligase domain-containing protein 1 (Lengsin) (Lens glutamine synthase-like). | |||||
|
EP300_HUMAN
|
||||||
| θ value | 1.38821 (rank : 54) | NC score | 0.011874 (rank : 65) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q09472 | Gene names | EP300, P300 | |||
|
Domain Architecture |

|
|||||
| Description | E1A-associated protein p300 (EC 2.3.1.48). | |||||
|
M3K2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 55) | NC score | -0.003036 (rank : 84) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 846 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q61083 | Gene names | Map3k2, Mekk2 | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase 2 (EC 2.7.11.25) (MAPK/ERK kinase kinase 2) (MEK kinase 2) (MEKK 2). | |||||
|
JPH2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 56) | NC score | 0.010112 (rank : 72) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9ET78, Q9ET79 | Gene names | Jph2, Jp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junctophilin-2 (Junctophilin type 2) (JP-2). | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 57) | NC score | 0.006578 (rank : 77) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
3BP1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 58) | NC score | 0.011619 (rank : 68) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 376 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y3L3, Q6IBZ2, Q6ZVL9, Q96HQ5, Q9NSQ9 | Gene names | SH3BP1 | |||
|
Domain Architecture |
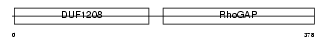
|
|||||
| Description | SH3 domain-binding protein 1 (3BP-1). | |||||
|
LIMK1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 59) | NC score | -0.004071 (rank : 86) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1008 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P53668 | Gene names | Limk1, Limk | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain kinase 1 (EC 2.7.11.1) (LIMK-1) (KIZ-1). | |||||
|
MYO15_MOUSE
|
||||||
| θ value | 3.0926 (rank : 60) | NC score | 0.005037 (rank : 78) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9QZZ4, O70395, Q9QWL6 | Gene names | Myo15a, Myo15 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-15 (Myosin XV) (Unconventional myosin-15). | |||||
|
NDST4_HUMAN
|
||||||
| θ value | 3.0926 (rank : 61) | NC score | 0.010755 (rank : 70) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H3R1 | Gene names | NDST4, HSST4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bifunctional heparan sulfate N-deacetylase/N-sulfotransferase 4 (EC 2.8.2.8) (Glucosaminyl N-deacetylase/N-sulfotransferase 4) (NDST- 4) (N-heparan sulfate sulfotransferase 4) (N-HSST 4) [Includes: Heparan sulfate N-deacetylase 4 (EC 3.-.-.-); Heparan sulfate N- sulfotransferase 4 (EC 2.8.2.-)]. | |||||
|
TF2L1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 62) | NC score | 0.035572 (rank : 50) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q3UNW5, Q99PF2 | Gene names | Tcfcp2l1, Crtr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor CP2-like protein 1 (CP2-related transcriptional repressor 1) (CRTR-1). | |||||
|
FOXN1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 63) | NC score | 0.010076 (rank : 73) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 377 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O15353, O15352 | Gene names | FOXN1, WHN | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein N1 (Transcription factor winged-helix nude). | |||||
|
PARM1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 64) | NC score | 0.013001 (rank : 64) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q923D3, Q3TTV0, Q3UFU4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein PARM-1 precursor. | |||||
|
WASF2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 65) | NC score | 0.009976 (rank : 74) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Y6W5, O60794, Q9UDY7 | Gene names | WASF2, WAVE2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein family member 2 (WASP-family protein member 2) (Protein WAVE-2) (Verprolin homology domain-containing protein 2). | |||||
|
WASF2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 66) | NC score | 0.017112 (rank : 55) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8BH43 | Gene names | Wasf2, Wave2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein family member 2 (WASP-family protein member 2) (Protein WAVE-2). | |||||
|
BCL9_MOUSE
|
||||||
| θ value | 5.27518 (rank : 67) | NC score | 0.013837 (rank : 61) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 770 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9D219, Q67FX9, Q8BUJ8, Q8VE74 | Gene names | Bcl9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9). | |||||
|
CPSF7_HUMAN
|
||||||
| θ value | 5.27518 (rank : 68) | NC score | 0.017502 (rank : 54) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8N684, Q7Z3H9, Q9H025, Q9H9V1 | Gene names | CPSF7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage and polyadenylation specificity factor 7 (Cleavage and polyadenylation specificity factor 59 kDa subunit) (CPSF 59 kDa subunit) (Pre-mRNA cleavage factor Im 59 kDa subunit). | |||||
|
CPSF7_MOUSE
|
||||||
| θ value | 5.27518 (rank : 69) | NC score | 0.017586 (rank : 53) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BTV2, Q3TNF1, Q8BKE7, Q8CFS8 | Gene names | Cpsf7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage and polyadenylation specificity factor 7. | |||||
|
LHX5_HUMAN
|
||||||
| θ value | 5.27518 (rank : 70) | NC score | 0.000327 (rank : 81) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 453 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H2C1 | Gene names | LHX5 | |||
|
Domain Architecture |
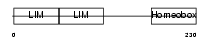
|
|||||
| Description | LIM/homeobox protein Lhx5. | |||||
|
PHC1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 71) | NC score | 0.007895 (rank : 76) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q64028, P70359, Q64307, Q8BZ80 | Gene names | Phc1, Edr, Edr1, Rae28 | |||
|
Domain Architecture |

