Please be patient as the page loads
|
SMOC2_HUMAN
|
||||||
| SwissProt Accessions | Q9H3U7, Q5TAT7, Q5TAT8, Q86VV9, Q96SF3, Q9H1L3, Q9H1L4, Q9H3U0, Q9H4F7, Q9HCV2 | Gene names | SMOC2, SMAP2 | |||
|
Domain Architecture |
|
|||||
| Description | SPARC-related modular calcium-binding protein 2 precursor (Secreted modular calcium-binding protein 2) (SMOC-2) (Smooth muscle-associated protein 2) (SMAP-2). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
SMOC2_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 88 | |
| SwissProt Accessions | Q9H3U7, Q5TAT7, Q5TAT8, Q86VV9, Q96SF3, Q9H1L3, Q9H1L4, Q9H3U0, Q9H4F7, Q9HCV2 | Gene names | SMOC2, SMAP2 | |||
|
Domain Architecture |

|
|||||
| Description | SPARC-related modular calcium-binding protein 2 precursor (Secreted modular calcium-binding protein 2) (SMOC-2) (Smooth muscle-associated protein 2) (SMAP-2). | |||||
|
SMOC2_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.996339 (rank : 2) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q8CD91, Q8VCM1, Q9D9K2, Q9ER95 | Gene names | Smoc2 | |||
|
Domain Architecture |
|
|||||
| Description | SPARC-related modular calcium-binding protein 2 precursor (Secreted modular calcium-binding protein 2) (SMOC-2). | |||||
|
SMOC1_HUMAN
|
||||||
| θ value | 2.28167e-128 (rank : 3) | NC score | 0.971624 (rank : 3) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9H4F8, Q96F78 | Gene names | SMOC1 | |||
|
Domain Architecture |
|
|||||
| Description | SPARC-related modular calcium-binding protein 1 precursor (Secreted modular calcium-binding protein 1) (SMOC-1). | |||||
|
SMOC1_MOUSE
|
||||||
| θ value | 1.17183e-124 (rank : 4) | NC score | 0.970263 (rank : 4) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q8BLY1, Q9WVN9 | Gene names | Smoc1, Srg | |||
|
Domain Architecture |
|
|||||
| Description | SPARC-related modular calcium-binding protein 1 precursor (Secreted modular calcium-binding protein 1) (SMOC-1) (SPARC-related gene protein). | |||||
|
NID2_MOUSE
|
||||||
| θ value | 1.52774e-15 (rank : 5) | NC score | 0.277989 (rank : 38) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 338 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O88322 | Gene names | Nid2 | |||
|
Domain Architecture |

|
|||||
| Description | Nidogen-2 precursor (NID-2) (Entactin-2). | |||||
|
NID2_HUMAN
|
||||||
| θ value | 1.99529e-15 (rank : 6) | NC score | 0.277443 (rank : 39) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 456 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q14112, O43710 | Gene names | NID2 | |||
|
Domain Architecture |

|
|||||
| Description | Nidogen-2 precursor (NID-2) (Osteonidogen). | |||||
|
THYG_HUMAN
|
||||||
| θ value | 1.99529e-15 (rank : 7) | NC score | 0.368028 (rank : 22) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P01266, O15274, O43899, Q15593, Q15948, Q9NYR1, Q9NYR2, Q9UMZ0, Q9UNY3 | Gene names | TG | |||
|
Domain Architecture |

|
|||||
| Description | Thyroglobulin precursor. | |||||
|
THYG_MOUSE
|
||||||
| θ value | 1.09485e-13 (rank : 8) | NC score | 0.351411 (rank : 28) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O08710, O88590, Q9QWY7 | Gene names | Tg, Tgn | |||
|
Domain Architecture |

|
|||||
| Description | Thyroglobulin precursor. | |||||
|
TICN2_MOUSE
|
||||||
| θ value | 1.47974e-10 (rank : 9) | NC score | 0.533786 (rank : 6) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9ER58 | Gene names | Spock2, Ticn2 | |||
|
Domain Architecture |
|
|||||
| Description | Testican-2 precursor (SPARC/osteonectin, CWCV, and Kazal-like domains proteoglycan 2). | |||||
|
TICN2_HUMAN
|
||||||
| θ value | 2.13673e-09 (rank : 10) | NC score | 0.531194 (rank : 7) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q92563, Q6UW87 | Gene names | SPOCK2, KIAA0275, TICN2 | |||
|
Domain Architecture |
|
|||||
| Description | Testican-2 precursor (SPARC/osteonectin, CWCV, and Kazal-like domains proteoglycan 2). | |||||
|
NID1_HUMAN
|
||||||
| θ value | 1.80886e-08 (rank : 11) | NC score | 0.247221 (rank : 42) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P14543, Q14942, Q59FL2, Q5TAF2, Q5TAF3, Q86XD7 | Gene names | NID1, NID | |||
|
Domain Architecture |

|
|||||
| Description | Nidogen-1 precursor (Entactin). | |||||
|
NID1_MOUSE
|
||||||
| θ value | 6.87365e-08 (rank : 12) | NC score | 0.243140 (rank : 43) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P10493, Q8BQI3, Q8C3U8, Q8C9P6 | Gene names | Nid1, Ent | |||
|
Domain Architecture |

|
|||||
| Description | Nidogen-1 precursor (Entactin). | |||||
|
HG2A_MOUSE
|
||||||
| θ value | 5.81887e-07 (rank : 13) | NC score | 0.537802 (rank : 5) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P04441, O19452 | Gene names | Cd74, Ii | |||
|
Domain Architecture |
|
|||||
| Description | H-2 class II histocompatibility antigen gamma chain (MHC class II- associated invariant chain) (Ia antigen-associated invariant chain) (Ii) (CD74 antigen). | |||||
|
TICN1_HUMAN
|
||||||
| θ value | 2.21117e-06 (rank : 14) | NC score | 0.496165 (rank : 8) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q08629 | Gene names | SPOCK1, SPOCK, TIC1, TICN1 | |||
|
Domain Architecture |
|
|||||
| Description | Testican-1 precursor (Protein SPOCK). | |||||
|
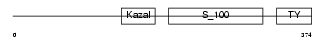
TICN1_MOUSE
|
||||||
| θ value | 2.21117e-06 (rank : 15) | NC score | 0.494463 (rank : 9) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q62288 | Gene names | Spock1, Spock, Ticn1 | |||
|
Domain Architecture |
|
|||||
| Description | Testican-1 precursor (Protein SPOCK). | |||||
|
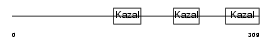
FST_HUMAN
|
||||||
| θ value | 2.88788e-06 (rank : 16) | NC score | 0.366060 (rank : 24) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P19883, Q9BTH0 | Gene names | FST | |||
|
Domain Architecture |
|
|||||
| Description | Follistatin precursor (FS) (Activin-binding protein). | |||||
|
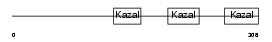
FST_MOUSE
|
||||||
| θ value | 2.88788e-06 (rank : 17) | NC score | 0.357866 (rank : 26) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P47931 | Gene names | Fst | |||
|
Domain Architecture |
|
|||||
| Description | Follistatin precursor (FS) (Activin-binding protein). | |||||
|
IBP3_HUMAN
|
||||||
| θ value | 8.40245e-06 (rank : 18) | NC score | 0.397432 (rank : 14) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P17936, Q2V509, Q6P1M6 | Gene names | IGFBP3, IBP3 | |||
|
Domain Architecture |

|
|||||
| Description | Insulin-like growth factor-binding protein 3 precursor (IGFBP-3) (IBP- 3) (IGF-binding protein 3). | |||||
|
HG2A_HUMAN
|
||||||
| θ value | 1.09739e-05 (rank : 19) | NC score | 0.492464 (rank : 10) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P04233, Q14597, Q29832, Q5U0J8, Q8WLP6 | Gene names | CD74, DHLAG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HLA class II histocompatibility antigen gamma chain (HLA-DR antigens- associated invariant chain) (Ia antigen-associated invariant chain) (Ii) (p33) (CD74 antigen). | |||||
|
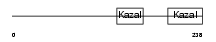
FSTL3_HUMAN
|
||||||
| θ value | 1.43324e-05 (rank : 20) | NC score | 0.352065 (rank : 27) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O95633 | Gene names | FSTL3, FLRG | |||
|
Domain Architecture |
|
|||||
| Description | Follistatin-related protein 3 precursor (Follistatin-like 3) (Follistatin-related gene protein). | |||||
|
SPRC_MOUSE
|
||||||
| θ value | 1.87187e-05 (rank : 21) | NC score | 0.370004 (rank : 20) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P07214 | Gene names | Sparc | |||
|
Domain Architecture |
|
|||||
| Description | SPARC precursor (Secreted protein acidic and rich in cysteine) (Osteonectin) (ON) (Basement-membrane protein 40) (BM-40). | |||||
|
IBP1_HUMAN
|
||||||
| θ value | 4.1701e-05 (rank : 22) | NC score | 0.399103 (rank : 13) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P08833, Q8IYP5 | Gene names | IGFBP1, IBP1 | |||
|
Domain Architecture |
|
|||||
| Description | Insulin-like growth factor-binding protein 1 precursor (IGFBP-1) (IBP- 1) (IGF-binding protein 1) (Placental protein 12) (PP12). | |||||
|
TICN3_HUMAN
|
||||||
| θ value | 4.1701e-05 (rank : 23) | NC score | 0.483504 (rank : 11) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9BQ16, O75705, Q6UW53, Q96Q26 | Gene names | SPOCK3, TICN3 | |||
|
Domain Architecture |
|
|||||
| Description | Testican-3 precursor (SPARC/osteonectin, CWCV, and Kazal-like domains proteoglycan 3). | |||||
|
SPRC_HUMAN
|
||||||
| θ value | 5.44631e-05 (rank : 24) | NC score | 0.368289 (rank : 21) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P09486 | Gene names | SPARC, ON | |||
|
Domain Architecture |
|
|||||
| Description | SPARC precursor (Secreted protein acidic and rich in cysteine) (Osteonectin) (ON) (Basement-membrane protein 40) (BM-40). | |||||
|
TICN3_MOUSE
|
||||||
| θ value | 5.44631e-05 (rank : 25) | NC score | 0.481074 (rank : 12) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8BKV0, Q9ER59 | Gene names | Spock3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testican-3 precursor (SPARC/osteonectin, CWCV, and Kazal-like domains proteoglycan 3). | |||||
|
IBP1_MOUSE
|
||||||
| θ value | 7.1131e-05 (rank : 26) | NC score | 0.374843 (rank : 18) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P47876, Q61732 | Gene names | Igfbp1, Igfbp-1 | |||
|
Domain Architecture |

