Please be patient as the page loads
|
SPT6H_HUMAN
|
||||||
| SwissProt Accessions | Q7KZ85, Q15737, Q6GMQ4, Q7KYW9, Q7LDK4, Q8N526, Q92775, Q96AH3, Q9BTH9, Q9BTI2 | Gene names | SUPT6H, KIAA0162, SPT6H | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor SPT6 (hSPT6) (Tat-cotransactivator 2 protein) (Tat-CT2 protein). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
SPT6H_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 104 | |
| SwissProt Accessions | Q7KZ85, Q15737, Q6GMQ4, Q7KYW9, Q7LDK4, Q8N526, Q92775, Q96AH3, Q9BTH9, Q9BTI2 | Gene names | SUPT6H, KIAA0162, SPT6H | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor SPT6 (hSPT6) (Tat-cotransactivator 2 protein) (Tat-CT2 protein). | |||||
|
SPT6H_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.989367 (rank : 2) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q62383, Q5SYM0, Q6GQS3, Q6ZQI0, Q8BQY6 | Gene names | Supt6h, Kiaa0162 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor SPT6. | |||||
|
LEO1_MOUSE
|
||||||
| θ value | 1.69304e-06 (rank : 3) | NC score | 0.154488 (rank : 3) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
LEO1_HUMAN
|
||||||
| θ value | 3.19293e-05 (rank : 4) | NC score | 0.151269 (rank : 4) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
MLL3_MOUSE
|
||||||
| θ value | 0.00869519 (rank : 5) | NC score | 0.060020 (rank : 34) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
CENPC_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 6) | NC score | 0.102327 (rank : 6) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q03188, Q9P0M5 | Gene names | CENPC1, CENPC, ICEN7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein C 1 (CENP-C) (Centromere autoantigen C) (Interphase centromere complex protein 7). | |||||
|
TCP1L_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 7) | NC score | 0.131854 (rank : 5) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8TDR4, Q96LN5 | Gene names | TCP10L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | T-complex protein 10A homolog 2 (T-complex protein 10A-2) (TCP10A-2) (TCP10-like). | |||||
|
ANKY2_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 8) | NC score | 0.061526 (rank : 32) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 316 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q3TPE9, Q8BK14, Q8BYW5, Q8R3N4, Q921J1 | Gene names | Ankmy2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat and MYND domain-containing protein 2. | |||||
|
ERG_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 9) | NC score | 0.047249 (rank : 52) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P81270, Q8C5L4, Q920K7, Q920K8, Q920K9 | Gene names | Erg, Erg-3 | |||
|
Domain Architecture |
|
|||||
| Description | Transcriptional regulator ERG. | |||||
|
IWS1_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 10) | NC score | 0.080698 (rank : 15) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
TCAL5_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 11) | NC score | 0.065042 (rank : 26) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 806 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q5H9L2 | Gene names | TCEAL5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 5 (TCEA-like protein 5) (Transcription elongation factor S-II protein-like 5). | |||||
|
RRP5_HUMAN
|
||||||
| θ value | 0.125558 (rank : 12) | NC score | 0.100606 (rank : 8) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q14690, Q86VQ8 | Gene names | PDCD11, KIAA0185 | |||
|
Domain Architecture |

|
|||||
| Description | RRP5 protein homolog (Programmed cell death protein 11). | |||||
|
CLGN_MOUSE
|
||||||
| θ value | 0.163984 (rank : 13) | NC score | 0.055649 (rank : 39) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P52194, Q9D2K5 | Gene names | Clgn, Meg1 | |||
|
Domain Architecture |
|
|||||
| Description | Calmegin precursor (MEG 1 antigen) (Calnexin-T) (A2/6). | |||||
|
NSBP1_MOUSE
|
||||||
| θ value | 0.163984 (rank : 14) | NC score | 0.100762 (rank : 7) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
TSH2_HUMAN
|
||||||
| θ value | 0.163984 (rank : 15) | NC score | 0.040531 (rank : 62) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9NRE2, Q4VXM4, Q6N003, Q8N260 | Gene names | TSHZ2, C20orf17, TSH2, ZNF218 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Teashirt homolog 2 (Zinc finger protein 218) (Ovarian cancer-related protein 10-2) (OVC10-2). | |||||
|
UBP47_HUMAN
|
||||||
| θ value | 0.163984 (rank : 16) | NC score | 0.063136 (rank : 28) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96K76, Q658U0, Q86Y73, Q8TEP6, Q9BWI0, Q9NWN1 | Gene names | USP47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 47 (EC 3.1.2.15) (Ubiquitin thioesterase 47) (Ubiquitin-specific-processing protease 47) (Deubiquitinating enzyme 47). | |||||
|
UBP47_MOUSE
|
||||||
| θ value | 0.163984 (rank : 17) | NC score | 0.061741 (rank : 30) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BY87, Q5EBP2, Q6KAR9, Q80V06, Q8BHU1, Q8BI15, Q8BI16, Q8BUW4, Q91X25 | Gene names | Usp47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 47 (EC 3.1.2.15) (Ubiquitin thioesterase 47) (Ubiquitin-specific-processing protease 47) (Deubiquitinating enzyme 47). | |||||
|
IWS1_MOUSE
|
||||||
| θ value | 0.21417 (rank : 18) | NC score | 0.077919 (rank : 17) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
MAP1A_HUMAN
|
||||||
| θ value | 0.21417 (rank : 19) | NC score | 0.052751 (rank : 43) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P78559, O95643, Q12973, Q15882, Q9UJT4 | Gene names | MAP1A | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 1A (MAP 1A) (Proliferation-related protein p80) [Contains: MAP1 light chain LC2]. | |||||
|
PEG3_MOUSE
|
||||||
| θ value | 0.21417 (rank : 20) | NC score | 0.019314 (rank : 81) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1101 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q3URU2, O54978, Q3TQ69, Q5EBP7, Q61138, Q6GQS0, Q80U47, Q8R5N0, Q9QX53 | Gene names | Peg3, Kiaa0287, Pw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Paternally expressed gene 3 protein (ASF-1). | |||||
|
DCTN2_MOUSE
|
||||||
| θ value | 0.279714 (rank : 21) | NC score | 0.082915 (rank : 13) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99KJ8 | Gene names | Dctn2 | |||
|
Domain Architecture |

|
|||||
| Description | Dynactin subunit 2 (Dynactin complex 50 kDa subunit) (50 kDa dynein- associated polypeptide) (p50 dynamitin) (DCTN-50) (Growth cone membrane protein 23-48K) (GMP23-48K). | |||||
|
DDX10_MOUSE
|
||||||
| θ value | 0.279714 (rank : 22) | NC score | 0.025555 (rank : 72) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q80Y44 | Gene names | Ddx10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX10 (EC 3.6.1.-) (DEAD box protein 10). | |||||
|
RED_HUMAN
|
||||||
| θ value | 0.279714 (rank : 23) | NC score | 0.097642 (rank : 10) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q13123 | Gene names | IK, RED, RER | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Red (Protein RER) (IK factor) (Cytokine IK). | |||||
|
RED_MOUSE
|
||||||
| θ value | 0.279714 (rank : 24) | NC score | 0.098561 (rank : 9) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Z1M8 | Gene names | Ik, Red, Rer | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Red (Protein RER) (IK factor) (Cytokine IK). | |||||
|
SEC63_HUMAN
|
||||||
| θ value | 0.365318 (rank : 25) | NC score | 0.077379 (rank : 18) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9UGP8, O95380, Q86VS9, Q8IWL0, Q9NTE0 | Gene names | SEC63, SEC63L | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
RBBP6_HUMAN
|
||||||
| θ value | 0.47712 (rank : 26) | NC score | 0.048997 (rank : 50) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
NEB1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 27) | NC score | 0.041093 (rank : 60) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 829 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9ULJ8, O76059, Q9NXT2 | Gene names | PPP1R9A, KIAA1222 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neurabin-1 (Neurabin-I) (Neural tissue-specific F-actin-binding protein I) (Protein phosphatase 1 regulatory subunit 9A). | |||||
|
PUM1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 28) | NC score | 0.039849 (rank : 64) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q14671, Q5VXY7, Q9HAN1 | Gene names | PUM1, KIAA0099, PUMH1 | |||
|
Domain Architecture |

|
|||||
| Description | Pumilio homolog 1 (Pumilio-1) (HsPUM). | |||||
|
PUM1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 29) | NC score | 0.039896 (rank : 63) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80U78, Q80X96, Q80YU8, Q8BPV7, Q9EPU6 | Gene names | Pum1, Kiaa0099 | |||
|
Domain Architecture |

|
|||||
| Description | Pumilio homolog 1. | |||||
|
ANKY2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 30) | NC score | 0.043847 (rank : 58) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 306 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8IV38, Q96BL3 | Gene names | ANKMY2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat and MYND domain-containing protein 2. | |||||
|
BANK1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 31) | NC score | 0.038198 (rank : 65) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NDB2, Q8N5K8, Q8NB56, Q8WYN5, Q9NWP2 | Gene names | BANK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell scaffold protein with ankyrin repeats. | |||||
|
CAMKV_HUMAN
|
||||||
| θ value | 1.06291 (rank : 32) | NC score | 0.009614 (rank : 105) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1084 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8NCB2, Q6FIB8, Q8NBS8, Q8NC85, Q8NDU4, Q8WTT8, Q9BQC9, Q9H0Q5 | Gene names | CAMKV | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CaM kinase-like vesicle-associated protein. | |||||
|
DCTN2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 33) | NC score | 0.060259 (rank : 33) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q13561, Q86YN2, Q9BW17 | Gene names | DCTN2, DCTN50 | |||
|
Domain Architecture |

