Please be patient as the page loads
|
REC8L_MOUSE
|
||||||
| SwissProt Accessions | Q8C5S7, Q3UIN9, Q9JK52 | Gene names | Rec8L1, Rec8 | |||
|
Domain Architecture |

|
|||||
| Description | Meiotic recombination protein REC8-like 1 (Cohesin Rec8p). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
REC8L_MOUSE
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8C5S7, Q3UIN9, Q9JK52 | Gene names | Rec8L1, Rec8 | |||
|
Domain Architecture |

|
|||||
| Description | Meiotic recombination protein REC8-like 1 (Cohesin Rec8p). | |||||
|
REC8L_HUMAN
|
||||||
| θ value | 5.9077e-185 (rank : 2) | NC score | 0.956914 (rank : 2) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O95072, Q8WUV8, Q9BTF2, Q9NVQ9 | Gene names | REC8L1, REC8 | |||
|
Domain Architecture |

|
|||||
| Description | Meiotic recombination protein REC8-like 1 (Cohesin Rec8p). | |||||
|
RAD21_MOUSE
|
||||||
| θ value | 1.87187e-05 (rank : 3) | NC score | 0.289241 (rank : 3) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q61550, P70219, Q91VB9, Q9DBU4 | Gene names | Rad21, Hr21 | |||
|
Domain Architecture |

|
|||||
| Description | Double-strand-break repair protein rad21 homolog (Pokeweed agglutinin- binding protein 29) (PW29) (SCC1 homolog). | |||||
|
RAD21_HUMAN
|
||||||
| θ value | 7.1131e-05 (rank : 4) | NC score | 0.285167 (rank : 4) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O60216, Q15001, Q99568 | Gene names | RAD21, HR21, KIAA0078, NXP1 | |||
|
Domain Architecture |

|
|||||
| Description | Double-strand-break repair protein rad21 homolog (hHR21) (Nuclear matrix protein 1) (NXP-1) (SCC1 homolog). | |||||
|
EP300_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 5) | NC score | 0.059044 (rank : 6) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q09472 | Gene names | EP300, P300 | |||
|
Domain Architecture |