|
|||||
| Description | Insulin-like growth factor-binding protein 1 precursor (IGFBP-1) (IBP- 1) (IGF-binding protein 1). | |||||
|
TEFF2_HUMAN
|
||||||
| θ value | 7.1131e-05 (rank : 27) | NC score | 0.295073 (rank : 34) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9UIK5, Q4ZFW4, Q53H90, Q53RE1, Q8N2R5, Q9NR15, Q9NSS5, Q9P2Y9, Q9UK65 | Gene names | TMEFF2, HPP1, TENB2, TPEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein with EGF-like and two follistatin-like domains 2 precursor (Transmembrane protein containing epidermal growth factor and follistatin domains) (Tomoregulin) (TR) (Hyperplastic polyposis protein 1). | |||||
|
TEFF2_MOUSE
|
||||||
| θ value | 7.1131e-05 (rank : 28) | NC score | 0.294968 (rank : 35) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9QYM9, Q3UY20, Q8CDH1, Q9JJE3 | Gene names | Tmeff2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein with EGF-like and two follistatin-like domains 2 precursor (Tomoregulin) (TR). | |||||
|
SPRL1_MOUSE
|
||||||
| θ value | 0.000270298 (rank : 29) | NC score | 0.290212 (rank : 36) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P70663, P97810, Q99L82 | Gene names | Sparcl1, Ecm2, Sc1 | |||
|
Domain Architecture |

|
|||||
| Description | SPARC-like protein 1 precursor (Matrix glycoprotein Sc1) (Extracellular matrix protein 2). | |||||
|
IBP2_MOUSE
|
||||||
| θ value | 0.00035302 (rank : 30) | NC score | 0.373028 (rank : 19) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P47877, Q61722 | Gene names | Igfbp2, Igfbp-2 | |||
|
Domain Architecture |

|
|||||
| Description | Insulin-like growth factor-binding protein 2 precursor (IGFBP-2) (IBP- 2) (IGF-binding protein 2). | |||||
|
IBP4_HUMAN
|
||||||
| θ value | 0.000461057 (rank : 31) | NC score | 0.380905 (rank : 17) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P22692 | Gene names | IGFBP4, IBP4 | |||
|
Domain Architecture |
|
|||||
| Description | Insulin-like growth factor-binding protein 4 precursor (IGFBP-4) (IBP- 4) (IGF-binding protein 4). | |||||
|
IBP4_MOUSE
|
||||||
| θ value | 0.000461057 (rank : 32) | NC score | 0.383085 (rank : 16) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P47879, O35666 | Gene names | Igfbp4, Igfbp-4 | |||
|
Domain Architecture |
|
|||||
| Description | Insulin-like growth factor-binding protein 4 precursor (IGFBP-4) (IBP- 4) (IGF-binding protein 4). | |||||
|
AGRIN_HUMAN
|
||||||
| θ value | 0.000602161 (rank : 33) | NC score | 0.163549 (rank : 46) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 582 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O00468, Q5SVA2, Q60FE1, Q7KYS8, Q8N4J5, Q96IC1, Q9BTD4 | Gene names | AGRIN, AGRN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Agrin precursor. | |||||
|
SPRL1_HUMAN
|
||||||
| θ value | 0.000602161 (rank : 34) | NC score | 0.314258 (rank : 33) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q14515, Q14800 | Gene names | SPARCL1 | |||
|
Domain Architecture |

|
|||||
| Description | SPARC-like protein 1 precursor (High endothelial venule protein) (Hevin) (MAST 9). | |||||
|
FSTL3_MOUSE
|
||||||
| θ value | 0.000786445 (rank : 35) | NC score | 0.326120 (rank : 32) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9EQC7 | Gene names | Fstl3, Flrg | |||
|
Domain Architecture |
|
|||||
| Description | Follistatin-related protein 3 precursor (Follistatin-like 3) (Follistatin-related gene protein). | |||||
|
IBP2_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 36) | NC score | 0.347509 (rank : 29) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P18065, Q14619, Q9UCL3 | Gene names | IGFBP2, BP2 | |||
|
Domain Architecture |
|
|||||
| Description | Insulin-like growth factor-binding protein 2 precursor (IGFBP-2) (IBP- 2) (IGF-binding protein 2). | |||||
|
IBP6_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 37) | NC score | 0.383824 (rank : 15) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P24592, Q14492 | Gene names | IGFBP6, IBP6 | |||
|
Domain Architecture |

|
|||||
| Description | Insulin-like growth factor-binding protein 6 precursor (IGFBP-6) (IBP- 6) (IGF-binding protein 6). | |||||
|
IBP6_MOUSE
|
||||||
| θ value | 0.00509761 (rank : 38) | NC score | 0.367457 (rank : 23) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P47880 | Gene names | Igfbp6, Igfbp-6 | |||
|
Domain Architecture |
|
|||||
| Description | Insulin-like growth factor-binding protein 6 precursor (IGFBP-6) (IBP- 6) (IGF-binding protein 6). | |||||
|
FSTL1_MOUSE
|
||||||
| θ value | 0.00869519 (rank : 39) | NC score | 0.274189 (rank : 40) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q62356, Q99JI9 | Gene names | Fstl1, Frp, Fstl, Tsc36 | |||
|
Domain Architecture |
|
|||||
| Description | Follistatin-related protein 1 precursor (Follistatin-like 1) (TGF- beta-inducible protein TSC-36). | |||||
|
IBP3_MOUSE
|
||||||
| θ value | 0.00869519 (rank : 40) | NC score | 0.364015 (rank : 25) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P47878 | Gene names | Igfbp3, Igfbp-3 | |||
|
Domain Architecture |
|
|||||
| Description | Insulin-like growth factor-binding protein 3 precursor (IGFBP-3) (IBP- 3) (IGF-binding protein 3). | |||||
|
FSTL1_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 41) | NC score | 0.274018 (rank : 41) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q12841 | Gene names | FSTL1, FRP | |||
|
Domain Architecture |
|
|||||
| Description | Follistatin-related protein 1 precursor (Follistatin-like 1). | |||||
|
GUC1C_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 42) | NC score | 0.074216 (rank : 61) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O95843, O95844, Q9UNM0 | Gene names | GUCA1C, GCAP3 | |||
|
Domain Architecture |
|
|||||
| Description | Guanylyl cyclase-activating protein 3 (GCAP 3) (Guanylate cyclase activator 1C). | |||||
|
IBP5_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 43) | NC score | 0.334640 (rank : 30) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P24593 | Gene names | IGFBP5, IBP5 | |||
|
Domain Architecture |

|
|||||
| Description | Insulin-like growth factor-binding protein 5 precursor (IGFBP-5) (IBP- 5) (IGF-binding protein 5). | |||||
|
KAZD1_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 44) | NC score | 0.159649 (rank : 47) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q96I82, Q6ZMB1, Q9BQ73 | Gene names | KAZALD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kazal-type serine protease inhibitor domain-containing protein 1 precursor. | |||||
|
KAZD1_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 45) | NC score | 0.156176 (rank : 48) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8BJ66 | Gene names | Kazald1, Bono1, Igfbprp10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kazal-type serine protease inhibitor domain-containing protein 1 precursor (IGFBP-related protein 10) (IGFBP-rP10) (Bone and odontoblast expressed protein 1). | |||||
|
IBP5_MOUSE
|
||||||
| θ value | 0.125558 (rank : 46) | NC score | 0.331498 (rank : 31) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q07079 | Gene names | Igfbp5, Igfbp-5 | |||
|
Domain Architecture |

|
|||||
| Description | Insulin-like growth factor-binding protein 5 precursor (IGFBP-5) (IBP- 5) (IGF-binding protein 5). | |||||
|
TACD1_HUMAN
|
||||||
| θ value | 0.125558 (rank : 47) | NC score | 0.289628 (rank : 37) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P16422, P18180, Q96C47 | Gene names | TACSTD1, GA733-2, M1S2, TROP1 | |||
|
Domain Architecture |
|
|||||
| Description | Tumor-associated calcium signal transducer 1 precursor (Major gastrointestinal tumor-associated protein GA733-2) (Epithelial cell surface antigen) (Epithelial glycoprotein) (EGP) (Adenocarcinoma- associated antigen) (KSA) (KS 1/4 antigen) (Cell surface glycoprotein Trop-1) (CD326 antigen). | |||||
|
PCY1B_MOUSE
|
||||||
| θ value | 0.279714 (rank : 48) | NC score | 0.042873 (rank : 87) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q811Q9, Q3UEW0, Q80Y63, Q811Q8, Q8BKD2, Q8C085 | Gene names | Pcyt1b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Choline-phosphate cytidylyltransferase B (EC 2.7.7.15) (Phosphorylcholine transferase B) (CTP:phosphocholine cytidylyltransferase B) (CT B) (CCT B) (CCT-beta). | |||||
|
FSTL5_MOUSE
|
||||||
| θ value | 0.365318 (rank : 49) | NC score | 0.101225 (rank : 52) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 355 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8BFR2, Q80TG3, Q8C4T3 | Gene names | Fstl5, Kiaa1263 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Follistatin-related protein 5 precursor (Follistatin-like 5) (m- D/Bsp120I 1-2). | |||||
|
PCY1B_HUMAN
|
||||||
| θ value | 0.47712 (rank : 50) | NC score | 0.041257 (rank : 90) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y5K3, O60621, Q86XC9 | Gene names | PCYT1B, CCTB | |||
|
Domain Architecture |
|
|||||
| Description | Choline-phosphate cytidylyltransferase B (EC 2.7.7.15) (Phosphorylcholine transferase B) (CTP:phosphocholine cytidylyltransferase B) (CT B) (CCT B) (CCT-beta). | |||||
|
FSTL5_HUMAN
|
||||||
| θ value | 0.62314 (rank : 51) | NC score | 0.101607 (rank : 51) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8N475, Q9NSW7, Q9ULF7 | Gene names | FSTL5, KIAA1263 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Follistatin-related protein 5 precursor (Follistatin-like 5). | |||||
|
GUC1A_MOUSE
|
||||||
| θ value | 0.813845 (rank : 52) | NC score | 0.055220 (rank : 69) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P43081 | Gene names | Guca1a, Gcap, Gcap1, Guca1 | |||
|
Domain Architecture |
|
|||||
| Description | Guanylyl cyclase-activating protein 1 (GCAP 1) (Guanylate cyclase activator 1A). | |||||
|
S10A9_HUMAN
|
||||||
| θ value | 0.813845 (rank : 53) | NC score | 0.036475 (rank : 91) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P06702, Q9NYM0, Q9UCJ1 | Gene names | S100A9, CAGB, MRP14 | |||
|
Domain Architecture |