|
|||||
| Description | Dynactin subunit 2 (Dynactin complex 50 kDa subunit) (50 kDa dynein- associated polypeptide) (p50 dynamitin) (DCTN-50). | |||||
|
MEPD_HUMAN
|
||||||
| θ value | 1.06291 (rank : 34) | NC score | 0.053361 (rank : 40) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P52888, Q9UCB3 | Gene names | THOP1 | |||
|
Domain Architecture |

|
|||||
| Description | Thimet oligopeptidase (EC 3.4.24.15) (Endopeptidase 24.15) (MP78). | |||||
|
SEC63_MOUSE
|
||||||
| θ value | 1.06291 (rank : 35) | NC score | 0.071815 (rank : 22) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 601 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8VHE0, Q8VEB9 | Gene names | Sec63, Sec63l | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
SMCA2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 36) | NC score | 0.023827 (rank : 75) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 503 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P51531 | Gene names | SMARCA2, BRM, SNF2A, SNF2L2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L2 (EC 3.6.1.-) (ATP- dependent helicase SMARCA2) (SNF2-alpha) (SWI/SNF-related matrix- associated actin-dependent regulator of chromatin subfamily A member 2) (hBRM). | |||||
|
TNF13_HUMAN
|
||||||
| θ value | 1.06291 (rank : 37) | NC score | 0.078262 (rank : 16) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O75888, Q96HV6, Q9P1M8, Q9P1M9 | Gene names | TNFSF13, APRIL, TALL2, ZTNF2 | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor ligand superfamily member 13 precursor (A proliferation-inducing ligand) (APRIL) (TNF- and APOL-related leukocyte expressed ligand 2) (TALL-2) (TNF-related death ligand 1) (TRDL-1) (CD256 antigen). | |||||
|
CC47_MOUSE
|
||||||
| θ value | 1.38821 (rank : 38) | NC score | 0.057301 (rank : 37) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9D024, Q3TSB5, Q3V1S7, Q8C5D9, Q920S6 | Gene names | Ccdc47, Asp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 47 precursor (Adipocyte-specific protein 4). | |||||
|
DMPK_HUMAN
|
||||||
| θ value | 1.81305 (rank : 39) | NC score | 0.005861 (rank : 116) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 944 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q09013, Q16205 | Gene names | DMPK, MDPK | |||
|
Domain Architecture |

|
|||||
| Description | Myotonin-protein kinase (EC 2.7.11.1) (Myotonic dystrophy protein kinase) (MDPK) (DM-kinase) (DMK) (DMPK) (MT-PK). | |||||
|
GOGB1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 40) | NC score | 0.021570 (rank : 78) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 2297 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q14789, Q14398 | Gene names | GOLGB1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily B member 1 (Giantin) (Macrogolgin) (372 kDa Golgi complex-associated protein) (GCP372). | |||||
|
HM2L1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 41) | NC score | 0.045925 (rank : 55) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UGU5, O75672, O75673, Q9UMT5 | Gene names | HMG2L1, HMGBCG | |||
|
Domain Architecture |

|
|||||
| Description | High mobility group protein 2-like 1 (Protein HMGBCG). | |||||
|
K1794_HUMAN
|
||||||
| θ value | 1.81305 (rank : 42) | NC score | 0.066836 (rank : 25) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NVI1, Q96JN1, Q96ST0, Q9BT96 | Gene names | KIAA1794 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1794. | |||||
|
MDC1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 43) | NC score | 0.030460 (rank : 69) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
MY18A_HUMAN
|
||||||
| θ value | 1.81305 (rank : 44) | NC score | 0.013272 (rank : 97) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1929 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q92614, Q8IXP8 | Gene names | MYO18A, KIAA0216, MYSPDZ | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
NEK1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 45) | NC score | 0.007673 (rank : 110) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P51954 | Gene names | Nek1 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Nek1 (EC 2.7.11.1) (NimA-related protein kinase 1). | |||||
|
NOP14_HUMAN
|
||||||
| θ value | 1.81305 (rank : 46) | NC score | 0.044063 (rank : 57) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 573 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P78316, Q7LGI5, Q7Z6K0, Q8TBR6 | Gene names | NOP14, C4orf9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable nucleolar complex protein 14. | |||||
|
SN1L2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 47) | NC score | 0.007501 (rank : 111) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 964 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9H0K1, O94878, Q6AZE2, Q76N03, Q8NCV7, Q96CZ8 | Gene names | SNF1LK2, KIAA0781, QIK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SNF1-like kinase 2 (EC 2.7.11.1) (Qin- induced kinase). | |||||
|
SOCS2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 48) | NC score | 0.051364 (rank : 46) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O14508, O14542, O95102, Q9UKS5 | Gene names | SOCS2, CIS2, SSI2, STATI2 | |||
|
Domain Architecture |
|
|||||
| Description | Suppressor of cytokine signaling 2 (SOCS-2) (Cytokine-inducible SH2 protein 2) (CIS-2) (STAT-induced STAT inhibitor 2) (SSI-2). | |||||
|
SOCS2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 49) | NC score | 0.052236 (rank : 45) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O35717 | Gene names | Socs2, Cis2, Cish2 | |||
|
Domain Architecture |
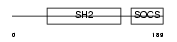
|
|||||
| Description | Suppressor of cytokine signaling 2 (SOCS-2) (Cytokine-inducible SH2 protein 2) (CIS-2). | |||||
|
GRAP2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 50) | NC score | 0.011287 (rank : 102) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 297 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O89100 | Gene names | Grap2, Gads, Grb2l, Grid, Mona | |||
|
Domain Architecture |
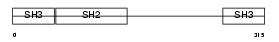
|
|||||
| Description | GRB2-related adaptor protein 2 (GADS protein) (Growth factor receptor- binding protein) (GRBLG) (GRB-2-like protein) (GRB2L) (Hematopoietic cell-associated adaptor protein GrpL) (GRB-2-related monocytic adapter protein) (Monocytic adapter) (MONA) (Adapter protein GRID). | |||||
|
MPP10_HUMAN
|
||||||
| θ value | 2.36792 (rank : 51) | NC score | 0.081141 (rank : 14) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 306 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O00566 | Gene names | MPHOSPH10, MPP10 | |||
|
Domain Architecture |

|
|||||
| Description | U3 small nucleolar ribonucleoprotein protein MPP10 (M phase phosphoprotein 10). | |||||
|
TCP10_HUMAN
|
||||||
| θ value | 2.36792 (rank : 52) | NC score | 0.061678 (rank : 31) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q12799 | Gene names | TCP10, TCP10A | |||
|
Domain Architecture |

|
|||||
| Description | T-complex protein 10A homolog. | |||||
|
CCDC4_HUMAN
|
||||||
| θ value | 3.0926 (rank : 53) | NC score | 0.058611 (rank : 35) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6ZU67, Q58A26, Q58A27 | Gene names | CCDC4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 4. | |||||
|
IFIT5_HUMAN
|
||||||
| θ value | 3.0926 (rank : 54) | NC score | 0.033741 (rank : 68) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13325 | Gene names | IFIT5, RI58 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon-induced protein with tetratricopeptide repeats 5 (IFIT-5) (Retinoic acid- and interferon-inducible 58 kDa protein). | |||||
|
SETD2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 55) | NC score | 0.047248 (rank : 53) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9BYW2, O75397, O75405, Q17RW8, Q5BKS9, Q5QGN2, Q69YI5, Q6IN64, Q6ZN53, Q6ZS25, Q8N3R0, Q8TCN0, Q9C0D1, Q9H696, Q9NZW9 | Gene names | SETD2, HIF1, HYPB, KIAA1732, SET2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36-specific (EC 2.1.1.43) (SET domain-containing protein 2) (hSET2) (Huntingtin- interacting protein HYPB) (Huntingtin-interacting protein 1) (HIF-1) (p231HBP). | |||||
|
SF04_MOUSE
|
||||||
| θ value | 3.0926 (rank : 56) | NC score | 0.044228 (rank : 56) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8CH02, Q8R094 | Gene names | Sf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor 4. | |||||
|
BICD2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 57) | NC score | 0.016398 (rank : 87) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 856 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8TD16, O75181, Q5TBQ2, Q5TBQ3, Q96LH2, Q9BT84, Q9H561 | Gene names | BICD2, KIAA0699 | |||
|
Domain Architecture |
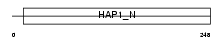
|
|||||
| Description | Cytoskeleton-like bicaudal D protein homolog 2. | |||||
|
BICD2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 58) | NC score | 0.016123 (rank : 88) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 864 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q921C5, Q80TU1, Q8BTE3, Q9DCL3 | Gene names | Bicd2, Kiaa0699 | |||
|
Domain Architecture |
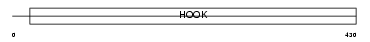
|
|||||
| Description | Cytoskeleton-like bicaudal D protein homolog 2. | |||||
|
CCD21_HUMAN
|
||||||
| θ value | 4.03905 (rank : 59) | NC score | 0.016899 (rank : 85) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1228 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q6P2H3, Q5VY68, Q5VY70, Q9H6Q1, Q9H828, Q9UF52 | Gene names | CCDC21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 21. | |||||
|
CE152_HUMAN
|
||||||
| θ value | 4.03905 (rank : 60) | NC score | 0.022896 (rank : 77) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 976 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O94986, Q6NTA0 | Gene names | CEP152, KIAA0912 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 152 kDa (Cep152 protein). | |||||
|
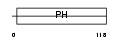
GAB2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 61) | NC score | 0.023767 (rank : 76) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Z1S8, Q9R1X3 | Gene names | Gab2 | |||
|
Domain Architecture |
|
|||||
| Description | GRB2-associated-binding protein 2 (GRB2-associated binder 2) (PH domain-containing adaptor molecule p97). | |||||
|
MPP10_MOUSE
|
||||||
| θ value | 4.03905 (rank : 62) | NC score | 0.058301 (rank : 36) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q810V0 | Gene names | Mphosph10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U3 small nucleolar ribonucleoprotein protein MPP10 (M phase phosphoprotein 10). | |||||
|
N4BP2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 63) | NC score | 0.028385 (rank : 70) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q86UW6, Q9NVK2, Q9P2D4 | Gene names | N4BP2, B3BP, KIAA1413 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-binding protein 2 (EC 3.-.-.-) (N4BP2) (BCL-3-binding protein). | |||||
|
NPFF_HUMAN
|
||||||
| θ value | 4.03905 (rank : 64) | NC score | 0.064494 (rank : 27) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 5 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O15130 | Gene names | NPFF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FMRFamide-related peptides precursor [Contains: Neuropeptide SF (NPSF); Neuropeptide FF (NPFF); Neuropeptide AF (NPAF)]. | |||||
|
RPGR_HUMAN
|
||||||
| θ value | 4.03905 (rank : 65) | NC score | 0.037409 (rank : 66) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q92834, O00702, O00737, Q8N5T6, Q93039, Q9HD29, Q9UMR1 | Gene names | RPGR, RP3, XLRP3 | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator. | |||||
|
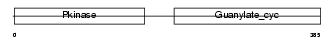
ANPRA_HUMAN
|
||||||
| θ value | 5.27518 (rank : 66) | NC score | 0.006549 (rank : 114) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 612 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P16066, Q6P4Q3 | Gene names | NPR1, ANPRA | |||
|
Domain Architecture |
|
|||||
| Description | Atrial natriuretic peptide receptor A precursor (ANP-A) (ANPRA) (GC-A) (Guanylate cyclase) (EC 4.6.1.2) (NPR-A) (Atrial natriuretic peptide A-type receptor). | |||||
|
ANPRA_MOUSE
|
||||||
| θ value | 5.27518 (rank : 67) | NC score | 0.006591 (rank : 113) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 600 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P18293 | Gene names | Npr1, Npra | |||
|
Domain Architecture |

