Please be patient as the page loads
|
TULP1_MOUSE
|
||||||
| SwissProt Accessions | Q9Z273 | Gene names | Tulp1 | |||
|
Domain Architecture |

|
|||||
| Description | Tubby-related protein 1 (Tubby-like protein 1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
TULP1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.962471 (rank : 2) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O00294, O43536 | Gene names | TULP1 | |||
|
Domain Architecture |

|
|||||
| Description | Tubby-related protein 1 (Tubby-like protein 1). | |||||
|
TULP1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 146 | |
| SwissProt Accessions | Q9Z273 | Gene names | Tulp1 | |||
|
Domain Architecture |

|
|||||
| Description | Tubby-related protein 1 (Tubby-like protein 1). | |||||
|
TUB_HUMAN
|
||||||
| θ value | 3.65967e-110 (rank : 3) | NC score | 0.954504 (rank : 3) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P50607, O00293 | Gene names | TUB | |||
|
Domain Architecture |

|
|||||
| Description | Tubby protein homolog. | |||||
|
TUB_MOUSE
|
||||||
| θ value | 4.77967e-110 (rank : 4) | NC score | 0.954057 (rank : 4) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P50586 | Gene names | Tub, Rd5 | |||
|
Domain Architecture |

|
|||||
| Description | Tubby protein. | |||||
|
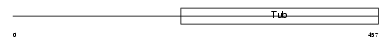
TULP3_MOUSE
|
||||||
| θ value | 1.48926e-103 (rank : 5) | NC score | 0.948106 (rank : 5) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O88413 | Gene names | Tulp3 | |||
|
Domain Architecture |
|
|||||
| Description | Tubby-related protein 3 (Tubby-like protein 3). | |||||
|
TULP3_HUMAN
|
||||||
| θ value | 6.7005e-96 (rank : 6) | NC score | 0.946875 (rank : 6) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O75386 | Gene names | TULP3 | |||
|
Domain Architecture |

|
|||||
| Description | Tubby-related protein 3 (Tubby-like protein 3). | |||||
|
TULP2_HUMAN
|
||||||
| θ value | 2.02684e-76 (rank : 7) | NC score | 0.933375 (rank : 7) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O00295 | Gene names | TULP2 | |||
|
Domain Architecture |

|
|||||
| Description | Tubby-related protein 2 (Tubby-like protein 2). | |||||
|
TULP4_MOUSE
|
||||||
| θ value | 2.43908e-13 (rank : 8) | NC score | 0.468495 (rank : 8) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9JIL5, Q8CA75, Q922C2 | Gene names | Tulp4, Tusp | |||
|
Domain Architecture |

|
|||||
| Description | Tubby-like protein 4 (Tubby superfamily protein). | |||||
|
TULP4_HUMAN
|
||||||
| θ value | 3.18553e-13 (rank : 9) | NC score | 0.428616 (rank : 9) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 426 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9NRJ4, Q9HD22, Q9P2F0 | Gene names | TULP4, KIAA1397, TUSP | |||
|
Domain Architecture |

|
|||||
| Description | Tubby-like protein 4 (Tubby superfamily protein). | |||||
|
NFH_MOUSE
|
||||||
| θ value | 5.26297e-08 (rank : 10) | NC score | 0.083632 (rank : 15) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 94 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
TCAL5_HUMAN
|
||||||
| θ value | 0.000786445 (rank : 11) | NC score | 0.081914 (rank : 16) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 806 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q5H9L2 | Gene names | TCEAL5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 5 (TCEA-like protein 5) (Transcription elongation factor S-II protein-like 5). | |||||
|
IWS1_MOUSE
|
||||||
| θ value | 0.00228821 (rank : 12) | NC score | 0.094043 (rank : 10) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
MPP8_HUMAN
|
||||||
| θ value | 0.00228821 (rank : 13) | NC score | 0.046348 (rank : 91) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 948 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q99549, Q86TK3, Q9BTP1 | Gene names | MPHOSPH8, MPP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | M-phase phosphoprotein 8. | |||||
|
MDC1_MOUSE
|
||||||
| θ value | 0.00869519 (rank : 14) | NC score | 0.065487 (rank : 36) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
VPS72_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 15) | NC score | 0.090607 (rank : 12) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q15906, Q53GJ2, Q5U0R4 | Gene names | VPS72, TCFL1, YL1 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 72 homolog (Transcription factor-like 1) (Protein YL-1). | |||||
|
MINT_MOUSE
|
||||||
| θ value | 0.0148317 (rank : 16) | NC score | 0.067980 (rank : 32) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
DCDC2_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 17) | NC score | 0.058718 (rank : 53) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q5DU00, Q5SZU0, Q80Y99 | Gene names | Dcdc2, Kiaa1154 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Doublecortin domain-containing protein 2. | |||||
|
VPS72_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 18) | NC score | 0.084857 (rank : 14) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q62481, Q3U2N4, Q810A9, Q99K81 | Gene names | Vps72, Tcfl1, Yl1 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 72 homolog (Transcription factor-like 1) (Protein YL-1). | |||||
|
IWS1_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 19) | NC score | 0.080458 (rank : 17) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
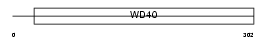
RBBP5_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 20) | NC score | 0.053013 (rank : 74) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 372 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q15291 | Gene names | RBBP5, RBQ3 | |||
|
Domain Architecture |
|
|||||
| Description | Retinoblastoma-binding protein 5 (RBBP-5) (Retinoblastoma-binding protein RBQ-3). | |||||
|
MYST3_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 21) | NC score | 0.054484 (rank : 68) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q92794 | Gene names | MYST3, MOZ, RUNXBP2, ZNF220 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Runt-related transcription factor-binding protein 2) (Monocytic leukemia zinc finger protein) (Zinc finger protein 220). | |||||
|
NIPBL_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 22) | NC score | 0.076402 (rank : 20) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
SNIP_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 23) | NC score | 0.048795 (rank : 89) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 319 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9QWI6, O70298 | Gene names | P140, Kiaa1684 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | p130Cas-associated protein (p140Cap) (SNAP-25-interacting protein) (SNIP). | |||||
|
TXLNA_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 24) | NC score | 0.025725 (rank : 126) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 982 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P40222, Q66K62, Q86T54, Q86T85, Q86T86, Q86Y86, Q86YW3, Q8N2Y3 | Gene names | TXLNA, TXLN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alpha-taxilin. | |||||
|
TXND2_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 25) | NC score | 0.075416 (rank : 21) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
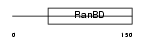
RANG_HUMAN
|
||||||
| θ value | 0.125558 (rank : 26) | NC score | 0.035582 (rank : 104) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P43487 | Gene names | RANBP1 | |||
|
Domain Architecture |
|
|||||
| Description | Ran-specific GTPase-activating protein (Ran-binding protein 1) (RanBP1). | |||||
|
DDX51_HUMAN
|
||||||
| θ value | 0.163984 (rank : 27) | NC score | 0.023588 (rank : 132) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8N8A6, Q5CZ71, Q8IXK5, Q96ED1 | Gene names | DDX51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX51 (EC 3.6.1.-) (DEAD box protein 51). | |||||
|
MY18A_MOUSE
|
||||||
| θ value | 0.163984 (rank : 28) | NC score | 0.018764 (rank : 145) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1989 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9JMH9, Q811D7 | Gene names | Myo18a, Myspdz | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
MDC1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 29) | NC score | 0.060811 (rank : 47) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
AHTF1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 30) | NC score | 0.050233 (rank : 82) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8CJF7, Q8BVJ5, Q8VD55 | Gene names | Ahctf1, Elys | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-hook-containing transcription factor 1 (Embryonic large molecule derived from yolk sac). | |||||
|
MSX1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 31) | NC score | 0.009283 (rank : 167) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P28360, Q96NY4 | Gene names | MSX1, HOX7 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein MSX-1 (Msh homeobox 1-like protein) (Hox-7). | |||||
|
NSD1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 32) | NC score | 0.032664 (rank : 112) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O88491, Q8C480, Q9CT70 | Gene names | Nsd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein). | |||||
|
TCAL3_MOUSE
|
||||||
| θ value | 0.365318 (rank : 33) | NC score | 0.090746 (rank : 11) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8R0A5, Q9CYK5, Q9D1S4 | Gene names | Tceal3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 3 (TCEA-like protein 3) (Transcription elongation factor S-II protein-like 3). | |||||
|
TTF1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 34) | NC score | 0.069116 (rank : 27) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q62187, Q9JKK5 | Gene names | Ttf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription termination factor 1 (TTF-1) (TTF-I) (mTFF-I) (RNA polymerase I termination factor). | |||||
|
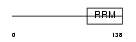
IF4B_HUMAN
|
||||||
| θ value | 0.47712 (rank : 35) | NC score | 0.049187 (rank : 87) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P23588 | Gene names | EIF4B | |||
|
Domain Architecture |
|
|||||
| Description | Eukaryotic translation initiation factor 4B (eIF-4B). | |||||
|
TCAL3_HUMAN
|
||||||
| θ value | 0.47712 (rank : 36) | NC score | 0.080451 (rank : 18) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q969E4 | Gene names | TCEAL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 3 (TCEA-like protein 3) (Transcription elongation factor S-II protein-like 3). | |||||
|
ALKB5_HUMAN
|
||||||
| θ value | 0.62314 (rank : 37) | NC score | 0.033289 (rank : 110) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6P6C2, Q9NXD6 | Gene names | ALKBH5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alkylated repair protein alkB homolog 5. | |||||
|
CDKL2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 38) | NC score | 0.003522 (rank : 181) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 854 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9QUK0, Q3UL60, Q3V3X7, Q9QYI1, Q9QYI2 | Gene names | Cdkl2, Kkiamre, Kkm | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-dependent kinase-like 2 (EC 2.7.11.22) (Serine/threonine- protein kinase KKIAMRE). | |||||
|
CJ012_HUMAN
|
||||||
| θ value | 0.62314 (rank : 39) | NC score | 0.070202 (rank : 24) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8N655, Q9H945, Q9Y457 | Gene names | C10orf12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf12. | |||||
|
MBB1A_HUMAN
|
||||||
| θ value | 0.62314 (rank : 40) | NC score | 0.057034 (rank : 59) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 665 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9BQG0, Q86VM3, Q9BW49, Q9P0V5, Q9UF99 | Gene names | MYBBP1A, P160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myb-binding protein 1A. | |||||
|
AFF4_MOUSE
|
||||||
| θ value | 0.813845 (rank : 41) | NC score | 0.071139 (rank : 23) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9ESC8, Q8C6K4, Q8C6W3, Q8CCH3 | Gene names | Aff4, Alf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4. | |||||
|
ELOA1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 42) | NC score | 0.044317 (rank : 93) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q14241, Q8IXH1 | Gene names | TCEB3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription elongation factor B polypeptide 3 (RNA polymerase II transcription factor SIII subunit A1) (SIII p110) (Elongin-A) (EloA) (Elongin 110 kDa subunit). | |||||
|
HAIR_MOUSE
|
||||||
| θ value | 0.813845 (rank : 43) | NC score | 0.026184 (rank : 125) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q61645, Q80Y47 | Gene names | Hr | |||
|
Domain Architecture |