|
|||||
| Description | Protein S100-A9 (S100 calcium-binding protein A9) (Calgranulin-B) (Migration inhibitory factor-related protein 14) (MRP-14) (P14) (Leukocyte L1 complex heavy chain) (Calprotectin L1H subunit). | |||||
|
TACD2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 54) | NC score | 0.240966 (rank : 44) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P09758, Q15658, Q96QD2 | Gene names | TACSTD2, GA733-1, M1S1, TROP2 | |||
|
Domain Architecture |
|
|||||
| Description | Tumor-associated calcium signal transducer 2 precursor (Pancreatic carcinoma marker protein GA733-1) (Cell surface glycoprotein Trop-2). | |||||
|
CFAI_MOUSE
|
||||||
| θ value | 1.06291 (rank : 55) | NC score | 0.028016 (rank : 93) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q61129, Q9WU07 | Gene names | Cfi, If | |||
|
Domain Architecture |

|
|||||
| Description | Complement factor I precursor (EC 3.4.21.45) (C3B/C4B inactivator) [Contains: Complement factor I heavy chain; Complement factor I light chain]. | |||||
|
ESCO1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 56) | NC score | 0.027632 (rank : 94) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5FWF5, Q69YG4, Q69YS3, Q6IMD7, Q8N3Z5, Q8NBG2, Q96PX7 | Gene names | ESCO1, EFO1, KIAA1911 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyltransferase ESCO1 (EC 2.3.1.-) (Establishment of cohesion 1 homolog 1) (ECO1 homolog 1) (ESO1 homolog 1) (Establishment factor- like protein 1) (EFO1p) (hEFO1) (CTF7 homolog 1). | |||||
|
ISK6_MOUSE
|
||||||
| θ value | 1.06291 (rank : 57) | NC score | 0.183157 (rank : 45) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8BT20 | Gene names | Spink6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine protease inhibitor Kazal-type 6 precursor. | |||||
|
HPCA_HUMAN
|
||||||
| θ value | 1.38821 (rank : 58) | NC score | 0.048466 (rank : 79) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P84074, P32076, P41211, P70510 | Gene names | HPCA, BDR2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuron-specific calcium-binding protein hippocalcin (Calcium-binding protein BDR-2). | |||||
|
HPCA_MOUSE
|
||||||
| θ value | 1.38821 (rank : 59) | NC score | 0.048466 (rank : 80) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P84075, P32076, P41211, P70510 | Gene names | Hpca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuron-specific calcium-binding protein hippocalcin. | |||||
|
MADCA_HUMAN
|
||||||
| θ value | 1.38821 (rank : 60) | NC score | 0.025696 (rank : 95) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q13477, O60222, O75867 | Gene names | MADCAM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucosal addressin cell adhesion molecule 1 precursor (MAdCAM-1) (hMAdCAM-1). | |||||
|
ANKY1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 61) | NC score | 0.028968 (rank : 92) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9P2S6, Q8IYX5, Q8NDK5, Q9NX10 | Gene names | ANKMY1, TSAL1, ZMYND13 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin repeat and MYND domain-containing protein 1 (Testis-specific ankyrin-like protein 1) (Zinc-finger MYND domain-containing protein 13). | |||||
|
GP152_HUMAN
|
||||||
| θ value | 2.36792 (rank : 62) | NC score | 0.012754 (rank : 103) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 636 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8TDT2, Q86SM0 | Gene names | GPR152, PGR5 | |||
|
Domain Architecture |

|
|||||
| Description | Probable G-protein coupled receptor 152 (G-protein coupled receptor PGR5). | |||||
|
HPCL1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 63) | NC score | 0.045814 (rank : 85) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P37235, Q969S5 | Gene names | HPCAL1, BDR1 | |||
|
Domain Architecture |
|
|||||
| Description | Hippocalcin-like protein 1 (Visinin-like protein 3) (VILIP-3) (Calcium-binding protein BDR-1) (HLP2). | |||||
|
HPCL1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 64) | NC score | 0.046180 (rank : 84) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P62748, P35333 | Gene names | Hpcal1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hippocalcin-like protein 1 (Visinin-like protein 3) (VILIP-3) (Neural visinin-like protein 3) (NVL-3) (NVP-3). | |||||
|
TULP1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 65) | NC score | 0.022318 (rank : 97) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Z273 | Gene names | Tulp1 | |||
|
Domain Architecture |

|
|||||
| Description | Tubby-related protein 1 (Tubby-like protein 1). | |||||
|
FSTL4_MOUSE
|
||||||
| θ value | 3.0926 (rank : 66) | NC score | 0.094566 (rank : 55) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q5STE3, Q5DU03 | Gene names | Fstl4, Kiaa1061 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Follistatin-related protein 4 precursor (Follistatin-like 4) (m- D/Bsp120I 1-1). | |||||
|
KCIP2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 67) | NC score | 0.050701 (rank : 76) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9JJ69, Q6DTJ2, Q8K1T8, Q8K1T9, Q8K1U0, Q8VHN4, Q8VHN5, Q8VHN6, Q9JJ68 | Gene names | Kcnip2, Kchip2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kv channel-interacting protein 2 (KChIP2) (A-type potassium channel modulatory protein 2) (Potassium channel-interacting protein 2). | |||||
|
LAMA3_MOUSE
|
||||||
| θ value | 3.0926 (rank : 68) | NC score | 0.023004 (rank : 96) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 859 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q61789, O08751, Q61788, Q61966, Q9JHQ7 | Gene names | Lama3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-3 chain precursor (Nicein subunit alpha). | |||||
|
PKP3_MOUSE
|
||||||
| θ value | 3.0926 (rank : 69) | NC score | 0.019409 (rank : 98) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9QY23 | Gene names | Pkp3 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-3. | |||||
|
KCIP2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 70) | NC score | 0.049557 (rank : 78) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9NS61, Q7Z6F1, Q96K86, Q96T41, Q96T42, Q96T43, Q96T44, Q9H0N4, Q9HD10, Q9HD11, Q9NS60, Q9NY10, Q9NZI1 | Gene names | KCNIP2, KCHIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kv channel-interacting protein 2 (KChIP2) (A-type potassium channel modulatory protein 2) (Potassium channel-interacting protein 2) (Cardiac voltage-gated potassium channel modulatory subunit). | |||||
|
KCIP4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 71) | NC score | 0.048041 (rank : 81) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6PIL6, Q4W5G8, Q8NEU0, Q9BWT2, Q9H294, Q9H2A4 | Gene names | KCNIP4, CALP, KCHIP4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kv channel-interacting protein 4 (KChIP4) (A-type potassium channel modulatory protein 4) (Potassium channel-interacting protein 4) (Calsenilin-like protein). | |||||
|
KCIP4_MOUSE
|
||||||
| θ value | 4.03905 (rank : 72) | NC score | 0.048030 (rank : 82) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6PHZ8, Q6DTJ3, Q8CAD0, Q8R4I2, Q9EQ01 | Gene names | Kcnip4, Calp, Kchip4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kv channel-interacting protein 4 (KChIP4) (A-type potassium channel modulatory protein 4) (Potassium channel-interacting protein 4) (Calsenilin-like protein). | |||||
|
NCALD_HUMAN
|
||||||
| θ value | 4.03905 (rank : 73) | NC score | 0.042298 (rank : 88) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P61601, P29554, Q8IYC3, Q9H0W2 | Gene names | NCALD | |||
|
Domain Architecture |
|
|||||
| Description | Neurocalcin delta. | |||||
|
NCALD_MOUSE
|
||||||
| θ value | 4.03905 (rank : 74) | NC score | 0.042298 (rank : 89) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q91X97, Q3TJS9, Q8BZN9 | Gene names | Ncald, D15Ertd412e | |||
|
Domain Architecture |
|
|||||
| Description | Neurocalcin delta. | |||||
|
PKP3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 75) | NC score | 0.015807 (rank : 101) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y446 | Gene names | PKP3 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-3. | |||||
|
SMCA2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 76) | NC score | 0.004409 (rank : 106) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 503 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P51531 | Gene names | SMARCA2, BRM, SNF2A, SNF2L2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L2 (EC 3.6.1.-) (ATP- dependent helicase SMARCA2) (SNF2-alpha) (SWI/SNF-related matrix- associated actin-dependent regulator of chromatin subfamily A member 2) (hBRM). | |||||
|
SUHW4_MOUSE
|
||||||
| θ value | 4.03905 (rank : 77) | NC score | 0.000900 (rank : 108) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 737 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q68FE8, Q3T9B1, Q8BI82 | Gene names | Suhw4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppressor of hairy wing homolog 4. | |||||
|
ACM3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 78) | NC score | -0.001803 (rank : 110) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P20309, Q4QRI3, Q5VXY2, Q9HB60 | Gene names | CHRM3 | |||
|
Domain Architecture |

|
|||||
| Description | Muscarinic acetylcholine receptor M3. | |||||
|
GUC1A_HUMAN
|
||||||
| θ value | 5.27518 (rank : 79) | NC score | 0.046336 (rank : 83) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P43080, Q9NU14 | Gene names | GUCA1A, GCAP, GCAP1, GUCA1 | |||
|
Domain Architecture |
|
|||||
| Description | Guanylyl cyclase-activating protein 1 (GCAP 1) (Guanylate cyclase activator 1A). | |||||
|
HTRA1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 80) | NC score | 0.097749 (rank : 54) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9R118 | Gene names | Htra1, Htra, Prss11 | |||
|
Domain Architecture |

|
|||||
| Description | Serine protease HTRA1 precursor (EC 3.4.21.-). | |||||
|
SO1A2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 81) | NC score | 0.017096 (rank : 99) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P46721, Q9UGP7, Q9UL38 | Gene names | SLCO1A2, OATP, SLC21A3 | |||
|
Domain Architecture |

