Please be patient as the page loads
|
RED_HUMAN
|
||||||
| SwissProt Accessions | Q13123 | Gene names | IK, RED, RER | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Red (Protein RER) (IK factor) (Cytokine IK). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
RED_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | Q13123 | Gene names | IK, RED, RER | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Red (Protein RER) (IK factor) (Cytokine IK). | |||||
|
RED_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.988722 (rank : 2) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9Z1M8 | Gene names | Ik, Red, Rer | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Red (Protein RER) (IK factor) (Cytokine IK). | |||||
|
TCRG1_HUMAN
|
||||||
| θ value | 6.43352e-06 (rank : 3) | NC score | 0.184942 (rank : 3) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 929 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O14776 | Gene names | TCERG1, CA150, TAF2S | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150). | |||||
|
TCRG1_MOUSE
|
||||||
| θ value | 6.43352e-06 (rank : 4) | NC score | 0.184061 (rank : 4) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8CGF7, Q61051, Q8C490, Q8CHT8, Q9R0R5 | Gene names | Tcerg1, Taf2s | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150) (p144) (Formin- binding protein 28) (FBP 28). | |||||
|
CCD55_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 5) | NC score | 0.132041 (rank : 5) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9H0G5, Q6FI71 | Gene names | CCDC55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 55. | |||||
|
HM20A_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 6) | NC score | 0.105726 (rank : 6) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9NP66, Q53G31, Q9NSF6 | Gene names | HMG20A, HMGX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | High mobility group protein 20A (HMG box-containing protein 20A) (HMG domain-containing protein HMGX1). | |||||
|
HM20A_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 7) | NC score | 0.105373 (rank : 7) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 348 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9DC33, Q3LSF9, Q6NV87, Q8BSK1, Q8C3C1, Q8CAA0, Q9CYG2 | Gene names | Hmg20a, Ibraf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | High mobility group protein 20A (HMG box-containing protein 20A) (iBRAF) (Inhibitor of BRAF35) (HMG domain protein HMGX1). | |||||
|
RAD50_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 8) | NC score | 0.073517 (rank : 26) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1317 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q92878, O43254, Q6GMT7, Q6P5X3, Q9UP86 | Gene names | RAD50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (hRAD50). | |||||
|
RCOR3_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 9) | NC score | 0.087927 (rank : 15) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9P2K3, Q5VT47, Q7L9I5, Q8N5U3, Q9NV83 | Gene names | RCOR3, KIAA1343 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 3. | |||||
|
DIDO1_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 10) | NC score | 0.088788 (rank : 14) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
RAD50_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 11) | NC score | 0.071351 (rank : 28) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1366 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P70388, Q6PEB0, Q8C2T7, Q9CU59 | Gene names | Rad50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (mRad50). | |||||
|
RPTN_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 12) | NC score | 0.068466 (rank : 30) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6XPR3 | Gene names | RPTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Repetin. | |||||
|
SNUT1_MOUSE
|
||||||
| θ value | 0.163984 (rank : 13) | NC score | 0.103994 (rank : 8) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Z315, Q8K155, Q9R1I9, Q9R270 | Gene names | Sart1, Haf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U4/U6.U5 tri-snRNP-associated protein 1 (Squamous cell carcinoma antigen recognized by T-cells 1) (SART-1) (mSART-1) (Hypoxia- associated factor). | |||||
|
RCOR3_MOUSE
|
||||||
| θ value | 0.279714 (rank : 14) | NC score | 0.079851 (rank : 22) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6PGA0 | Gene names | Rcor3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 3. | |||||
|
SPT6H_HUMAN
|
||||||
| θ value | 0.279714 (rank : 15) | NC score | 0.097642 (rank : 10) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q7KZ85, Q15737, Q6GMQ4, Q7KYW9, Q7LDK4, Q8N526, Q92775, Q96AH3, Q9BTH9, Q9BTI2 | Gene names | SUPT6H, KIAA0162, SPT6H | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor SPT6 (hSPT6) (Tat-cotransactivator 2 protein) (Tat-CT2 protein). | |||||
|
TR150_MOUSE
|
||||||
| θ value | 0.279714 (rank : 16) | NC score | 0.098797 (rank : 9) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 468 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q569Z6 | Gene names | Thrap3, Trap150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid hormone receptor-associated protein 3 (Thyroid hormone receptor-associated protein complex 150 kDa component) (Trap150). | |||||
|
HOOK3_HUMAN
|
||||||
| θ value | 0.365318 (rank : 17) | NC score | 0.049405 (rank : 65) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1070 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q86VS8, Q8NBH0, Q9BY13 | Gene names | HOOK3 | |||
|
Domain Architecture |

|
|||||
| Description | Hook homolog 3 (hHK3). | |||||
|
TR150_HUMAN
|
||||||
| θ value | 0.365318 (rank : 18) | NC score | 0.087783 (rank : 16) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Y2W1, Q5VTK6 | Gene names | THRAP3, TRAP150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid hormone receptor-associated protein 3 (Thyroid hormone receptor-associated protein complex 150 kDa component) (Trap150). | |||||
|
DNJC8_HUMAN
|
||||||
| θ value | 0.47712 (rank : 19) | NC score | 0.093207 (rank : 11) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O75937, Q6IBA4, Q8N4Z5, Q9P051, Q9P067 | Gene names | DNAJC8, SPF31 | |||
|
Domain Architecture |
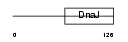
|
|||||
| Description | DnaJ homolog subfamily C member 8 (Splicing protein spf31). | |||||
|
DNJC8_MOUSE
|
||||||
| θ value | 0.47712 (rank : 20) | NC score | 0.092598 (rank : 12) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6NZB0, Q6PG72, Q8C2M6 | Gene names | Dnajc8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 8. | |||||
|
PHF20_HUMAN
|
||||||
| θ value | 0.47712 (rank : 21) | NC score | 0.076409 (rank : 24) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9BVI0, Q5JWY9, Q9BWV4, Q9BXA3, Q9BZW3, Q9H421, Q9H4J6, Q9NZ22 | Gene names | PHF20, C20orf104, GLEA2, HCA58, NZF, TZP | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58) (Glioma-expressed antigen 2) (Transcription factor TZP) (Novel zinc finger protein). | |||||
|
CLGN_MOUSE
|
||||||
| θ value | 0.62314 (rank : 22) | NC score | 0.045627 (rank : 71) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P52194, Q9D2K5 | Gene names | Clgn, Meg1 | |||
|
Domain Architecture |
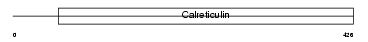
|
|||||
| Description | Calmegin precursor (MEG 1 antigen) (Calnexin-T) (A2/6). | |||||
|
CALU_HUMAN
|
||||||
| θ value | 0.813845 (rank : 23) | NC score | 0.039564 (rank : 79) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O43852, O60456, Q6FHB9, Q96RL3, Q9NR43 | Gene names | CALU | |||
|
Domain Architecture |

|
|||||
| Description | Calumenin precursor (Crocalbin) (IEF SSP 9302). | |||||
|
CALU_MOUSE
|
||||||
| θ value | 0.813845 (rank : 24) | NC score | 0.039742 (rank : 78) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O35887 | Gene names | Calu | |||
|
Domain Architecture |