|
|||||
| Description | E1A-associated protein p300 (EC 2.3.1.48). | |||||
|
AFF2_MOUSE
|
||||||
| θ value | 0.125558 (rank : 6) | NC score | 0.053637 (rank : 9) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O55112 | Gene names | Aff2, Fmr2, Ox19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation protein 2 homolog) (Protein FMR-2) (FMR2P) (Protein Ox19). | |||||
|
UN13A_HUMAN
|
||||||
| θ value | 0.21417 (rank : 7) | NC score | 0.041763 (rank : 12) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UPW8 | Gene names | UNC13A, KIAA1032 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Unc-13 homolog A (Munc13-1). | |||||
|
HCN4_HUMAN
|
||||||
| θ value | 0.47712 (rank : 8) | NC score | 0.025150 (rank : 23) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y3Q4, Q9UMQ7 | Gene names | HCN4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 4. | |||||
|
RGS3_HUMAN
|
||||||
| θ value | 0.47712 (rank : 9) | NC score | 0.032245 (rank : 14) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
PRR12_HUMAN
|
||||||
| θ value | 0.62314 (rank : 10) | NC score | 0.054054 (rank : 8) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9ULL5, Q8N4J6 | Gene names | PRR12, KIAA1205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 12. | |||||
|
EP400_HUMAN
|
||||||
| θ value | 1.06291 (rank : 11) | NC score | 0.031576 (rank : 15) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q96L91, O15411, Q6P2F5, Q8N8Q7, Q8NE05, Q96JK7, Q9P230 | Gene names | EP400, CAGH32, KIAA1498, KIAA1818, TNRC12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E1A-binding protein p400 (EC 3.6.1.-) (p400 kDa SWI2/SNF2-related protein) (Domino homolog) (hDomino) (CAG repeat protein 32) (Trinucleotide repeat-containing gene 12 protein). | |||||
|
APXL_HUMAN
|
||||||
| θ value | 1.38821 (rank : 12) | NC score | 0.036675 (rank : 13) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 553 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q13796 | Gene names | APXL | |||
|
Domain Architecture |
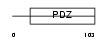
|
|||||
| Description | Apical-like protein (Protein APXL). | |||||
|
BRD4_HUMAN
|
||||||
| θ value | 1.38821 (rank : 13) | NC score | 0.031111 (rank : 17) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O60885, O60433, Q96PD3 | Gene names | BRD4, HUNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 4 (HUNK1 protein). | |||||
|
FOXP1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 14) | NC score | 0.016866 (rank : 29) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9H334, Q9H332, Q9H333, Q9P0R1 | Gene names | FOXP1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P1. | |||||
|
1B52_HUMAN
|
||||||
| θ value | 1.81305 (rank : 15) | NC score | 0.006622 (rank : 39) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P30490, Q9MY75, Q9TPT4 | Gene names | HLA-B, HLAB | |||
|
Domain Architecture |
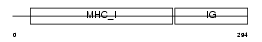
|
|||||
| Description | HLA class I histocompatibility antigen, B-52 alpha chain precursor (MHC class I antigen B*52) (Bw-52) (B-5). | |||||
|
CUTL1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 16) | NC score | 0.014056 (rank : 31) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P53564, O08994, P70301, Q571L6, Q91ZD2 | Gene names | Cutl1, Cux, Cux1, Kiaa4047 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP) (Homeobox protein Cux). | |||||
|
PELP1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 17) | NC score | 0.046721 (rank : 10) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 596 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8IZL8, O15450, Q5EGN3, Q6NTE6, Q96FT1, Q9BU60 | Gene names | PELP1, HMX3, MNAR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor) (Transcription factor HMX3). | |||||
|
ZCHC5_HUMAN
|
||||||
| θ value | 1.81305 (rank : 18) | NC score | 0.054574 (rank : 7) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8N8U3 | Gene names | ZCCHC5 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger CCHC domain-containing protein 5. | |||||
|
RAPH1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 19) | NC score | 0.030222 (rank : 18) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q70E73, Q96Q37, Q9C0I2 | Gene names | RAPH1, ALS2CR9, KIAA1681, LPD, PREL2, RMO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras-associated and pleckstrin homology domains-containing protein 1 (RAPH1) (Lamellipodin) (Proline-rich EVH1 ligand 2) (PREL-2) (Protein RMO1) (Amyotrophic lateral sclerosis 2 chromosomal region candidate 9 gene protein). | |||||
|
ZN544_HUMAN
|
||||||
| θ value | 2.36792 (rank : 20) | NC score | 0.001293 (rank : 44) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6NX49, Q9UEX4 | Gene names | ZNF544 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 544. | |||||
|
PCLO_HUMAN
|
||||||
| θ value | 3.0926 (rank : 21) | NC score | 0.023402 (rank : 25) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
PELP1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 22) | NC score | 0.043594 (rank : 11) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9DBD5, Q5F2E2, Q6PEM0, Q91YM9 | Gene names | Pelp1, Mnar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor). | |||||
|
DDEF1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 23) | NC score | 0.012925 (rank : 33) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 592 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9QWY8, O08612, Q80TG8, Q80UV6, Q99LV8, Q9Z2B6 | Gene names | Ddef1, Asap1, Kiaa1249, Shag1 | |||
|
Domain Architecture |

|
|||||
| Description | 130 kDa phosphatidylinositol 4,5-biphosphate-dependent ARF1 GTPase- activating protein (PIP2-dependent ARF1 GAP) (ADP-ribosylation factor- directed GTPase-activating protein 1) (ARF GTPase-activating protein 1) (Development and differentiation-enhancing factor 1) (Differentiation-enhancing factor 1) (DEF-1). | |||||
|
BUB1B_HUMAN
|
||||||
| θ value | 5.27518 (rank : 24) | NC score | 0.019673 (rank : 27) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O60566, O60501, O60627, O60758, O75389, Q96KM4 | Gene names | BUB1B, BUBR1, MAD3L | |||
|
Domain Architecture |

|
|||||
| Description | Mitotic checkpoint serine/threonine-protein kinase BUB1 beta (EC 2.7.11.1) (hBUBR1) (MAD3/BUB1-related protein kinase) (Mitotic checkpoint kinase MAD3L) (SSK1). | |||||
|
CDT1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 25) | NC score | 0.026284 (rank : 22) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H211, Q86XX9, Q96CJ5, Q96GK5, Q96H67, Q96HE6, Q9BWM0 | Gene names | CDT1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA replication factor Cdt1 (Double parked homolog) (DUP). | |||||
|
CN102_MOUSE
|
||||||
| θ value | 5.27518 (rank : 26) | NC score | 0.031552 (rank : 16) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80XC6, Q8R3D7, Q99LT9 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C14orf102 homolog. | |||||
|
DCC_MOUSE
|
||||||
| θ value | 5.27518 (rank : 27) | NC score | 0.008399 (rank : 37) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P70211 | Gene names | Dcc | |||
|
Domain Architecture |