|
|||||
| Description | Protein hairless. | |||||
|
NPBL_MOUSE
|
||||||
| θ value | 0.813845 (rank : 44) | NC score | 0.076696 (rank : 19) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
IF4E3_HUMAN
|
||||||
| θ value | 1.06291 (rank : 45) | NC score | 0.039361 (rank : 98) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O60573, O75349 | Gene names | EIF4EL3 | |||
|
Domain Architecture |
|
|||||
| Description | Eukaryotic translation initiation factor 4E type 3 (eIF4E type 3) (eIF-4E type 3) (mRNA cap-binding protein type 3) (Eukaryotic translation initiation factor 4E-like 3) (Eukaryotic translation initiation factor 4E homologous protein) (mRNA cap-binding protein 4EHP) (eIF4E-like protein 4E-LP). | |||||
|
IF4E3_MOUSE
|
||||||
| θ value | 1.06291 (rank : 46) | NC score | 0.039379 (rank : 97) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BMB3, O88503, Q3UCK2 | Gene names | Eif4el3 | |||
|
Domain Architecture |
|
|||||
| Description | Eukaryotic translation initiation factor 4E type 3 (eIF4E type 3) (eIF-4E type 3) (mRNA cap-binding protein type 3) (Eukaryotic translation initiation factor 4E-like 3) (eIF4E-like protein 4E-LP). | |||||
|
MLX_HUMAN
|
||||||
| θ value | 1.06291 (rank : 47) | NC score | 0.033407 (rank : 109) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UH92, Q53XM6, Q96FL2, Q9H2V0, Q9H2V1, Q9H2V2, Q9NXN3 | Gene names | MLX, TCFL4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
TBC10_HUMAN
|
||||||
| θ value | 1.06291 (rank : 48) | NC score | 0.018574 (rank : 146) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9BXI6, O76053, Q543A2 | Gene names | TBC1D10A, EPI64, TBC1D10 | |||
|
Domain Architecture |
|
|||||
| Description | TBC1 domain family member 10A (EBP50-PDX interactor of 64 kDa) (EPI64 protein). | |||||
|
WDFY3_HUMAN
|
||||||
| θ value | 1.06291 (rank : 49) | NC score | 0.013274 (rank : 155) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 348 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8IZQ1, Q4W5K5, Q6P0Q5, Q8N1T2, Q96BS7, Q96N85, Q9Y2J7 | Gene names | WDFY3, KIAA0993 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat and FYVE domain-containing protein 3 (Autophagy-linked FYVE protein) (Alfy). | |||||
|
XYLT1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 50) | NC score | 0.021592 (rank : 138) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q811B1, Q9EPL1 | Gene names | Xylt1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Xylosyltransferase 1 (EC 2.4.2.26) (Xylosyltransferase I) (Peptide O- xylosyltransferase 1). | |||||
|
CTRO_HUMAN
|
||||||
| θ value | 1.38821 (rank : 51) | NC score | 0.006343 (rank : 174) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 2092 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O14578, Q6XUH8, Q86UQ9, Q9UPZ7 | Gene names | CIT, KIAA0949, STK21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Citron Rho-interacting kinase (EC 2.7.11.1) (CRIK) (Rho-interacting, serine/threonine-protein kinase 21). | |||||
|
GP179_HUMAN
|
||||||
| θ value | 1.38821 (rank : 52) | NC score | 0.045001 (rank : 92) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 372 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q6PRD1 | Gene names | GPR179, GPR158L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 179 precursor (Probable G-protein coupled receptor 158-like 1). | |||||
|
NFH_HUMAN
|
||||||
| θ value | 1.38821 (rank : 53) | NC score | 0.053073 (rank : 73) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
RBBP6_MOUSE
|
||||||
| θ value | 1.38821 (rank : 54) | NC score | 0.064690 (rank : 38) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
SEC63_MOUSE
|
||||||
| θ value | 1.38821 (rank : 55) | NC score | 0.030818 (rank : 113) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 601 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8VHE0, Q8VEB9 | Gene names | Sec63, Sec63l | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
SMOC2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 56) | NC score | 0.023394 (rank : 133) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CD91, Q8VCM1, Q9D9K2, Q9ER95 | Gene names | Smoc2 | |||
|
Domain Architecture |
|
|||||
| Description | SPARC-related modular calcium-binding protein 2 precursor (Secreted modular calcium-binding protein 2) (SMOC-2). | |||||
|
STK39_HUMAN
|
||||||
| θ value | 1.38821 (rank : 57) | NC score | 0.004804 (rank : 178) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 882 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UEW8, O14774, Q9UER4 | Gene names | STK39, SPAK | |||
|
Domain Architecture |
|
|||||
| Description | STE20/SPS1-related proline-alanine-rich protein kinase (EC 2.7.11.1) (Ste-20-related kinase) (Serine/threonine-protein kinase 39) (DCHT). | |||||
|
STK39_MOUSE
|
||||||
| θ value | 1.38821 (rank : 58) | NC score | 0.005032 (rank : 177) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 915 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Z1W9, Q80W13 | Gene names | Stk39, Spak | |||
|
Domain Architecture |
|
|||||
| Description | STE20/SPS1-related proline-alanine-rich protein kinase (EC 2.7.11.1) (Ste-20-related kinase) (Serine/threonine-protein kinase 39). | |||||
|
SYTL1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 59) | NC score | 0.016924 (rank : 150) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 314 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q99N80, Q99J26 | Gene names | Sytl1, Slp1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 1 (Exophilin-7). | |||||
|
ADDB_MOUSE
|
||||||
| θ value | 1.81305 (rank : 60) | NC score | 0.042762 (rank : 95) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 574 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9QYB8, Q3U0E1, Q80VH9, Q8C1C4, Q9CXE3, Q9JLE5, Q9QYB7 | Gene names | Add2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-adducin (Erythrocyte adducin subunit beta) (Add97). | |||||
|
MYLK_MOUSE
|
||||||
| θ value | 1.81305 (rank : 61) | NC score | 0.006983 (rank : 172) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1489 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q6PDN3, Q3TSJ7, Q80UX0, Q80YN7, Q80YN8, Q8K026, Q924D2, Q9ERD3 | Gene names | Mylk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin light chain kinase, smooth muscle (EC 2.7.11.18) (MLCK) (Telokin) (Kinase-related protein) (KRP). | |||||
|
PRB4S_HUMAN
|
||||||
| θ value | 1.81305 (rank : 62) | NC score | 0.035169 (rank : 106) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P10163, P02813 | Gene names | PRB4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 4 allele S precursor (Salivary proline-rich protein Po) (Parotid o protein) [Contains: Protein N1; Glycosylated protein A]. | |||||
|
SV2B_MOUSE
|
||||||
| θ value | 1.81305 (rank : 63) | NC score | 0.012060 (rank : 159) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BG39, Q80TT1, Q9ES95 | Gene names | Sv2b, Kiaa0735 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptic vesicle glycoprotein 2B (Synaptic vesicle protein 2B). | |||||
|
TGON2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 64) | NC score | 0.057281 (rank : 58) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
TOX_MOUSE
|
||||||
| θ value | 1.81305 (rank : 65) | NC score | 0.022415 (rank : 135) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q66JW3, Q8BKH9, Q8BYQ5, Q8R4H0 | Gene names | Tox | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thymus high mobility group box protein TOX. | |||||
|
ZAR1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 66) | NC score | 0.035680 (rank : 103) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q86SH2 | Gene names | ZAR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zygote arrest 1 (Oocyte-specific maternal effect factor). | |||||
|
ARI5B_HUMAN
|
||||||
| θ value | 2.36792 (rank : 67) | NC score | 0.036930 (rank : 101) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q14865, Q7Z3M4, Q8N421, Q9H786 | Gene names | ARID5B, DESRT, MRF2 | |||
|
Domain Architecture |
|
|||||
| Description | AT-rich interactive domain-containing protein 5B (ARID domain- containing protein 5B) (Mrf1-like) (Modulator recognition factor 2) (MRF-2). | |||||
|
CCD55_MOUSE
|
||||||
| θ value | 2.36792 (rank : 68) | NC score | 0.044068 (rank : 94) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q5NCR9 | Gene names | Ccdc55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 55. | |||||
|
DDX21_MOUSE
|
||||||
| θ value | 2.36792 (rank : 69) | NC score | 0.029511 (rank : 120) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9JIK5, Q9WV45 | Gene names | Ddx21 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar RNA helicase 2 (EC 3.6.1.-) (Nucleolar RNA helicase II) (Nucleolar RNA helicase Gu) (RH II/Gu) (Gu-alpha) (DEAD box protein 21). | |||||
|
FLNA_HUMAN
|
||||||
| θ value | 2.36792 (rank : 70) | NC score | 0.015392 (rank : 153) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P21333 | Gene names | FLNA, FLN, FLN1 | |||
|
Domain Architecture |

|
|||||
| Description | Filamin-A (Alpha-filamin) (Filamin-1) (Endothelial actin-binding protein) (Actin-binding protein 280) (ABP-280) (Nonmuscle filamin). | |||||
|
FLNA_MOUSE
|
||||||
| θ value | 2.36792 (rank : 71) | NC score | 0.015521 (rank : 152) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BTM8, O54934, Q7TQI1, Q8BLK1, Q8BTN7, Q8VHX5, Q8VHX8, Q99KQ2 | Gene names | Flna, Fln, Fln1 | |||
|
Domain Architecture |

|
|||||
| Description | Filamin-A (Alpha-filamin) (Filamin-1) (Endothelial actin-binding protein) (Actin-binding protein 280) (ABP-280) (Nonmuscle filamin). | |||||
|
FYB_HUMAN
|
||||||
| θ value | 2.36792 (rank : 72) | NC score | 0.049038 (rank : 88) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 379 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O15117, O00359 | Gene names | FYB, SLAP130 | |||
|
Domain Architecture |

|
|||||
| Description | FYN-binding protein (FYN-T-binding protein) (FYB-120/130) (p120/p130) (SLP-76-associated phosphoprotein) (SLAP-130). | |||||
|
GCX1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 73) | NC score | 0.049476 (rank : 86) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96NM4, Q5TE33, Q5TE34, Q5TE35, Q96IC9, Q9BQN5 | Gene names | GCX1, C20orf100 | |||
|
Domain Architecture |

|
|||||
| Description | Granulosa cell HMG box protein 1 (GCX-1). | |||||
|
LAD1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 74) | NC score | 0.064354 (rank : 41) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O00515, O95614 | Gene names | LAD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ladinin 1 (Lad-1) (120 kDa linear IgA bullous dermatosis antigen) (97 kDa linear IgA bullous dermatosis antigen) (Linear IgA disease antigen homolog) (LadA). | |||||
|
MLX_MOUSE
|
||||||
| θ value | 2.36792 (rank : 75) | NC score | 0.030106 (rank : 116) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O08609, Q9EQJ7, Q9EQJ8 | Gene names | Mlx, Tcfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
SFR12_MOUSE
|
||||||
| θ value | 2.36792 (rank : 76) | NC score | 0.064595 (rank : 39) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8BZX4, Q8BLM8, Q8BWK7, Q8BYK0, Q8BYZ4 | Gene names | Sfrs12, Srrp86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86). | |||||
|
SMOC2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 77) | NC score | 0.022318 (rank : 136) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H3U7, Q5TAT7, Q5TAT8, Q86VV9, Q96SF3, Q9H1L3, Q9H1L4, Q9H3U0, Q9H4F7, Q9HCV2 | Gene names | SMOC2, SMAP2 | |||
|
Domain Architecture |
|
|||||
| Description | SPARC-related modular calcium-binding protein 2 precursor (Secreted modular calcium-binding protein 2) (SMOC-2) (Smooth muscle-associated protein 2) (SMAP-2). | |||||
|
TACC2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 78) | NC score | 0.024787 (rank : 128) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 856 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O95359, Q9NZ41, Q9NZR5 | Gene names | TACC2 | |||
|
Domain Architecture |

|
|||||
| Description | Transforming acidic coiled-coil-containing protein 2 (Anti Zuai-1) (AZU-1). | |||||
|
TBX2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 79) | NC score | 0.012431 (rank : 157) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q60707 | Gene names | Tbx2 | |||
|
Domain Architecture |

|
|||||
| Description | T-box transcription factor TBX2 (T-box protein 2). | |||||
|
TRIPC_HUMAN
|
||||||
| θ value | 2.36792 (rank : 80) | NC score | 0.027568 (rank : 124) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 319 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q14669, Q15644 | Gene names | TRIP12, KIAA0045 | |||
|
Domain Architecture |

|
|||||
| Description | Thyroid receptor-interacting protein 12 (TRIP12). | |||||
|
CFDP1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 81) | NC score | 0.033618 (rank : 107) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UEE9, O00393, O00404, Q9UEF0, Q9UEF1, Q9UEF8 | Gene names | CFDP1, BCNT | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Craniofacial development protein 1 (Bucentaur). | |||||
|
CGRE1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 82) | NC score | 0.028076 (rank : 123) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q99674 | Gene names | CGREF1, CGR11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell growth regulator with EF hand domain 1 (Cell growth regulatory gene 11 protein). | |||||
|
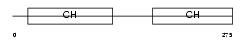
CLMN_HUMAN
|
||||||
| θ value | 3.0926 (rank : 83) | NC score | 0.020307 (rank : 143) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q96JQ2, Q9H713, Q9HA23, Q9HA57, Q9UFP4, Q9ULN2 | Gene names | CLMN, KIAA1188 | |||
|
Domain Architecture |
|
|||||
| Description | Calmin. | |||||
|
K1143_MOUSE
|
||||||
| θ value | 3.0926 (rank : 84) | NC score | 0.029529 (rank : 119) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8K039, Q9CSR4, Q9D0W8 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1143 homolog. | |||||
|
MINK1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 85) | NC score | 0.005861 (rank : 175) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8N4C8, Q9P1X1, Q9P2R8 | Gene names | MINK1, MAP4K6, MINK | |||
|
Domain Architecture |

|
|||||
| Description | Misshapen-like kinase 1 (EC 2.7.11.1) (Mitogen-activated protein kinase kinase kinase kinase 6) (MAPK/ERK kinase kinase kinase 6) (MEK kinase kinase 6) (MEKKK 6) (Misshapen/NIK-related kinase) (GCK family kinase MiNK). | |||||
|
PEG3_MOUSE
|
||||||
| θ value | 3.0926 (rank : 86) | NC score | 0.006595 (rank : 173) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1101 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q3URU2, O54978, Q3TQ69, Q5EBP7, Q61138, Q6GQS0, Q80U47, Q8R5N0, Q9QX53 | Gene names | Peg3, Kiaa0287, Pw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Paternally expressed gene 3 protein (ASF-1). | |||||
|
RAPH1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 87) | NC score | 0.037223 (rank : 100) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q70E73, Q96Q37, Q9C0I2 | Gene names | RAPH1, ALS2CR9, KIAA1681, LPD, PREL2, RMO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras-associated and pleckstrin homology domains-containing protein 1 (RAPH1) (Lamellipodin) (Proline-rich EVH1 ligand 2) (PREL-2) (Protein RMO1) (Amyotrophic lateral sclerosis 2 chromosomal region candidate 9 gene protein). | |||||
|
SETBP_MOUSE
|
||||||
| θ value | 3.0926 (rank : 88) | NC score | 0.029549 (rank : 118) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Z180, Q66JL8 | Gene names | Setbp1, Kiaa0437 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SET-binding protein (SEB). | |||||
|
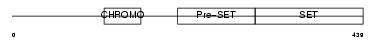
SUV92_MOUSE
|
||||||
| θ value | 3.0926 (rank : 89) | NC score | 0.017243 (rank : 149) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9EQQ0, Q9CUK3, Q9JLP7 | Gene names | Suv39h2 | |||
|
Domain Architecture |
|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 2 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 2) (H3-K9-HMTase 2) (Suppressor of variegation 3-9 homolog 2) (Su(var)3-9 homolog 2). | |||||
|
CF010_HUMAN
|
||||||
| θ value | 4.03905 (rank : 90) | NC score | 0.066673 (rank : 33) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q5SRN2, Q5SPI9, Q5SPJ0, Q5SPK9, Q5SPL0, Q5SRN3, Q5TG25, Q5TG26, Q8N4B6 | Gene names | C6orf10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C6orf10. | |||||
|
DUET_HUMAN
|
||||||
| θ value | 4.03905 (rank : 91) | NC score | 0.002947 (rank : 182) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1220 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Y2A5, Q8TBQ5, Q9NSZ4 | Gene names | DUET, TRAD | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Duet (EC 2.7.11.1) (Serine/threonine kinase with Dbl- and pleckstrin homology domain). | |||||
|
HNRPU_HUMAN
|
||||||
| θ value | 4.03905 (rank : 92) | NC score | 0.020611 (rank : 142) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q00839, O75507, Q96HY9 | Gene names | HNRPU, SAFA, U21.1 | |||
|
Domain Architecture |