|
|||||
| Description | Calumenin precursor (Crocalbin). | |||||
|
LEO1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 25) | NC score | 0.053088 (rank : 58) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
NOP14_HUMAN
|
||||||
| θ value | 1.06291 (rank : 26) | NC score | 0.055880 (rank : 47) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 573 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P78316, Q7LGI5, Q7Z6K0, Q8TBR6 | Gene names | NOP14, C4orf9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable nucleolar complex protein 14. | |||||
|
PPIL4_HUMAN
|
||||||
| θ value | 1.06291 (rank : 27) | NC score | 0.038826 (rank : 81) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8WUA2, Q7Z3Q5 | Gene names | PPIL4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peptidyl-prolyl cis-trans isomerase-like 4 (EC 5.2.1.8) (PPIase) (Rotamase) (Cyclophilin-like protein PPIL4). | |||||
|
TAF4_HUMAN
|
||||||
| θ value | 1.06291 (rank : 28) | NC score | 0.057019 (rank : 42) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 687 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O00268, Q5TBP6, Q99721, Q9BR40, Q9BX42 | Gene names | TAF4, TAF2C, TAF2C1, TAF4A, TAFII130, TAFII135 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 4 (TBP-associated factor 4) (Transcription initiation factor TFIID 135 kDa subunit) (TAF(II)135) (TAFII-135) (TAFII135) (TAFII-130) (TAFII130). | |||||
|
MRCKB_MOUSE
|
||||||
| θ value | 1.38821 (rank : 29) | NC score | 0.026347 (rank : 98) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 2180 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q7TT50, Q69ZR5, Q80W33 | Gene names | Cdc42bpb, Kiaa1124 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK beta (EC 2.7.11.1) (CDC42-binding protein kinase beta) (Myotonic dystrophy kinase-related CDC42-binding kinase beta) (Myotonic dystrophy protein kinase-like beta) (MRCK beta) (DMPK-like beta). | |||||
|
PHF3_HUMAN
|
||||||
| θ value | 1.38821 (rank : 30) | NC score | 0.080137 (rank : 21) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q92576, Q5T1T6, Q9NQ16, Q9UI45 | Gene names | PHF3, KIAA0244 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 3. | |||||
|
SPT6H_MOUSE
|
||||||
| θ value | 1.38821 (rank : 31) | NC score | 0.089227 (rank : 13) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q62383, Q5SYM0, Q6GQS3, Q6ZQI0, Q8BQY6 | Gene names | Supt6h, Kiaa0162 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor SPT6. | |||||
|
GOGA4_MOUSE
|
||||||
| θ value | 1.81305 (rank : 32) | NC score | 0.044246 (rank : 73) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1939 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q91VW5, O70365, Q8CGH6 | Gene names | Golga4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (tGolgin-1). | |||||
|
MEPE_HUMAN
|
||||||
| θ value | 1.81305 (rank : 33) | NC score | 0.071255 (rank : 29) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NQ76 | Gene names | MEPE | |||
|
Domain Architecture |
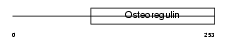
|
|||||
| Description | Matrix extracellular phosphoglycoprotein precursor (Osteoblast/osteocyte factor 45) (OF45). | |||||
|
MYO6_HUMAN
|
||||||
| θ value | 1.81305 (rank : 34) | NC score | 0.038517 (rank : 82) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 727 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9UM54, Q9BZZ7, Q9UEG2 | Gene names | MYO6, KIAA0389 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-6 (Myosin VI). | |||||
|
PRGC1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 35) | NC score | 0.048093 (rank : 66) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O70343 | Gene names | Ppargc1a, Pgc1, Pgc1a, Ppargc1 | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisome proliferator-activated receptor gamma coactivator 1-alpha (PPAR gamma coactivator 1-alpha) (PPARGC-1-alpha) (PGC-1-alpha). | |||||
|
STIM1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 36) | NC score | 0.055109 (rank : 50) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 665 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q13586 | Gene names | STIM1, GOK | |||
|
Domain Architecture |
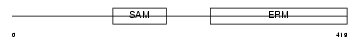
|
|||||
| Description | Stromal interaction molecule 1 precursor. | |||||
|
STIM1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 37) | NC score | 0.056172 (rank : 45) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 660 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P70302 | Gene names | Stim1, Sim | |||
|
Domain Architecture |
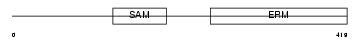
|
|||||
| Description | Stromal interaction molecule 1 precursor. | |||||
|
ZN192_HUMAN
|
||||||
| θ value | 1.81305 (rank : 38) | NC score | 0.002137 (rank : 109) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 789 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15776, Q4VAR1, Q4VAR2, Q4VAR3, Q9H4T1 | Gene names | ZNF192 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 192 (LD5-1). | |||||
|
C8AP2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 39) | NC score | 0.052554 (rank : 60) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 548 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UKL3, Q6PH76, Q7LCQ7, Q86YD9, Q9NUQ4, Q9NZV9, Q9P2N1, Q9Y563 | Gene names | CASP8AP2, FLASH, KIAA1315, RIP25 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
DESP_HUMAN
|
||||||
| θ value | 2.36792 (rank : 40) | NC score | 0.043814 (rank : 74) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P15924, O75993, Q14189, Q9UHN4 | Gene names | DSP | |||
|
Domain Architecture |

|
|||||
| Description | Desmoplakin (DP) (250/210 kDa paraneoplastic pemphigus antigen). | |||||
|
CTRO_MOUSE
|
||||||
| θ value | 3.0926 (rank : 41) | NC score | 0.025104 (rank : 99) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 2344 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P49025, O88528, O88937, O88938, Q3UM99, Q8CIJ1 | Gene names | Cit | |||
|
Domain Architecture |

|
|||||
| Description | Citron Rho-interacting kinase (EC 2.7.11.1) (CRIK) (Rho-interacting, serine/threonine-protein kinase 21). | |||||
|
MYO5B_HUMAN
|
||||||
| θ value | 3.0926 (rank : 42) | NC score | 0.030298 (rank : 94) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 911 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9ULV0 | Gene names | MYO5B, KIAA1119 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5B (Myosin Vb). | |||||
|
NEST_HUMAN
|
||||||
| θ value | 3.0926 (rank : 43) | NC score | 0.044698 (rank : 72) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1305 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P48681, O00552 | Gene names | NES | |||
|
Domain Architecture |

|
|||||
| Description | Nestin. | |||||
|
NUCB1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 44) | NC score | 0.047417 (rank : 69) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q02818, Q15838, Q7Z4J7, Q9BUR1 | Gene names | NUCB1, NUC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleobindin-1 precursor (CALNUC). | |||||
|
RAI14_MOUSE
|
||||||
| θ value | 3.0926 (rank : 45) | NC score | 0.033553 (rank : 88) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1505 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9EP71, Q3URT3, Q6ZPT6 | Gene names | Rai14, Kiaa1334, Norpeg | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankycorbin (Ankyrin repeat and coiled-coil structure-containing protein) (Retinoic acid-induced protein 14) (Novel retinal pigment epithelial cell protein) (p125). | |||||
|
SEC63_HUMAN
|
||||||
| θ value | 3.0926 (rank : 46) | NC score | 0.056720 (rank : 43) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UGP8, O95380, Q86VS9, Q8IWL0, Q9NTE0 | Gene names | SEC63, SEC63L | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
SNUT1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 47) | NC score | 0.077519 (rank : 23) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O43290, Q53GB5 | Gene names | SART1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U4/U6.U5 tri-snRNP-associated protein 1 (U4/U6.U5 tri-snRNP-associated 110 kDa protein) (Squamous cell carcinoma antigen recognized by T cells 1) (SART-1) (hSART-1) (hSnu66). | |||||
|
TAOK1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 48) | NC score | 0.018631 (rank : 104) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1734 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q7L7X3, Q96L75, Q9H2K7, Q9H7S5, Q9P2I6 | Gene names | TAOK1, KIAA1361, MAP3K16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase TAO1 (EC 2.7.11.1) (Thousand and one amino acid protein 1) (STE20-like kinase PSK2) (Kinase from chicken homolog B) (hKFC-B). | |||||
|
TAOK1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 49) | NC score | 0.019007 (rank : 103) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1720 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q5F2E8, Q69ZL2, Q8JZX2, Q91VG7 | Gene names | Taok1, Kiaa1361 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase TAO1 (EC 2.7.11.1) (Thousand and one amino acid protein 1). | |||||
|
ANR12_HUMAN
|
||||||
| θ value | 4.03905 (rank : 50) | NC score | 0.052612 (rank : 59) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1081 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q6UB98, O94951, Q658K1, Q6QMF7, Q9H231, Q9H784 | Gene names | ANKRD12, ANCO2, KIAA0874 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 12 (Ankyrin repeat-containing cofactor 2) (GAC-1 protein). | |||||
|
CCD21_HUMAN
|
||||||
| θ value | 4.03905 (rank : 51) | NC score | 0.046955 (rank : 70) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1228 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q6P2H3, Q5VY68, Q5VY70, Q9H6Q1, Q9H828, Q9UF52 | Gene names | CCDC21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 21. | |||||
|
CROCC_MOUSE
|
||||||
| θ value | 4.03905 (rank : 52) | NC score | 0.042771 (rank : 75) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1460 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8CJ40, Q7TQL2, Q80U01, Q8R0B9 | Gene names | Crocc, Kiaa0445 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rootletin (Ciliary rootlet coiled-coil protein). | |||||
|
HOOK3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 53) | NC score | 0.039510 (rank : 80) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1007 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8BUK6, Q8BV48, Q8BW57, Q8BY41 | Gene names | Hook3 | |||
|
Domain Architecture |
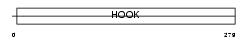
|
|||||
| Description | Hook homolog 3. | |||||
|
ITSN1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 54) | NC score | 0.042417 (rank : 76) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1757 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q15811, O95216, Q9UET5, Q9UK60, Q9UNK1, Q9UNK2, Q9UQ92 | Gene names | ITSN1, ITSN, SH3D1A | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-1 (SH3 domain-containing protein 1A) (SH3P17). | |||||
|
MY18A_MOUSE
|
||||||
| θ value | 4.03905 (rank : 55) | NC score | 0.038472 (rank : 83) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1989 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9JMH9, Q811D7 | Gene names | Myo18a, Myspdz | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
PHF20_MOUSE
|
||||||
| θ value | 4.03905 (rank : 56) | NC score | 0.056093 (rank : 46) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8BLG0, Q8BMA2, Q8BYR4, Q921N1 | Gene names | Phf20, Hca58 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58 homolog). | |||||
|
TPR_HUMAN
|
||||||
| θ value | 4.03905 (rank : 57) | NC score | 0.040706 (rank : 77) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
ZHX2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 58) | NC score | 0.022578 (rank : 100) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8C0C0 | Gene names | Zhx2, Afr1, Raf | |||
|
Domain Architecture |

