Please be patient as the page loads
|
B4GN3_HUMAN
|
||||||
| SwissProt Accessions | Q6L9W6, Q6ZNC1, Q8N7T6 | Gene names | B4GALNT3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N- acetylgalactosaminyltransferase 2 (EC 2.4.1.244) (NGalNAc-T2) (Beta- 1,4-N-acetylgalactosaminyltransferase III) (Beta4GalNAc-T3) (Beta4GalNAcT3). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
B4GN3_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 128 | |
| SwissProt Accessions | Q6L9W6, Q6ZNC1, Q8N7T6 | Gene names | B4GALNT3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N- acetylgalactosaminyltransferase 2 (EC 2.4.1.244) (NGalNAc-T2) (Beta- 1,4-N-acetylgalactosaminyltransferase III) (Beta4GalNAc-T3) (Beta4GalNAcT3). | |||||
|
B4GN3_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.977199 (rank : 2) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6L8S8 | Gene names | B4galnt3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N- acetylgalactosaminyltransferase 2 (EC 2.4.1.244) (NGalNAc-T2) (Beta- 1,4-N-acetylgalactosaminyltransferase III) (Beta4GalNAc-T3) (Beta4GalNAcT3). | |||||
|
B4GN4_MOUSE
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.931433 (rank : 3) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q766D5, Q8CHV2 | Gene names | B4galnt4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N- acetylgalactosaminyltransferase 1 (EC 2.4.1.244) (NGalNAc-T1) (Beta- 1,4-N-acetylgalactosaminyltransferase IV) (Beta4GalNAc-T4) (Beta4GalNAcT4). | |||||
|
B4GN4_HUMAN
|
||||||
| θ value | 4.31301e-119 (rank : 4) | NC score | 0.906427 (rank : 4) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q76KP1, Q96LV2 | Gene names | B4GALNT4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N- acetylgalactosaminyltransferase 1 (EC 2.4.1.244) (NGalNAc-T1) (Beta- 1,4-N-acetylgalactosaminyltransferase IV) (Beta4GalNAc-T4) (Beta4GalNAcT4). | |||||
|
CGAT1_HUMAN
|
||||||
| θ value | 2.5182e-18 (rank : 5) | NC score | 0.585491 (rank : 5) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8TDX6, Q6P9G6, Q8IUF9, Q9NSQ7, Q9NUM9 | Gene names | GALNACT1, CHGN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin beta-1,4-N-acetylgalactosaminyltransferase 1 (EC 2.4.1.174) (beta4GalNAcT-1). | |||||
|
CGAT1_MOUSE
|
||||||
| θ value | 5.60996e-18 (rank : 6) | NC score | 0.582826 (rank : 6) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BJQ9, Q3UZX6, Q8BWV9, Q8BZU7, Q8C195, Q8R0M6 | Gene names | Galnact1, Chgn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin beta-1,4-N-acetylgalactosaminyltransferase 1 (EC 2.4.1.174) (beta4GalNAcT-1). | |||||
|
CGAT2_HUMAN
|
||||||
| θ value | 5.25075e-16 (rank : 7) | NC score | 0.567510 (rank : 7) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8N6G5, Q6MZJ5, Q6MZP6, Q8TCH4, Q9P1I6 | Gene names | GALNACT2, CHGN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin beta-1,4-N-acetylgalactosaminyltransferase 2 (EC 2.4.1.174) (GalNAcT-2) (beta4GalNAcT-2). | |||||
|
CGAT2_MOUSE
|
||||||
| θ value | 2.60593e-15 (rank : 8) | NC score | 0.561094 (rank : 8) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8C1F4, Q6PAP6, Q8R5A2, Q9D2M1 | Gene names | Galnact2, Chgn2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin beta-1,4-N-acetylgalactosaminyltransferase 2 (EC 2.4.1.174) (GalNAcT-2) (beta4GalNAcT-2). | |||||
|
CHSS1_MOUSE
|
||||||
| θ value | 1.58096e-12 (rank : 9) | NC score | 0.477405 (rank : 10) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6ZQ11 | Gene names | Chsy1, Kiaa0990 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate synthase 1 (EC 2.4.1.175) (Glucuronosyl-N- acetylgalactosaminyl-proteoglycan 4-beta-N- acetylgalactosaminyltransferase) (Chondroitin sulfate synthase 1) (N- acetylgalactosaminyl-proteoglycan 3-beta-glucuronosyltransferase) (EC 2.4.1.226) (Chondroitin glucuronyltransferase II) (N- acetylgalactosaminyltransferase II). | |||||
|
CHSS1_HUMAN
|
||||||
| θ value | 1.02475e-11 (rank : 10) | NC score | 0.478495 (rank : 9) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q86X52, Q6UX38, Q7LFU5, Q9Y2J5 | Gene names | CHSY1, CHSY, CSS1, KIAA0990 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate synthase 1 (EC 2.4.1.175) (Glucuronosyl-N- acetylgalactosaminyl-proteoglycan 4-beta-N- acetylgalactosaminyltransferase 1) (Chondroitin sulfate synthase 1) (N-acetylgalactosaminyl-proteoglycan 3-beta-glucuronosyltransferase 1) (EC 2.4.1.226) (Chondroitin glucuronyltransferase II) (N- acetylgalactosaminyltransferase II). | |||||
|
CHSS3_HUMAN
|
||||||
| θ value | 6.64225e-11 (rank : 11) | NC score | 0.460866 (rank : 11) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q70JA7, Q76L22, Q86Y52 | Gene names | CSS3, CHSY2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate synthase 3 (EC 2.4.1.175) (Glucuronosyl-N- acetylgalactosaminyl-proteoglycan 4-beta-N- acetylgalactosaminyltransferase II) (Chondroitin synthase 2) (N- acetylgalactosaminyl-proteoglycan 3-beta-glucuronosyltransferase II) (EC 2.4.1.226) (Chondroitin glucuronyltransferase II) (N- acetylgalactosaminyltransferase II) (Carbohydrate synthase 2). | |||||
|
MLL3_MOUSE
|
||||||
| θ value | 0.000461057 (rank : 12) | NC score | 0.059398 (rank : 20) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
SH2D3_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 13) | NC score | 0.041444 (rank : 31) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9QZS8, Q9JME1 | Gene names | Sh2d3c, Chat, Shep1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH2 domain-containing protein 3C (SH2 domain-containing Eph receptor- binding protein 1) (Cas/HEF1-associated signal transducer). | |||||
|
PCLO_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 14) | NC score | 0.042678 (rank : 28) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
MILK1_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 15) | NC score | 0.032007 (rank : 42) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8N3F8, Q5TI16, Q7RTP5, Q8N3N8, Q9BVL9, Q9BY92, Q9UH43, Q9UH44, Q9UH45 | Gene names | MIRAB13, KIAA1668 | |||
|
Domain Architecture |
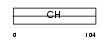
|
|||||
| Description | Molecule interacting with Rab13 (MIRab13) (MICAL-like protein 1). | |||||
|
SYNJ1_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 16) | NC score | 0.033711 (rank : 39) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O43426, O43425, O94984 | Gene names | SYNJ1, KIAA0910 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptojanin-1 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 1). | |||||
|
CR054_MOUSE
|
||||||
| θ value | 0.125558 (rank : 17) | NC score | 0.056342 (rank : 22) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CB14, Q8BNS5, Q8C2X0 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C18orf54 homolog precursor. | |||||
|
JPH2_MOUSE
|
||||||
| θ value | 0.163984 (rank : 18) | NC score | 0.027486 (rank : 57) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9ET78, Q9ET79 | Gene names | Jph2, Jp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junctophilin-2 (Junctophilin type 2) (JP-2). | |||||
|
DCC_HUMAN
|
||||||
| θ value | 0.279714 (rank : 19) | NC score | 0.012577 (rank : 96) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 583 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P43146 | Gene names | DCC | |||
|
Domain Architecture |

|
|||||
| Description | Netrin receptor DCC precursor (Tumor suppressor protein DCC) (Colorectal cancer suppressor). | |||||
|
LARG2_HUMAN
|
||||||
| θ value | 0.279714 (rank : 20) | NC score | 0.038850 (rank : 32) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8N3Y3, Q8N8Y6, Q8NAK3, Q8WY62 | Gene names | GYLTL1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glycosyltransferase-like protein LARGE2 (EC 2.4.-.-) (Glycosyltransferase-like 1B). | |||||
|
BRD4_MOUSE
|
||||||
| θ value | 0.365318 (rank : 21) | NC score | 0.030366 (rank : 48) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 845 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9ESU6, Q8VHF7, Q8VHF8 | Gene names | Brd4, Mcap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 4 (Mitotic chromosome-associated protein) (MCAP). | |||||
|
CHD2_HUMAN
|
||||||
| θ value | 0.365318 (rank : 22) | NC score | 0.014030 (rank : 92) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O14647 | Gene names | CHD2 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 2 (EC 3.6.1.-) (ATP- dependent helicase CHD2) (CHD-2). | |||||
|
PARM1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 23) | NC score | 0.046147 (rank : 27) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q923D3, Q3TTV0, Q3UFU4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein PARM-1 precursor. | |||||
|
RP1L1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 24) | NC score | 0.031221 (rank : 43) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1072 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8IWN7, Q86SQ1, Q8IWN8, Q8IWN9, Q8IWP0, Q8IWP1, Q8IWP2 | Gene names | RP1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinitis pigmentosa 1-like 1 protein. | |||||
|
SRRM1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 25) | NC score | 0.058493 (rank : 21) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
CD2_HUMAN
|
||||||
| θ value | 0.47712 (rank : 26) | NC score | 0.061209 (rank : 19) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P06729, Q96TE5 | Gene names | CD2 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell surface antigen CD2 precursor (T-cell surface antigen T11/Leu- 5) (LFA-2) (LFA-3 receptor) (Erythrocyte receptor) (Rosette receptor). | |||||
|
SOX17_MOUSE
|
||||||
| θ value | 0.47712 (rank : 27) | NC score | 0.016763 (rank : 82) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q61473, Q61472, Q62248 | Gene names | Sox17, Sox-17 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-17. | |||||
|
G3BP_MOUSE
|
||||||
| θ value | 0.62314 (rank : 28) | NC score | 0.038621 (rank : 33) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P97855 | Gene names | G3bp, G3bp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ras-GTPase-activating protein-binding protein 1 (EC 3.6.1.-) (ATP- dependent DNA helicase VIII) (GAP SH3-domain-binding protein 1) (G3BP- 1) (HDH-VIII). | |||||
|
LARG2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 29) | NC score | 0.036254 (rank : 35) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5XPT3, Q5XPT2, Q8BJZ8, Q8K253 | Gene names | Gyltl1b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glycosyltransferase-like protein LARGE2 (EC 2.4.-.-) (Glycosyltransferase-like 1B). | |||||
|
PRB4S_HUMAN
|
||||||
| θ value | 0.813845 (rank : 30) | NC score | 0.035611 (rank : 37) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P10163, P02813 | Gene names | PRB4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 4 allele S precursor (Salivary proline-rich protein Po) (Parotid o protein) [Contains: Protein N1; Glycosylated protein A]. | |||||
|
PS1C2_MOUSE
|
||||||
| θ value | 0.813845 (rank : 31) | NC score | 0.066079 (rank : 17) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q80ZC9 | Gene names | Psors1c2, Spr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Psoriasis susceptibility 1 candidate gene 2 protein homolog precursor (SPR1 protein). | |||||
|
SRRM1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 32) | NC score | 0.063324 (rank : 18) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
G3BP_HUMAN
|
||||||
| θ value | 1.06291 (rank : 33) | NC score | 0.037144 (rank : 34) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q13283 | Gene names | G3BP, G3BP1 | |||
|
Domain Architecture |