|
|||||
| Description | Polyhomeotic-like protein 1 (mPH1) (Early development regulatory protein 1) (RAE-28). | |||||
|
RTN1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 72) | NC score | 0.011806 (rank : 67) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q16799, Q16800, Q16801 | Gene names | RTN1, NSP | |||
|
Domain Architecture |

|
|||||
| Description | Reticulon-1 (Neuroendocrine-specific protein). | |||||
|
BCL9_HUMAN
|
||||||
| θ value | 6.88961 (rank : 73) | NC score | 0.015847 (rank : 57) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1008 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O00512 | Gene names | BCL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9) (Legless homolog). | |||||
|
FUBP3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 74) | NC score | 0.010209 (rank : 71) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96I24, Q92946, Q9BVB6 | Gene names | FUBP3 | |||
|
Domain Architecture |

|
|||||
| Description | Far upstream element-binding protein 3 (FUSE-binding protein 3). | |||||
|
SOX10_MOUSE
|
||||||
| θ value | 6.88961 (rank : 75) | NC score | 0.002533 (rank : 80) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q04888, O08518, O09141, O54856, P70416 | Gene names | Sox10, Sox-10, Sox21 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-10 (SOX-21) (Transcription factor SOX-M). | |||||
|
TCF7_MOUSE
|
||||||
| θ value | 6.88961 (rank : 76) | NC score | 0.011308 (rank : 69) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q00417 | Gene names | Tcf7, Tcf-1, Tcf1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 7 (T-cell-specific transcription factor 1) (TCF- 1) (T-cell factor 1). | |||||
|
BAZ2A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 77) | NC score | 0.004601 (rank : 79) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q91YE5 | Gene names | Baz2a, Tip5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5). | |||||
|
CELR3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 78) | NC score | -0.001720 (rank : 83) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 687 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q91ZI0, Q9ESD0 | Gene names | Celsr3 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin EGF LAG seven-pass G-type receptor 3 precursor. | |||||
|
GP137_HUMAN
|
||||||
| θ value | 8.99809 (rank : 79) | NC score | 0.013345 (rank : 62) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q14444, Q15074 | Gene names | M11S1, GPIP137 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GPI-anchored protein p137 (p137GPI) (Membrane component chromosome 11 surface marker 1). | |||||
|
KLH17_HUMAN
|
||||||
| θ value | 8.99809 (rank : 80) | NC score | -0.000871 (rank : 82) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6TDP4 | Gene names | KLHL17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kelch-like protein 17 (Actinfilin). | |||||
|
M3K2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 81) | NC score | -0.003890 (rank : 85) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 849 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y2U5, Q53QL9, Q53S75, Q59GZ6, Q8NC32, Q9NYK3 | Gene names | MAP3K2, MAPKKK2, MEKK2 | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase 2 (EC 2.7.11.25) (MAPK/ERK kinase kinase 2) (MEK kinase 2) (MEKK 2). | |||||
|
MDC1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 82) | NC score | 0.011872 (rank : 66) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
PHLA1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 83) | NC score | 0.015440 (rank : 60) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q62392, Q3TY48, Q3UG87 | Gene names | Phlda1, Tdag51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family A member 1 (T-cell death- associated gene 51 protein) (Proline- and glutamine-rich protein) (PQR protein). | |||||
|
REC8L_MOUSE
|
||||||
| θ value | 8.99809 (rank : 84) | NC score | 0.008056 (rank : 75) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8C5S7, Q3UIN9, Q9JK52 | Gene names | Rec8L1, Rec8 | |||
|
Domain Architecture |