|
|||||
| Description | Solute carrier organic anion transporter family member 1A2 (Solute carrier family 21 member 3) (Sodium-independent organic anion transporter) (Organic anion-transporting polypeptide 1) (OATP1) (OATP- A). | |||||
|
CABP5_MOUSE
|
||||||
| θ value | 6.88961 (rank : 82) | NC score | 0.011526 (rank : 104) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9JLK3 | Gene names | Cabp5 | |||
|
Domain Architecture |
|
|||||
| Description | Calcium-binding protein 5 (CaBP5). | |||||
|
CSEN_HUMAN
|
||||||
| θ value | 6.88961 (rank : 83) | NC score | 0.045712 (rank : 86) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Y2W7, Q53TJ5, Q96T40, Q9UJ84, Q9UJ85 | Gene names | KCNIP3, CSEN, DREAM, KCHIP3 | |||
|
Domain Architecture |

|
|||||
| Description | Calsenilin (DRE-antagonist modulator) (DREAM) (Kv channel-interacting protein 3) (KChIP3) (A-type potassium channel modulatory protein 3). | |||||
|
CYGB_MOUSE
|
||||||
| θ value | 6.88961 (rank : 84) | NC score | 0.015369 (rank : 102) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9CX80 | Gene names | Cygb | |||
|
Domain Architecture |
|
|||||
| Description | Cytoglobin (Histoglobin) (HGb). | |||||
|
LAMA2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 85) | NC score | 0.016784 (rank : 100) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 1092 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P24043, Q14736, Q93022 | Gene names | LAMA2, LAMM | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-2 chain precursor (Laminin M chain) (Merosin heavy chain). | |||||
|
PRDM2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 86) | NC score | -0.000934 (rank : 109) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q13029, Q13149, Q14550, Q5VUL9 | Gene names | PRDM2, RIZ | |||
|
Domain Architecture |

|
|||||
| Description | PR domain zinc finger protein 2 (PR domain-containing protein 2) (Retinoblastoma protein-interacting zinc-finger protein) (Zinc finger protein RIZ) (MTE-binding protein) (MTB-ZF) (GATA-3-binding protein G3B). | |||||
|
MINT_MOUSE
|
||||||
| θ value | 8.99809 (rank : 87) | NC score | 0.001649 (rank : 107) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
RNF31_HUMAN
|
||||||
| θ value | 8.99809 (rank : 88) | NC score | 0.008025 (rank : 105) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96EP0, Q86VI2, Q8TEI0, Q96GB4, Q96NF1, Q9H5F1, Q9NWD2 | Gene names | RNF31, ZIBRA | |||
|
Domain Architecture |
|
|||||
| Description | RING finger protein 31 (Zinc in-between-RING-finger ubiquitin- associated domain protein). | |||||
|
EGF_HUMAN
|
||||||
| θ value | θ > 10 (rank : 89) | NC score | 0.054640 (rank : 70) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P01133 | Gene names | EGF | |||
|
Domain Architecture |

|
|||||
| Description | Pro-epidermal growth factor precursor (EGF) [Contains: Epidermal growth factor (Urogastrone)]. | |||||
|
EGF_MOUSE
|
||||||
| θ value | θ > 10 (rank : 90) | NC score | 0.054491 (rank : 71) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 358 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P01132, Q6P9J2 | Gene names | Egf | |||
|
Domain Architecture |

|
|||||
| Description | Pro-epidermal growth factor precursor (EGF) [Contains: Epidermal growth factor]. | |||||
|
FSTL4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 91) | NC score | 0.094541 (rank : 56) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q6MZW2, Q8TBU0, Q9UPU1 | Gene names | FSTL4, KIAA1061 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Follistatin-related protein 4 precursor (Follistatin-like 4). | |||||
|
HTRA1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 92) | NC score | 0.082267 (rank : 58) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q92743, Q9UNS5 | Gene names | HTRA1, HTRA, PRSS11 | |||
|
Domain Architecture |

|
|||||
| Description | Serine protease HTRA1 precursor (EC 3.4.21.-) (L56). | |||||
|
IBP7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 93) | NC score | 0.081820 (rank : 59) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q16270, Q07822 | Gene names | IGFBP7, MAC25, PSF | |||
|
Domain Architecture |
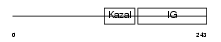
|
|||||
| Description | Insulin-like growth factor-binding protein 7 precursor (IGFBP-7) (IBP- 7) (IGF-binding protein 7) (MAC25 protein) (Prostacyclin-stimulating factor) (PGI2-stimulating factor) (IGFBP-rP1). | |||||
|
IBP7_MOUSE
|
||||||
| θ value | θ > 10 (rank : 94) | NC score | 0.076390 (rank : 60) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q61581, O88812, Q9EQW0 | Gene names | Igfbp7, Mac25 | |||
|
Domain Architecture |
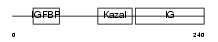
|
|||||
| Description | Insulin-like growth factor-binding protein 7 precursor (IGFBP-7) (IBP- 7) (IGF-binding protein 7) (MAC25 protein). | |||||
|
IPK1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 95) | NC score | 0.072427 (rank : 62) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P00995 | Gene names | SPINK1, PSTI | |||
|
Domain Architecture |
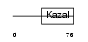
|
|||||
| Description | Pancreatic secretory trypsin inhibitor precursor (Tumor-associated trypsin inhibitor) (TATI) (Serine protease inhibitor Kazal-type 1). | |||||
|
IPK2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 96) | NC score | 0.053452 (rank : 72) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P20155 | Gene names | SPINK2 | |||
|
Domain Architecture |
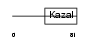
|
|||||
| Description | Serine protease inhibitor Kazal-type 2 precursor (Acrosin-trypsin inhibitor) (HUSI-II). | |||||
|
ISK3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 97) | NC score | 0.050012 (rank : 77) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P09036, Q5M9M3 | Gene names | Spink3 | |||
|
Domain Architecture |
|
|||||
| Description | Serine protease inhibitor Kazal-type 3 precursor (Prostatic secretory glycoprotein) (P12). | |||||
|
ISK4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 98) | NC score | 0.094088 (rank : 57) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O60575 | Gene names | SPINK4 | |||
|
Domain Architecture |
|
|||||
| Description | Serine protease inhibitor Kazal-type 4 precursor (Peptide PEC-60 homolog). | |||||
|
ISK4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 99) | NC score | 0.100197 (rank : 53) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O35679 | Gene names | Spink4, Mpgc60 | |||
|
Domain Architecture |
|
|||||
| Description | Serine protease inhibitor Kazal-type 4 precursor (Peptide PEC-60 homolog) (MPGC60 protein). | |||||
|
ISK5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.104052 (rank : 50) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9NQ38, O75770, Q96PP2, Q96PP3 | Gene names | SPINK5 | |||
|
Domain Architecture |

|
|||||
| Description | Serine protease inhibitor Kazal-type 5 precursor (Lympho-epithelial Kazal-type-related inhibitor) (LEKTI) [Contains: Hemofiltrate peptide HF6478; Hemofiltrate peptide HF7665]. | |||||
|
ISK6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.110871 (rank : 49) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q6UWN8, Q8N5P0 | Gene names | SPINK6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine protease inhibitor Kazal-type 6 precursor. | |||||
|
ISK7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.065718 (rank : 64) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P58062 | Gene names | SPINK7, ECG2 | |||
|
Domain Architecture |
|
|||||
| Description | Serine protease inhibitor Kazal-type 7 precursor (Esophagus cancer- related gene 2 protein) (ECRG-2). | |||||
|
ISK7_MOUSE
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.068464 (rank : 63) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6IE32 | Gene names | Spink7, Ecg2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine protease inhibitor Kazal-type 7 precursor (Esophagus cancer- related gene 2 protein) (ECRG-2). | |||||
|
LRP5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.058310 (rank : 65) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 224 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O75197, Q96TD6, Q9UP66 | Gene names | LRP5, LRP7 | |||
|
Domain Architecture |

|
|||||
| Description | Low-density lipoprotein receptor-related protein 5 precursor. | |||||
|
LRP5_MOUSE
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.058061 (rank : 66) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q91VN0, O88883, Q9R208 | Gene names | Lrp5, Lrp7 | |||
|
Domain Architecture |

|
|||||
| Description | Low-density lipoprotein receptor-related protein 5 precursor (LRP7) (Lr3). | |||||
|
LRP6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 106) | NC score | 0.056441 (rank : 67) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O75581 | Gene names | LRP6 | |||
|
Domain Architecture |

|
|||||
| Description | Low-density lipoprotein receptor-related protein 6 precursor. | |||||
|
LRP6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 107) | NC score | 0.056389 (rank : 68) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O88572 | Gene names | Lrp6 | |||
|
Domain Architecture |

|
|||||
| Description | Low-density lipoprotein receptor-related protein 6 precursor. | |||||
|
RECK_HUMAN
|
||||||
| θ value | θ > 10 (rank : 108) | NC score | 0.051166 (rank : 74) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O95980, Q8WX37 | Gene names | RECK | |||
|
Domain Architecture |

|
|||||
| Description | Reversion-inducing cysteine-rich protein with Kazal motifs precursor (hRECK) (Suppressor of tumorigenicity 15) (ST15). | |||||
|
TECTA_HUMAN
|
||||||
| θ value | θ > 10 (rank : 109) | NC score | 0.053281 (rank : 73) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O75443 | Gene names | TECTA | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-tectorin precursor. | |||||
|
TECTA_MOUSE
|
||||||
| θ value | θ > 10 (rank : 110) | NC score | 0.051103 (rank : 75) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 244 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O08523 | Gene names | Tecta | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-tectorin precursor. | |||||
|
SMOC2_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 88 | |
| SwissProt Accessions | Q9H3U7, Q5TAT7, Q5TAT8, Q86VV9, Q96SF3, Q9H1L3, Q9H1L4, Q9H3U0, Q9H4F7, Q9HCV2 | Gene names | SMOC2, SMAP2 | |||
|
Domain Architecture |

|
|||||
| Description | SPARC-related modular calcium-binding protein 2 precursor (Secreted modular calcium-binding protein 2) (SMOC-2) (Smooth muscle-associated protein 2) (SMAP-2). | |||||
|
SMOC2_MOUSE
|
||||||
| NC score | 0.996339 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q8CD91, Q8VCM1, Q9D9K2, Q9ER95 | Gene names | Smoc2 | |||
|
Domain Architecture |