|
|||||
| Description | Zinc fingers and homeoboxes protein 2 (Zinc finger and homeodomain protein 2) (Alpha-fetoprotein regulator 1) (AFP regulator 1) (Regulator of AFP). | |||||
|
BPA1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 59) | NC score | 0.031947 (rank : 90) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q91ZU6, Q91ZU7 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 1/2/3/4 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
CEP55_HUMAN
|
||||||
| θ value | 5.27518 (rank : 60) | NC score | 0.031303 (rank : 92) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q53EZ4, Q32WF5, Q3MV20, Q5VY28, Q6N034, Q96H32, Q9NVS7 | Gene names | CEP55, C10orf3, URCC6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome protein 55 (Up-regulated in colon cancer 6). | |||||
|
CJ006_MOUSE
|
||||||
| θ value | 5.27518 (rank : 61) | NC score | 0.030793 (rank : 93) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6P9P0, Q8C041, Q8C0J5 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf6 homolog. | |||||
|
DSG3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 62) | NC score | 0.004714 (rank : 108) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P32926 | Gene names | DSG3 | |||
|
Domain Architecture |
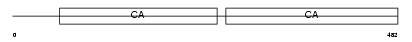
|
|||||
| Description | Desmoglein-3 precursor (130 kDa pemphigus vulgaris antigen) (PVA). | |||||
|
ELOA1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 63) | NC score | 0.038295 (rank : 84) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q14241, Q8IXH1 | Gene names | TCEB3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription elongation factor B polypeptide 3 (RNA polymerase II transcription factor SIII subunit A1) (SIII p110) (Elongin-A) (EloA) (Elongin 110 kDa subunit). | |||||
|
KIF4A_HUMAN
|
||||||
| θ value | 5.27518 (rank : 64) | NC score | 0.027986 (rank : 96) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1159 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O95239, Q86TN3, Q86XX7, Q9NNY6, Q9NY24, Q9UMW3 | Gene names | KIF4A, KIF4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromosome-associated kinesin KIF4A (Chromokinesin). | |||||
|
STAM2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 65) | NC score | 0.012331 (rank : 106) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O88811, Q8C8Y4 | Gene names | Stam2, Hbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal transducing adapter molecule 2 (STAM-2) (Hrs-binding protein). | |||||
|
CCD21_MOUSE
|
||||||
| θ value | 6.88961 (rank : 66) | NC score | 0.047462 (rank : 68) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8BMK0, Q8BUF1, Q8K0E6 | Gene names | Ccdc21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 21. | |||||
|
GEM_HUMAN
|
||||||
| θ value | 6.88961 (rank : 67) | NC score | 0.008358 (rank : 107) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P55040 | Gene names | GEM, KIR | |||
|
Domain Architecture |
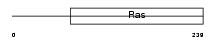
|
|||||
| Description | GTP-binding protein GEM (GTP-binding mitogen-induced T-cell protein) (RAS-like protein KIR). | |||||
|
MDC1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 68) | NC score | 0.019937 (rank : 102) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
MPP8_HUMAN
|
||||||
| θ value | 6.88961 (rank : 69) | NC score | 0.032237 (rank : 89) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 948 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q99549, Q86TK3, Q9BTP1 | Gene names | MPHOSPH8, MPP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | M-phase phosphoprotein 8. | |||||
|
MY18A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 70) | NC score | 0.035596 (rank : 85) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1929 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q92614, Q8IXP8 | Gene names | MYO18A, KIAA0216, MYSPDZ | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
MYH9_MOUSE
|
||||||
| θ value | 6.88961 (rank : 71) | NC score | 0.035485 (rank : 86) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1829 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q8VDD5 | Gene names | Myh9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
SEC63_MOUSE
|
||||||
| θ value | 6.88961 (rank : 72) | NC score | 0.055262 (rank : 49) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 601 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8VHE0, Q8VEB9 | Gene names | Sec63, Sec63l | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
CN120_MOUSE
|
||||||
| θ value | 8.99809 (rank : 73) | NC score | 0.027865 (rank : 97) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9DB96, Q8CIJ2 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | Protein C14orf120 homolog. | |||||
|
LEO1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 74) | NC score | 0.048062 (rank : 67) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
MYH9_HUMAN
|
||||||
| θ value | 8.99809 (rank : 75) | NC score | 0.035218 (rank : 87) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P35579, O60805 | Gene names | MYH9 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
RILP_MOUSE
|
||||||
| θ value | 8.99809 (rank : 76) | NC score | 0.028109 (rank : 95) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q5ND29, Q80WB8, Q8CHY9 | Gene names | Rilp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab-interacting lysosomal protein. | |||||
|
SBP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 77) | NC score | 0.020580 (rank : 101) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13228 | Gene names | SELENBP1, SBP | |||
|
Domain Architecture |
|
|||||
| Description | Selenium-binding protein 1. | |||||
|
SDCG8_HUMAN
|
||||||
| θ value | 8.99809 (rank : 78) | NC score | 0.031329 (rank : 91) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1002 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q86SQ7, O60527, Q3ZCR6, Q8N5F2, Q9P0F1 | Gene names | SDCCAG8, CCCAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serologically defined colon cancer antigen 8 (Centrosomal colon cancer autoantigen protein) (hCCCAP) (Antigen NY-CO-8). | |||||
|
SRBS1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 79) | NC score | 0.013073 (rank : 105) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 396 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9BX66, Q7LBE5, Q8IVK0, Q8IVQ4, Q96KF3, Q96KF4, Q9BX64, Q9BX65, Q9P2Q0, Q9UFT2, Q9UHN7, Q9Y338 | Gene names | SORBS1, KIAA1296, SH3D5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sorbin and SH3 domain-containing protein 1 (Ponsin) (c-Cbl-associated protein) (CAP) (SH3 domain protein 5) (SH3P12). | |||||
|
ACINU_HUMAN
|
||||||
| θ value | θ > 10 (rank : 80) | NC score | 0.056334 (rank : 44) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9UKV3, O75158, Q9UG91, Q9UKV1, Q9UKV2 | Gene names | ACIN1, ACINUS, KIAA0670 | |||
|
Domain Architecture |

|
|||||
| Description | Apoptotic chromatin condensation inducer in the nucleus (Acinus). | |||||
|
ACINU_MOUSE
|
||||||
| θ value | θ > 10 (rank : 81) | NC score | 0.062419 (rank : 35) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 712 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9JIX8, Q9CSN7, Q9CSR9, Q9CSX7, Q9R046, Q9R047 | Gene names | Acin1, Acinus | |||
|
Domain Architecture |

|
|||||
| Description | Apoptotic chromatin condensation inducer in the nucleus (Acinus). | |||||
|
CAF1A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 82) | NC score | 0.050866 (rank : 62) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 736 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9QWF0 | Gene names | Chaf1a, Caip150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromatin assembly factor 1 subunit A (CAF-1 subunit A) (Chromatin assembly factor I p150 subunit) (CAF-I 150 kDa subunit) (CAF-Ip150). | |||||
|
CALD1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 83) | NC score | 0.062091 (rank : 37) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 959 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q05682, Q13978, Q13979, Q14741, Q14742 | Gene names | CALD1, CAD, CDM | |||
|
Domain Architecture |