|
|||||
| Description | Meiotic recombination protein REC8-like 1 (Cohesin Rec8p). | |||||
|
WIRE_MOUSE
|
||||||
| θ value | 8.99809 (rank : 85) | NC score | 0.015564 (rank : 59) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6PEV3 | Gene names | Wire | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WIP-related protein (WASP-interacting protein-related protein). | |||||
|
ZN645_HUMAN
|
||||||
| θ value | 8.99809 (rank : 86) | NC score | 0.013217 (rank : 63) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 265 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8N7E2, Q6DJY9 | Gene names | ZNF645 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 645. | |||||
|
ERG_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 86 | |
| SwissProt Accessions | P11308, Q16113 | Gene names | ERG | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ERG (Transforming protein ERG). | |||||
|
ERG_MOUSE
|
||||||
| NC score | 0.997316 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | P81270, Q8C5L4, Q920K7, Q920K8, Q920K9 | Gene names | Erg, Erg-3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ERG. | |||||
|
FLI1_MOUSE
|
||||||
| NC score | 0.991845 (rank : 3) | θ value | 5.55514e-167 (rank : 4) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P26323 | Gene names | Fli1, Fli-1 | |||
|
Domain Architecture |

|
|||||
| Description | Friend leukemia integration 1 transcription factor (Retroviral integration site protein Fli-1). | |||||
|
FLI1_HUMAN
|
||||||
| NC score | 0.991702 (rank : 4) | θ value | 5.54228e-175 (rank : 3) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q01543, Q14319, Q92480, Q9UE07 | Gene names | FLI1 | |||
|
Domain Architecture |

|
|||||
| Description | Friend leukemia integration 1 transcription factor (Fli-1 proto- oncogene) (ERGB transcription factor). | |||||
|
GABPA_MOUSE
|
||||||
| NC score | 0.924419 (rank : 5) | θ value | 1.62093e-41 (rank : 5) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q00422 | Gene names | Gabpa, E4tf1a | |||
|
Domain Architecture |

|
|||||
| Description | GA-binding protein alpha chain (GABP-subunit alpha). | |||||
|
GABPA_HUMAN
|
||||||
| NC score | 0.922535 (rank : 6) | θ value | 2.11701e-41 (rank : 6) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q06546, Q12939 | Gene names | GABPA, E4TF1A | |||
|
Domain Architecture |

|
|||||
| Description | GA-binding protein alpha chain (GABP-subunit alpha) (Transcription factor E4TF1-60) (Nuclear respiratory factor 2 subunit alpha). | |||||
|
ETV2_HUMAN
|
||||||
| NC score | 0.905631 (rank : 7) | θ value | 2.51237e-26 (rank : 17) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O00321, Q9UEA0 | Gene names | ETV2, ER71, ETSRP71 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 2 (Ets-related protein 71). | |||||
|
ETV2_MOUSE
|
||||||
| NC score | 0.903625 (rank : 8) | θ value | 3.28125e-26 (rank : 18) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P41163 | Gene names | Etv2, Er71, Etsrp71 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 2 (Ets-related protein 71). | |||||
|
ETS1_HUMAN
|
||||||
| NC score | 0.895053 (rank : 9) | θ value | 7.54701e-31 (rank : 7) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P14921, Q14278, Q16080 | Gene names | ETS1 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-1 protein (p54). | |||||
|
ETS1_MOUSE
|
||||||
| NC score | 0.894970 (rank : 10) | θ value | 7.54701e-31 (rank : 8) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P27577, Q61403 | Gene names | Ets1, Ets-1 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-1 protein (p54). | |||||
|
ETS2_HUMAN
|
||||||
| NC score | 0.894663 (rank : 11) | θ value | 7.54701e-31 (rank : 9) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P15036 | Gene names | ETS2 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-2 protein. | |||||
|
ETS2_MOUSE
|
||||||
| NC score | 0.893488 (rank : 12) | θ value | 7.54701e-31 (rank : 10) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P15037 | Gene names | Ets2 | |||
|
Domain Architecture |

