Please be patient as the page loads
|
PITM3_HUMAN
|
||||||
| SwissProt Accessions | Q9BZ71, Q59GH9, Q9NPQ4 | Gene names | PITPNM3, NIR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated phosphatidylinositol transfer protein 3 (Phosphatidylinositol transfer protein, membrane-associated 3) (Pyk2 N-terminal domain-interacting receptor 1) (NIR-1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
PITM1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.886809 (rank : 6) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O00562, Q6T7X3, Q8TBN3, Q9BZ73 | Gene names | PITPNM1, DRES9, NIR2, PITPNM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated phosphatidylinositol transfer protein 1 (Phosphatidylinositol transfer protein, membrane-associated 1) (Pyk2 N-terminal domain-interacting receptor 2) (NIR-2) (Drosophila retinal degeneration B homolog). | |||||
|
PITM1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.896057 (rank : 5) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O35954 | Gene names | Pitpnm1, Dres9, Mpt1, Nir2, Pitpnm | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated phosphatidylinositol transfer protein 1 (Phosphatidylinositol transfer protein, membrane-associated 1) (PITPnm 1) (Mpt-1) (Pyk2 N-terminal domain-interacting receptor 2) (NIR-2) (Drosophila retinal degeneration B homolog 1) (RdgB1). | |||||
|
PITM2_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.898959 (rank : 3) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9BZ72, Q9P271 | Gene names | PITPNM2, KIAA1457, NIR3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated phosphatidylinositol transfer protein 2 (Phosphatidylinositol transfer protein, membrane-associated 2) (Pyk2 N-terminal domain-interacting receptor 3) (NIR-3). | |||||
|
PITM2_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.897424 (rank : 4) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6ZPQ6, Q9R1P5 | Gene names | Pitpnm2, Kiaa1457, Nir3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated phosphatidylinositol transfer protein 2 (Phosphatidylinositol transfer protein, membrane-associated 2) (Pyk2 N-terminal domain-interacting receptor 3) (NIR-3) (Drosophila retinal degeneration B homolog 2) (RdgB2). | |||||
|
PITM3_HUMAN
|
||||||
| θ value | 0 (rank : 5) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9BZ71, Q59GH9, Q9NPQ4 | Gene names | PITPNM3, NIR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated phosphatidylinositol transfer protein 3 (Phosphatidylinositol transfer protein, membrane-associated 3) (Pyk2 N-terminal domain-interacting receptor 1) (NIR-1). | |||||
|
PITM3_MOUSE
|
||||||
| θ value | 0 (rank : 6) | NC score | 0.999278 (rank : 2) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q3UHE1, Q3UH22, Q5RIT9 | Gene names | Pitpnm3, Nir1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated phosphatidylinositol transfer protein 3 (Phosphatidylinositol transfer protein, membrane-associated 3) (Pyk2 N-terminal domain-interacting receptor 1) (NIR-1). | |||||
|
DDHD1_MOUSE
|
||||||
| θ value | 1.29631e-06 (rank : 7) | NC score | 0.392073 (rank : 11) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q80YA3, Q8VEF2 | Gene names | Ddhd1 | |||
|
Domain Architecture |
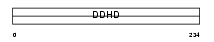
|
|||||
| Description | Probable phospholipase DDHD1 (EC 3.1.1.-) (DDHD domain protein 1) (Phosphatidic acid-preferring phospholipase A1 homolog) (PA-PLA1). | |||||
|
DDHD1_HUMAN
|
||||||
| θ value | 1.69304e-06 (rank : 8) | NC score | 0.376992 (rank : 12) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8NEL9, Q8WVH3, Q96LL2, Q9C0F8 | Gene names | DDHD1, KIAA1705 | |||
|
Domain Architecture |
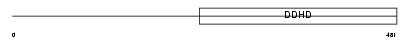
|
|||||
| Description | Probable phospholipase DDHD1 (EC 3.1.1.-) (DDHD domain protein 1) (Phosphatidic acid-preferring phospholipase A1 homolog) (PA-PLA1). | |||||
|
S23IP_MOUSE
|
||||||
| θ value | 4.92598e-06 (rank : 9) | NC score | 0.321608 (rank : 14) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 363 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6NZC7, Q8BXY6 | Gene names | Sec23ip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SEC23-interacting protein. | |||||
