Please be patient as the page loads
|
CABL1_MOUSE
|
||||||
| SwissProt Accessions | Q9ESJ1, Q9EPR8 | Gene names | Cables1, Cables | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CDK5 and ABL1 enzyme substrate 1 (Interactor with CDK3 1) (Ik3-1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
CABL1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.984846 (rank : 2) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8TDN4, Q8N3Y8, Q8NA22, Q9BTG1 | Gene names | CABLES1, CABLES | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CDK5 and ABL1 enzyme substrate 1 (Interactor with CDK3 1) (Ik3-1). | |||||
|
CABL1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9ESJ1, Q9EPR8 | Gene names | Cables1, Cables | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CDK5 and ABL1 enzyme substrate 1 (Interactor with CDK3 1) (Ik3-1). | |||||
|
CABL2_HUMAN
|
||||||
| θ value | 2.22025e-107 (rank : 3) | NC score | 0.944199 (rank : 4) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BTV7, Q5JWL0, Q9BYK0 | Gene names | CABLES2, C20orf150 | |||
|
Domain Architecture |

|
|||||
| Description | CDK5 and ABL1 enzyme substrate 2 (Interactor with CDK3 2) (Ik3-2). | |||||
|
CABL2_MOUSE
|
||||||
| θ value | 9.32813e-106 (rank : 4) | NC score | 0.950447 (rank : 3) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8K3M5 | Gene names | Cables2 | |||
|
Domain Architecture |

|
|||||
| Description | CDK5 and ABL1 enzyme substrate 2 (Interactor with CDK3 2) (Ik3-2). | |||||
|
IRS2_MOUSE
|
||||||
| θ value | 0.21417 (rank : 5) | NC score | 0.050326 (rank : 8) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P81122 | Gene names | Irs2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Insulin receptor substrate 2 (IRS-2) (4PS). | |||||
|
CCNT2_HUMAN
|
||||||
| θ value | 0.62314 (rank : 6) | NC score | 0.068007 (rank : 6) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O60583, O60582, Q5I1Y0 | Gene names | CCNT2 | |||
|
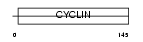
Domain Architecture |
|
|||||
| Description | Cyclin-T2 (CycT2). | |||||
|
IRS2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 7) | NC score | 0.043993 (rank : 9) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y4H2, Q96RR2, Q9BZG0, Q9Y6I5 | Gene names | IRS2 | |||
|
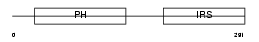
Domain Architecture |
|
|||||
| Description | Insulin receptor substrate 2 (IRS-2). | |||||
|
K1543_HUMAN
|
||||||
| θ value | 1.38821 (rank : 8) | NC score | 0.026751 (rank : 12) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9P1Y5, Q8NDF1 | Gene names | KIAA1543 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1543. | |||||
|
ASPP2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 9) | NC score | 0.013781 (rank : 19) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1156 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q13625, Q12892, Q86X75, Q96KQ3 | Gene names | TP53BP2, ASPP2, BBP | |||
|
Domain Architecture |