|
|||||
| Description | Atrial natriuretic peptide receptor A precursor (ANP-A) (ANPRA) (GC-A) (Guanylate cyclase) (EC 4.6.1.2) (NPR-A) (Atrial natriuretic peptide A-type receptor). | |||||
|
DLX3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 68) | NC score | 0.008630 (rank : 108) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 367 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q64205 | Gene names | Dlx3 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein DLX-3. | |||||
|
DMXL2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 69) | NC score | 0.015508 (rank : 90) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8TDJ6, O94938 | Gene names | DMXL2, KIAA0856 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DmX-like 2 (Rabconnectin-3). | |||||
|
HIPK2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 70) | NC score | 0.006289 (rank : 115) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 871 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9QZR5, O88905, Q99P45, Q99P46, Q9D2E6, Q9D474, Q9EQL2, Q9QZR4 | Gene names | Hipk2, Nbak1, Stank | |||
|
Domain Architecture |

|
|||||
| Description | Homeodomain-interacting protein kinase 2 (EC 2.7.11.1) (Nuclear body- associated kinase 1) (Sialophorin tail-associated nuclear serine/threonine-protein kinase). | |||||
|
KTN1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 71) | NC score | 0.013061 (rank : 98) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1324 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q61595, Q8BG49, Q8BHF4, Q8BHM8, Q8C9Y5, Q8CG51, Q8CG52, Q8CG53, Q8CG54, Q8CG55, Q8CG56, Q8CG57, Q8CG58, Q8CG59, Q8CG60, Q8CG61, Q8CG62, Q8CG63 | Gene names | Ktn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinectin. | |||||
|
NASP_HUMAN
|
||||||
| θ value | 5.27518 (rank : 72) | NC score | 0.046584 (rank : 54) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P49321, Q96A69, Q9BTW2 | Gene names | NASP | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear autoantigenic sperm protein (NASP). | |||||
|
NEST_HUMAN
|
||||||
| θ value | 5.27518 (rank : 73) | NC score | 0.026006 (rank : 71) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1305 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P48681, O00552 | Gene names | NES | |||
|
Domain Architecture |

|
|||||
| Description | Nestin. | |||||
|
PRRT3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 74) | NC score | 0.013001 (rank : 99) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q5FWE3, Q49AD0, Q6UXY6, Q8NBC9 | Gene names | PRRT3, UNQ5823/PRO19642 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
SM1L2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 75) | NC score | 0.018213 (rank : 83) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1019 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8NDV3, Q5TIC3, Q9Y3G5 | Gene names | SMC1L2 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 2 protein (SMC1beta protein). | |||||
|
AKAP2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 76) | NC score | 0.016777 (rank : 86) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O54931, O54932, O54933 | Gene names | Akap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 2 (Protein kinase A-anchoring protein 2) (PRKA2) (AKAP-2) (AKAP expressed in kidney and lung) (AKAP-KL). | |||||
|
CASP_HUMAN
|
||||||
| θ value | 6.88961 (rank : 77) | NC score | 0.014012 (rank : 95) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1131 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q13948, Q53GU9, Q8TBS3 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein CASP. | |||||
|
CCD16_MOUSE
|
||||||
| θ value | 6.88961 (rank : 78) | NC score | 0.019954 (rank : 80) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8R1N0, Q3TR52, Q9CWV9, Q9CYI6 | Gene names | Ccdc16, Omcg1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 16 (Ovus mutant candidate gene 1 protein). | |||||
|
CTRO_HUMAN
|
||||||
| θ value | 6.88961 (rank : 79) | NC score | 0.009981 (rank : 103) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 2092 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O14578, Q6XUH8, Q86UQ9, Q9UPZ7 | Gene names | CIT, KIAA0949, STK21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Citron Rho-interacting kinase (EC 2.7.11.1) (CRIK) (Rho-interacting, serine/threonine-protein kinase 21). | |||||
|
CUTL1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 80) | NC score | 0.014736 (rank : 93) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1574 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P39880, Q6NYH4, Q9UEV5 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP). | |||||
|
ERIC1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 81) | NC score | 0.076851 (rank : 19) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q86X53, Q9P063 | Gene names | ERICH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate-rich protein 1. | |||||
|
FA21C_HUMAN
|
||||||
| θ value | 6.88961 (rank : 82) | NC score | 0.050217 (rank : 49) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9Y4E1, Q5SQU4, Q5SQU5, Q7L521, Q9UG79 | Gene names | FAM21C, KIAA0592 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM21C. | |||||
|
GRP78_HUMAN
|
||||||
| θ value | 6.88961 (rank : 83) | NC score | 0.008794 (rank : 106) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P11021, Q2EF78, Q9NPF1, Q9UK02 | Gene names | HSPA5, GRP78 | |||
|
Domain Architecture |

|
|||||
| Description | 78 kDa glucose-regulated protein precursor (GRP 78) (Heat shock 70 kDa protein 5) (Immunoglobulin heavy chain-binding protein) (BiP) (Endoplasmic reticulum lumenal Ca(2+)-binding protein grp78). | |||||
|
GRP78_MOUSE
|
||||||
| θ value | 6.88961 (rank : 84) | NC score | 0.008660 (rank : 107) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P20029, O35642, Q3UFF2, Q61630 | Gene names | Hspa5, Grp78 | |||
|
Domain Architecture |

|
|||||
| Description | 78 kDa glucose-regulated protein precursor (GRP 78) (Heat shock 70 kDa protein 5) (Immunoglobulin heavy chain-binding protein) (BiP). | |||||
|
GTF2I_HUMAN
|
||||||
| θ value | 6.88961 (rank : 85) | NC score | 0.014744 (rank : 92) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P78347, O14743, O15359, O43546, O43588, O43589, Q9BSZ4 | Gene names | GTF2I, BAP135, WBSCR6 | |||
|
Domain Architecture |

|
|||||
| Description | General transcription factor II-I (GTFII-I) (TFII-I) (Bruton tyrosine kinase-associated protein 135) (BTK-associated protein 135) (BAP-135) (SRF-Phox1-interacting protein) (SPIN) (Williams-Beuren syndrome chromosome region 6 protein). | |||||
|
K1543_HUMAN
|
||||||
| θ value | 6.88961 (rank : 86) | NC score | 0.025379 (rank : 73) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9P1Y5, Q8NDF1 | Gene names | KIAA1543 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1543. | |||||
|
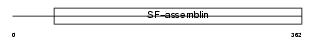
K1H1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 87) | NC score | 0.006976 (rank : 112) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q15323 | Gene names | KRTHA1, HHA1, HKA1 | |||
|
Domain Architecture |
|
|||||
| Description | Keratin, type I cuticular Ha1 (Hair keratin, type I Ha1). | |||||
|
MLH1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 88) | NC score | 0.014681 (rank : 94) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9JK91, Q62454 | Gene names | Mlh1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA mismatch repair protein Mlh1 (MutL protein homolog 1). | |||||
|
P85B_HUMAN
|
||||||
| θ value | 6.88961 (rank : 89) | NC score | 0.016994 (rank : 84) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O00459, Q9UPH9 | Gene names | PIK3R2 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 3-kinase regulatory subunit beta (PI3-kinase p85- subunit beta) (PtdIns-3-kinase p85-beta). | |||||
|
PDK4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 90) | NC score | 0.015912 (rank : 89) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O70571 | Gene names | Pdk4 | |||
|
Domain Architecture |