|
S23IP_HUMAN
|
||||||
| θ value | 1.43324e-05 (rank : 10) | NC score | 0.329003 (rank : 13) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y6Y8, Q8IXH5, Q9BUK5 | Gene names | SEC23IP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SEC23-interacting protein (p125). | |||||
|
IL1R2_MOUSE
|
||||||
| θ value | 0.813845 (rank : 11) | NC score | 0.031957 (rank : 16) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P27931 | Gene names | Il1r2, Il-1r2, Il1rb | |||
|
Domain Architecture |
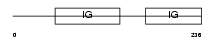
|
|||||
| Description | Interleukin-1 receptor type II precursor (IL-1R-2) (IL-1R-beta) (CD121b antigen). | |||||
|
SDK1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 12) | NC score | 0.011790 (rank : 26) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 495 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q3UH53, Q3UFD1, Q6PAL2, Q6V3A4, Q8BMC2, Q8BZI1 | Gene names | Sdk1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein sidekick-1 precursor. | |||||
|
LPIN3_MOUSE
|
||||||
| θ value | 1.38821 (rank : 13) | NC score | 0.025791 (rank : 19) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99PI4, Q3TQ75, Q8C7R9 | Gene names | Lpin3 | |||
|
Domain Architecture |
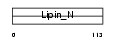
|
|||||
| Description | Lipin-3. | |||||
|
CLAP2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 14) | NC score | 0.014893 (rank : 22) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O75122, Q7L8F6, Q8N6R6, Q9BQT3, Q9BQT4, Q9H7A3, Q9NSZ2 | Gene names | CLASP2, KIAA0627 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CLIP-associating protein 2 (Cytoplasmic linker-associated protein 2). | |||||
|
FOG2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 15) | NC score | 0.009815 (rank : 27) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 731 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8WW38, Q9NPL7, Q9NPS4, Q9UNI5 | Gene names | ZFPM2, FOG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein ZFPM2 (Zinc finger protein multitype 2) (Friend of GATA protein 2) (FOG-2) (hFOG-2). | |||||
|
SDK1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 16) | NC score | 0.009218 (rank : 28) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 511 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7Z5N4, Q8TEN9, Q8TEP5, Q96N44 | Gene names | SDK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein sidekick-1 precursor. | |||||
|
SFR17_HUMAN
|
||||||
| θ value | 2.36792 (rank : 17) | NC score | 0.028420 (rank : 18) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8TF01, Q5T064, Q6P2B4, Q6P2N4, Q6PJ93, Q6PK36, Q7Z640, Q8TF00, Q96K10, Q96SM5, Q9P076, Q9P0C0, Q9Y4N3 | Gene names | SRRP130, C6orf111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 130 (Serine-arginine-rich- splicing regulatory protein 130) (SRrp130) (SR-rich protein) (SR- related protein). | |||||
|
B4GN3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 18) | NC score | 0.018339 (rank : 21) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6L9W6, Q6ZNC1, Q8N7T6 | Gene names | B4GALNT3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N- acetylgalactosaminyltransferase 2 (EC 2.4.1.244) (NGalNAc-T2) (Beta- 1,4-N-acetylgalactosaminyltransferase III) (Beta4GalNAc-T3) (Beta4GalNAcT3). | |||||
|
IRAK1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 19) | NC score | 0.004791 (rank : 33) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 820 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P51617, Q7Z5V4, Q96RL2 | Gene names | IRAK1, IRAK | |||
|
Domain Architecture |
|
|||||
| Description | Interleukin-1 receptor-associated kinase 1 (EC 2.7.11.1) (IRAK-1). | |||||
|
CC036_HUMAN
|
||||||
| θ value | 4.03905 (rank : 20) | NC score | 0.047109 (rank : 15) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q3SXR2, Q3SXR3, Q9H6K8 | Gene names | C3orf36 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C3orf36. | |||||
|