|
|||||
| Description | Ras-GTPase-activating protein-binding protein 1 (EC 3.6.1.-) (ATP- dependent DNA helicase VIII) (GAP SH3-domain-binding protein 1) (G3BP- 1) (HDH-VIII). | |||||
|
LR37B_HUMAN
|
||||||
| θ value | 1.06291 (rank : 34) | NC score | 0.028087 (rank : 55) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q96QE4, Q5YKG6 | Gene names | LRRC37B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 37B precursor (C66 SLIT-like testicular protein). | |||||
|
NCOR2_MOUSE
|
||||||
| θ value | 1.06291 (rank : 35) | NC score | 0.024159 (rank : 66) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 790 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9WU42, Q9WU43, Q9WUC1 | Gene names | Ncor2, Smrt | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC). | |||||
|
PRB3_HUMAN
|
||||||
| θ value | 1.06291 (rank : 36) | NC score | 0.033240 (rank : 40) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q04118, Q15188, Q7M4M9 | Gene names | PRB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 3 precursor (Parotid salivary glycoprotein G1) (Proline-rich protein G1). | |||||
|
AMELX_HUMAN
|
||||||
| θ value | 1.38821 (rank : 37) | NC score | 0.030631 (rank : 45) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99217, Q96NW6, Q9UCA7 | Gene names | AMELX, AMG, AMGX | |||
|
Domain Architecture |

|
|||||
| Description | Amelogenin, X isoform precursor. | |||||
|
BRD4_HUMAN
|
||||||
| θ value | 1.38821 (rank : 38) | NC score | 0.032709 (rank : 41) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O60885, O60433, Q96PD3 | Gene names | BRD4, HUNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 4 (HUNK1 protein). | |||||
|
CMGA_HUMAN
|
||||||
| θ value | 1.38821 (rank : 39) | NC score | 0.029677 (rank : 51) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P10645, Q96E84, Q96GL7, Q9BQB5 | Gene names | CHGA | |||
|
Domain Architecture |

|
|||||
| Description | Chromogranin A precursor (CgA) (Pituitary secretory protein I) (SP-I) [Contains: Vasostatin-1 (Vasostatin I); Vasostatin-2 (Vasostatin II); EA-92; ES-43; Pancreastatin; SS-18; WA-8; WE-14; LF-19; AL-11; GV-19; GR-44; ER-37]. | |||||
|
FHOD1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 40) | NC score | 0.023177 (rank : 70) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Y613, Q59F76, Q6Y1F2, Q76MS8, Q8N521 | Gene names | FHOD1, FHOS | |||
|
Domain Architecture |

|
|||||
| Description | FH1/FH2 domain-containing protein (Formin homolog overexpressed in spleen) (FHOS) (Formin homology 2 domain-containing protein 1). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | 1.38821 (rank : 41) | NC score | 0.046645 (rank : 26) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
PRP2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 42) | NC score | 0.035812 (rank : 36) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P05142 | Gene names | Prh1, Prp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein MP-2 precursor. | |||||
|
PS1C2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 43) | NC score | 0.052331 (rank : 24) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UIG4 | Gene names | PSORS1C2, C6orf17, SPR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Psoriasis susceptibility 1 candidate gene 2 protein precursor (SPR1 protein). | |||||
|
WNK4_HUMAN
|
||||||
| θ value | 1.38821 (rank : 44) | NC score | 0.004736 (rank : 123) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1043 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96J92, Q8N8X3, Q8N8Z2, Q96DT8, Q9BYS5 | Gene names | WNK4, PRKWNK4 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK4 (EC 2.7.11.1) (Protein kinase with no lysine 4) (Protein kinase, lysine-deficient 4). | |||||
|
ABCA4_HUMAN
|
||||||
| θ value | 1.81305 (rank : 45) | NC score | 0.010925 (rank : 102) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P78363, O15112, O60438, O60915 | Gene names | ABCA4, ABCR | |||
|
Domain Architecture |

|
|||||
| Description | Retinal-specific ATP-binding cassette transporter (ATP-binding cassette sub-family A member 4) (RIM ABC transporter) (RIM protein) (RmP) (Stargardt disease protein). | |||||
|
DDEF2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 46) | NC score | 0.010929 (rank : 101) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 625 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q7SIG6, Q501K1, Q66JN2 | Gene names | Ddef2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Development and differentiation-enhancing factor 2 (Pyk2 C-terminus- associated protein) (PAP) (Paxillin-associated protein with ARFGAP activity 3) (PAG3). | |||||
|
EMSY_HUMAN
|
||||||
| θ value | 1.81305 (rank : 47) | NC score | 0.028758 (rank : 53) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q7Z589, Q4G109, Q8NBU6, Q8TE50, Q9H8I9, Q9NRH0 | Gene names | EMSY, C11orf30 | |||
|
Domain Architecture |
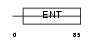
|
|||||
| Description | Protein EMSY. | |||||
|
PHLA1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 48) | NC score | 0.028966 (rank : 52) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q62392, Q3TY48, Q3UG87 | Gene names | Phlda1, Tdag51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family A member 1 (T-cell death- associated gene 51 protein) (Proline- and glutamine-rich protein) (PQR protein). | |||||
|
R3HD2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 49) | NC score | 0.029854 (rank : 50) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q80TM6, Q80YB1, Q8BLS5, Q9CW50 | Gene names | R3hdm2, Kiaa1002 | |||
|
Domain Architecture |

|
|||||
| Description | R3H domain-containing protein 2. | |||||
|
SFPQ_HUMAN
|
||||||
| θ value | 1.81305 (rank : 50) | NC score | 0.027776 (rank : 56) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 986 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P23246, P30808 | Gene names | SFPQ, PSF | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, proline- and glutamine-rich (Polypyrimidine tract- binding protein-associated-splicing factor) (PTB-associated-splicing factor) (PSF) (DNA-binding p52/p100 complex, 100 kDa subunit) (100 kDa DNA-pairing protein) (hPOMp100). | |||||
|
B4GT2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 51) | NC score | 0.074044 (rank : 15) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O60909, O60511, Q5T4X5, Q5T4Y5, Q9BUP6, Q9NSY7 | Gene names | B4GALT2 | |||
|
Domain Architecture |
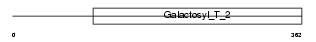
|
|||||
| Description | Beta-1,4-galactosyltransferase 2 (EC 2.4.1.-) (Beta-1,4-GalTase 2) (Beta4Gal-T2) (b4Gal-T2) (UDP-galactose:beta-N-acetylglucosamine beta- 1,4-galactosyltransferase 2) (UDP-Gal:beta-GlcNAc beta-1,4- galactosyltransferase 2) [Includes: Lactose synthase A protein (EC 2.4.1.22); N-acetyllactosamine synthase (EC 2.4.1.90) (Nal synthetase); Beta-N-acetylglucosaminylglycopeptide beta-1,4- galactosyltransferase (EC 2.4.1.38); Beta-N-acetylglucosaminyl- glycolipid beta-1,4-galactosyltransferase (EC 2.4.1.-)]. | |||||
|
SPTN2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 52) | NC score | 0.009147 (rank : 108) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 535 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O15020, O14872, O14873 | Gene names | SPTBN2, KIAA0302 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin beta chain, brain 2 (Spectrin, non-erythroid beta chain 2) (Beta-III spectrin). | |||||
|
ASPP1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 53) | NC score | 0.014961 (rank : 86) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1244 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q62415 | Gene names | Ppp1r13b, Aspp1 | |||
|
Domain Architecture |

|
|||||
| Description | Apoptosis-stimulating of p53 protein 1 (Protein phosphatase 1 regulatory subunit 13B). | |||||
|
B4GT2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 54) | NC score | 0.073230 (rank : 16) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9Z2Y2, Q3TMP2 | Gene names | B4galt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-1,4-galactosyltransferase 2 (EC 2.4.1.-) (Beta-1,4-GalTase 2) (Beta4Gal-T2) (b4Gal-T2) (UDP-galactose:beta-N-acetylglucosamine beta- 1,4-galactosyltransferase 2) (UDP-Gal:beta-GlcNAc beta-1,4- galactosyltransferase 2) [Includes: Lactose synthase A protein (EC 2.4.1.22); N-acetyllactosamine synthase (EC 2.4.1.90) (Nal synthetase); Beta-N-acetylglucosaminylglycopeptide beta-1,4- galactosyltransferase (EC 2.4.1.38); Beta-N-acetylglucosaminyl- glycolipid beta-1,4-galactosyltransferase (EC 2.4.1.-)]. | |||||
|
FOXJ3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 55) | NC score | 0.007372 (rank : 115) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UPW0, Q9NSS7 | Gene names | FOXJ3, KIAA1041 | |||
|
Domain Architecture |
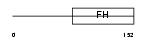
|
|||||
| Description | Forkhead box protein J3. | |||||
|
GALT3_MOUSE
|
||||||
| θ value | 3.0926 (rank : 56) | NC score | 0.026295 (rank : 60) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P70419, Q80V55 | Gene names | Galnt3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 3 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 3) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 3) (Polypeptide GalNAc transferase 3) (GalNAc-T3) (pp-GaNTase 3). | |||||
|
JADE2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 57) | NC score | 0.014609 (rank : 89) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NQC1, Q6IE80, Q8TEK0, Q92513, Q96GQ6 | Gene names | PHF15, JADE2, KIAA0239 | |||
|
Domain Architecture |

|
|||||
| Description | Protein Jade-2 (PHD finger protein 15). | |||||
|
K1802_HUMAN
|
||||||
| θ value | 3.0926 (rank : 58) | NC score | 0.018451 (rank : 76) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1378 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q96JM3, Q6P181, Q8NC88, Q9BST0 | Gene names | KIAA1802, C13orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
MLL2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 59) | NC score | 0.031084 (rank : 44) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
NSBP1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 60) | NC score | 0.014238 (rank : 90) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
PEG3_MOUSE
|
||||||
| θ value | 3.0926 (rank : 61) | NC score | 0.003071 (rank : 127) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1101 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q3URU2, O54978, Q3TQ69, Q5EBP7, Q61138, Q6GQS0, Q80U47, Q8R5N0, Q9QX53 | Gene names | Peg3, Kiaa0287, Pw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Paternally expressed gene 3 protein (ASF-1). | |||||
|
PITM3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 62) | NC score | 0.018339 (rank : 77) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9BZ71, Q59GH9, Q9NPQ4 | Gene names | PITPNM3, NIR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated phosphatidylinositol transfer protein 3 (Phosphatidylinositol transfer protein, membrane-associated 3) (Pyk2 N-terminal domain-interacting receptor 1) (NIR-1). | |||||
|
PITM3_MOUSE
|
||||||
| θ value | 3.0926 (rank : 63) | NC score | 0.018125 (rank : 79) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q3UHE1, Q3UH22, Q5RIT9 | Gene names | Pitpnm3, Nir1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated phosphatidylinositol transfer protein 3 (Phosphatidylinositol transfer protein, membrane-associated 3) (Pyk2 N-terminal domain-interacting receptor 1) (NIR-1). | |||||
|
REXO1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 64) | NC score | 0.019227 (rank : 74) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q7TT28, Q3UMP1, Q69ZR0, Q6NSQ6, Q6PI95, Q9DA29 | Gene names | Rexo1, Kiaa1138, Tceb3bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA exonuclease 1 homolog (EC 3.1.-.-) (Transcription elongation factor B polypeptide 3-binding protein 1). | |||||
|
TIF1A_MOUSE
|
||||||
| θ value | 3.0926 (rank : 65) | NC score | 0.017284 (rank : 80) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q64127, Q64126 | Gene names | Trim24, Tif1, Tif1a | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-alpha (TIF1-alpha) (Tripartite motif-containing protein 24). | |||||
|
CHGUT_HUMAN
|
||||||
| θ value | 4.03905 (rank : 66) | NC score | 0.157892 (rank : 14) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9P2E5, Q6P2I4, Q6UXD2 | Gene names | CSGLCAT, KIAA1402 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate glucuronyltransferase (EC 2.4.1.226) (N- acetylgalactosaminyl-proteoglycan 3-beta-glucuronosyltransferase) (Chondroitin glucuronyltransferase II) (CSGlcA-T). | |||||
|
HD_HUMAN
|
||||||
| θ value | 4.03905 (rank : 67) | NC score | 0.016821 (rank : 81) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P42858, Q9UQB7 | Gene names | HD, IT15 | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin (Huntington disease protein) (HD protein). | |||||
|
HGS_HUMAN
|
||||||
| θ value | 4.03905 (rank : 68) | NC score | 0.014658 (rank : 88) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 517 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O14964, Q9NR36 | Gene names | HGS, HRS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate (Protein pp110) (Hrs). | |||||
|
KI67_HUMAN
|
||||||
| θ value | 4.03905 (rank : 69) | NC score | 0.030397 (rank : 47) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P46013 | Gene names | MKI67 | |||
|
Domain Architecture |

