Please be patient as the page loads
|
PKP3_MOUSE
|
||||||
| SwissProt Accessions | Q9QY23 | Gene names | Pkp3 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-3. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
PKP3_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.997772 (rank : 2) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9Y446 | Gene names | PKP3 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-3. | |||||
|
PKP3_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9QY23 | Gene names | Pkp3 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-3. | |||||
|
PKP1_MOUSE
|
||||||
| θ value | 2.65329e-68 (rank : 3) | NC score | 0.927583 (rank : 3) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P97350 | Gene names | Pkp1 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-1. | |||||
|
PKP4_MOUSE
|
||||||
| θ value | 2.93355e-67 (rank : 4) | NC score | 0.900334 (rank : 10) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q68FH0, Q640N0, Q68G56, Q8BK47, Q8BVH1, Q9CRE3 | Gene names | Pkp4, Armrp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plakophilin-4 (Armadillo-related protein). | |||||
|
PKP1_HUMAN
|
||||||
| θ value | 7.98882e-65 (rank : 5) | NC score | 0.924419 (rank : 4) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q13835, O00645, Q15152 | Gene names | PKP1 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-1 (Band-6 protein) (B6P). | |||||
|
ARVC_HUMAN
|
||||||
| θ value | 8.83269e-64 (rank : 6) | NC score | 0.909052 (rank : 7) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O00192 | Gene names | ARVCF | |||
|
Domain Architecture |

|
|||||
| Description | Armadillo repeat protein deleted in velo-cardio-facial syndrome. | |||||
|
CTND2_HUMAN
|
||||||
| θ value | 8.83269e-64 (rank : 7) | NC score | 0.874562 (rank : 13) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UQB3, O00379, O15390, O43206, O43840, Q13589, Q9UM66, Q9UPM3 | Gene names | CTNND2, NPRAP | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-2 (Delta-catenin) (Neural plakophilin-related ARM-repeat protein) (NPRAP) (Neurojungin) (GT24). | |||||
|
PKP4_HUMAN
|
||||||
| θ value | 1.50663e-63 (rank : 8) | NC score | 0.898165 (rank : 11) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q99569 | Gene names | PKP4 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-4 (p0071). | |||||
|
ARVC_MOUSE
|
||||||
| θ value | 5.7252e-63 (rank : 9) | NC score | 0.911166 (rank : 6) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P98203, Q6PGJ6, Q8BQ36, Q8BRF2, Q8C3U7, Q924L2, Q924L3, Q924L4, Q924L5 | Gene names | Arvcf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat protein deleted in velo-cardio-facial syndrome homolog. | |||||
|
CTND1_HUMAN
|
||||||
| θ value | 1.84173e-61 (rank : 10) | NC score | 0.906203 (rank : 9) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O60716, O15088, O60713, O60714, O60715, O60935, Q6RBX8, Q9UP71, Q9UP72, Q9UP73 | Gene names | CTNND1, KIAA0384 | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-1 (p120 catenin) (p120(ctn)) (Cadherin-associated Src substrate) (CAS) (p120(cas)). | |||||
|
CTND1_MOUSE
|
||||||
| θ value | 2.03627e-60 (rank : 11) | NC score | 0.906293 (rank : 8) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P30999 | Gene names | Ctnnd1, Catns | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-1 (p120 catenin) (p120(ctn)) (Cadherin-associated Src substrate) (CAS) (p120(cas)). | |||||
|
CTND2_MOUSE
|
||||||
| θ value | 3.47335e-60 (rank : 12) | NC score | 0.884846 (rank : 12) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O35927 | Gene names | Ctnnd2, Catnd2, Nprap | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-2 (Neural plakophilin-related ARM-repeat protein) (NPRAP) (Neurojungin). | |||||
|
PKP2_HUMAN
|
||||||
| θ value | 1.23536e-57 (rank : 13) | NC score | 0.922045 (rank : 5) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q99959, Q99960 | Gene names | PKP2 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-2. | |||||
|
PLAK_MOUSE
|
||||||
| θ value | 0.00298849 (rank : 14) | NC score | 0.239405 (rank : 17) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q02257, Q8CGD3 | Gene names | Jup | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junction plakoglobin (Desmoplakin-3) (Desmoplakin III). | |||||
|
PLAK_HUMAN
|
||||||
| θ value | 0.00665767 (rank : 15) | NC score | 0.207855 (rank : 18) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P14923, Q9HCX9 | Gene names | JUP, DP3 | |||
|
Domain Architecture |

|
|||||
| Description | Junction plakoglobin (Desmoplakin-3) (Desmoplakin III). | |||||
|
ARMC4_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 16) | NC score | 0.337480 (rank : 14) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q5T2S8, Q9H0C0 | Gene names | ARMC4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing protein 4. | |||||
|
CTNB1_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 17) | NC score | 0.254992 (rank : 16) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P35222, Q8NEW9, Q8NI94, Q9H391 | Gene names | CTNNB1, CTNNB | |||
|
Domain Architecture |

