Please be patient as the page loads
|
AHTF1_MOUSE
|
||||||
| SwissProt Accessions | Q8CJF7, Q8BVJ5, Q8VD55 | Gene names | Ahctf1, Elys | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-hook-containing transcription factor 1 (Embryonic large molecule derived from yolk sac). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
AHTF1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.911494 (rank : 2) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8WYP5, Q7Z4E3, Q8IZA4, Q96EH9, Q9Y4Q6 | Gene names | AHCTF1, ELYS, TMBS62 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-hook-containing transcription factor 1 (Embryonic large molecule derived from yolk sac). | |||||
|
AHTF1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 176 | |
| SwissProt Accessions | Q8CJF7, Q8BVJ5, Q8VD55 | Gene names | Ahctf1, Elys | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-hook-containing transcription factor 1 (Embryonic large molecule derived from yolk sac). | |||||
|
PRG4_MOUSE
|
||||||
| θ value | 0.00175202 (rank : 3) | NC score | 0.112472 (rank : 6) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9JM99, Q3UEL1, Q3V198 | Gene names | Prg4, Msf, Szp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
SNPC4_HUMAN
|
||||||
| θ value | 0.00175202 (rank : 4) | NC score | 0.088073 (rank : 17) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q5SXM2, Q9Y6P7 | Gene names | SNAPC4, SNAP190 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | snRNA-activating protein complex subunit 4 (SNAPc subunit 4) (snRNA- activating protein complex 190 kDa subunit) (SNAPc 190 kDa subunit) (Proximal sequence element-binding transcription factor subunit alpha) (PSE-binding factor subunit alpha) (PTF subunit alpha). | |||||
|
MAP4_HUMAN
|
||||||
| θ value | 0.00228821 (rank : 5) | NC score | 0.099435 (rank : 10) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 428 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P27816, Q13082, Q96A76 | Gene names | MAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 4 (MAP 4). | |||||
|
PCLO_MOUSE
|
||||||
| θ value | 0.00509761 (rank : 6) | NC score | 0.087093 (rank : 19) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 92 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
NIPBL_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 7) | NC score | 0.087708 (rank : 18) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
M3K9_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 8) | NC score | 0.012799 (rank : 172) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1123 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P80192, Q6EH31, Q9H2N5 | Gene names | MAP3K9, MLK1, PRKE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase 9 (EC 2.7.11.25) (Mixed lineage kinase 1). | |||||
|
MDC1_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 9) | NC score | 0.097777 (rank : 11) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
UBP31_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 10) | NC score | 0.050840 (rank : 63) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 293 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q70CQ4, Q6AW97, Q6ZTC0, Q6ZTN2, Q9ULL7 | Gene names | USP31, KIAA1203 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 31 (EC 3.1.2.15) (Ubiquitin thioesterase 31) (Ubiquitin-specific-processing protease 31) (Deubiquitinating enzyme 31). | |||||
|
MAGD2_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 11) | NC score | 0.032586 (rank : 100) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UNF1, O76058, Q5BJF3, Q8NAL6, Q9H218, Q9P0U9, Q9UM52 | Gene names | MAGED2, BCG1 | |||
|
Domain Architecture |

|
|||||
| Description | Melanoma-associated antigen D2 (MAGE-D2 antigen) (MAGE-D) (Breast cancer-associated gene 1 protein) (BCG-1) (11B6) (Hepatocellular carcinoma-associated protein JCL-1). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 12) | NC score | 0.088609 (rank : 16) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 87 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
KI67_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 13) | NC score | 0.117883 (rank : 5) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P46013 | Gene names | MKI67 | |||
|
Domain Architecture |

|
|||||
| Description | Antigen KI-67. | |||||
|
LPXN_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 14) | NC score | 0.025909 (rank : 126) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99N69 | Gene names | Lpxn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leupaxin. | |||||
|
PCLO_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 15) | NC score | 0.097305 (rank : 12) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 87 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
SOCS6_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 16) | NC score | 0.043350 (rank : 78) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9JLY0, Q9D2A0 | Gene names | Socs6, Cis4, Cish4, Socs4 | |||
|
Domain Architecture |

|
|||||
| Description | Suppressor of cytokine signaling 6 (SOCS-6) (Suppressor of cytokine signaling 4) (SOCS-4) (Cytokine-inducible SH2 protein 4) (CIS-4). | |||||
|
RTEL1_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 17) | NC score | 0.050543 (rank : 67) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NZ71, Q5JTV4, Q9BW37, Q9H402, Q9H4X6, Q9NX25, Q9NZ73, Q9UPR4, Q9Y4R6 | Gene names | RTEL1, C20orf41, KIAA1088, NHL | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of telomere elongation helicase 1 (EC 3.6.1.-) (Helicase- like protein NHL). | |||||
|
SETX_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 18) | NC score | 0.059424 (rank : 41) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q7Z333, O75120, Q3KQX4, Q68DW5, Q6AZD7, Q7Z3J6, Q8WX33, Q9H9D1, Q9NVP9 | Gene names | SETX, ALS4, KIAA0625, SCAR1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable helicase senataxin (EC 3.6.1.-) (SEN1 homolog). | |||||
|
FHOD1_MOUSE
|
||||||
| θ value | 0.125558 (rank : 19) | NC score | 0.063678 (rank : 34) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6P9Q4, Q8BMK2 | Gene names | Fhod1, Fhos1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FH1/FH2 domain-containing protein (Formin homolog overexpressed in spleen) (FHOS) (Formin homology 2 domain-containing protein 1). | |||||
|
PAF49_HUMAN
|
||||||
| θ value | 0.163984 (rank : 20) | NC score | 0.090714 (rank : 15) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O15446, Q32N11, Q7Z5U2, Q9UPF6 | Gene names | CD3EAP, ASE1, CAST, PAF49 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase I-associated factor PAF49 (Anti-sense to ERCC-1 protein) (ASE-1) (CD3-epsilon-associated protein) (CD3E-associated protein) (CAST). | |||||
|
PHF20_MOUSE
|
||||||
| θ value | 0.163984 (rank : 21) | NC score | 0.082915 (rank : 20) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8BLG0, Q8BMA2, Q8BYR4, Q921N1 | Gene names | Phf20, Hca58 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58 homolog). | |||||
|
SALL2_MOUSE
|
||||||
| θ value | 0.163984 (rank : 22) | NC score | 0.014458 (rank : 166) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 836 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9QX96 | Gene names | Sall2, Sal2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sal-like protein 2 (Spalt-like protein 2) (MSal-2). | |||||
|
JIP3_HUMAN
|
||||||
| θ value | 0.21417 (rank : 23) | NC score | 0.041299 (rank : 83) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1039 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9UPT6, Q96RY4, Q9H4I4, Q9H7P1, Q9NUG0 | Gene names | MAPK8IP3, JIP3, KIAA1066 | |||
|
Domain Architecture |

|
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 3 (JNK-interacting protein 3) (JIP-3) (JNK MAP kinase scaffold protein 3) (Mitogen- activated protein kinase 8-interacting protein 3). | |||||
|
SFR16_MOUSE
|
||||||
| θ value | 0.21417 (rank : 24) | NC score | 0.124176 (rank : 3) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8CFC7, Q8CFC8, Q9Z2N3 | Gene names | Sfrs16, Clasp, Swap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 16 (Suppressor of white-apricot homolog 2) (Clk4-associating SR-related protein). | |||||
|
CKAP2_HUMAN
|
||||||
| θ value | 0.279714 (rank : 25) | NC score | 0.060787 (rank : 39) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8WWK9, Q3KRA5, Q5VXB4, Q8IWV5, Q8IWV6, Q96FH9, Q9H012, Q9H0D0, Q9H988, Q9HC49, Q9NVG4 | Gene names | CKAP2, LB1, TMAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-associated protein 2 (Tumor-associated microtubule- associated protein) (CTCL tumor antigen se20-10). | |||||
|
FHOD1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 26) | NC score | 0.063196 (rank : 35) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9Y613, Q59F76, Q6Y1F2, Q76MS8, Q8N521 | Gene names | FHOD1, FHOS | |||
|
Domain Architecture |

|
|||||
| Description | FH1/FH2 domain-containing protein (Formin homolog overexpressed in spleen) (FHOS) (Formin homology 2 domain-containing protein 1). | |||||
|
NFH_HUMAN
|
||||||
| θ value | 0.279714 (rank : 27) | NC score | 0.049774 (rank : 71) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
PHAR2_HUMAN
|
||||||
| θ value | 0.279714 (rank : 28) | NC score | 0.063903 (rank : 33) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O75167, Q68DM2 | Gene names | PHACTR2, C6orf56, KIAA0680 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatase and actin regulator 2. | |||||
|
TULP1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 29) | NC score | 0.051360 (rank : 60) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O00294, O43536 | Gene names | TULP1 | |||
|
Domain Architecture |

|
|||||
| Description | Tubby-related protein 1 (Tubby-like protein 1). | |||||
|
SFR16_HUMAN
|
||||||
| θ value | 0.365318 (rank : 30) | NC score | 0.117947 (rank : 4) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8N2M8, O96026, Q6UW71, Q96DX2 | Gene names | SFRS16, SWAP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 16 (Suppressor of white-apricot homolog 2). | |||||
|
SPEG_MOUSE
|
||||||
| θ value | 0.365318 (rank : 31) | NC score | 0.021089 (rank : 137) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | 0.365318 (rank : 32) | NC score | 0.102197 (rank : 8) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
TULP1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 33) | NC score | 0.050233 (rank : 70) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Z273 | Gene names | Tulp1 | |||
|
Domain Architecture |

|
|||||
| Description | Tubby-related protein 1 (Tubby-like protein 1). | |||||
|
APXL_HUMAN
|
||||||
| θ value | 0.47712 (rank : 34) | NC score | 0.058326 (rank : 44) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 553 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q13796 | Gene names | APXL | |||
|
Domain Architecture |
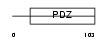
|
|||||
| Description | Apical-like protein (Protein APXL). | |||||
|
RBBP6_HUMAN
|
||||||
| θ value | 0.47712 (rank : 35) | NC score | 0.057343 (rank : 51) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
CHSS1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 36) | NC score | 0.032994 (rank : 98) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q86X52, Q6UX38, Q7LFU5, Q9Y2J5 | Gene names | CHSY1, CHSY, CSS1, KIAA0990 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate synthase 1 (EC 2.4.1.175) (Glucuronosyl-N- acetylgalactosaminyl-proteoglycan 4-beta-N- acetylgalactosaminyltransferase 1) (Chondroitin sulfate synthase 1) (N-acetylgalactosaminyl-proteoglycan 3-beta-glucuronosyltransferase 1) (EC 2.4.1.226) (Chondroitin glucuronyltransferase II) (N- acetylgalactosaminyltransferase II). | |||||
|
GCC2_HUMAN
|
||||||
| θ value | 0.62314 (rank : 37) | NC score | 0.020952 (rank : 139) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1558 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8IWJ2, O15045, Q8TDH3, Q9H2G8 | Gene names | GCC2, KIAA0336 | |||
|
Domain Architecture |

|
|||||
| Description | GRIP and coiled-coil domain-containing protein 2 (Golgi coiled coil protein GCC185) (CTCL tumor antigen se1-1) (CLL-associated antigen KW- 11) (NY-REN-53 antigen). | |||||
|
INCE_MOUSE
|
||||||
| θ value | 0.62314 (rank : 38) | NC score | 0.036825 (rank : 88) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9WU62, Q7TN28, Q8BGN4, Q8CGI4 | Gene names | Incenp | |||
|
Domain Architecture |

|
|||||
| Description | Inner centromere protein. | |||||
|
MADCA_HUMAN
|
||||||
| θ value | 0.62314 (rank : 39) | NC score | 0.065955 (rank : 29) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q13477, O60222, O75867 | Gene names | MADCAM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucosal addressin cell adhesion molecule 1 precursor (MAdCAM-1) (hMAdCAM-1). | |||||
|
RBP24_HUMAN
|
||||||
| θ value | 0.62314 (rank : 40) | NC score | 0.020843 (rank : 140) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q4ZG46 | Gene names | RANBP2L4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 2-like 4 (RanBP2L4). | |||||
|
RGS3_MOUSE
|
||||||
| θ value | 0.62314 (rank : 41) | NC score | 0.047689 (rank : 72) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
SRRM1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 42) | NC score | 0.101841 (rank : 9) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
STB5L_HUMAN
|
||||||
| θ value | 0.62314 (rank : 43) | NC score | 0.052832 (rank : 57) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y2K9, Q4G1B4, Q6PIC3 | Gene names | STXBP5L, KIAA1006, LLGL4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntaxin-binding protein 5-like (Tomosyn-2) (Lethal(2) giant larvae protein homolog 4). | |||||
|
TMC1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 44) | NC score | 0.028802 (rank : 116) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8TDI8 | Gene names | TMC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane cochlear-expressed protein 1. | |||||
|
AL2S4_MOUSE
|
||||||
| θ value | 0.813845 (rank : 45) | NC score | 0.063049 (rank : 36) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q3V0J1, Q3TIS2 | Gene names | Als2cr4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyotrophic lateral sclerosis 2 chromosomal region candidate gene 4 protein homolog. | |||||
|
CLAP1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 46) | NC score | 0.031677 (rank : 106) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q7Z460, O75118, Q8N5B8, Q9BQT5 | Gene names | CLASP1, KIAA0622, MAST1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CLIP-associating protein 1 (Cytoplasmic linker-associated protein 1) (Multiple asters homolog 1). | |||||
|
K0460_HUMAN
|
||||||
| θ value | 0.813845 (rank : 47) | NC score | 0.043419 (rank : 77) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q5VT52, O75048, Q5VT53, Q6MZL4, Q86XD2 | Gene names | KIAA0460 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0460. | |||||
|
LAP2B_MOUSE
|
||||||
| θ value | 0.813845 (rank : 48) | NC score | 0.034482 (rank : 94) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q61029, Q61030, Q61031, Q61032 | Gene names | Tmpo, Lap2 | |||
|
Domain Architecture |

|
|||||
| Description | Lamina-associated polypeptide 2 isoforms beta/delta/epsilon/gamma (Thymopoietin isoforms beta/delta/epsilon/gamma) (TP beta/delta/epsilon/gamma). | |||||
|
RON_MOUSE
|
||||||
| θ value | 0.813845 (rank : 49) | NC score | 0.007679 (rank : 184) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 836 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q62190, Q62555 | Gene names | Mst1r, Ron, Stk | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage-stimulating protein receptor precursor (EC 2.7.10.1) (MSP receptor) (p185-Ron) (Stem cell-derived tyrosine kinase) (CDw136 antigen) [Contains: Macrophage-stimulating protein receptor alpha chain; Macrophage-stimulating protein receptor beta chain]. | |||||
|
STB5L_MOUSE
|
||||||
| θ value | 0.813845 (rank : 50) | NC score | 0.051274 (rank : 61) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q5DQR4, Q3TSM3, Q3UVG2, Q5DQR1, Q5DQR2, Q5DQR3, Q8BS52 | Gene names | Stxbp5l, Llgl4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntaxin-binding protein 5-like (Tomosyn-2) (Lethal(2) giant larvae protein homolog 4). | |||||
|
ALMS1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 51) | NC score | 0.047177 (rank : 73) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8TCU4, Q53S05, Q580Q8, Q86VP9, Q9Y4G4 | Gene names | ALMS1, KIAA0328 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alstrom syndrome protein 1. | |||||
|
GRIN1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 52) | NC score | 0.064109 (rank : 32) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q7Z2K8, Q8ND74, Q96PZ4 | Gene names | GPRIN1, KIAA1893 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
NFH_MOUSE
|
||||||
| θ value | 1.06291 (rank : 53) | NC score | 0.060153 (rank : 40) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 97 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
NPBL_MOUSE
|
||||||
| θ value | 1.06291 (rank : 54) | NC score | 0.077439 (rank : 21) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
PCQAP_MOUSE
|
||||||
| θ value | 1.06291 (rank : 55) | NC score | 0.062684 (rank : 37) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q924H2 | Gene names | Pcqap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Positive cofactor 2 glutamine/Q-rich-associated protein (PC2 glutamine/Q-rich-associated protein) (mPcqap). | |||||
|
TARA_HUMAN
|
||||||
| θ value | 1.06291 (rank : 56) | NC score | 0.065567 (rank : 30) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | Q9H2D6, O94797, Q2PZW8, Q2Q3Z9, Q2Q400, Q5R3M6, Q96DW1, Q9BT77, Q9BTL7, Q9BY98, Q9Y3L4 | Gene names | TRIOBP, KIAA1662, TARA | |||
|
Domain Architecture |