|
|||||
| Description | [Pyruvate dehydrogenase [lipoamide]] kinase isozyme 4, mitochondrial precursor (EC 2.7.11.2) (Pyruvate dehydrogenase kinase isoform 4). | |||||
|
PQBP1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 91) | NC score | 0.048460 (rank : 51) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O60828, Q4VY25, Q4VY26, Q4VY27, Q4VY29, Q4VY30, Q4VY34, Q4VY35, Q4VY36, Q4VY37, Q4VY38, Q9GZP2, Q9GZU4, Q9GZZ4 | Gene names | PQBP1, NPW38 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyglutamine-binding protein 1 (Polyglutamine tract-binding protein 1) (PQBP-1) (38 kDa nuclear protein containing a WW domain) (Npw38). | |||||
|
SF04_HUMAN
|
||||||
| θ value | 6.88961 (rank : 92) | NC score | 0.034846 (rank : 67) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8IWZ8, O60378, Q6P3X9, Q8TCQ4, Q8WWT4, Q8WWT5, Q9NTG3 | Gene names | SF4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor 4 (RNA-binding protein RBP). | |||||
|
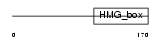
SOX9_HUMAN
|
||||||
| θ value | 6.88961 (rank : 93) | NC score | 0.011351 (rank : 101) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P48436 | Gene names | SOX9 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor SOX-9. | |||||
|
SRCA_HUMAN
|
||||||
| θ value | 6.88961 (rank : 94) | NC score | 0.043639 (rank : 59) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q86TD4 | Gene names | SRL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcalumenin precursor. | |||||
|
TIF1G_HUMAN
|
||||||
| θ value | 6.88961 (rank : 95) | NC score | 0.009832 (rank : 104) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 410 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UPN9, O95855, Q9C017, Q9UJ79 | Gene names | TRIM33, KIAA1113, RFG7, TIF1G | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-gamma (TIF1-gamma) (Tripartite motif-containing protein 33) (RET-fused gene 7 protein) (Rfg7 protein). | |||||
|
CMAH_MOUSE
|
||||||
| θ value | 8.99809 (rank : 96) | NC score | 0.013434 (rank : 96) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q61419, Q7TMR9, Q8JZM9 | Gene names | Cmah | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytidine monophosphate-N-acetylneuraminic acid hydroxylase precursor (EC 1.14.18.2) (CMP-N-acetylneuraminate monooxygenase) (CMP-N- acetylneuraminic acid hydroxylase) (CMP-NeuAc hydroxylase) (CMP-Neu5Ac hydroxylase). | |||||
|
IFT74_MOUSE
|
||||||
| θ value | 8.99809 (rank : 97) | NC score | 0.012622 (rank : 100) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 939 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8BKE9, Q80ZI3, Q9CSY1, Q9CUS0, Q9D9T5 | Gene names | Ift74, Ccdc2, Cmg1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intraflagellar transport 74 homolog (Coiled-coil domain-containing protein 2) (Capillary morphogenesis protein 1) (CMG-1). | |||||
|
M3K9_HUMAN
|
||||||
| θ value | 8.99809 (rank : 98) | NC score | 0.004070 (rank : 117) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1123 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P80192, Q6EH31, Q9H2N5 | Gene names | MAP3K9, MLK1, PRKE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase 9 (EC 2.7.11.25) (Mixed lineage kinase 1). | |||||
|
NKTR_HUMAN
|
||||||
| θ value | 8.99809 (rank : 99) | NC score | 0.024351 (rank : 74) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 658 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P30414 | Gene names | NKTR | |||
|
Domain Architecture |

|
|||||
| Description | NK-tumor recognition protein (Natural-killer cells cyclophilin-related protein) (NK-TR protein). | |||||
|
PF21B_HUMAN
|
||||||
| θ value | 8.99809 (rank : 100) | NC score | 0.014759 (rank : 91) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96EK2, Q5TFL2, Q6ICC0 | Gene names | PHF21B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
PININ_MOUSE
|
||||||
| θ value | 8.99809 (rank : 101) | NC score | 0.040646 (rank : 61) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 777 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O35691, Q8CD89, Q8CGU3 | Gene names | Pnn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pinin. | |||||
|
TITIN_HUMAN
|
||||||
| θ value | 8.99809 (rank : 102) | NC score | 0.008556 (rank : 109) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 2923 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8WZ42, Q10465, Q10466, Q15598, Q2XUS3, Q32Q60, Q4U1Z6, Q6NSG0, Q6PDB1, Q6PJP0, Q7KYM2, Q7KYN4, Q7KYN5, Q7LDM3, Q7Z2X3, Q8TCG8, Q8WZ51, Q8WZ52, Q8WZ53, Q8WZB3, Q92761, Q92762, Q9UD97, Q9UP84, Q9Y6L9 | Gene names | TTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Titin (EC 2.7.11.1) (Connectin) (Rhabdomyosarcoma antigen MU-RMS- 40.14). | |||||
|
TULP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 103) | NC score | 0.021359 (rank : 79) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9Z273 | Gene names | Tulp1 | |||
|
Domain Architecture |

|
|||||
| Description | Tubby-related protein 1 (Tubby-like protein 1). | |||||
|
VPS72_MOUSE
|
||||||
| θ value | 8.99809 (rank : 104) | NC score | 0.018907 (rank : 82) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q62481, Q3U2N4, Q810A9, Q99K81 | Gene names | Vps72, Tcfl1, Yl1 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 72 homolog (Transcription factor-like 1) (Protein YL-1). | |||||
|
CYLC2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.053143 (rank : 42) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q14093 | Gene names | CYLC2, CYL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-2 (Cylicin II) (Multiple-band polypeptide II). | |||||
|
DMP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 106) | NC score | 0.051027 (rank : 47) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q13316, O43265 | Gene names | DMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1). | |||||
|
DMP1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 107) | NC score | 0.050582 (rank : 48) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O55188 | Gene names | Dmp1, Dmp | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1) (AG1). | |||||
|
DSPP_MOUSE
|
||||||
| θ value | θ > 10 (rank : 108) | NC score | 0.053267 (rank : 41) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
HTSF1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 109) | NC score | 0.052408 (rank : 44) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8BGC0, Q1WWK0, Q9CT41, Q9DAU3 | Gene names | Htatsf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIV Tat-specific factor 1 homolog. | |||||
|
NSBP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 110) | NC score | 0.072454 (rank : 21) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P82970 | Gene names | NSBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1. | |||||
|
NUCKS_HUMAN
|
||||||
| θ value | θ > 10 (rank : 111) | NC score | 0.062120 (rank : 29) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9H1E3, Q9H1D6, Q9H723 | Gene names | NUCKS1, NUCKS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (P1). | |||||
|
NUCKS_MOUSE
|
||||||
| θ value | θ > 10 (rank : 112) | NC score | 0.067860 (rank : 24) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q80XU3, Q8BVD8 | Gene names | Nucks1, Nucks | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (JC7). | |||||
|
PTMA_HUMAN
|
||||||
| θ value | θ > 10 (rank : 113) | NC score | 0.094918 (rank : 11) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P06454, Q15249, Q15592 | Gene names | PTMA, TMSA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha-1]. | |||||
|
PTMA_MOUSE
|
||||||
| θ value | θ > 10 (rank : 114) | NC score | 0.092844 (rank : 12) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P26350, Q3UQV6 | Gene names | Ptma | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha]. | |||||
|
PTMS_HUMAN
|
||||||
| θ value | θ > 10 (rank : 115) | NC score | 0.069624 (rank : 23) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P20962 | Gene names | PTMS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parathymosin. | |||||
|
PTMS_MOUSE
|
||||||
| θ value | θ > 10 (rank : 116) | NC score | 0.073877 (rank : 20) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9D0J8 | Gene names | Ptms | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parathymosin. | |||||
|
SRCH_HUMAN
|
||||||
| θ value | θ > 10 (rank : 117) | NC score | 0.056108 (rank : 38) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P23327, Q504Y6 | Gene names | HRC, HCP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcoplasmic reticulum histidine-rich calcium-binding protein precursor. | |||||
|
SPT6H_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 104 | |
| SwissProt Accessions | Q7KZ85, Q15737, Q6GMQ4, Q7KYW9, Q7LDK4, Q8N526, Q92775, Q96AH3, Q9BTH9, Q9BTI2 | Gene names | SUPT6H, KIAA0162, SPT6H | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor SPT6 (hSPT6) (Tat-cotransactivator 2 protein) (Tat-CT2 protein). | |||||
|
SPT6H_MOUSE
|
||||||
| NC score | 0.989367 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q62383, Q5SYM0, Q6GQS3, Q6ZQI0, Q8BQY6 | Gene names | Supt6h, Kiaa0162 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor SPT6. | |||||
|
LEO1_MOUSE
|
||||||
| NC score | 0.154488 (rank : 3) | θ value | 1.69304e-06 (rank : 3) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
LEO1_HUMAN
|
||||||
| NC score | 0.151269 (rank : 4) | θ value | 3.19293e-05 (rank : 4) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
TCP1L_HUMAN
|
||||||
| NC score | 0.131854 (rank : 5) | θ value | 0.0563607 (rank : 7) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8TDR4, Q96LN5 | Gene names | TCP10L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | T-complex protein 10A homolog 2 (T-complex protein 10A-2) (TCP10A-2) (TCP10-like). | |||||
|
CENPC_HUMAN
|
||||||
| NC score | 0.102327 (rank : 6) | θ value | 0.0431538 (rank : 6) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q03188, Q9P0M5 | Gene names | CENPC1, CENPC, ICEN7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein C 1 (CENP-C) (Centromere autoantigen C) (Interphase centromere complex protein 7). | |||||
|
NSBP1_MOUSE
|
||||||
| NC score | 0.100762 (rank : 7) | θ value | 0.163984 (rank : 14) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
RRP5_HUMAN
|
||||||
| NC score | 0.100606 (rank : 8) | θ value | 0.125558 (rank : 12) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q14690, Q86VQ8 | Gene names | PDCD11, KIAA0185 | |||
|
Domain Architecture |