|
|||||
| Description | Caldesmon (CDM). | |||||
|
CCD11_MOUSE
|
||||||
| θ value | θ > 10 (rank : 84) | NC score | 0.055517 (rank : 48) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9D439 | Gene names | Ccdc11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 11. | |||||
|
CCD55_MOUSE
|
||||||
| θ value | θ > 10 (rank : 85) | NC score | 0.081457 (rank : 20) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q5NCR9 | Gene names | Ccdc55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 55. | |||||
|
CROP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 86) | NC score | 0.061671 (rank : 39) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O95232, Q6PHR9, Q9NUY0, Q9P2S7 | Gene names | CROP, CREAP1, O48 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cisplatin resistance-associated overexpressed protein (cAMP regulatory element-associated protein 1) (CRE-associated protein 1) (CREAP-1) (Luc7A) (Okadaic acid-inducible phosphoprotein OA48-18). | |||||
|
CROP_MOUSE
|
||||||
| θ value | θ > 10 (rank : 87) | NC score | 0.061382 (rank : 41) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q5SUF2, Q3U9D5, Q8BUJ5, Q921Z3, Q9CRS7, Q9CTY4 | Gene names | Crop | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cisplatin resistance-associated overexpressed protein. | |||||
|
IF2P_HUMAN
|
||||||
| θ value | θ > 10 (rank : 88) | NC score | 0.054061 (rank : 55) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 889 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | O60841, O95805, Q9UF81, Q9UMN7 | Gene names | EIF5B, IF2, KIAA0741 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 5B (eIF-5B) (Translation initiation factor IF-2). | |||||
|
IF3A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 89) | NC score | 0.054339 (rank : 53) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 875 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q14152, O00653 | Gene names | EIF3S10, KIAA0139 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 10 (eIF-3 theta) (eIF3 p167) (eIF3 p180) (eIF3 p185) (eIF3a). | |||||
|
IF3A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 90) | NC score | 0.054307 (rank : 54) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 871 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P23116, Q60697, Q62162 | Gene names | Eif3s10, Csma, Eif3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 10 (eIF-3 theta) (eIF3 p167) (eIF3 p180) (eIF3 p185) (eIF3a) (p162 protein) (Centrosomin). | |||||
|
INCE_HUMAN
|
||||||
| θ value | θ > 10 (rank : 91) | NC score | 0.061977 (rank : 38) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 978 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9NQS7, Q5Y192 | Gene names | INCENP | |||
|
Domain Architecture |

|
|||||
| Description | Inner centromere protein. | |||||
|
INCE_MOUSE
|
||||||
| θ value | θ > 10 (rank : 92) | NC score | 0.061436 (rank : 40) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9WU62, Q7TN28, Q8BGN4, Q8CGI4 | Gene names | Incenp | |||
|
Domain Architecture |

|
|||||
| Description | Inner centromere protein. | |||||
|
NUCB1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 93) | NC score | 0.050079 (rank : 64) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q02819, Q3TU42, Q9CWA1, Q9D8J4, Q9DBL3 | Gene names | Nucb1, Nuc, Nucb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleobindin-1 precursor (CALNUC). | |||||
|
PRP40_HUMAN
|
||||||
| θ value | θ > 10 (rank : 94) | NC score | 0.054838 (rank : 51) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 442 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O75400, O43856, O75404, Q8TBQ1, Q9H782, Q9NWU9, Q9P0Q2, Q9Y5A8 | Gene names | PRPF40A, FLAF1, FNBP3, HYPA | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Huntingtin yeast partner A) (Huntingtin-interacting protein HYPA/FBP11) (Fas ligand-associated factor 1) (NY-REN-6 antigen). | |||||
|
PRP40_MOUSE
|
||||||
| θ value | θ > 10 (rank : 95) | NC score | 0.064026 (rank : 32) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9R1C7, Q61049, Q8BQ76, Q8BRW4, Q8C2U1 | Gene names | Prpf40a, Fbp11, Fnbp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Formin-binding protein 11) (FBP-11). | |||||
|
RBM25_HUMAN
|
||||||
| θ value | θ > 10 (rank : 96) | NC score | 0.084423 (rank : 17) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P49756, Q9UEQ5, Q9UIE9 | Gene names | RBM25, RNPC7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable RNA-binding protein 25 (RNA-binding motif protein 25) (RNA- binding region-containing protein 7) (Protein S164). | |||||
|
REN3B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 97) | NC score | 0.063008 (rank : 34) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9BZI7, Q9H1J0 | Gene names | UPF3B, RENT3B, UPF3X | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of nonsense transcripts 3B (Nonsense mRNA reducing factor 3B) (Up-frameshift suppressor 3 homolog B) (hUpf3B) (hUpf3p-X). | |||||
|
RENT2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 98) | NC score | 0.062272 (rank : 36) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 618 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9HAU5, Q8N8U1, Q9H1J2, Q9NWL1, Q9P2D9, Q9Y4M9 | Gene names | UPF2, KIAA1408, RENT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of nonsense transcripts 2 (Nonsense mRNA reducing factor 2) (Up-frameshift suppressor 2 homolog) (hUpf2). | |||||
|
SFR11_HUMAN
|
||||||
| θ value | θ > 10 (rank : 99) | NC score | 0.064672 (rank : 31) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q05519 | Gene names | SFRS11 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor arginine/serine-rich 11 (Arginine-rich 54 kDa nuclear protein) (p54). | |||||
|
SFR12_HUMAN
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.083596 (rank : 18) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 553 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q8WXA9, Q86X37 | Gene names | SFRS12, SRRP86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86) (Splicing regulatory protein 508) (SRrp508). | |||||
|
SFR12_MOUSE
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.075064 (rank : 25) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q8BZX4, Q8BLM8, Q8BWK7, Q8BYK0, Q8BYZ4 | Gene names | Sfrs12, Srrp86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86). | |||||
|
SFR17_HUMAN
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.053971 (rank : 56) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8TF01, Q5T064, Q6P2B4, Q6P2N4, Q6PJ93, Q6PK36, Q7Z640, Q8TF00, Q96K10, Q96SM5, Q9P076, Q9P0C0, Q9Y4N3 | Gene names | SRRP130, C6orf111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 130 (Serine-arginine-rich- splicing regulatory protein 130) (SRrp130) (SR-rich protein) (SR- related protein). | |||||
|
T2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.072706 (rank : 27) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q06666 | Gene names | Srst, T2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Octapeptide-repeat protein T2. | |||||
|
TAF3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.052378 (rank : 61) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 568 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q5VWG9, Q6GMS5, Q6P6B5, Q86VY6, Q9BQS9, Q9UFI8 | Gene names | TAF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
TAF3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.050773 (rank : 63) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 463 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q5HZG4, Q3U490, Q3UWX2, Q8BIU8, Q99JH4 | Gene names | Taf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
TRHY_HUMAN
|
||||||
| θ value | θ > 10 (rank : 106) | NC score | 0.063022 (rank : 33) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1385 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q07283 | Gene names | THH, THL, TRHY | |||
|
Domain Architecture |