|
|||||
| Description | Apoptosis-stimulating of p53 protein 2 (Tumor suppressor p53-binding protein 2) (p53-binding protein 2) (p53BP2) (53BP2) (Bcl2-binding protein) (Bbp) (NY-REN-51 antigen). | |||||
|
CCNC_HUMAN
|
||||||
| θ value | 1.81305 (rank : 10) | NC score | 0.069716 (rank : 5) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P24863, Q9H543 | Gene names | CCNC | |||
|
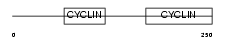
Domain Architecture |
|
|||||
| Description | Cyclin-C. | |||||
|
CCNC_MOUSE
|
||||||
| θ value | 1.81305 (rank : 11) | NC score | 0.067822 (rank : 7) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q62447, Q9WUZ4 | Gene names | Ccnc | |||
|
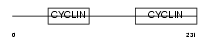
Domain Architecture |
|
|||||
| Description | Cyclin-C. | |||||
|
ZN385_HUMAN
|
||||||
| θ value | 1.81305 (rank : 12) | NC score | 0.019774 (rank : 16) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 614 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96PM9, Q5VH53, Q9H7R6, Q9UFU3 | Gene names | ZNF385, HZF, RZF | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 385 (Hematopoietic zinc finger protein) (Retinal zinc finger protein). | |||||
|
ZN541_HUMAN
|
||||||
| θ value | 1.81305 (rank : 13) | NC score | 0.027299 (rank : 11) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H0D2, Q8NDK8 | Gene names | ZNF541 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 541. | |||||
|
LRP12_HUMAN
|
||||||
| θ value | 2.36792 (rank : 14) | NC score | 0.011040 (rank : 24) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y561 | Gene names | LRP12, ST7 | |||
|
Domain Architecture |
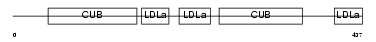
|
|||||
| Description | Low-density lipoprotein receptor-related protein 12 precursor (Suppressor of tumorigenicity protein 7). | |||||
|
PTRF_HUMAN
|
||||||
| θ value | 2.36792 (rank : 15) | NC score | 0.021441 (rank : 14) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6NZI2, O00535, Q6GMY1, Q96H74, Q9BT85, Q9HAP4 | Gene names | PTRF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polymerase I and transcript release factor (PTRF protein). | |||||
|
SPEG_HUMAN
|
||||||
| θ value | 2.36792 (rank : 16) | NC score | 0.004000 (rank : 28) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q15772, Q27J74, Q695L1, Q6FGA6, Q6ZQW1, Q6ZTL8, Q9P2P9 | Gene names | SPEG, APEG1, KIAA1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle preferentially expressed protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
AHTF1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 17) | NC score | 0.021021 (rank : 15) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CJF7, Q8BVJ5, Q8VD55 | Gene names | Ahctf1, Elys | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-hook-containing transcription factor 1 (Embryonic large molecule derived from yolk sac). | |||||
|
B4GT1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 18) | NC score | 0.011133 (rank : 23) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P15291, Q12909, Q12910, Q12911, Q14456, Q14509, Q14523 | Gene names | B4GALT1, GGTB2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-1,4-galactosyltransferase 1 (EC 2.4.1.-) (Beta-1,4-GalTase 1) (Beta4Gal-T1) (b4Gal-T1) (UDP-galactose:beta-N-acetylglucosamine beta- 1,4-galactosyltransferase 1) (UDP-Gal:beta-GlcNAc beta-1,4- galactosyltransferase 1) [Includes: Lactose synthase A protein (EC 2.4.1.22); N-acetyllactosamine synthase (EC 2.4.1.90) (Nal synthetase); Beta-N-acetylglucosaminylglycopeptide beta-1,4- galactosyltransferase (EC 2.4.1.38); Beta-N-acetylglucosaminyl- glycolipid beta-1,4-galactosyltransferase (EC 2.4.1.-)]. | |||||
|
SAPS1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 19) | NC score | 0.024273 (rank : 13) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UPN7, Q504V2, Q6NVJ6, Q9BU97 | Gene names | SAPS1, KIAA1115 | |||
|
Domain Architecture |