ARHG1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 21) | NC score | 0.012951 (rank : 25) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q61210, O89074, Q80YE8, Q80YE9, Q91VL3 | Gene names | Arhgef1, Lbcl2, Lsc | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 1 (Lymphoid blast crisis-like 2) (Lbc's second cousin). | |||||
|
BRD4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 22) | NC score | 0.009036 (rank : 29) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O60885, O60433, Q96PD3 | Gene names | BRD4, HUNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 4 (HUNK1 protein). | |||||
|
LPIN3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 23) | NC score | 0.019409 (rank : 20) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9BQK8, Q5TDB9, Q9NPY8, Q9UJE5 | Gene names | LPIN3, LIPN3L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipin-3 (Lipin 3-like). | |||||
|
HXD11_MOUSE
|
||||||
| θ value | 6.88961 (rank : 24) | NC score | 0.004289 (rank : 34) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 376 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P23813 | Gene names | Hoxd11, Hox-4.6, Hoxd-11 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-D11 (Hox-4.6) (Hox-5.5). | |||||
|
NCOR2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 25) | NC score | 0.009001 (rank : 30) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y618, O00613, O15416, Q13354, Q9Y5U0 | Gene names | NCOR2, CTG26 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC) (CTG repeat protein 26) (SMAP270). | |||||
|
TTC28_MOUSE
|
||||||
| θ value | 6.88961 (rank : 26) | NC score | 0.007921 (rank : 31) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80XJ3, Q6P9J6, Q80TL5, Q8BV10, Q8C0F2 | Gene names | Ttc28, Kiaa1043 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tetratricopeptide repeat protein 28 (TPR repeat protein 28). | |||||
|
CABL1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 27) | NC score | 0.007556 (rank : 32) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9ESJ1, Q9EPR8 | Gene names | Cables1, Cables | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CDK5 and ABL1 enzyme substrate 1 (Interactor with CDK3 1) (Ik3-1). | |||||
|
REPS2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 28) | NC score | 0.029698 (rank : 17) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8NFH8, O43428, Q8NFI5 | Gene names | REPS2, POB1 | |||
|
Domain Architecture |

|
|||||
| Description | RalBP1-associated Eps domain-containing protein 2 (RalBP1-interacting protein 2) (Partner of RalBP1). | |||||
|
RN103_HUMAN
|
||||||
| θ value | 8.99809 (rank : 29) | NC score | 0.014635 (rank : 24) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O00237, Q8IVB9 | Gene names | RNF103 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 103 (Zinc finger protein 103 homolog) (Zfp-103) (KF-1) (hKF-1). | |||||
|
RN103_MOUSE
|
||||||
| θ value | 8.99809 (rank : 30) | NC score | 0.014642 (rank : 23) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9R1W3, O08670, O08883 | Gene names | Rnf103, Zfp103 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 103 (Zinc finger protein 103) (Zfp-103) (KF-1) (mKF-1). | |||||
|
PIPNA_HUMAN
|
||||||
| θ value | θ > 10 (rank : 31) | NC score | 0.443959 (rank : 8) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q00169 | Gene names | PITPNA, PITPN | |||
|
Domain Architecture |
|
|||||
| Description | Phosphatidylinositol transfer protein alpha isoform (PtdIns transfer protein alpha) (PtdInsTP) (PI-TP-alpha). | |||||
|
PIPNA_MOUSE
|
||||||
| θ value | θ > 10 (rank : 32) | NC score | 0.445791 (rank : 7) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P53810 | Gene names | Pitpna, Pitpn | |||
|
Domain Architecture |
|
|||||
| Description | Phosphatidylinositol transfer protein alpha isoform (PtdIns transfer protein alpha) (PtdInsTP) (PI-TP-alpha). | |||||
|
PIPNB_HUMAN
|
||||||
| θ value | θ > 10 (rank : 33) | NC score | 0.438443 (rank : 10) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P48739, Q8N5W1 | Gene names | PITPNB | |||
|
Domain Architecture |
|
|||||