|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein U (hnRNP U) (Scaffold attachment factor A) (SAF-A) (p120) (pp120). | |||||
|
IF2B2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 93) | NC score | 0.013143 (rank : 156) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q5SF07, Q3TCU4, Q7TQF9 | Gene names | Igf2bp2, Imp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Insulin-like growth factor 2 mRNA-binding protein 2 (IGF-II mRNA- binding protein 2) (IMP-2). | |||||
|
LRRF1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 94) | NC score | 0.061242 (rank : 44) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 440 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q3UZ39, O70323, Q3TS94 | Gene names | Lrrfip1, Flap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat flightless-interacting protein 1 (LRR FLII- interacting protein 1) (FLI-LRR-associated protein 1) (Flap-1) (H186 FLAP). | |||||
|
MBB1A_MOUSE
|
||||||
| θ value | 4.03905 (rank : 95) | NC score | 0.059398 (rank : 52) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q7TPV4, O35851, Q80Y66, Q8R4X2, Q99KP0 | Gene names | Mybbp1a, P160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myb-binding protein 1A (Myb-binding protein of 160 kDa). | |||||
|
PCLO_HUMAN
|
||||||
| θ value | 4.03905 (rank : 96) | NC score | 0.041754 (rank : 96) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
REEP4_MOUSE
|
||||||
| θ value | 4.03905 (rank : 97) | NC score | 0.010124 (rank : 165) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8K072 | Gene names | Reep4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Receptor expression-enhancing protein 4. | |||||
|
AATF_HUMAN
|
||||||
| θ value | 5.27518 (rank : 98) | NC score | 0.021189 (rank : 140) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NY61, Q9P0A4, Q9UNX5 | Gene names | AATF, CHE1, DED | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein AATF (Apoptosis-antagonizing transcription factor) (Rb-binding protein Che-1). | |||||
|
ANK2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 99) | NC score | 0.022788 (rank : 134) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
CCNK_MOUSE
|
||||||
| θ value | 5.27518 (rank : 100) | NC score | 0.030029 (rank : 117) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O88874, Q8R068 | Gene names | Ccnk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-K. | |||||
|
CMTA1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 101) | NC score | 0.012189 (rank : 158) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y6Y1, Q5VUE1 | Gene names | CAMTA1, KIAA0833 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calmodulin-binding transcription activator 1. | |||||
|
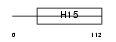
H12_HUMAN
|
||||||
| θ value | 5.27518 (rank : 102) | NC score | 0.025441 (rank : 127) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P16403 | Gene names | HIST1H1C, H1F2 | |||
|
Domain Architecture |
|
|||||
| Description | Histone H1.2 (Histone H1d). | |||||
|
H6ST3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 103) | NC score | 0.011138 (rank : 162) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IZP7, Q5W0L0, Q68CW6 | Gene names | HS6ST3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heparan-sulfate 6-O-sulfotransferase 3 (EC 2.8.2.-) (HS6ST-3). | |||||
|
HS105_MOUSE
|
||||||
| θ value | 5.27518 (rank : 104) | NC score | 0.007354 (rank : 171) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q61699, Q3TNS2, Q3UIY8, Q62578, Q62579, Q6A0A5, Q8C430, Q8VCW6 | Gene names | Hsph1, Hsp105, Hsp110, Kiaa0201 | |||
|
Domain Architecture |

|
|||||
| Description | Heat-shock protein 105 kDa (Heat shock-related 100 kDa protein E7I) (HSP-E7I) (Heat shock 110 kDa protein) (42 degrees C-HSP). | |||||
|
MLTK_HUMAN
|
||||||
| θ value | 5.27518 (rank : 105) | NC score | 0.001417 (rank : 184) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 911 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9NYL2, Q53SX1, Q580W8, Q59GY5, Q86YW8, Q9HCC4, Q9HCC5, Q9HDD2, Q9NYE9 | Gene names | MLTK, ZAK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase MLT (EC 2.7.11.25) (MLK-like mitogen-activated protein triple kinase) (Leucine zipper- and sterile alpha motif-containing kinase) (Sterile alpha motif- and leucine zipper-containing kinase AZK) (Mixed lineage kinase-related kinase) (MLK-related kinase) (MRK) (Cervical cancer suppressor gene 4 protein). | |||||
|
MY18A_HUMAN
|
||||||
| θ value | 5.27518 (rank : 106) | NC score | 0.014740 (rank : 154) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1929 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q92614, Q8IXP8 | Gene names | MYO18A, KIAA0216, MYSPDZ | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
MY18B_HUMAN
|
||||||
| θ value | 5.27518 (rank : 107) | NC score | 0.021858 (rank : 137) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1299 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q8IUG5, Q8NDI8, Q8TE65, Q8WWS0, Q96KH2, Q96KR8, Q96KR9 | Gene names | MYO18B | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18B (Myosin XVIIIb). | |||||
|
NSBP1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 108) | NC score | 0.052879 (rank : 76) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
SASH1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 109) | NC score | 0.015742 (rank : 151) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P59808 | Gene names | Sash1 | |||
|
Domain Architecture |

|
|||||
| Description | SAM and SH3 domain-containing protein 1. | |||||
|
SFPQ_HUMAN
|
||||||
| θ value | 5.27518 (rank : 110) | NC score | 0.024180 (rank : 130) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 986 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P23246, P30808 | Gene names | SFPQ, PSF | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, proline- and glutamine-rich (Polypyrimidine tract- binding protein-associated-splicing factor) (PTB-associated-splicing factor) (PSF) (DNA-binding p52/p100 complex, 100 kDa subunit) (100 kDa DNA-pairing protein) (hPOMp100). | |||||
|
AFF4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 111) | NC score | 0.070124 (rank : 25) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 611 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9UHB7, Q498B2, Q59FB3, Q6P592, Q8TDR1, Q9P0E4 | Gene names | AFF4, AF5Q31, MCEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4 (ALL1-fused gene from chromosome 5q31) (Major CDK9 elongation factor-associated protein). | |||||
|
ASCC2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 112) | NC score | 0.011567 (rank : 161) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9H1I8, Q4TT54, Q8TAZ0, Q9H711, Q9H9D6 | Gene names | ASCC2, ASC1P100 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Activating signal cointegrator 1 complex subunit 2 (ASC-1 complex subunit p100) (Trip4 complex subunit p100). | |||||
|
CFDP1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 113) | NC score | 0.032935 (rank : 111) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O88271, O70565, Q9JMA5 | Gene names | Cfdp1, Bcnt, Cfdp, Cp27 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Craniofacial development protein 1 (Bucentaur) (27 kDa craniofacial protein) (Protein Cp27). | |||||
|
DNMT1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 114) | NC score | 0.065422 (rank : 37) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P26358, Q9UHG5, Q9ULA2, Q9UMZ6 | Gene names | DNMT1, AIM, DNMT | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 1 (EC 2.1.1.37) (Dnmt1) (DNA methyltransferase HsaI) (DNA MTase HsaI) (MCMT) (M.HsaI). | |||||
|
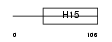
H13_MOUSE
|
||||||
| θ value | 6.88961 (rank : 115) | NC score | 0.030633 (rank : 115) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P43277, Q8C6M4 | Gene names | Hist1h1d, H1f3 | |||
|
Domain Architecture |
|
|||||
| Description | Histone H1.3 (H1 VAR.4) (H1d). | |||||
|
MADCA_HUMAN
|
||||||
| θ value | 6.88961 (rank : 116) | NC score | 0.033465 (rank : 108) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q13477, O60222, O75867 | Gene names | MADCAM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucosal addressin cell adhesion molecule 1 precursor (MAdCAM-1) (hMAdCAM-1). | |||||
|
MLL2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 117) | NC score | 0.035290 (rank : 105) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
NCOR2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 118) | NC score | 0.036356 (rank : 102) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 790 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9WU42, Q9WU43, Q9WUC1 | Gene names | Ncor2, Smrt | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC). | |||||
|
PER3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 119) | NC score | 0.011020 (rank : 163) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P56645, Q5H8X4, Q969K6, Q96S77, Q96S78, Q9C0J3, Q9NSP9, Q9UGU8 | Gene names | PER3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (hPER3). | |||||
|
PHF20_MOUSE
|
||||||
| θ value | 6.88961 (rank : 120) | NC score | 0.039322 (rank : 99) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8BLG0, Q8BMA2, Q8BYR4, Q921N1 | Gene names | Phf20, Hca58 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58 homolog). | |||||
|
PRDM9_HUMAN
|
||||||
| θ value | 6.88961 (rank : 121) | NC score | -0.003037 (rank : 187) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 750 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NQV7 | Gene names | PRDM9, PFM6 | |||
|
Domain Architecture |
|
|||||
| Description | PR domain zinc finger protein 9 (PR domain-containing protein 9). | |||||
|
ROCK2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 122) | NC score | 0.004504 (rank : 179) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 2073 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O75116, Q9UQN5 | Gene names | ROCK2, KIAA0619 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 2 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 2) (p164 ROCK-2) (Rho kinase 2). | |||||
|
SEC63_HUMAN
|
||||||
| θ value | 6.88961 (rank : 123) | NC score | 0.028900 (rank : 121) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UGP8, O95380, Q86VS9, Q8IWL0, Q9NTE0 | Gene names | SEC63, SEC63L | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
SMC2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 124) | NC score | 0.012026 (rank : 160) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O95347, Q9P1P2 | Gene names | SMC2L1, CAPE, SMC2 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 2-like 1 protein (Chromosome- associated protein E) (hCAP-E) (XCAP-E homolog). | |||||
|
TOP2A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 125) | NC score | 0.017531 (rank : 148) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P11388, Q71UN1, Q9HB24, Q9HB25, Q9HB26, Q9UP44, Q9UQP9 | Gene names | TOP2A, TOP2 | |||
|
Domain Architecture |

|
|||||
| Description | DNA topoisomerase 2-alpha (EC 5.99.1.3) (DNA topoisomerase II, alpha isozyme). | |||||
|
TXND2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 126) | NC score | 0.058603 (rank : 55) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
UBN1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 127) | NC score | 0.024678 (rank : 129) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9NPG3, Q13079, Q9P1P7 | Gene names | UBN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubinuclein (Ubiquitously expressed nuclear protein) (VT4). | |||||
|
CEBPD_MOUSE
|
||||||
| θ value | 8.99809 (rank : 128) | NC score | 0.007369 (rank : 170) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q00322 | Gene names | Cebpd, Crp3 | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein delta (C/EBP delta) (C/EBP-related protein 3). | |||||
|
CT2NL_MOUSE
|
||||||
| θ value | 8.99809 (rank : 129) | NC score | 0.010853 (rank : 164) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 902 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q99LJ0, Q6ZPR0, Q8BSV1 | Gene names | Cttnbp2nl, Kiaa1433 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CTTNBP2 N-terminal-like protein. | |||||
|
DCNL4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 130) | NC score | 0.008240 (rank : 169) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CCA0 | Gene names | Dcun1d4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DCN1-like protein 4 (Defective in cullin neddylation protein 1-like protein 4) (DCUN1 domain-containing protein 4). | |||||
|
DOT1L_HUMAN
|
||||||
| θ value | 8.99809 (rank : 131) | NC score | 0.019080 (rank : 144) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 660 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8TEK3, O60379, Q96JL1 | Gene names | DOT1L, KIAA1814 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-79 specific (EC 2.1.1.43) (Histone H3-K79 methyltransferase) (H3-K79-HMTase) (DOT1-like protein). | |||||
|
FOXGB_MOUSE
|
||||||
| θ value | 8.99809 (rank : 132) | NC score | 0.010088 (rank : 166) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q60987, Q80VP3 | Gene names | Foxg1b, Fkhl1, Foxg1, Hfhbf1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein G1B (Forkhead-related protein FKHL1) (Transcription factor BF-1) (Brain factor 1) (BF1). | |||||
|
GP73_MOUSE
|
||||||
| θ value | 8.99809 (rank : 133) | NC score | 0.023758 (rank : 131) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q91XA2 | Gene names | Golph2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi phosphoprotein 2 (Golgi membrane protein GP73). | |||||
|
HAIR_HUMAN
|
||||||
| θ value | 8.99809 (rank : 134) | NC score | 0.017870 (rank : 147) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O43593, Q96H33, Q9NPE1 | Gene names | HR | |||
|
Domain Architecture |