|
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
CD4_HUMAN
|
||||||
| θ value | 1.38821 (rank : 57) | NC score | 0.034287 (rank : 95) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P01730, Q4ZGK2, Q5U066 | Gene names | CD4 | |||
|
Domain Architecture |
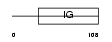
|
|||||
| Description | T-cell surface glycoprotein CD4 precursor (T-cell surface antigen T4/Leu-3). | |||||
|
FOG1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 58) | NC score | 0.016772 (rank : 154) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 981 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8IX07 | Gene names | ZFPM1, FOG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein ZFPM1 (Zinc finger protein multitype 1) (Friend of GATA protein 1) (Friend of GATA-1) (FOG-1). | |||||
|
MACF4_HUMAN
|
||||||
| θ value | 1.38821 (rank : 59) | NC score | 0.014968 (rank : 163) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1321 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q96PK2, Q8WXY1 | Gene names | MACF1, ABP620, ACF7, KIAA0465, KIAA1251 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1, isoform 4. | |||||
|
PCQAP_HUMAN
|
||||||
| θ value | 1.38821 (rank : 60) | NC score | 0.058700 (rank : 42) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 549 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q96RN5, O15413, Q6IC31, Q8NF16, Q96CT0, Q96IH7, Q9P1T3 | Gene names | PCQAP, ARC105, CTG7A, TIG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Positive cofactor 2 glutamine/Q-rich-associated protein (PC2 glutamine/Q-rich-associated protein) (TPA-inducible gene 1 protein) (TIG-1) (Activator-recruited cofactor 105 kDa component) (ARC105) (CTG repeat protein 7a). | |||||
|
PRG4_HUMAN
|
||||||
| θ value | 1.38821 (rank : 61) | NC score | 0.094553 (rank : 13) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
SAMN1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 62) | NC score | 0.028318 (rank : 118) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P57725, Q8C6S2 | Gene names | Samsn1 | |||
|
Domain Architecture |

|
|||||
| Description | SAM domain-containing protein SAMSN-1 (SAM domain, SH3 domain and nuclear localization signals protein 1). | |||||
|
SPEG_HUMAN
|
||||||
| θ value | 1.38821 (rank : 63) | NC score | 0.017923 (rank : 149) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q15772, Q27J74, Q695L1, Q6FGA6, Q6ZQW1, Q6ZTL8, Q9P2P9 | Gene names | SPEG, APEG1, KIAA1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle preferentially expressed protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
WASIP_HUMAN
|
||||||
| θ value | 1.38821 (rank : 64) | NC score | 0.058631 (rank : 43) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O43516, Q15220, Q6MZU9, Q9BU37, Q9UNP1 | Gene names | WASPIP, WIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein) (PRPL-2 protein). | |||||
|
ZEP1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 65) | NC score | 0.018311 (rank : 148) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1063 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q03172 | Gene names | Hivep1, Cryabp1, Znf40 | |||
|
Domain Architecture |
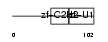
|
|||||
| Description | Zinc finger protein 40 (Transcription factor alphaA-CRYBP1) (Alpha A- crystallin-binding protein I) (Alpha A-CRYBP1). | |||||
|
CALD1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 66) | NC score | 0.031994 (rank : 104) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 959 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q05682, Q13978, Q13979, Q14741, Q14742 | Gene names | CALD1, CAD, CDM | |||
|
Domain Architecture |

|
|||||
| Description | Caldesmon (CDM). | |||||
|
KTNB1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 67) | NC score | 0.016483 (rank : 158) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8BG40, Q8CD18, Q8R1J0, Q9CWV2 | Gene names | Katnb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Katanin p80 WD40-containing subunit B1 (Katanin p80 subunit B1) (p80 katanin). | |||||
|
MELPH_MOUSE
|
||||||
| θ value | 1.81305 (rank : 68) | NC score | 0.029166 (rank : 114) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q91V27, Q99N53 | Gene names | Mlph, Ln, Slac2a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanophilin (Exophilin-3) (Leaden protein) (Synaptotagmin-like protein 2a) (Slac2-a) (Slp homolog lacking C2 domains a). | |||||
|
NCOA7_HUMAN
|
||||||
| θ value | 1.81305 (rank : 69) | NC score | 0.026775 (rank : 123) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8NI08, Q3LID6, Q4G0V1, Q5TF95, Q6IPQ4, Q6NE83, Q86T89, Q8N1W4 | Gene names | NCOA7, ERAP140, ESNA1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 7 (140 kDa estrogen receptor-associated protein) (Estrogen nuclear receptor coactivator 1). | |||||
|
RERE_MOUSE
|
||||||
| θ value | 1.81305 (rank : 70) | NC score | 0.043160 (rank : 79) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q80TZ9 | Gene names | Rere, Atr2, Kiaa0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-2). | |||||
|
RIMS2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 71) | NC score | 0.035226 (rank : 91) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 376 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9EQZ7, Q8C433, Q8CCK2 | Gene names | Rims2, Rab3ip2, Rim2 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 2 (Rab3-interacting molecule 2) (RIM 2) (Rab3-interacting protein 2). | |||||
|
SCYBG_MOUSE
|
||||||
| θ value | 1.81305 (rank : 72) | NC score | 0.043755 (rank : 75) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BSU2, Q8VE25, Q9EPB3 | Gene names | Cxcl16, Srpsox | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Small inducible cytokine B16 precursor (Transmembrane chemokine CXCL16) (SR-PSOX) (Scavenger receptor for phosphatidylserine and oxidized low density lipoprotein). | |||||
|
WASIP_MOUSE
|
||||||
| θ value | 1.81305 (rank : 73) | NC score | 0.057827 (rank : 48) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8K1I7, Q3U0U8 | Gene names | Waspip, Wip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein). | |||||
|
WDR7_HUMAN
|
||||||
| θ value | 1.81305 (rank : 74) | NC score | 0.032164 (rank : 103) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 244 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Y4E6, Q86UX5, Q86VP2, Q96PS7 | Gene names | WDR7, KIAA0541, TRAG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 7 (TGF-beta resistance-associated protein TRAG) (Rabconnectin-3 beta). | |||||
|
CHSS1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 75) | NC score | 0.028214 (rank : 120) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6ZQ11 | Gene names | Chsy1, Kiaa0990 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate synthase 1 (EC 2.4.1.175) (Glucuronosyl-N- acetylgalactosaminyl-proteoglycan 4-beta-N- acetylgalactosaminyltransferase) (Chondroitin sulfate synthase 1) (N- acetylgalactosaminyl-proteoglycan 3-beta-glucuronosyltransferase) (EC 2.4.1.226) (Chondroitin glucuronyltransferase II) (N- acetylgalactosaminyltransferase II). | |||||
|
GNPTA_MOUSE
|
||||||
| θ value | 2.36792 (rank : 76) | NC score | 0.024131 (rank : 130) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q69ZN6, Q3U3K6, Q3US34 | Gene names | Gnptab, Gnpta, Kiaa1208 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylglucosamine-1-phosphotransferase subunits alpha/beta precursor (EC 2.7.8.17) (GlcNAc-1-phosphotransferase alpha/beta subunits) (UDP- N-acetylglucosamine-1-phosphotransferase alpha/beta subunits) (Stealth protein GNPTAB) [Contains: N-acetylglucosamine-1-phosphotransferase subunit alpha; N-acetylglucosamine-1-phosphotransferase subunit beta]. | |||||
|
MUC2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 77) | NC score | 0.038053 (rank : 87) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q02817, Q14878 | Gene names | MUC2, SMUC | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-2 precursor (Intestinal mucin-2). | |||||
|
OC90_HUMAN
|
||||||
| θ value | 2.36792 (rank : 78) | NC score | 0.023138 (rank : 132) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q02509 | Gene names | OC90, PLA2L | |||
|
Domain Architecture |

|
|||||
| Description | Otoconin 90 precursor (Oc90) (Phospholipase A2 homolog). | |||||
|
PP2CG_HUMAN
|
||||||
| θ value | 2.36792 (rank : 79) | NC score | 0.022019 (rank : 135) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O15355 | Gene names | PPM1G, PPM1C | |||
|
Domain Architecture |

|
|||||
| Description | Protein phosphatase 2C isoform gamma (EC 3.1.3.16) (PP2C-gamma) (Protein phosphatase magnesium-dependent 1 gamma) (Protein phosphatase 1C). | |||||
|
PRRT3_HUMAN
|
||||||
| θ value | 2.36792 (rank : 80) | NC score | 0.051212 (rank : 62) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q5FWE3, Q49AD0, Q6UXY6, Q8NBC9 | Gene names | PRRT3, UNQ5823/PRO19642 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
RPGR_MOUSE
|
||||||
| θ value | 2.36792 (rank : 81) | NC score | 0.026574 (rank : 124) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9R0X5, O88408, Q9CU92 | Gene names | Rpgr | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator (mRpgr). | |||||
|
SFRIP_HUMAN
|
||||||
| θ value | 2.36792 (rank : 82) | NC score | 0.068082 (rank : 28) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 530 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q99590, Q8IW59 | Gene names | SFRS2IP, CASP11, SIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SFRS2-interacting protein (Splicing factor, arginine/serine-rich 2- interacting protein) (SC35-interacting protein 1) (CTD-associated SR protein 11) (Splicing regulatory protein 129) (SRrp129) (NY-REN-40 antigen). | |||||
|
SGOL2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 83) | NC score | 0.030903 (rank : 108) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q562F6, Q53RR9, Q53T20, Q86XY4, Q8IWK2, Q8IZK1, Q8N1Q5, Q96LQ3 | Gene names | SGOL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Shugoshin-like 2 (Tripin). | |||||
|
SNW1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 84) | NC score | 0.036572 (rank : 89) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9CSN1 | Gene names | Snw1, Skiip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SNW domain-containing protein 1 (Nuclear protein SkiP) (Ski- interacting protein). | |||||
|
SPN90_HUMAN
|
||||||
| θ value | 2.36792 (rank : 85) | NC score | 0.025299 (rank : 127) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NZQ3, Q9UGM8 | Gene names | NCKIPSD, AF3P21, SPIN90 | |||
|
Domain Architecture |
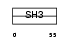
|
|||||
| Description | SH3 adapter protein SPIN90 (NCK-interacting protein with SH3 domain) (SH3 protein interacting with Nck, 90 kDa) (VacA-interacting protein, 54 kDa) (VIP54) (AF3p21) (Diaphanous protein-interacting protein) (Dia-interacting protein 1) (DIP-1). | |||||
|
SRRM1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 86) | NC score | 0.102534 (rank : 7) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
ANK2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 87) | NC score | 0.031804 (rank : 105) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 87 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
CABL1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 88) | NC score | 0.021021 (rank : 138) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9ESJ1, Q9EPR8 | Gene names | Cables1, Cables | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CDK5 and ABL1 enzyme substrate 1 (Interactor with CDK3 1) (Ik3-1). | |||||
|
CN155_HUMAN
|
||||||
| θ value | 3.0926 (rank : 89) | NC score | 0.058129 (rank : 45) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q5H9T9, Q5H9U7, Q86YI2, Q9H0J3 | Gene names | C14orf155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf155. | |||||
|
DNM3B_MOUSE
|
||||||
| θ value | 3.0926 (rank : 90) | NC score | 0.017336 (rank : 151) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O88509, O88510, O88511 | Gene names | Dnmt3b | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 3B (EC 2.1.1.37) (Dnmt3b) (DNA methyltransferase MmuIIIB) (DNA MTase MmuIIIB) (M.MmuIIIB). | |||||
|
HSF2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 91) | NC score | 0.019366 (rank : 145) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q03933, Q9H445 | Gene names | HSF2, HSTF2 | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock factor protein 2 (HSF 2) (Heat shock transcription factor 2) (HSTF 2). | |||||
|
PHLPP_HUMAN
|
||||||
| θ value | 3.0926 (rank : 92) | NC score | 0.013158 (rank : 171) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 604 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O60346, Q641Q7, Q6P4C4, Q6PJI6, Q86TN6, Q96FK2, Q9NUY1 | Gene names | PHLPP, KIAA0606, PLEKHE1, SCOP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH domain leucine-rich repeat-containing protein phosphatase (EC 3.1.3.16) (PH domain leucine-rich repeat protein phosphatase) (Pleckstrin homology domain-containing family E protein 1) (Suprachiasmatic nucleus circadian oscillatory protein) (hSCOP). | |||||
|
STXB5_HUMAN
|
||||||
| θ value | 3.0926 (rank : 93) | NC score | 0.042994 (rank : 81) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5T5C0, Q5T5C1, Q5T5C2, Q8NBG8, Q96NG9 | Gene names | STXBP5, LLGL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntaxin-binding protein 5 (Tomosyn-1) (Lethal(2) giant larvae protein homolog 3). | |||||
|
STXB5_MOUSE
|
||||||
| θ value | 3.0926 (rank : 94) | NC score | 0.043094 (rank : 80) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8K400, Q80TM5, Q8K401, Q9D2F3 | Gene names | Stxbp5, Kiaa4253, Llgl3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntaxin-binding protein 5 (Tomosyn-1) (Lethal(2) giant larvae protein homolog 3). | |||||
|
TSH3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 95) | NC score | 0.018648 (rank : 147) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 564 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q63HK5, Q9H0G6, Q9P254 | Gene names | TSHZ3, KIAA1474, TSH3, ZNF537 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Teashirt homolog 3 (Zinc finger protein 537). | |||||
|
WDR7_MOUSE
|
||||||
| θ value | 3.0926 (rank : 96) | NC score | 0.030163 (rank : 110) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 226 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q920I9, Q80TY3, Q920J0 | Gene names | Wdr7, Kiaa0541, Trag | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 7 (TGF-beta resistance-associated protein TRAG). | |||||
|
ZN292_HUMAN
|
||||||
| θ value | 3.0926 (rank : 97) | NC score | 0.019423 (rank : 144) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 918 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O60281, Q9H8G3, Q9H8J4 | Gene names | ZNF292, KIAA0530 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 292. | |||||
|
ZN406_MOUSE
|
||||||
| θ value | 3.0926 (rank : 98) | NC score | 0.002375 (rank : 196) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 800 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q7TS63 | Gene names | Znf406, Zfp406 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 406. | |||||
|
BAZ2A_MOUSE
|
||||||
| θ value | 4.03905 (rank : 99) | NC score | 0.028178 (rank : 122) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q91YE5 | Gene names | Baz2a, Tip5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5). | |||||
|
CDC20_HUMAN
|
||||||
| θ value | 4.03905 (rank : 100) | NC score | 0.024470 (rank : 129) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q12834, Q9BW56, Q9UQI9 | Gene names | CDC20 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle protein 20 homolog (p55CDC). | |||||
|
CDC20_MOUSE
|
||||||
| θ value | 4.03905 (rank : 101) | NC score | 0.024702 (rank : 128) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9JJ66, Q3TGP1, Q8BPG4, Q99LK3 | Gene names | Cdc20 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle protein 20 homolog (p55CDC) (mmCdc20). | |||||
|
COG8_HUMAN
|
||||||
| θ value | 4.03905 (rank : 102) | NC score | 0.022096 (rank : 134) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96MW5, Q8WVV6, Q9H6F8 | Gene names | COG8 | |||
|
Domain Architecture |
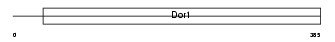
|
|||||
| Description | Conserved oligomeric Golgi complex component 8. | |||||
|
FOXI1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 103) | NC score | 0.006914 (rank : 187) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q922I5, Q5SRI5, Q9D299 | Gene names | Foxi1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein I1. | |||||
|
GEMI5_MOUSE
|
||||||
| θ value | 4.03905 (rank : 104) | NC score | 0.032557 (rank : 101) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 424 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8BX17 | Gene names | Gemin5 | |||
|
Domain Architecture |