|
|||||
| Description | Netrin receptor DCC precursor (Tumor suppressor protein DCC). | |||||
|
EP400_MOUSE
|
||||||
| θ value | 5.27518 (rank : 28) | NC score | 0.027142 (rank : 20) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 592 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8CHI8, Q80TC8, Q8BXI5, Q8BYW3, Q8C0P6, Q8CHI7, Q8VDF4, Q9DA54 | Gene names | Ep400, Kiaa1498 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E1A-binding protein p400 (EC 3.6.1.-) (p400 kDa SWI2/SNF2-related protein) (Domino homolog) (mDomino). | |||||
|
JAG2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 29) | NC score | 0.004383 (rank : 42) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y219, Q9UE17, Q9UE99, Q9UNK8, Q9Y6P9, Q9Y6Q0 | Gene names | JAG2 | |||
|
Domain Architecture |

|
|||||
| Description | Jagged-2 precursor (Jagged2) (HJ2). | |||||
|
KLF13_MOUSE
|
||||||
| θ value | 5.27518 (rank : 30) | NC score | 0.002716 (rank : 43) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 757 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9JJZ6, Q9ESX3, Q9JHF8 | Gene names | Klf13, Bteb3, Fklf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Krueppel-like factor 13 (Transcription factor BTEB3) (Basic transcription element-binding protein 3) (BTE-binding protein 3) (RANTES factor of late activated T-lymphocytes 1) (RFLAT-1) (Erythroid transcription factor FKLF-2). | |||||
|
MRE11_MOUSE
|
||||||
| θ value | 5.27518 (rank : 31) | NC score | 0.021928 (rank : 26) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61216, Q62430 | Gene names | Mre11a, Mre11 | |||
|
Domain Architecture |
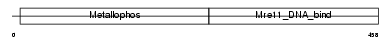
|
|||||
| Description | Double-strand break repair protein MRE11A (MRE11 homolog 1) (MRE11 meiotic recombination 11 homolog A) (MmMRE11A). | |||||
|
SDCG8_MOUSE
|
||||||
| θ value | 5.27518 (rank : 32) | NC score | 0.010372 (rank : 36) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 886 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q80UF4 | Gene names | Sdccag8, Cccap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serologically defined colon cancer antigen 8 (Centrosomal colon cancer autoantigen protein) (mCCCAP). | |||||
|
SRRM1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 33) | NC score | 0.024763 (rank : 24) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
TCAL5_HUMAN
|
||||||
| θ value | 5.27518 (rank : 34) | NC score | 0.013778 (rank : 32) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 806 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q5H9L2 | Gene names | TCEAL5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 5 (TCEA-like protein 5) (Transcription elongation factor S-II protein-like 5). | |||||
|
1B37_HUMAN
|
||||||
| θ value | 6.88961 (rank : 35) | NC score | 0.005041 (rank : 40) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P18463, O19627, Q95HA3, Q95HA8, Q95HM9, Q9GJ31 | Gene names | HLA-B, HLAB | |||
|
Domain Architecture |
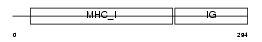
|
|||||
| Description | HLA class I histocompatibility antigen, B-37 alpha chain precursor (MHC class I antigen B*37). | |||||
|
CN102_HUMAN
|
||||||
| θ value | 6.88961 (rank : 36) | NC score | 0.029864 (rank : 19) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H7Z3, Q9NWH6 | Gene names | C14orf102 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C14orf102. | |||||
|
ERCC5_MOUSE
|
||||||
| θ value | 6.88961 (rank : 37) | NC score | 0.018550 (rank : 28) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P35689, Q61528, Q64248 | Gene names | Ercc5, Ercc-5, Xpg | |||
|
Domain Architecture |