|
|||||
| Description | Protein hairless. | |||||
|
HME2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 135) | NC score | 0.002866 (rank : 183) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P09066 | Gene names | En2, En-2 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein engrailed-2 (Mo-En-2). | |||||
|
HXC10_HUMAN
|
||||||
| θ value | 8.99809 (rank : 136) | NC score | 0.005355 (rank : 176) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NYD6, O15219, O15220, Q9BVD5 | Gene names | HOXC10 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-C10. | |||||
|
INOC1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 137) | NC score | 0.004257 (rank : 180) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9ULG1, Q9NTG6 | Gene names | INOC1, KIAA1259 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative DNA helicase INO80 complex homolog 1 (EC 3.6.1.-) (hINO80). | |||||
|
PTN13_MOUSE
|
||||||
| θ value | 8.99809 (rank : 138) | NC score | 0.001288 (rank : 185) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 426 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q64512, Q61494, Q62135, Q64499 | Gene names | Ptpn13, Ptp14 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 13 (EC 3.1.3.48) (Protein tyrosine phosphatase PTP-BL) (Protein-tyrosine phosphatase RIP) (protein tyrosine phosphatase DPZPTP) (PTP36). | |||||
|
RBBP6_HUMAN
|
||||||
| θ value | 8.99809 (rank : 139) | NC score | 0.047599 (rank : 90) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
SDPR_HUMAN
|
||||||
| θ value | 8.99809 (rank : 140) | NC score | 0.008498 (rank : 168) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O95810 | Gene names | SDPR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serum deprivation-response protein (Phosphatidylserine-binding protein) (PS-p68). | |||||
|
SFR12_HUMAN
|
||||||
| θ value | 8.99809 (rank : 141) | NC score | 0.059819 (rank : 48) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 553 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8WXA9, Q86X37 | Gene names | SFRS12, SRRP86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86) (Splicing regulatory protein 508) (SRrp508). | |||||
|
SPT6H_HUMAN
|
||||||
| θ value | 8.99809 (rank : 142) | NC score | 0.021359 (rank : 139) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q7KZ85, Q15737, Q6GMQ4, Q7KYW9, Q7LDK4, Q8N526, Q92775, Q96AH3, Q9BTH9, Q9BTI2 | Gene names | SUPT6H, KIAA0162, SPT6H | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor SPT6 (hSPT6) (Tat-cotransactivator 2 protein) (Tat-CT2 protein). | |||||
|
SPT6H_MOUSE
|
||||||
| θ value | 8.99809 (rank : 143) | NC score | 0.021077 (rank : 141) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q62383, Q5SYM0, Q6GQS3, Q6ZQI0, Q8BQY6 | Gene names | Supt6h, Kiaa0162 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor SPT6. | |||||
|
TM131_MOUSE
|
||||||
| θ value | 8.99809 (rank : 144) | NC score | 0.030697 (rank : 114) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O70472 | Gene names | Tmem131, D1Bwg0491e, Rw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 131 (Protein RW1). | |||||
|
ZCPW1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 145) | NC score | 0.028587 (rank : 122) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6IR42 | Gene names | Zcwpw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CW-type PWWP domain protein 1 homolog. | |||||
|
ZN509_MOUSE
|
||||||
| θ value | 8.99809 (rank : 146) | NC score | -0.001916 (rank : 186) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 990 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8BXX2, Q8CID4, Q8K2Z6 | Gene names | Znf509, Zfp509 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 509. | |||||
|
AFF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 147) | NC score | 0.056157 (rank : 63) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P51825 | Gene names | AFF1, AF4, FEL, MLLT2 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4) (Protein FEL). | |||||
|
BASP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 148) | NC score | 0.075004 (rank : 22) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
BASP_MOUSE
|
||||||
| θ value | θ > 10 (rank : 149) | NC score | 0.068588 (rank : 28) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
CN155_HUMAN
|
||||||
| θ value | θ > 10 (rank : 150) | NC score | 0.050011 (rank : 85) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q5H9T9, Q5H9U7, Q86YI2, Q9H0J3 | Gene names | C14orf155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf155. | |||||
|
CYLC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 151) | NC score | 0.052683 (rank : 77) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P35663, Q5JQQ9 | Gene names | CYLC1, CYL, CYL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-1 (Cylicin I) (Multiple-band polypeptide I). | |||||
|
CYLC2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 152) | NC score | 0.059591 (rank : 49) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q14093 | Gene names | CYLC2, CYL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-2 (Cylicin II) (Multiple-band polypeptide II). | |||||
|
DEK_MOUSE
|
||||||
| θ value | θ > 10 (rank : 153) | NC score | 0.055919 (rank : 64) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q7TNV0, Q80VC5, Q8BZV6 | Gene names | Dek | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DEK. | |||||
|
ENL_HUMAN
|
||||||
| θ value | θ > 10 (rank : 154) | NC score | 0.087949 (rank : 13) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q03111, Q14768 | Gene names | MLLT1, ENL, LTG19, YEATS1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein ENL (YEATS domain-containing protein 1). | |||||
|
GNAS3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 155) | NC score | 0.052304 (rank : 78) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O95467, O95417 | Gene names | GNAS, GNAS1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuroendocrine secretory protein 55 (NESP55) [Contains: LHAL tetrapeptide; GPIPIRRH peptide]. | |||||
|
GPTC8_HUMAN
|
||||||
| θ value | θ > 10 (rank : 156) | NC score | 0.058187 (rank : 56) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 366 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9UKJ3, O60300 | Gene names | GPATC8, KIAA0553 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 8. | |||||
|
HIRP3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 157) | NC score | 0.058640 (rank : 54) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9BW71, O75707, O75708 | Gene names | HIRIP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIRA-interacting protein 3. | |||||
|
LAD1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 158) | NC score | 0.053716 (rank : 70) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 391 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P57016 | Gene names | Lad1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ladinin 1 (Lad-1) (Linear IgA disease autoantigen). | |||||
|
MARCS_HUMAN
|
||||||
| θ value | θ > 10 (rank : 159) | NC score | 0.050418 (rank : 81) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P29966, Q2LA83, Q5TDB7 | Gene names | MARCKS, MACS, PRKCSL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS) (Protein kinase C substrate, 80 kDa protein, light chain) (PKCSL) (80K-L protein). | |||||
|
MARCS_MOUSE
|
||||||
| θ value | θ > 10 (rank : 160) | NC score | 0.051628 (rank : 79) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P26645 | Gene names | Marcks, Macs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS). | |||||
|
MINT_HUMAN
|
||||||
| θ value | θ > 10 (rank : 161) | NC score | 0.052936 (rank : 75) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1291 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q96T58, Q9H9A8, Q9NWH5, Q9UQ01, Q9Y556 | Gene names | SPEN, KIAA0929, MINT, SHARP | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
NEUM_HUMAN
|
||||||
| θ value | θ > 10 (rank : 162) | NC score | 0.053361 (rank : 71) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P17677 | Gene names | GAP43 | |||
|
Domain Architecture |
|
|||||
| Description | Neuromodulin (Axonal membrane protein GAP-43) (Growth-associated protein 43) (PP46) (Neural phosphoprotein B-50). | |||||
|
NOLC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 163) | NC score | 0.069194 (rank : 26) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q14978, Q15030 | Gene names | NOLC1, KIAA0035 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar phosphoprotein p130 (Nucleolar 130 kDa protein) (140 kDa nucleolar phosphoprotein) (Nopp140) (Nucleolar and coiled-body phosphoprotein 1). | |||||
|
NSBP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 164) | NC score | 0.061032 (rank : 46) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P82970 | Gene names | NSBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1. | |||||
|
NUCKS_HUMAN
|
||||||
| θ value | θ > 10 (rank : 165) | NC score | 0.061063 (rank : 45) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9H1E3, Q9H1D6, Q9H723 | Gene names | NUCKS1, NUCKS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (P1). | |||||
|
NUCKS_MOUSE
|
||||||
| θ value | θ > 10 (rank : 166) | NC score | 0.056976 (rank : 60) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q80XU3, Q8BVD8 | Gene names | Nucks1, Nucks | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (JC7). | |||||
|
PTMA_HUMAN
|
||||||
| θ value | θ > 10 (rank : 167) | NC score | 0.055012 (rank : 66) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P06454, Q15249, Q15592 | Gene names | PTMA, TMSA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha-1]. | |||||
|
PTMA_MOUSE
|
||||||
| θ value | θ > 10 (rank : 168) | NC score | 0.055162 (rank : 65) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P26350, Q3UQV6 | Gene names | Ptma | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha]. | |||||
|
RP1L1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 169) | NC score | 0.053722 (rank : 69) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1072 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8IWN7, Q86SQ1, Q8IWN8, Q8IWN9, Q8IWP0, Q8IWP1, Q8IWP2 | Gene names | RP1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinitis pigmentosa 1-like 1 protein. | |||||
|
SFR11_HUMAN
|
||||||
| θ value | θ > 10 (rank : 170) | NC score | 0.068254 (rank : 29) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q05519 | Gene names | SFRS11 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor arginine/serine-rich 11 (Arginine-rich 54 kDa nuclear protein) (p54). | |||||
|
SRRM1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 171) | NC score | 0.058114 (rank : 57) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
SRRM1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 172) | NC score | 0.050955 (rank : 80) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 173) | NC score | 0.059453 (rank : 51) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 174) | NC score | 0.050220 (rank : 83) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
T2FA_HUMAN
|
||||||
| θ value | θ > 10 (rank : 175) | NC score | 0.059556 (rank : 50) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P35269, Q9BWN0 | Gene names | GTF2F1, RAP74 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor IIF alpha subunit (EC 2.7.11.1) (TFIIF-alpha) (Transcription initiation factor RAP74) (General transcription factor IIF polypeptide 1 74 kDa subunit protein). | |||||
|
TCAL2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 176) | NC score | 0.054544 (rank : 67) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9H3H9 | Gene names | TCEAL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 2 (TCEA-like protein 2) (Transcription elongation factor S-II protein-like 2). | |||||
|
TCAL4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 177) | NC score | 0.053107 (rank : 72) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q96EI5, Q8WY12, Q9H2H1, Q9H775 | Gene names | TCEAL4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 4 (TCEA-like protein 4) (Transcription elongation factor S-II protein-like 4). | |||||
|
TCAL5_MOUSE
|
||||||
| θ value | θ > 10 (rank : 178) | NC score | 0.066105 (rank : 34) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8CCT4, Q497R6 | Gene names | Tceal5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 5 (TCEA-like protein 5) (Transcription elongation factor S-II protein-like 5). | |||||
|
TCAL6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 179) | NC score | 0.068142 (rank : 31) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6IPX3, Q5H9J8 | Gene names | TCEAL6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 6 (TCEA-like protein 6) (Transcription elongation factor S-II protein-like 6). | |||||
|
TCOF_HUMAN
|
||||||
| θ value | θ > 10 (rank : 180) | NC score | 0.064414 (rank : 40) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q13428, Q99408, Q99860 | Gene names | TCOF1 | |||
|
Domain Architecture |

|
|||||
| Description | Treacle protein (Treacher Collins syndrome protein). | |||||
|
TGON2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 181) | NC score | 0.050202 (rank : 84) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q62314 | Gene names | Tgoln2, Ttgn2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (TGN38B). | |||||
|
TRDN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 182) | NC score | 0.062581 (rank : 42) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q13061 | Gene names | TRDN | |||
|
Domain Architecture |

|
|||||
| Description | Triadin. | |||||
|
VCX1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 183) | NC score | 0.062581 (rank : 43) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9H320, Q9P0H3 | Gene names | VCX, VCX1, VCX10R, VCXB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Variable charge X-linked protein 1 (VCX-B1 protein) (Variably charged protein X-B1) (Variable charge protein on X with ten repeats) (VCX- 10r). | |||||
|
VCX3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 184) | NC score | 0.065955 (rank : 35) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9NNX9, Q9P0H4 | Gene names | VCX3A, VCX3, VCX8R, VCXA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Variable charge X-linked protein 3 (VCX-A protein) (Variably charged protein X-A) (Variable charge protein on X with eight repeats) (VCX- 8r). | |||||
|
VCXC_HUMAN
|
||||||
| θ value | θ > 10 (rank : 185) | NC score | 0.068219 (rank : 30) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9H321 | Gene names | VCXC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | VCX-C protein (Variably charged protein X-C). | |||||
|
WDR35_HUMAN
|
||||||
| θ value | θ > 10 (rank : 186) | NC score | 0.056808 (rank : 62) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9P2L0, Q8NE11 | Gene names | WDR35, KIAA1336 | |||
|
Domain Architecture |
|
|||||
| Description | WD repeat protein 35. | |||||
|
WDR35_MOUSE
|
||||||
| θ value | θ > 10 (rank : 187) | NC score | 0.056961 (rank : 61) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BND3, Q8CDL4 | Gene names | Wdr35 | |||
|
Domain Architecture |
|
|||||
| Description | WD repeat protein 35. | |||||
|
TULP1_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 146 | |
| SwissProt Accessions | Q9Z273 | Gene names | Tulp1 | |||
|
Domain Architecture |

|
|||||
| Description | Tubby-related protein 1 (Tubby-like protein 1). | |||||
|
TULP1_HUMAN
|
||||||
| NC score | 0.962471 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O00294, O43536 | Gene names | TULP1 | |||
|
Domain Architecture |

|
|||||
| Description | Tubby-related protein 1 (Tubby-like protein 1). | |||||
|
TUB_HUMAN
|
||||||
| NC score | 0.954504 (rank : 3) | θ value | 3.65967e-110 (rank : 3) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P50607, O00293 | Gene names | TUB | |||
|
Domain Architecture |

|
|||||
| Description | Tubby protein homolog. | |||||
|
TUB_MOUSE
|
||||||
| NC score | 0.954057 (rank : 4) | θ value | 4.77967e-110 (rank : 4) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P50586 | Gene names | Tub, Rd5 | |||
|
Domain Architecture |

|
|||||
| Description | Tubby protein. | |||||
|
TULP3_MOUSE
|
||||||
| NC score | 0.948106 (rank : 5) | θ value | 1.48926e-103 (rank : 5) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O88413 | Gene names | Tulp3 | |||
|
Domain Architecture |

|
|||||
| Description | Tubby-related protein 3 (Tubby-like protein 3). | |||||
|
TULP3_HUMAN
|
||||||
| NC score | 0.946875 (rank : 6) | θ value | 6.7005e-96 (rank : 6) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O75386 | Gene names | TULP3 | |||
|
Domain Architecture |

|
|||||
| Description | Tubby-related protein 3 (Tubby-like protein 3). | |||||
|
TULP2_HUMAN
|
||||||
| NC score | 0.933375 (rank : 7) | θ value | 2.02684e-76 (rank : 7) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O00295 | Gene names | TULP2 | |||
|
Domain Architecture |