|
|||||
| Description | SPARC-related modular calcium-binding protein 2 precursor (Secreted modular calcium-binding protein 2) (SMOC-2). | |||||
|
SMOC1_HUMAN
|
||||||
| NC score | 0.971624 (rank : 3) | θ value | 2.28167e-128 (rank : 3) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q9H4F8, Q96F78 | Gene names | SMOC1 | |||
|
Domain Architecture |

|
|||||
| Description | SPARC-related modular calcium-binding protein 1 precursor (Secreted modular calcium-binding protein 1) (SMOC-1). | |||||
|
SMOC1_MOUSE
|
||||||
| NC score | 0.970263 (rank : 4) | θ value | 1.17183e-124 (rank : 4) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q8BLY1, Q9WVN9 | Gene names | Smoc1, Srg | |||
|
Domain Architecture |

|
|||||
| Description | SPARC-related modular calcium-binding protein 1 precursor (Secreted modular calcium-binding protein 1) (SMOC-1) (SPARC-related gene protein). | |||||
|
HG2A_MOUSE
|
||||||
| NC score | 0.537802 (rank : 5) | θ value | 5.81887e-07 (rank : 13) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P04441, O19452 | Gene names | Cd74, Ii | |||
|
Domain Architecture |

|
|||||
| Description | H-2 class II histocompatibility antigen gamma chain (MHC class II- associated invariant chain) (Ia antigen-associated invariant chain) (Ii) (CD74 antigen). | |||||
|
TICN2_MOUSE
|
||||||
| NC score | 0.533786 (rank : 6) | θ value | 1.47974e-10 (rank : 9) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9ER58 | Gene names | Spock2, Ticn2 | |||
|
Domain Architecture |

|
|||||
| Description | Testican-2 precursor (SPARC/osteonectin, CWCV, and Kazal-like domains proteoglycan 2). | |||||
|
TICN2_HUMAN
|
||||||
| NC score | 0.531194 (rank : 7) | θ value | 2.13673e-09 (rank : 10) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q92563, Q6UW87 | Gene names | SPOCK2, KIAA0275, TICN2 | |||
|
Domain Architecture |

|
|||||
| Description | Testican-2 precursor (SPARC/osteonectin, CWCV, and Kazal-like domains proteoglycan 2). | |||||
|
TICN1_HUMAN
|
||||||
| NC score | 0.496165 (rank : 8) | θ value | 2.21117e-06 (rank : 14) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q08629 | Gene names | SPOCK1, SPOCK, TIC1, TICN1 | |||
|
Domain Architecture |

|
|||||
| Description | Testican-1 precursor (Protein SPOCK). | |||||
|
TICN1_MOUSE
|
||||||
| NC score | 0.494463 (rank : 9) | θ value | 2.21117e-06 (rank : 15) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q62288 | Gene names | Spock1, Spock, Ticn1 | |||
|
Domain Architecture |

|
|||||
| Description | Testican-1 precursor (Protein SPOCK). | |||||
|
HG2A_HUMAN
|
||||||
| NC score | 0.492464 (rank : 10) | θ value | 1.09739e-05 (rank : 19) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P04233, Q14597, Q29832, Q5U0J8, Q8WLP6 | Gene names | CD74, DHLAG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HLA class II histocompatibility antigen gamma chain (HLA-DR antigens- associated invariant chain) (Ia antigen-associated invariant chain) (Ii) (p33) (CD74 antigen). | |||||
|
TICN3_HUMAN
|
||||||
| NC score | 0.483504 (rank : 11) | θ value | 4.1701e-05 (rank : 23) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9BQ16, O75705, Q6UW53, Q96Q26 | Gene names | SPOCK3, TICN3 | |||
|
Domain Architecture |

|
|||||
| Description | Testican-3 precursor (SPARC/osteonectin, CWCV, and Kazal-like domains proteoglycan 3). | |||||
|
TICN3_MOUSE
|
||||||
| NC score | 0.481074 (rank : 12) | θ value | 5.44631e-05 (rank : 25) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8BKV0, Q9ER59 | Gene names | Spock3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testican-3 precursor (SPARC/osteonectin, CWCV, and Kazal-like domains proteoglycan 3). | |||||
|
IBP1_HUMAN
|
||||||
| NC score | 0.399103 (rank : 13) | θ value | 4.1701e-05 (rank : 22) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P08833, Q8IYP5 | Gene names | IGFBP1, IBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Insulin-like growth factor-binding protein 1 precursor (IGFBP-1) (IBP- 1) (IGF-binding protein 1) (Placental protein 12) (PP12). | |||||
|
IBP3_HUMAN
|
||||||
| NC score | 0.397432 (rank : 14) | θ value | 8.40245e-06 (rank : 18) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P17936, Q2V509, Q6P1M6 | Gene names | IGFBP3, IBP3 | |||
|
Domain Architecture |

|
|||||
| Description | Insulin-like growth factor-binding protein 3 precursor (IGFBP-3) (IBP- 3) (IGF-binding protein 3). | |||||
|
IBP6_HUMAN
|
||||||
| NC score | 0.383824 (rank : 15) | θ value | 0.00509761 (rank : 37) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P24592, Q14492 | Gene names | IGFBP6, IBP6 | |||
|
Domain Architecture |

|
|||||
| Description | Insulin-like growth factor-binding protein 6 precursor (IGFBP-6) (IBP- 6) (IGF-binding protein 6). | |||||
|
IBP4_MOUSE
|
||||||
| NC score | 0.383085 (rank : 16) | θ value | 0.000461057 (rank : 32) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P47879, O35666 | Gene names | Igfbp4, Igfbp-4 | |||
|
Domain Architecture |

|
|||||
| Description | Insulin-like growth factor-binding protein 4 precursor (IGFBP-4) (IBP- 4) (IGF-binding protein 4). | |||||
|
IBP4_HUMAN
|
||||||
| NC score | 0.380905 (rank : 17) | θ value | 0.000461057 (rank : 31) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P22692 | Gene names | IGFBP4, IBP4 | |||
|
Domain Architecture |

|
|||||
| Description | Insulin-like growth factor-binding protein 4 precursor (IGFBP-4) (IBP- 4) (IGF-binding protein 4). | |||||
|
IBP1_MOUSE
|
||||||
| NC score | 0.374843 (rank : 18) | θ value | 7.1131e-05 (rank : 26) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P47876, Q61732 | Gene names | Igfbp1, Igfbp-1 | |||
|
Domain Architecture |

|
|||||
| Description | Insulin-like growth factor-binding protein 1 precursor (IGFBP-1) (IBP- 1) (IGF-binding protein 1). | |||||
|
IBP2_MOUSE
|
||||||
| NC score | 0.373028 (rank : 19) | θ value | 0.00035302 (rank : 30) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P47877, Q61722 | Gene names | Igfbp2, Igfbp-2 | |||
|
Domain Architecture |

|
|||||
| Description | Insulin-like growth factor-binding protein 2 precursor (IGFBP-2) (IBP- 2) (IGF-binding protein 2). | |||||
|
SPRC_MOUSE
|
||||||
| NC score | 0.370004 (rank : 20) | θ value | 1.87187e-05 (rank : 21) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P07214 | Gene names | Sparc | |||
|
Domain Architecture |

|
|||||
| Description | SPARC precursor (Secreted protein acidic and rich in cysteine) (Osteonectin) (ON) (Basement-membrane protein 40) (BM-40). | |||||
|
SPRC_HUMAN
|
||||||
| NC score | 0.368289 (rank : 21) | θ value | 5.44631e-05 (rank : 24) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P09486 | Gene names | SPARC, ON | |||
|
Domain Architecture |

|
|||||
| Description | SPARC precursor (Secreted protein acidic and rich in cysteine) (Osteonectin) (ON) (Basement-membrane protein 40) (BM-40). | |||||
|
THYG_HUMAN
|
||||||
| NC score | 0.368028 (rank : 22) | θ value | 1.99529e-15 (rank : 7) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P01266, O15274, O43899, Q15593, Q15948, Q9NYR1, Q9NYR2, Q9UMZ0, Q9UNY3 | Gene names | TG | |||
|
Domain Architecture |

|
|||||
| Description | Thyroglobulin precursor. | |||||
|
IBP6_MOUSE
|
||||||
| NC score | 0.367457 (rank : 23) | θ value | 0.00509761 (rank : 38) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P47880 | Gene names | Igfbp6, Igfbp-6 | |||
|
Domain Architecture |

|
|||||
| Description | Insulin-like growth factor-binding protein 6 precursor (IGFBP-6) (IBP- 6) (IGF-binding protein 6). | |||||
|
FST_HUMAN
|
||||||
| NC score | 0.366060 (rank : 24) | θ value | 2.88788e-06 (rank : 16) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P19883, Q9BTH0 | Gene names | FST | |||
|
Domain Architecture |

|
|||||
| Description | Follistatin precursor (FS) (Activin-binding protein). | |||||
|
IBP3_MOUSE
|
||||||
| NC score | 0.364015 (rank : 25) | θ value | 0.00869519 (rank : 40) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P47878 | Gene names | Igfbp3, Igfbp-3 | |||
|
Domain Architecture |

|
|||||
| Description | Insulin-like growth factor-binding protein 3 precursor (IGFBP-3) (IBP- 3) (IGF-binding protein 3). | |||||
|
FST_MOUSE
|
||||||
| NC score | 0.357866 (rank : 26) | θ value | 2.88788e-06 (rank : 17) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P47931 | Gene names | Fst | |||
|
Domain Architecture |

|
|||||
| Description | Follistatin precursor (FS) (Activin-binding protein). | |||||
|
FSTL3_HUMAN
|
||||||
| NC score | 0.352065 (rank : 27) | θ value | 1.43324e-05 (rank : 20) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O95633 | Gene names | FSTL3, FLRG | |||
|
Domain Architecture |

|
|||||
| Description | Follistatin-related protein 3 precursor (Follistatin-like 3) (Follistatin-related gene protein). | |||||
|
THYG_MOUSE
|
||||||
| NC score | 0.351411 (rank : 28) | θ value | 1.09485e-13 (rank : 8) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O08710, O88590, Q9QWY7 | Gene names | Tg, Tgn | |||
|
Domain Architecture |