| Description | Phosphatidylinositol transfer protein beta isoform (PtdIns transfer protein beta) (PtdInsTP) (PI-TP-beta). | |||||
|
PIPNB_MOUSE
|
||||||
| θ value | θ > 10 (rank : 34) | NC score | 0.439566 (rank : 9) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P53811 | Gene names | Pitpnb | |||
|
Domain Architecture |
|
|||||
| Description | Phosphatidylinositol transfer protein beta isoform (PtdIns transfer protein beta) (PtdInsTP) (PI-TP-beta). | |||||
|
PITM3_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 5) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9BZ71, Q59GH9, Q9NPQ4 | Gene names | PITPNM3, NIR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated phosphatidylinositol transfer protein 3 (Phosphatidylinositol transfer protein, membrane-associated 3) (Pyk2 N-terminal domain-interacting receptor 1) (NIR-1). | |||||
|
PITM3_MOUSE
|
||||||
| NC score | 0.999278 (rank : 2) | θ value | 0 (rank : 6) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q3UHE1, Q3UH22, Q5RIT9 | Gene names | Pitpnm3, Nir1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated phosphatidylinositol transfer protein 3 (Phosphatidylinositol transfer protein, membrane-associated 3) (Pyk2 N-terminal domain-interacting receptor 1) (NIR-1). | |||||
|
PITM2_HUMAN
|
||||||
| NC score | 0.898959 (rank : 3) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9BZ72, Q9P271 | Gene names | PITPNM2, KIAA1457, NIR3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated phosphatidylinositol transfer protein 2 (Phosphatidylinositol transfer protein, membrane-associated 2) (Pyk2 N-terminal domain-interacting receptor 3) (NIR-3). | |||||
|
PITM2_MOUSE
|
||||||
| NC score | 0.897424 (rank : 4) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6ZPQ6, Q9R1P5 | Gene names | Pitpnm2, Kiaa1457, Nir3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated phosphatidylinositol transfer protein 2 (Phosphatidylinositol transfer protein, membrane-associated 2) (Pyk2 N-terminal domain-interacting receptor 3) (NIR-3) (Drosophila retinal degeneration B homolog 2) (RdgB2). | |||||
|
PITM1_MOUSE
|
||||||
| NC score | 0.896057 (rank : 5) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O35954 | Gene names | Pitpnm1, Dres9, Mpt1, Nir2, Pitpnm | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated phosphatidylinositol transfer protein 1 (Phosphatidylinositol transfer protein, membrane-associated 1) (PITPnm 1) (Mpt-1) (Pyk2 N-terminal domain-interacting receptor 2) (NIR-2) (Drosophila retinal degeneration B homolog 1) (RdgB1). | |||||
|
PITM1_HUMAN
|
||||||
| NC score | 0.886809 (rank : 6) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O00562, Q6T7X3, Q8TBN3, Q9BZ73 | Gene names | PITPNM1, DRES9, NIR2, PITPNM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated phosphatidylinositol transfer protein 1 (Phosphatidylinositol transfer protein, membrane-associated 1) (Pyk2 N-terminal domain-interacting receptor 2) (NIR-2) (Drosophila retinal degeneration B homolog). | |||||
|
PIPNA_MOUSE
|
||||||
| NC score | 0.445791 (rank : 7) | θ value | θ > 10 (rank : 32) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P53810 | Gene names | Pitpna, Pitpn | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol transfer protein alpha isoform (PtdIns transfer protein alpha) (PtdInsTP) (PI-TP-alpha). | |||||
|
PIPNA_HUMAN
|
||||||
| NC score | 0.443959 (rank : 8) | θ value | θ > 10 (rank : 31) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q00169 | Gene names | PITPNA, PITPN | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol transfer protein alpha isoform (PtdIns transfer protein alpha) (PtdInsTP) (PI-TP-alpha). | |||||
|
PIPNB_MOUSE
|
||||||
| NC score | 0.439566 (rank : 9) | θ value | θ > 10 (rank : 34) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P53811 | Gene names | Pitpnb | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol transfer protein beta isoform (PtdIns transfer protein beta) (PtdInsTP) (PI-TP-beta). | |||||
|
PIPNB_HUMAN
|
||||||