|
|||||
| Description | Tubby-related protein 2 (Tubby-like protein 2). | |||||
|
TULP4_MOUSE
|
||||||
| NC score | 0.468495 (rank : 8) | θ value | 2.43908e-13 (rank : 8) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9JIL5, Q8CA75, Q922C2 | Gene names | Tulp4, Tusp | |||
|
Domain Architecture |

|
|||||
| Description | Tubby-like protein 4 (Tubby superfamily protein). | |||||
|
TULP4_HUMAN
|
||||||
| NC score | 0.428616 (rank : 9) | θ value | 3.18553e-13 (rank : 9) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 426 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9NRJ4, Q9HD22, Q9P2F0 | Gene names | TULP4, KIAA1397, TUSP | |||
|
Domain Architecture |

|
|||||
| Description | Tubby-like protein 4 (Tubby superfamily protein). | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.094043 (rank : 10) | θ value | 0.00228821 (rank : 12) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
TCAL3_MOUSE
|
||||||
| NC score | 0.090746 (rank : 11) | θ value | 0.365318 (rank : 33) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8R0A5, Q9CYK5, Q9D1S4 | Gene names | Tceal3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 3 (TCEA-like protein 3) (Transcription elongation factor S-II protein-like 3). | |||||
|
VPS72_HUMAN
|
||||||
| NC score | 0.090607 (rank : 12) | θ value | 0.00869519 (rank : 15) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q15906, Q53GJ2, Q5U0R4 | Gene names | VPS72, TCFL1, YL1 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 72 homolog (Transcription factor-like 1) (Protein YL-1). | |||||
|
ENL_HUMAN
|
||||||
| NC score | 0.087949 (rank : 13) | θ value | θ > 10 (rank : 154) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q03111, Q14768 | Gene names | MLLT1, ENL, LTG19, YEATS1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein ENL (YEATS domain-containing protein 1). | |||||
|
VPS72_MOUSE
|
||||||
| NC score | 0.084857 (rank : 14) | θ value | 0.0330416 (rank : 18) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q62481, Q3U2N4, Q810A9, Q99K81 | Gene names | Vps72, Tcfl1, Yl1 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 72 homolog (Transcription factor-like 1) (Protein YL-1). | |||||
|
NFH_MOUSE
|
||||||
| NC score | 0.083632 (rank : 15) | θ value | 5.26297e-08 (rank : 10) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 94 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
TCAL5_HUMAN
|
||||||
| NC score | 0.081914 (rank : 16) | θ value | 0.000786445 (rank : 11) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 806 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q5H9L2 | Gene names | TCEAL5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 5 (TCEA-like protein 5) (Transcription elongation factor S-II protein-like 5). | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.080458 (rank : 17) | θ value | 0.0431538 (rank : 19) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
TCAL3_HUMAN
|
||||||
| NC score | 0.080451 (rank : 18) | θ value | 0.47712 (rank : 36) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q969E4 | Gene names | TCEAL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 3 (TCEA-like protein 3) (Transcription elongation factor S-II protein-like 3). | |||||
|
NPBL_MOUSE
|
||||||
| NC score | 0.076696 (rank : 19) | θ value | 0.813845 (rank : 44) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
NIPBL_HUMAN
|
||||||
| NC score | 0.076402 (rank : 20) | θ value | 0.0736092 (rank : 22) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
TXND2_HUMAN
|
||||||
| NC score | 0.075416 (rank : 21) | θ value | 0.0961366 (rank : 25) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
BASP_HUMAN
|
||||||
| NC score | 0.075004 (rank : 22) | θ value | θ > 10 (rank : 148) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
AFF4_MOUSE
|
||||||
| NC score | 0.071139 (rank : 23) | θ value | 0.813845 (rank : 41) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9ESC8, Q8C6K4, Q8C6W3, Q8CCH3 | Gene names | Aff4, Alf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4. | |||||
|
CJ012_HUMAN
|
||||||
| NC score | 0.070202 (rank : 24) | θ value | 0.62314 (rank : 39) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8N655, Q9H945, Q9Y457 | Gene names | C10orf12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf12. | |||||
|
AFF4_HUMAN
|
||||||
| NC score | 0.070124 (rank : 25) | θ value | 6.88961 (rank : 111) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 611 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q9UHB7, Q498B2, Q59FB3, Q6P592, Q8TDR1, Q9P0E4 | Gene names | AFF4, AF5Q31, MCEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4 (ALL1-fused gene from chromosome 5q31) (Major CDK9 elongation factor-associated protein). | |||||
|
NOLC1_HUMAN
|
||||||
| NC score | 0.069194 (rank : 26) | θ value | θ > 10 (rank : 163) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q14978, Q15030 | Gene names | NOLC1, KIAA0035 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar phosphoprotein p130 (Nucleolar 130 kDa protein) (140 kDa nucleolar phosphoprotein) (Nopp140) (Nucleolar and coiled-body phosphoprotein 1). | |||||
|
TTF1_MOUSE
|
||||||
| NC score | 0.069116 (rank : 27) | θ value | 0.365318 (rank : 34) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q62187, Q9JKK5 | Gene names | Ttf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription termination factor 1 (TTF-1) (TTF-I) (mTFF-I) (RNA polymerase I termination factor). | |||||
|
BASP_MOUSE
|
||||||
| NC score | 0.068588 (rank : 28) | θ value | θ > 10 (rank : 149) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
SFR11_HUMAN
|
||||||
| NC score | 0.068254 (rank : 29) | θ value | θ > 10 (rank : 170) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q05519 | Gene names | SFRS11 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor arginine/serine-rich 11 (Arginine-rich 54 kDa nuclear protein) (p54). | |||||
|
VCXC_HUMAN
|
||||||
| NC score | 0.068219 (rank : 30) | θ value | θ > 10 (rank : 185) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9H321 | Gene names | VCXC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | VCX-C protein (Variably charged protein X-C). | |||||
|
TCAL6_HUMAN
|
||||||
| NC score | 0.068142 (rank : 31) | θ value | θ > 10 (rank : 179) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6IPX3, Q5H9J8 | Gene names | TCEAL6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 6 (TCEA-like protein 6) (Transcription elongation factor S-II protein-like 6). | |||||
|
MINT_MOUSE
|
||||||
| NC score | 0.067980 (rank : 32) | θ value | 0.0148317 (rank : 16) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
CF010_HUMAN
|
||||||
| NC score | 0.066673 (rank : 33) | θ value | 4.03905 (rank : 90) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q5SRN2, Q5SPI9, Q5SPJ0, Q5SPK9, Q5SPL0, Q5SRN3, Q5TG25, Q5TG26, Q8N4B6 | Gene names | C6orf10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C6orf10. | |||||
|
TCAL5_MOUSE
|
||||||
| NC score | 0.066105 (rank : 34) | θ value | θ > 10 (rank : 178) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8CCT4, Q497R6 | Gene names | Tceal5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 5 (TCEA-like protein 5) (Transcription elongation factor S-II protein-like 5). | |||||
|
VCX3_HUMAN
|
||||||
| NC score | 0.065955 (rank : 35) | θ value | θ > 10 (rank : 184) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9NNX9, Q9P0H4 | Gene names | VCX3A, VCX3, VCX8R, VCXA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Variable charge X-linked protein 3 (VCX-A protein) (Variably charged protein X-A) (Variable charge protein on X with eight repeats) (VCX- 8r). | |||||
|
MDC1_MOUSE
|
||||||
| NC score | 0.065487 (rank : 36) | θ value | 0.00869519 (rank : 14) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
DNMT1_HUMAN
|
||||||
| NC score | 0.065422 (rank : 37) | θ value | 6.88961 (rank : 114) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P26358, Q9UHG5, Q9ULA2, Q9UMZ6 | Gene names | DNMT1, AIM, DNMT | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 1 (EC 2.1.1.37) (Dnmt1) (DNA methyltransferase HsaI) (DNA MTase HsaI) (MCMT) (M.HsaI). | |||||
|
RBBP6_MOUSE
|
||||||
| NC score | 0.064690 (rank : 38) | θ value | 1.38821 (rank : 54) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
SFR12_MOUSE
|
||||||
| NC score | 0.064595 (rank : 39) | θ value | 2.36792 (rank : 76) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8BZX4, Q8BLM8, Q8BWK7, Q8BYK0, Q8BYZ4 | Gene names | Sfrs12, Srrp86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86). | |||||
|
TCOF_HUMAN
|
||||||
| NC score | 0.064414 (rank : 40) | θ value | θ > 10 (rank : 180) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q13428, Q99408, Q99860 | Gene names | TCOF1 | |||
|
Domain Architecture |

|
|||||
| Description | Treacle protein (Treacher Collins syndrome protein). | |||||
|
LAD1_HUMAN
|
||||||
| NC score | 0.064354 (rank : 41) | θ value | 2.36792 (rank : 74) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O00515, O95614 | Gene names | LAD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ladinin 1 (Lad-1) (120 kDa linear IgA bullous dermatosis antigen) (97 kDa linear IgA bullous dermatosis antigen) (Linear IgA disease antigen homolog) (LadA). | |||||
|
TRDN_HUMAN
|
||||||
| NC score | 0.062581 (rank : 42) | θ value | θ > 10 (rank : 182) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q13061 | Gene names | TRDN | |||
|
Domain Architecture |

|
|||||
| Description | Triadin. | |||||
|
VCX1_HUMAN
|
||||||
| NC score | 0.062581 (rank : 43) | θ value | θ > 10 (rank : 183) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9H320, Q9P0H3 | Gene names | VCX, VCX1, VCX10R, VCXB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Variable charge X-linked protein 1 (VCX-B1 protein) (Variably charged protein X-B1) (Variable charge protein on X with ten repeats) (VCX- 10r). | |||||
|
LRRF1_MOUSE
|
||||||
| NC score | 0.061242 (rank : 44) | θ value | 4.03905 (rank : 94) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 440 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q3UZ39, O70323, Q3TS94 | Gene names | Lrrfip1, Flap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat flightless-interacting protein 1 (LRR FLII- interacting protein 1) (FLI-LRR-associated protein 1) (Flap-1) (H186 FLAP). | |||||
|
NUCKS_HUMAN
|
||||||
| NC score | 0.061063 (rank : 45) | θ value | θ > 10 (rank : 165) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9H1E3, Q9H1D6, Q9H723 | Gene names | NUCKS1, NUCKS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (P1). | |||||
|
NSBP1_HUMAN
|
||||||
| NC score | 0.061032 (rank : 46) | θ value | θ > 10 (rank : 164) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P82970 | Gene names | NSBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1. | |||||
|
MDC1_HUMAN
|
||||||
| NC score | 0.060811 (rank : 47) | θ value | 0.279714 (rank : 29) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
SFR12_HUMAN
|
||||||
| NC score | 0.059819 (rank : 48) | θ value | 8.99809 (rank : 141) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 553 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8WXA9, Q86X37 | Gene names | SFRS12, SRRP86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86) (Splicing regulatory protein 508) (SRrp508). | |||||
|
CYLC2_HUMAN
|
||||||
| NC score | 0.059591 (rank : 49) | θ value | θ > 10 (rank : 152) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q14093 | Gene names | CYLC2, CYL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-2 (Cylicin II) (Multiple-band polypeptide II). | |||||
|
T2FA_HUMAN
|
||||||
| NC score | 0.059556 (rank : 50) | θ value | θ > 10 (rank : 175) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P35269, Q9BWN0 | Gene names | GTF2F1, RAP74 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor IIF alpha subunit (EC 2.7.11.1) (TFIIF-alpha) (Transcription initiation factor RAP74) (General transcription factor IIF polypeptide 1 74 kDa subunit protein). | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.059453 (rank : 51) | θ value | θ > 10 (rank : 173) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
MBB1A_MOUSE
|
||||||
| NC score | 0.059398 (rank : 52) | θ value | 4.03905 (rank : 95) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q7TPV4, O35851, Q80Y66, Q8R4X2, Q99KP0 | Gene names | Mybbp1a, P160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myb-binding protein 1A (Myb-binding protein of 160 kDa). | |||||
|
DCDC2_MOUSE
|
||||||
| NC score | 0.058718 (rank : 53) | θ value | 0.0330416 (rank : 17) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q5DU00, Q5SZU0, Q80Y99 | Gene names | Dcdc2, Kiaa1154 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Doublecortin domain-containing protein 2. | |||||
|
HIRP3_HUMAN
|
||||||
| NC score | 0.058640 (rank : 54) | θ value | θ > 10 (rank : 157) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9BW71, O75707, O75708 | Gene names | HIRIP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIRA-interacting protein 3. | |||||
|
TXND2_MOUSE
|
||||||
| NC score | 0.058603 (rank : 55) | θ value | 6.88961 (rank : 126) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
GPTC8_HUMAN
|
||||||
| NC score | 0.058187 (rank : 56) | θ value | θ > 10 (rank : 156) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 366 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9UKJ3, O60300 | Gene names | GPATC8, KIAA0553 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 8. | |||||
|
SRRM1_HUMAN
|
||||||
| NC score | 0.058114 (rank : 57) | θ value | θ > 10 (rank : 171) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
TGON2_HUMAN
|
||||||
| NC score | 0.057281 (rank : 58) | θ value | 1.81305 (rank : 64) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
MBB1A_HUMAN
|
||||||
| NC score | 0.057034 (rank : 59) | θ value | 0.62314 (rank : 40) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 665 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9BQG0, Q86VM3, Q9BW49, Q9P0V5, Q9UF99 | Gene names | MYBBP1A, P160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myb-binding protein 1A. | |||||
|
NUCKS_MOUSE
|
||||||
| NC score | 0.056976 (rank : 60) | θ value | θ > 10 (rank : 166) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q80XU3, Q8BVD8 | Gene names | Nucks1, Nucks | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (JC7). | |||||
|
WDR35_MOUSE
|
||||||
| NC score | 0.056961 (rank : 61) | θ value | θ > 10 (rank : 187) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BND3, Q8CDL4 | Gene names | Wdr35 | |||
|
Domain Architecture |