|
|||||
| Description | Gem-associated protein 5 (Gemin5). | |||||
|
JADE3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 105) | NC score | 0.012400 (rank : 175) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q6IE82, Q6ZQG2 | Gene names | Phf16, Jade3, Kiaa0215 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-3 (PHD finger protein 16). | |||||
|
KLK2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 106) | NC score | 0.002684 (rank : 195) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P20151, Q15946, Q9UJZ9 | Gene names | KLK2 | |||
|
Domain Architecture |
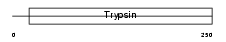
|
|||||
| Description | Kallikrein-2 precursor (EC 3.4.21.35) (Tissue kallikrein-2) (Glandular kallikrein-1) (hGK-1). | |||||
|
PTPRN_MOUSE
|
||||||
| θ value | 4.03905 (rank : 107) | NC score | 0.009565 (rank : 181) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q60673, Q62129 | Gene names | Ptprn, Ptp35 | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase-like N precursor (R-PTP-N) (PTP IA-2). | |||||
|
RP1L1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 108) | NC score | 0.040328 (rank : 85) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1072 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q8IWN7, Q86SQ1, Q8IWN8, Q8IWN9, Q8IWP0, Q8IWP1, Q8IWP2 | Gene names | RP1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinitis pigmentosa 1-like 1 protein. | |||||
|
SIGL6_HUMAN
|
||||||
| θ value | 4.03905 (rank : 109) | NC score | 0.011728 (rank : 178) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O43699, O15388, O43700 | Gene names | SIGLEC6, CD33L, CD33L1, OBBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sialic acid-binding Ig-like lectin 6 precursor (Siglec-6) (Obesity- binding protein 1) (OB-BP1) (CD33 antigen-like 1) (CDw327 antigen). | |||||
|
TEX10_HUMAN
|
||||||
| θ value | 4.03905 (rank : 110) | NC score | 0.029012 (rank : 115) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NXF1, Q5T722, Q5T723, Q8NCN8, Q8TDY0 | Gene names | TEX10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-expressed sequence 10 protein. | |||||
|
ZC11A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 111) | NC score | 0.069456 (rank : 26) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O75152, Q6AHY4, Q6AHY9, Q6AW79, Q6AWA1, Q6PJK4, Q86XZ7 | Gene names | ZC3H11A, KIAA0663 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 11A. | |||||
|
CT096_HUMAN
|
||||||
| θ value | 5.27518 (rank : 112) | NC score | 0.020351 (rank : 143) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 438 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NUD7, Q8N840, Q8NAX5 | Gene names | C20orf96 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C20orf96. | |||||
|
CTR9_HUMAN
|
||||||
| θ value | 5.27518 (rank : 113) | NC score | 0.028651 (rank : 117) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 383 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q6PD62, Q15015 | Gene names | CTR9, KIAA0155, SH2BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein CTR9 homolog (SH2 domain-binding protein 1). | |||||
|
DCC_MOUSE
|
||||||
| θ value | 5.27518 (rank : 114) | NC score | 0.015873 (rank : 161) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P70211 | Gene names | Dcc | |||
|
Domain Architecture |

|
|||||
| Description | Netrin receptor DCC precursor (Tumor suppressor protein DCC). | |||||
|
EMSY_HUMAN
|
||||||
| θ value | 5.27518 (rank : 115) | NC score | 0.034594 (rank : 92) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q7Z589, Q4G109, Q8NBU6, Q8TE50, Q9H8I9, Q9NRH0 | Gene names | EMSY, C11orf30 | |||
|
Domain Architecture |
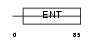
|
|||||
| Description | Protein EMSY. | |||||
|
FOG1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 116) | NC score | 0.019072 (rank : 146) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1237 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O35615 | Gene names | Zfpm1, Fog, Fog1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein ZFPM1 (Zinc finger protein multitype 1) (Friend of GATA protein 1) (Friend of GATA-1) (FOG-1). | |||||
|
HRX_HUMAN
|
||||||
| θ value | 5.27518 (rank : 117) | NC score | 0.034006 (rank : 96) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
JHD2C_HUMAN
|
||||||
| θ value | 5.27518 (rank : 118) | NC score | 0.017364 (rank : 150) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q15652, Q5SQZ8, Q5SQZ9, Q5SR00, Q7Z3E7, Q8N3U0, Q96KB9, Q9P2G7 | Gene names | JMJD1C, JHDM2C, KIAA1380, TRIP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable JmjC domain-containing histone demethylation protein 2C (EC 1.14.11.-) (Jumonji domain-containing protein 1C) (Thyroid receptor-interacting protein 8) (TRIP-8). | |||||
|
LIRB3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 119) | NC score | 0.007470 (rank : 185) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O75022, O15471, Q86U49 | Gene names | LILRB3, ILT5, LIR3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leukocyte immunoglobulin-like receptor subfamily B member 3 precursor (Leukocyte immunoglobulin-like receptor 3) (LIR-3) (Immunoglobulin- like transcript 5) (ILT-5) (Monocyte inhibitory receptor HL9) (CD85a antigen). | |||||
|
M3K12_HUMAN
|
||||||
| θ value | 5.27518 (rank : 120) | NC score | 0.005263 (rank : 189) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 940 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q12852, Q86VQ5, Q8WY25 | Gene names | MAP3K12, ZPK | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase 12 (EC 2.7.11.25) (Mixed lineage kinase) (Leucine-zipper protein kinase) (ZPK) (Dual leucine zipper bearing kinase) (DLK) (MAPK-upstream kinase) (MUK). | |||||
|
PALB2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 121) | NC score | 0.022527 (rank : 133) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q86YC2, Q8N7Y6, Q8ND31, Q9H6W1 | Gene names | PALB2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Partner and localizer of BRCA2. | |||||
|
PSD4_MOUSE
|
||||||
| θ value | 5.27518 (rank : 122) | NC score | 0.012398 (rank : 176) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8BLR5, Q3TE33, Q3TQF5, Q3UD59, Q3UFH9, Q80V44 | Gene names | Psd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH and SEC7 domain-containing protein 4 (Pleckstrin homology and SEC7 domain-containing protein 4). | |||||
|
PTN12_HUMAN
|
||||||
| θ value | 5.27518 (rank : 123) | NC score | 0.007184 (rank : 186) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q05209, Q16130, Q59FD6, Q75MN8, Q86XU4 | Gene names | PTPN12 | |||
|
Domain Architecture |
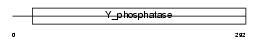
|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 12 (EC 3.1.3.48) (Protein-tyrosine phosphatase G1) (PTPG1). | |||||
|
RIF1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 124) | NC score | 0.030117 (rank : 111) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q6PR54, Q6T8D9, Q7TPD8, Q8BTU2, Q8C408, Q9CS77 | Gene names | Rif1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Telomere-associated protein RIF1 (Rap1-interacting factor 1 homolog) (mRif1). | |||||
|
RIMB2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 125) | NC score | 0.014538 (rank : 165) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O15034, Q96ID2 | Gene names | RIMBP2, KIAA0318, RBP2 | |||
|
Domain Architecture |

|
|||||
| Description | RIM-binding protein 2 (RIM-BP2). | |||||
|
RIMS1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 126) | NC score | 0.036280 (rank : 90) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 437 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q86UR5, O15048, Q8TDY9, Q8TDZ5, Q9HBA1, Q9HBA2, Q9HBA3, Q9HBA4, Q9HBA5, Q9HBA6 | Gene names | RIMS1, KIAA0340, RIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 1 (Rab3-interacting molecule 1) (RIM 1). | |||||
|
RSPO1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 127) | NC score | 0.012708 (rank : 173) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Z132, Q3V1S3 | Gene names | Rspo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | R-spondin-1 precursor (Roof plate-specific spondin-1) (Cysteine-rich and single thrombospondin domain-containing protein 3) (Cristin-3) (mCristin-3). | |||||
|
SNW1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 128) | NC score | 0.031671 (rank : 107) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q13573, Q13483, Q32N03, Q5D0D6 | Gene names | SNW1, SKIIP, SKIP | |||
|
Domain Architecture |
|
|||||
| Description | SNW domain-containing protein 1 (Nuclear protein SkiP) (Ski- interacting protein) (Nuclear receptor coactivator NCoA-62). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 129) | NC score | 0.092760 (rank : 14) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
TSH3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 130) | NC score | 0.021104 (rank : 136) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 590 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8CGV9, Q5DTX4 | Gene names | Tshz3, Kiaa1474, Tsh3, Zfp537, Znf537 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Teashirt homolog 3 (Zinc finger protein 537). | |||||
|
UBP2L_MOUSE
|
||||||
| θ value | 5.27518 (rank : 131) | NC score | 0.032239 (rank : 102) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q80X50, Q8BIT6, Q8BIW4, Q8BJ01, Q8CIG7, Q8K102, Q9CRT6 | Gene names | Ubap2l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-associated protein 2-like. | |||||
|
WDFY1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 132) | NC score | 0.026519 (rank : 125) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8IWB7, Q9H9D5, Q9P2B3 | Gene names | WDFY1, KIAA1435, WDF1, ZFYVE17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat and FYVE domain-containing protein 1 (WD40- and FYVE domain- containing protein 1) (Phosphoinositide-binding protein 1) (FENS-1) (Zinc finger FYVE domain-containing protein 17). | |||||
|
WDR69_HUMAN
|
||||||
| θ value | 5.27518 (rank : 133) | NC score | 0.010237 (rank : 179) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 324 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8N136, Q6ZRY1, Q8N776 | Gene names | WDR69 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 69. | |||||
|
ZN592_HUMAN
|
||||||
| θ value | 5.27518 (rank : 134) | NC score | 0.004818 (rank : 190) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 762 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q92610 | Gene names | ZNF592, KIAA0211 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 592. | |||||
|
ADA19_MOUSE
|
||||||
| θ value | 6.88961 (rank : 135) | NC score | 0.003915 (rank : 191) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O35674 | Gene names | Adam19, Mltnb | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 19 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 19) (Meltrin beta). | |||||
|
ATF7_HUMAN
|
||||||
| θ value | 6.88961 (rank : 136) | NC score | 0.016498 (rank : 157) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P17544, Q13814 | Gene names | ATF7, ATFA | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-7 (Activating transcription factor 7) (Transcription factor ATF-A). | |||||
|
BAT2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 137) | NC score | 0.043694 (rank : 76) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
BRCA2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 138) | NC score | 0.032724 (rank : 99) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P51587, O00183, O15008, Q13879 | Gene names | BRCA2, FANCD1 | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer type 2 susceptibility protein (Fanconi anemia group D1 protein). | |||||
|
CNGB3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 139) | NC score | 0.014112 (rank : 168) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9NQW8, Q9NRE9 | Gene names | CNGB3 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic nucleotide-gated cation channel beta 3 (CNG channel beta 3) (Cyclic nucleotide-gated channel beta 3) (Cone photoreceptor cGMP- gated channel subunit beta) (Cyclic nucleotide-gated cation channel modulatory subunit). | |||||
|
CSKI1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 140) | NC score | 0.016683 (rank : 155) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1031 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q6P9K8, Q6ZPU2, Q8BWU2, Q8BX99, Q9CXH0 | Gene names | Caskin1, Kiaa1306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-1 (CASK-interacting protein 1). | |||||
|
CT2NL_HUMAN
|
||||||
| θ value | 6.88961 (rank : 141) | NC score | 0.015667 (rank : 162) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1059 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9P2B4, Q96B40 | Gene names | CTTNBP2NL, KIAA1433 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CTTNBP2 N-terminal-like protein. | |||||
|
DSG2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 142) | NC score | 0.002739 (rank : 194) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O55111, Q8K069, Q8R517 | Gene names | Dsg2 | |||
|
Domain Architecture |

|
|||||
| Description | Desmoglein-2 precursor. | |||||
|
FA47A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 143) | NC score | 0.052108 (rank : 59) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 451 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q5JRC9, Q8TAA0 | Gene names | FAM47A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM47A. | |||||
|
IQGA1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 144) | NC score | 0.009460 (rank : 183) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9JKF1 | Gene names | Iqgap1 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP1. | |||||
|
JIP4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 145) | NC score | 0.028234 (rank : 119) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q58A65, Q3UH77, Q3UHF0, Q58VQ4, Q5NC70, Q5NC78, Q6A057, Q6PAS3, Q8CJC2 | Gene names | Spag9, Jip4, Jsap2, Kiaa0516, Mapk8ip4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 4 (JNK-interacting protein 4) (JIP-4) (JNK-associated leucine-zipper protein) (JLP) (Sperm-associated antigen 9) (Mitogen-activated protein kinase 8- interacting protein 4) (JNK/SAPK-associated protein 2) (JSAP2). | |||||
|
L2GL2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 146) | NC score | 0.029891 (rank : 112) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6P1M3, Q14521, Q9BR62 | Gene names | LLGL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lethal(2) giant larvae protein homolog 2 (HGL). | |||||
|
MDC1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 147) | NC score | 0.075827 (rank : 22) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
MYST3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 148) | NC score | 0.028182 (rank : 121) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q92794 | Gene names | MYST3, MOZ, RUNXBP2, ZNF220 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Runt-related transcription factor-binding protein 2) (Monocytic leukemia zinc finger protein) (Zinc finger protein 220). | |||||
|
RSNL2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 149) | NC score | 0.012651 (rank : 174) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CI96, Q8BW09, Q921Q4, Q9D2S6 | Gene names | Rsnl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Restin-like protein 2. | |||||
|
TAF4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 150) | NC score | 0.045775 (rank : 74) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 687 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O00268, Q5TBP6, Q99721, Q9BR40, Q9BX42 | Gene names | TAF4, TAF2C, TAF2C1, TAF4A, TAFII130, TAFII135 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 4 (TBP-associated factor 4) (Transcription initiation factor TFIID 135 kDa subunit) (TAF(II)135) (TAFII-135) (TAFII135) (TAFII-130) (TAFII130). | |||||
|
TAF5L_MOUSE
|
||||||
| θ value | 6.88961 (rank : 151) | NC score | 0.016656 (rank : 156) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q91WQ5, Q8BTL5 | Gene names | Taf5l, Paf65b | |||
|
Domain Architecture |