|
|||||
| Description | Antigen KI-67. | |||||
|
NAT6_MOUSE
|
||||||
| θ value | 4.03905 (rank : 70) | NC score | 0.021663 (rank : 72) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9R123, Q8QZY5, Q8R014, Q9ERM0 | Gene names | Nat6, Fus2 | |||
|
Domain Architecture |

|
|||||
| Description | N-acetyltransferase 6 (EC 2.3.1.-) (Fus-2 protein) (Fusion 2 protein). | |||||
|
SMG5_MOUSE
|
||||||
| θ value | 4.03905 (rank : 71) | NC score | 0.015781 (rank : 84) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6ZPY2, Q497S2, Q8R3S4 | Gene names | Smg5, Est1b, Kiaa1089 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein SMG5 (SMG-5 homolog) (EST1-like protein B). | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 72) | NC score | 0.024164 (rank : 65) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
UBIP1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 73) | NC score | 0.013141 (rank : 94) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q811S7, Q3UPR3, Q3US11, Q60786, Q8C514 | Gene names | Ubp1, Cp2b, Nf2d9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Upstream-binding protein 1 (Nuclear factor 2d9) (NF2d9). | |||||
|
ABCA4_MOUSE
|
||||||
| θ value | 5.27518 (rank : 74) | NC score | 0.009343 (rank : 106) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O35600 | Gene names | Abca4, Abcr | |||
|
Domain Architecture |

|
|||||
| Description | Retinal-specific ATP-binding cassette transporter (ATP-binding cassette sub-family A member 4) (RIM ABC transporter) (RIM protein) (RmP). | |||||
|
AFF1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 75) | NC score | 0.018922 (rank : 75) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P51825 | Gene names | AFF1, AF4, FEL, MLLT2 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4) (Protein FEL). | |||||
|
ANR47_MOUSE
|
||||||
| θ value | 5.27518 (rank : 76) | NC score | 0.005899 (rank : 120) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Z1P7 | Gene names | Ankrd47, D17Ertd288e, Ng28 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 47. | |||||
|
DSCL1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 77) | NC score | 0.006229 (rank : 119) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8TD84, Q76MU9, Q8IZY3, Q8IZY4, Q8WXU7, Q9ULT7 | Gene names | DSCAML1, DSCAM2, KIAA1132 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Down syndrome cell adhesion molecule-like protein 1 precursor (Down syndrome cell adhesion molecule 2). | |||||
|
E41L2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 78) | NC score | 0.009005 (rank : 109) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 459 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O70318 | Gene names | Epb41l2, Epb4.1l2 | |||
|
Domain Architecture |

|
|||||
| Description | Band 4.1-like protein 2 (Generally expressed protein 4.1) (4.1G). | |||||
|
G3BP2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 79) | NC score | 0.030612 (rank : 46) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UN86, O60606, O75149, Q9UPA1 | Gene names | G3BP2, KIAA0660 | |||
|
Domain Architecture |

|
|||||
| Description | Ras-GTPase-activating protein-binding protein 2 (GAP SH3-domain- binding protein 2) (G3BP-2). | |||||
|
GALT3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 80) | NC score | 0.024101 (rank : 67) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14435, Q7Z476 | Gene names | GALNT3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 3 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 3) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 3) (Polypeptide GalNAc transferase 3) (GalNAc-T3) (pp-GaNTase 3). | |||||
|
GALT6_HUMAN
|
||||||
| θ value | 5.27518 (rank : 81) | NC score | 0.023270 (rank : 69) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8NCL4, Q8IYH4, Q9H6G2, Q9UIV5 | Gene names | GALNT6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 6 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 6) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 6) (Polypeptide GalNAc transferase 6) (GalNAc-T6) (pp-GaNTase 6). | |||||
|
MOR2A_MOUSE
|
||||||
| θ value | 5.27518 (rank : 82) | NC score | 0.012064 (rank : 98) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q69ZX6, Q5QNQ7, Q6P547 | Gene names | Morc2a, Kiaa0852, Zcwcc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORC family CW-type zinc finger protein 2A (Zinc finger CW-type coiled-coil domain protein 1). | |||||
|
SI1L3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 83) | NC score | 0.013949 (rank : 93) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O60292, Q2TV87 | Gene names | SIPA1L3, KIAA0545, SPAL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal-induced proliferation-associated 1-like protein 3 (SPA-1-like protein 3). | |||||
|
SPTA3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 84) | NC score | 0.030136 (rank : 49) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9D9T6, Q80ZS4, Q9D5L5, Q9DAF5 | Gene names | Spata3, Tsarg1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Spermatogenesis-associated protein 3 (Testis spermatocyte apoptosis- related protein 1) (Testis and spermatogenesis cell-related protein 1) (mTSARG1). | |||||
|
WIRE_MOUSE
|
||||||
| θ value | 5.27518 (rank : 85) | NC score | 0.024931 (rank : 63) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6PEV3 | Gene names | Wire | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WIP-related protein (WASP-interacting protein-related protein). | |||||
|
AMOL1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 86) | NC score | 0.009163 (rank : 107) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9D4H4, Q571F1 | Gene names | Amotl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin-like protein 1. | |||||
|
ARI1A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 87) | NC score | 0.023015 (rank : 71) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 787 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O14497, Q53FK9, Q5T0W1, Q5T0W2, Q5T0W3, Q8NFD6, Q96T89, Q9BY33, Q9HBJ5, Q9UPZ1 | Gene names | ARID1A, C1orf4, OSA1, SMARCF1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 1A (ARID domain- containing protein 1A) (SWI/SNF-related, matrix-associated, actin- dependent regulator of chromatin subfamily F member 1) (SWI-SNF complex protein p270) (B120) (SWI-like protein) (Osa homolog 1) (hOSA1) (hELD) (BRG1-associated factor 250) (BAF250) (BRG1-associated factor 250a) (BAF250A). | |||||
|
ATX2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 88) | NC score | 0.016499 (rank : 83) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 375 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q99700, Q6ZQZ7, Q99493 | Gene names | ATXN2, ATX2, SCA2, TNRC13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2 (Spinocerebellar ataxia type 2 protein) (Trinucleotide repeat-containing gene 13 protein). | |||||
|
BAT2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 89) | NC score | 0.025337 (rank : 62) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
DRD2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 90) | NC score | 0.000113 (rank : 131) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P61168, P13953 | Gene names | Drd2 | |||
|
Domain Architecture |
|
|||||
| Description | D(2) dopamine receptor (Dopamine D2 receptor). | |||||
|
FATH_HUMAN
|
||||||
| θ value | 6.88961 (rank : 91) | NC score | 0.000300 (rank : 130) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 570 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14517 | Gene names | FAT | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin-related tumor suppressor homolog precursor (Protein fat homolog). | |||||
|
HXA3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 92) | NC score | 0.003577 (rank : 126) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 535 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O43365 | Gene names | HOXA3, HOX1E | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-A3 (Hox-1E). | |||||
|
KCNC3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 93) | NC score | 0.004868 (rank : 122) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q14003 | Gene names | KCNC3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 3 (Voltage-gated potassium channel subunit Kv3.3) (KSHIIID). | |||||
|
NCA11_MOUSE
|
||||||
| θ value | 6.88961 (rank : 94) | NC score | 0.007054 (rank : 116) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 661 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P13595, Q61949 | Gene names | Ncam1, Ncam | |||
|
Domain Architecture |
|
|||||
| Description | Neural cell adhesion molecule 1, 180 kDa isoform precursor (N-CAM 180) (NCAM-180). | |||||
|
NEK1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 95) | NC score | -0.001430 (rank : 132) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P51954 | Gene names | Nek1 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Nek1 (EC 2.7.11.1) (NimA-related protein kinase 1). | |||||
|
PCLO_HUMAN
|
||||||
| θ value | 6.88961 (rank : 96) | NC score | 0.033808 (rank : 38) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
PLS1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 97) | NC score | 0.008924 (rank : 110) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O15162 | Gene names | PLSCR1 | |||
|
Domain Architecture |
|
|||||
| Description | Phospholipid scramblase 1 (PL scramblase 1) (Ca(2+)-dependent phospholipid scramblase 1) (Erythrocyte phospholipid scramblase) (MmTRA1b). | |||||
|
PRG4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 98) | NC score | 0.019933 (rank : 73) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
PRR8_HUMAN
|
||||||
| θ value | 6.88961 (rank : 99) | NC score | 0.026598 (rank : 59) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9NSV0 | Gene names | PRR8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 8. | |||||
|
RB15B_HUMAN
|
||||||
| θ value | 6.88961 (rank : 100) | NC score | 0.010457 (rank : 104) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8NDT2, Q6QE19, Q9BV96 | Gene names | RBM15B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative RNA-binding protein 15B (RNA-binding motif protein 15B). | |||||
|
RNZ2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 101) | NC score | 0.012531 (rank : 97) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BQ52, Q9HAS8, Q9HAS9, Q9NVT1 | Gene names | ELAC2, HPC2 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc phosphodiesterase ELAC protein 2 (EC 3.1.26.11) (Ribonuclease Z 2) (RNase Z 2) (tRNase Z 2) (tRNA 3 endonuclease 2) (ElaC homolog protein 2) (Heredity prostate cancer protein 2). | |||||
|
SFR15_HUMAN
|
||||||
| θ value | 6.88961 (rank : 102) | NC score | 0.025694 (rank : 61) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O95104, Q6P1M5, Q8N3I8, Q9UFM1, Q9ULP8 | Gene names | SFRS15, KIAA1172 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 15 (CTD-binding SR-like protein RA4). | |||||
|
SNIP1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 103) | NC score | 0.014704 (rank : 87) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BIZ6, Q3V106, Q8BIZ4 | Gene names | Snip1 | |||
|
Domain Architecture |

|
|||||
| Description | Smad nuclear-interacting protein 1. | |||||
|
TAF1C_HUMAN
|
||||||
| θ value | 6.88961 (rank : 104) | NC score | 0.013065 (rank : 95) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q15572, Q59F67, Q8N6V3 | Gene names | TAF1C | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TATA box-binding protein-associated factor RNA polymerase I subunit C (TATA box-binding protein-associated factor 1C) (TBP-associated factor 1C) (TBP-associated factor RNA polymerase I 110 kDa) (TAFI110). | |||||
|
TLE4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 105) | NC score | 0.007397 (rank : 114) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 477 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q04727, Q5T1Y2, Q9BZ07, Q9BZ08, Q9BZ09, Q9NSL3, Q9ULF9 | Gene names | TLE4, KIAA1261 | |||
|
Domain Architecture |

|
|||||
| Description | Transducin-like enhancer protein 4. | |||||
|
TLE4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 106) | NC score | 0.007411 (rank : 113) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 482 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q62441, Q9JKQ9 | Gene names | Tle4, Grg4 | |||
|
Domain Architecture |