|
|||||
| Description | SAPS domain family member 1. | |||||
|
GRIPE_MOUSE
|
||||||
| θ value | 5.27518 (rank : 20) | NC score | 0.014762 (rank : 18) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6GYP7, Q6GYP6, Q6ZQ28, Q8BNJ5, Q8C9G8, Q8CIW4, Q9JMC4 | Gene names | Garnl1, Kiaa0884, Tulip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GTPase-activating RapGAP domain-like 1 (GAP-related-interacting partner to E12) (GRIPE) (Tuberin-like protein 1). | |||||
|
PELP1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 21) | NC score | 0.015337 (rank : 17) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9DBD5, Q5F2E2, Q6PEM0, Q91YM9 | Gene names | Pelp1, Mnar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor). | |||||
|
ZN217_HUMAN
|
||||||
| θ value | 5.27518 (rank : 22) | NC score | 0.003073 (rank : 29) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 905 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O75362 | Gene names | ZNF217, ZABC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 217. | |||||
|
BCAR3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 23) | NC score | 0.009531 (rank : 25) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O75815, Q6UW40, Q9BR50 | Gene names | BCAR3, NSP2, SH2D3B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Breast cancer anti-estrogen resistance protein 3 (SH2 domain- containing protein 3B) (Novel SH2-containing protein 2). | |||||
|
DGCR8_HUMAN
|
||||||
| θ value | 6.88961 (rank : 24) | NC score | 0.031349 (rank : 10) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8WYQ5, Q6DCB2, Q6MZE9, Q6Y2L0, Q96G39, Q96GP8, Q9H6L8, Q9H6T7, Q9NRW2 | Gene names | DGCR8, C22orf12, DGCRK6 | |||
|
Domain Architecture |
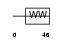
|
|||||
| Description | DGCR8 protein (DiGeorge syndrome critical region 8). | |||||
|
PKN3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 25) | NC score | 0.000866 (rank : 31) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1009 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8K045 | Gene names | Pkn3, Pknbeta | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase N3 (EC 2.7.11.13) (Protein kinase PKN- beta) (Protein-kinase C-related kinase 3). | |||||
|
TECT1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 26) | NC score | 0.013722 (rank : 20) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BZ64, Q3UMP0, Q8BUE2 | Gene names | Tect1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tectonic-1 precursor. | |||||
|
ZN516_MOUSE
|
||||||
| θ value | 6.88961 (rank : 27) | NC score | 0.002882 (rank : 30) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1197 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q7TSH3 | Gene names | Znf516, Kiaa0222, Zfp516 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 516. | |||||
|
MLL3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 28) | NC score | 0.005722 (rank : 27) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
PITM3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 29) | NC score | 0.007556 (rank : 26) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9BZ71, Q59GH9, Q9NPQ4 | Gene names | PITPNM3, NIR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated phosphatidylinositol transfer protein 3 (Phosphatidylinositol transfer protein, membrane-associated 3) (Pyk2 N-terminal domain-interacting receptor 1) (NIR-1). | |||||
|
TF2H1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 30) | NC score | 0.012738 (rank : 21) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P32780, Q9H5K5, Q9NQD9 | Gene names | GTF2H1, BTF2 | |||
|
Domain Architecture |
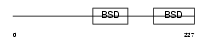
|
|||||
| Description | TFIIH basal transcription factor complex p62 subunit (Basic transcription factor 62 kDa subunit) (BTF2-p62) (General transcription factor IIH polypeptide 1). | |||||
|
XRCC1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 31) | NC score | 0.011209 (rank : 22) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q60596, Q3THC5, Q5U435, Q7TNQ5 | Gene names | Xrcc1, Xrcc-1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-repair protein XRCC1 (X-ray repair cross-complementing protein 1). | |||||
|
CABL1_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9ESJ1, Q9EPR8 | Gene names | Cables1, Cables | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CDK5 and ABL1 enzyme substrate 1 (Interactor with CDK3 1) (Ik3-1). | |||||
|
CABL1_HUMAN
|
||||||
| NC score | 0.984846 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8TDN4, Q8N3Y8, Q8NA22, Q9BTG1 | Gene names | CABLES1, CABLES | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CDK5 and ABL1 enzyme substrate 1 (Interactor with CDK3 1) (Ik3-1). | |||||
|
CABL2_MOUSE
|
||||||
| NC score | 0.950447 (rank : 3) | θ value | 9.32813e-106 (rank : 4) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8K3M5 | Gene names | Cables2 | |||
|
Domain Architecture |

|
|||||
| Description | CDK5 and ABL1 enzyme substrate 2 (Interactor with CDK3 2) (Ik3-2). | |||||
|
CABL2_HUMAN
|
||||||
| NC score | 0.944199 (rank : 4) | θ value | 2.22025e-107 (rank : 3) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BTV7, Q5JWL0, Q9BYK0 | Gene names | CABLES2, C20orf150 | |||
|
Domain Architecture |

|
|||||
| Description | CDK5 and ABL1 enzyme substrate 2 (Interactor with CDK3 2) (Ik3-2). | |||||
|
CCNC_HUMAN
|
||||||
| NC score | 0.069716 (rank : 5) | θ value | 1.81305 (rank : 10) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P24863, Q9H543 | Gene names | CCNC | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-C. | |||||
|
CCNT2_HUMAN
|
||||||
| NC score | 0.068007 (rank : 6) | θ value | 0.62314 (rank : 6) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O60583, O60582, Q5I1Y0 | Gene names | CCNT2 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-T2 (CycT2). | |||||
|
CCNC_MOUSE
|
||||||
| NC score | 0.067822 (rank : 7) | θ value | 1.81305 (rank : 11) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q62447, Q9WUZ4 | Gene names | Ccnc | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-C. | |||||
|
IRS2_MOUSE
|
||||||
| NC score | 0.050326 (rank : 8) | θ value | 0.21417 (rank : 5) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P81122 | Gene names | Irs2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Insulin receptor substrate 2 (IRS-2) (4PS). | |||||
|
IRS2_HUMAN
|
||||||
| NC score | 0.043993 (rank : 9) | θ value | 1.38821 (rank : 7) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y4H2, Q96RR2, Q9BZG0, Q9Y6I5 | Gene names | IRS2 | |||
|
Domain Architecture |