|
|||||
| Description | RRP5 protein homolog (Programmed cell death protein 11). | |||||
|
RED_MOUSE
|
||||||
| NC score | 0.098561 (rank : 9) | θ value | 0.279714 (rank : 24) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Z1M8 | Gene names | Ik, Red, Rer | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Red (Protein RER) (IK factor) (Cytokine IK). | |||||
|
RED_HUMAN
|
||||||
| NC score | 0.097642 (rank : 10) | θ value | 0.279714 (rank : 23) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q13123 | Gene names | IK, RED, RER | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Red (Protein RER) (IK factor) (Cytokine IK). | |||||
|
PTMA_HUMAN
|
||||||
| NC score | 0.094918 (rank : 11) | θ value | θ > 10 (rank : 113) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P06454, Q15249, Q15592 | Gene names | PTMA, TMSA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha-1]. | |||||
|
PTMA_MOUSE
|
||||||
| NC score | 0.092844 (rank : 12) | θ value | θ > 10 (rank : 114) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P26350, Q3UQV6 | Gene names | Ptma | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha]. | |||||
|
DCTN2_MOUSE
|
||||||
| NC score | 0.082915 (rank : 13) | θ value | 0.279714 (rank : 21) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99KJ8 | Gene names | Dctn2 | |||
|
Domain Architecture |

|
|||||
| Description | Dynactin subunit 2 (Dynactin complex 50 kDa subunit) (50 kDa dynein- associated polypeptide) (p50 dynamitin) (DCTN-50) (Growth cone membrane protein 23-48K) (GMP23-48K). | |||||
|
MPP10_HUMAN
|
||||||
| NC score | 0.081141 (rank : 14) | θ value | 2.36792 (rank : 51) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 306 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O00566 | Gene names | MPHOSPH10, MPP10 | |||
|
Domain Architecture |

|
|||||
| Description | U3 small nucleolar ribonucleoprotein protein MPP10 (M phase phosphoprotein 10). | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.080698 (rank : 15) | θ value | 0.0736092 (rank : 10) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
TNF13_HUMAN
|
||||||
| NC score | 0.078262 (rank : 16) | θ value | 1.06291 (rank : 37) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O75888, Q96HV6, Q9P1M8, Q9P1M9 | Gene names | TNFSF13, APRIL, TALL2, ZTNF2 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor ligand superfamily member 13 precursor (A proliferation-inducing ligand) (APRIL) (TNF- and APOL-related leukocyte expressed ligand 2) (TALL-2) (TNF-related death ligand 1) (TRDL-1) (CD256 antigen). | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.077919 (rank : 17) | θ value | 0.21417 (rank : 18) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
SEC63_HUMAN
|
||||||
| NC score | 0.077379 (rank : 18) | θ value | 0.365318 (rank : 25) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9UGP8, O95380, Q86VS9, Q8IWL0, Q9NTE0 | Gene names | SEC63, SEC63L | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
ERIC1_HUMAN
|
||||||
| NC score | 0.076851 (rank : 19) | θ value | 6.88961 (rank : 81) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q86X53, Q9P063 | Gene names | ERICH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate-rich protein 1. | |||||
|
PTMS_MOUSE
|
||||||
| NC score | 0.073877 (rank : 20) | θ value | θ > 10 (rank : 116) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9D0J8 | Gene names | Ptms | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parathymosin. | |||||
|
NSBP1_HUMAN
|
||||||
| NC score | 0.072454 (rank : 21) | θ value | θ > 10 (rank : 110) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P82970 | Gene names | NSBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1. | |||||
|
SEC63_MOUSE
|
||||||
| NC score | 0.071815 (rank : 22) | θ value | 1.06291 (rank : 35) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 601 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8VHE0, Q8VEB9 | Gene names | Sec63, Sec63l | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
PTMS_HUMAN
|
||||||
| NC score | 0.069624 (rank : 23) | θ value | θ > 10 (rank : 115) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P20962 | Gene names | PTMS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parathymosin. | |||||
|
NUCKS_MOUSE
|
||||||
| NC score | 0.067860 (rank : 24) | θ value | θ > 10 (rank : 112) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q80XU3, Q8BVD8 | Gene names | Nucks1, Nucks | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (JC7). | |||||
|
K1794_HUMAN
|
||||||
| NC score | 0.066836 (rank : 25) | θ value | 1.81305 (rank : 42) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NVI1, Q96JN1, Q96ST0, Q9BT96 | Gene names | KIAA1794 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1794. | |||||
|
TCAL5_HUMAN
|
||||||
| NC score | 0.065042 (rank : 26) | θ value | 0.0736092 (rank : 11) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 806 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q5H9L2 | Gene names | TCEAL5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 5 (TCEA-like protein 5) (Transcription elongation factor S-II protein-like 5). | |||||
|
NPFF_HUMAN
|
||||||
| NC score | 0.064494 (rank : 27) | θ value | 4.03905 (rank : 64) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 5 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O15130 | Gene names | NPFF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FMRFamide-related peptides precursor [Contains: Neuropeptide SF (NPSF); Neuropeptide FF (NPFF); Neuropeptide AF (NPAF)]. | |||||
|
UBP47_HUMAN
|
||||||
| NC score | 0.063136 (rank : 28) | θ value | 0.163984 (rank : 16) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96K76, Q658U0, Q86Y73, Q8TEP6, Q9BWI0, Q9NWN1 | Gene names | USP47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 47 (EC 3.1.2.15) (Ubiquitin thioesterase 47) (Ubiquitin-specific-processing protease 47) (Deubiquitinating enzyme 47). | |||||
|
NUCKS_HUMAN
|
||||||
| NC score | 0.062120 (rank : 29) | θ value | θ > 10 (rank : 111) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9H1E3, Q9H1D6, Q9H723 | Gene names | NUCKS1, NUCKS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (P1). | |||||
|
UBP47_MOUSE
|
||||||
| NC score | 0.061741 (rank : 30) | θ value | 0.163984 (rank : 17) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BY87, Q5EBP2, Q6KAR9, Q80V06, Q8BHU1, Q8BI15, Q8BI16, Q8BUW4, Q91X25 | Gene names | Usp47 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 47 (EC 3.1.2.15) (Ubiquitin thioesterase 47) (Ubiquitin-specific-processing protease 47) (Deubiquitinating enzyme 47). | |||||
|
TCP10_HUMAN
|
||||||
| NC score | 0.061678 (rank : 31) | θ value | 2.36792 (rank : 52) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q12799 | Gene names | TCP10, TCP10A | |||
|
Domain Architecture |

|
|||||
| Description | T-complex protein 10A homolog. | |||||
|
ANKY2_MOUSE
|
||||||
| NC score | 0.061526 (rank : 32) | θ value | 0.0736092 (rank : 8) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 316 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q3TPE9, Q8BK14, Q8BYW5, Q8R3N4, Q921J1 | Gene names | Ankmy2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat and MYND domain-containing protein 2. | |||||
|
DCTN2_HUMAN
|
||||||
| NC score | 0.060259 (rank : 33) | θ value | 1.06291 (rank : 33) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q13561, Q86YN2, Q9BW17 | Gene names | DCTN2, DCTN50 | |||
|
Domain Architecture |

|
|||||
| Description | Dynactin subunit 2 (Dynactin complex 50 kDa subunit) (50 kDa dynein- associated polypeptide) (p50 dynamitin) (DCTN-50). | |||||
|
MLL3_MOUSE
|
||||||
| NC score | 0.060020 (rank : 34) | θ value | 0.00869519 (rank : 5) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
CCDC4_HUMAN
|
||||||
| NC score | 0.058611 (rank : 35) | θ value | 3.0926 (rank : 53) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6ZU67, Q58A26, Q58A27 | Gene names | CCDC4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 4. | |||||
|
MPP10_MOUSE
|
||||||
| NC score | 0.058301 (rank : 36) | θ value | 4.03905 (rank : 62) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q810V0 | Gene names | Mphosph10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U3 small nucleolar ribonucleoprotein protein MPP10 (M phase phosphoprotein 10). | |||||
|
CC47_MOUSE
|
||||||
| NC score | 0.057301 (rank : 37) | θ value | 1.38821 (rank : 38) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9D024, Q3TSB5, Q3V1S7, Q8C5D9, Q920S6 | Gene names | Ccdc47, Asp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 47 precursor (Adipocyte-specific protein 4). | |||||
|
SRCH_HUMAN
|
||||||
| NC score | 0.056108 (rank : 38) | θ value | θ > 10 (rank : 117) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P23327, Q504Y6 | Gene names | HRC, HCP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcoplasmic reticulum histidine-rich calcium-binding protein precursor. | |||||
|
CLGN_MOUSE
|
||||||
| NC score | 0.055649 (rank : 39) | θ value | 0.163984 (rank : 13) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P52194, Q9D2K5 | Gene names | Clgn, Meg1 | |||
|
Domain Architecture |