|
|||||
| Description | TAF5-like RNA polymerase II p300/CBP-associated factor-associated factor 65 kDa subunit 5L (PCAF-associated factor 65 beta) (PAF65- beta). | |||||
|
UT14A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 152) | NC score | 0.013311 (rank : 170) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9BVJ6, Q5JYF1 | Gene names | UTP14A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U3 small nucleolar RNA-associated protein 14 homolog A (Antigen NY-CO- 16). | |||||
|
WDR46_MOUSE
|
||||||
| θ value | 6.88961 (rank : 153) | NC score | 0.033052 (rank : 97) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9Z0H1 | Gene names | Wdr46, Bing4 | |||
|
Domain Architecture |
|
|||||
| Description | WD repeat protein 46 (WD repeat protein BING4). | |||||
|
BTK_HUMAN
|
||||||
| θ value | 8.99809 (rank : 154) | NC score | 0.003546 (rank : 193) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1026 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q06187 | Gene names | BTK, AGMX1, ATK, BPK | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein kinase BTK (EC 2.7.10.2) (Bruton tyrosine kinase) (Agammaglobulinaemia tyrosine kinase) (ATK) (B cell progenitor kinase) (BPK). | |||||
|
BTK_MOUSE
|
||||||
| θ value | 8.99809 (rank : 155) | NC score | 0.003605 (rank : 192) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1017 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P35991, Q61365 | Gene names | Btk, Bpk | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein kinase BTK (EC 2.7.10.2) (Bruton tyrosine kinase) (Agammaglobulinaemia tyrosine kinase) (ATK) (B cell progenitor kinase) (BPK) (Kinase EMB). | |||||
|
CD2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 156) | NC score | 0.029371 (rank : 113) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P06729, Q96TE5 | Gene names | CD2 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell surface antigen CD2 precursor (T-cell surface antigen T11/Leu- 5) (LFA-2) (LFA-3 receptor) (Erythrocyte receptor) (Rosette receptor). | |||||
|
CUTL1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 157) | NC score | 0.016894 (rank : 152) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P53564, O08994, P70301, Q571L6, Q91ZD2 | Gene names | Cutl1, Cux, Cux1, Kiaa4047 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP) (Homeobox protein Cux). | |||||
|
GPTC8_HUMAN
|
||||||
| θ value | 8.99809 (rank : 158) | NC score | 0.038707 (rank : 86) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 366 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9UKJ3, O60300 | Gene names | GPATC8, KIAA0553 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 8. | |||||
|
HELZ_HUMAN
|
||||||
| θ value | 8.99809 (rank : 159) | NC score | 0.016014 (rank : 160) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P42694 | Gene names | HELZ, KIAA0054 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable helicase with zinc-finger domain (EC 3.6.1.-). | |||||
|
HMGA1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 160) | NC score | 0.040596 (rank : 84) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P17096, P10910, Q9UKB0 | Gene names | HMGA1, HMGIY | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | High mobility group protein HMG-I/HMG-Y (HMG-I(Y)) (High mobility group AT-hook protein 1) (High mobility group protein A1) (High mobility group protein-R). | |||||
|
KSR1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 161) | NC score | 0.005427 (rank : 188) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q61097, Q61648, Q78DX8 | Gene names | Ksr1, Ksr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinase suppressor of ras-1 (Kinase suppressor of ras) (mKSR1) (Hb protein). | |||||
|
NCOR2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 162) | NC score | 0.034566 (rank : 93) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 790 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9WU42, Q9WU43, Q9WUC1 | Gene names | Ncor2, Smrt | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC). | |||||
|
PHF20_HUMAN
|
||||||
| θ value | 8.99809 (rank : 163) | NC score | 0.042628 (rank : 82) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9BVI0, Q5JWY9, Q9BWV4, Q9BXA3, Q9BZW3, Q9H421, Q9H4J6, Q9NZ22 | Gene names | PHF20, C20orf104, GLEA2, HCA58, NZF, TZP | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58) (Glioma-expressed antigen 2) (Transcription factor TZP) (Novel zinc finger protein). | |||||
|
PKHA4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 164) | NC score | 0.020517 (rank : 142) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9H4M7, Q8N4M8, Q8N658 | Gene names | PLEKHA4, PEPP1 | |||
|
Domain Architecture |
|
|||||
| Description | Pleckstrin homology domain-containing family A member 4 (Phosphoinositol 3-phosphate-binding protein 1) (PEPP-1). | |||||
|
PRCC_HUMAN
|
||||||
| θ value | 8.99809 (rank : 165) | NC score | 0.030738 (rank : 109) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q92733, O00665, O00724 | Gene names | PRCC, TPRC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein PRCC (Papillary renal cell carcinoma translocation-associated gene protein). | |||||
|
PRPF3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 166) | NC score | 0.014746 (rank : 164) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q922U1, Q3TJH4, Q9D6C6 | Gene names | Prpf3 | |||
|
Domain Architecture |
|
|||||
| Description | U4/U6 small nuclear ribonucleoprotein Prp3 (Pre-mRNA-splicing factor 3). | |||||
|
SENP7_HUMAN
|
||||||
| θ value | 8.99809 (rank : 167) | NC score | 0.014451 (rank : 167) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9BQF6, Q96PS5, Q9C0F6, Q9HBT5 | Gene names | SENP7, KIAA1707, SSP2, SUSP2 | |||
|
Domain Architecture |

|
|||||
| Description | Sentrin-specific protease 7 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP7) (SUMO-1-specific protease 2). | |||||
|
SOS2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 168) | NC score | 0.009771 (rank : 180) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q07890, Q15503 | Gene names | SOS2 | |||
|
Domain Architecture |

|
|||||
| Description | Son of sevenless homolog 2 (SOS-2). | |||||
|
SYN3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 169) | NC score | 0.020839 (rank : 141) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O14994 | Gene names | SYN3 | |||
|
Domain Architecture |
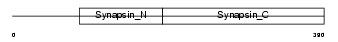
|
|||||
| Description | Synapsin-3 (Synapsin III). | |||||
|
THAP3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 170) | NC score | 0.013818 (rank : 169) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8WTV1, Q569K1, Q5TH66, Q5TH67, Q8N8T6, Q9BSC7, Q9Y3H2, Q9Y3H3 | Gene names | THAP3 | |||
|
Domain Architecture |
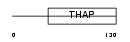
|
|||||
| Description | THAP domain-containing protein 3. | |||||
|
TITIN_HUMAN
|
||||||
| θ value | 8.99809 (rank : 171) | NC score | 0.016170 (rank : 159) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 2923 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | Q8WZ42, Q10465, Q10466, Q15598, Q2XUS3, Q32Q60, Q4U1Z6, Q6NSG0, Q6PDB1, Q6PJP0, Q7KYM2, Q7KYN4, Q7KYN5, Q7LDM3, Q7Z2X3, Q8TCG8, Q8WZ51, Q8WZ52, Q8WZ53, Q8WZB3, Q92761, Q92762, Q9UD97, Q9UP84, Q9Y6L9 | Gene names | TTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Titin (EC 2.7.11.1) (Connectin) (Rhabdomyosarcoma antigen MU-RMS- 40.14). | |||||
|
TNR8_MOUSE
|
||||||
| θ value | 8.99809 (rank : 172) | NC score | 0.016826 (rank : 153) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q60846 | Gene names | Tnfrsf8 | |||
|
Domain Architecture |
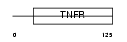
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 8 precursor (CD30L receptor) (Lymphocyte activation antigen CD30). | |||||
|
TXND2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 173) | NC score | 0.069421 (rank : 27) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
UROL1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 174) | NC score | 0.009463 (rank : 182) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5DID0, Q5DIC9, Q6LA40, Q6LA41, Q8N216 | Gene names | UMODL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uromodulin-like 1 precursor (Olfactorin). | |||||
|
WD51A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 175) | NC score | 0.012024 (rank : 177) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8NBT0, Q2TAK6, Q96IK6, Q9UFJ8 | Gene names | WDR51A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 51A. | |||||
|
ZN318_MOUSE
|
||||||
| θ value | 8.99809 (rank : 176) | NC score | 0.023234 (rank : 131) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 642 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q99PP2, Q3TZL5, Q9JJ01 | Gene names | Znf318, Tzf, Zfp318 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 318 (Testicular zinc finger protein). | |||||
|
AFF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 177) | NC score | 0.054319 (rank : 56) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P51825 | Gene names | AFF1, AF4, FEL, MLLT2 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4) (Protein FEL). | |||||
|
AKA12_HUMAN
|
||||||
| θ value | θ > 10 (rank : 178) | NC score | 0.050324 (rank : 69) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q02952, O00310, O00498, Q99970 | Gene names | AKAP12, AKAP250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 12 (A-kinase anchor protein 250 kDa) (AKAP 250) (Myasthenia gravis autoantigen gravin). | |||||
|
BSN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 179) | NC score | 0.050748 (rank : 64) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
BSN_MOUSE
|
||||||
| θ value | θ > 10 (rank : 180) | NC score | 0.057913 (rank : 47) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
CJ012_HUMAN
|
||||||
| θ value | θ > 10 (rank : 181) | NC score | 0.050648 (rank : 65) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q8N655, Q9H945, Q9Y457 | Gene names | C10orf12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf12. | |||||
|
CK024_MOUSE
|
||||||
| θ value | θ > 10 (rank : 182) | NC score | 0.056237 (rank : 53) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9D8N1, Q8VCP2 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C11orf24 homolog precursor. | |||||
|
GRIN1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 183) | NC score | 0.055246 (rank : 54) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q3UNH4, Q6PGF9, Q9QZY2 | Gene names | Gprin1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
LAD1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 184) | NC score | 0.072841 (rank : 24) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 391 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P57016 | Gene names | Lad1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ladinin 1 (Lad-1) (Linear IgA disease autoantigen). | |||||
|
MAP1B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 185) | NC score | 0.062369 (rank : 38) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | P46821 | Gene names | MAP1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) [Contains: MAP1 light chain LC1]. | |||||
|
MAP1B_MOUSE
|
||||||
| θ value | θ > 10 (rank : 186) | NC score | 0.050633 (rank : 66) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 977 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | P14873 | Gene names | Map1b, Mtap1b, Mtap5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) (MAP1.2) (MAP1(X)) [Contains: MAP1 light chain LC1]. | |||||
|
MAP4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 187) | NC score | 0.069896 (rank : 25) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P27546 | Gene names | Map4, Mtap4 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 4 (MAP 4). | |||||
|
NOLC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 188) | NC score | 0.065078 (rank : 31) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q14978, Q15030 | Gene names | NOLC1, KIAA0035 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar phosphoprotein p130 (Nucleolar 130 kDa protein) (140 kDa nucleolar phosphoprotein) (Nopp140) (Nucleolar and coiled-body phosphoprotein 1). | |||||
|
RBBP6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 189) | NC score | 0.055000 (rank : 55) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
RL1D1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 190) | NC score | 0.050406 (rank : 68) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O76021, Q6PL22, Q8IWS7, Q8WUZ1, Q9HDA9, Q9Y3Z9 | Gene names | RSL1D1, CATX11, CSIG, PBK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ribosomal L1 domain-containing protein 1 (Cellular senescence- inhibited gene protein) (Protein PBK1) (CATX-11). | |||||
|
RSF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 191) | NC score | 0.057529 (rank : 50) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 539 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q96T23, Q86X86, Q9H3L8, Q9NVZ8, Q9NYU0 | Gene names | RSF1, HBXAP, XAP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Remodeling and spacing factor 1 (Rsf-1) (Hepatitis B virus X- associated protein) (HBV pX-associated protein 8) (p325 subunit of RSF chromatin remodelling complex). | |||||
|
SON_HUMAN
|
||||||
| θ value | θ > 10 (rank : 192) | NC score | 0.052224 (rank : 58) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
TGON2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 193) | NC score | 0.073455 (rank : 23) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
TRDN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 194) | NC score | 0.057656 (rank : 49) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q13061 | Gene names | TRDN | |||
|
Domain Architecture |

|
|||||
| Description | Triadin. | |||||
|
TXND2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 195) | NC score | 0.057953 (rank : 46) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
ZCH10_MOUSE
|
||||||
| θ value | θ > 10 (rank : 196) | NC score | 0.056282 (rank : 52) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9CX48 | Gene names | Zcchc10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 10. | |||||
|
AHTF1_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 176 | |
| SwissProt Accessions | Q8CJF7, Q8BVJ5, Q8VD55 | Gene names | Ahctf1, Elys | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-hook-containing transcription factor 1 (Embryonic large molecule derived from yolk sac). | |||||
|
AHTF1_HUMAN
|
||||||
| NC score | 0.911494 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8WYP5, Q7Z4E3, Q8IZA4, Q96EH9, Q9Y4Q6 | Gene names | AHCTF1, ELYS, TMBS62 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-hook-containing transcription factor 1 (Embryonic large molecule derived from yolk sac). | |||||
|
SFR16_MOUSE
|
||||||
| NC score | 0.124176 (rank : 3) | θ value | 0.21417 (rank : 24) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8CFC7, Q8CFC8, Q9Z2N3 | Gene names | Sfrs16, Clasp, Swap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 16 (Suppressor of white-apricot homolog 2) (Clk4-associating SR-related protein). | |||||
|
SFR16_HUMAN
|
||||||
| NC score | 0.117947 (rank : 4) | θ value | 0.365318 (rank : 30) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8N2M8, O96026, Q6UW71, Q96DX2 | Gene names | SFRS16, SWAP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 16 (Suppressor of white-apricot homolog 2). | |||||
|
KI67_HUMAN
|
||||||
| NC score | 0.117883 (rank : 5) | θ value | 0.0563607 (rank : 13) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P46013 | Gene names | MKI67 | |||
|
Domain Architecture |