|
|||||
| Description | Trichohyalin. | |||||
|
WDR60_MOUSE
|
||||||
| θ value | θ > 10 (rank : 107) | NC score | 0.054430 (rank : 52) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8C761, Q3UNM4 | Gene names | Wdr60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 60. | |||||
|
XE7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 108) | NC score | 0.053970 (rank : 57) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 738 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q02040, Q02832 | Gene names | CXYorf3, DXYS155E, XE7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-lymphocyte antigen precursor (B-lymphocyte surface antigen) (721P) (Protein XE7). | |||||
|
ZC313_HUMAN
|
||||||
| θ value | θ > 10 (rank : 109) | NC score | 0.082374 (rank : 19) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q5T200, O94936, Q5T1Z9, Q7Z7J3, Q8NDT6, Q9H0L6 | Gene names | ZC3H13, KIAA0853 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 13. | |||||
|
RED_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | Q13123 | Gene names | IK, RED, RER | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Red (Protein RER) (IK factor) (Cytokine IK). | |||||
|
RED_MOUSE
|
||||||
| NC score | 0.988722 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q9Z1M8 | Gene names | Ik, Red, Rer | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Red (Protein RER) (IK factor) (Cytokine IK). | |||||
|
TCRG1_HUMAN
|
||||||
| NC score | 0.184942 (rank : 3) | θ value | 6.43352e-06 (rank : 3) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 929 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O14776 | Gene names | TCERG1, CA150, TAF2S | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150). | |||||
|
TCRG1_MOUSE
|
||||||
| NC score | 0.184061 (rank : 4) | θ value | 6.43352e-06 (rank : 4) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8CGF7, Q61051, Q8C490, Q8CHT8, Q9R0R5 | Gene names | Tcerg1, Taf2s | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation regulator 1 (TATA box-binding protein- associated factor 2S) (Transcription factor CA150) (p144) (Formin- binding protein 28) (FBP 28). | |||||
|
CCD55_HUMAN
|
||||||
| NC score | 0.132041 (rank : 5) | θ value | 0.00869519 (rank : 5) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9H0G5, Q6FI71 | Gene names | CCDC55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 55. | |||||
|
HM20A_HUMAN
|
||||||
| NC score | 0.105726 (rank : 6) | θ value | 0.0193708 (rank : 6) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9NP66, Q53G31, Q9NSF6 | Gene names | HMG20A, HMGX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | High mobility group protein 20A (HMG box-containing protein 20A) (HMG domain-containing protein HMGX1). | |||||
|
HM20A_MOUSE
|
||||||
| NC score | 0.105373 (rank : 7) | θ value | 0.0193708 (rank : 7) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 348 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9DC33, Q3LSF9, Q6NV87, Q8BSK1, Q8C3C1, Q8CAA0, Q9CYG2 | Gene names | Hmg20a, Ibraf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | High mobility group protein 20A (HMG box-containing protein 20A) (iBRAF) (Inhibitor of BRAF35) (HMG domain protein HMGX1). | |||||
|
SNUT1_MOUSE
|
||||||
| NC score | 0.103994 (rank : 8) | θ value | 0.163984 (rank : 13) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Z315, Q8K155, Q9R1I9, Q9R270 | Gene names | Sart1, Haf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U4/U6.U5 tri-snRNP-associated protein 1 (Squamous cell carcinoma antigen recognized by T-cells 1) (SART-1) (mSART-1) (Hypoxia- associated factor). | |||||
|
TR150_MOUSE
|
||||||
| NC score | 0.098797 (rank : 9) | θ value | 0.279714 (rank : 16) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 468 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q569Z6 | Gene names | Thrap3, Trap150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid hormone receptor-associated protein 3 (Thyroid hormone receptor-associated protein complex 150 kDa component) (Trap150). | |||||
|
SPT6H_HUMAN
|
||||||
| NC score | 0.097642 (rank : 10) | θ value | 0.279714 (rank : 15) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q7KZ85, Q15737, Q6GMQ4, Q7KYW9, Q7LDK4, Q8N526, Q92775, Q96AH3, Q9BTH9, Q9BTI2 | Gene names | SUPT6H, KIAA0162, SPT6H | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor SPT6 (hSPT6) (Tat-cotransactivator 2 protein) (Tat-CT2 protein). | |||||
|
DNJC8_HUMAN
|
||||||
| NC score | 0.093207 (rank : 11) | θ value | 0.47712 (rank : 19) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O75937, Q6IBA4, Q8N4Z5, Q9P051, Q9P067 | Gene names | DNAJC8, SPF31 | |||
|
Domain Architecture |

|
|||||
| Description | DnaJ homolog subfamily C member 8 (Splicing protein spf31). | |||||
|
DNJC8_MOUSE
|
||||||
| NC score | 0.092598 (rank : 12) | θ value | 0.47712 (rank : 20) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q6NZB0, Q6PG72, Q8C2M6 | Gene names | Dnajc8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DnaJ homolog subfamily C member 8. | |||||
|
SPT6H_MOUSE
|
||||||
| NC score | 0.089227 (rank : 13) | θ value | 1.38821 (rank : 31) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q62383, Q5SYM0, Q6GQS3, Q6ZQI0, Q8BQY6 | Gene names | Supt6h, Kiaa0162 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor SPT6. | |||||
|
DIDO1_HUMAN
|
||||||
| NC score | 0.088788 (rank : 14) | θ value | 0.0563607 (rank : 10) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
RCOR3_HUMAN
|
||||||
| NC score | 0.087927 (rank : 15) | θ value | 0.0330416 (rank : 9) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9P2K3, Q5VT47, Q7L9I5, Q8N5U3, Q9NV83 | Gene names | RCOR3, KIAA1343 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 3. | |||||
|
TR150_HUMAN
|
||||||
| NC score | 0.087783 (rank : 16) | θ value | 0.365318 (rank : 18) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Y2W1, Q5VTK6 | Gene names | THRAP3, TRAP150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid hormone receptor-associated protein 3 (Thyroid hormone receptor-associated protein complex 150 kDa component) (Trap150). | |||||
|
RBM25_HUMAN
|
||||||
| NC score | 0.084423 (rank : 17) | θ value | θ > 10 (rank : 96) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P49756, Q9UEQ5, Q9UIE9 | Gene names | RBM25, RNPC7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable RNA-binding protein 25 (RNA-binding motif protein 25) (RNA- binding region-containing protein 7) (Protein S164). | |||||
|
SFR12_HUMAN
|
||||||
| NC score | 0.083596 (rank : 18) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 553 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q8WXA9, Q86X37 | Gene names | SFRS12, SRRP86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86) (Splicing regulatory protein 508) (SRrp508). | |||||
|
ZC313_HUMAN
|
||||||
| NC score | 0.082374 (rank : 19) | θ value | θ > 10 (rank : 109) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q5T200, O94936, Q5T1Z9, Q7Z7J3, Q8NDT6, Q9H0L6 | Gene names | ZC3H13, KIAA0853 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 13. | |||||
|
CCD55_MOUSE
|
||||||
| NC score | 0.081457 (rank : 20) | θ value | θ > 10 (rank : 85) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q5NCR9 | Gene names | Ccdc55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 55. | |||||
|
PHF3_HUMAN
|
||||||
| NC score | 0.080137 (rank : 21) | θ value | 1.38821 (rank : 30) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q92576, Q5T1T6, Q9NQ16, Q9UI45 | Gene names | PHF3, KIAA0244 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 3. | |||||
|
RCOR3_MOUSE
|
||||||
| NC score | 0.079851 (rank : 22) | θ value | 0.279714 (rank : 14) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6PGA0 | Gene names | Rcor3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 3. | |||||
|
SNUT1_HUMAN
|
||||||
| NC score | 0.077519 (rank : 23) | θ value | 3.0926 (rank : 47) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O43290, Q53GB5 | Gene names | SART1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U4/U6.U5 tri-snRNP-associated protein 1 (U4/U6.U5 tri-snRNP-associated 110 kDa protein) (Squamous cell carcinoma antigen recognized by T cells 1) (SART-1) (hSART-1) (hSnu66). | |||||
|
PHF20_HUMAN
|
||||||
| NC score | 0.076409 (rank : 24) | θ value | 0.47712 (rank : 21) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9BVI0, Q5JWY9, Q9BWV4, Q9BXA3, Q9BZW3, Q9H421, Q9H4J6, Q9NZ22 | Gene names | PHF20, C20orf104, GLEA2, HCA58, NZF, TZP | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58) (Glioma-expressed antigen 2) (Transcription factor TZP) (Novel zinc finger protein). | |||||
|
SFR12_MOUSE
|
||||||
| NC score | 0.075064 (rank : 25) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q8BZX4, Q8BLM8, Q8BWK7, Q8BYK0, Q8BYZ4 | Gene names | Sfrs12, Srrp86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86). | |||||
|
RAD50_HUMAN
|
||||||
| NC score | 0.073517 (rank : 26) | θ value | 0.0193708 (rank : 8) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1317 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q92878, O43254, Q6GMT7, Q6P5X3, Q9UP86 | Gene names | RAD50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (hRAD50). | |||||
|
T2_MOUSE
|
||||||
| NC score | 0.072706 (rank : 27) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q06666 | Gene names | Srst, T2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Octapeptide-repeat protein T2. | |||||
|
RAD50_MOUSE
|
||||||
| NC score | 0.071351 (rank : 28) | θ value | 0.0563607 (rank : 11) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1366 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P70388, Q6PEB0, Q8C2T7, Q9CU59 | Gene names | Rad50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (mRad50). | |||||
|
MEPE_HUMAN
|
||||||
| NC score | 0.071255 (rank : 29) | θ value | 1.81305 (rank : 33) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NQ76 | Gene names | MEPE | |||
|
Domain Architecture |

|
|||||
| Description | Matrix extracellular phosphoglycoprotein precursor (Osteoblast/osteocyte factor 45) (OF45). | |||||
|
RPTN_HUMAN
|
||||||
| NC score | 0.068466 (rank : 30) | θ value | 0.0961366 (rank : 12) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6XPR3 | Gene names | RPTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Repetin. | |||||
|
SFR11_HUMAN
|
||||||
| NC score | 0.064672 (rank : 31) | θ value | θ > 10 (rank : 99) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q05519 | Gene names | SFRS11 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor arginine/serine-rich 11 (Arginine-rich 54 kDa nuclear protein) (p54). | |||||
|
PRP40_MOUSE
|
||||||
| NC score | 0.064026 (rank : 32) | θ value | θ > 10 (rank : 95) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9R1C7, Q61049, Q8BQ76, Q8BRW4, Q8C2U1 | Gene names | Prpf40a, Fbp11, Fnbp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Formin-binding protein 11) (FBP-11). | |||||
|
TRHY_HUMAN
|
||||||
| NC score | 0.063022 (rank : 33) | θ value | θ > 10 (rank : 106) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1385 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q07283 | Gene names | THH, THL, TRHY | |||
|
Domain Architecture |