|
|||||
| Description | C-ets-2 protein. | |||||
|
ELK1_HUMAN
|
||||||
| NC score | 0.882059 (rank : 13) | θ value | 1.16704e-23 (rank : 25) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P19419, O75606, O95058, Q969X8, Q9UJM4 | Gene names | ELK1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-1. | |||||
|
ELK1_MOUSE
|
||||||
| NC score | 0.879676 (rank : 14) | θ value | 8.93572e-24 (rank : 24) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P41969 | Gene names | Elk1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-1. | |||||
|
ELK4_MOUSE
|
||||||
| NC score | 0.876739 (rank : 15) | θ value | 6.18819e-25 (rank : 21) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P41158 | Gene names | Elk4, Sap1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-4 (Serum response factor accessory protein 1) (SAP-1). | |||||
|
ERF_HUMAN
|
||||||
| NC score | 0.876605 (rank : 16) | θ value | 1.2049e-28 (rank : 11) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P50548, Q9UPI7 | Gene names | ERF | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing transcription factor ERF (Ets2 repressor factor). | |||||
|
ELK3_HUMAN
|
||||||
| NC score | 0.876407 (rank : 17) | θ value | 4.58923e-20 (rank : 29) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P41970, Q6FG57, Q6GU29, Q9UD17 | Gene names | ELK3, NET, SAP2 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-3 (ETS-related protein NET) (ETS- related protein ERP) (SRF accessory protein 2) (SAP-2). | |||||
|
ERF_MOUSE
|
||||||
| NC score | 0.876063 (rank : 18) | θ value | 1.2049e-28 (rank : 12) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P70459 | Gene names | Erf | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing transcription factor ERF. | |||||
|
ELK3_MOUSE
|
||||||
| NC score | 0.873928 (rank : 19) | θ value | 4.58923e-20 (rank : 30) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P41971, P97747, Q62346 | Gene names | Elk3, Erp, Net | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-3 (ETS-related protein NET) (ETS- related protein ERP). | |||||
|
ETV3_MOUSE
|
||||||
| NC score | 0.869177 (rank : 20) | θ value | 1.02001e-27 (rank : 13) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q8R4Z4, Q9QZW1 | Gene names | Etv3, Mets, Pe1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
ETV7_HUMAN
|
||||||
| NC score | 0.867063 (rank : 21) | θ value | 9.87957e-23 (rank : 27) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9Y603, Q9NZ65, Q9NZ66, Q9NZ68, Q9NZR8, Q9UNJ7, Q9Y5K4, Q9Y604 | Gene names | ETV7, TEL2, TELB, TREF | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor ETV7 (Transcription factor Tel-2) (ETS-related protein Tel2) (Tel-related Ets factor). | |||||
|
ELK4_HUMAN
|
||||||
| NC score | 0.860759 (rank : 22) | θ value | 1.16704e-23 (rank : 26) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P28324, P28323 | Gene names | ELK4, SAP1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-4 (Serum response factor accessory protein 1) (SAP-1). | |||||
|
ETV1_MOUSE
|
||||||
| NC score | 0.849210 (rank : 23) | θ value | 8.63488e-27 (rank : 16) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P41164 | Gene names | Etv1, Er81, Etsrp81 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 1 (ER81 protein). | |||||
|
ETV1_HUMAN
|
||||||
| NC score | 0.848320 (rank : 24) | θ value | 8.63488e-27 (rank : 15) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P50549, O75849, Q9UQ71, Q9Y636 | Gene names | ETV1, ER81 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 1 (ER81 protein). | |||||
|
SPDEF_MOUSE
|
||||||
| NC score | 0.844817 (rank : 25) | θ value | 2.69047e-20 (rank : 28) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9WTP3 | Gene names | Spdef, Pdef, Pse | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SAM pointed domain-containing Ets transcription factor (Prostate- derived Ets factor) (Prostate epithelium-specific Ets transcription factor) (Prostate-specific Ets). | |||||
|
ETV4_MOUSE
|
||||||
| NC score | 0.843414 (rank : 26) | θ value | 1.05554e-24 (rank : 23) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P28322 | Gene names | Etv4, Pea-3, Pea3 | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 4 (Polyomavirus enhancer activator 3) (Protein PEA3). | |||||
|
ETV4_HUMAN
|
||||||
| NC score | 0.843056 (rank : 27) | θ value | 1.05554e-24 (rank : 22) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P43268, Q96AW9 | Gene names | ETV4, E1AF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 4 (Adenovirus E1A enhancer-binding protein) (E1A-F). | |||||
|
ETV3_HUMAN
|
||||||
| NC score | 0.842441 (rank : 28) | θ value | 5.06226e-27 (rank : 14) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P41162, Q8TAC8, Q9BX30 | Gene names | ETV3, METS, PE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 3 (ETS-domain transcriptional repressor PE1) (PE-1) (Mitogenic Ets transcriptional suppressor). | |||||
|
ETV5_HUMAN
|
||||||
| NC score | 0.841862 (rank : 29) | θ value | 1.24688e-25 (rank : 19) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P41161 | Gene names | ETV5, ERM | |||
|
Domain Architecture |