| NC score | 0.438443 (rank : 10) | θ value | θ > 10 (rank : 33) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P48739, Q8N5W1 | Gene names | PITPNB | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol transfer protein beta isoform (PtdIns transfer protein beta) (PtdInsTP) (PI-TP-beta). | |||||
|
DDHD1_MOUSE
|
||||||
| NC score | 0.392073 (rank : 11) | θ value | 1.29631e-06 (rank : 7) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q80YA3, Q8VEF2 | Gene names | Ddhd1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipase DDHD1 (EC 3.1.1.-) (DDHD domain protein 1) (Phosphatidic acid-preferring phospholipase A1 homolog) (PA-PLA1). | |||||
|
DDHD1_HUMAN
|
||||||
| NC score | 0.376992 (rank : 12) | θ value | 1.69304e-06 (rank : 8) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8NEL9, Q8WVH3, Q96LL2, Q9C0F8 | Gene names | DDHD1, KIAA1705 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phospholipase DDHD1 (EC 3.1.1.-) (DDHD domain protein 1) (Phosphatidic acid-preferring phospholipase A1 homolog) (PA-PLA1). | |||||
|
S23IP_HUMAN
|
||||||
| NC score | 0.329003 (rank : 13) | θ value | 1.43324e-05 (rank : 10) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y6Y8, Q8IXH5, Q9BUK5 | Gene names | SEC23IP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SEC23-interacting protein (p125). | |||||
|
S23IP_MOUSE
|
||||||
| NC score | 0.321608 (rank : 14) | θ value | 4.92598e-06 (rank : 9) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 363 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6NZC7, Q8BXY6 | Gene names | Sec23ip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SEC23-interacting protein. | |||||
|
CC036_HUMAN
|
||||||
| NC score | 0.047109 (rank : 15) | θ value | 4.03905 (rank : 20) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q3SXR2, Q3SXR3, Q9H6K8 | Gene names | C3orf36 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C3orf36. | |||||
|
IL1R2_MOUSE
|
||||||
| NC score | 0.031957 (rank : 16) | θ value | 0.813845 (rank : 11) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P27931 | Gene names | Il1r2, Il-1r2, Il1rb | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin-1 receptor type II precursor (IL-1R-2) (IL-1R-beta) (CD121b antigen). | |||||
|
REPS2_HUMAN
|
||||||
| NC score | 0.029698 (rank : 17) | θ value | 8.99809 (rank : 28) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8NFH8, O43428, Q8NFI5 | Gene names | REPS2, POB1 | |||
|
Domain Architecture |

|
|||||
| Description | RalBP1-associated Eps domain-containing protein 2 (RalBP1-interacting protein 2) (Partner of RalBP1). | |||||
|
SFR17_HUMAN
|
||||||
| NC score | 0.028420 (rank : 18) | θ value | 2.36792 (rank : 17) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8TF01, Q5T064, Q6P2B4, Q6P2N4, Q6PJ93, Q6PK36, Q7Z640, Q8TF00, Q96K10, Q96SM5, Q9P076, Q9P0C0, Q9Y4N3 | Gene names | SRRP130, C6orf111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 130 (Serine-arginine-rich- splicing regulatory protein 130) (SRrp130) (SR-rich protein) (SR- related protein). | |||||
|
LPIN3_MOUSE
|
||||||
| NC score | 0.025791 (rank : 19) | θ value | 1.38821 (rank : 13) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99PI4, Q3TQ75, Q8C7R9 | Gene names | Lpin3 | |||
|
Domain Architecture |

|
|||||
| Description | Lipin-3. | |||||
|
LPIN3_HUMAN
|
||||||
| NC score | 0.019409 (rank : 20) | θ value | 5.27518 (rank : 23) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9BQK8, Q5TDB9, Q9NPY8, Q9UJE5 | Gene names | LPIN3, LIPN3L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipin-3 (Lipin 3-like). | |||||
|
B4GN3_HUMAN
|
||||||
| NC score | 0.018339 (rank : 21) | θ value | 3.0926 (rank : 18) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6L9W6, Q6ZNC1, Q8N7T6 | Gene names | B4GALNT3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N- acetylgalactosaminyltransferase 2 (EC 2.4.1.244) (NGalNAc-T2) (Beta- 1,4-N-acetylgalactosaminyltransferase III) (Beta4GalNAc-T3) (Beta4GalNAcT3). | |||||
|
CLAP2_HUMAN
|
||||||
| NC score | 0.014893 (rank : 22) | θ value | 2.36792 (rank : 14) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O75122, Q7L8F6, Q8N6R6, Q9BQT3, Q9BQT4, Q9H7A3, Q9NSZ2 | Gene names | CLASP2, KIAA0627 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CLIP-associating protein 2 (Cytoplasmic linker-associated protein 2). | |||||