|
|||||
| Description | WD repeat protein 35. | |||||
|
WDR35_HUMAN
|
||||||
| NC score | 0.056808 (rank : 62) | θ value | θ > 10 (rank : 186) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9P2L0, Q8NE11 | Gene names | WDR35, KIAA1336 | |||
|
Domain Architecture |

|
|||||
| Description | WD repeat protein 35. | |||||
|
AFF1_HUMAN
|
||||||
| NC score | 0.056157 (rank : 63) | θ value | θ > 10 (rank : 147) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P51825 | Gene names | AFF1, AF4, FEL, MLLT2 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4) (Protein FEL). | |||||
|
DEK_MOUSE
|
||||||
| NC score | 0.055919 (rank : 64) | θ value | θ > 10 (rank : 153) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q7TNV0, Q80VC5, Q8BZV6 | Gene names | Dek | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DEK. | |||||
|
PTMA_MOUSE
|
||||||
| NC score | 0.055162 (rank : 65) | θ value | θ > 10 (rank : 168) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P26350, Q3UQV6 | Gene names | Ptma | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha]. | |||||
|
PTMA_HUMAN
|
||||||
| NC score | 0.055012 (rank : 66) | θ value | θ > 10 (rank : 167) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P06454, Q15249, Q15592 | Gene names | PTMA, TMSA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha-1]. | |||||
|
TCAL2_HUMAN
|
||||||
| NC score | 0.054544 (rank : 67) | θ value | θ > 10 (rank : 176) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9H3H9 | Gene names | TCEAL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 2 (TCEA-like protein 2) (Transcription elongation factor S-II protein-like 2). | |||||
|
MYST3_HUMAN
|
||||||
| NC score | 0.054484 (rank : 68) | θ value | 0.0736092 (rank : 21) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q92794 | Gene names | MYST3, MOZ, RUNXBP2, ZNF220 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Runt-related transcription factor-binding protein 2) (Monocytic leukemia zinc finger protein) (Zinc finger protein 220). | |||||
|
RP1L1_HUMAN
|
||||||
| NC score | 0.053722 (rank : 69) | θ value | θ > 10 (rank : 169) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1072 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8IWN7, Q86SQ1, Q8IWN8, Q8IWN9, Q8IWP0, Q8IWP1, Q8IWP2 | Gene names | RP1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinitis pigmentosa 1-like 1 protein. | |||||
|
LAD1_MOUSE
|
||||||
| NC score | 0.053716 (rank : 70) | θ value | θ > 10 (rank : 158) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 391 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P57016 | Gene names | Lad1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ladinin 1 (Lad-1) (Linear IgA disease autoantigen). | |||||
|
NEUM_HUMAN
|
||||||
| NC score | 0.053361 (rank : 71) | θ value | θ > 10 (rank : 162) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P17677 | Gene names | GAP43 | |||
|
Domain Architecture |

|
|||||
| Description | Neuromodulin (Axonal membrane protein GAP-43) (Growth-associated protein 43) (PP46) (Neural phosphoprotein B-50). | |||||
|
TCAL4_HUMAN
|
||||||
| NC score | 0.053107 (rank : 72) | θ value | θ > 10 (rank : 177) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q96EI5, Q8WY12, Q9H2H1, Q9H775 | Gene names | TCEAL4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 4 (TCEA-like protein 4) (Transcription elongation factor S-II protein-like 4). | |||||
|
NFH_HUMAN
|
||||||
| NC score | 0.053073 (rank : 73) | θ value | 1.38821 (rank : 53) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
RBBP5_HUMAN
|
||||||
| NC score | 0.053013 (rank : 74) | θ value | 0.0431538 (rank : 20) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 372 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q15291 | Gene names | RBBP5, RBQ3 | |||
|
Domain Architecture |

|
|||||
| Description | Retinoblastoma-binding protein 5 (RBBP-5) (Retinoblastoma-binding protein RBQ-3). | |||||
|
MINT_HUMAN
|
||||||
| NC score | 0.052936 (rank : 75) | θ value | θ > 10 (rank : 161) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1291 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q96T58, Q9H9A8, Q9NWH5, Q9UQ01, Q9Y556 | Gene names | SPEN, KIAA0929, MINT, SHARP | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
NSBP1_MOUSE
|
||||||
| NC score | 0.052879 (rank : 76) | θ value | 5.27518 (rank : 108) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
CYLC1_HUMAN
|
||||||
| NC score | 0.052683 (rank : 77) | θ value | θ > 10 (rank : 151) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P35663, Q5JQQ9 | Gene names | CYLC1, CYL, CYL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-1 (Cylicin I) (Multiple-band polypeptide I). | |||||
|
GNAS3_HUMAN
|
||||||
| NC score | 0.052304 (rank : 78) | θ value | θ > 10 (rank : 155) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O95467, O95417 | Gene names | GNAS, GNAS1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuroendocrine secretory protein 55 (NESP55) [Contains: LHAL tetrapeptide; GPIPIRRH peptide]. | |||||
|
MARCS_MOUSE
|
||||||
| NC score | 0.051628 (rank : 79) | θ value | θ > 10 (rank : 160) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P26645 | Gene names | Marcks, Macs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS). | |||||
|
SRRM1_MOUSE
|
||||||
| NC score | 0.050955 (rank : 80) | θ value | θ > 10 (rank : 172) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
MARCS_HUMAN
|
||||||
| NC score | 0.050418 (rank : 81) | θ value | θ > 10 (rank : 159) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P29966, Q2LA83, Q5TDB7 | Gene names | MARCKS, MACS, PRKCSL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS) (Protein kinase C substrate, 80 kDa protein, light chain) (PKCSL) (80K-L protein). | |||||
|
AHTF1_MOUSE
|
||||||
| NC score | 0.050233 (rank : 82) | θ value | 0.365318 (rank : 30) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8CJF7, Q8BVJ5, Q8VD55 | Gene names | Ahctf1, Elys | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-hook-containing transcription factor 1 (Embryonic large molecule derived from yolk sac). | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.050220 (rank : 83) | θ value | θ > 10 (rank : 174) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
TGON2_MOUSE
|
||||||
| NC score | 0.050202 (rank : 84) | θ value | θ > 10 (rank : 181) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q62314 | Gene names | Tgoln2, Ttgn2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (TGN38B). | |||||
|
CN155_HUMAN
|
||||||
| NC score | 0.050011 (rank : 85) | θ value | θ > 10 (rank : 150) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q5H9T9, Q5H9U7, Q86YI2, Q9H0J3 | Gene names | C14orf155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf155. | |||||
|
GCX1_HUMAN
|
||||||
| NC score | 0.049476 (rank : 86) | θ value | 2.36792 (rank : 73) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96NM4, Q5TE33, Q5TE34, Q5TE35, Q96IC9, Q9BQN5 | Gene names | GCX1, C20orf100 | |||
|
Domain Architecture |

|
|||||
| Description | Granulosa cell HMG box protein 1 (GCX-1). | |||||
|
IF4B_HUMAN
|
||||||
| NC score | 0.049187 (rank : 87) | θ value | 0.47712 (rank : 35) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P23588 | Gene names | EIF4B | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4B (eIF-4B). | |||||
|
FYB_HUMAN
|
||||||
| NC score | 0.049038 (rank : 88) | θ value | 2.36792 (rank : 72) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 379 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O15117, O00359 | Gene names | FYB, SLAP130 | |||
|
Domain Architecture |

|
|||||
| Description | FYN-binding protein (FYN-T-binding protein) (FYB-120/130) (p120/p130) (SLP-76-associated phosphoprotein) (SLAP-130). | |||||
|
SNIP_MOUSE
|
||||||
| NC score | 0.048795 (rank : 89) | θ value | 0.0961366 (rank : 23) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 319 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9QWI6, O70298 | Gene names | P140, Kiaa1684 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | p130Cas-associated protein (p140Cap) (SNAP-25-interacting protein) (SNIP). | |||||
|
RBBP6_HUMAN
|
||||||
| NC score | 0.047599 (rank : 90) | θ value | 8.99809 (rank : 139) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
MPP8_HUMAN
|
||||||
| NC score | 0.046348 (rank : 91) | θ value | 0.00228821 (rank : 13) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 948 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q99549, Q86TK3, Q9BTP1 | Gene names | MPHOSPH8, MPP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | M-phase phosphoprotein 8. | |||||
|
GP179_HUMAN
|
||||||
| NC score | 0.045001 (rank : 92) | θ value | 1.38821 (rank : 52) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 372 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q6PRD1 | Gene names | GPR179, GPR158L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 179 precursor (Probable G-protein coupled receptor 158-like 1). | |||||
|
ELOA1_HUMAN
|
||||||
| NC score | 0.044317 (rank : 93) | θ value | 0.813845 (rank : 42) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q14241, Q8IXH1 | Gene names | TCEB3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription elongation factor B polypeptide 3 (RNA polymerase II transcription factor SIII subunit A1) (SIII p110) (Elongin-A) (EloA) (Elongin 110 kDa subunit). | |||||
|
CCD55_MOUSE
|
||||||
| NC score | 0.044068 (rank : 94) | θ value | 2.36792 (rank : 68) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q5NCR9 | Gene names | Ccdc55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 55. | |||||
|
ADDB_MOUSE
|
||||||
| NC score | 0.042762 (rank : 95) | θ value | 1.81305 (rank : 60) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 574 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9QYB8, Q3U0E1, Q80VH9, Q8C1C4, Q9CXE3, Q9JLE5, Q9QYB7 | Gene names | Add2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-adducin (Erythrocyte adducin subunit beta) (Add97). | |||||
|
PCLO_HUMAN
|
||||||
| NC score | 0.041754 (rank : 96) | θ value | 4.03905 (rank : 96) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
IF4E3_MOUSE
|
||||||
| NC score | 0.039379 (rank : 97) | θ value | 1.06291 (rank : 46) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BMB3, O88503, Q3UCK2 | Gene names | Eif4el3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4E type 3 (eIF4E type 3) (eIF-4E type 3) (mRNA cap-binding protein type 3) (Eukaryotic translation initiation factor 4E-like 3) (eIF4E-like protein 4E-LP). | |||||
|
IF4E3_HUMAN
|
||||||
| NC score | 0.039361 (rank : 98) | θ value | 1.06291 (rank : 45) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O60573, O75349 | Gene names | EIF4EL3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4E type 3 (eIF4E type 3) (eIF-4E type 3) (mRNA cap-binding protein type 3) (Eukaryotic translation initiation factor 4E-like 3) (Eukaryotic translation initiation factor 4E homologous protein) (mRNA cap-binding protein 4EHP) (eIF4E-like protein 4E-LP). | |||||
|
PHF20_MOUSE
|
||||||
| NC score | 0.039322 (rank : 99) | θ value | 6.88961 (rank : 120) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8BLG0, Q8BMA2, Q8BYR4, Q921N1 | Gene names | Phf20, Hca58 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58 homolog). | |||||
|
RAPH1_HUMAN
|
||||||
| NC score | 0.037223 (rank : 100) | θ value | 3.0926 (rank : 87) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q70E73, Q96Q37, Q9C0I2 | Gene names | RAPH1, ALS2CR9, KIAA1681, LPD, PREL2, RMO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras-associated and pleckstrin homology domains-containing protein 1 (RAPH1) (Lamellipodin) (Proline-rich EVH1 ligand 2) (PREL-2) (Protein RMO1) (Amyotrophic lateral sclerosis 2 chromosomal region candidate 9 gene protein). | |||||
|
ARI5B_HUMAN
|
||||||
| NC score | 0.036930 (rank : 101) | θ value | 2.36792 (rank : 67) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q14865, Q7Z3M4, Q8N421, Q9H786 | Gene names | ARID5B, DESRT, MRF2 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 5B (ARID domain- containing protein 5B) (Mrf1-like) (Modulator recognition factor 2) (MRF-2). | |||||
|
NCOR2_MOUSE
|
||||||
| NC score | 0.036356 (rank : 102) | θ value | 6.88961 (rank : 118) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 790 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9WU42, Q9WU43, Q9WUC1 | Gene names | Ncor2, Smrt | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC). | |||||
|
ZAR1_HUMAN
|
||||||
| NC score | 0.035680 (rank : 103) | θ value | 1.81305 (rank : 66) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q86SH2 | Gene names | ZAR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zygote arrest 1 (Oocyte-specific maternal effect factor). | |||||
|
RANG_HUMAN
|
||||||
| NC score | 0.035582 (rank : 104) | θ value | 0.125558 (rank : 26) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P43487 | Gene names | RANBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Ran-specific GTPase-activating protein (Ran-binding protein 1) (RanBP1). | |||||
|
MLL2_HUMAN
|
||||||
| NC score | 0.035290 (rank : 105) | θ value | 6.88961 (rank : 117) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
PRB4S_HUMAN
|
||||||
| NC score | 0.035169 (rank : 106) | θ value | 1.81305 (rank : 62) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P10163, P02813 | Gene names | PRB4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 4 allele S precursor (Salivary proline-rich protein Po) (Parotid o protein) [Contains: Protein N1; Glycosylated protein A]. | |||||
|
CFDP1_HUMAN
|
||||||
| NC score | 0.033618 (rank : 107) | θ value | 3.0926 (rank : 81) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UEE9, O00393, O00404, Q9UEF0, Q9UEF1, Q9UEF8 | Gene names | CFDP1, BCNT | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Craniofacial development protein 1 (Bucentaur). | |||||
|
MADCA_HUMAN
|
||||||
| NC score | 0.033465 (rank : 108) | θ value | 6.88961 (rank : 116) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q13477, O60222, O75867 | Gene names | MADCAM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucosal addressin cell adhesion molecule 1 precursor (MAdCAM-1) (hMAdCAM-1). | |||||
|
MLX_HUMAN
|
||||||
| NC score | 0.033407 (rank : 109) | θ value | 1.06291 (rank : 47) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UH92, Q53XM6, Q96FL2, Q9H2V0, Q9H2V1, Q9H2V2, Q9NXN3 | Gene names | MLX, TCFL4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
ALKB5_HUMAN
|
||||||
| NC score | 0.033289 (rank : 110) | θ value | 0.62314 (rank : 37) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6P6C2, Q9NXD6 | Gene names | ALKBH5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alkylated repair protein alkB homolog 5. | |||||
|
CFDP1_MOUSE
|
||||||
| NC score | 0.032935 (rank : 111) | θ value | 6.88961 (rank : 113) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O88271, O70565, Q9JMA5 | Gene names | Cfdp1, Bcnt, Cfdp, Cp27 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Craniofacial development protein 1 (Bucentaur) (27 kDa craniofacial protein) (Protein Cp27). | |||||
|
NSD1_MOUSE
|
||||||
| NC score | 0.032664 (rank : 112) | θ value | 0.365318 (rank : 32) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O88491, Q8C480, Q9CT70 | Gene names | Nsd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein). | |||||
|
SEC63_MOUSE
|
||||||
| NC score | 0.030818 (rank : 113) | θ value | 1.38821 (rank : 55) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 601 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8VHE0, Q8VEB9 | Gene names | Sec63, Sec63l | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
TM131_MOUSE
|
||||||
| NC score | 0.030697 (rank : 114) | θ value | 8.99809 (rank : 144) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O70472 | Gene names | Tmem131, D1Bwg0491e, Rw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 131 (Protein RW1). | |||||
|
H13_MOUSE
|
||||||
| NC score | 0.030633 (rank : 115) | θ value | 6.88961 (rank : 115) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P43277, Q8C6M4 | Gene names | Hist1h1d, H1f3 | |||
|
Domain Architecture |