|
|||||
| Description | ETS translocation variant 5 (Ets-related protein ERM). | |||||
|
SPDEF_HUMAN
|
||||||
| NC score | 0.839816 (rank : 30) | θ value | 5.43371e-13 (rank : 40) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O95238 | Gene names | SPDEF, PDEF, PSE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SAM pointed domain-containing Ets transcription factor (Prostate- derived Ets factor) (Prostate epithelium-specific Ets transcription factor) (Prostate-specific Ets). | |||||
|
ETV5_MOUSE
|
||||||
| NC score | 0.823837 (rank : 31) | θ value | 1.24688e-25 (rank : 20) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9CXC9, Q3TG49, Q8C0F3, Q9JHB1 | Gene names | Etv5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS translocation variant 5. | |||||
|
ELF5_MOUSE
|
||||||
| NC score | 0.804917 (rank : 32) | θ value | 1.42992e-13 (rank : 39) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8VDK3, Q921H5, Q9Z2K6 | Gene names | Elf5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-5 (E74-like factor 5). | |||||
|
ETV6_MOUSE
|
||||||
| NC score | 0.804678 (rank : 33) | θ value | 2.20605e-14 (rank : 38) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P97360 | Gene names | Etv6, Tel, Tel1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor ETV6 (ETS-related protein Tel1) (Tel) (ETS translocation variant 6). | |||||
|
ELF5_HUMAN
|
||||||
| NC score | 0.801301 (rank : 34) | θ value | 6.64225e-11 (rank : 41) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9UKW6, O95175, Q8N2K9, Q96QY3, Q9UKW5 | Gene names | ELF5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-5 (E74-like factor 5) (Epithelium-specific Ets transcription factor 2) (ESE-2) (Epithelium- restricted ESE-1-related Ets factor). | |||||
|
ETV6_HUMAN
|
||||||
| NC score | 0.799950 (rank : 35) | θ value | 2.20605e-14 (rank : 37) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P41212, Q9UMF6 | Gene names | ETV6, TEL, TEL1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor ETV6 (ETS-related protein Tel1) (Tel) (ETS translocation variant 6). | |||||
|
ELF2_MOUSE
|
||||||
| NC score | 0.798358 (rank : 36) | θ value | 4.44505e-15 (rank : 34) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9JHC9, Q6NST2, Q8BTX8, Q9JHC7, Q9JHC8, Q9JHD0 | Gene names | Elf2, Nerf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-2 (E74-like factor 2) (New ETS- related factor). | |||||
|
ELF2_HUMAN
|
||||||
| NC score | 0.796180 (rank : 37) | θ value | 5.8054e-15 (rank : 36) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q15723, Q15724, Q15725, Q6P1K5 | Gene names | ELF2, NERF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-2 (E74-like factor 2) (New ETS- related factor). | |||||
|
ELF1_HUMAN
|
||||||
| NC score | 0.792988 (rank : 38) | θ value | 1.99529e-15 (rank : 31) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P32519, Q9UDE1 | Gene names | ELF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-1 (E74-like factor 1). | |||||
|
ELF4_HUMAN
|
||||||
| NC score | 0.791369 (rank : 39) | θ value | 4.44505e-15 (rank : 35) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q99607, O60435 | Gene names | ELF4, ELFR, MEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-4 (E74-like factor 4) (Myeloid Elf-1-like factor). | |||||
|
ELF1_MOUSE
|
||||||
| NC score | 0.786126 (rank : 40) | θ value | 1.99529e-15 (rank : 32) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q60775 | Gene names | Elf1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS-related transcription factor Elf-1 (E74-like factor 1). | |||||
|
ELF4_MOUSE
|
||||||
| NC score | 0.771411 (rank : 41) | θ value | 3.40345e-15 (rank : 33) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9Z2U4 | Gene names | Elf4, Mef | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ETS-related transcription factor Elf-4 (E74-like factor 4) (Myeloid Elf-1-like factor). | |||||
|
SPIB_HUMAN
|
||||||
| NC score | 0.631467 (rank : 42) | θ value | 1.99992e-07 (rank : 42) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q01892, Q15359 | Gene names | SPIB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor Spi-B. | |||||
|
SPI1_HUMAN
|
||||||
| NC score | 0.612181 (rank : 43) | θ value | 0.000786445 (rank : 45) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P17947 | Gene names | SPI1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor PU.1 (31 kDa transforming protein). | |||||
|
SPI1_MOUSE
|
||||||
| NC score | 0.611772 (rank : 44) | θ value | 0.000786445 (rank : 46) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P17433, Q99L57 | Gene names | Spi1, Sfpi-1, Sfpi1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor PU.1 (31 kDa transforming protein) (SFFV proviral integration 1 protein). | |||||
|
SPIB_MOUSE
|
||||||
| NC score | 0.608833 (rank : 45) | θ value | 0.000461057 (rank : 44) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O35906, O35907, O35909, O55199 | Gene names | Spib | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor Spi-B. | |||||
|
SPIC_HUMAN
|
||||||
| NC score | 0.590881 (rank : 46) | θ value | 3.19293e-05 (rank : 43) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8N5J4 | Gene names | SPIC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor Spi-C. | |||||
|
SPIC_MOUSE
|
||||||
| NC score | 0.560360 (rank : 47) | θ value | 0.000786445 (rank : 47) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q6P3D7, Q9Z0Y0 | Gene names | Spic | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor Spi-C (Pu.1-related factor) (Prf). | |||||
|
TFCP2_MOUSE
|
||||||
| NC score | 0.052792 (rank : 48) | θ value | 0.0113563 (rank : 49) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9ERA0, Q3UGJ4, Q8VC13 | Gene names | Tcfcp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alpha-globin transcription factor CP2. | |||||