|
|||||
| Description | Antigen KI-67. | |||||
|
PRG4_MOUSE
|
||||||
| NC score | 0.112472 (rank : 6) | θ value | 0.00175202 (rank : 3) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9JM99, Q3UEL1, Q3V198 | Gene names | Prg4, Msf, Szp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
SRRM1_MOUSE
|
||||||
| NC score | 0.102534 (rank : 7) | θ value | 2.36792 (rank : 86) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.102197 (rank : 8) | θ value | 0.365318 (rank : 32) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
SRRM1_HUMAN
|
||||||
| NC score | 0.101841 (rank : 9) | θ value | 0.62314 (rank : 42) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
MAP4_HUMAN
|
||||||
| NC score | 0.099435 (rank : 10) | θ value | 0.00228821 (rank : 5) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 428 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P27816, Q13082, Q96A76 | Gene names | MAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 4 (MAP 4). | |||||
|
MDC1_HUMAN
|
||||||
| NC score | 0.097777 (rank : 11) | θ value | 0.0330416 (rank : 9) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
PCLO_HUMAN
|
||||||
| NC score | 0.097305 (rank : 12) | θ value | 0.0563607 (rank : 15) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 87 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
PRG4_HUMAN
|
||||||
| NC score | 0.094553 (rank : 13) | θ value | 1.38821 (rank : 61) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.092760 (rank : 14) | θ value | 5.27518 (rank : 129) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
PAF49_HUMAN
|
||||||
| NC score | 0.090714 (rank : 15) | θ value | 0.163984 (rank : 20) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O15446, Q32N11, Q7Z5U2, Q9UPF6 | Gene names | CD3EAP, ASE1, CAST, PAF49 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase I-associated factor PAF49 (Anti-sense to ERCC-1 protein) (ASE-1) (CD3-epsilon-associated protein) (CD3E-associated protein) (CAST). | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.088609 (rank : 16) | θ value | 0.0431538 (rank : 12) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 87 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
SNPC4_HUMAN
|
||||||
| NC score | 0.088073 (rank : 17) | θ value | 0.00175202 (rank : 4) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q5SXM2, Q9Y6P7 | Gene names | SNAPC4, SNAP190 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | snRNA-activating protein complex subunit 4 (SNAPc subunit 4) (snRNA- activating protein complex 190 kDa subunit) (SNAPc 190 kDa subunit) (Proximal sequence element-binding transcription factor subunit alpha) (PSE-binding factor subunit alpha) (PTF subunit alpha). | |||||
|
NIPBL_HUMAN
|
||||||
| NC score | 0.087708 (rank : 18) | θ value | 0.0113563 (rank : 7) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
PCLO_MOUSE
|
||||||
| NC score | 0.087093 (rank : 19) | θ value | 0.00509761 (rank : 6) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 92 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
PHF20_MOUSE
|
||||||
| NC score | 0.082915 (rank : 20) | θ value | 0.163984 (rank : 21) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q8BLG0, Q8BMA2, Q8BYR4, Q921N1 | Gene names | Phf20, Hca58 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58 homolog). | |||||
|
NPBL_MOUSE
|
||||||
| NC score | 0.077439 (rank : 21) | θ value | 1.06291 (rank : 54) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
MDC1_MOUSE
|
||||||
| NC score | 0.075827 (rank : 22) | θ value | 6.88961 (rank : 147) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
TGON2_HUMAN
|
||||||
| NC score | 0.073455 (rank : 23) | θ value | θ > 10 (rank : 193) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
LAD1_MOUSE
|
||||||
| NC score | 0.072841 (rank : 24) | θ value | θ > 10 (rank : 184) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 391 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P57016 | Gene names | Lad1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ladinin 1 (Lad-1) (Linear IgA disease autoantigen). | |||||
|
MAP4_MOUSE
|
||||||
| NC score | 0.069896 (rank : 25) | θ value | θ > 10 (rank : 187) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P27546 | Gene names | Map4, Mtap4 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 4 (MAP 4). | |||||
|
ZC11A_HUMAN
|
||||||
| NC score | 0.069456 (rank : 26) | θ value | 4.03905 (rank : 111) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O75152, Q6AHY4, Q6AHY9, Q6AW79, Q6AWA1, Q6PJK4, Q86XZ7 | Gene names | ZC3H11A, KIAA0663 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 11A. | |||||
|
TXND2_MOUSE
|
||||||
| NC score | 0.069421 (rank : 27) | θ value | 8.99809 (rank : 173) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
SFRIP_HUMAN
|
||||||
| NC score | 0.068082 (rank : 28) | θ value | 2.36792 (rank : 82) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 530 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q99590, Q8IW59 | Gene names | SFRS2IP, CASP11, SIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SFRS2-interacting protein (Splicing factor, arginine/serine-rich 2- interacting protein) (SC35-interacting protein 1) (CTD-associated SR protein 11) (Splicing regulatory protein 129) (SRrp129) (NY-REN-40 antigen). | |||||
|
MADCA_HUMAN
|
||||||
| NC score | 0.065955 (rank : 29) | θ value | 0.62314 (rank : 39) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q13477, O60222, O75867 | Gene names | MADCAM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucosal addressin cell adhesion molecule 1 precursor (MAdCAM-1) (hMAdCAM-1). | |||||
|
TARA_HUMAN
|
||||||
| NC score | 0.065567 (rank : 30) | θ value | 1.06291 (rank : 56) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | Q9H2D6, O94797, Q2PZW8, Q2Q3Z9, Q2Q400, Q5R3M6, Q96DW1, Q9BT77, Q9BTL7, Q9BY98, Q9Y3L4 | Gene names | TRIOBP, KIAA1662, TARA | |||
|
Domain Architecture |

|
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
NOLC1_HUMAN
|
||||||
| NC score | 0.065078 (rank : 31) | θ value | θ > 10 (rank : 188) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q14978, Q15030 | Gene names | NOLC1, KIAA0035 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar phosphoprotein p130 (Nucleolar 130 kDa protein) (140 kDa nucleolar phosphoprotein) (Nopp140) (Nucleolar and coiled-body phosphoprotein 1). | |||||
|
GRIN1_HUMAN
|
||||||
| NC score | 0.064109 (rank : 32) | θ value | 1.06291 (rank : 52) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q7Z2K8, Q8ND74, Q96PZ4 | Gene names | GPRIN1, KIAA1893 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
PHAR2_HUMAN
|
||||||
| NC score | 0.063903 (rank : 33) | θ value | 0.279714 (rank : 28) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O75167, Q68DM2 | Gene names | PHACTR2, C6orf56, KIAA0680 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatase and actin regulator 2. | |||||
|
FHOD1_MOUSE
|
||||||
| NC score | 0.063678 (rank : 34) | θ value | 0.125558 (rank : 19) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6P9Q4, Q8BMK2 | Gene names | Fhod1, Fhos1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FH1/FH2 domain-containing protein (Formin homolog overexpressed in spleen) (FHOS) (Formin homology 2 domain-containing protein 1). | |||||
|
FHOD1_HUMAN
|
||||||
| NC score | 0.063196 (rank : 35) | θ value | 0.279714 (rank : 26) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9Y613, Q59F76, Q6Y1F2, Q76MS8, Q8N521 | Gene names | FHOD1, FHOS | |||
|
Domain Architecture |

|
|||||
| Description | FH1/FH2 domain-containing protein (Formin homolog overexpressed in spleen) (FHOS) (Formin homology 2 domain-containing protein 1). | |||||
|
AL2S4_MOUSE
|
||||||
| NC score | 0.063049 (rank : 36) | θ value | 0.813845 (rank : 45) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q3V0J1, Q3TIS2 | Gene names | Als2cr4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyotrophic lateral sclerosis 2 chromosomal region candidate gene 4 protein homolog. | |||||
|
PCQAP_MOUSE
|
||||||
| NC score | 0.062684 (rank : 37) | θ value | 1.06291 (rank : 55) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q924H2 | Gene names | Pcqap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Positive cofactor 2 glutamine/Q-rich-associated protein (PC2 glutamine/Q-rich-associated protein) (mPcqap). | |||||
|
MAP1B_HUMAN
|
||||||
| NC score | 0.062369 (rank : 38) | θ value | θ > 10 (rank : 185) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | P46821 | Gene names | MAP1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) [Contains: MAP1 light chain LC1]. | |||||
|
CKAP2_HUMAN
|
||||||
| NC score | 0.060787 (rank : 39) | θ value | 0.279714 (rank : 25) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8WWK9, Q3KRA5, Q5VXB4, Q8IWV5, Q8IWV6, Q96FH9, Q9H012, Q9H0D0, Q9H988, Q9HC49, Q9NVG4 | Gene names | CKAP2, LB1, TMAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-associated protein 2 (Tumor-associated microtubule- associated protein) (CTCL tumor antigen se20-10). | |||||
|
NFH_MOUSE
|
||||||
| NC score | 0.060153 (rank : 40) | θ value | 1.06291 (rank : 53) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 97 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
SETX_HUMAN
|
||||||
| NC score | 0.059424 (rank : 41) | θ value | 0.0961366 (rank : 18) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q7Z333, O75120, Q3KQX4, Q68DW5, Q6AZD7, Q7Z3J6, Q8WX33, Q9H9D1, Q9NVP9 | Gene names | SETX, ALS4, KIAA0625, SCAR1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable helicase senataxin (EC 3.6.1.-) (SEN1 homolog). | |||||
|
PCQAP_HUMAN
|
||||||
| NC score | 0.058700 (rank : 42) | θ value | 1.38821 (rank : 60) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 549 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q96RN5, O15413, Q6IC31, Q8NF16, Q96CT0, Q96IH7, Q9P1T3 | Gene names | PCQAP, ARC105, CTG7A, TIG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Positive cofactor 2 glutamine/Q-rich-associated protein (PC2 glutamine/Q-rich-associated protein) (TPA-inducible gene 1 protein) (TIG-1) (Activator-recruited cofactor 105 kDa component) (ARC105) (CTG repeat protein 7a). | |||||
|
WASIP_HUMAN
|
||||||
| NC score | 0.058631 (rank : 43) | θ value | 1.38821 (rank : 64) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O43516, Q15220, Q6MZU9, Q9BU37, Q9UNP1 | Gene names | WASPIP, WIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein) (PRPL-2 protein). | |||||
|
APXL_HUMAN
|
||||||
| NC score | 0.058326 (rank : 44) | θ value | 0.47712 (rank : 34) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 553 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q13796 | Gene names | APXL | |||
|
Domain Architecture |

|
|||||
| Description | Apical-like protein (Protein APXL). | |||||
|
CN155_HUMAN
|
||||||
| NC score | 0.058129 (rank : 45) | θ value | 3.0926 (rank : 89) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q5H9T9, Q5H9U7, Q86YI2, Q9H0J3 | Gene names | C14orf155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf155. | |||||
|
TXND2_HUMAN
|
||||||
| NC score | 0.057953 (rank : 46) | θ value | θ > 10 (rank : 195) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
BSN_MOUSE
|
||||||
| NC score | 0.057913 (rank : 47) | θ value | θ > 10 (rank : 180) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
WASIP_MOUSE
|
||||||
| NC score | 0.057827 (rank : 48) | θ value | 1.81305 (rank : 73) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8K1I7, Q3U0U8 | Gene names | Waspip, Wip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein). | |||||
|
TRDN_HUMAN
|
||||||
| NC score | 0.057656 (rank : 49) | θ value | θ > 10 (rank : 194) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q13061 | Gene names | TRDN | |||
|
Domain Architecture |

|
|||||
| Description | Triadin. | |||||
|
RSF1_HUMAN
|
||||||
| NC score | 0.057529 (rank : 50) | θ value | θ > 10 (rank : 191) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 539 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q96T23, Q86X86, Q9H3L8, Q9NVZ8, Q9NYU0 | Gene names | RSF1, HBXAP, XAP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Remodeling and spacing factor 1 (Rsf-1) (Hepatitis B virus X- associated protein) (HBV pX-associated protein 8) (p325 subunit of RSF chromatin remodelling complex). | |||||
|
RBBP6_HUMAN
|
||||||
| NC score | 0.057343 (rank : 51) | θ value | 0.47712 (rank : 35) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
ZCH10_MOUSE
|
||||||
| NC score | 0.056282 (rank : 52) | θ value | θ > 10 (rank : 196) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9CX48 | Gene names | Zcchc10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 10. | |||||
|
CK024_MOUSE
|
||||||
| NC score | 0.056237 (rank : 53) | θ value | θ > 10 (rank : 182) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9D8N1, Q8VCP2 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C11orf24 homolog precursor. | |||||
|
GRIN1_MOUSE
|
||||||
| NC score | 0.055246 (rank : 54) | θ value | θ > 10 (rank : 183) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q3UNH4, Q6PGF9, Q9QZY2 | Gene names | Gprin1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
RBBP6_MOUSE
|
||||||
| NC score | 0.055000 (rank : 55) | θ value | θ > 10 (rank : 189) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
AFF1_HUMAN
|
||||||
| NC score | 0.054319 (rank : 56) | θ value | θ > 10 (rank : 177) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P51825 | Gene names | AFF1, AF4, FEL, MLLT2 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4) (Protein FEL). | |||||
|
STB5L_HUMAN
|
||||||
| NC score | 0.052832 (rank : 57) | θ value | 0.62314 (rank : 43) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y2K9, Q4G1B4, Q6PIC3 | Gene names | STXBP5L, KIAA1006, LLGL4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntaxin-binding protein 5-like (Tomosyn-2) (Lethal(2) giant larvae protein homolog 4). | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.052224 (rank : 58) | θ value | θ > 10 (rank : 192) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
FA47A_HUMAN
|
||||||
| NC score | 0.052108 (rank : 59) | θ value | 6.88961 (rank : 143) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 451 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q5JRC9, Q8TAA0 | Gene names | FAM47A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM47A. | |||||
|
TULP1_HUMAN
|
||||||
| NC score | 0.051360 (rank : 60) | θ value | 0.279714 (rank : 29) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O00294, O43536 | Gene names | TULP1 | |||
|
Domain Architecture |

|
|||||
| Description | Tubby-related protein 1 (Tubby-like protein 1). | |||||
|
STB5L_MOUSE
|
||||||
| NC score | 0.051274 (rank : 61) | θ value | 0.813845 (rank : 50) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q5DQR4, Q3TSM3, Q3UVG2, Q5DQR1, Q5DQR2, Q5DQR3, Q8BS52 | Gene names | Stxbp5l, Llgl4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntaxin-binding protein 5-like (Tomosyn-2) (Lethal(2) giant larvae protein homolog 4). | |||||
|
PRRT3_HUMAN
|
||||||
| NC score | 0.051212 (rank : 62) | θ value | 2.36792 (rank : 80) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q5FWE3, Q49AD0, Q6UXY6, Q8NBC9 | Gene names | PRRT3, UNQ5823/PRO19642 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
UBP31_HUMAN
|
||||||
| NC score | 0.050840 (rank : 63) | θ value | 0.0330416 (rank : 10) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 293 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q70CQ4, Q6AW97, Q6ZTC0, Q6ZTN2, Q9ULL7 | Gene names | USP31, KIAA1203 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 31 (EC 3.1.2.15) (Ubiquitin thioesterase 31) (Ubiquitin-specific-processing protease 31) (Deubiquitinating enzyme 31). | |||||
|
BSN_HUMAN
|
||||||
| NC score | 0.050748 (rank : 64) | θ value | θ > 10 (rank : 179) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
CJ012_HUMAN
|
||||||
| NC score | 0.050648 (rank : 65) | θ value | θ > 10 (rank : 181) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q8N655, Q9H945, Q9Y457 | Gene names | C10orf12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf12. | |||||
|
MAP1B_MOUSE
|
||||||
| NC score | 0.050633 (rank : 66) | θ value | θ > 10 (rank : 186) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 977 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | P14873 | Gene names | Map1b, Mtap1b, Mtap5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) (MAP1.2) (MAP1(X)) [Contains: MAP1 light chain LC1]. | |||||
|
RTEL1_HUMAN
|
||||||
| NC score | 0.050543 (rank : 67) | θ value | 0.0961366 (rank : 17) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NZ71, Q5JTV4, Q9BW37, Q9H402, Q9H4X6, Q9NX25, Q9NZ73, Q9UPR4, Q9Y4R6 | Gene names | RTEL1, C20orf41, KIAA1088, NHL | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of telomere elongation helicase 1 (EC 3.6.1.-) (Helicase- like protein NHL). | |||||
|
RL1D1_HUMAN
|
||||||
| NC score | 0.050406 (rank : 68) | θ value | θ > 10 (rank : 190) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O76021, Q6PL22, Q8IWS7, Q8WUZ1, Q9HDA9, Q9Y3Z9 | Gene names | RSL1D1, CATX11, CSIG, PBK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ribosomal L1 domain-containing protein 1 (Cellular senescence- inhibited gene protein) (Protein PBK1) (CATX-11). | |||||
|
AKA12_HUMAN
|
||||||
| NC score | 0.050324 (rank : 69) | θ value | θ > 10 (rank : 178) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q02952, O00310, O00498, Q99970 | Gene names | AKAP12, AKAP250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 12 (A-kinase anchor protein 250 kDa) (AKAP 250) (Myasthenia gravis autoantigen gravin). | |||||
|
TULP1_MOUSE
|
||||||
| NC score | 0.050233 (rank : 70) | θ value | 0.365318 (rank : 33) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Z273 | Gene names | Tulp1 | |||
|
Domain Architecture |