|
RN103_MOUSE
|
||||||
| NC score | 0.014642 (rank : 23) | θ value | 8.99809 (rank : 30) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9R1W3, O08670, O08883 | Gene names | Rnf103, Zfp103 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 103 (Zinc finger protein 103) (Zfp-103) (KF-1) (mKF-1). | |||||
|
RN103_HUMAN
|
||||||
| NC score | 0.014635 (rank : 24) | θ value | 8.99809 (rank : 29) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O00237, Q8IVB9 | Gene names | RNF103 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 103 (Zinc finger protein 103 homolog) (Zfp-103) (KF-1) (hKF-1). | |||||
|
ARHG1_MOUSE
|
||||||
| NC score | 0.012951 (rank : 25) | θ value | 5.27518 (rank : 21) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q61210, O89074, Q80YE8, Q80YE9, Q91VL3 | Gene names | Arhgef1, Lbcl2, Lsc | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 1 (Lymphoid blast crisis-like 2) (Lbc's second cousin). | |||||
|
SDK1_MOUSE
|
||||||
| NC score | 0.011790 (rank : 26) | θ value | 1.06291 (rank : 12) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 495 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q3UH53, Q3UFD1, Q6PAL2, Q6V3A4, Q8BMC2, Q8BZI1 | Gene names | Sdk1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein sidekick-1 precursor. | |||||
|
FOG2_HUMAN
|
||||||
| NC score | 0.009815 (rank : 27) | θ value | 2.36792 (rank : 15) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 731 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8WW38, Q9NPL7, Q9NPS4, Q9UNI5 | Gene names | ZFPM2, FOG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein ZFPM2 (Zinc finger protein multitype 2) (Friend of GATA protein 2) (FOG-2) (hFOG-2). | |||||
|
SDK1_HUMAN
|
||||||
| NC score | 0.009218 (rank : 28) | θ value | 2.36792 (rank : 16) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 511 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7Z5N4, Q8TEN9, Q8TEP5, Q96N44 | Gene names | SDK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein sidekick-1 precursor. | |||||
|
BRD4_HUMAN
|
||||||
| NC score | 0.009036 (rank : 29) | θ value | 5.27518 (rank : 22) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O60885, O60433, Q96PD3 | Gene names | BRD4, HUNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 4 (HUNK1 protein). | |||||
|
NCOR2_HUMAN
|
||||||
| NC score | 0.009001 (rank : 30) | θ value | 6.88961 (rank : 25) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y618, O00613, O15416, Q13354, Q9Y5U0 | Gene names | NCOR2, CTG26 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC) (CTG repeat protein 26) (SMAP270). | |||||
|
TTC28_MOUSE
|
||||||
| NC score | 0.007921 (rank : 31) | θ value | 6.88961 (rank : 26) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80XJ3, Q6P9J6, Q80TL5, Q8BV10, Q8C0F2 | Gene names | Ttc28, Kiaa1043 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tetratricopeptide repeat protein 28 (TPR repeat protein 28). | |||||
|
CABL1_MOUSE
|
||||||
| NC score | 0.007556 (rank : 32) | θ value | 8.99809 (rank : 27) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9ESJ1, Q9EPR8 | Gene names | Cables1, Cables | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CDK5 and ABL1 enzyme substrate 1 (Interactor with CDK3 1) (Ik3-1). | |||||
|
IRAK1_HUMAN
|
||||||
| NC score | 0.004791 (rank : 33) | θ value | 3.0926 (rank : 19) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 820 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P51617, Q7Z5V4, Q96RL2 | Gene names | IRAK1, IRAK | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin-1 receptor-associated kinase 1 (EC 2.7.11.1) (IRAK-1). | |||||
|
HXD11_MOUSE
|
||||||
| NC score | 0.004289 (rank : 34) | θ value | 6.88961 (rank : 24) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 376 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P23813 | Gene names | Hoxd11, Hox-4.6, Hoxd-11 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-D11 (Hox-4.6) (Hox-5.5). | |||||