|
TFCP2_HUMAN
|
||||||
| NC score | 0.052777 (rank : 49) | θ value | 0.0113563 (rank : 48) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q12800, Q12801, Q9UD77 | Gene names | TFCP2, LSF, SEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alpha-globin transcription factor CP2 (Transcription factor LSF) (SAA3 enhancer factor). | |||||
|
TF2L1_MOUSE
|
||||||
| NC score | 0.035572 (rank : 50) | θ value | 3.0926 (rank : 62) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q3UNW5, Q99PF2 | Gene names | Tcfcp2l1, Crtr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor CP2-like protein 1 (CP2-related transcriptional repressor 1) (CRTR-1). | |||||
|
WIRE_HUMAN
|
||||||
| NC score | 0.021357 (rank : 51) | θ value | 0.813845 (rank : 52) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8TF74, Q658J8, Q71RE1, Q8TE44 | Gene names | WIRE, WICH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WIP-related protein (WASP-interacting protein-related protein) (WIP- and CR16-homologous protein). | |||||
|
GLUD1_MOUSE
|
||||||
| NC score | 0.020717 (rank : 52) | θ value | 1.06291 (rank : 53) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CIX8 | Gene names | Gluld1, Lgs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate--ammonia ligase domain-containing protein 1 (Lengsin) (Lens glutamine synthase-like). | |||||
|
CPSF7_MOUSE
|
||||||
| NC score | 0.017586 (rank : 53) | θ value | 5.27518 (rank : 69) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BTV2, Q3TNF1, Q8BKE7, Q8CFS8 | Gene names | Cpsf7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage and polyadenylation specificity factor 7. | |||||
|
CPSF7_HUMAN
|
||||||
| NC score | 0.017502 (rank : 54) | θ value | 5.27518 (rank : 68) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8N684, Q7Z3H9, Q9H025, Q9H9V1 | Gene names | CPSF7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cleavage and polyadenylation specificity factor 7 (Cleavage and polyadenylation specificity factor 59 kDa subunit) (CPSF 59 kDa subunit) (Pre-mRNA cleavage factor Im 59 kDa subunit). | |||||
|
WASF2_MOUSE
|
||||||
| NC score | 0.017112 (rank : 55) | θ value | 4.03905 (rank : 66) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8BH43 | Gene names | Wasf2, Wave2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein family member 2 (WASP-family protein member 2) (Protein WAVE-2). | |||||
|
NDST4_MOUSE
|
||||||
| NC score | 0.016239 (rank : 56) | θ value | 0.62314 (rank : 51) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9EQW8 | Gene names | Ndst4, Hsst4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bifunctional heparan sulfate N-deacetylase/N-sulfotransferase 4 (EC 2.8.2.8) (Glucosaminyl N-deacetylase/N-sulfotransferase 4) (NDST- 4) (N-heparan sulfate sulfotransferase 4) (N-HSST 4) [Includes: Heparan sulfate N-deacetylase 4 (EC 3.-.-.-); Heparan sulfate N- sulfotransferase 4 (EC 2.8.2.-)]. | |||||
|
BCL9_HUMAN
|
||||||
| NC score | 0.015847 (rank : 57) | θ value | 6.88961 (rank : 73) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1008 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O00512 | Gene names | BCL9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9) (Legless homolog). | |||||
|
AHI1_HUMAN
|
||||||
| NC score | 0.015716 (rank : 58) | θ value | 0.125558 (rank : 50) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 537 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8N157, Q504T3, Q5TCP9, Q6P098, Q6PIT6, Q8NDX0, Q9H0H2 | Gene names | AHI1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Jouberin (Abelson helper integration site 1 protein homolog) (AHI-1). | |||||
|
WIRE_MOUSE
|
||||||
| NC score | 0.015564 (rank : 59) | θ value | 8.99809 (rank : 85) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6PEV3 | Gene names | Wire | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WIP-related protein (WASP-interacting protein-related protein). | |||||
|
PHLA1_MOUSE
|
||||||
| NC score | 0.015440 (rank : 60) | θ value | 8.99809 (rank : 83) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q62392, Q3TY48, Q3UG87 | Gene names | Phlda1, Tdag51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family A member 1 (T-cell death- associated gene 51 protein) (Proline- and glutamine-rich protein) (PQR protein). | |||||
|
BCL9_MOUSE
|
||||||
| NC score | 0.013837 (rank : 61) | θ value | 5.27518 (rank : 67) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 770 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9D219, Q67FX9, Q8BUJ8, Q8VE74 | Gene names | Bcl9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma 9 protein (Bcl-9). | |||||
|
GP137_HUMAN
|
||||||
| NC score | 0.013345 (rank : 62) | θ value | 8.99809 (rank : 79) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q14444, Q15074 | Gene names | M11S1, GPIP137 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GPI-anchored protein p137 (p137GPI) (Membrane component chromosome 11 surface marker 1). | |||||
|
ZN645_HUMAN
|
||||||
| NC score | 0.013217 (rank : 63) | θ value | 8.99809 (rank : 86) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 265 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8N7E2, Q6DJY9 | Gene names | ZNF645 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 645. | |||||
|
PARM1_MOUSE
|
||||||
| NC score | 0.013001 (rank : 64) | θ value | 4.03905 (rank : 64) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q923D3, Q3TTV0, Q3UFU4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein PARM-1 precursor. | |||||
|
EP300_HUMAN
|
||||||
| NC score | 0.011874 (rank : 65) | θ value | 1.38821 (rank : 54) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q09472 | Gene names | EP300, P300 | |||
|
Domain Architecture |