|
|||||
| Description | Transducin-like enhancer protein 4 (Groucho-related protein 4) (Grg- 4). | |||||
|
B4GT6_HUMAN
|
||||||
| θ value | 8.99809 (rank : 107) | NC score | 0.042125 (rank : 29) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UBX8, O60514 | Gene names | B4GALT6 | |||
|
Domain Architecture |
|
|||||
| Description | Beta-1,4-galactosyltransferase 6 (EC 2.4.1.-) (Beta-1,4-GalTase 6) (Beta4Gal-T6) (b4Gal-T6) (UDP-galactose:beta-N-acetylglucosamine beta- 1,4-galactosyltransferase 6) (UDP-Gal:beta-GlcNAc beta-1,4- galactosyltransferase 6) [Includes: Lactosylceramide synthase (EC 2.4.1.-) (LacCer synthase) (UDP-Gal:glucosylceramide beta-1,4- galactosyltransferase)]. | |||||
|
B4GT6_MOUSE
|
||||||
| θ value | 8.99809 (rank : 108) | NC score | 0.041757 (rank : 30) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9WVK5 | Gene names | B4galt6 | |||
|
Domain Architecture |
|
|||||
| Description | Beta-1,4-galactosyltransferase 6 (EC 2.4.1.-) (Beta-1,4-GalTase 6) (Beta4Gal-T6) (b4Gal-T6) (UDP-galactose:beta-N-acetylglucosamine beta- 1,4-galactosyltransferase 6) (UDP-Gal:beta-GlcNAc beta-1,4- galactosyltransferase 6) [Includes: Lactosylceramide synthase (EC 2.4.1.-) (LacCer synthase) (UDP-Gal:glucosylceramide beta-1,4- galactosyltransferase)]. | |||||
|
BSN_HUMAN
|
||||||
| θ value | 8.99809 (rank : 109) | NC score | 0.023639 (rank : 68) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
CAR10_HUMAN
|
||||||
| θ value | 8.99809 (rank : 110) | NC score | 0.004346 (rank : 124) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 569 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BWT7, Q14CQ8, Q5TFG6, Q9UGR5, Q9UGR6, Q9Y3H0 | Gene names | CARD10, CARMA3 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 10 (CARD-containing MAGUK protein 3) (Carma 3). | |||||
|
DIDO1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 111) | NC score | 0.026600 (rank : 58) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
EVPL_MOUSE
|
||||||
| θ value | 8.99809 (rank : 112) | NC score | 0.002323 (rank : 128) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 952 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9D952 | Gene names | Evpl | |||
|
Domain Architecture |

|
|||||
| Description | Envoplakin (p210) (210 kDa cornified envelope precursor protein). | |||||
|
G3BP2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 113) | NC score | 0.028331 (rank : 54) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P97379, Q9R1B8 | Gene names | G3bp2 | |||
|
Domain Architecture |

|
|||||
| Description | Ras-GTPase-activating protein-binding protein 2 (GAP SH3-domain- binding protein 2) (G3BP-2). | |||||
|
GCP6_HUMAN
|
||||||
| θ value | 8.99809 (rank : 114) | NC score | 0.011248 (rank : 100) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 295 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96RT7, Q5JZ80, Q6PJ40, Q86YE9, Q9BY91, Q9UGX3, Q9UGX4 | Gene names | TUBGCP6, GCP6, KIAA1669 | |||
|
Domain Architecture |

|
|||||
| Description | Gamma-tubulin complex component 6 (GCP-6). | |||||
|
OBSCN_HUMAN
|
||||||
| θ value | 8.99809 (rank : 115) | NC score | 0.002301 (rank : 129) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1931 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q5VST9, Q2A664, Q5T7G8, Q5T7G9, Q5VSU2, Q86YC7, Q8NHN0, Q8NHN1, Q8NHN2, Q8NHN4, Q8NHN5, Q8NHN6, Q8NHN7, Q8NHN8, Q8NHN9, Q96AA2, Q9HCD3, Q9HCL6 | Gene names | OBSCN, KIAA1556, KIAA1639 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Obscurin (Obscurin-myosin light chain kinase) (Obscurin-MLCK) (Obscurin-RhoGEF). | |||||
|
PER2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 116) | NC score | 0.007036 (rank : 117) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O15055 | Gene names | PER2, KIAA0347 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2. | |||||
|
PININ_HUMAN
|
||||||
| θ value | 8.99809 (rank : 117) | NC score | 0.006277 (rank : 118) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 753 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9H307, O60899, Q53EM7, Q6P5X4, Q7KYL1, Q99738, Q9UHZ9, Q9UQR9 | Gene names | PNN, DRS, MEMA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pinin (140 kDa nuclear and cell adhesion-related phosphoprotein) (Domain-rich serine protein) (DRS-protein) (DRSP) (Melanoma metastasis clone A protein) (Desmosome-associated protein) (SR-like protein) (Nuclear protein SDK3). | |||||
|
PO6F2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 118) | NC score | 0.007419 (rank : 112) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 618 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P78424, P78425, Q75ME8, Q86UM6, Q9UDS7 | Gene names | POU6F2, RPF1 | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 6, transcription factor 2 (Retina-derived POU-domain factor 1) (RPF-1). | |||||
|
RTEL1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 119) | NC score | 0.009551 (rank : 105) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NZ71, Q5JTV4, Q9BW37, Q9H402, Q9H4X6, Q9NX25, Q9NZ73, Q9UPR4, Q9Y4R6 | Gene names | RTEL1, C20orf41, KIAA1088, NHL | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of telomere elongation helicase 1 (EC 3.6.1.-) (Helicase- like protein NHL). | |||||
|
SHC1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 120) | NC score | 0.008392 (rank : 111) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P29353, O15290, Q8N4K5, Q96CL1 | Gene names | SHC1, SHC, SHCA | |||
|
Domain Architecture |

|
|||||
| Description | SHC-transforming protein 1 (SH2 domain protein C1) (Src homology 2 domain-containing-transforming protein C1). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 121) | NC score | 0.024692 (rank : 64) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
TAU_HUMAN
|
||||||
| θ value | 8.99809 (rank : 122) | NC score | 0.018312 (rank : 78) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 448 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P10636, P18518, Q14799, Q15549, Q15550, Q15551, Q9UDJ3, Q9UMH0, Q9UQ96 | Gene names | MAPT, MTBT1, TAU | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein tau (Neurofibrillary tangle protein) (Paired helical filament-tau) (PHF-tau). | |||||
|
TCOF_MOUSE
|
||||||
| θ value | 8.99809 (rank : 123) | NC score | 0.015326 (rank : 85) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O08784, O08857 | Gene names | Tcof1 | |||
|
Domain Architecture |

|
|||||
| Description | Treacle protein (Treacher Collins syndrome protein homolog). | |||||
|
TIF1A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 124) | NC score | 0.014079 (rank : 91) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O15164, O95854 | Gene names | TRIM24, RNF82, TIF1, TIF1A | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-alpha (TIF1-alpha) (Tripartite motif-containing protein 24) (RING finger protein 82). | |||||
|
UBIP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 125) | NC score | 0.010916 (rank : 103) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NZI7, Q68CT0, Q86Y57, Q9H8V0, Q9UD76, Q9UD78 | Gene names | UBP1, LBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Upstream-binding protein 1 (LBP-1). | |||||
|
WBS14_MOUSE
|
||||||
| θ value | 8.99809 (rank : 126) | NC score | 0.011286 (rank : 99) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q99MZ3, Q99MY9, Q99MZ0, Q99MZ1, Q99MZ2, Q9JLM5 | Gene names | Mlxipl, Mio, Wbscr14 | |||
|
Domain Architecture |