|
|||||
| Description | Tubby-related protein 1 (Tubby-like protein 1). | |||||
|
NFH_HUMAN
|
||||||
| NC score | 0.049774 (rank : 71) | θ value | 0.279714 (rank : 27) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
RGS3_MOUSE
|
||||||
| NC score | 0.047689 (rank : 72) | θ value | 0.62314 (rank : 41) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
ALMS1_HUMAN
|
||||||
| NC score | 0.047177 (rank : 73) | θ value | 1.06291 (rank : 51) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8TCU4, Q53S05, Q580Q8, Q86VP9, Q9Y4G4 | Gene names | ALMS1, KIAA0328 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alstrom syndrome protein 1. | |||||
|
TAF4_HUMAN
|
||||||
| NC score | 0.045775 (rank : 74) | θ value | 6.88961 (rank : 150) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 687 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | O00268, Q5TBP6, Q99721, Q9BR40, Q9BX42 | Gene names | TAF4, TAF2C, TAF2C1, TAF4A, TAFII130, TAFII135 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 4 (TBP-associated factor 4) (Transcription initiation factor TFIID 135 kDa subunit) (TAF(II)135) (TAFII-135) (TAFII135) (TAFII-130) (TAFII130). | |||||
|
SCYBG_MOUSE
|
||||||
| NC score | 0.043755 (rank : 75) | θ value | 1.81305 (rank : 72) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BSU2, Q8VE25, Q9EPB3 | Gene names | Cxcl16, Srpsox | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Small inducible cytokine B16 precursor (Transmembrane chemokine CXCL16) (SR-PSOX) (Scavenger receptor for phosphatidylserine and oxidized low density lipoprotein). | |||||
|
BAT2_HUMAN
|
||||||
| NC score | 0.043694 (rank : 76) | θ value | 6.88961 (rank : 137) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
K0460_HUMAN
|
||||||
| NC score | 0.043419 (rank : 77) | θ value | 0.813845 (rank : 47) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q5VT52, O75048, Q5VT53, Q6MZL4, Q86XD2 | Gene names | KIAA0460 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0460. | |||||
|
SOCS6_MOUSE
|
||||||
| NC score | 0.043350 (rank : 78) | θ value | 0.0736092 (rank : 16) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9JLY0, Q9D2A0 | Gene names | Socs6, Cis4, Cish4, Socs4 | |||
|
Domain Architecture |

|
|||||
| Description | Suppressor of cytokine signaling 6 (SOCS-6) (Suppressor of cytokine signaling 4) (SOCS-4) (Cytokine-inducible SH2 protein 4) (CIS-4). | |||||
|
RERE_MOUSE
|
||||||
| NC score | 0.043160 (rank : 79) | θ value | 1.81305 (rank : 70) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 785 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q80TZ9 | Gene names | Rere, Atr2, Kiaa0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-2). | |||||
|
STXB5_MOUSE
|
||||||
| NC score | 0.043094 (rank : 80) | θ value | 3.0926 (rank : 94) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8K400, Q80TM5, Q8K401, Q9D2F3 | Gene names | Stxbp5, Kiaa4253, Llgl3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntaxin-binding protein 5 (Tomosyn-1) (Lethal(2) giant larvae protein homolog 3). | |||||
|
STXB5_HUMAN
|
||||||
| NC score | 0.042994 (rank : 81) | θ value | 3.0926 (rank : 93) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5T5C0, Q5T5C1, Q5T5C2, Q8NBG8, Q96NG9 | Gene names | STXBP5, LLGL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntaxin-binding protein 5 (Tomosyn-1) (Lethal(2) giant larvae protein homolog 3). | |||||
|
PHF20_HUMAN
|
||||||
| NC score | 0.042628 (rank : 82) | θ value | 8.99809 (rank : 163) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9BVI0, Q5JWY9, Q9BWV4, Q9BXA3, Q9BZW3, Q9H421, Q9H4J6, Q9NZ22 | Gene names | PHF20, C20orf104, GLEA2, HCA58, NZF, TZP | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58) (Glioma-expressed antigen 2) (Transcription factor TZP) (Novel zinc finger protein). | |||||
|
JIP3_HUMAN
|
||||||
| NC score | 0.041299 (rank : 83) | θ value | 0.21417 (rank : 23) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1039 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9UPT6, Q96RY4, Q9H4I4, Q9H7P1, Q9NUG0 | Gene names | MAPK8IP3, JIP3, KIAA1066 | |||
|
Domain Architecture |

|
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 3 (JNK-interacting protein 3) (JIP-3) (JNK MAP kinase scaffold protein 3) (Mitogen- activated protein kinase 8-interacting protein 3). | |||||
|
HMGA1_HUMAN
|
||||||
| NC score | 0.040596 (rank : 84) | θ value | 8.99809 (rank : 160) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P17096, P10910, Q9UKB0 | Gene names | HMGA1, HMGIY | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | High mobility group protein HMG-I/HMG-Y (HMG-I(Y)) (High mobility group AT-hook protein 1) (High mobility group protein A1) (High mobility group protein-R). | |||||
|
RP1L1_HUMAN
|
||||||
| NC score | 0.040328 (rank : 85) | θ value | 4.03905 (rank : 108) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1072 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q8IWN7, Q86SQ1, Q8IWN8, Q8IWN9, Q8IWP0, Q8IWP1, Q8IWP2 | Gene names | RP1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinitis pigmentosa 1-like 1 protein. | |||||
|
GPTC8_HUMAN
|
||||||
| NC score | 0.038707 (rank : 86) | θ value | 8.99809 (rank : 158) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 366 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9UKJ3, O60300 | Gene names | GPATC8, KIAA0553 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 8. | |||||
|
MUC2_HUMAN
|
||||||
| NC score | 0.038053 (rank : 87) | θ value | 2.36792 (rank : 77) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q02817, Q14878 | Gene names | MUC2, SMUC | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-2 precursor (Intestinal mucin-2). | |||||
|
INCE_MOUSE
|
||||||
| NC score | 0.036825 (rank : 88) | θ value | 0.62314 (rank : 38) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9WU62, Q7TN28, Q8BGN4, Q8CGI4 | Gene names | Incenp | |||
|
Domain Architecture |

|
|||||
| Description | Inner centromere protein. | |||||
|
SNW1_MOUSE
|
||||||
| NC score | 0.036572 (rank : 89) | θ value | 2.36792 (rank : 84) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9CSN1 | Gene names | Snw1, Skiip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SNW domain-containing protein 1 (Nuclear protein SkiP) (Ski- interacting protein). | |||||
|
RIMS1_HUMAN
|
||||||
| NC score | 0.036280 (rank : 90) | θ value | 5.27518 (rank : 126) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 437 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q86UR5, O15048, Q8TDY9, Q8TDZ5, Q9HBA1, Q9HBA2, Q9HBA3, Q9HBA4, Q9HBA5, Q9HBA6 | Gene names | RIMS1, KIAA0340, RIM1 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 1 (Rab3-interacting molecule 1) (RIM 1). | |||||
|
RIMS2_MOUSE
|
||||||
| NC score | 0.035226 (rank : 91) | θ value | 1.81305 (rank : 71) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 376 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9EQZ7, Q8C433, Q8CCK2 | Gene names | Rims2, Rab3ip2, Rim2 | |||
|
Domain Architecture |

|
|||||
| Description | Regulating synaptic membrane exocytosis protein 2 (Rab3-interacting molecule 2) (RIM 2) (Rab3-interacting protein 2). | |||||
|
EMSY_HUMAN
|
||||||
| NC score | 0.034594 (rank : 92) | θ value | 5.27518 (rank : 115) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q7Z589, Q4G109, Q8NBU6, Q8TE50, Q9H8I9, Q9NRH0 | Gene names | EMSY, C11orf30 | |||
|
Domain Architecture |

|
|||||
| Description | Protein EMSY. | |||||
|
NCOR2_MOUSE
|
||||||
| NC score | 0.034566 (rank : 93) | θ value | 8.99809 (rank : 162) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 790 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q9WU42, Q9WU43, Q9WUC1 | Gene names | Ncor2, Smrt | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC). | |||||
|
LAP2B_MOUSE
|
||||||
| NC score | 0.034482 (rank : 94) | θ value | 0.813845 (rank : 48) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q61029, Q61030, Q61031, Q61032 | Gene names | Tmpo, Lap2 | |||
|
Domain Architecture |

|
|||||
| Description | Lamina-associated polypeptide 2 isoforms beta/delta/epsilon/gamma (Thymopoietin isoforms beta/delta/epsilon/gamma) (TP beta/delta/epsilon/gamma). | |||||
|
CD4_HUMAN
|
||||||
| NC score | 0.034287 (rank : 95) | θ value | 1.38821 (rank : 57) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P01730, Q4ZGK2, Q5U066 | Gene names | CD4 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell surface glycoprotein CD4 precursor (T-cell surface antigen T4/Leu-3). | |||||
|
HRX_HUMAN
|
||||||
| NC score | 0.034006 (rank : 96) | θ value | 5.27518 (rank : 117) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
WDR46_MOUSE
|
||||||
| NC score | 0.033052 (rank : 97) | θ value | 6.88961 (rank : 153) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9Z0H1 | Gene names | Wdr46, Bing4 | |||
|
Domain Architecture |

|
|||||
| Description | WD repeat protein 46 (WD repeat protein BING4). | |||||
|
CHSS1_HUMAN
|
||||||
| NC score | 0.032994 (rank : 98) | θ value | 0.62314 (rank : 36) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q86X52, Q6UX38, Q7LFU5, Q9Y2J5 | Gene names | CHSY1, CHSY, CSS1, KIAA0990 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate synthase 1 (EC 2.4.1.175) (Glucuronosyl-N- acetylgalactosaminyl-proteoglycan 4-beta-N- acetylgalactosaminyltransferase 1) (Chondroitin sulfate synthase 1) (N-acetylgalactosaminyl-proteoglycan 3-beta-glucuronosyltransferase 1) (EC 2.4.1.226) (Chondroitin glucuronyltransferase II) (N- acetylgalactosaminyltransferase II). | |||||
|
BRCA2_HUMAN
|
||||||
| NC score | 0.032724 (rank : 99) | θ value | 6.88961 (rank : 138) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P51587, O00183, O15008, Q13879 | Gene names | BRCA2, FANCD1 | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer type 2 susceptibility protein (Fanconi anemia group D1 protein). | |||||
|
MAGD2_HUMAN
|
||||||
| NC score | 0.032586 (rank : 100) | θ value | 0.0431538 (rank : 11) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UNF1, O76058, Q5BJF3, Q8NAL6, Q9H218, Q9P0U9, Q9UM52 | Gene names | MAGED2, BCG1 | |||
|
Domain Architecture |

|
|||||
| Description | Melanoma-associated antigen D2 (MAGE-D2 antigen) (MAGE-D) (Breast cancer-associated gene 1 protein) (BCG-1) (11B6) (Hepatocellular carcinoma-associated protein JCL-1). | |||||
|
GEMI5_MOUSE
|
||||||
| NC score | 0.032557 (rank : 101) | θ value | 4.03905 (rank : 104) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 424 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8BX17 | Gene names | Gemin5 | |||
|
Domain Architecture |

|
|||||
| Description | Gem-associated protein 5 (Gemin5). | |||||
|
UBP2L_MOUSE
|
||||||
| NC score | 0.032239 (rank : 102) | θ value | 5.27518 (rank : 131) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q80X50, Q8BIT6, Q8BIW4, Q8BJ01, Q8CIG7, Q8K102, Q9CRT6 | Gene names | Ubap2l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-associated protein 2-like. | |||||
|
WDR7_HUMAN
|
||||||
| NC score | 0.032164 (rank : 103) | θ value | 1.81305 (rank : 74) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 244 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Y4E6, Q86UX5, Q86VP2, Q96PS7 | Gene names | WDR7, KIAA0541, TRAG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 7 (TGF-beta resistance-associated protein TRAG) (Rabconnectin-3 beta). | |||||
|
CALD1_HUMAN
|
||||||
| NC score | 0.031994 (rank : 104) | θ value | 1.81305 (rank : 66) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 959 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q05682, Q13978, Q13979, Q14741, Q14742 | Gene names | CALD1, CAD, CDM | |||
|
Domain Architecture |

|
|||||
| Description | Caldesmon (CDM). | |||||
|
ANK2_HUMAN
|
||||||
| NC score | 0.031804 (rank : 105) | θ value | 3.0926 (rank : 87) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 87 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
CLAP1_HUMAN
|
||||||
| NC score | 0.031677 (rank : 106) | θ value | 0.813845 (rank : 46) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q7Z460, O75118, Q8N5B8, Q9BQT5 | Gene names | CLASP1, KIAA0622, MAST1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CLIP-associating protein 1 (Cytoplasmic linker-associated protein 1) (Multiple asters homolog 1). | |||||
|
SNW1_HUMAN
|
||||||
| NC score | 0.031671 (rank : 107) | θ value | 5.27518 (rank : 128) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q13573, Q13483, Q32N03, Q5D0D6 | Gene names | SNW1, SKIIP, SKIP | |||
|
Domain Architecture |