|
|||||
| Description | Trichohyalin. | |||||
|
REN3B_HUMAN
|
||||||
| NC score | 0.063008 (rank : 34) | θ value | θ > 10 (rank : 97) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9BZI7, Q9H1J0 | Gene names | UPF3B, RENT3B, UPF3X | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of nonsense transcripts 3B (Nonsense mRNA reducing factor 3B) (Up-frameshift suppressor 3 homolog B) (hUpf3B) (hUpf3p-X). | |||||
|
ACINU_MOUSE
|
||||||
| NC score | 0.062419 (rank : 35) | θ value | θ > 10 (rank : 81) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 712 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9JIX8, Q9CSN7, Q9CSR9, Q9CSX7, Q9R046, Q9R047 | Gene names | Acin1, Acinus | |||
|
Domain Architecture |

|
|||||
| Description | Apoptotic chromatin condensation inducer in the nucleus (Acinus). | |||||
|
RENT2_HUMAN
|
||||||
| NC score | 0.062272 (rank : 36) | θ value | θ > 10 (rank : 98) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 618 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9HAU5, Q8N8U1, Q9H1J2, Q9NWL1, Q9P2D9, Q9Y4M9 | Gene names | UPF2, KIAA1408, RENT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Regulator of nonsense transcripts 2 (Nonsense mRNA reducing factor 2) (Up-frameshift suppressor 2 homolog) (hUpf2). | |||||
|
CALD1_HUMAN
|
||||||
| NC score | 0.062091 (rank : 37) | θ value | θ > 10 (rank : 83) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 959 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q05682, Q13978, Q13979, Q14741, Q14742 | Gene names | CALD1, CAD, CDM | |||
|
Domain Architecture |

|
|||||
| Description | Caldesmon (CDM). | |||||
|
INCE_HUMAN
|
||||||
| NC score | 0.061977 (rank : 38) | θ value | θ > 10 (rank : 91) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 978 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9NQS7, Q5Y192 | Gene names | INCENP | |||
|
Domain Architecture |

|
|||||
| Description | Inner centromere protein. | |||||
|
CROP_HUMAN
|
||||||
| NC score | 0.061671 (rank : 39) | θ value | θ > 10 (rank : 86) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | O95232, Q6PHR9, Q9NUY0, Q9P2S7 | Gene names | CROP, CREAP1, O48 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cisplatin resistance-associated overexpressed protein (cAMP regulatory element-associated protein 1) (CRE-associated protein 1) (CREAP-1) (Luc7A) (Okadaic acid-inducible phosphoprotein OA48-18). | |||||
|
INCE_MOUSE
|
||||||
| NC score | 0.061436 (rank : 40) | θ value | θ > 10 (rank : 92) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q9WU62, Q7TN28, Q8BGN4, Q8CGI4 | Gene names | Incenp | |||
|
Domain Architecture |

|
|||||
| Description | Inner centromere protein. | |||||
|
CROP_MOUSE
|
||||||
| NC score | 0.061382 (rank : 41) | θ value | θ > 10 (rank : 87) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q5SUF2, Q3U9D5, Q8BUJ5, Q921Z3, Q9CRS7, Q9CTY4 | Gene names | Crop | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cisplatin resistance-associated overexpressed protein. | |||||
|
TAF4_HUMAN
|
||||||
| NC score | 0.057019 (rank : 42) | θ value | 1.06291 (rank : 28) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 687 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O00268, Q5TBP6, Q99721, Q9BR40, Q9BX42 | Gene names | TAF4, TAF2C, TAF2C1, TAF4A, TAFII130, TAFII135 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 4 (TBP-associated factor 4) (Transcription initiation factor TFIID 135 kDa subunit) (TAF(II)135) (TAFII-135) (TAFII135) (TAFII-130) (TAFII130). | |||||
|
SEC63_HUMAN
|
||||||
| NC score | 0.056720 (rank : 43) | θ value | 3.0926 (rank : 46) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UGP8, O95380, Q86VS9, Q8IWL0, Q9NTE0 | Gene names | SEC63, SEC63L | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
ACINU_HUMAN
|
||||||
| NC score | 0.056334 (rank : 44) | θ value | θ > 10 (rank : 80) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9UKV3, O75158, Q9UG91, Q9UKV1, Q9UKV2 | Gene names | ACIN1, ACINUS, KIAA0670 | |||
|
Domain Architecture |

|
|||||
| Description | Apoptotic chromatin condensation inducer in the nucleus (Acinus). | |||||
|
STIM1_MOUSE
|
||||||
| NC score | 0.056172 (rank : 45) | θ value | 1.81305 (rank : 37) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 660 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P70302 | Gene names | Stim1, Sim | |||
|
Domain Architecture |

|
|||||
| Description | Stromal interaction molecule 1 precursor. | |||||
|
PHF20_MOUSE
|
||||||
| NC score | 0.056093 (rank : 46) | θ value | 4.03905 (rank : 56) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8BLG0, Q8BMA2, Q8BYR4, Q921N1 | Gene names | Phf20, Hca58 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 20 (Hepatocellular carcinoma-associated antigen 58 homolog). | |||||
|
NOP14_HUMAN
|
||||||
| NC score | 0.055880 (rank : 47) | θ value | 1.06291 (rank : 26) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 573 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P78316, Q7LGI5, Q7Z6K0, Q8TBR6 | Gene names | NOP14, C4orf9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable nucleolar complex protein 14. | |||||
|
CCD11_MOUSE
|
||||||
| NC score | 0.055517 (rank : 48) | θ value | θ > 10 (rank : 84) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9D439 | Gene names | Ccdc11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 11. | |||||
|
SEC63_MOUSE
|
||||||
| NC score | 0.055262 (rank : 49) | θ value | 6.88961 (rank : 72) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 601 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8VHE0, Q8VEB9 | Gene names | Sec63, Sec63l | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
STIM1_HUMAN
|
||||||
| NC score | 0.055109 (rank : 50) | θ value | 1.81305 (rank : 36) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 665 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q13586 | Gene names | STIM1, GOK | |||
|
Domain Architecture |

|
|||||
| Description | Stromal interaction molecule 1 precursor. | |||||
|
PRP40_HUMAN
|
||||||
| NC score | 0.054838 (rank : 51) | θ value | θ > 10 (rank : 94) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 442 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O75400, O43856, O75404, Q8TBQ1, Q9H782, Q9NWU9, Q9P0Q2, Q9Y5A8 | Gene names | PRPF40A, FLAF1, FNBP3, HYPA | |||
|
Domain Architecture |