|
|||||
| Description | Williams-Beuren syndrome chromosome region 14 protein homolog (Mlx interactor) (MLX-interacting protein-like). | |||||
|
WDR47_MOUSE
|
||||||
| θ value | 8.99809 (rank : 127) | NC score | 0.003720 (rank : 125) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 295 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8CGF6, Q80TP4 | Gene names | Wdr47, Kiaa0893 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 47. | |||||
|
ZFHX2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 128) | NC score | 0.004918 (rank : 121) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1178 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9C0A1 | Gene names | ZFHX2, KIAA1762 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger homeobox protein 2. | |||||
|
B3GLT_HUMAN
|
||||||
| θ value | θ > 10 (rank : 129) | NC score | 0.051733 (rank : 25) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6Y288, Q5W0H2, Q6NUI3 | Gene names | B3GALTL, B3GTL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-1,3-glucosyltransferase (EC 2.4.1.-) (Beta3Glc-T) (Beta-3- glycosyltransferase-like). | |||||
|
B3GLT_MOUSE
|
||||||
| θ value | θ > 10 (rank : 130) | NC score | 0.055897 (rank : 23) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BHT6, Q0PCR7, Q149S5 | Gene names | B3galtl, Gm1057 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-1,3-glucosyltransferase (EC 2.4.1.-) (Beta3Glc-T) (Beta-3- glycosyltransferase-like). | |||||
|
CHSS2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 131) | NC score | 0.180529 (rank : 12) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8IZ52, Q6UXD6, Q7L4G1, Q9H0F8, Q9H618 | Gene names | CSS2, CHPF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate synthase 2 (EC 2.4.1.175) (Glucuronosyl-N- acetylgalactosaminyl-proteoglycan 4-beta-N- acetylgalactosaminyltransferase II) (N-acetylgalactosaminyl- proteoglycan 3-beta-glucuronosyltransferase II) (EC 2.4.1.226) (Chondroitin glucuronyltransferase II) (N- acetylgalactosaminyltransferase) (Chondroitin-polymerizing factor). | |||||
|
CHSS2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 132) | NC score | 0.179987 (rank : 13) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6IQX7 | Gene names | Css2, D1Bwg1363e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate synthase 2 (EC 2.4.1.175) (Glucuronosyl-N- acetylgalactosaminyl-proteoglycan 4-beta-N- acetylgalactosaminyltransferase II) (N-acetylgalactosaminyl- proteoglycan 3-beta-glucuronosyltransferase II) (EC 2.4.1.226) (Chondroitin glucuronyltransferase II) (N- acetylgalactosaminyltransferase). | |||||
|
B4GN3_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 128 | |
| SwissProt Accessions | Q6L9W6, Q6ZNC1, Q8N7T6 | Gene names | B4GALNT3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N- acetylgalactosaminyltransferase 2 (EC 2.4.1.244) (NGalNAc-T2) (Beta- 1,4-N-acetylgalactosaminyltransferase III) (Beta4GalNAc-T3) (Beta4GalNAcT3). | |||||
|
B4GN3_MOUSE
|
||||||
| NC score | 0.977199 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6L8S8 | Gene names | B4galnt3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N- acetylgalactosaminyltransferase 2 (EC 2.4.1.244) (NGalNAc-T2) (Beta- 1,4-N-acetylgalactosaminyltransferase III) (Beta4GalNAc-T3) (Beta4GalNAcT3). | |||||
|
B4GN4_MOUSE
|
||||||
| NC score | 0.931433 (rank : 3) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q766D5, Q8CHV2 | Gene names | B4galnt4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N- acetylgalactosaminyltransferase 1 (EC 2.4.1.244) (NGalNAc-T1) (Beta- 1,4-N-acetylgalactosaminyltransferase IV) (Beta4GalNAc-T4) (Beta4GalNAcT4). | |||||
|
B4GN4_HUMAN
|
||||||
| NC score | 0.906427 (rank : 4) | θ value | 4.31301e-119 (rank : 4) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q76KP1, Q96LV2 | Gene names | B4GALNT4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N- acetylgalactosaminyltransferase 1 (EC 2.4.1.244) (NGalNAc-T1) (Beta- 1,4-N-acetylgalactosaminyltransferase IV) (Beta4GalNAc-T4) (Beta4GalNAcT4). | |||||
|
CGAT1_HUMAN
|
||||||
| NC score | 0.585491 (rank : 5) | θ value | 2.5182e-18 (rank : 5) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8TDX6, Q6P9G6, Q8IUF9, Q9NSQ7, Q9NUM9 | Gene names | GALNACT1, CHGN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin beta-1,4-N-acetylgalactosaminyltransferase 1 (EC 2.4.1.174) (beta4GalNAcT-1). | |||||
|
CGAT1_MOUSE
|
||||||
| NC score | 0.582826 (rank : 6) | θ value | 5.60996e-18 (rank : 6) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BJQ9, Q3UZX6, Q8BWV9, Q8BZU7, Q8C195, Q8R0M6 | Gene names | Galnact1, Chgn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin beta-1,4-N-acetylgalactosaminyltransferase 1 (EC 2.4.1.174) (beta4GalNAcT-1). | |||||
|
CGAT2_HUMAN
|
||||||
| NC score | 0.567510 (rank : 7) | θ value | 5.25075e-16 (rank : 7) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8N6G5, Q6MZJ5, Q6MZP6, Q8TCH4, Q9P1I6 | Gene names | GALNACT2, CHGN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin beta-1,4-N-acetylgalactosaminyltransferase 2 (EC 2.4.1.174) (GalNAcT-2) (beta4GalNAcT-2). | |||||
|
CGAT2_MOUSE
|
||||||
| NC score | 0.561094 (rank : 8) | θ value | 2.60593e-15 (rank : 8) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8C1F4, Q6PAP6, Q8R5A2, Q9D2M1 | Gene names | Galnact2, Chgn2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin beta-1,4-N-acetylgalactosaminyltransferase 2 (EC 2.4.1.174) (GalNAcT-2) (beta4GalNAcT-2). | |||||
|
CHSS1_HUMAN
|
||||||
| NC score | 0.478495 (rank : 9) | θ value | 1.02475e-11 (rank : 10) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q86X52, Q6UX38, Q7LFU5, Q9Y2J5 | Gene names | CHSY1, CHSY, CSS1, KIAA0990 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate synthase 1 (EC 2.4.1.175) (Glucuronosyl-N- acetylgalactosaminyl-proteoglycan 4-beta-N- acetylgalactosaminyltransferase 1) (Chondroitin sulfate synthase 1) (N-acetylgalactosaminyl-proteoglycan 3-beta-glucuronosyltransferase 1) (EC 2.4.1.226) (Chondroitin glucuronyltransferase II) (N- acetylgalactosaminyltransferase II). | |||||
|
CHSS1_MOUSE
|
||||||
| NC score | 0.477405 (rank : 10) | θ value | 1.58096e-12 (rank : 9) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6ZQ11 | Gene names | Chsy1, Kiaa0990 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate synthase 1 (EC 2.4.1.175) (Glucuronosyl-N- acetylgalactosaminyl-proteoglycan 4-beta-N- acetylgalactosaminyltransferase) (Chondroitin sulfate synthase 1) (N- acetylgalactosaminyl-proteoglycan 3-beta-glucuronosyltransferase) (EC 2.4.1.226) (Chondroitin glucuronyltransferase II) (N- acetylgalactosaminyltransferase II). | |||||
|
CHSS3_HUMAN
|
||||||
| NC score | 0.460866 (rank : 11) | θ value | 6.64225e-11 (rank : 11) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q70JA7, Q76L22, Q86Y52 | Gene names | CSS3, CHSY2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate synthase 3 (EC 2.4.1.175) (Glucuronosyl-N- acetylgalactosaminyl-proteoglycan 4-beta-N- acetylgalactosaminyltransferase II) (Chondroitin synthase 2) (N- acetylgalactosaminyl-proteoglycan 3-beta-glucuronosyltransferase II) (EC 2.4.1.226) (Chondroitin glucuronyltransferase II) (N- acetylgalactosaminyltransferase II) (Carbohydrate synthase 2). | |||||
|
CHSS2_HUMAN
|
||||||
| NC score | 0.180529 (rank : 12) | θ value | θ > 10 (rank : 131) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8IZ52, Q6UXD6, Q7L4G1, Q9H0F8, Q9H618 | Gene names | CSS2, CHPF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate synthase 2 (EC 2.4.1.175) (Glucuronosyl-N- acetylgalactosaminyl-proteoglycan 4-beta-N- acetylgalactosaminyltransferase II) (N-acetylgalactosaminyl- proteoglycan 3-beta-glucuronosyltransferase II) (EC 2.4.1.226) (Chondroitin glucuronyltransferase II) (N- acetylgalactosaminyltransferase) (Chondroitin-polymerizing factor). | |||||
|
CHSS2_MOUSE
|
||||||
| NC score | 0.179987 (rank : 13) | θ value | θ > 10 (rank : 132) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6IQX7 | Gene names | Css2, D1Bwg1363e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate synthase 2 (EC 2.4.1.175) (Glucuronosyl-N- acetylgalactosaminyl-proteoglycan 4-beta-N- acetylgalactosaminyltransferase II) (N-acetylgalactosaminyl- proteoglycan 3-beta-glucuronosyltransferase II) (EC 2.4.1.226) (Chondroitin glucuronyltransferase II) (N- acetylgalactosaminyltransferase). | |||||
|
CHGUT_HUMAN
|
||||||
| NC score | 0.157892 (rank : 14) | θ value | 4.03905 (rank : 66) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9P2E5, Q6P2I4, Q6UXD2 | Gene names | CSGLCAT, KIAA1402 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate glucuronyltransferase (EC 2.4.1.226) (N- acetylgalactosaminyl-proteoglycan 3-beta-glucuronosyltransferase) (Chondroitin glucuronyltransferase II) (CSGlcA-T). | |||||
|
B4GT2_HUMAN
|
||||||
| NC score | 0.074044 (rank : 15) | θ value | 2.36792 (rank : 51) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O60909, O60511, Q5T4X5, Q5T4Y5, Q9BUP6, Q9NSY7 | Gene names | B4GALT2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-1,4-galactosyltransferase 2 (EC 2.4.1.-) (Beta-1,4-GalTase 2) (Beta4Gal-T2) (b4Gal-T2) (UDP-galactose:beta-N-acetylglucosamine beta- 1,4-galactosyltransferase 2) (UDP-Gal:beta-GlcNAc beta-1,4- galactosyltransferase 2) [Includes: Lactose synthase A protein (EC 2.4.1.22); N-acetyllactosamine synthase (EC 2.4.1.90) (Nal synthetase); Beta-N-acetylglucosaminylglycopeptide beta-1,4- galactosyltransferase (EC 2.4.1.38); Beta-N-acetylglucosaminyl- glycolipid beta-1,4-galactosyltransferase (EC 2.4.1.-)]. | |||||
|
B4GT2_MOUSE
|
||||||
| NC score | 0.073230 (rank : 16) | θ value | 3.0926 (rank : 54) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9Z2Y2, Q3TMP2 | Gene names | B4galt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-1,4-galactosyltransferase 2 (EC 2.4.1.-) (Beta-1,4-GalTase 2) (Beta4Gal-T2) (b4Gal-T2) (UDP-galactose:beta-N-acetylglucosamine beta- 1,4-galactosyltransferase 2) (UDP-Gal:beta-GlcNAc beta-1,4- galactosyltransferase 2) [Includes: Lactose synthase A protein (EC 2.4.1.22); N-acetyllactosamine synthase (EC 2.4.1.90) (Nal synthetase); Beta-N-acetylglucosaminylglycopeptide beta-1,4- galactosyltransferase (EC 2.4.1.38); Beta-N-acetylglucosaminyl- glycolipid beta-1,4-galactosyltransferase (EC 2.4.1.-)]. | |||||
|
PS1C2_MOUSE
|
||||||
| NC score | 0.066079 (rank : 17) | θ value | 0.813845 (rank : 31) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q80ZC9 | Gene names | Psors1c2, Spr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Psoriasis susceptibility 1 candidate gene 2 protein homolog precursor (SPR1 protein). | |||||
|
SRRM1_MOUSE
|
||||||
| NC score | 0.063324 (rank : 18) | θ value | 0.813845 (rank : 32) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
CD2_HUMAN
|
||||||
| NC score | 0.061209 (rank : 19) | θ value | 0.47712 (rank : 26) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P06729, Q96TE5 | Gene names | CD2 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell surface antigen CD2 precursor (T-cell surface antigen T11/Leu- 5) (LFA-2) (LFA-3 receptor) (Erythrocyte receptor) (Rosette receptor). | |||||
|
MLL3_MOUSE
|
||||||
| NC score | 0.059398 (rank : 20) | θ value | 0.000461057 (rank : 12) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
SRRM1_HUMAN
|
||||||
| NC score | 0.058493 (rank : 21) | θ value | 0.365318 (rank : 25) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
CR054_MOUSE
|
||||||
| NC score | 0.056342 (rank : 22) | θ value | 0.125558 (rank : 17) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CB14, Q8BNS5, Q8C2X0 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C18orf54 homolog precursor. | |||||
|
B3GLT_MOUSE
|
||||||
| NC score | 0.055897 (rank : 23) | θ value | θ > 10 (rank : 130) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BHT6, Q0PCR7, Q149S5 | Gene names | B3galtl, Gm1057 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-1,3-glucosyltransferase (EC 2.4.1.-) (Beta3Glc-T) (Beta-3- glycosyltransferase-like). | |||||
|
PS1C2_HUMAN
|
||||||
| NC score | 0.052331 (rank : 24) | θ value | 1.38821 (rank : 43) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UIG4 | Gene names | PSORS1C2, C6orf17, SPR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Psoriasis susceptibility 1 candidate gene 2 protein precursor (SPR1 protein). | |||||
|
B3GLT_HUMAN
|
||||||
| NC score | 0.051733 (rank : 25) | θ value | θ > 10 (rank : 129) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6Y288, Q5W0H2, Q6NUI3 | Gene names | B3GALTL, B3GTL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-1,3-glucosyltransferase (EC 2.4.1.-) (Beta3Glc-T) (Beta-3- glycosyltransferase-like). | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.046645 (rank : 26) | θ value | 1.38821 (rank : 41) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
PARM1_MOUSE
|
||||||
| NC score | 0.046147 (rank : 27) | θ value | 0.365318 (rank : 23) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q923D3, Q3TTV0, Q3UFU4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein PARM-1 precursor. | |||||
|
PCLO_MOUSE
|
||||||
| NC score | 0.042678 (rank : 28) | θ value | 0.0736092 (rank : 14) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
B4GT6_HUMAN
|
||||||
| NC score | 0.042125 (rank : 29) | θ value | 8.99809 (rank : 107) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UBX8, O60514 | Gene names | B4GALT6 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-1,4-galactosyltransferase 6 (EC 2.4.1.-) (Beta-1,4-GalTase 6) (Beta4Gal-T6) (b4Gal-T6) (UDP-galactose:beta-N-acetylglucosamine beta- 1,4-galactosyltransferase 6) (UDP-Gal:beta-GlcNAc beta-1,4- galactosyltransferase 6) [Includes: Lactosylceramide synthase (EC 2.4.1.-) (LacCer synthase) (UDP-Gal:glucosylceramide beta-1,4- galactosyltransferase)]. | |||||
|
B4GT6_MOUSE
|
||||||
| NC score | 0.041757 (rank : 30) | θ value | 8.99809 (rank : 108) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9WVK5 | Gene names | B4galt6 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-1,4-galactosyltransferase 6 (EC 2.4.1.-) (Beta-1,4-GalTase 6) (Beta4Gal-T6) (b4Gal-T6) (UDP-galactose:beta-N-acetylglucosamine beta- 1,4-galactosyltransferase 6) (UDP-Gal:beta-GlcNAc beta-1,4- galactosyltransferase 6) [Includes: Lactosylceramide synthase (EC 2.4.1.-) (LacCer synthase) (UDP-Gal:glucosylceramide beta-1,4- galactosyltransferase)]. | |||||
|
SH2D3_MOUSE
|
||||||
| NC score | 0.041444 (rank : 31) | θ value | 0.0113563 (rank : 13) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9QZS8, Q9JME1 | Gene names | Sh2d3c, Chat, Shep1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH2 domain-containing protein 3C (SH2 domain-containing Eph receptor- binding protein 1) (Cas/HEF1-associated signal transducer). | |||||
|
LARG2_HUMAN
|
||||||
| NC score | 0.038850 (rank : 32) | θ value | 0.279714 (rank : 20) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8N3Y3, Q8N8Y6, Q8NAK3, Q8WY62 | Gene names | GYLTL1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glycosyltransferase-like protein LARGE2 (EC 2.4.-.-) (Glycosyltransferase-like 1B). | |||||
|
G3BP_MOUSE
|
||||||
| NC score | 0.038621 (rank : 33) | θ value | 0.62314 (rank : 28) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P97855 | Gene names | G3bp, G3bp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ras-GTPase-activating protein-binding protein 1 (EC 3.6.1.-) (ATP- dependent DNA helicase VIII) (GAP SH3-domain-binding protein 1) (G3BP- 1) (HDH-VIII). | |||||
|
G3BP_HUMAN
|
||||||
| NC score | 0.037144 (rank : 34) | θ value | 1.06291 (rank : 33) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q13283 | Gene names | G3BP, G3BP1 | |||
|
Domain Architecture |