|
|||||
| Description | Thyroglobulin precursor. | |||||
|
IBP2_HUMAN
|
||||||
| NC score | 0.347509 (rank : 29) | θ value | 0.00390308 (rank : 36) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P18065, Q14619, Q9UCL3 | Gene names | IGFBP2, BP2 | |||
|
Domain Architecture |

|
|||||
| Description | Insulin-like growth factor-binding protein 2 precursor (IGFBP-2) (IBP- 2) (IGF-binding protein 2). | |||||
|
IBP5_HUMAN
|
||||||
| NC score | 0.334640 (rank : 30) | θ value | 0.0736092 (rank : 43) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P24593 | Gene names | IGFBP5, IBP5 | |||
|
Domain Architecture |

|
|||||
| Description | Insulin-like growth factor-binding protein 5 precursor (IGFBP-5) (IBP- 5) (IGF-binding protein 5). | |||||
|
IBP5_MOUSE
|
||||||
| NC score | 0.331498 (rank : 31) | θ value | 0.125558 (rank : 46) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q07079 | Gene names | Igfbp5, Igfbp-5 | |||
|
Domain Architecture |

|
|||||
| Description | Insulin-like growth factor-binding protein 5 precursor (IGFBP-5) (IBP- 5) (IGF-binding protein 5). | |||||
|
FSTL3_MOUSE
|
||||||
| NC score | 0.326120 (rank : 32) | θ value | 0.000786445 (rank : 35) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9EQC7 | Gene names | Fstl3, Flrg | |||
|
Domain Architecture |

|
|||||
| Description | Follistatin-related protein 3 precursor (Follistatin-like 3) (Follistatin-related gene protein). | |||||
|
SPRL1_HUMAN
|
||||||
| NC score | 0.314258 (rank : 33) | θ value | 0.000602161 (rank : 34) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q14515, Q14800 | Gene names | SPARCL1 | |||
|
Domain Architecture |

|
|||||
| Description | SPARC-like protein 1 precursor (High endothelial venule protein) (Hevin) (MAST 9). | |||||
|
TEFF2_HUMAN
|
||||||
| NC score | 0.295073 (rank : 34) | θ value | 7.1131e-05 (rank : 27) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9UIK5, Q4ZFW4, Q53H90, Q53RE1, Q8N2R5, Q9NR15, Q9NSS5, Q9P2Y9, Q9UK65 | Gene names | TMEFF2, HPP1, TENB2, TPEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein with EGF-like and two follistatin-like domains 2 precursor (Transmembrane protein containing epidermal growth factor and follistatin domains) (Tomoregulin) (TR) (Hyperplastic polyposis protein 1). | |||||
|
TEFF2_MOUSE
|
||||||
| NC score | 0.294968 (rank : 35) | θ value | 7.1131e-05 (rank : 28) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9QYM9, Q3UY20, Q8CDH1, Q9JJE3 | Gene names | Tmeff2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein with EGF-like and two follistatin-like domains 2 precursor (Tomoregulin) (TR). | |||||
|
SPRL1_MOUSE
|
||||||
| NC score | 0.290212 (rank : 36) | θ value | 0.000270298 (rank : 29) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P70663, P97810, Q99L82 | Gene names | Sparcl1, Ecm2, Sc1 | |||
|
Domain Architecture |

|
|||||
| Description | SPARC-like protein 1 precursor (Matrix glycoprotein Sc1) (Extracellular matrix protein 2). | |||||
|
TACD1_HUMAN
|
||||||
| NC score | 0.289628 (rank : 37) | θ value | 0.125558 (rank : 47) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P16422, P18180, Q96C47 | Gene names | TACSTD1, GA733-2, M1S2, TROP1 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor-associated calcium signal transducer 1 precursor (Major gastrointestinal tumor-associated protein GA733-2) (Epithelial cell surface antigen) (Epithelial glycoprotein) (EGP) (Adenocarcinoma- associated antigen) (KSA) (KS 1/4 antigen) (Cell surface glycoprotein Trop-1) (CD326 antigen). | |||||
|
NID2_MOUSE
|
||||||
| NC score | 0.277989 (rank : 38) | θ value | 1.52774e-15 (rank : 5) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 338 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O88322 | Gene names | Nid2 | |||
|
Domain Architecture |

|
|||||
| Description | Nidogen-2 precursor (NID-2) (Entactin-2). | |||||
|
NID2_HUMAN
|
||||||
| NC score | 0.277443 (rank : 39) | θ value | 1.99529e-15 (rank : 6) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 456 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q14112, O43710 | Gene names | NID2 | |||
|
Domain Architecture |

|
|||||
| Description | Nidogen-2 precursor (NID-2) (Osteonidogen). | |||||
|
FSTL1_MOUSE
|
||||||
| NC score | 0.274189 (rank : 40) | θ value | 0.00869519 (rank : 39) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q62356, Q99JI9 | Gene names | Fstl1, Frp, Fstl, Tsc36 | |||
|
Domain Architecture |

|
|||||
| Description | Follistatin-related protein 1 precursor (Follistatin-like 1) (TGF- beta-inducible protein TSC-36). | |||||
|
FSTL1_HUMAN
|
||||||
| NC score | 0.274018 (rank : 41) | θ value | 0.0113563 (rank : 41) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q12841 | Gene names | FSTL1, FRP | |||
|
Domain Architecture |

|
|||||
| Description | Follistatin-related protein 1 precursor (Follistatin-like 1). | |||||
|
NID1_HUMAN
|
||||||
| NC score | 0.247221 (rank : 42) | θ value | 1.80886e-08 (rank : 11) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P14543, Q14942, Q59FL2, Q5TAF2, Q5TAF3, Q86XD7 | Gene names | NID1, NID | |||
|
Domain Architecture |

|
|||||
| Description | Nidogen-1 precursor (Entactin). | |||||
|
NID1_MOUSE
|
||||||
| NC score | 0.243140 (rank : 43) | θ value | 6.87365e-08 (rank : 12) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P10493, Q8BQI3, Q8C3U8, Q8C9P6 | Gene names | Nid1, Ent | |||
|
Domain Architecture |

|
|||||
| Description | Nidogen-1 precursor (Entactin). | |||||
|
TACD2_HUMAN
|
||||||
| NC score | 0.240966 (rank : 44) | θ value | 0.813845 (rank : 54) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P09758, Q15658, Q96QD2 | Gene names | TACSTD2, GA733-1, M1S1, TROP2 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor-associated calcium signal transducer 2 precursor (Pancreatic carcinoma marker protein GA733-1) (Cell surface glycoprotein Trop-2). | |||||
|
ISK6_MOUSE
|
||||||
| NC score | 0.183157 (rank : 45) | θ value | 1.06291 (rank : 57) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8BT20 | Gene names | Spink6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine protease inhibitor Kazal-type 6 precursor. | |||||
|
AGRIN_HUMAN
|
||||||
| NC score | 0.163549 (rank : 46) | θ value | 0.000602161 (rank : 33) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 582 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O00468, Q5SVA2, Q60FE1, Q7KYS8, Q8N4J5, Q96IC1, Q9BTD4 | Gene names | AGRIN, AGRN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Agrin precursor. | |||||
|
KAZD1_HUMAN
|
||||||
| NC score | 0.159649 (rank : 47) | θ value | 0.0961366 (rank : 44) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q96I82, Q6ZMB1, Q9BQ73 | Gene names | KAZALD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kazal-type serine protease inhibitor domain-containing protein 1 precursor. | |||||
|
KAZD1_MOUSE
|
||||||
| NC score | 0.156176 (rank : 48) | θ value | 0.0961366 (rank : 45) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8BJ66 | Gene names | Kazald1, Bono1, Igfbprp10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kazal-type serine protease inhibitor domain-containing protein 1 precursor (IGFBP-related protein 10) (IGFBP-rP10) (Bone and odontoblast expressed protein 1). | |||||
|
ISK6_HUMAN
|
||||||
| NC score | 0.110871 (rank : 49) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q6UWN8, Q8N5P0 | Gene names | SPINK6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine protease inhibitor Kazal-type 6 precursor. | |||||
|
ISK5_HUMAN
|
||||||
| NC score | 0.104052 (rank : 50) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9NQ38, O75770, Q96PP2, Q96PP3 | Gene names | SPINK5 | |||
|
Domain Architecture |

|
|||||
| Description | Serine protease inhibitor Kazal-type 5 precursor (Lympho-epithelial Kazal-type-related inhibitor) (LEKTI) [Contains: Hemofiltrate peptide HF6478; Hemofiltrate peptide HF7665]. | |||||
|
FSTL5_HUMAN
|
||||||
| NC score | 0.101607 (rank : 51) | θ value | 0.62314 (rank : 51) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8N475, Q9NSW7, Q9ULF7 | Gene names | FSTL5, KIAA1263 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Follistatin-related protein 5 precursor (Follistatin-like 5). | |||||
|
FSTL5_MOUSE
|
||||||
| NC score | 0.101225 (rank : 52) | θ value | 0.365318 (rank : 49) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 355 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8BFR2, Q80TG3, Q8C4T3 | Gene names | Fstl5, Kiaa1263 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Follistatin-related protein 5 precursor (Follistatin-like 5) (m- D/Bsp120I 1-2). | |||||
|
ISK4_MOUSE
|
||||||
| NC score | 0.100197 (rank : 53) | θ value | θ > 10 (rank : 99) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O35679 | Gene names | Spink4, Mpgc60 | |||
|
Domain Architecture |

|
|||||
| Description | Serine protease inhibitor Kazal-type 4 precursor (Peptide PEC-60 homolog) (MPGC60 protein). | |||||
|
HTRA1_MOUSE
|
||||||
| NC score | 0.097749 (rank : 54) | θ value | 5.27518 (rank : 80) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9R118 | Gene names | Htra1, Htra, Prss11 | |||
|
Domain Architecture |