|
|||||
| Description | E1A-associated protein p300 (EC 2.3.1.48). | |||||
|
MDC1_HUMAN
|
||||||
| NC score | 0.011872 (rank : 66) | θ value | 8.99809 (rank : 82) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
RTN1_HUMAN
|
||||||
| NC score | 0.011806 (rank : 67) | θ value | 5.27518 (rank : 72) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q16799, Q16800, Q16801 | Gene names | RTN1, NSP | |||
|
Domain Architecture |

|
|||||
| Description | Reticulon-1 (Neuroendocrine-specific protein). | |||||
|
3BP1_HUMAN
|
||||||
| NC score | 0.011619 (rank : 68) | θ value | 3.0926 (rank : 58) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 376 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y3L3, Q6IBZ2, Q6ZVL9, Q96HQ5, Q9NSQ9 | Gene names | SH3BP1 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 domain-binding protein 1 (3BP-1). | |||||
|
TCF7_MOUSE
|
||||||
| NC score | 0.011308 (rank : 69) | θ value | 6.88961 (rank : 76) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q00417 | Gene names | Tcf7, Tcf-1, Tcf1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 7 (T-cell-specific transcription factor 1) (TCF- 1) (T-cell factor 1). | |||||
|
NDST4_HUMAN
|
||||||
| NC score | 0.010755 (rank : 70) | θ value | 3.0926 (rank : 61) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H3R1 | Gene names | NDST4, HSST4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bifunctional heparan sulfate N-deacetylase/N-sulfotransferase 4 (EC 2.8.2.8) (Glucosaminyl N-deacetylase/N-sulfotransferase 4) (NDST- 4) (N-heparan sulfate sulfotransferase 4) (N-HSST 4) [Includes: Heparan sulfate N-deacetylase 4 (EC 3.-.-.-); Heparan sulfate N- sulfotransferase 4 (EC 2.8.2.-)]. | |||||
|
FUBP3_HUMAN
|
||||||
| NC score | 0.010209 (rank : 71) | θ value | 6.88961 (rank : 74) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96I24, Q92946, Q9BVB6 | Gene names | FUBP3 | |||
|
Domain Architecture |