|
|||||
| Description | SNW domain-containing protein 1 (Nuclear protein SkiP) (Ski- interacting protein) (Nuclear receptor coactivator NCoA-62). | |||||
|
SGOL2_HUMAN
|
||||||
| NC score | 0.030903 (rank : 108) | θ value | 2.36792 (rank : 83) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q562F6, Q53RR9, Q53T20, Q86XY4, Q8IWK2, Q8IZK1, Q8N1Q5, Q96LQ3 | Gene names | SGOL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Shugoshin-like 2 (Tripin). | |||||
|
PRCC_HUMAN
|
||||||
| NC score | 0.030738 (rank : 109) | θ value | 8.99809 (rank : 165) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q92733, O00665, O00724 | Gene names | PRCC, TPRC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein PRCC (Papillary renal cell carcinoma translocation-associated gene protein). | |||||
|
WDR7_MOUSE
|
||||||
| NC score | 0.030163 (rank : 110) | θ value | 3.0926 (rank : 96) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 226 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q920I9, Q80TY3, Q920J0 | Gene names | Wdr7, Kiaa0541, Trag | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 7 (TGF-beta resistance-associated protein TRAG). | |||||
|
RIF1_MOUSE
|
||||||
| NC score | 0.030117 (rank : 111) | θ value | 5.27518 (rank : 124) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q6PR54, Q6T8D9, Q7TPD8, Q8BTU2, Q8C408, Q9CS77 | Gene names | Rif1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Telomere-associated protein RIF1 (Rap1-interacting factor 1 homolog) (mRif1). | |||||
|
L2GL2_HUMAN
|
||||||
| NC score | 0.029891 (rank : 112) | θ value | 6.88961 (rank : 146) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6P1M3, Q14521, Q9BR62 | Gene names | LLGL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lethal(2) giant larvae protein homolog 2 (HGL). | |||||
|
CD2_HUMAN
|
||||||
| NC score | 0.029371 (rank : 113) | θ value | 8.99809 (rank : 156) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P06729, Q96TE5 | Gene names | CD2 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell surface antigen CD2 precursor (T-cell surface antigen T11/Leu- 5) (LFA-2) (LFA-3 receptor) (Erythrocyte receptor) (Rosette receptor). | |||||
|
MELPH_MOUSE
|
||||||
| NC score | 0.029166 (rank : 114) | θ value | 1.81305 (rank : 68) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q91V27, Q99N53 | Gene names | Mlph, Ln, Slac2a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanophilin (Exophilin-3) (Leaden protein) (Synaptotagmin-like protein 2a) (Slac2-a) (Slp homolog lacking C2 domains a). | |||||
|
TEX10_HUMAN
|
||||||
| NC score | 0.029012 (rank : 115) | θ value | 4.03905 (rank : 110) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NXF1, Q5T722, Q5T723, Q8NCN8, Q8TDY0 | Gene names | TEX10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-expressed sequence 10 protein. | |||||
|
TMC1_HUMAN
|
||||||
| NC score | 0.028802 (rank : 116) | θ value | 0.62314 (rank : 44) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8TDI8 | Gene names | TMC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane cochlear-expressed protein 1. | |||||
|
CTR9_HUMAN
|
||||||
| NC score | 0.028651 (rank : 117) | θ value | 5.27518 (rank : 113) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 383 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q6PD62, Q15015 | Gene names | CTR9, KIAA0155, SH2BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein CTR9 homolog (SH2 domain-binding protein 1). | |||||
|
SAMN1_MOUSE
|
||||||
| NC score | 0.028318 (rank : 118) | θ value | 1.38821 (rank : 62) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P57725, Q8C6S2 | Gene names | Samsn1 | |||
|
Domain Architecture |

|
|||||
| Description | SAM domain-containing protein SAMSN-1 (SAM domain, SH3 domain and nuclear localization signals protein 1). | |||||
|
JIP4_MOUSE
|
||||||
| NC score | 0.028234 (rank : 119) | θ value | 6.88961 (rank : 145) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q58A65, Q3UH77, Q3UHF0, Q58VQ4, Q5NC70, Q5NC78, Q6A057, Q6PAS3, Q8CJC2 | Gene names | Spag9, Jip4, Jsap2, Kiaa0516, Mapk8ip4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | C-jun-amino-terminal kinase-interacting protein 4 (JNK-interacting protein 4) (JIP-4) (JNK-associated leucine-zipper protein) (JLP) (Sperm-associated antigen 9) (Mitogen-activated protein kinase 8- interacting protein 4) (JNK/SAPK-associated protein 2) (JSAP2). | |||||
|
CHSS1_MOUSE
|
||||||
| NC score | 0.028214 (rank : 120) | θ value | 2.36792 (rank : 75) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6ZQ11 | Gene names | Chsy1, Kiaa0990 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate synthase 1 (EC 2.4.1.175) (Glucuronosyl-N- acetylgalactosaminyl-proteoglycan 4-beta-N- acetylgalactosaminyltransferase) (Chondroitin sulfate synthase 1) (N- acetylgalactosaminyl-proteoglycan 3-beta-glucuronosyltransferase) (EC 2.4.1.226) (Chondroitin glucuronyltransferase II) (N- acetylgalactosaminyltransferase II). | |||||
|
MYST3_HUMAN
|
||||||
| NC score | 0.028182 (rank : 121) | θ value | 6.88961 (rank : 148) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q92794 | Gene names | MYST3, MOZ, RUNXBP2, ZNF220 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Runt-related transcription factor-binding protein 2) (Monocytic leukemia zinc finger protein) (Zinc finger protein 220). | |||||
|
BAZ2A_MOUSE
|
||||||
| NC score | 0.028178 (rank : 122) | θ value | 4.03905 (rank : 99) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q91YE5 | Gene names | Baz2a, Tip5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5). | |||||
|
NCOA7_HUMAN
|
||||||
| NC score | 0.026775 (rank : 123) | θ value | 1.81305 (rank : 69) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8NI08, Q3LID6, Q4G0V1, Q5TF95, Q6IPQ4, Q6NE83, Q86T89, Q8N1W4 | Gene names | NCOA7, ERAP140, ESNA1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 7 (140 kDa estrogen receptor-associated protein) (Estrogen nuclear receptor coactivator 1). | |||||
|
RPGR_MOUSE
|
||||||
| NC score | 0.026574 (rank : 124) | θ value | 2.36792 (rank : 81) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9R0X5, O88408, Q9CU92 | Gene names | Rpgr | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator (mRpgr). | |||||
|
WDFY1_HUMAN
|
||||||
| NC score | 0.026519 (rank : 125) | θ value | 5.27518 (rank : 132) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8IWB7, Q9H9D5, Q9P2B3 | Gene names | WDFY1, KIAA1435, WDF1, ZFYVE17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat and FYVE domain-containing protein 1 (WD40- and FYVE domain- containing protein 1) (Phosphoinositide-binding protein 1) (FENS-1) (Zinc finger FYVE domain-containing protein 17). | |||||
|
LPXN_MOUSE
|
||||||
| NC score | 0.025909 (rank : 126) | θ value | 0.0563607 (rank : 14) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99N69 | Gene names | Lpxn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leupaxin. | |||||
|
SPN90_HUMAN
|
||||||
| NC score | 0.025299 (rank : 127) | θ value | 2.36792 (rank : 85) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NZQ3, Q9UGM8 | Gene names | NCKIPSD, AF3P21, SPIN90 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 adapter protein SPIN90 (NCK-interacting protein with SH3 domain) (SH3 protein interacting with Nck, 90 kDa) (VacA-interacting protein, 54 kDa) (VIP54) (AF3p21) (Diaphanous protein-interacting protein) (Dia-interacting protein 1) (DIP-1). | |||||
|
CDC20_MOUSE
|
||||||
| NC score | 0.024702 (rank : 128) | θ value | 4.03905 (rank : 101) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9JJ66, Q3TGP1, Q8BPG4, Q99LK3 | Gene names | Cdc20 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle protein 20 homolog (p55CDC) (mmCdc20). | |||||
|
CDC20_HUMAN
|
||||||
| NC score | 0.024470 (rank : 129) | θ value | 4.03905 (rank : 100) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q12834, Q9BW56, Q9UQI9 | Gene names | CDC20 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle protein 20 homolog (p55CDC). | |||||
|
GNPTA_MOUSE
|
||||||
| NC score | 0.024131 (rank : 130) | θ value | 2.36792 (rank : 76) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q69ZN6, Q3U3K6, Q3US34 | Gene names | Gnptab, Gnpta, Kiaa1208 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylglucosamine-1-phosphotransferase subunits alpha/beta precursor (EC 2.7.8.17) (GlcNAc-1-phosphotransferase alpha/beta subunits) (UDP- N-acetylglucosamine-1-phosphotransferase alpha/beta subunits) (Stealth protein GNPTAB) [Contains: N-acetylglucosamine-1-phosphotransferase subunit alpha; N-acetylglucosamine-1-phosphotransferase subunit beta]. | |||||
|
ZN318_MOUSE
|
||||||
| NC score | 0.023234 (rank : 131) | θ value | 8.99809 (rank : 176) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 642 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q99PP2, Q3TZL5, Q9JJ01 | Gene names | Znf318, Tzf, Zfp318 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 318 (Testicular zinc finger protein). | |||||
|
OC90_HUMAN
|
||||||
| NC score | 0.023138 (rank : 132) | θ value | 2.36792 (rank : 78) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q02509 | Gene names | OC90, PLA2L | |||
|
Domain Architecture |

|
|||||
| Description | Otoconin 90 precursor (Oc90) (Phospholipase A2 homolog). | |||||
|
PALB2_HUMAN
|
||||||
| NC score | 0.022527 (rank : 133) | θ value | 5.27518 (rank : 121) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q86YC2, Q8N7Y6, Q8ND31, Q9H6W1 | Gene names | PALB2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Partner and localizer of BRCA2. | |||||
|
COG8_HUMAN
|
||||||
| NC score | 0.022096 (rank : 134) | θ value | 4.03905 (rank : 102) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96MW5, Q8WVV6, Q9H6F8 | Gene names | COG8 | |||
|
Domain Architecture |

|
|||||
| Description | Conserved oligomeric Golgi complex component 8. | |||||
|
PP2CG_HUMAN
|
||||||
| NC score | 0.022019 (rank : 135) | θ value | 2.36792 (rank : 79) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O15355 | Gene names | PPM1G, PPM1C | |||
|
Domain Architecture |

|
|||||
| Description | Protein phosphatase 2C isoform gamma (EC 3.1.3.16) (PP2C-gamma) (Protein phosphatase magnesium-dependent 1 gamma) (Protein phosphatase 1C). | |||||
|
TSH3_MOUSE
|
||||||
| NC score | 0.021104 (rank : 136) | θ value | 5.27518 (rank : 130) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 590 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8CGV9, Q5DTX4 | Gene names | Tshz3, Kiaa1474, Tsh3, Zfp537, Znf537 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Teashirt homolog 3 (Zinc finger protein 537). | |||||
|
SPEG_MOUSE
|
||||||
| NC score | 0.021089 (rank : 137) | θ value | 0.365318 (rank : 31) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
CABL1_MOUSE
|
||||||
| NC score | 0.021021 (rank : 138) | θ value | 3.0926 (rank : 88) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9ESJ1, Q9EPR8 | Gene names | Cables1, Cables | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CDK5 and ABL1 enzyme substrate 1 (Interactor with CDK3 1) (Ik3-1). | |||||
|
GCC2_HUMAN
|
||||||
| NC score | 0.020952 (rank : 139) | θ value | 0.62314 (rank : 37) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1558 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8IWJ2, O15045, Q8TDH3, Q9H2G8 | Gene names | GCC2, KIAA0336 | |||
|
Domain Architecture |

|
|||||
| Description | GRIP and coiled-coil domain-containing protein 2 (Golgi coiled coil protein GCC185) (CTCL tumor antigen se1-1) (CLL-associated antigen KW- 11) (NY-REN-53 antigen). | |||||
|
RBP24_HUMAN
|
||||||
| NC score | 0.020843 (rank : 140) | θ value | 0.62314 (rank : 40) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q4ZG46 | Gene names | RANBP2L4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 2-like 4 (RanBP2L4). | |||||
|
SYN3_HUMAN
|
||||||
| NC score | 0.020839 (rank : 141) | θ value | 8.99809 (rank : 169) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O14994 | Gene names | SYN3 | |||
|
Domain Architecture |

|
|||||
| Description | Synapsin-3 (Synapsin III). | |||||
|
PKHA4_HUMAN
|
||||||
| NC score | 0.020517 (rank : 142) | θ value | 8.99809 (rank : 164) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9H4M7, Q8N4M8, Q8N658 | Gene names | PLEKHA4, PEPP1 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 4 (Phosphoinositol 3-phosphate-binding protein 1) (PEPP-1). | |||||
|
CT096_HUMAN
|
||||||
| NC score | 0.020351 (rank : 143) | θ value | 5.27518 (rank : 112) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 438 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NUD7, Q8N840, Q8NAX5 | Gene names | C20orf96 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C20orf96. | |||||
|
ZN292_HUMAN
|
||||||
| NC score | 0.019423 (rank : 144) | θ value | 3.0926 (rank : 97) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 918 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O60281, Q9H8G3, Q9H8J4 | Gene names | ZNF292, KIAA0530 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 292. | |||||
|
HSF2_HUMAN
|
||||||
| NC score | 0.019366 (rank : 145) | θ value | 3.0926 (rank : 91) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q03933, Q9H445 | Gene names | HSF2, HSTF2 | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock factor protein 2 (HSF 2) (Heat shock transcription factor 2) (HSTF 2). | |||||
|
FOG1_MOUSE
|
||||||
| NC score | 0.019072 (rank : 146) | θ value | 5.27518 (rank : 116) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1237 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O35615 | Gene names | Zfpm1, Fog, Fog1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein ZFPM1 (Zinc finger protein multitype 1) (Friend of GATA protein 1) (Friend of GATA-1) (FOG-1). | |||||
|
TSH3_HUMAN
|
||||||
| NC score | 0.018648 (rank : 147) | θ value | 3.0926 (rank : 95) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 564 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q63HK5, Q9H0G6, Q9P254 | Gene names | TSHZ3, KIAA1474, TSH3, ZNF537 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Teashirt homolog 3 (Zinc finger protein 537). | |||||
|
ZEP1_MOUSE
|
||||||
| NC score | 0.018311 (rank : 148) | θ value | 1.38821 (rank : 65) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1063 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q03172 | Gene names | Hivep1, Cryabp1, Znf40 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 40 (Transcription factor alphaA-CRYBP1) (Alpha A- crystallin-binding protein I) (Alpha A-CRYBP1). | |||||
|
SPEG_HUMAN
|
||||||
| NC score | 0.017923 (rank : 149) | θ value | 1.38821 (rank : 63) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q15772, Q27J74, Q695L1, Q6FGA6, Q6ZQW1, Q6ZTL8, Q9P2P9 | Gene names | SPEG, APEG1, KIAA1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle preferentially expressed protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
JHD2C_HUMAN
|
||||||
| NC score | 0.017364 (rank : 150) | θ value | 5.27518 (rank : 118) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q15652, Q5SQZ8, Q5SQZ9, Q5SR00, Q7Z3E7, Q8N3U0, Q96KB9, Q9P2G7 | Gene names | JMJD1C, JHDM2C, KIAA1380, TRIP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable JmjC domain-containing histone demethylation protein 2C (EC 1.14.11.-) (Jumonji domain-containing protein 1C) (Thyroid receptor-interacting protein 8) (TRIP-8). | |||||
|
DNM3B_MOUSE
|
||||||
| NC score | 0.017336 (rank : 151) | θ value | 3.0926 (rank : 90) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O88509, O88510, O88511 | Gene names | Dnmt3b | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 3B (EC 2.1.1.37) (Dnmt3b) (DNA methyltransferase MmuIIIB) (DNA MTase MmuIIIB) (M.MmuIIIB). | |||||
|
CUTL1_MOUSE
|
||||||
| NC score | 0.016894 (rank : 152) | θ value | 8.99809 (rank : 157) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P53564, O08994, P70301, Q571L6, Q91ZD2 | Gene names | Cutl1, Cux, Cux1, Kiaa4047 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP) (Homeobox protein Cux). | |||||
|
TNR8_MOUSE
|
||||||
| NC score | 0.016826 (rank : 153) | θ value | 8.99809 (rank : 172) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q60846 | Gene names | Tnfrsf8 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 8 precursor (CD30L receptor) (Lymphocyte activation antigen CD30). | |||||
|
FOG1_HUMAN
|
||||||
| NC score | 0.016772 (rank : 154) | θ value | 1.38821 (rank : 58) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 981 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8IX07 | Gene names | ZFPM1, FOG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein ZFPM1 (Zinc finger protein multitype 1) (Friend of GATA protein 1) (Friend of GATA-1) (FOG-1). | |||||
|
CSKI1_MOUSE
|
||||||
| NC score | 0.016683 (rank : 155) | θ value | 6.88961 (rank : 140) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1031 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q6P9K8, Q6ZPU2, Q8BWU2, Q8BX99, Q9CXH0 | Gene names | Caskin1, Kiaa1306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-1 (CASK-interacting protein 1). | |||||
|
TAF5L_MOUSE
|
||||||
| NC score | 0.016656 (rank : 156) | θ value | 6.88961 (rank : 151) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q91WQ5, Q8BTL5 | Gene names | Taf5l, Paf65b | |||
|
Domain Architecture |