|
|||||
| Description | DNA-repair protein complementing XP-G cells homolog (Xeroderma pigmentosum group G-complementing protein homolog) (DNA excision repair protein ERCC-5). | |||||
|
KIF4A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 38) | NC score | 0.004511 (rank : 41) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 1159 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O95239, Q86TN3, Q86XX7, Q9NNY6, Q9NY24, Q9UMW3 | Gene names | KIF4A, KIF4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromosome-associated kinesin KIF4A (Chromokinesin). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | 6.88961 (rank : 39) | NC score | 0.026689 (rank : 21) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
SNRPA_HUMAN
|
||||||
| θ value | 6.88961 (rank : 40) | NC score | 0.015666 (rank : 30) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P09012 | Gene names | SNRPA | |||
|
Domain Architecture |

|
|||||
| Description | U1 small nuclear ribonucleoprotein A (U1 snRNP protein A) (U1A protein) (U1-A). | |||||
|
B3GA3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 41) | NC score | 0.012266 (rank : 34) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P58158 | Gene names | B3gat3 | |||
|
Domain Architecture |
|
|||||
| Description | Galactosylgalactosylxylosylprotein 3-beta-glucuronosyltransferase 3 (EC 2.4.1.135) (Beta-1,3-glucuronyltransferase 3) (Glucuronosyltransferase-I) (GlcAT-I) (UDP-GlcUA:Gal Beta-1,3-Gal-R glucuronyltransferase) (GlcUAT-I). | |||||
|
ERG_HUMAN
|
||||||
| θ value | 8.99809 (rank : 42) | NC score | 0.008056 (rank : 38) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P11308, Q16113 | Gene names | ERG | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ERG (Transforming protein ERG). | |||||
|
MORC4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 43) | NC score | 0.011574 (rank : 35) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 244 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8TE76, Q5JUK7, Q96MZ2, Q9HAI7 | Gene names | MORC4, ZCWCC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORC family CW-type zinc finger protein 4 (Zinc finger CW-type coiled- coil domain protein 2). | |||||
|
PRAX_HUMAN
|
||||||
| θ value | 8.99809 (rank : 44) | NC score | 0.074394 (rank : 5) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9BXM0, Q9BXL9, Q9HCF2 | Gene names | PRX, KIAA1620 | |||
|
Domain Architecture |
|
|||||
| Description | Periaxin. | |||||
|
REC8L_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8C5S7, Q3UIN9, Q9JK52 | Gene names | Rec8L1, Rec8 | |||
|
Domain Architecture |

|
|||||
| Description | Meiotic recombination protein REC8-like 1 (Cohesin Rec8p). | |||||
|
REC8L_HUMAN
|
||||||
| NC score | 0.956914 (rank : 2) | θ value | 5.9077e-185 (rank : 2) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O95072, Q8WUV8, Q9BTF2, Q9NVQ9 | Gene names | REC8L1, REC8 | |||
|
Domain Architecture |

|
|||||
| Description | Meiotic recombination protein REC8-like 1 (Cohesin Rec8p). | |||||
|
RAD21_MOUSE
|
||||||
| NC score | 0.289241 (rank : 3) | θ value | 1.87187e-05 (rank : 3) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q61550, P70219, Q91VB9, Q9DBU4 | Gene names | Rad21, Hr21 | |||
|
Domain Architecture |

|
|||||
| Description | Double-strand-break repair protein rad21 homolog (Pokeweed agglutinin- binding protein 29) (PW29) (SCC1 homolog). | |||||
|
RAD21_HUMAN
|
||||||
| NC score | 0.285167 (rank : 4) | θ value | 7.1131e-05 (rank : 4) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O60216, Q15001, Q99568 | Gene names | RAD21, HR21, KIAA0078, NXP1 | |||
|
Domain Architecture |

|
|||||
| Description | Double-strand-break repair protein rad21 homolog (hHR21) (Nuclear matrix protein 1) (NXP-1) (SCC1 homolog). | |||||
|
PRAX_HUMAN
|
||||||
| NC score | 0.074394 (rank : 5) | θ value | 8.99809 (rank : 44) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9BXM0, Q9BXL9, Q9HCF2 | Gene names | PRX, KIAA1620 | |||
|
Domain Architecture |

|
|||||
| Description | Periaxin. | |||||
|
EP300_HUMAN
|
||||||
| NC score | 0.059044 (rank : 6) | θ value | 0.0148317 (rank : 5) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q09472 | Gene names | EP300, P300 | |||
|
Domain Architecture |