|
|||||
| Description | Histone H1.3 (H1 VAR.4) (H1d). | |||||
|
MLX_MOUSE
|
||||||
| NC score | 0.030106 (rank : 116) | θ value | 2.36792 (rank : 75) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O08609, Q9EQJ7, Q9EQJ8 | Gene names | Mlx, Tcfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Max-like protein X (Max-like bHLHZip protein) (BigMax protein) (Protein Mlx) (Transcription factor-like protein 4). | |||||
|
CCNK_MOUSE
|
||||||
| NC score | 0.030029 (rank : 117) | θ value | 5.27518 (rank : 100) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O88874, Q8R068 | Gene names | Ccnk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-K. | |||||
|
SETBP_MOUSE
|
||||||
| NC score | 0.029549 (rank : 118) | θ value | 3.0926 (rank : 88) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Z180, Q66JL8 | Gene names | Setbp1, Kiaa0437 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SET-binding protein (SEB). | |||||
|
K1143_MOUSE
|
||||||
| NC score | 0.029529 (rank : 119) | θ value | 3.0926 (rank : 84) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8K039, Q9CSR4, Q9D0W8 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1143 homolog. | |||||
|
DDX21_MOUSE
|
||||||
| NC score | 0.029511 (rank : 120) | θ value | 2.36792 (rank : 69) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9JIK5, Q9WV45 | Gene names | Ddx21 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar RNA helicase 2 (EC 3.6.1.-) (Nucleolar RNA helicase II) (Nucleolar RNA helicase Gu) (RH II/Gu) (Gu-alpha) (DEAD box protein 21). | |||||
|
SEC63_HUMAN
|
||||||
| NC score | 0.028900 (rank : 121) | θ value | 6.88961 (rank : 123) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UGP8, O95380, Q86VS9, Q8IWL0, Q9NTE0 | Gene names | SEC63, SEC63L | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
ZCPW1_MOUSE
|
||||||
| NC score | 0.028587 (rank : 122) | θ value | 8.99809 (rank : 145) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6IR42 | Gene names | Zcwpw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CW-type PWWP domain protein 1 homolog. | |||||
|
CGRE1_HUMAN
|
||||||
| NC score | 0.028076 (rank : 123) | θ value | 3.0926 (rank : 82) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q99674 | Gene names | CGREF1, CGR11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell growth regulator with EF hand domain 1 (Cell growth regulatory gene 11 protein). | |||||
|
TRIPC_HUMAN
|
||||||
| NC score | 0.027568 (rank : 124) | θ value | 2.36792 (rank : 80) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 319 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q14669, Q15644 | Gene names | TRIP12, KIAA0045 | |||
|
Domain Architecture |

|
|||||
| Description | Thyroid receptor-interacting protein 12 (TRIP12). | |||||
|
HAIR_MOUSE
|
||||||
| NC score | 0.026184 (rank : 125) | θ value | 0.813845 (rank : 43) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q61645, Q80Y47 | Gene names | Hr | |||
|
Domain Architecture |

|
|||||
| Description | Protein hairless. | |||||
|
TXLNA_HUMAN
|
||||||
| NC score | 0.025725 (rank : 126) | θ value | 0.0961366 (rank : 24) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 982 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P40222, Q66K62, Q86T54, Q86T85, Q86T86, Q86Y86, Q86YW3, Q8N2Y3 | Gene names | TXLNA, TXLN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alpha-taxilin. | |||||
|
H12_HUMAN
|
||||||
| NC score | 0.025441 (rank : 127) | θ value | 5.27518 (rank : 102) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P16403 | Gene names | HIST1H1C, H1F2 | |||
|
Domain Architecture |

|
|||||
| Description | Histone H1.2 (Histone H1d). | |||||
|
TACC2_HUMAN
|
||||||
| NC score | 0.024787 (rank : 128) | θ value | 2.36792 (rank : 78) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 856 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O95359, Q9NZ41, Q9NZR5 | Gene names | TACC2 | |||
|
Domain Architecture |

|
|||||
| Description | Transforming acidic coiled-coil-containing protein 2 (Anti Zuai-1) (AZU-1). | |||||
|
UBN1_HUMAN
|
||||||
| NC score | 0.024678 (rank : 129) | θ value | 6.88961 (rank : 127) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9NPG3, Q13079, Q9P1P7 | Gene names | UBN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubinuclein (Ubiquitously expressed nuclear protein) (VT4). | |||||
|
SFPQ_HUMAN
|
||||||
| NC score | 0.024180 (rank : 130) | θ value | 5.27518 (rank : 110) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 986 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P23246, P30808 | Gene names | SFPQ, PSF | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, proline- and glutamine-rich (Polypyrimidine tract- binding protein-associated-splicing factor) (PTB-associated-splicing factor) (PSF) (DNA-binding p52/p100 complex, 100 kDa subunit) (100 kDa DNA-pairing protein) (hPOMp100). | |||||
|
GP73_MOUSE
|
||||||
| NC score | 0.023758 (rank : 131) | θ value | 8.99809 (rank : 133) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q91XA2 | Gene names | Golph2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi phosphoprotein 2 (Golgi membrane protein GP73). | |||||
|
DDX51_HUMAN
|
||||||
| NC score | 0.023588 (rank : 132) | θ value | 0.163984 (rank : 27) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8N8A6, Q5CZ71, Q8IXK5, Q96ED1 | Gene names | DDX51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX51 (EC 3.6.1.-) (DEAD box protein 51). | |||||
|
SMOC2_MOUSE
|
||||||
| NC score | 0.023394 (rank : 133) | θ value | 1.38821 (rank : 56) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CD91, Q8VCM1, Q9D9K2, Q9ER95 | Gene names | Smoc2 | |||
|
Domain Architecture |

|
|||||
| Description | SPARC-related modular calcium-binding protein 2 precursor (Secreted modular calcium-binding protein 2) (SMOC-2). | |||||
|
ANK2_HUMAN
|
||||||
| NC score | 0.022788 (rank : 134) | θ value | 5.27518 (rank : 99) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
TOX_MOUSE
|
||||||
| NC score | 0.022415 (rank : 135) | θ value | 1.81305 (rank : 65) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q66JW3, Q8BKH9, Q8BYQ5, Q8R4H0 | Gene names | Tox | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thymus high mobility group box protein TOX. | |||||
|
SMOC2_HUMAN
|
||||||
| NC score | 0.022318 (rank : 136) | θ value | 2.36792 (rank : 77) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H3U7, Q5TAT7, Q5TAT8, Q86VV9, Q96SF3, Q9H1L3, Q9H1L4, Q9H3U0, Q9H4F7, Q9HCV2 | Gene names | SMOC2, SMAP2 | |||
|
Domain Architecture |

|
|||||
| Description | SPARC-related modular calcium-binding protein 2 precursor (Secreted modular calcium-binding protein 2) (SMOC-2) (Smooth muscle-associated protein 2) (SMAP-2). | |||||
|
MY18B_HUMAN
|
||||||
| NC score | 0.021858 (rank : 137) | θ value | 5.27518 (rank : 107) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1299 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q8IUG5, Q8NDI8, Q8TE65, Q8WWS0, Q96KH2, Q96KR8, Q96KR9 | Gene names | MYO18B | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18B (Myosin XVIIIb). | |||||
|
XYLT1_MOUSE
|
||||||
| NC score | 0.021592 (rank : 138) | θ value | 1.06291 (rank : 50) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q811B1, Q9EPL1 | Gene names | Xylt1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Xylosyltransferase 1 (EC 2.4.2.26) (Xylosyltransferase I) (Peptide O- xylosyltransferase 1). | |||||
|
SPT6H_HUMAN
|
||||||
| NC score | 0.021359 (rank : 139) | θ value | 8.99809 (rank : 142) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q7KZ85, Q15737, Q6GMQ4, Q7KYW9, Q7LDK4, Q8N526, Q92775, Q96AH3, Q9BTH9, Q9BTI2 | Gene names | SUPT6H, KIAA0162, SPT6H | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor SPT6 (hSPT6) (Tat-cotransactivator 2 protein) (Tat-CT2 protein). | |||||
|
AATF_HUMAN
|
||||||
| NC score | 0.021189 (rank : 140) | θ value | 5.27518 (rank : 98) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NY61, Q9P0A4, Q9UNX5 | Gene names | AATF, CHE1, DED | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein AATF (Apoptosis-antagonizing transcription factor) (Rb-binding protein Che-1). | |||||
|
SPT6H_MOUSE
|
||||||
| NC score | 0.021077 (rank : 141) | θ value | 8.99809 (rank : 143) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q62383, Q5SYM0, Q6GQS3, Q6ZQI0, Q8BQY6 | Gene names | Supt6h, Kiaa0162 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor SPT6. | |||||
|
HNRPU_HUMAN
|
||||||
| NC score | 0.020611 (rank : 142) | θ value | 4.03905 (rank : 92) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q00839, O75507, Q96HY9 | Gene names | HNRPU, SAFA, U21.1 | |||
|
Domain Architecture |

|
|||||
| Description | Heterogeneous nuclear ribonucleoprotein U (hnRNP U) (Scaffold attachment factor A) (SAF-A) (p120) (pp120). | |||||
|
CLMN_HUMAN
|
||||||
| NC score | 0.020307 (rank : 143) | θ value | 3.0926 (rank : 83) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q96JQ2, Q9H713, Q9HA23, Q9HA57, Q9UFP4, Q9ULN2 | Gene names | CLMN, KIAA1188 | |||
|
Domain Architecture |

|
|||||
| Description | Calmin. | |||||
|
DOT1L_HUMAN
|
||||||
| NC score | 0.019080 (rank : 144) | θ value | 8.99809 (rank : 131) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 660 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8TEK3, O60379, Q96JL1 | Gene names | DOT1L, KIAA1814 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-79 specific (EC 2.1.1.43) (Histone H3-K79 methyltransferase) (H3-K79-HMTase) (DOT1-like protein). | |||||
|
MY18A_MOUSE
|
||||||
| NC score | 0.018764 (rank : 145) | θ value | 0.163984 (rank : 28) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1989 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9JMH9, Q811D7 | Gene names | Myo18a, Myspdz | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
TBC10_HUMAN
|
||||||
| NC score | 0.018574 (rank : 146) | θ value | 1.06291 (rank : 48) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9BXI6, O76053, Q543A2 | Gene names | TBC1D10A, EPI64, TBC1D10 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 10A (EBP50-PDX interactor of 64 kDa) (EPI64 protein). | |||||
|
HAIR_HUMAN
|
||||||
| NC score | 0.017870 (rank : 147) | θ value | 8.99809 (rank : 134) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O43593, Q96H33, Q9NPE1 | Gene names | HR | |||
|
Domain Architecture |

|
|||||
| Description | Protein hairless. | |||||
|
TOP2A_HUMAN
|
||||||
| NC score | 0.017531 (rank : 148) | θ value | 6.88961 (rank : 125) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P11388, Q71UN1, Q9HB24, Q9HB25, Q9HB26, Q9UP44, Q9UQP9 | Gene names | TOP2A, TOP2 | |||
|
Domain Architecture |

|
|||||
| Description | DNA topoisomerase 2-alpha (EC 5.99.1.3) (DNA topoisomerase II, alpha isozyme). | |||||
|
SUV92_MOUSE
|
||||||
| NC score | 0.017243 (rank : 149) | θ value | 3.0926 (rank : 89) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9EQQ0, Q9CUK3, Q9JLP7 | Gene names | Suv39h2 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-9 specific 2 (EC 2.1.1.43) (Histone H3-K9 methyltransferase 2) (H3-K9-HMTase 2) (Suppressor of variegation 3-9 homolog 2) (Su(var)3-9 homolog 2). | |||||
|
SYTL1_MOUSE
|
||||||
| NC score | 0.016924 (rank : 150) | θ value | 1.38821 (rank : 59) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 314 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q99N80, Q99J26 | Gene names | Sytl1, Slp1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 1 (Exophilin-7). | |||||
|
SASH1_MOUSE
|
||||||
| NC score | 0.015742 (rank : 151) | θ value | 5.27518 (rank : 109) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P59808 | Gene names | Sash1 | |||
|
Domain Architecture |