|
|||||
| Description | Far upstream element-binding protein 3 (FUSE-binding protein 3). | |||||
|
JPH2_MOUSE
|
||||||
| NC score | 0.010112 (rank : 72) | θ value | 2.36792 (rank : 56) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9ET78, Q9ET79 | Gene names | Jph2, Jp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junctophilin-2 (Junctophilin type 2) (JP-2). | |||||
|
FOXN1_HUMAN
|
||||||
| NC score | 0.010076 (rank : 73) | θ value | 4.03905 (rank : 63) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 377 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O15353, O15352 | Gene names | FOXN1, WHN | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein N1 (Transcription factor winged-helix nude). | |||||
|
WASF2_HUMAN
|
||||||
| NC score | 0.009976 (rank : 74) | θ value | 4.03905 (rank : 65) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Y6W5, O60794, Q9UDY7 | Gene names | WASF2, WAVE2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein family member 2 (WASP-family protein member 2) (Protein WAVE-2) (Verprolin homology domain-containing protein 2). | |||||
|
REC8L_MOUSE
|
||||||
| NC score | 0.008056 (rank : 75) | θ value | 8.99809 (rank : 84) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8C5S7, Q3UIN9, Q9JK52 | Gene names | Rec8L1, Rec8 | |||
|
Domain Architecture |

|
|||||
| Description | Meiotic recombination protein REC8-like 1 (Cohesin Rec8p). | |||||
|
PHC1_MOUSE
|
||||||
| NC score | 0.007895 (rank : 76) | θ value | 5.27518 (rank : 71) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q64028, P70359, Q64307, Q8BZ80 | Gene names | Phc1, Edr, Edr1, Rae28 | |||
|
Domain Architecture |

|
|||||
| Description | Polyhomeotic-like protein 1 (mPH1) (Early development regulatory protein 1) (RAE-28). | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.006578 (rank : 77) | θ value | 2.36792 (rank : 57) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
MYO15_MOUSE
|
||||||
| NC score | 0.005037 (rank : 78) | θ value | 3.0926 (rank : 60) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9QZZ4, O70395, Q9QWL6 | Gene names | Myo15a, Myo15 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-15 (Myosin XV) (Unconventional myosin-15). | |||||
|
BAZ2A_MOUSE
|
||||||
| NC score | 0.004601 (rank : 79) | θ value | 8.99809 (rank : 77) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q91YE5 | Gene names | Baz2a, Tip5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5). | |||||
|
SOX10_MOUSE
|
||||||
| NC score | 0.002533 (rank : 80) | θ value | 6.88961 (rank : 75) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q04888, O08518, O09141, O54856, P70416 | Gene names | Sox10, Sox-10, Sox21 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-10 (SOX-21) (Transcription factor SOX-M). | |||||
|
LHX5_HUMAN
|
||||||
| NC score | 0.000327 (rank : 81) | θ value | 5.27518 (rank : 70) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 453 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H2C1 | Gene names | LHX5 | |||
|
Domain Architecture |

|
|||||
| Description | LIM/homeobox protein Lhx5. | |||||
|
KLH17_HUMAN
|
||||||
| NC score | -0.000871 (rank : 82) | θ value | 8.99809 (rank : 80) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 201 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6TDP4 | Gene names | KLHL17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kelch-like protein 17 (Actinfilin). | |||||
|
CELR3_MOUSE
|
||||||
| NC score | -0.001720 (rank : 83) | θ value | 8.99809 (rank : 78) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 687 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q91ZI0, Q9ESD0 | Gene names | Celsr3 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin EGF LAG seven-pass G-type receptor 3 precursor. | |||||
|
M3K2_MOUSE
|
||||||
| NC score | -0.003036 (rank : 84) | θ value | 1.81305 (rank : 55) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 846 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q61083 | Gene names | Map3k2, Mekk2 | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase 2 (EC 2.7.11.25) (MAPK/ERK kinase kinase 2) (MEK kinase 2) (MEKK 2). | |||||
|
M3K2_HUMAN
|
||||||
| NC score | -0.003890 (rank : 85) | θ value | 8.99809 (rank : 81) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 849 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y2U5, Q53QL9, Q53S75, Q59GZ6, Q8NC32, Q9NYK3 | Gene names | MAP3K2, MAPKKK2, MEKK2 | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase 2 (EC 2.7.11.25) (MAPK/ERK kinase kinase 2) (MEK kinase 2) (MEKK 2). | |||||
|
LIMK1_MOUSE
|
||||||
| NC score | -0.004071 (rank : 86) | θ value | 3.0926 (rank : 59) | |||
| Query Neighborhood Hits | 86 | Target Neighborhood Hits | 1008 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P53668 | Gene names | Limk1, Limk | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain kinase 1 (EC 2.7.11.1) (LIMK-1) (KIZ-1). | |||||