|
|||||
| Description | Insulin receptor substrate 2 (IRS-2). | |||||
|
DGCR8_HUMAN
|
||||||
| NC score | 0.031349 (rank : 10) | θ value | 6.88961 (rank : 24) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8WYQ5, Q6DCB2, Q6MZE9, Q6Y2L0, Q96G39, Q96GP8, Q9H6L8, Q9H6T7, Q9NRW2 | Gene names | DGCR8, C22orf12, DGCRK6 | |||
|
Domain Architecture |

|
|||||
| Description | DGCR8 protein (DiGeorge syndrome critical region 8). | |||||
|
ZN541_HUMAN
|
||||||
| NC score | 0.027299 (rank : 11) | θ value | 1.81305 (rank : 13) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H0D2, Q8NDK8 | Gene names | ZNF541 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 541. | |||||
|
K1543_HUMAN
|
||||||
| NC score | 0.026751 (rank : 12) | θ value | 1.38821 (rank : 8) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9P1Y5, Q8NDF1 | Gene names | KIAA1543 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1543. | |||||
|
SAPS1_HUMAN
|
||||||
| NC score | 0.024273 (rank : 13) | θ value | 4.03905 (rank : 19) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UPN7, Q504V2, Q6NVJ6, Q9BU97 | Gene names | SAPS1, KIAA1115 | |||
|
Domain Architecture |

|
|||||
| Description | SAPS domain family member 1. | |||||
|
PTRF_HUMAN
|
||||||
| NC score | 0.021441 (rank : 14) | θ value | 2.36792 (rank : 15) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6NZI2, O00535, Q6GMY1, Q96H74, Q9BT85, Q9HAP4 | Gene names | PTRF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polymerase I and transcript release factor (PTRF protein). | |||||
|
AHTF1_MOUSE
|
||||||
| NC score | 0.021021 (rank : 15) | θ value | 3.0926 (rank : 17) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CJF7, Q8BVJ5, Q8VD55 | Gene names | Ahctf1, Elys | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-hook-containing transcription factor 1 (Embryonic large molecule derived from yolk sac). | |||||
|
ZN385_HUMAN
|
||||||
| NC score | 0.019774 (rank : 16) | θ value | 1.81305 (rank : 12) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 614 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96PM9, Q5VH53, Q9H7R6, Q9UFU3 | Gene names | ZNF385, HZF, RZF | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 385 (Hematopoietic zinc finger protein) (Retinal zinc finger protein). | |||||
|
PELP1_MOUSE
|
||||||
| NC score | 0.015337 (rank : 17) | θ value | 5.27518 (rank : 21) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9DBD5, Q5F2E2, Q6PEM0, Q91YM9 | Gene names | Pelp1, Mnar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor). | |||||
|
GRIPE_MOUSE
|
||||||
| NC score | 0.014762 (rank : 18) | θ value | 5.27518 (rank : 20) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6GYP7, Q6GYP6, Q6ZQ28, Q8BNJ5, Q8C9G8, Q8CIW4, Q9JMC4 | Gene names | Garnl1, Kiaa0884, Tulip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GTPase-activating RapGAP domain-like 1 (GAP-related-interacting partner to E12) (GRIPE) (Tuberin-like protein 1). | |||||
|
ASPP2_HUMAN
|
||||||
| NC score | 0.013781 (rank : 19) | θ value | 1.81305 (rank : 9) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1156 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q13625, Q12892, Q86X75, Q96KQ3 | Gene names | TP53BP2, ASPP2, BBP | |||
|
Domain Architecture |

|
|||||
| Description | Apoptosis-stimulating of p53 protein 2 (Tumor suppressor p53-binding protein 2) (p53-binding protein 2) (p53BP2) (53BP2) (Bcl2-binding protein) (Bbp) (NY-REN-51 antigen). | |||||
|
TECT1_MOUSE
|
||||||
| NC score | 0.013722 (rank : 20) | θ value | 6.88961 (rank : 26) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BZ64, Q3UMP0, Q8BUE2 | Gene names | Tect1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tectonic-1 precursor. | |||||
|
TF2H1_HUMAN
|
||||||
| NC score | 0.012738 (rank : 21) | θ value | 8.99809 (rank : 30) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P32780, Q9H5K5, Q9NQD9 | Gene names | GTF2H1, BTF2 | |||
|
Domain Architecture |