|
|||||
| Description | Calmegin precursor (MEG 1 antigen) (Calnexin-T) (A2/6). | |||||
|
MEPD_HUMAN
|
||||||
| NC score | 0.053361 (rank : 40) | θ value | 1.06291 (rank : 34) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P52888, Q9UCB3 | Gene names | THOP1 | |||
|
Domain Architecture |

|
|||||
| Description | Thimet oligopeptidase (EC 3.4.24.15) (Endopeptidase 24.15) (MP78). | |||||
|
DSPP_MOUSE
|
||||||
| NC score | 0.053267 (rank : 41) | θ value | θ > 10 (rank : 108) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
CYLC2_HUMAN
|
||||||
| NC score | 0.053143 (rank : 42) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q14093 | Gene names | CYLC2, CYL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-2 (Cylicin II) (Multiple-band polypeptide II). | |||||
|
MAP1A_HUMAN
|
||||||
| NC score | 0.052751 (rank : 43) | θ value | 0.21417 (rank : 19) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P78559, O95643, Q12973, Q15882, Q9UJT4 | Gene names | MAP1A | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 1A (MAP 1A) (Proliferation-related protein p80) [Contains: MAP1 light chain LC2]. | |||||
|
HTSF1_MOUSE
|
||||||
| NC score | 0.052408 (rank : 44) | θ value | θ > 10 (rank : 109) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8BGC0, Q1WWK0, Q9CT41, Q9DAU3 | Gene names | Htatsf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIV Tat-specific factor 1 homolog. | |||||
|
SOCS2_MOUSE
|
||||||
| NC score | 0.052236 (rank : 45) | θ value | 1.81305 (rank : 49) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O35717 | Gene names | Socs2, Cis2, Cish2 | |||
|
Domain Architecture |

|
|||||
| Description | Suppressor of cytokine signaling 2 (SOCS-2) (Cytokine-inducible SH2 protein 2) (CIS-2). | |||||
|
SOCS2_HUMAN
|
||||||
| NC score | 0.051364 (rank : 46) | θ value | 1.81305 (rank : 48) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O14508, O14542, O95102, Q9UKS5 | Gene names | SOCS2, CIS2, SSI2, STATI2 | |||
|
Domain Architecture |

|
|||||
| Description | Suppressor of cytokine signaling 2 (SOCS-2) (Cytokine-inducible SH2 protein 2) (CIS-2) (STAT-induced STAT inhibitor 2) (SSI-2). | |||||
|
DMP1_HUMAN
|
||||||
| NC score | 0.051027 (rank : 47) | θ value | θ > 10 (rank : 106) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q13316, O43265 | Gene names | DMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1). | |||||
|
DMP1_MOUSE
|
||||||
| NC score | 0.050582 (rank : 48) | θ value | θ > 10 (rank : 107) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O55188 | Gene names | Dmp1, Dmp | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1) (AG1). | |||||
|
FA21C_HUMAN
|
||||||
| NC score | 0.050217 (rank : 49) | θ value | 6.88961 (rank : 82) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9Y4E1, Q5SQU4, Q5SQU5, Q7L521, Q9UG79 | Gene names | FAM21C, KIAA0592 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM21C. | |||||
|
RBBP6_HUMAN
|
||||||
| NC score | 0.048997 (rank : 50) | θ value | 0.47712 (rank : 26) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
PQBP1_HUMAN
|
||||||
| NC score | 0.048460 (rank : 51) | θ value | 6.88961 (rank : 91) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O60828, Q4VY25, Q4VY26, Q4VY27, Q4VY29, Q4VY30, Q4VY34, Q4VY35, Q4VY36, Q4VY37, Q4VY38, Q9GZP2, Q9GZU4, Q9GZZ4 | Gene names | PQBP1, NPW38 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyglutamine-binding protein 1 (Polyglutamine tract-binding protein 1) (PQBP-1) (38 kDa nuclear protein containing a WW domain) (Npw38). | |||||
|
ERG_MOUSE
|
||||||
| NC score | 0.047249 (rank : 52) | θ value | 0.0736092 (rank : 9) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P81270, Q8C5L4, Q920K7, Q920K8, Q920K9 | Gene names | Erg, Erg-3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ERG. | |||||
|
SETD2_HUMAN
|
||||||
| NC score | 0.047248 (rank : 53) | θ value | 3.0926 (rank : 55) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9BYW2, O75397, O75405, Q17RW8, Q5BKS9, Q5QGN2, Q69YI5, Q6IN64, Q6ZN53, Q6ZS25, Q8N3R0, Q8TCN0, Q9C0D1, Q9H696, Q9NZW9 | Gene names | SETD2, HIF1, HYPB, KIAA1732, SET2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36-specific (EC 2.1.1.43) (SET domain-containing protein 2) (hSET2) (Huntingtin- interacting protein HYPB) (Huntingtin-interacting protein 1) (HIF-1) (p231HBP). | |||||
|
NASP_HUMAN
|
||||||
| NC score | 0.046584 (rank : 54) | θ value | 5.27518 (rank : 72) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P49321, Q96A69, Q9BTW2 | Gene names | NASP | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear autoantigenic sperm protein (NASP). | |||||
|
HM2L1_HUMAN
|
||||||
| NC score | 0.045925 (rank : 55) | θ value | 1.81305 (rank : 41) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UGU5, O75672, O75673, Q9UMT5 | Gene names | HMG2L1, HMGBCG | |||
|
Domain Architecture |

|
|||||
| Description | High mobility group protein 2-like 1 (Protein HMGBCG). | |||||
|
SF04_MOUSE
|
||||||
| NC score | 0.044228 (rank : 56) | θ value | 3.0926 (rank : 56) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8CH02, Q8R094 | Gene names | Sf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor 4. | |||||
|
NOP14_HUMAN
|
||||||
| NC score | 0.044063 (rank : 57) | θ value | 1.81305 (rank : 46) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 573 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P78316, Q7LGI5, Q7Z6K0, Q8TBR6 | Gene names | NOP14, C4orf9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable nucleolar complex protein 14. | |||||
|
ANKY2_HUMAN
|
||||||
| NC score | 0.043847 (rank : 58) | θ value | 1.06291 (rank : 30) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 306 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8IV38, Q96BL3 | Gene names | ANKMY2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat and MYND domain-containing protein 2. | |||||
|
SRCA_HUMAN
|
||||||
| NC score | 0.043639 (rank : 59) | θ value | 6.88961 (rank : 94) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q86TD4 | Gene names | SRL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcalumenin precursor. | |||||
|
NEB1_HUMAN
|
||||||
| NC score | 0.041093 (rank : 60) | θ value | 0.62314 (rank : 27) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 829 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9ULJ8, O76059, Q9NXT2 | Gene names | PPP1R9A, KIAA1222 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neurabin-1 (Neurabin-I) (Neural tissue-specific F-actin-binding protein I) (Protein phosphatase 1 regulatory subunit 9A). | |||||
|
PININ_MOUSE
|
||||||
| NC score | 0.040646 (rank : 61) | θ value | 8.99809 (rank : 101) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 777 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O35691, Q8CD89, Q8CGU3 | Gene names | Pnn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pinin. | |||||
|
TSH2_HUMAN
|
||||||
| NC score | 0.040531 (rank : 62) | θ value | 0.163984 (rank : 15) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9NRE2, Q4VXM4, Q6N003, Q8N260 | Gene names | TSHZ2, C20orf17, TSH2, ZNF218 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Teashirt homolog 2 (Zinc finger protein 218) (Ovarian cancer-related protein 10-2) (OVC10-2). | |||||
|
PUM1_MOUSE
|
||||||
| NC score | 0.039896 (rank : 63) | θ value | 0.813845 (rank : 29) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80U78, Q80X96, Q80YU8, Q8BPV7, Q9EPU6 | Gene names | Pum1, Kiaa0099 | |||
|
Domain Architecture |

|
|||||
| Description | Pumilio homolog 1. | |||||
|
PUM1_HUMAN
|
||||||
| NC score | 0.039849 (rank : 64) | θ value | 0.813845 (rank : 28) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q14671, Q5VXY7, Q9HAN1 | Gene names | PUM1, KIAA0099, PUMH1 | |||
|
Domain Architecture |