|
|||||
| Description | Catenin beta-1 (Beta-catenin). | |||||
|
CTNB1_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 18) | NC score | 0.255047 (rank : 15) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q02248, Q922W1, Q9D335 | Gene names | Ctnnb1, Catnb | |||
|
Domain Architecture |

|
|||||
| Description | Catenin beta-1 (Beta-catenin). | |||||
|
NFKB1_HUMAN
|
||||||
| θ value | 0.21417 (rank : 19) | NC score | 0.018046 (rank : 38) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 306 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P19838, Q68D84, Q86V43, Q8N4X7, Q9NZC0 | Gene names | NFKB1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor NF-kappa-B p105 subunit (DNA-binding factor KBF1) (EBP- 1) [Contains: Nuclear factor NF-kappa-B p50 subunit]. | |||||
|
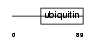
UBL7_HUMAN
|
||||||
| θ value | 0.62314 (rank : 20) | NC score | 0.029659 (rank : 32) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96S82, Q96I03 | Gene names | UBL7, BMSCUBP | |||
|
Domain Architecture |
|
|||||
| Description | Ubiquitin-like protein 7 (Ubiquitin-like protein SB132) (Bone marrow stromal cell ubiquitin-like protein) (BMSC-UbP). | |||||
|
SYNE1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 21) | NC score | 0.011261 (rank : 39) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1484 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8NF91, O94890, Q5JV19, Q5JV22, Q8N9P7, Q8TCP1, Q8WWW6, Q8WWW7, Q8WXF6, Q96N17, Q9C0A7, Q9H525, Q9H526, Q9NS36, Q9NU50, Q9UJ06, Q9UJ07, Q9ULF8 | Gene names | SYNE1, KIAA0796, KIAA1262, KIAA1756, MYNE1 | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-1 (Nuclear envelope spectrin repeat protein 1) (Synaptic nuclear envelope protein 1) (Syne-1) (Myocyte nuclear envelope protein 1) (Myne-1) (Enaptin). | |||||
|
AIRE_MOUSE
|
||||||
| θ value | 1.06291 (rank : 22) | NC score | 0.018514 (rank : 37) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Z0E3, Q9JLW0, Q9JLW1, Q9JLW2, Q9JLW3, Q9JLW4, Q9JLW5, Q9JLW6, Q9JLW7, Q9JLW8, Q9JLW9, Q9JLX0 | Gene names | Aire | |||
|
Domain Architecture |

|
|||||
| Description | Autoimmune regulator (Autoimmune polyendocrinopathy candidiasis ectodermal dystrophy protein homolog) (APECED protein homolog). | |||||
|
APC_HUMAN
|
||||||
| θ value | 1.06291 (rank : 23) | NC score | 0.172223 (rank : 20) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 603 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P25054, Q15162, Q15163, Q93042 | Gene names | APC, DP2.5 | |||
|
Domain Architecture |

|
|||||
| Description | Adenomatous polyposis coli protein (Protein APC). | |||||
|
APC_MOUSE
|
||||||
| θ value | 1.06291 (rank : 24) | NC score | 0.177151 (rank : 19) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q61315, Q62044 | Gene names | Apc | |||
|
Domain Architecture |

|
|||||
| Description | Adenomatous polyposis coli protein (Protein APC) (mAPC). | |||||
|
ERC2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 25) | NC score | 0.009993 (rank : 44) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1156 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O15083, Q86TK4 | Gene names | ERC2, KIAA0378 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ERC protein 2. | |||||
|
ERC2_MOUSE
|
||||||
| θ value | 1.06291 (rank : 26) | NC score | 0.010013 (rank : 43) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1166 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6PH08, Q80U20, Q8CCP1 | Gene names | Erc2, Cast1, D14Ertd171e, Kiaa0378 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ERC protein 2 (CAZ-associated structural protein 1) (CAST1). | |||||
|
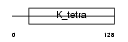
ZBT7A_MOUSE
|
||||||
| θ value | 1.81305 (rank : 27) | NC score | 0.000398 (rank : 51) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 795 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O88939, Q9CRJ0 | Gene names | Zbtb7a, Lrf, Zbtb7 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger and BTB domain-containing protein 7A (Leukemia/lymphoma- related factor). | |||||
|
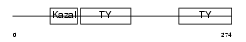
SMOC2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 28) | NC score | 0.019409 (rank : 36) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H3U7, Q5TAT7, Q5TAT8, Q86VV9, Q96SF3, Q9H1L3, Q9H1L4, Q9H3U0, Q9H4F7, Q9HCV2 | Gene names | SMOC2, SMAP2 | |||
|
Domain Architecture |
|
|||||
| Description | SPARC-related modular calcium-binding protein 2 precursor (Secreted modular calcium-binding protein 2) (SMOC-2) (Smooth muscle-associated protein 2) (SMAP-2). | |||||
|
TNPO2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 29) | NC score | 0.032914 (rank : 30) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O14787, O14655 | Gene names | TNPO2 | |||
|
Domain Architecture |