|
|||||
| Description | Pre-mRNA-processing factor 40 homolog A (Formin-binding protein 3) (Huntingtin yeast partner A) (Huntingtin-interacting protein HYPA/FBP11) (Fas ligand-associated factor 1) (NY-REN-6 antigen). | |||||
|
WDR60_MOUSE
|
||||||
| NC score | 0.054430 (rank : 52) | θ value | θ > 10 (rank : 107) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 344 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8C761, Q3UNM4 | Gene names | Wdr60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat protein 60. | |||||
|
IF3A_HUMAN
|
||||||
| NC score | 0.054339 (rank : 53) | θ value | θ > 10 (rank : 89) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 875 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q14152, O00653 | Gene names | EIF3S10, KIAA0139 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 10 (eIF-3 theta) (eIF3 p167) (eIF3 p180) (eIF3 p185) (eIF3a). | |||||
|
IF3A_MOUSE
|
||||||
| NC score | 0.054307 (rank : 54) | θ value | θ > 10 (rank : 90) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 871 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P23116, Q60697, Q62162 | Gene names | Eif3s10, Csma, Eif3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 10 (eIF-3 theta) (eIF3 p167) (eIF3 p180) (eIF3 p185) (eIF3a) (p162 protein) (Centrosomin). | |||||
|
IF2P_HUMAN
|
||||||
| NC score | 0.054061 (rank : 55) | θ value | θ > 10 (rank : 88) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 889 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | O60841, O95805, Q9UF81, Q9UMN7 | Gene names | EIF5B, IF2, KIAA0741 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 5B (eIF-5B) (Translation initiation factor IF-2). | |||||
|
SFR17_HUMAN
|
||||||
| NC score | 0.053971 (rank : 56) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8TF01, Q5T064, Q6P2B4, Q6P2N4, Q6PJ93, Q6PK36, Q7Z640, Q8TF00, Q96K10, Q96SM5, Q9P076, Q9P0C0, Q9Y4N3 | Gene names | SRRP130, C6orf111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 130 (Serine-arginine-rich- splicing regulatory protein 130) (SRrp130) (SR-rich protein) (SR- related protein). | |||||
|
XE7_HUMAN
|
||||||
| NC score | 0.053970 (rank : 57) | θ value | θ > 10 (rank : 108) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 738 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q02040, Q02832 | Gene names | CXYorf3, DXYS155E, XE7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-lymphocyte antigen precursor (B-lymphocyte surface antigen) (721P) (Protein XE7). | |||||
|
LEO1_HUMAN
|
||||||
| NC score | 0.053088 (rank : 58) | θ value | 0.813845 (rank : 25) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
ANR12_HUMAN
|
||||||
| NC score | 0.052612 (rank : 59) | θ value | 4.03905 (rank : 50) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1081 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q6UB98, O94951, Q658K1, Q6QMF7, Q9H231, Q9H784 | Gene names | ANKRD12, ANCO2, KIAA0874 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 12 (Ankyrin repeat-containing cofactor 2) (GAC-1 protein). | |||||
|
C8AP2_HUMAN
|
||||||
| NC score | 0.052554 (rank : 60) | θ value | 2.36792 (rank : 39) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 548 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UKL3, Q6PH76, Q7LCQ7, Q86YD9, Q9NUQ4, Q9NZV9, Q9P2N1, Q9Y563 | Gene names | CASP8AP2, FLASH, KIAA1315, RIP25 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
TAF3_HUMAN
|
||||||
| NC score | 0.052378 (rank : 61) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 568 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q5VWG9, Q6GMS5, Q6P6B5, Q86VY6, Q9BQS9, Q9UFI8 | Gene names | TAF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
CAF1A_MOUSE
|
||||||
| NC score | 0.050866 (rank : 62) | θ value | θ > 10 (rank : 82) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 736 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q9QWF0 | Gene names | Chaf1a, Caip150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromatin assembly factor 1 subunit A (CAF-1 subunit A) (Chromatin assembly factor I p150 subunit) (CAF-I 150 kDa subunit) (CAF-Ip150). | |||||
|
TAF3_MOUSE
|
||||||
| NC score | 0.050773 (rank : 63) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 463 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q5HZG4, Q3U490, Q3UWX2, Q8BIU8, Q99JH4 | Gene names | Taf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
NUCB1_MOUSE
|
||||||
| NC score | 0.050079 (rank : 64) | θ value | θ > 10 (rank : 93) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q02819, Q3TU42, Q9CWA1, Q9D8J4, Q9DBL3 | Gene names | Nucb1, Nuc, Nucb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleobindin-1 precursor (CALNUC). | |||||
|
HOOK3_HUMAN
|
||||||
| NC score | 0.049405 (rank : 65) | θ value | 0.365318 (rank : 17) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1070 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q86VS8, Q8NBH0, Q9BY13 | Gene names | HOOK3 | |||
|
Domain Architecture |

|
|||||
| Description | Hook homolog 3 (hHK3). | |||||
|
PRGC1_MOUSE
|
||||||
| NC score | 0.048093 (rank : 66) | θ value | 1.81305 (rank : 35) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O70343 | Gene names | Ppargc1a, Pgc1, Pgc1a, Ppargc1 | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisome proliferator-activated receptor gamma coactivator 1-alpha (PPAR gamma coactivator 1-alpha) (PPARGC-1-alpha) (PGC-1-alpha). | |||||
|
LEO1_MOUSE
|
||||||
| NC score | 0.048062 (rank : 67) | θ value | 8.99809 (rank : 74) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
CCD21_MOUSE
|
||||||
| NC score | 0.047462 (rank : 68) | θ value | 6.88961 (rank : 66) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8BMK0, Q8BUF1, Q8K0E6 | Gene names | Ccdc21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 21. | |||||
|
NUCB1_HUMAN
|
||||||
| NC score | 0.047417 (rank : 69) | θ value | 3.0926 (rank : 44) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q02818, Q15838, Q7Z4J7, Q9BUR1 | Gene names | NUCB1, NUC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleobindin-1 precursor (CALNUC). | |||||
|
CCD21_HUMAN
|
||||||
| NC score | 0.046955 (rank : 70) | θ value | 4.03905 (rank : 51) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1228 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q6P2H3, Q5VY68, Q5VY70, Q9H6Q1, Q9H828, Q9UF52 | Gene names | CCDC21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 21. | |||||
|
CLGN_MOUSE
|
||||||
| NC score | 0.045627 (rank : 71) | θ value | 0.62314 (rank : 22) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P52194, Q9D2K5 | Gene names | Clgn, Meg1 | |||
|
Domain Architecture |

|
|||||
| Description | Calmegin precursor (MEG 1 antigen) (Calnexin-T) (A2/6). | |||||
|
NEST_HUMAN
|
||||||
| NC score | 0.044698 (rank : 72) | θ value | 3.0926 (rank : 43) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1305 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P48681, O00552 | Gene names | NES | |||
|
Domain Architecture |

|
|||||
| Description | Nestin. | |||||
|
GOGA4_MOUSE
|
||||||
| NC score | 0.044246 (rank : 73) | θ value | 1.81305 (rank : 32) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1939 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q91VW5, O70365, Q8CGH6 | Gene names | Golga4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (tGolgin-1). | |||||
|
DESP_HUMAN
|
||||||
| NC score | 0.043814 (rank : 74) | θ value | 2.36792 (rank : 40) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P15924, O75993, Q14189, Q9UHN4 | Gene names | DSP | |||
|
Domain Architecture |

|
|||||
| Description | Desmoplakin (DP) (250/210 kDa paraneoplastic pemphigus antigen). | |||||
|
CROCC_MOUSE
|
||||||
| NC score | 0.042771 (rank : 75) | θ value | 4.03905 (rank : 52) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1460 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8CJ40, Q7TQL2, Q80U01, Q8R0B9 | Gene names | Crocc, Kiaa0445 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rootletin (Ciliary rootlet coiled-coil protein). | |||||
|
ITSN1_HUMAN
|
||||||
| NC score | 0.042417 (rank : 76) | θ value | 4.03905 (rank : 54) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1757 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q15811, O95216, Q9UET5, Q9UK60, Q9UNK1, Q9UNK2, Q9UQ92 | Gene names | ITSN1, ITSN, SH3D1A | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-1 (SH3 domain-containing protein 1A) (SH3P17). | |||||
|
TPR_HUMAN
|
||||||
| NC score | 0.040706 (rank : 77) | θ value | 4.03905 (rank : 57) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
CALU_MOUSE
|
||||||
| NC score | 0.039742 (rank : 78) | θ value | 0.813845 (rank : 24) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O35887 | Gene names | Calu | |||
|
Domain Architecture |

|
|||||
| Description | Calumenin precursor (Crocalbin). | |||||
|
CALU_HUMAN
|
||||||
| NC score | 0.039564 (rank : 79) | θ value | 0.813845 (rank : 23) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O43852, O60456, Q6FHB9, Q96RL3, Q9NR43 | Gene names | CALU | |||
|
Domain Architecture |

|
|||||
| Description | Calumenin precursor (Crocalbin) (IEF SSP 9302). | |||||
|
HOOK3_MOUSE
|
||||||
| NC score | 0.039510 (rank : 80) | θ value | 4.03905 (rank : 53) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1007 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8BUK6, Q8BV48, Q8BW57, Q8BY41 | Gene names | Hook3 | |||
|
Domain Architecture |