|
|||||
| Description | Ras-GTPase-activating protein-binding protein 1 (EC 3.6.1.-) (ATP- dependent DNA helicase VIII) (GAP SH3-domain-binding protein 1) (G3BP- 1) (HDH-VIII). | |||||
|
LARG2_MOUSE
|
||||||
| NC score | 0.036254 (rank : 35) | θ value | 0.62314 (rank : 29) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5XPT3, Q5XPT2, Q8BJZ8, Q8K253 | Gene names | Gyltl1b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glycosyltransferase-like protein LARGE2 (EC 2.4.-.-) (Glycosyltransferase-like 1B). | |||||
|
PRP2_MOUSE
|
||||||
| NC score | 0.035812 (rank : 36) | θ value | 1.38821 (rank : 42) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P05142 | Gene names | Prh1, Prp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein MP-2 precursor. | |||||
|
PRB4S_HUMAN
|
||||||
| NC score | 0.035611 (rank : 37) | θ value | 0.813845 (rank : 30) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P10163, P02813 | Gene names | PRB4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 4 allele S precursor (Salivary proline-rich protein Po) (Parotid o protein) [Contains: Protein N1; Glycosylated protein A]. | |||||
|
PCLO_HUMAN
|
||||||
| NC score | 0.033808 (rank : 38) | θ value | 6.88961 (rank : 96) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
SYNJ1_HUMAN
|
||||||
| NC score | 0.033711 (rank : 39) | θ value | 0.0961366 (rank : 16) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O43426, O43425, O94984 | Gene names | SYNJ1, KIAA0910 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptojanin-1 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 1). | |||||
|
PRB3_HUMAN
|
||||||
| NC score | 0.033240 (rank : 40) | θ value | 1.06291 (rank : 36) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q04118, Q15188, Q7M4M9 | Gene names | PRB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 3 precursor (Parotid salivary glycoprotein G1) (Proline-rich protein G1). | |||||
|
BRD4_HUMAN
|
||||||
| NC score | 0.032709 (rank : 41) | θ value | 1.38821 (rank : 38) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O60885, O60433, Q96PD3 | Gene names | BRD4, HUNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 4 (HUNK1 protein). | |||||
|
MILK1_HUMAN
|
||||||
| NC score | 0.032007 (rank : 42) | θ value | 0.0961366 (rank : 15) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8N3F8, Q5TI16, Q7RTP5, Q8N3N8, Q9BVL9, Q9BY92, Q9UH43, Q9UH44, Q9UH45 | Gene names | MIRAB13, KIAA1668 | |||
|
Domain Architecture |

|
|||||
| Description | Molecule interacting with Rab13 (MIRab13) (MICAL-like protein 1). | |||||
|
RP1L1_HUMAN
|
||||||
| NC score | 0.031221 (rank : 43) | θ value | 0.365318 (rank : 24) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1072 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8IWN7, Q86SQ1, Q8IWN8, Q8IWN9, Q8IWP0, Q8IWP1, Q8IWP2 | Gene names | RP1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinitis pigmentosa 1-like 1 protein. | |||||
|
MLL2_HUMAN
|
||||||
| NC score | 0.031084 (rank : 44) | θ value | 3.0926 (rank : 59) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
AMELX_HUMAN
|
||||||
| NC score | 0.030631 (rank : 45) | θ value | 1.38821 (rank : 37) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99217, Q96NW6, Q9UCA7 | Gene names | AMELX, AMG, AMGX | |||
|
Domain Architecture |

|
|||||
| Description | Amelogenin, X isoform precursor. | |||||
|
G3BP2_HUMAN
|
||||||
| NC score | 0.030612 (rank : 46) | θ value | 5.27518 (rank : 79) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9UN86, O60606, O75149, Q9UPA1 | Gene names | G3BP2, KIAA0660 | |||
|
Domain Architecture |

|
|||||
| Description | Ras-GTPase-activating protein-binding protein 2 (GAP SH3-domain- binding protein 2) (G3BP-2). | |||||
|
KI67_HUMAN
|
||||||
| NC score | 0.030397 (rank : 47) | θ value | 4.03905 (rank : 69) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P46013 | Gene names | MKI67 | |||
|
Domain Architecture |

|
|||||
| Description | Antigen KI-67. | |||||
|
BRD4_MOUSE
|
||||||
| NC score | 0.030366 (rank : 48) | θ value | 0.365318 (rank : 21) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 845 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9ESU6, Q8VHF7, Q8VHF8 | Gene names | Brd4, Mcap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 4 (Mitotic chromosome-associated protein) (MCAP). | |||||
|
SPTA3_MOUSE
|
||||||
| NC score | 0.030136 (rank : 49) | θ value | 5.27518 (rank : 84) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9D9T6, Q80ZS4, Q9D5L5, Q9DAF5 | Gene names | Spata3, Tsarg1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Spermatogenesis-associated protein 3 (Testis spermatocyte apoptosis- related protein 1) (Testis and spermatogenesis cell-related protein 1) (mTSARG1). | |||||
|
R3HD2_MOUSE
|
||||||
| NC score | 0.029854 (rank : 50) | θ value | 1.81305 (rank : 49) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q80TM6, Q80YB1, Q8BLS5, Q9CW50 | Gene names | R3hdm2, Kiaa1002 | |||
|
Domain Architecture |

|
|||||
| Description | R3H domain-containing protein 2. | |||||
|
CMGA_HUMAN
|
||||||
| NC score | 0.029677 (rank : 51) | θ value | 1.38821 (rank : 39) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P10645, Q96E84, Q96GL7, Q9BQB5 | Gene names | CHGA | |||
|
Domain Architecture |

|
|||||
| Description | Chromogranin A precursor (CgA) (Pituitary secretory protein I) (SP-I) [Contains: Vasostatin-1 (Vasostatin I); Vasostatin-2 (Vasostatin II); EA-92; ES-43; Pancreastatin; SS-18; WA-8; WE-14; LF-19; AL-11; GV-19; GR-44; ER-37]. | |||||
|
PHLA1_MOUSE
|
||||||
| NC score | 0.028966 (rank : 52) | θ value | 1.81305 (rank : 48) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q62392, Q3TY48, Q3UG87 | Gene names | Phlda1, Tdag51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family A member 1 (T-cell death- associated gene 51 protein) (Proline- and glutamine-rich protein) (PQR protein). | |||||
|
EMSY_HUMAN
|
||||||
| NC score | 0.028758 (rank : 53) | θ value | 1.81305 (rank : 47) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q7Z589, Q4G109, Q8NBU6, Q8TE50, Q9H8I9, Q9NRH0 | Gene names | EMSY, C11orf30 | |||
|
Domain Architecture |

|
|||||
| Description | Protein EMSY. | |||||
|
G3BP2_MOUSE
|
||||||
| NC score | 0.028331 (rank : 54) | θ value | 8.99809 (rank : 113) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P97379, Q9R1B8 | Gene names | G3bp2 | |||
|
Domain Architecture |

|
|||||
| Description | Ras-GTPase-activating protein-binding protein 2 (GAP SH3-domain- binding protein 2) (G3BP-2). | |||||
|
LR37B_HUMAN
|
||||||
| NC score | 0.028087 (rank : 55) | θ value | 1.06291 (rank : 34) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q96QE4, Q5YKG6 | Gene names | LRRC37B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 37B precursor (C66 SLIT-like testicular protein). | |||||
|
SFPQ_HUMAN
|
||||||
| NC score | 0.027776 (rank : 56) | θ value | 1.81305 (rank : 50) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 986 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P23246, P30808 | Gene names | SFPQ, PSF | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, proline- and glutamine-rich (Polypyrimidine tract- binding protein-associated-splicing factor) (PTB-associated-splicing factor) (PSF) (DNA-binding p52/p100 complex, 100 kDa subunit) (100 kDa DNA-pairing protein) (hPOMp100). | |||||
|
JPH2_MOUSE
|
||||||
| NC score | 0.027486 (rank : 57) | θ value | 0.163984 (rank : 18) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9ET78, Q9ET79 | Gene names | Jph2, Jp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junctophilin-2 (Junctophilin type 2) (JP-2). | |||||
|
DIDO1_HUMAN
|
||||||
| NC score | 0.026600 (rank : 58) | θ value | 8.99809 (rank : 111) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
PRR8_HUMAN
|
||||||
| NC score | 0.026598 (rank : 59) | θ value | 6.88961 (rank : 99) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9NSV0 | Gene names | PRR8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 8. | |||||
|
GALT3_MOUSE
|
||||||
| NC score | 0.026295 (rank : 60) | θ value | 3.0926 (rank : 56) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P70419, Q80V55 | Gene names | Galnt3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 3 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 3) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 3) (Polypeptide GalNAc transferase 3) (GalNAc-T3) (pp-GaNTase 3). | |||||
|
SFR15_HUMAN
|
||||||
| NC score | 0.025694 (rank : 61) | θ value | 6.88961 (rank : 102) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O95104, Q6P1M5, Q8N3I8, Q9UFM1, Q9ULP8 | Gene names | SFRS15, KIAA1172 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 15 (CTD-binding SR-like protein RA4). | |||||
|
BAT2_HUMAN
|
||||||
| NC score | 0.025337 (rank : 62) | θ value | 6.88961 (rank : 89) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
WIRE_MOUSE
|
||||||
| NC score | 0.024931 (rank : 63) | θ value | 5.27518 (rank : 85) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6PEV3 | Gene names | Wire | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WIP-related protein (WASP-interacting protein-related protein). | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.024692 (rank : 64) | θ value | 8.99809 (rank : 121) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.024164 (rank : 65) | θ value | 4.03905 (rank : 72) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
NCOR2_MOUSE
|
||||||
| NC score | 0.024159 (rank : 66) | θ value | 1.06291 (rank : 35) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 790 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9WU42, Q9WU43, Q9WUC1 | Gene names | Ncor2, Smrt | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC). | |||||
|
GALT3_HUMAN
|
||||||
| NC score | 0.024101 (rank : 67) | θ value | 5.27518 (rank : 80) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14435, Q7Z476 | Gene names | GALNT3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 3 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 3) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 3) (Polypeptide GalNAc transferase 3) (GalNAc-T3) (pp-GaNTase 3). | |||||
|
BSN_HUMAN
|
||||||
| NC score | 0.023639 (rank : 68) | θ value | 8.99809 (rank : 109) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
GALT6_HUMAN
|
||||||
| NC score | 0.023270 (rank : 69) | θ value | 5.27518 (rank : 81) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8NCL4, Q8IYH4, Q9H6G2, Q9UIV5 | Gene names | GALNT6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 6 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 6) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 6) (Polypeptide GalNAc transferase 6) (GalNAc-T6) (pp-GaNTase 6). | |||||
|
FHOD1_HUMAN
|
||||||
| NC score | 0.023177 (rank : 70) | θ value | 1.38821 (rank : 40) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Y613, Q59F76, Q6Y1F2, Q76MS8, Q8N521 | Gene names | FHOD1, FHOS | |||
|
Domain Architecture |