|
|||||
| Description | SAM and SH3 domain-containing protein 1. | |||||
|
FLNA_MOUSE
|
||||||
| NC score | 0.015521 (rank : 152) | θ value | 2.36792 (rank : 71) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BTM8, O54934, Q7TQI1, Q8BLK1, Q8BTN7, Q8VHX5, Q8VHX8, Q99KQ2 | Gene names | Flna, Fln, Fln1 | |||
|
Domain Architecture |

|
|||||
| Description | Filamin-A (Alpha-filamin) (Filamin-1) (Endothelial actin-binding protein) (Actin-binding protein 280) (ABP-280) (Nonmuscle filamin). | |||||
|
FLNA_HUMAN
|
||||||
| NC score | 0.015392 (rank : 153) | θ value | 2.36792 (rank : 70) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P21333 | Gene names | FLNA, FLN, FLN1 | |||
|
Domain Architecture |

|
|||||
| Description | Filamin-A (Alpha-filamin) (Filamin-1) (Endothelial actin-binding protein) (Actin-binding protein 280) (ABP-280) (Nonmuscle filamin). | |||||
|
MY18A_HUMAN
|
||||||
| NC score | 0.014740 (rank : 154) | θ value | 5.27518 (rank : 106) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1929 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q92614, Q8IXP8 | Gene names | MYO18A, KIAA0216, MYSPDZ | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
WDFY3_HUMAN
|
||||||
| NC score | 0.013274 (rank : 155) | θ value | 1.06291 (rank : 49) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 348 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8IZQ1, Q4W5K5, Q6P0Q5, Q8N1T2, Q96BS7, Q96N85, Q9Y2J7 | Gene names | WDFY3, KIAA0993 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat and FYVE domain-containing protein 3 (Autophagy-linked FYVE protein) (Alfy). | |||||
|
IF2B2_MOUSE
|
||||||
| NC score | 0.013143 (rank : 156) | θ value | 4.03905 (rank : 93) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q5SF07, Q3TCU4, Q7TQF9 | Gene names | Igf2bp2, Imp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Insulin-like growth factor 2 mRNA-binding protein 2 (IGF-II mRNA- binding protein 2) (IMP-2). | |||||
|
TBX2_MOUSE
|
||||||
| NC score | 0.012431 (rank : 157) | θ value | 2.36792 (rank : 79) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q60707 | Gene names | Tbx2 | |||
|
Domain Architecture |

|
|||||
| Description | T-box transcription factor TBX2 (T-box protein 2). | |||||
|
CMTA1_HUMAN
|
||||||
| NC score | 0.012189 (rank : 158) | θ value | 5.27518 (rank : 101) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y6Y1, Q5VUE1 | Gene names | CAMTA1, KIAA0833 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calmodulin-binding transcription activator 1. | |||||
|
SV2B_MOUSE
|
||||||
| NC score | 0.012060 (rank : 159) | θ value | 1.81305 (rank : 63) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BG39, Q80TT1, Q9ES95 | Gene names | Sv2b, Kiaa0735 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptic vesicle glycoprotein 2B (Synaptic vesicle protein 2B). | |||||
|
SMC2_HUMAN
|
||||||
| NC score | 0.012026 (rank : 160) | θ value | 6.88961 (rank : 124) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O95347, Q9P1P2 | Gene names | SMC2L1, CAPE, SMC2 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 2-like 1 protein (Chromosome- associated protein E) (hCAP-E) (XCAP-E homolog). | |||||
|
ASCC2_HUMAN
|
||||||
| NC score | 0.011567 (rank : 161) | θ value | 6.88961 (rank : 112) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9H1I8, Q4TT54, Q8TAZ0, Q9H711, Q9H9D6 | Gene names | ASCC2, ASC1P100 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Activating signal cointegrator 1 complex subunit 2 (ASC-1 complex subunit p100) (Trip4 complex subunit p100). | |||||
|
H6ST3_HUMAN
|
||||||
| NC score | 0.011138 (rank : 162) | θ value | 5.27518 (rank : 103) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IZP7, Q5W0L0, Q68CW6 | Gene names | HS6ST3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heparan-sulfate 6-O-sulfotransferase 3 (EC 2.8.2.-) (HS6ST-3). | |||||
|
PER3_HUMAN
|
||||||
| NC score | 0.011020 (rank : 163) | θ value | 6.88961 (rank : 119) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P56645, Q5H8X4, Q969K6, Q96S77, Q96S78, Q9C0J3, Q9NSP9, Q9UGU8 | Gene names | PER3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (hPER3). | |||||
|
CT2NL_MOUSE
|
||||||
| NC score | 0.010853 (rank : 164) | θ value | 8.99809 (rank : 129) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 902 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q99LJ0, Q6ZPR0, Q8BSV1 | Gene names | Cttnbp2nl, Kiaa1433 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CTTNBP2 N-terminal-like protein. | |||||
|
REEP4_MOUSE
|
||||||
| NC score | 0.010124 (rank : 165) | θ value | 4.03905 (rank : 97) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8K072 | Gene names | Reep4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Receptor expression-enhancing protein 4. | |||||
|
FOXGB_MOUSE
|
||||||
| NC score | 0.010088 (rank : 166) | θ value | 8.99809 (rank : 132) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q60987, Q80VP3 | Gene names | Foxg1b, Fkhl1, Foxg1, Hfhbf1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein G1B (Forkhead-related protein FKHL1) (Transcription factor BF-1) (Brain factor 1) (BF1). | |||||
|
MSX1_HUMAN
|
||||||
| NC score | 0.009283 (rank : 167) | θ value | 0.365318 (rank : 31) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P28360, Q96NY4 | Gene names | MSX1, HOX7 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein MSX-1 (Msh homeobox 1-like protein) (Hox-7). | |||||
|
SDPR_HUMAN
|
||||||
| NC score | 0.008498 (rank : 168) | θ value | 8.99809 (rank : 140) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O95810 | Gene names | SDPR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serum deprivation-response protein (Phosphatidylserine-binding protein) (PS-p68). | |||||
|
DCNL4_MOUSE
|
||||||
| NC score | 0.008240 (rank : 169) | θ value | 8.99809 (rank : 130) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CCA0 | Gene names | Dcun1d4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DCN1-like protein 4 (Defective in cullin neddylation protein 1-like protein 4) (DCUN1 domain-containing protein 4). | |||||
|
CEBPD_MOUSE
|
||||||
| NC score | 0.007369 (rank : 170) | θ value | 8.99809 (rank : 128) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q00322 | Gene names | Cebpd, Crp3 | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein delta (C/EBP delta) (C/EBP-related protein 3). | |||||
|
HS105_MOUSE
|
||||||
| NC score | 0.007354 (rank : 171) | θ value | 5.27518 (rank : 104) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q61699, Q3TNS2, Q3UIY8, Q62578, Q62579, Q6A0A5, Q8C430, Q8VCW6 | Gene names | Hsph1, Hsp105, Hsp110, Kiaa0201 | |||
|
Domain Architecture |

|
|||||
| Description | Heat-shock protein 105 kDa (Heat shock-related 100 kDa protein E7I) (HSP-E7I) (Heat shock 110 kDa protein) (42 degrees C-HSP). | |||||
|
MYLK_MOUSE
|
||||||
| NC score | 0.006983 (rank : 172) | θ value | 1.81305 (rank : 61) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1489 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q6PDN3, Q3TSJ7, Q80UX0, Q80YN7, Q80YN8, Q8K026, Q924D2, Q9ERD3 | Gene names | Mylk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin light chain kinase, smooth muscle (EC 2.7.11.18) (MLCK) (Telokin) (Kinase-related protein) (KRP). | |||||
|
PEG3_MOUSE
|
||||||
| NC score | 0.006595 (rank : 173) | θ value | 3.0926 (rank : 86) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1101 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q3URU2, O54978, Q3TQ69, Q5EBP7, Q61138, Q6GQS0, Q80U47, Q8R5N0, Q9QX53 | Gene names | Peg3, Kiaa0287, Pw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Paternally expressed gene 3 protein (ASF-1). | |||||
|
CTRO_HUMAN
|
||||||
| NC score | 0.006343 (rank : 174) | θ value | 1.38821 (rank : 51) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 2092 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O14578, Q6XUH8, Q86UQ9, Q9UPZ7 | Gene names | CIT, KIAA0949, STK21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Citron Rho-interacting kinase (EC 2.7.11.1) (CRIK) (Rho-interacting, serine/threonine-protein kinase 21). | |||||
|
MINK1_HUMAN
|
||||||
| NC score | 0.005861 (rank : 175) | θ value | 3.0926 (rank : 85) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8N4C8, Q9P1X1, Q9P2R8 | Gene names | MINK1, MAP4K6, MINK | |||
|
Domain Architecture |

|
|||||
| Description | Misshapen-like kinase 1 (EC 2.7.11.1) (Mitogen-activated protein kinase kinase kinase kinase 6) (MAPK/ERK kinase kinase kinase 6) (MEK kinase kinase 6) (MEKKK 6) (Misshapen/NIK-related kinase) (GCK family kinase MiNK). | |||||
|
HXC10_HUMAN
|
||||||
| NC score | 0.005355 (rank : 176) | θ value | 8.99809 (rank : 136) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NYD6, O15219, O15220, Q9BVD5 | Gene names | HOXC10 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-C10. | |||||
|
STK39_MOUSE
|
||||||
| NC score | 0.005032 (rank : 177) | θ value | 1.38821 (rank : 58) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 915 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Z1W9, Q80W13 | Gene names | Stk39, Spak | |||
|
Domain Architecture |

|
|||||
| Description | STE20/SPS1-related proline-alanine-rich protein kinase (EC 2.7.11.1) (Ste-20-related kinase) (Serine/threonine-protein kinase 39). | |||||
|
STK39_HUMAN
|
||||||
| NC score | 0.004804 (rank : 178) | θ value | 1.38821 (rank : 57) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 882 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UEW8, O14774, Q9UER4 | Gene names | STK39, SPAK | |||
|
Domain Architecture |

|
|||||
| Description | STE20/SPS1-related proline-alanine-rich protein kinase (EC 2.7.11.1) (Ste-20-related kinase) (Serine/threonine-protein kinase 39) (DCHT). | |||||
|
ROCK2_HUMAN
|
||||||
| NC score | 0.004504 (rank : 179) | θ value | 6.88961 (rank : 122) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 2073 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O75116, Q9UQN5 | Gene names | ROCK2, KIAA0619 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 2 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 2) (p164 ROCK-2) (Rho kinase 2). | |||||
|
INOC1_HUMAN
|
||||||
| NC score | 0.004257 (rank : 180) | θ value | 8.99809 (rank : 137) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9ULG1, Q9NTG6 | Gene names | INOC1, KIAA1259 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative DNA helicase INO80 complex homolog 1 (EC 3.6.1.-) (hINO80). | |||||
|
CDKL2_MOUSE
|
||||||
| NC score | 0.003522 (rank : 181) | θ value | 0.62314 (rank : 38) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 854 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9QUK0, Q3UL60, Q3V3X7, Q9QYI1, Q9QYI2 | Gene names | Cdkl2, Kkiamre, Kkm | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-dependent kinase-like 2 (EC 2.7.11.22) (Serine/threonine- protein kinase KKIAMRE). | |||||
|
DUET_HUMAN
|
||||||
| NC score | 0.002947 (rank : 182) | θ value | 4.03905 (rank : 91) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1220 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Y2A5, Q8TBQ5, Q9NSZ4 | Gene names | DUET, TRAD | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Duet (EC 2.7.11.1) (Serine/threonine kinase with Dbl- and pleckstrin homology domain). | |||||
|
HME2_MOUSE
|
||||||
| NC score | 0.002866 (rank : 183) | θ value | 8.99809 (rank : 135) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P09066 | Gene names | En2, En-2 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein engrailed-2 (Mo-En-2). | |||||
|
MLTK_HUMAN
|
||||||
| NC score | 0.001417 (rank : 184) | θ value | 5.27518 (rank : 105) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 911 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9NYL2, Q53SX1, Q580W8, Q59GY5, Q86YW8, Q9HCC4, Q9HCC5, Q9HDD2, Q9NYE9 | Gene names | MLTK, ZAK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase MLT (EC 2.7.11.25) (MLK-like mitogen-activated protein triple kinase) (Leucine zipper- and sterile alpha motif-containing kinase) (Sterile alpha motif- and leucine zipper-containing kinase AZK) (Mixed lineage kinase-related kinase) (MLK-related kinase) (MRK) (Cervical cancer suppressor gene 4 protein). | |||||
|
PTN13_MOUSE
|
||||||
| NC score | 0.001288 (rank : 185) | θ value | 8.99809 (rank : 138) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 426 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q64512, Q61494, Q62135, Q64499 | Gene names | Ptpn13, Ptp14 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 13 (EC 3.1.3.48) (Protein tyrosine phosphatase PTP-BL) (Protein-tyrosine phosphatase RIP) (protein tyrosine phosphatase DPZPTP) (PTP36). | |||||
|
ZN509_MOUSE
|
||||||
| NC score | -0.001916 (rank : 186) | θ value | 8.99809 (rank : 146) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 990 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8BXX2, Q8CID4, Q8K2Z6 | Gene names | Znf509, Zfp509 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 509. | |||||
|
PRDM9_HUMAN
|
||||||
| NC score | -0.003037 (rank : 187) | θ value | 6.88961 (rank : 121) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 750 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NQV7 | Gene names | PRDM9, PFM6 | |||
|
Domain Architecture |

|
|||||
| Description | PR domain zinc finger protein 9 (PR domain-containing protein 9). | |||||