|
|||||
| Description | TFIIH basal transcription factor complex p62 subunit (Basic transcription factor 62 kDa subunit) (BTF2-p62) (General transcription factor IIH polypeptide 1). | |||||
|
XRCC1_MOUSE
|
||||||
| NC score | 0.011209 (rank : 22) | θ value | 8.99809 (rank : 31) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q60596, Q3THC5, Q5U435, Q7TNQ5 | Gene names | Xrcc1, Xrcc-1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-repair protein XRCC1 (X-ray repair cross-complementing protein 1). | |||||
|
B4GT1_HUMAN
|
||||||
| NC score | 0.011133 (rank : 23) | θ value | 4.03905 (rank : 18) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P15291, Q12909, Q12910, Q12911, Q14456, Q14509, Q14523 | Gene names | B4GALT1, GGTB2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-1,4-galactosyltransferase 1 (EC 2.4.1.-) (Beta-1,4-GalTase 1) (Beta4Gal-T1) (b4Gal-T1) (UDP-galactose:beta-N-acetylglucosamine beta- 1,4-galactosyltransferase 1) (UDP-Gal:beta-GlcNAc beta-1,4- galactosyltransferase 1) [Includes: Lactose synthase A protein (EC 2.4.1.22); N-acetyllactosamine synthase (EC 2.4.1.90) (Nal synthetase); Beta-N-acetylglucosaminylglycopeptide beta-1,4- galactosyltransferase (EC 2.4.1.38); Beta-N-acetylglucosaminyl- glycolipid beta-1,4-galactosyltransferase (EC 2.4.1.-)]. | |||||
|
LRP12_HUMAN
|
||||||
| NC score | 0.011040 (rank : 24) | θ value | 2.36792 (rank : 14) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y561 | Gene names | LRP12, ST7 | |||
|
Domain Architecture |

|
|||||
| Description | Low-density lipoprotein receptor-related protein 12 precursor (Suppressor of tumorigenicity protein 7). | |||||
|
BCAR3_HUMAN
|
||||||
| NC score | 0.009531 (rank : 25) | θ value | 6.88961 (rank : 23) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O75815, Q6UW40, Q9BR50 | Gene names | BCAR3, NSP2, SH2D3B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Breast cancer anti-estrogen resistance protein 3 (SH2 domain- containing protein 3B) (Novel SH2-containing protein 2). | |||||
|
PITM3_HUMAN
|
||||||
| NC score | 0.007556 (rank : 26) | θ value | 8.99809 (rank : 29) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9BZ71, Q59GH9, Q9NPQ4 | Gene names | PITPNM3, NIR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated phosphatidylinositol transfer protein 3 (Phosphatidylinositol transfer protein, membrane-associated 3) (Pyk2 N-terminal domain-interacting receptor 1) (NIR-1). | |||||
|
MLL3_HUMAN
|
||||||
| NC score | 0.005722 (rank : 27) | θ value | 8.99809 (rank : 28) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
SPEG_HUMAN
|
||||||
| NC score | 0.004000 (rank : 28) | θ value | 2.36792 (rank : 16) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q15772, Q27J74, Q695L1, Q6FGA6, Q6ZQW1, Q6ZTL8, Q9P2P9 | Gene names | SPEG, APEG1, KIAA1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle preferentially expressed protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
ZN217_HUMAN
|
||||||
| NC score | 0.003073 (rank : 29) | θ value | 5.27518 (rank : 22) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 905 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O75362 | Gene names | ZNF217, ZABC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 217. | |||||
|
ZN516_MOUSE
|
||||||
| NC score | 0.002882 (rank : 30) | θ value | 6.88961 (rank : 27) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1197 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q7TSH3 | Gene names | Znf516, Kiaa0222, Zfp516 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 516. | |||||
|
PKN3_MOUSE
|
||||||
| NC score | 0.000866 (rank : 31) | θ value | 6.88961 (rank : 25) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1009 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8K045 | Gene names | Pkn3, Pknbeta | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase N3 (EC 2.7.11.13) (Protein kinase PKN- beta) (Protein-kinase C-related kinase 3). | |||||