|
|||||
| Description | FH1/FH2 domain-containing protein (Formin homolog overexpressed in spleen) (FHOS) (Formin homology 2 domain-containing protein 1). | |||||
|
ARI1A_HUMAN
|
||||||
| NC score | 0.023015 (rank : 71) | θ value | 6.88961 (rank : 87) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 787 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O14497, Q53FK9, Q5T0W1, Q5T0W2, Q5T0W3, Q8NFD6, Q96T89, Q9BY33, Q9HBJ5, Q9UPZ1 | Gene names | ARID1A, C1orf4, OSA1, SMARCF1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 1A (ARID domain- containing protein 1A) (SWI/SNF-related, matrix-associated, actin- dependent regulator of chromatin subfamily F member 1) (SWI-SNF complex protein p270) (B120) (SWI-like protein) (Osa homolog 1) (hOSA1) (hELD) (BRG1-associated factor 250) (BAF250) (BRG1-associated factor 250a) (BAF250A). | |||||
|
NAT6_MOUSE
|
||||||
| NC score | 0.021663 (rank : 72) | θ value | 4.03905 (rank : 70) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9R123, Q8QZY5, Q8R014, Q9ERM0 | Gene names | Nat6, Fus2 | |||
|
Domain Architecture |

|
|||||
| Description | N-acetyltransferase 6 (EC 2.3.1.-) (Fus-2 protein) (Fusion 2 protein). | |||||
|
PRG4_HUMAN
|
||||||
| NC score | 0.019933 (rank : 73) | θ value | 6.88961 (rank : 98) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
REXO1_MOUSE
|
||||||
| NC score | 0.019227 (rank : 74) | θ value | 3.0926 (rank : 64) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q7TT28, Q3UMP1, Q69ZR0, Q6NSQ6, Q6PI95, Q9DA29 | Gene names | Rexo1, Kiaa1138, Tceb3bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA exonuclease 1 homolog (EC 3.1.-.-) (Transcription elongation factor B polypeptide 3-binding protein 1). | |||||
|
AFF1_HUMAN
|
||||||
| NC score | 0.018922 (rank : 75) | θ value | 5.27518 (rank : 75) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P51825 | Gene names | AFF1, AF4, FEL, MLLT2 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4) (Protein FEL). | |||||
|
K1802_HUMAN
|
||||||
| NC score | 0.018451 (rank : 76) | θ value | 3.0926 (rank : 58) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1378 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q96JM3, Q6P181, Q8NC88, Q9BST0 | Gene names | KIAA1802, C13orf8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
PITM3_HUMAN
|
||||||
| NC score | 0.018339 (rank : 77) | θ value | 3.0926 (rank : 62) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9BZ71, Q59GH9, Q9NPQ4 | Gene names | PITPNM3, NIR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated phosphatidylinositol transfer protein 3 (Phosphatidylinositol transfer protein, membrane-associated 3) (Pyk2 N-terminal domain-interacting receptor 1) (NIR-1). | |||||
|
TAU_HUMAN
|
||||||
| NC score | 0.018312 (rank : 78) | θ value | 8.99809 (rank : 122) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 448 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P10636, P18518, Q14799, Q15549, Q15550, Q15551, Q9UDJ3, Q9UMH0, Q9UQ96 | Gene names | MAPT, MTBT1, TAU | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein tau (Neurofibrillary tangle protein) (Paired helical filament-tau) (PHF-tau). | |||||
|
PITM3_MOUSE
|
||||||
| NC score | 0.018125 (rank : 79) | θ value | 3.0926 (rank : 63) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q3UHE1, Q3UH22, Q5RIT9 | Gene names | Pitpnm3, Nir1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane-associated phosphatidylinositol transfer protein 3 (Phosphatidylinositol transfer protein, membrane-associated 3) (Pyk2 N-terminal domain-interacting receptor 1) (NIR-1). | |||||
|
TIF1A_MOUSE
|
||||||
| NC score | 0.017284 (rank : 80) | θ value | 3.0926 (rank : 65) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q64127, Q64126 | Gene names | Trim24, Tif1, Tif1a | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-alpha (TIF1-alpha) (Tripartite motif-containing protein 24). | |||||
|
HD_HUMAN
|
||||||
| NC score | 0.016821 (rank : 81) | θ value | 4.03905 (rank : 67) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P42858, Q9UQB7 | Gene names | HD, IT15 | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin (Huntington disease protein) (HD protein). | |||||
|
SOX17_MOUSE
|
||||||
| NC score | 0.016763 (rank : 82) | θ value | 0.47712 (rank : 27) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q61473, Q61472, Q62248 | Gene names | Sox17, Sox-17 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-17. | |||||
|
ATX2_HUMAN
|
||||||
| NC score | 0.016499 (rank : 83) | θ value | 6.88961 (rank : 88) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 375 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q99700, Q6ZQZ7, Q99493 | Gene names | ATXN2, ATX2, SCA2, TNRC13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2 (Spinocerebellar ataxia type 2 protein) (Trinucleotide repeat-containing gene 13 protein). | |||||
|
SMG5_MOUSE
|
||||||
| NC score | 0.015781 (rank : 84) | θ value | 4.03905 (rank : 71) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6ZPY2, Q497S2, Q8R3S4 | Gene names | Smg5, Est1b, Kiaa1089 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein SMG5 (SMG-5 homolog) (EST1-like protein B). | |||||
|
TCOF_MOUSE
|
||||||
| NC score | 0.015326 (rank : 85) | θ value | 8.99809 (rank : 123) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O08784, O08857 | Gene names | Tcof1 | |||
|
Domain Architecture |

|
|||||
| Description | Treacle protein (Treacher Collins syndrome protein homolog). | |||||
|
ASPP1_MOUSE
|
||||||
| NC score | 0.014961 (rank : 86) | θ value | 3.0926 (rank : 53) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1244 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q62415 | Gene names | Ppp1r13b, Aspp1 | |||
|
Domain Architecture |

|
|||||
| Description | Apoptosis-stimulating of p53 protein 1 (Protein phosphatase 1 regulatory subunit 13B). | |||||
|
SNIP1_MOUSE
|
||||||
| NC score | 0.014704 (rank : 87) | θ value | 6.88961 (rank : 103) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BIZ6, Q3V106, Q8BIZ4 | Gene names | Snip1 | |||
|
Domain Architecture |

|
|||||
| Description | Smad nuclear-interacting protein 1. | |||||
|
HGS_HUMAN
|
||||||
| NC score | 0.014658 (rank : 88) | θ value | 4.03905 (rank : 68) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 517 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O14964, Q9NR36 | Gene names | HGS, HRS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatocyte growth factor-regulated tyrosine kinase substrate (Protein pp110) (Hrs). | |||||
|
JADE2_HUMAN
|
||||||
| NC score | 0.014609 (rank : 89) | θ value | 3.0926 (rank : 57) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NQC1, Q6IE80, Q8TEK0, Q92513, Q96GQ6 | Gene names | PHF15, JADE2, KIAA0239 | |||
|
Domain Architecture |

|
|||||
| Description | Protein Jade-2 (PHD finger protein 15). | |||||
|
NSBP1_MOUSE
|
||||||
| NC score | 0.014238 (rank : 90) | θ value | 3.0926 (rank : 60) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
TIF1A_HUMAN
|
||||||
| NC score | 0.014079 (rank : 91) | θ value | 8.99809 (rank : 124) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O15164, O95854 | Gene names | TRIM24, RNF82, TIF1, TIF1A | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-alpha (TIF1-alpha) (Tripartite motif-containing protein 24) (RING finger protein 82). | |||||
|
CHD2_HUMAN
|
||||||
| NC score | 0.014030 (rank : 92) | θ value | 0.365318 (rank : 22) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O14647 | Gene names | CHD2 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 2 (EC 3.6.1.-) (ATP- dependent helicase CHD2) (CHD-2). | |||||
|
SI1L3_HUMAN
|
||||||
| NC score | 0.013949 (rank : 93) | θ value | 5.27518 (rank : 83) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O60292, Q2TV87 | Gene names | SIPA1L3, KIAA0545, SPAL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal-induced proliferation-associated 1-like protein 3 (SPA-1-like protein 3). | |||||
|
UBIP1_MOUSE
|
||||||
| NC score | 0.013141 (rank : 94) | θ value | 4.03905 (rank : 73) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q811S7, Q3UPR3, Q3US11, Q60786, Q8C514 | Gene names | Ubp1, Cp2b, Nf2d9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Upstream-binding protein 1 (Nuclear factor 2d9) (NF2d9). | |||||
|
TAF1C_HUMAN
|
||||||
| NC score | 0.013065 (rank : 95) | θ value | 6.88961 (rank : 104) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q15572, Q59F67, Q8N6V3 | Gene names | TAF1C | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TATA box-binding protein-associated factor RNA polymerase I subunit C (TATA box-binding protein-associated factor 1C) (TBP-associated factor 1C) (TBP-associated factor RNA polymerase I 110 kDa) (TAFI110). | |||||
|
DCC_HUMAN
|
||||||
| NC score | 0.012577 (rank : 96) | θ value | 0.279714 (rank : 19) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 583 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P43146 | Gene names | DCC | |||
|
Domain Architecture |

|
|||||
| Description | Netrin receptor DCC precursor (Tumor suppressor protein DCC) (Colorectal cancer suppressor). | |||||
|
RNZ2_HUMAN
|
||||||
| NC score | 0.012531 (rank : 97) | θ value | 6.88961 (rank : 101) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BQ52, Q9HAS8, Q9HAS9, Q9NVT1 | Gene names | ELAC2, HPC2 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc phosphodiesterase ELAC protein 2 (EC 3.1.26.11) (Ribonuclease Z 2) (RNase Z 2) (tRNase Z 2) (tRNA 3 endonuclease 2) (ElaC homolog protein 2) (Heredity prostate cancer protein 2). | |||||
|
MOR2A_MOUSE
|
||||||
| NC score | 0.012064 (rank : 98) | θ value | 5.27518 (rank : 82) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q69ZX6, Q5QNQ7, Q6P547 | Gene names | Morc2a, Kiaa0852, Zcwcc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORC family CW-type zinc finger protein 2A (Zinc finger CW-type coiled-coil domain protein 1). | |||||
|
WBS14_MOUSE
|
||||||
| NC score | 0.011286 (rank : 99) | θ value | 8.99809 (rank : 126) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q99MZ3, Q99MY9, Q99MZ0, Q99MZ1, Q99MZ2, Q9JLM5 | Gene names | Mlxipl, Mio, Wbscr14 | |||
|
Domain Architecture |

|
|||||
| Description | Williams-Beuren syndrome chromosome region 14 protein homolog (Mlx interactor) (MLX-interacting protein-like). | |||||
|
GCP6_HUMAN
|
||||||
| NC score | 0.011248 (rank : 100) | θ value | 8.99809 (rank : 114) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 295 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96RT7, Q5JZ80, Q6PJ40, Q86YE9, Q9BY91, Q9UGX3, Q9UGX4 | Gene names | TUBGCP6, GCP6, KIAA1669 | |||
|
Domain Architecture |