|
|||||
| Description | Hook homolog 3. | |||||
|
PPIL4_HUMAN
|
||||||
| NC score | 0.038826 (rank : 81) | θ value | 1.06291 (rank : 27) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8WUA2, Q7Z3Q5 | Gene names | PPIL4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peptidyl-prolyl cis-trans isomerase-like 4 (EC 5.2.1.8) (PPIase) (Rotamase) (Cyclophilin-like protein PPIL4). | |||||
|
MYO6_HUMAN
|
||||||
| NC score | 0.038517 (rank : 82) | θ value | 1.81305 (rank : 34) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 727 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q9UM54, Q9BZZ7, Q9UEG2 | Gene names | MYO6, KIAA0389 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-6 (Myosin VI). | |||||
|
MY18A_MOUSE
|
||||||
| NC score | 0.038472 (rank : 83) | θ value | 4.03905 (rank : 55) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1989 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9JMH9, Q811D7 | Gene names | Myo18a, Myspdz | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
ELOA1_HUMAN
|
||||||
| NC score | 0.038295 (rank : 84) | θ value | 5.27518 (rank : 63) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q14241, Q8IXH1 | Gene names | TCEB3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription elongation factor B polypeptide 3 (RNA polymerase II transcription factor SIII subunit A1) (SIII p110) (Elongin-A) (EloA) (Elongin 110 kDa subunit). | |||||
|
MY18A_HUMAN
|
||||||
| NC score | 0.035596 (rank : 85) | θ value | 6.88961 (rank : 70) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1929 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q92614, Q8IXP8 | Gene names | MYO18A, KIAA0216, MYSPDZ | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
MYH9_MOUSE
|
||||||
| NC score | 0.035485 (rank : 86) | θ value | 6.88961 (rank : 71) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1829 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q8VDD5 | Gene names | Myh9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
MYH9_HUMAN
|
||||||
| NC score | 0.035218 (rank : 87) | θ value | 8.99809 (rank : 75) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P35579, O60805 | Gene names | MYH9 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
RAI14_MOUSE
|
||||||
| NC score | 0.033553 (rank : 88) | θ value | 3.0926 (rank : 45) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1505 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9EP71, Q3URT3, Q6ZPT6 | Gene names | Rai14, Kiaa1334, Norpeg | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankycorbin (Ankyrin repeat and coiled-coil structure-containing protein) (Retinoic acid-induced protein 14) (Novel retinal pigment epithelial cell protein) (p125). | |||||
|
MPP8_HUMAN
|
||||||
| NC score | 0.032237 (rank : 89) | θ value | 6.88961 (rank : 69) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 948 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q99549, Q86TK3, Q9BTP1 | Gene names | MPHOSPH8, MPP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | M-phase phosphoprotein 8. | |||||
|
BPA1_MOUSE
|
||||||
| NC score | 0.031947 (rank : 90) | θ value | 5.27518 (rank : 59) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q91ZU6, Q91ZU7 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 1/2/3/4 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
SDCG8_HUMAN
|
||||||
| NC score | 0.031329 (rank : 91) | θ value | 8.99809 (rank : 78) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1002 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q86SQ7, O60527, Q3ZCR6, Q8N5F2, Q9P0F1 | Gene names | SDCCAG8, CCCAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serologically defined colon cancer antigen 8 (Centrosomal colon cancer autoantigen protein) (hCCCAP) (Antigen NY-CO-8). | |||||
|
CEP55_HUMAN
|
||||||
| NC score | 0.031303 (rank : 92) | θ value | 5.27518 (rank : 60) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q53EZ4, Q32WF5, Q3MV20, Q5VY28, Q6N034, Q96H32, Q9NVS7 | Gene names | CEP55, C10orf3, URCC6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosome protein 55 (Up-regulated in colon cancer 6). | |||||
|
CJ006_MOUSE
|
||||||
| NC score | 0.030793 (rank : 93) | θ value | 5.27518 (rank : 61) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6P9P0, Q8C041, Q8C0J5 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf6 homolog. | |||||
|
MYO5B_HUMAN
|
||||||
| NC score | 0.030298 (rank : 94) | θ value | 3.0926 (rank : 42) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 911 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9ULV0 | Gene names | MYO5B, KIAA1119 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-5B (Myosin Vb). | |||||
|
RILP_MOUSE
|
||||||
| NC score | 0.028109 (rank : 95) | θ value | 8.99809 (rank : 76) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 257 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q5ND29, Q80WB8, Q8CHY9 | Gene names | Rilp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab-interacting lysosomal protein. | |||||
|
KIF4A_HUMAN
|
||||||
| NC score | 0.027986 (rank : 96) | θ value | 5.27518 (rank : 64) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1159 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O95239, Q86TN3, Q86XX7, Q9NNY6, Q9NY24, Q9UMW3 | Gene names | KIF4A, KIF4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromosome-associated kinesin KIF4A (Chromokinesin). | |||||
|
CN120_MOUSE
|
||||||
| NC score | 0.027865 (rank : 97) | θ value | 8.99809 (rank : 73) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9DB96, Q8CIJ2 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | Protein C14orf120 homolog. | |||||
|
MRCKB_MOUSE
|
||||||
| NC score | 0.026347 (rank : 98) | θ value | 1.38821 (rank : 29) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 2180 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q7TT50, Q69ZR5, Q80W33 | Gene names | Cdc42bpb, Kiaa1124 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK beta (EC 2.7.11.1) (CDC42-binding protein kinase beta) (Myotonic dystrophy kinase-related CDC42-binding kinase beta) (Myotonic dystrophy protein kinase-like beta) (MRCK beta) (DMPK-like beta). | |||||
|
CTRO_MOUSE
|
||||||
| NC score | 0.025104 (rank : 99) | θ value | 3.0926 (rank : 41) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 2344 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P49025, O88528, O88937, O88938, Q3UM99, Q8CIJ1 | Gene names | Cit | |||
|
Domain Architecture |

|
|||||
| Description | Citron Rho-interacting kinase (EC 2.7.11.1) (CRIK) (Rho-interacting, serine/threonine-protein kinase 21). | |||||
|
ZHX2_MOUSE
|
||||||
| NC score | 0.022578 (rank : 100) | θ value | 4.03905 (rank : 58) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 435 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8C0C0 | Gene names | Zhx2, Afr1, Raf | |||
|
Domain Architecture |

|
|||||
| Description | Zinc fingers and homeoboxes protein 2 (Zinc finger and homeodomain protein 2) (Alpha-fetoprotein regulator 1) (AFP regulator 1) (Regulator of AFP). | |||||
|
SBP1_HUMAN
|
||||||
| NC score | 0.020580 (rank : 101) | θ value | 8.99809 (rank : 77) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13228 | Gene names | SELENBP1, SBP | |||
|
Domain Architecture |

|
|||||
| Description | Selenium-binding protein 1. | |||||
|
MDC1_HUMAN
|
||||||
| NC score | 0.019937 (rank : 102) | θ value | 6.88961 (rank : 68) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
TAOK1_MOUSE
|
||||||
| NC score | 0.019007 (rank : 103) | θ value | 3.0926 (rank : 49) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1720 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q5F2E8, Q69ZL2, Q8JZX2, Q91VG7 | Gene names | Taok1, Kiaa1361 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase TAO1 (EC 2.7.11.1) (Thousand and one amino acid protein 1). | |||||
|
TAOK1_HUMAN
|
||||||
| NC score | 0.018631 (rank : 104) | θ value | 3.0926 (rank : 48) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 1734 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q7L7X3, Q96L75, Q9H2K7, Q9H7S5, Q9P2I6 | Gene names | TAOK1, KIAA1361, MAP3K16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase TAO1 (EC 2.7.11.1) (Thousand and one amino acid protein 1) (STE20-like kinase PSK2) (Kinase from chicken homolog B) (hKFC-B). | |||||
|
SRBS1_HUMAN
|
||||||
| NC score | 0.013073 (rank : 105) | θ value | 8.99809 (rank : 79) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 396 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9BX66, Q7LBE5, Q8IVK0, Q8IVQ4, Q96KF3, Q96KF4, Q9BX64, Q9BX65, Q9P2Q0, Q9UFT2, Q9UHN7, Q9Y338 | Gene names | SORBS1, KIAA1296, SH3D5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sorbin and SH3 domain-containing protein 1 (Ponsin) (c-Cbl-associated protein) (CAP) (SH3 domain protein 5) (SH3P12). | |||||
|
STAM2_MOUSE
|
||||||
| NC score | 0.012331 (rank : 106) | θ value | 5.27518 (rank : 65) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O88811, Q8C8Y4 | Gene names | Stam2, Hbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal transducing adapter molecule 2 (STAM-2) (Hrs-binding protein). | |||||
|
GEM_HUMAN
|
||||||
| NC score | 0.008358 (rank : 107) | θ value | 6.88961 (rank : 67) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P55040 | Gene names | GEM, KIR | |||
|
Domain Architecture |

|
|||||
| Description | GTP-binding protein GEM (GTP-binding mitogen-induced T-cell protein) (RAS-like protein KIR). | |||||
|
DSG3_HUMAN
|
||||||
| NC score | 0.004714 (rank : 108) | θ value | 5.27518 (rank : 62) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P32926 | Gene names | DSG3 | |||
|
Domain Architecture |

|
|||||
| Description | Desmoglein-3 precursor (130 kDa pemphigus vulgaris antigen) (PVA). | |||||
|
ZN192_HUMAN
|
||||||
| NC score | 0.002137 (rank : 109) | θ value | 1.81305 (rank : 38) | |||
| Query Neighborhood Hits | 79 | Target Neighborhood Hits | 789 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15776, Q4VAR1, Q4VAR2, Q4VAR3, Q9H4T1 | Gene names | ZNF192 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 192 (LD5-1). | |||||