|
|||||
| Description | TAF5-like RNA polymerase II p300/CBP-associated factor-associated factor 65 kDa subunit 5L (PCAF-associated factor 65 beta) (PAF65- beta). | |||||
|
ATF7_HUMAN
|
||||||
| NC score | 0.016498 (rank : 157) | θ value | 6.88961 (rank : 136) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P17544, Q13814 | Gene names | ATF7, ATFA | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-7 (Activating transcription factor 7) (Transcription factor ATF-A). | |||||
|
KTNB1_MOUSE
|
||||||
| NC score | 0.016483 (rank : 158) | θ value | 1.81305 (rank : 67) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8BG40, Q8CD18, Q8R1J0, Q9CWV2 | Gene names | Katnb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Katanin p80 WD40-containing subunit B1 (Katanin p80 subunit B1) (p80 katanin). | |||||
|
TITIN_HUMAN
|
||||||
| NC score | 0.016170 (rank : 159) | θ value | 8.99809 (rank : 171) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 2923 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | Q8WZ42, Q10465, Q10466, Q15598, Q2XUS3, Q32Q60, Q4U1Z6, Q6NSG0, Q6PDB1, Q6PJP0, Q7KYM2, Q7KYN4, Q7KYN5, Q7LDM3, Q7Z2X3, Q8TCG8, Q8WZ51, Q8WZ52, Q8WZ53, Q8WZB3, Q92761, Q92762, Q9UD97, Q9UP84, Q9Y6L9 | Gene names | TTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Titin (EC 2.7.11.1) (Connectin) (Rhabdomyosarcoma antigen MU-RMS- 40.14). | |||||
|
HELZ_HUMAN
|
||||||
| NC score | 0.016014 (rank : 160) | θ value | 8.99809 (rank : 159) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P42694 | Gene names | HELZ, KIAA0054 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable helicase with zinc-finger domain (EC 3.6.1.-). | |||||
|
DCC_MOUSE
|
||||||
| NC score | 0.015873 (rank : 161) | θ value | 5.27518 (rank : 114) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P70211 | Gene names | Dcc | |||
|
Domain Architecture |

|
|||||
| Description | Netrin receptor DCC precursor (Tumor suppressor protein DCC). | |||||
|
CT2NL_HUMAN
|
||||||
| NC score | 0.015667 (rank : 162) | θ value | 6.88961 (rank : 141) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1059 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9P2B4, Q96B40 | Gene names | CTTNBP2NL, KIAA1433 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CTTNBP2 N-terminal-like protein. | |||||
|
MACF4_HUMAN
|
||||||
| NC score | 0.014968 (rank : 163) | θ value | 1.38821 (rank : 59) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1321 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q96PK2, Q8WXY1 | Gene names | MACF1, ABP620, ACF7, KIAA0465, KIAA1251 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1, isoform 4. | |||||
|
PRPF3_MOUSE
|
||||||
| NC score | 0.014746 (rank : 164) | θ value | 8.99809 (rank : 166) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q922U1, Q3TJH4, Q9D6C6 | Gene names | Prpf3 | |||
|
Domain Architecture |

|
|||||
| Description | U4/U6 small nuclear ribonucleoprotein Prp3 (Pre-mRNA-splicing factor 3). | |||||
|
RIMB2_HUMAN
|
||||||
| NC score | 0.014538 (rank : 165) | θ value | 5.27518 (rank : 125) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O15034, Q96ID2 | Gene names | RIMBP2, KIAA0318, RBP2 | |||
|
Domain Architecture |

|
|||||
| Description | RIM-binding protein 2 (RIM-BP2). | |||||
|
SALL2_MOUSE
|
||||||
| NC score | 0.014458 (rank : 166) | θ value | 0.163984 (rank : 22) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 836 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9QX96 | Gene names | Sall2, Sal2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sal-like protein 2 (Spalt-like protein 2) (MSal-2). | |||||
|
SENP7_HUMAN
|
||||||
| NC score | 0.014451 (rank : 167) | θ value | 8.99809 (rank : 167) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9BQF6, Q96PS5, Q9C0F6, Q9HBT5 | Gene names | SENP7, KIAA1707, SSP2, SUSP2 | |||
|
Domain Architecture |

|
|||||
| Description | Sentrin-specific protease 7 (EC 3.4.22.-) (Sentrin/SUMO-specific protease SENP7) (SUMO-1-specific protease 2). | |||||
|
CNGB3_HUMAN
|
||||||
| NC score | 0.014112 (rank : 168) | θ value | 6.88961 (rank : 139) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9NQW8, Q9NRE9 | Gene names | CNGB3 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic nucleotide-gated cation channel beta 3 (CNG channel beta 3) (Cyclic nucleotide-gated channel beta 3) (Cone photoreceptor cGMP- gated channel subunit beta) (Cyclic nucleotide-gated cation channel modulatory subunit). | |||||
|
THAP3_HUMAN
|
||||||
| NC score | 0.013818 (rank : 169) | θ value | 8.99809 (rank : 170) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8WTV1, Q569K1, Q5TH66, Q5TH67, Q8N8T6, Q9BSC7, Q9Y3H2, Q9Y3H3 | Gene names | THAP3 | |||
|
Domain Architecture |

|
|||||
| Description | THAP domain-containing protein 3. | |||||
|
UT14A_HUMAN
|
||||||
| NC score | 0.013311 (rank : 170) | θ value | 6.88961 (rank : 152) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9BVJ6, Q5JYF1 | Gene names | UTP14A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U3 small nucleolar RNA-associated protein 14 homolog A (Antigen NY-CO- 16). | |||||
|
PHLPP_HUMAN
|
||||||
| NC score | 0.013158 (rank : 171) | θ value | 3.0926 (rank : 92) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 604 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O60346, Q641Q7, Q6P4C4, Q6PJI6, Q86TN6, Q96FK2, Q9NUY1 | Gene names | PHLPP, KIAA0606, PLEKHE1, SCOP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH domain leucine-rich repeat-containing protein phosphatase (EC 3.1.3.16) (PH domain leucine-rich repeat protein phosphatase) (Pleckstrin homology domain-containing family E protein 1) (Suprachiasmatic nucleus circadian oscillatory protein) (hSCOP). | |||||
|
M3K9_HUMAN
|
||||||
| NC score | 0.012799 (rank : 172) | θ value | 0.0252991 (rank : 8) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1123 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P80192, Q6EH31, Q9H2N5 | Gene names | MAP3K9, MLK1, PRKE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase 9 (EC 2.7.11.25) (Mixed lineage kinase 1). | |||||
|
RSPO1_MOUSE
|
||||||
| NC score | 0.012708 (rank : 173) | θ value | 5.27518 (rank : 127) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Z132, Q3V1S3 | Gene names | Rspo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | R-spondin-1 precursor (Roof plate-specific spondin-1) (Cysteine-rich and single thrombospondin domain-containing protein 3) (Cristin-3) (mCristin-3). | |||||
|
RSNL2_MOUSE
|
||||||
| NC score | 0.012651 (rank : 174) | θ value | 6.88961 (rank : 149) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CI96, Q8BW09, Q921Q4, Q9D2S6 | Gene names | Rsnl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Restin-like protein 2. | |||||
|
JADE3_MOUSE
|
||||||
| NC score | 0.012400 (rank : 175) | θ value | 4.03905 (rank : 105) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q6IE82, Q6ZQG2 | Gene names | Phf16, Jade3, Kiaa0215 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-3 (PHD finger protein 16). | |||||
|
PSD4_MOUSE
|
||||||
| NC score | 0.012398 (rank : 176) | θ value | 5.27518 (rank : 122) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8BLR5, Q3TE33, Q3TQF5, Q3UD59, Q3UFH9, Q80V44 | Gene names | Psd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PH and SEC7 domain-containing protein 4 (Pleckstrin homology and SEC7 domain-containing protein 4). | |||||
|
WD51A_HUMAN
|
||||||
| NC score | 0.012024 (rank : 177) | θ value | 8.99809 (rank : 175) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8NBT0, Q2TAK6, Q96IK6, Q9UFJ8 | Gene names | WDR51A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 51A. | |||||
|
SIGL6_HUMAN
|
||||||
| NC score | 0.011728 (rank : 178) | θ value | 4.03905 (rank : 109) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O43699, O15388, O43700 | Gene names | SIGLEC6, CD33L, CD33L1, OBBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sialic acid-binding Ig-like lectin 6 precursor (Siglec-6) (Obesity- binding protein 1) (OB-BP1) (CD33 antigen-like 1) (CDw327 antigen). | |||||
|
WDR69_HUMAN
|
||||||
| NC score | 0.010237 (rank : 179) | θ value | 5.27518 (rank : 133) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 324 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8N136, Q6ZRY1, Q8N776 | Gene names | WDR69 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 69. | |||||
|
SOS2_HUMAN
|
||||||
| NC score | 0.009771 (rank : 180) | θ value | 8.99809 (rank : 168) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q07890, Q15503 | Gene names | SOS2 | |||
|
Domain Architecture |

|
|||||
| Description | Son of sevenless homolog 2 (SOS-2). | |||||
|
PTPRN_MOUSE
|
||||||
| NC score | 0.009565 (rank : 181) | θ value | 4.03905 (rank : 107) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q60673, Q62129 | Gene names | Ptprn, Ptp35 | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase-like N precursor (R-PTP-N) (PTP IA-2). | |||||
|
UROL1_HUMAN
|
||||||
| NC score | 0.009463 (rank : 182) | θ value | 8.99809 (rank : 174) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5DID0, Q5DIC9, Q6LA40, Q6LA41, Q8N216 | Gene names | UMODL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uromodulin-like 1 precursor (Olfactorin). | |||||
|
IQGA1_MOUSE
|
||||||
| NC score | 0.009460 (rank : 183) | θ value | 6.88961 (rank : 144) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9JKF1 | Gene names | Iqgap1 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP1. | |||||
|
RON_MOUSE
|
||||||
| NC score | 0.007679 (rank : 184) | θ value | 0.813845 (rank : 49) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 836 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q62190, Q62555 | Gene names | Mst1r, Ron, Stk | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage-stimulating protein receptor precursor (EC 2.7.10.1) (MSP receptor) (p185-Ron) (Stem cell-derived tyrosine kinase) (CDw136 antigen) [Contains: Macrophage-stimulating protein receptor alpha chain; Macrophage-stimulating protein receptor beta chain]. | |||||
|
LIRB3_HUMAN
|
||||||
| NC score | 0.007470 (rank : 185) | θ value | 5.27518 (rank : 119) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O75022, O15471, Q86U49 | Gene names | LILRB3, ILT5, LIR3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leukocyte immunoglobulin-like receptor subfamily B member 3 precursor (Leukocyte immunoglobulin-like receptor 3) (LIR-3) (Immunoglobulin- like transcript 5) (ILT-5) (Monocyte inhibitory receptor HL9) (CD85a antigen). | |||||
|
PTN12_HUMAN
|
||||||
| NC score | 0.007184 (rank : 186) | θ value | 5.27518 (rank : 123) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q05209, Q16130, Q59FD6, Q75MN8, Q86XU4 | Gene names | PTPN12 | |||
|
Domain Architecture |
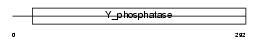
|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 12 (EC 3.1.3.48) (Protein-tyrosine phosphatase G1) (PTPG1). | |||||
|
FOXI1_MOUSE
|
||||||
| NC score | 0.006914 (rank : 187) | θ value | 4.03905 (rank : 103) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q922I5, Q5SRI5, Q9D299 | Gene names | Foxi1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein I1. | |||||
|
KSR1_MOUSE
|
||||||
| NC score | 0.005427 (rank : 188) | θ value | 8.99809 (rank : 161) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q61097, Q61648, Q78DX8 | Gene names | Ksr1, Ksr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinase suppressor of ras-1 (Kinase suppressor of ras) (mKSR1) (Hb protein). | |||||
|
M3K12_HUMAN
|
||||||
| NC score | 0.005263 (rank : 189) | θ value | 5.27518 (rank : 120) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 940 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q12852, Q86VQ5, Q8WY25 | Gene names | MAP3K12, ZPK | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase 12 (EC 2.7.11.25) (Mixed lineage kinase) (Leucine-zipper protein kinase) (ZPK) (Dual leucine zipper bearing kinase) (DLK) (MAPK-upstream kinase) (MUK). | |||||
|
ZN592_HUMAN
|
||||||
| NC score | 0.004818 (rank : 190) | θ value | 5.27518 (rank : 134) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 762 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q92610 | Gene names | ZNF592, KIAA0211 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 592. | |||||
|
ADA19_MOUSE
|
||||||
| NC score | 0.003915 (rank : 191) | θ value | 6.88961 (rank : 135) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O35674 | Gene names | Adam19, Mltnb | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 19 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 19) (Meltrin beta). | |||||
|
BTK_MOUSE
|
||||||
| NC score | 0.003605 (rank : 192) | θ value | 8.99809 (rank : 155) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1017 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P35991, Q61365 | Gene names | Btk, Bpk | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein kinase BTK (EC 2.7.10.2) (Bruton tyrosine kinase) (Agammaglobulinaemia tyrosine kinase) (ATK) (B cell progenitor kinase) (BPK) (Kinase EMB). | |||||
|
BTK_HUMAN
|
||||||
| NC score | 0.003546 (rank : 193) | θ value | 8.99809 (rank : 154) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 1026 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q06187 | Gene names | BTK, AGMX1, ATK, BPK | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein kinase BTK (EC 2.7.10.2) (Bruton tyrosine kinase) (Agammaglobulinaemia tyrosine kinase) (ATK) (B cell progenitor kinase) (BPK). | |||||
|
DSG2_MOUSE
|
||||||
| NC score | 0.002739 (rank : 194) | θ value | 6.88961 (rank : 142) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O55111, Q8K069, Q8R517 | Gene names | Dsg2 | |||
|
Domain Architecture |

|
|||||
| Description | Desmoglein-2 precursor. | |||||
|
KLK2_HUMAN
|
||||||
| NC score | 0.002684 (rank : 195) | θ value | 4.03905 (rank : 106) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P20151, Q15946, Q9UJZ9 | Gene names | KLK2 | |||
|
Domain Architecture |

|
|||||
| Description | Kallikrein-2 precursor (EC 3.4.21.35) (Tissue kallikrein-2) (Glandular kallikrein-1) (hGK-1). | |||||
|
ZN406_MOUSE
|
||||||
| NC score | 0.002375 (rank : 196) | θ value | 3.0926 (rank : 98) | |||
| Query Neighborhood Hits | 176 | Target Neighborhood Hits | 800 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q7TS63 | Gene names | Znf406, Zfp406 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 406. | |||||