|
|||||
| Description | Gamma-tubulin complex component 6 (GCP-6). | |||||
|
DDEF2_MOUSE
|
||||||
| NC score | 0.010929 (rank : 101) | θ value | 1.81305 (rank : 46) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 625 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q7SIG6, Q501K1, Q66JN2 | Gene names | Ddef2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Development and differentiation-enhancing factor 2 (Pyk2 C-terminus- associated protein) (PAP) (Paxillin-associated protein with ARFGAP activity 3) (PAG3). | |||||
|
ABCA4_HUMAN
|
||||||
| NC score | 0.010925 (rank : 102) | θ value | 1.81305 (rank : 45) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P78363, O15112, O60438, O60915 | Gene names | ABCA4, ABCR | |||
|
Domain Architecture |

|
|||||
| Description | Retinal-specific ATP-binding cassette transporter (ATP-binding cassette sub-family A member 4) (RIM ABC transporter) (RIM protein) (RmP) (Stargardt disease protein). | |||||
|
UBIP1_HUMAN
|
||||||
| NC score | 0.010916 (rank : 103) | θ value | 8.99809 (rank : 125) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NZI7, Q68CT0, Q86Y57, Q9H8V0, Q9UD76, Q9UD78 | Gene names | UBP1, LBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Upstream-binding protein 1 (LBP-1). | |||||
|
RB15B_HUMAN
|
||||||
| NC score | 0.010457 (rank : 104) | θ value | 6.88961 (rank : 100) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8NDT2, Q6QE19, Q9BV96 | Gene names | RBM15B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative RNA-binding protein 15B (RNA-binding motif protein 15B). | |||||
|
RTEL1_HUMAN
|
||||||
| NC score | 0.009551 (rank : 105) | θ value | 8.99809 (rank : 119) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NZ71, Q5JTV4, Q9BW37, Q9H402, Q9H4X6, Q9NX25, Q9NZ73, Q9UPR4, Q9Y4R6 | Gene names | RTEL1, C20orf41, KIAA1088, NHL | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of telomere elongation helicase 1 (EC 3.6.1.-) (Helicase- like protein NHL). | |||||
|
ABCA4_MOUSE
|
||||||
| NC score | 0.009343 (rank : 106) | θ value | 5.27518 (rank : 74) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O35600 | Gene names | Abca4, Abcr | |||
|
Domain Architecture |

|
|||||
| Description | Retinal-specific ATP-binding cassette transporter (ATP-binding cassette sub-family A member 4) (RIM ABC transporter) (RIM protein) (RmP). | |||||
|
AMOL1_MOUSE
|
||||||
| NC score | 0.009163 (rank : 107) | θ value | 6.88961 (rank : 86) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9D4H4, Q571F1 | Gene names | Amotl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin-like protein 1. | |||||
|
SPTN2_HUMAN
|
||||||
| NC score | 0.009147 (rank : 108) | θ value | 2.36792 (rank : 52) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 535 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O15020, O14872, O14873 | Gene names | SPTBN2, KIAA0302 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin beta chain, brain 2 (Spectrin, non-erythroid beta chain 2) (Beta-III spectrin). | |||||
|
E41L2_MOUSE
|
||||||
| NC score | 0.009005 (rank : 109) | θ value | 5.27518 (rank : 78) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 459 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O70318 | Gene names | Epb41l2, Epb4.1l2 | |||
|
Domain Architecture |

|
|||||
| Description | Band 4.1-like protein 2 (Generally expressed protein 4.1) (4.1G). | |||||
|
PLS1_HUMAN
|
||||||
| NC score | 0.008924 (rank : 110) | θ value | 6.88961 (rank : 97) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O15162 | Gene names | PLSCR1 | |||
|
Domain Architecture |

|
|||||
| Description | Phospholipid scramblase 1 (PL scramblase 1) (Ca(2+)-dependent phospholipid scramblase 1) (Erythrocyte phospholipid scramblase) (MmTRA1b). | |||||
|
SHC1_HUMAN
|
||||||
| NC score | 0.008392 (rank : 111) | θ value | 8.99809 (rank : 120) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P29353, O15290, Q8N4K5, Q96CL1 | Gene names | SHC1, SHC, SHCA | |||
|
Domain Architecture |

|
|||||
| Description | SHC-transforming protein 1 (SH2 domain protein C1) (Src homology 2 domain-containing-transforming protein C1). | |||||
|
PO6F2_HUMAN
|
||||||
| NC score | 0.007419 (rank : 112) | θ value | 8.99809 (rank : 118) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 618 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P78424, P78425, Q75ME8, Q86UM6, Q9UDS7 | Gene names | POU6F2, RPF1 | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 6, transcription factor 2 (Retina-derived POU-domain factor 1) (RPF-1). | |||||
|
TLE4_MOUSE
|
||||||
| NC score | 0.007411 (rank : 113) | θ value | 6.88961 (rank : 106) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 482 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q62441, Q9JKQ9 | Gene names | Tle4, Grg4 | |||
|
Domain Architecture |

|
|||||
| Description | Transducin-like enhancer protein 4 (Groucho-related protein 4) (Grg- 4). | |||||
|
TLE4_HUMAN
|
||||||
| NC score | 0.007397 (rank : 114) | θ value | 6.88961 (rank : 105) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 477 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q04727, Q5T1Y2, Q9BZ07, Q9BZ08, Q9BZ09, Q9NSL3, Q9ULF9 | Gene names | TLE4, KIAA1261 | |||
|
Domain Architecture |

|
|||||
| Description | Transducin-like enhancer protein 4. | |||||
|
FOXJ3_HUMAN
|
||||||
| NC score | 0.007372 (rank : 115) | θ value | 3.0926 (rank : 55) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UPW0, Q9NSS7 | Gene names | FOXJ3, KIAA1041 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein J3. | |||||
|
NCA11_MOUSE
|
||||||
| NC score | 0.007054 (rank : 116) | θ value | 6.88961 (rank : 94) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 661 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P13595, Q61949 | Gene names | Ncam1, Ncam | |||
|
Domain Architecture |

|
|||||
| Description | Neural cell adhesion molecule 1, 180 kDa isoform precursor (N-CAM 180) (NCAM-180). | |||||
|
PER2_HUMAN
|
||||||
| NC score | 0.007036 (rank : 117) | θ value | 8.99809 (rank : 116) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O15055 | Gene names | PER2, KIAA0347 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 2. | |||||
|
PININ_HUMAN
|
||||||
| NC score | 0.006277 (rank : 118) | θ value | 8.99809 (rank : 117) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 753 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9H307, O60899, Q53EM7, Q6P5X4, Q7KYL1, Q99738, Q9UHZ9, Q9UQR9 | Gene names | PNN, DRS, MEMA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pinin (140 kDa nuclear and cell adhesion-related phosphoprotein) (Domain-rich serine protein) (DRS-protein) (DRSP) (Melanoma metastasis clone A protein) (Desmosome-associated protein) (SR-like protein) (Nuclear protein SDK3). | |||||
|
DSCL1_HUMAN
|
||||||
| NC score | 0.006229 (rank : 119) | θ value | 5.27518 (rank : 77) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 589 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8TD84, Q76MU9, Q8IZY3, Q8IZY4, Q8WXU7, Q9ULT7 | Gene names | DSCAML1, DSCAM2, KIAA1132 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Down syndrome cell adhesion molecule-like protein 1 precursor (Down syndrome cell adhesion molecule 2). | |||||
|
ANR47_MOUSE
|
||||||
| NC score | 0.005899 (rank : 120) | θ value | 5.27518 (rank : 76) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Z1P7 | Gene names | Ankrd47, D17Ertd288e, Ng28 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 47. | |||||
|
ZFHX2_HUMAN
|
||||||
| NC score | 0.004918 (rank : 121) | θ value | 8.99809 (rank : 128) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1178 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9C0A1 | Gene names | ZFHX2, KIAA1762 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger homeobox protein 2. | |||||
|
KCNC3_HUMAN
|
||||||
| NC score | 0.004868 (rank : 122) | θ value | 6.88961 (rank : 93) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q14003 | Gene names | KCNC3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 3 (Voltage-gated potassium channel subunit Kv3.3) (KSHIIID). | |||||
|
WNK4_HUMAN
|
||||||
| NC score | 0.004736 (rank : 123) | θ value | 1.38821 (rank : 44) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1043 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96J92, Q8N8X3, Q8N8Z2, Q96DT8, Q9BYS5 | Gene names | WNK4, PRKWNK4 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK4 (EC 2.7.11.1) (Protein kinase with no lysine 4) (Protein kinase, lysine-deficient 4). | |||||
|
CAR10_HUMAN
|
||||||
| NC score | 0.004346 (rank : 124) | θ value | 8.99809 (rank : 110) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 569 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BWT7, Q14CQ8, Q5TFG6, Q9UGR5, Q9UGR6, Q9Y3H0 | Gene names | CARD10, CARMA3 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 10 (CARD-containing MAGUK protein 3) (Carma 3). | |||||
|
WDR47_MOUSE
|
||||||
| NC score | 0.003720 (rank : 125) | θ value | 8.99809 (rank : 127) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 295 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8CGF6, Q80TP4 | Gene names | Wdr47, Kiaa0893 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 47. | |||||
|
HXA3_HUMAN
|
||||||
| NC score | 0.003577 (rank : 126) | θ value | 6.88961 (rank : 92) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 535 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O43365 | Gene names | HOXA3, HOX1E | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-A3 (Hox-1E). | |||||
|
PEG3_MOUSE
|
||||||
| NC score | 0.003071 (rank : 127) | θ value | 3.0926 (rank : 61) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1101 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q3URU2, O54978, Q3TQ69, Q5EBP7, Q61138, Q6GQS0, Q80U47, Q8R5N0, Q9QX53 | Gene names | Peg3, Kiaa0287, Pw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Paternally expressed gene 3 protein (ASF-1). | |||||
|
EVPL_MOUSE
|
||||||
| NC score | 0.002323 (rank : 128) | θ value | 8.99809 (rank : 112) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 952 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9D952 | Gene names | Evpl | |||
|
Domain Architecture |

|
|||||
| Description | Envoplakin (p210) (210 kDa cornified envelope precursor protein). | |||||
|
OBSCN_HUMAN
|
||||||
| NC score | 0.002301 (rank : 129) | θ value | 8.99809 (rank : 115) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1931 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q5VST9, Q2A664, Q5T7G8, Q5T7G9, Q5VSU2, Q86YC7, Q8NHN0, Q8NHN1, Q8NHN2, Q8NHN4, Q8NHN5, Q8NHN6, Q8NHN7, Q8NHN8, Q8NHN9, Q96AA2, Q9HCD3, Q9HCL6 | Gene names | OBSCN, KIAA1556, KIAA1639 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Obscurin (Obscurin-myosin light chain kinase) (Obscurin-MLCK) (Obscurin-RhoGEF). | |||||
|
FATH_HUMAN
|
||||||
| NC score | 0.000300 (rank : 130) | θ value | 6.88961 (rank : 91) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 570 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14517 | Gene names | FAT | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin-related tumor suppressor homolog precursor (Protein fat homolog). | |||||
|
DRD2_MOUSE
|
||||||
| NC score | 0.000113 (rank : 131) | θ value | 6.88961 (rank : 90) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P61168, P13953 | Gene names | Drd2 | |||
|
Domain Architecture |

|
|||||
| Description | D(2) dopamine receptor (Dopamine D2 receptor). | |||||
|
NEK1_MOUSE
|
||||||
| NC score | -0.001430 (rank : 132) | θ value | 6.88961 (rank : 95) | |||
| Query Neighborhood Hits | 128 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P51954 | Gene names | Nek1 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Nek1 (EC 2.7.11.1) (NimA-related protein kinase 1). | |||||