|
|||||
| Description | E1A-associated protein p300 (EC 2.3.1.48). | |||||
|
ZCHC5_HUMAN
|
||||||
| NC score | 0.054574 (rank : 7) | θ value | 1.81305 (rank : 18) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8N8U3 | Gene names | ZCCHC5 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger CCHC domain-containing protein 5. | |||||
|
PRR12_HUMAN
|
||||||
| NC score | 0.054054 (rank : 8) | θ value | 0.62314 (rank : 10) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9ULL5, Q8N4J6 | Gene names | PRR12, KIAA1205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 12. | |||||
|
AFF2_MOUSE
|
||||||
| NC score | 0.053637 (rank : 9) | θ value | 0.125558 (rank : 6) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O55112 | Gene names | Aff2, Fmr2, Ox19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation protein 2 homolog) (Protein FMR-2) (FMR2P) (Protein Ox19). | |||||
|
PELP1_HUMAN
|
||||||
| NC score | 0.046721 (rank : 10) | θ value | 1.81305 (rank : 17) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 596 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8IZL8, O15450, Q5EGN3, Q6NTE6, Q96FT1, Q9BU60 | Gene names | PELP1, HMX3, MNAR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor) (Transcription factor HMX3). | |||||
|
PELP1_MOUSE
|
||||||
| NC score | 0.043594 (rank : 11) | θ value | 3.0926 (rank : 22) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9DBD5, Q5F2E2, Q6PEM0, Q91YM9 | Gene names | Pelp1, Mnar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor). | |||||
|
UN13A_HUMAN
|
||||||
| NC score | 0.041763 (rank : 12) | θ value | 0.21417 (rank : 7) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UPW8 | Gene names | UNC13A, KIAA1032 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Unc-13 homolog A (Munc13-1). | |||||
|
APXL_HUMAN
|
||||||
| NC score | 0.036675 (rank : 13) | θ value | 1.38821 (rank : 12) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 553 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q13796 | Gene names | APXL | |||
|
Domain Architecture |

|
|||||
| Description | Apical-like protein (Protein APXL). | |||||
|
RGS3_HUMAN
|
||||||
| NC score | 0.032245 (rank : 14) | θ value | 0.47712 (rank : 9) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
EP400_HUMAN
|
||||||
| NC score | 0.031576 (rank : 15) | θ value | 1.06291 (rank : 11) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q96L91, O15411, Q6P2F5, Q8N8Q7, Q8NE05, Q96JK7, Q9P230 | Gene names | EP400, CAGH32, KIAA1498, KIAA1818, TNRC12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E1A-binding protein p400 (EC 3.6.1.-) (p400 kDa SWI2/SNF2-related protein) (Domino homolog) (hDomino) (CAG repeat protein 32) (Trinucleotide repeat-containing gene 12 protein). | |||||
|
CN102_MOUSE
|
||||||
| NC score | 0.031552 (rank : 16) | θ value | 5.27518 (rank : 26) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80XC6, Q8R3D7, Q99LT9 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C14orf102 homolog. | |||||
|
BRD4_HUMAN
|
||||||
| NC score | 0.031111 (rank : 17) | θ value | 1.38821 (rank : 13) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O60885, O60433, Q96PD3 | Gene names | BRD4, HUNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 4 (HUNK1 protein). | |||||
|
RAPH1_HUMAN
|
||||||
| NC score | 0.030222 (rank : 18) | θ value | 2.36792 (rank : 19) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q70E73, Q96Q37, Q9C0I2 | Gene names | RAPH1, ALS2CR9, KIAA1681, LPD, PREL2, RMO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras-associated and pleckstrin homology domains-containing protein 1 (RAPH1) (Lamellipodin) (Proline-rich EVH1 ligand 2) (PREL-2) (Protein RMO1) (Amyotrophic lateral sclerosis 2 chromosomal region candidate 9 gene protein). | |||||
|
CN102_HUMAN
|
||||||
| NC score | 0.029864 (rank : 19) | θ value | 6.88961 (rank : 36) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H7Z3, Q9NWH6 | Gene names | C14orf102 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C14orf102. | |||||
|
EP400_MOUSE
|
||||||
| NC score | 0.027142 (rank : 20) | θ value | 5.27518 (rank : 28) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 592 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8CHI8, Q80TC8, Q8BXI5, Q8BYW3, Q8C0P6, Q8CHI7, Q8VDF4, Q9DA54 | Gene names | Ep400, Kiaa1498 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E1A-binding protein p400 (EC 3.6.1.-) (p400 kDa SWI2/SNF2-related protein) (Domino homolog) (mDomino). | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.026689 (rank : 21) | θ value | 6.88961 (rank : 39) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
CDT1_HUMAN
|
||||||
| NC score | 0.026284 (rank : 22) | θ value | 5.27518 (rank : 25) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H211, Q86XX9, Q96CJ5, Q96GK5, Q96H67, Q96HE6, Q9BWM0 | Gene names | CDT1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA replication factor Cdt1 (Double parked homolog) (DUP). | |||||
|
HCN4_HUMAN
|
||||||
| NC score | 0.025150 (rank : 23) | θ value | 0.47712 (rank : 8) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y3Q4, Q9UMQ7 | Gene names | HCN4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 4. | |||||
|
SRRM1_MOUSE
|
||||||
| NC score | 0.024763 (rank : 24) | θ value | 5.27518 (rank : 33) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
PCLO_HUMAN
|
||||||
| NC score | 0.023402 (rank : 25) | θ value | 3.0926 (rank : 21) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
MRE11_MOUSE
|
||||||
| NC score | 0.021928 (rank : 26) | θ value | 5.27518 (rank : 31) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61216, Q62430 | Gene names | Mre11a, Mre11 | |||
|
Domain Architecture |