|
|||||
| Description | Pumilio homolog 1 (Pumilio-1) (HsPUM). | |||||
|
BANK1_HUMAN
|
||||||
| NC score | 0.038198 (rank : 65) | θ value | 1.06291 (rank : 31) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NDB2, Q8N5K8, Q8NB56, Q8WYN5, Q9NWP2 | Gene names | BANK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell scaffold protein with ankyrin repeats. | |||||
|
RPGR_HUMAN
|
||||||
| NC score | 0.037409 (rank : 66) | θ value | 4.03905 (rank : 65) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q92834, O00702, O00737, Q8N5T6, Q93039, Q9HD29, Q9UMR1 | Gene names | RPGR, RP3, XLRP3 | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator. | |||||
|
SF04_HUMAN
|
||||||
| NC score | 0.034846 (rank : 67) | θ value | 6.88961 (rank : 92) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8IWZ8, O60378, Q6P3X9, Q8TCQ4, Q8WWT4, Q8WWT5, Q9NTG3 | Gene names | SF4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor 4 (RNA-binding protein RBP). | |||||
|
IFIT5_HUMAN
|
||||||
| NC score | 0.033741 (rank : 68) | θ value | 3.0926 (rank : 54) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13325 | Gene names | IFIT5, RI58 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon-induced protein with tetratricopeptide repeats 5 (IFIT-5) (Retinoic acid- and interferon-inducible 58 kDa protein). | |||||
|
MDC1_HUMAN
|
||||||
| NC score | 0.030460 (rank : 69) | θ value | 1.81305 (rank : 43) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
N4BP2_HUMAN
|
||||||
| NC score | 0.028385 (rank : 70) | θ value | 4.03905 (rank : 63) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q86UW6, Q9NVK2, Q9P2D4 | Gene names | N4BP2, B3BP, KIAA1413 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD4-binding protein 2 (EC 3.-.-.-) (N4BP2) (BCL-3-binding protein). | |||||
|
NEST_HUMAN
|
||||||
| NC score | 0.026006 (rank : 71) | θ value | 5.27518 (rank : 73) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1305 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P48681, O00552 | Gene names | NES | |||
|
Domain Architecture |

|
|||||
| Description | Nestin. | |||||
|
DDX10_MOUSE
|
||||||
| NC score | 0.025555 (rank : 72) | θ value | 0.279714 (rank : 22) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q80Y44 | Gene names | Ddx10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX10 (EC 3.6.1.-) (DEAD box protein 10). | |||||
|
K1543_HUMAN
|
||||||
| NC score | 0.025379 (rank : 73) | θ value | 6.88961 (rank : 86) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9P1Y5, Q8NDF1 | Gene names | KIAA1543 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1543. | |||||
|
NKTR_HUMAN
|
||||||
| NC score | 0.024351 (rank : 74) | θ value | 8.99809 (rank : 99) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 658 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P30414 | Gene names | NKTR | |||
|
Domain Architecture |

|
|||||
| Description | NK-tumor recognition protein (Natural-killer cells cyclophilin-related protein) (NK-TR protein). | |||||
|
SMCA2_HUMAN
|
||||||
| NC score | 0.023827 (rank : 75) | θ value | 1.06291 (rank : 36) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 503 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P51531 | Gene names | SMARCA2, BRM, SNF2A, SNF2L2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L2 (EC 3.6.1.-) (ATP- dependent helicase SMARCA2) (SNF2-alpha) (SWI/SNF-related matrix- associated actin-dependent regulator of chromatin subfamily A member 2) (hBRM). | |||||
|
GAB2_MOUSE
|
||||||
| NC score | 0.023767 (rank : 76) | θ value | 4.03905 (rank : 61) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Z1S8, Q9R1X3 | Gene names | Gab2 | |||
|
Domain Architecture |

|
|||||
| Description | GRB2-associated-binding protein 2 (GRB2-associated binder 2) (PH domain-containing adaptor molecule p97). | |||||
|
CE152_HUMAN
|
||||||
| NC score | 0.022896 (rank : 77) | θ value | 4.03905 (rank : 60) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 976 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O94986, Q6NTA0 | Gene names | CEP152, KIAA0912 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 152 kDa (Cep152 protein). | |||||
|
GOGB1_HUMAN
|
||||||
| NC score | 0.021570 (rank : 78) | θ value | 1.81305 (rank : 40) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 2297 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q14789, Q14398 | Gene names | GOLGB1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily B member 1 (Giantin) (Macrogolgin) (372 kDa Golgi complex-associated protein) (GCP372). | |||||
|
TULP1_MOUSE
|
||||||
| NC score | 0.021359 (rank : 79) | θ value | 8.99809 (rank : 103) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9Z273 | Gene names | Tulp1 | |||
|
Domain Architecture |

|
|||||
| Description | Tubby-related protein 1 (Tubby-like protein 1). | |||||
|
CCD16_MOUSE
|
||||||
| NC score | 0.019954 (rank : 80) | θ value | 6.88961 (rank : 78) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8R1N0, Q3TR52, Q9CWV9, Q9CYI6 | Gene names | Ccdc16, Omcg1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 16 (Ovus mutant candidate gene 1 protein). | |||||
|
PEG3_MOUSE
|
||||||
| NC score | 0.019314 (rank : 81) | θ value | 0.21417 (rank : 20) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1101 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q3URU2, O54978, Q3TQ69, Q5EBP7, Q61138, Q6GQS0, Q80U47, Q8R5N0, Q9QX53 | Gene names | Peg3, Kiaa0287, Pw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Paternally expressed gene 3 protein (ASF-1). | |||||
|
VPS72_MOUSE
|
||||||
| NC score | 0.018907 (rank : 82) | θ value | 8.99809 (rank : 104) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q62481, Q3U2N4, Q810A9, Q99K81 | Gene names | Vps72, Tcfl1, Yl1 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 72 homolog (Transcription factor-like 1) (Protein YL-1). | |||||
|
SM1L2_HUMAN
|
||||||
| NC score | 0.018213 (rank : 83) | θ value | 5.27518 (rank : 75) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1019 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8NDV3, Q5TIC3, Q9Y3G5 | Gene names | SMC1L2 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 2 protein (SMC1beta protein). | |||||
|
P85B_HUMAN
|
||||||
| NC score | 0.016994 (rank : 84) | θ value | 6.88961 (rank : 89) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O00459, Q9UPH9 | Gene names | PIK3R2 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 3-kinase regulatory subunit beta (PI3-kinase p85- subunit beta) (PtdIns-3-kinase p85-beta). | |||||
|
CCD21_HUMAN
|
||||||
| NC score | 0.016899 (rank : 85) | θ value | 4.03905 (rank : 59) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1228 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q6P2H3, Q5VY68, Q5VY70, Q9H6Q1, Q9H828, Q9UF52 | Gene names | CCDC21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 21. | |||||
|
AKAP2_MOUSE
|
||||||
| NC score | 0.016777 (rank : 86) | θ value | 6.88961 (rank : 76) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O54931, O54932, O54933 | Gene names | Akap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 2 (Protein kinase A-anchoring protein 2) (PRKA2) (AKAP-2) (AKAP expressed in kidney and lung) (AKAP-KL). | |||||
|
BICD2_HUMAN
|
||||||
| NC score | 0.016398 (rank : 87) | θ value | 4.03905 (rank : 57) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 856 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8TD16, O75181, Q5TBQ2, Q5TBQ3, Q96LH2, Q9BT84, Q9H561 | Gene names | BICD2, KIAA0699 | |||
|
Domain Architecture |

|
|||||
| Description | Cytoskeleton-like bicaudal D protein homolog 2. | |||||
|
BICD2_MOUSE
|
||||||
| NC score | 0.016123 (rank : 88) | θ value | 4.03905 (rank : 58) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 864 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q921C5, Q80TU1, Q8BTE3, Q9DCL3 | Gene names | Bicd2, Kiaa0699 | |||
|
Domain Architecture |

|
|||||
| Description | Cytoskeleton-like bicaudal D protein homolog 2. | |||||
|
PDK4_MOUSE
|
||||||
| NC score | 0.015912 (rank : 89) | θ value | 6.88961 (rank : 90) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O70571 | Gene names | Pdk4 | |||
|
Domain Architecture |

|
|||||
| Description | [Pyruvate dehydrogenase [lipoamide]] kinase isozyme 4, mitochondrial precursor (EC 2.7.11.2) (Pyruvate dehydrogenase kinase isoform 4). | |||||
|
DMXL2_HUMAN
|
||||||
| NC score | 0.015508 (rank : 90) | θ value | 5.27518 (rank : 69) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8TDJ6, O94938 | Gene names | DMXL2, KIAA0856 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DmX-like 2 (Rabconnectin-3). | |||||
|
PF21B_HUMAN
|
||||||
| NC score | 0.014759 (rank : 91) | θ value | 8.99809 (rank : 100) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96EK2, Q5TFL2, Q6ICC0 | Gene names | PHF21B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 21B. | |||||
|
GTF2I_HUMAN
|
||||||
| NC score | 0.014744 (rank : 92) | θ value | 6.88961 (rank : 85) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P78347, O14743, O15359, O43546, O43588, O43589, Q9BSZ4 | Gene names | GTF2I, BAP135, WBSCR6 | |||
|
Domain Architecture |