|
|||||
| Description | Serine protease HTRA1 precursor (EC 3.4.21.-). | |||||
|
FSTL4_MOUSE
|
||||||
| NC score | 0.094566 (rank : 55) | θ value | 3.0926 (rank : 66) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q5STE3, Q5DU03 | Gene names | Fstl4, Kiaa1061 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Follistatin-related protein 4 precursor (Follistatin-like 4) (m- D/Bsp120I 1-1). | |||||
|
FSTL4_HUMAN
|
||||||
| NC score | 0.094541 (rank : 56) | θ value | θ > 10 (rank : 91) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q6MZW2, Q8TBU0, Q9UPU1 | Gene names | FSTL4, KIAA1061 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Follistatin-related protein 4 precursor (Follistatin-like 4). | |||||
|
ISK4_HUMAN
|
||||||
| NC score | 0.094088 (rank : 57) | θ value | θ > 10 (rank : 98) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O60575 | Gene names | SPINK4 | |||
|
Domain Architecture |

|
|||||
| Description | Serine protease inhibitor Kazal-type 4 precursor (Peptide PEC-60 homolog). | |||||
|
HTRA1_HUMAN
|
||||||
| NC score | 0.082267 (rank : 58) | θ value | θ > 10 (rank : 92) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q92743, Q9UNS5 | Gene names | HTRA1, HTRA, PRSS11 | |||
|
Domain Architecture |

|
|||||
| Description | Serine protease HTRA1 precursor (EC 3.4.21.-) (L56). | |||||
|
IBP7_HUMAN
|
||||||
| NC score | 0.081820 (rank : 59) | θ value | θ > 10 (rank : 93) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q16270, Q07822 | Gene names | IGFBP7, MAC25, PSF | |||
|
Domain Architecture |

|
|||||
| Description | Insulin-like growth factor-binding protein 7 precursor (IGFBP-7) (IBP- 7) (IGF-binding protein 7) (MAC25 protein) (Prostacyclin-stimulating factor) (PGI2-stimulating factor) (IGFBP-rP1). | |||||
|
IBP7_MOUSE
|
||||||
| NC score | 0.076390 (rank : 60) | θ value | θ > 10 (rank : 94) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q61581, O88812, Q9EQW0 | Gene names | Igfbp7, Mac25 | |||
|
Domain Architecture |

|
|||||
| Description | Insulin-like growth factor-binding protein 7 precursor (IGFBP-7) (IBP- 7) (IGF-binding protein 7) (MAC25 protein). | |||||
|
GUC1C_HUMAN
|
||||||
| NC score | 0.074216 (rank : 61) | θ value | 0.0563607 (rank : 42) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O95843, O95844, Q9UNM0 | Gene names | GUCA1C, GCAP3 | |||
|
Domain Architecture |

|
|||||
| Description | Guanylyl cyclase-activating protein 3 (GCAP 3) (Guanylate cyclase activator 1C). | |||||
|
IPK1_HUMAN
|
||||||
| NC score | 0.072427 (rank : 62) | θ value | θ > 10 (rank : 95) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P00995 | Gene names | SPINK1, PSTI | |||
|
Domain Architecture |

|
|||||
| Description | Pancreatic secretory trypsin inhibitor precursor (Tumor-associated trypsin inhibitor) (TATI) (Serine protease inhibitor Kazal-type 1). | |||||
|
ISK7_MOUSE
|
||||||
| NC score | 0.068464 (rank : 63) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6IE32 | Gene names | Spink7, Ecg2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine protease inhibitor Kazal-type 7 precursor (Esophagus cancer- related gene 2 protein) (ECRG-2). | |||||
|
ISK7_HUMAN
|
||||||
| NC score | 0.065718 (rank : 64) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P58062 | Gene names | SPINK7, ECG2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine protease inhibitor Kazal-type 7 precursor (Esophagus cancer- related gene 2 protein) (ECRG-2). | |||||
|
LRP5_HUMAN
|
||||||
| NC score | 0.058310 (rank : 65) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 224 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O75197, Q96TD6, Q9UP66 | Gene names | LRP5, LRP7 | |||
|
Domain Architecture |

|
|||||
| Description | Low-density lipoprotein receptor-related protein 5 precursor. | |||||
|
LRP5_MOUSE
|
||||||
| NC score | 0.058061 (rank : 66) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q91VN0, O88883, Q9R208 | Gene names | Lrp5, Lrp7 | |||
|
Domain Architecture |

|
|||||
| Description | Low-density lipoprotein receptor-related protein 5 precursor (LRP7) (Lr3). | |||||
|
LRP6_HUMAN
|
||||||
| NC score | 0.056441 (rank : 67) | θ value | θ > 10 (rank : 106) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O75581 | Gene names | LRP6 | |||
|
Domain Architecture |

|
|||||
| Description | Low-density lipoprotein receptor-related protein 6 precursor. | |||||
|
LRP6_MOUSE
|
||||||
| NC score | 0.056389 (rank : 68) | θ value | θ > 10 (rank : 107) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O88572 | Gene names | Lrp6 | |||
|
Domain Architecture |

|
|||||
| Description | Low-density lipoprotein receptor-related protein 6 precursor. | |||||
|
GUC1A_MOUSE
|
||||||
| NC score | 0.055220 (rank : 69) | θ value | 0.813845 (rank : 52) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P43081 | Gene names | Guca1a, Gcap, Gcap1, Guca1 | |||
|
Domain Architecture |

|
|||||
| Description | Guanylyl cyclase-activating protein 1 (GCAP 1) (Guanylate cyclase activator 1A). | |||||
|
EGF_HUMAN
|
||||||
| NC score | 0.054640 (rank : 70) | θ value | θ > 10 (rank : 89) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P01133 | Gene names | EGF | |||
|
Domain Architecture |

|
|||||
| Description | Pro-epidermal growth factor precursor (EGF) [Contains: Epidermal growth factor (Urogastrone)]. | |||||
|
EGF_MOUSE
|
||||||
| NC score | 0.054491 (rank : 71) | θ value | θ > 10 (rank : 90) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 358 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P01132, Q6P9J2 | Gene names | Egf | |||
|
Domain Architecture |

|
|||||
| Description | Pro-epidermal growth factor precursor (EGF) [Contains: Epidermal growth factor]. | |||||
|
IPK2_HUMAN
|
||||||
| NC score | 0.053452 (rank : 72) | θ value | θ > 10 (rank : 96) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P20155 | Gene names | SPINK2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine protease inhibitor Kazal-type 2 precursor (Acrosin-trypsin inhibitor) (HUSI-II). | |||||
|
TECTA_HUMAN
|
||||||
| NC score | 0.053281 (rank : 73) | θ value | θ > 10 (rank : 109) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O75443 | Gene names | TECTA | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-tectorin precursor. | |||||
|
RECK_HUMAN
|
||||||
| NC score | 0.051166 (rank : 74) | θ value | θ > 10 (rank : 108) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O95980, Q8WX37 | Gene names | RECK | |||
|
Domain Architecture |

|
|||||
| Description | Reversion-inducing cysteine-rich protein with Kazal motifs precursor (hRECK) (Suppressor of tumorigenicity 15) (ST15). | |||||
|
TECTA_MOUSE
|
||||||
| NC score | 0.051103 (rank : 75) | θ value | θ > 10 (rank : 110) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 244 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O08523 | Gene names | Tecta | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-tectorin precursor. | |||||
|
KCIP2_MOUSE
|
||||||
| NC score | 0.050701 (rank : 76) | θ value | 3.0926 (rank : 67) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9JJ69, Q6DTJ2, Q8K1T8, Q8K1T9, Q8K1U0, Q8VHN4, Q8VHN5, Q8VHN6, Q9JJ68 | Gene names | Kcnip2, Kchip2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kv channel-interacting protein 2 (KChIP2) (A-type potassium channel modulatory protein 2) (Potassium channel-interacting protein 2). | |||||
|
ISK3_MOUSE
|
||||||
| NC score | 0.050012 (rank : 77) | θ value | θ > 10 (rank : 97) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P09036, Q5M9M3 | Gene names | Spink3 | |||
|
Domain Architecture |

|
|||||
| Description | Serine protease inhibitor Kazal-type 3 precursor (Prostatic secretory glycoprotein) (P12). | |||||
|
KCIP2_HUMAN
|
||||||
| NC score | 0.049557 (rank : 78) | θ value | 4.03905 (rank : 70) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9NS61, Q7Z6F1, Q96K86, Q96T41, Q96T42, Q96T43, Q96T44, Q9H0N4, Q9HD10, Q9HD11, Q9NS60, Q9NY10, Q9NZI1 | Gene names | KCNIP2, KCHIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kv channel-interacting protein 2 (KChIP2) (A-type potassium channel modulatory protein 2) (Potassium channel-interacting protein 2) (Cardiac voltage-gated potassium channel modulatory subunit). | |||||
|
HPCA_HUMAN
|
||||||
| NC score | 0.048466 (rank : 79) | θ value | 1.38821 (rank : 58) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P84074, P32076, P41211, P70510 | Gene names | HPCA, BDR2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuron-specific calcium-binding protein hippocalcin (Calcium-binding protein BDR-2). | |||||
|
HPCA_MOUSE
|
||||||
| NC score | 0.048466 (rank : 80) | θ value | 1.38821 (rank : 59) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P84075, P32076, P41211, P70510 | Gene names | Hpca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuron-specific calcium-binding protein hippocalcin. | |||||
|
KCIP4_HUMAN
|
||||||
| NC score | 0.048041 (rank : 81) | θ value | 4.03905 (rank : 71) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6PIL6, Q4W5G8, Q8NEU0, Q9BWT2, Q9H294, Q9H2A4 | Gene names | KCNIP4, CALP, KCHIP4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kv channel-interacting protein 4 (KChIP4) (A-type potassium channel modulatory protein 4) (Potassium channel-interacting protein 4) (Calsenilin-like protein). | |||||
|
KCIP4_MOUSE
|
||||||
| NC score | 0.048030 (rank : 82) | θ value | 4.03905 (rank : 72) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6PHZ8, Q6DTJ3, Q8CAD0, Q8R4I2, Q9EQ01 | Gene names | Kcnip4, Calp, Kchip4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kv channel-interacting protein 4 (KChIP4) (A-type potassium channel modulatory protein 4) (Potassium channel-interacting protein 4) (Calsenilin-like protein). | |||||
|
GUC1A_HUMAN
|
||||||
| NC score | 0.046336 (rank : 83) | θ value | 5.27518 (rank : 79) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P43080, Q9NU14 | Gene names | GUCA1A, GCAP, GCAP1, GUCA1 | |||
|
Domain Architecture |