|
|||||
| Description | Double-strand break repair protein MRE11A (MRE11 homolog 1) (MRE11 meiotic recombination 11 homolog A) (MmMRE11A). | |||||
|
BUB1B_HUMAN
|
||||||
| NC score | 0.019673 (rank : 27) | θ value | 5.27518 (rank : 24) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O60566, O60501, O60627, O60758, O75389, Q96KM4 | Gene names | BUB1B, BUBR1, MAD3L | |||
|
Domain Architecture |

|
|||||
| Description | Mitotic checkpoint serine/threonine-protein kinase BUB1 beta (EC 2.7.11.1) (hBUBR1) (MAD3/BUB1-related protein kinase) (Mitotic checkpoint kinase MAD3L) (SSK1). | |||||
|
ERCC5_MOUSE
|
||||||
| NC score | 0.018550 (rank : 28) | θ value | 6.88961 (rank : 37) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P35689, Q61528, Q64248 | Gene names | Ercc5, Ercc-5, Xpg | |||
|
Domain Architecture |

|
|||||
| Description | DNA-repair protein complementing XP-G cells homolog (Xeroderma pigmentosum group G-complementing protein homolog) (DNA excision repair protein ERCC-5). | |||||
|
FOXP1_HUMAN
|
||||||
| NC score | 0.016866 (rank : 29) | θ value | 1.38821 (rank : 14) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9H334, Q9H332, Q9H333, Q9P0R1 | Gene names | FOXP1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P1. | |||||
|
SNRPA_HUMAN
|
||||||
| NC score | 0.015666 (rank : 30) | θ value | 6.88961 (rank : 40) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P09012 | Gene names | SNRPA | |||
|
Domain Architecture |

|
|||||
| Description | U1 small nuclear ribonucleoprotein A (U1 snRNP protein A) (U1A protein) (U1-A). | |||||
|
CUTL1_MOUSE
|
||||||
| NC score | 0.014056 (rank : 31) | θ value | 1.81305 (rank : 16) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P53564, O08994, P70301, Q571L6, Q91ZD2 | Gene names | Cutl1, Cux, Cux1, Kiaa4047 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP) (Homeobox protein Cux). | |||||
|
TCAL5_HUMAN
|
||||||
| NC score | 0.013778 (rank : 32) | θ value | 5.27518 (rank : 34) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 806 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q5H9L2 | Gene names | TCEAL5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 5 (TCEA-like protein 5) (Transcription elongation factor S-II protein-like 5). | |||||
|
DDEF1_MOUSE
|
||||||
| NC score | 0.012925 (rank : 33) | θ value | 4.03905 (rank : 23) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 592 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9QWY8, O08612, Q80TG8, Q80UV6, Q99LV8, Q9Z2B6 | Gene names | Ddef1, Asap1, Kiaa1249, Shag1 | |||
|
Domain Architecture |

|
|||||
| Description | 130 kDa phosphatidylinositol 4,5-biphosphate-dependent ARF1 GTPase- activating protein (PIP2-dependent ARF1 GAP) (ADP-ribosylation factor- directed GTPase-activating protein 1) (ARF GTPase-activating protein 1) (Development and differentiation-enhancing factor 1) (Differentiation-enhancing factor 1) (DEF-1). | |||||
|
B3GA3_MOUSE
|
||||||
| NC score | 0.012266 (rank : 34) | θ value | 8.99809 (rank : 41) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P58158 | Gene names | B3gat3 | |||
|
Domain Architecture |