|
|||||
| Description | General transcription factor II-I (GTFII-I) (TFII-I) (Bruton tyrosine kinase-associated protein 135) (BTK-associated protein 135) (BAP-135) (SRF-Phox1-interacting protein) (SPIN) (Williams-Beuren syndrome chromosome region 6 protein). | |||||
|
CUTL1_HUMAN
|
||||||
| NC score | 0.014736 (rank : 93) | θ value | 6.88961 (rank : 80) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1574 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P39880, Q6NYH4, Q9UEV5 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP). | |||||
|
MLH1_MOUSE
|
||||||
| NC score | 0.014681 (rank : 94) | θ value | 6.88961 (rank : 88) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9JK91, Q62454 | Gene names | Mlh1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA mismatch repair protein Mlh1 (MutL protein homolog 1). | |||||
|
CASP_HUMAN
|
||||||
| NC score | 0.014012 (rank : 95) | θ value | 6.88961 (rank : 77) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1131 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q13948, Q53GU9, Q8TBS3 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein CASP. | |||||
|
CMAH_MOUSE
|
||||||
| NC score | 0.013434 (rank : 96) | θ value | 8.99809 (rank : 96) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q61419, Q7TMR9, Q8JZM9 | Gene names | Cmah | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytidine monophosphate-N-acetylneuraminic acid hydroxylase precursor (EC 1.14.18.2) (CMP-N-acetylneuraminate monooxygenase) (CMP-N- acetylneuraminic acid hydroxylase) (CMP-NeuAc hydroxylase) (CMP-Neu5Ac hydroxylase). | |||||
|
MY18A_HUMAN
|
||||||
| NC score | 0.013272 (rank : 97) | θ value | 1.81305 (rank : 44) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1929 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q92614, Q8IXP8 | Gene names | MYO18A, KIAA0216, MYSPDZ | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
KTN1_MOUSE
|
||||||
| NC score | 0.013061 (rank : 98) | θ value | 5.27518 (rank : 71) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1324 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q61595, Q8BG49, Q8BHF4, Q8BHM8, Q8C9Y5, Q8CG51, Q8CG52, Q8CG53, Q8CG54, Q8CG55, Q8CG56, Q8CG57, Q8CG58, Q8CG59, Q8CG60, Q8CG61, Q8CG62, Q8CG63 | Gene names | Ktn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinectin. | |||||
|
PRRT3_HUMAN
|
||||||
| NC score | 0.013001 (rank : 99) | θ value | 5.27518 (rank : 74) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q5FWE3, Q49AD0, Q6UXY6, Q8NBC9 | Gene names | PRRT3, UNQ5823/PRO19642 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
IFT74_MOUSE
|
||||||
| NC score | 0.012622 (rank : 100) | θ value | 8.99809 (rank : 97) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 939 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8BKE9, Q80ZI3, Q9CSY1, Q9CUS0, Q9D9T5 | Gene names | Ift74, Ccdc2, Cmg1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intraflagellar transport 74 homolog (Coiled-coil domain-containing protein 2) (Capillary morphogenesis protein 1) (CMG-1). | |||||
|
SOX9_HUMAN
|
||||||
| NC score | 0.011351 (rank : 101) | θ value | 6.88961 (rank : 93) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P48436 | Gene names | SOX9 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-9. | |||||
|
GRAP2_MOUSE
|
||||||
| NC score | 0.011287 (rank : 102) | θ value | 2.36792 (rank : 50) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 297 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O89100 | Gene names | Grap2, Gads, Grb2l, Grid, Mona | |||
|
Domain Architecture |

|
|||||
| Description | GRB2-related adaptor protein 2 (GADS protein) (Growth factor receptor- binding protein) (GRBLG) (GRB-2-like protein) (GRB2L) (Hematopoietic cell-associated adaptor protein GrpL) (GRB-2-related monocytic adapter protein) (Monocytic adapter) (MONA) (Adapter protein GRID). | |||||
|
CTRO_HUMAN
|
||||||
| NC score | 0.009981 (rank : 103) | θ value | 6.88961 (rank : 79) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 2092 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O14578, Q6XUH8, Q86UQ9, Q9UPZ7 | Gene names | CIT, KIAA0949, STK21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Citron Rho-interacting kinase (EC 2.7.11.1) (CRIK) (Rho-interacting, serine/threonine-protein kinase 21). | |||||
|
TIF1G_HUMAN
|
||||||
| NC score | 0.009832 (rank : 104) | θ value | 6.88961 (rank : 95) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 410 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UPN9, O95855, Q9C017, Q9UJ79 | Gene names | TRIM33, KIAA1113, RFG7, TIF1G | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-gamma (TIF1-gamma) (Tripartite motif-containing protein 33) (RET-fused gene 7 protein) (Rfg7 protein). | |||||
|
CAMKV_HUMAN
|
||||||
| NC score | 0.009614 (rank : 105) | θ value | 1.06291 (rank : 32) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1084 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8NCB2, Q6FIB8, Q8NBS8, Q8NC85, Q8NDU4, Q8WTT8, Q9BQC9, Q9H0Q5 | Gene names | CAMKV | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CaM kinase-like vesicle-associated protein. | |||||
|
GRP78_HUMAN
|
||||||
| NC score | 0.008794 (rank : 106) | θ value | 6.88961 (rank : 83) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P11021, Q2EF78, Q9NPF1, Q9UK02 | Gene names | HSPA5, GRP78 | |||
|
Domain Architecture |

|
|||||
| Description | 78 kDa glucose-regulated protein precursor (GRP 78) (Heat shock 70 kDa protein 5) (Immunoglobulin heavy chain-binding protein) (BiP) (Endoplasmic reticulum lumenal Ca(2+)-binding protein grp78). | |||||
|
GRP78_MOUSE
|
||||||
| NC score | 0.008660 (rank : 107) | θ value | 6.88961 (rank : 84) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P20029, O35642, Q3UFF2, Q61630 | Gene names | Hspa5, Grp78 | |||
|
Domain Architecture |

|
|||||
| Description | 78 kDa glucose-regulated protein precursor (GRP 78) (Heat shock 70 kDa protein 5) (Immunoglobulin heavy chain-binding protein) (BiP). | |||||
|
DLX3_MOUSE
|
||||||
| NC score | 0.008630 (rank : 108) | θ value | 5.27518 (rank : 68) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 367 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q64205 | Gene names | Dlx3 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein DLX-3. | |||||
|
TITIN_HUMAN
|
||||||
| NC score | 0.008556 (rank : 109) | θ value | 8.99809 (rank : 102) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 2923 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8WZ42, Q10465, Q10466, Q15598, Q2XUS3, Q32Q60, Q4U1Z6, Q6NSG0, Q6PDB1, Q6PJP0, Q7KYM2, Q7KYN4, Q7KYN5, Q7LDM3, Q7Z2X3, Q8TCG8, Q8WZ51, Q8WZ52, Q8WZ53, Q8WZB3, Q92761, Q92762, Q9UD97, Q9UP84, Q9Y6L9 | Gene names | TTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Titin (EC 2.7.11.1) (Connectin) (Rhabdomyosarcoma antigen MU-RMS- 40.14). | |||||
|
NEK1_MOUSE
|
||||||
| NC score | 0.007673 (rank : 110) | θ value | 1.81305 (rank : 45) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P51954 | Gene names | Nek1 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Nek1 (EC 2.7.11.1) (NimA-related protein kinase 1). | |||||
|
SN1L2_HUMAN
|
||||||
| NC score | 0.007501 (rank : 111) | θ value | 1.81305 (rank : 47) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 964 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9H0K1, O94878, Q6AZE2, Q76N03, Q8NCV7, Q96CZ8 | Gene names | SNF1LK2, KIAA0781, QIK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SNF1-like kinase 2 (EC 2.7.11.1) (Qin- induced kinase). | |||||
|
K1H1_HUMAN
|
||||||
| NC score | 0.006976 (rank : 112) | θ value | 6.88961 (rank : 87) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q15323 | Gene names | KRTHA1, HHA1, HKA1 | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type I cuticular Ha1 (Hair keratin, type I Ha1). | |||||
|
ANPRA_MOUSE
|
||||||
| NC score | 0.006591 (rank : 113) | θ value | 5.27518 (rank : 67) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 600 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P18293 | Gene names | Npr1, Npra | |||
|
Domain Architecture |

|
|||||
| Description | Atrial natriuretic peptide receptor A precursor (ANP-A) (ANPRA) (GC-A) (Guanylate cyclase) (EC 4.6.1.2) (NPR-A) (Atrial natriuretic peptide A-type receptor). | |||||
|
ANPRA_HUMAN
|
||||||
| NC score | 0.006549 (rank : 114) | θ value | 5.27518 (rank : 66) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 612 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P16066, Q6P4Q3 | Gene names | NPR1, ANPRA | |||
|
Domain Architecture |

|
|||||
| Description | Atrial natriuretic peptide receptor A precursor (ANP-A) (ANPRA) (GC-A) (Guanylate cyclase) (EC 4.6.1.2) (NPR-A) (Atrial natriuretic peptide A-type receptor). | |||||
|
HIPK2_MOUSE
|
||||||
| NC score | 0.006289 (rank : 115) | θ value | 5.27518 (rank : 70) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 871 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9QZR5, O88905, Q99P45, Q99P46, Q9D2E6, Q9D474, Q9EQL2, Q9QZR4 | Gene names | Hipk2, Nbak1, Stank | |||
|
Domain Architecture |

|
|||||
| Description | Homeodomain-interacting protein kinase 2 (EC 2.7.11.1) (Nuclear body- associated kinase 1) (Sialophorin tail-associated nuclear serine/threonine-protein kinase). | |||||
|
DMPK_HUMAN
|
||||||
| NC score | 0.005861 (rank : 116) | θ value | 1.81305 (rank : 39) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 944 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q09013, Q16205 | Gene names | DMPK, MDPK | |||
|
Domain Architecture |

|
|||||
| Description | Myotonin-protein kinase (EC 2.7.11.1) (Myotonic dystrophy protein kinase) (MDPK) (DM-kinase) (DMK) (DMPK) (MT-PK). | |||||
|
M3K9_HUMAN
|
||||||
| NC score | 0.004070 (rank : 117) | θ value | 8.99809 (rank : 98) | |||
| Query Neighborhood Hits | 104 | Target Neighborhood Hits | 1123 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P80192, Q6EH31, Q9H2N5 | Gene names | MAP3K9, MLK1, PRKE1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitogen-activated protein kinase kinase kinase 9 (EC 2.7.11.25) (Mixed lineage kinase 1). | |||||