|
|||||
| Description | Guanylyl cyclase-activating protein 1 (GCAP 1) (Guanylate cyclase activator 1A). | |||||
|
HPCL1_MOUSE
|
||||||
| NC score | 0.046180 (rank : 84) | θ value | 2.36792 (rank : 64) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P62748, P35333 | Gene names | Hpcal1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hippocalcin-like protein 1 (Visinin-like protein 3) (VILIP-3) (Neural visinin-like protein 3) (NVL-3) (NVP-3). | |||||
|
HPCL1_HUMAN
|
||||||
| NC score | 0.045814 (rank : 85) | θ value | 2.36792 (rank : 63) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P37235, Q969S5 | Gene names | HPCAL1, BDR1 | |||
|
Domain Architecture |

|
|||||
| Description | Hippocalcin-like protein 1 (Visinin-like protein 3) (VILIP-3) (Calcium-binding protein BDR-1) (HLP2). | |||||
|
CSEN_HUMAN
|
||||||
| NC score | 0.045712 (rank : 86) | θ value | 6.88961 (rank : 83) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Y2W7, Q53TJ5, Q96T40, Q9UJ84, Q9UJ85 | Gene names | KCNIP3, CSEN, DREAM, KCHIP3 | |||
|
Domain Architecture |

|
|||||
| Description | Calsenilin (DRE-antagonist modulator) (DREAM) (Kv channel-interacting protein 3) (KChIP3) (A-type potassium channel modulatory protein 3). | |||||
|
PCY1B_MOUSE
|
||||||
| NC score | 0.042873 (rank : 87) | θ value | 0.279714 (rank : 48) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q811Q9, Q3UEW0, Q80Y63, Q811Q8, Q8BKD2, Q8C085 | Gene names | Pcyt1b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Choline-phosphate cytidylyltransferase B (EC 2.7.7.15) (Phosphorylcholine transferase B) (CTP:phosphocholine cytidylyltransferase B) (CT B) (CCT B) (CCT-beta). | |||||
|
NCALD_HUMAN
|
||||||
| NC score | 0.042298 (rank : 88) | θ value | 4.03905 (rank : 73) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P61601, P29554, Q8IYC3, Q9H0W2 | Gene names | NCALD | |||
|
Domain Architecture |

|
|||||
| Description | Neurocalcin delta. | |||||
|
NCALD_MOUSE
|
||||||
| NC score | 0.042298 (rank : 89) | θ value | 4.03905 (rank : 74) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q91X97, Q3TJS9, Q8BZN9 | Gene names | Ncald, D15Ertd412e | |||
|
Domain Architecture |

|
|||||
| Description | Neurocalcin delta. | |||||
|
PCY1B_HUMAN
|
||||||
| NC score | 0.041257 (rank : 90) | θ value | 0.47712 (rank : 50) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y5K3, O60621, Q86XC9 | Gene names | PCYT1B, CCTB | |||
|
Domain Architecture |

|
|||||
| Description | Choline-phosphate cytidylyltransferase B (EC 2.7.7.15) (Phosphorylcholine transferase B) (CTP:phosphocholine cytidylyltransferase B) (CT B) (CCT B) (CCT-beta). | |||||
|
S10A9_HUMAN
|
||||||
| NC score | 0.036475 (rank : 91) | θ value | 0.813845 (rank : 53) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P06702, Q9NYM0, Q9UCJ1 | Gene names | S100A9, CAGB, MRP14 | |||
|
Domain Architecture |

|
|||||
| Description | Protein S100-A9 (S100 calcium-binding protein A9) (Calgranulin-B) (Migration inhibitory factor-related protein 14) (MRP-14) (P14) (Leukocyte L1 complex heavy chain) (Calprotectin L1H subunit). | |||||
|
ANKY1_HUMAN
|
||||||
| NC score | 0.028968 (rank : 92) | θ value | 2.36792 (rank : 61) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9P2S6, Q8IYX5, Q8NDK5, Q9NX10 | Gene names | ANKMY1, TSAL1, ZMYND13 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin repeat and MYND domain-containing protein 1 (Testis-specific ankyrin-like protein 1) (Zinc-finger MYND domain-containing protein 13). | |||||
|
CFAI_MOUSE
|
||||||
| NC score | 0.028016 (rank : 93) | θ value | 1.06291 (rank : 55) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q61129, Q9WU07 | Gene names | Cfi, If | |||
|
Domain Architecture |

|
|||||
| Description | Complement factor I precursor (EC 3.4.21.45) (C3B/C4B inactivator) [Contains: Complement factor I heavy chain; Complement factor I light chain]. | |||||
|
ESCO1_HUMAN
|
||||||
| NC score | 0.027632 (rank : 94) | θ value | 1.06291 (rank : 56) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5FWF5, Q69YG4, Q69YS3, Q6IMD7, Q8N3Z5, Q8NBG2, Q96PX7 | Gene names | ESCO1, EFO1, KIAA1911 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyltransferase ESCO1 (EC 2.3.1.-) (Establishment of cohesion 1 homolog 1) (ECO1 homolog 1) (ESO1 homolog 1) (Establishment factor- like protein 1) (EFO1p) (hEFO1) (CTF7 homolog 1). | |||||
|
MADCA_HUMAN
|
||||||
| NC score | 0.025696 (rank : 95) | θ value | 1.38821 (rank : 60) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q13477, O60222, O75867 | Gene names | MADCAM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucosal addressin cell adhesion molecule 1 precursor (MAdCAM-1) (hMAdCAM-1). | |||||
|
LAMA3_MOUSE
|
||||||
| NC score | 0.023004 (rank : 96) | θ value | 3.0926 (rank : 68) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 859 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q61789, O08751, Q61788, Q61966, Q9JHQ7 | Gene names | Lama3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-3 chain precursor (Nicein subunit alpha). | |||||
|
TULP1_MOUSE
|
||||||
| NC score | 0.022318 (rank : 97) | θ value | 2.36792 (rank : 65) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Z273 | Gene names | Tulp1 | |||
|
Domain Architecture |

|
|||||
| Description | Tubby-related protein 1 (Tubby-like protein 1). | |||||
|
PKP3_MOUSE
|
||||||
| NC score | 0.019409 (rank : 98) | θ value | 3.0926 (rank : 69) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9QY23 | Gene names | Pkp3 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-3. | |||||
|
SO1A2_HUMAN
|
||||||
| NC score | 0.017096 (rank : 99) | θ value | 5.27518 (rank : 81) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P46721, Q9UGP7, Q9UL38 | Gene names | SLCO1A2, OATP, SLC21A3 | |||
|
Domain Architecture |

|
|||||
| Description | Solute carrier organic anion transporter family member 1A2 (Solute carrier family 21 member 3) (Sodium-independent organic anion transporter) (Organic anion-transporting polypeptide 1) (OATP1) (OATP- A). | |||||
|
LAMA2_HUMAN
|
||||||
| NC score | 0.016784 (rank : 100) | θ value | 6.88961 (rank : 85) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 1092 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P24043, Q14736, Q93022 | Gene names | LAMA2, LAMM | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-2 chain precursor (Laminin M chain) (Merosin heavy chain). | |||||
|
PKP3_HUMAN
|
||||||
| NC score | 0.015807 (rank : 101) | θ value | 4.03905 (rank : 75) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y446 | Gene names | PKP3 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-3. | |||||
|
CYGB_MOUSE
|
||||||
| NC score | 0.015369 (rank : 102) | θ value | 6.88961 (rank : 84) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9CX80 | Gene names | Cygb | |||
|
Domain Architecture |

|
|||||
| Description | Cytoglobin (Histoglobin) (HGb). | |||||
|
GP152_HUMAN
|
||||||
| NC score | 0.012754 (rank : 103) | θ value | 2.36792 (rank : 62) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 636 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8TDT2, Q86SM0 | Gene names | GPR152, PGR5 | |||
|
Domain Architecture |

|
|||||
| Description | Probable G-protein coupled receptor 152 (G-protein coupled receptor PGR5). | |||||
|
CABP5_MOUSE
|
||||||
| NC score | 0.011526 (rank : 104) | θ value | 6.88961 (rank : 82) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9JLK3 | Gene names | Cabp5 | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-binding protein 5 (CaBP5). | |||||
|
RNF31_HUMAN
|
||||||
| NC score | 0.008025 (rank : 105) | θ value | 8.99809 (rank : 88) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96EP0, Q86VI2, Q8TEI0, Q96GB4, Q96NF1, Q9H5F1, Q9NWD2 | Gene names | RNF31, ZIBRA | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 31 (Zinc in-between-RING-finger ubiquitin- associated domain protein). | |||||
|
SMCA2_HUMAN
|
||||||
| NC score | 0.004409 (rank : 106) | θ value | 4.03905 (rank : 76) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 503 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P51531 | Gene names | SMARCA2, BRM, SNF2A, SNF2L2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L2 (EC 3.6.1.-) (ATP- dependent helicase SMARCA2) (SNF2-alpha) (SWI/SNF-related matrix- associated actin-dependent regulator of chromatin subfamily A member 2) (hBRM). | |||||
|
MINT_MOUSE
|
||||||
| NC score | 0.001649 (rank : 107) | θ value | 8.99809 (rank : 87) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
SUHW4_MOUSE
|
||||||
| NC score | 0.000900 (rank : 108) | θ value | 4.03905 (rank : 77) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 737 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q68FE8, Q3T9B1, Q8BI82 | Gene names | Suhw4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppressor of hairy wing homolog 4. | |||||
|
PRDM2_HUMAN
|
||||||
| NC score | -0.000934 (rank : 109) | θ value | 6.88961 (rank : 86) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q13029, Q13149, Q14550, Q5VUL9 | Gene names | PRDM2, RIZ | |||
|
Domain Architecture |

|
|||||
| Description | PR domain zinc finger protein 2 (PR domain-containing protein 2) (Retinoblastoma protein-interacting zinc-finger protein) (Zinc finger protein RIZ) (MTE-binding protein) (MTB-ZF) (GATA-3-binding protein G3B). | |||||
|
ACM3_HUMAN
|
||||||
| NC score | -0.001803 (rank : 110) | θ value | 5.27518 (rank : 78) | |||
| Query Neighborhood Hits | 88 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P20309, Q4QRI3, Q5VXY2, Q9HB60 | Gene names | CHRM3 | |||
|
Domain Architecture |

|
|||||
| Description | Muscarinic acetylcholine receptor M3. | |||||