|
|||||
| Description | Galactosylgalactosylxylosylprotein 3-beta-glucuronosyltransferase 3 (EC 2.4.1.135) (Beta-1,3-glucuronyltransferase 3) (Glucuronosyltransferase-I) (GlcAT-I) (UDP-GlcUA:Gal Beta-1,3-Gal-R glucuronyltransferase) (GlcUAT-I). | |||||
|
MORC4_HUMAN
|
||||||
| NC score | 0.011574 (rank : 35) | θ value | 8.99809 (rank : 43) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 244 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8TE76, Q5JUK7, Q96MZ2, Q9HAI7 | Gene names | MORC4, ZCWCC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORC family CW-type zinc finger protein 4 (Zinc finger CW-type coiled- coil domain protein 2). | |||||
|
SDCG8_MOUSE
|
||||||
| NC score | 0.010372 (rank : 36) | θ value | 5.27518 (rank : 32) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 886 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q80UF4 | Gene names | Sdccag8, Cccap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serologically defined colon cancer antigen 8 (Centrosomal colon cancer autoantigen protein) (mCCCAP). | |||||
|
DCC_MOUSE
|
||||||
| NC score | 0.008399 (rank : 37) | θ value | 5.27518 (rank : 27) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P70211 | Gene names | Dcc | |||
|
Domain Architecture |

|
|||||
| Description | Netrin receptor DCC precursor (Tumor suppressor protein DCC). | |||||
|
ERG_HUMAN
|
||||||
| NC score | 0.008056 (rank : 38) | θ value | 8.99809 (rank : 42) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P11308, Q16113 | Gene names | ERG | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ERG (Transforming protein ERG). | |||||
|
1B52_HUMAN
|
||||||
| NC score | 0.006622 (rank : 39) | θ value | 1.81305 (rank : 15) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P30490, Q9MY75, Q9TPT4 | Gene names | HLA-B, HLAB | |||
|
Domain Architecture |

|
|||||
| Description | HLA class I histocompatibility antigen, B-52 alpha chain precursor (MHC class I antigen B*52) (Bw-52) (B-5). | |||||
|
1B37_HUMAN
|
||||||
| NC score | 0.005041 (rank : 40) | θ value | 6.88961 (rank : 35) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P18463, O19627, Q95HA3, Q95HA8, Q95HM9, Q9GJ31 | Gene names | HLA-B, HLAB | |||
|
Domain Architecture |

|
|||||
| Description | HLA class I histocompatibility antigen, B-37 alpha chain precursor (MHC class I antigen B*37). | |||||
|
KIF4A_HUMAN
|
||||||
| NC score | 0.004511 (rank : 41) | θ value | 6.88961 (rank : 38) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 1159 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O95239, Q86TN3, Q86XX7, Q9NNY6, Q9NY24, Q9UMW3 | Gene names | KIF4A, KIF4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromosome-associated kinesin KIF4A (Chromokinesin). | |||||
|
JAG2_HUMAN
|
||||||
| NC score | 0.004383 (rank : 42) | θ value | 5.27518 (rank : 29) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y219, Q9UE17, Q9UE99, Q9UNK8, Q9Y6P9, Q9Y6Q0 | Gene names | JAG2 | |||
|
Domain Architecture |

|
|||||
| Description | Jagged-2 precursor (Jagged2) (HJ2). | |||||
|
KLF13_MOUSE
|
||||||
| NC score | 0.002716 (rank : 43) | θ value | 5.27518 (rank : 30) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 757 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9JJZ6, Q9ESX3, Q9JHF8 | Gene names | Klf13, Bteb3, Fklf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Krueppel-like factor 13 (Transcription factor BTEB3) (Basic transcription element-binding protein 3) (BTE-binding protein 3) (RANTES factor of late activated T-lymphocytes 1) (RFLAT-1) (Erythroid transcription factor FKLF-2). | |||||
|
ZN544_HUMAN
|
||||||
| NC score | 0.001293 (rank : 44) | θ value | 2.36792 (rank : 20) | |||
| Query Neighborhood Hits | 44 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6NX49, Q9UEX4 | Gene names | ZNF544 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 544. | |||||