|
|||||
| Description | Transportin-2 (Karyopherin beta-2b). | |||||
|
TNPO2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 30) | NC score | 0.032877 (rank : 31) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q99LG2, Q8C0U9 | Gene names | Tnpo2 | |||
|
Domain Architecture |

|
|||||
| Description | Transportin-2 (Karyopherin beta-2b). | |||||
|
ZBT7A_HUMAN
|
||||||
| θ value | 3.0926 (rank : 31) | NC score | 0.000086 (rank : 52) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 823 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O95365, O00456 | Gene names | ZBTB7A, FBI1, LRF, ZBTB7 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 7A (Leukemia/lymphoma- related factor) (Factor that binds to inducer of short transcripts protein 1) (Factor binding IST protein 1) (FBI-1) (HIV-1 1st-binding protein 1) (TTF-I-interacting peptide 21) (TIP21). | |||||
|
GDS1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 32) | NC score | 0.133943 (rank : 21) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P52306, Q9NYM2, Q9NZA8 | Gene names | RAP1GDS1 | |||
|
Domain Architecture |

|
|||||
| Description | Rap1 GTPase-GDP dissociation stimulator 1 (SMG P21 stimulatory GDP/GTP exchange protein) (SMG GDS protein) (Exchange factor smgGDS). | |||||
|
RB6I2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 33) | NC score | 0.005876 (rank : 47) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1584 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IUD2, Q6NVK2, Q8IUD3, Q8IUD4, Q8IUD5, Q8NAS1, Q9NXN5, Q9UIK7, Q9UPS1 | Gene names | ERC1, ELKS, KIAA1081, RAB6IP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ELKS/RAB6-interacting/CAST family member 1 (RAB6-interacting protein 2) (ERC protein 1). | |||||
|
TIF1A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 34) | NC score | 0.010332 (rank : 42) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O15164, O95854 | Gene names | TRIM24, RNF82, TIF1, TIF1A | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-alpha (TIF1-alpha) (Tripartite motif-containing protein 24) (RING finger protein 82). | |||||
|
TIF1A_MOUSE
|
||||||
| θ value | 4.03905 (rank : 35) | NC score | 0.010519 (rank : 41) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q64127, Q64126 | Gene names | Trim24, Tif1, Tif1a | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-alpha (TIF1-alpha) (Tripartite motif-containing protein 24). | |||||
|
TNPO1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 36) | NC score | 0.029301 (rank : 33) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q92973, Q92957, Q92975 | Gene names | TNPO1, KPNB2, MIP1, TRN | |||
|
Domain Architecture |

|
|||||
| Description | Transportin-1 (Importin beta-2) (Karyopherin beta-2) (M9 region interaction protein) (MIP). | |||||
|
TNPO1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 37) | NC score | 0.028084 (rank : 34) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BFY9, Q8C0S3, Q8K2E7 | Gene names | Tnpo1, Kpnb2 | |||
|
Domain Architecture |

|
|||||
| Description | Transportin-1 (Importin beta-2) (Karyopherin beta-2). | |||||
|
TTF1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 38) | NC score | 0.025934 (rank : 35) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q62187, Q9JKK5 | Gene names | Ttf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription termination factor 1 (TTF-1) (TTF-I) (mTFF-I) (RNA polymerase I termination factor). | |||||
|
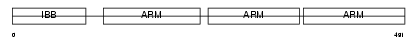
IMA5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 39) | NC score | 0.066877 (rank : 23) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O15131 | Gene names | KPNA5 | |||
|
Domain Architecture |
|
|||||
| Description | Importin alpha-6 subunit (Karyopherin alpha-5 subunit). | |||||
|
KCNK7_MOUSE
|
||||||
| θ value | 6.88961 (rank : 40) | NC score | 0.007206 (rank : 46) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Z2T1, Q9QXY0, Q9QYE8, Q9R1V1, Q9R242 | Gene names | Kcnk7, Dpkch3, Kcnk6, Kcnk8, Knot1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 7 (Putative potassium channel DP3) (Double-pore K(+) channel 3) (Neuromuscular two p domain potassium channel). | |||||
|
MSX1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 41) | NC score | 0.001216 (rank : 50) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P28360, Q96NY4 | Gene names | MSX1, HOX7 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein MSX-1 (Msh homeobox 1-like protein) (Hox-7). | |||||
|
YTHD3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 42) | NC score | 0.007405 (rank : 45) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BYK6, Q3UVI5, Q6NXJ6, Q6NXJ8, Q8BKB6 | Gene names | Ythdf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YTH domain family protein 3. | |||||
|
FUT4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 43) | NC score | 0.005660 (rank : 48) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q11127 | Gene names | Fut4, Elft | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-(1,3)-fucosyltransferase (EC 2.4.1.-) (Galactoside 3-L- fucosyltransferase) (Fucosyltransferase 4) (FUCT-IV) (Fuc-TIV). | |||||
|
IMA3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 44) | NC score | 0.050492 (rank : 29) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O35344 | Gene names | Kpna3 | |||
|
Domain Architecture |
|
|||||
| Description | Importin alpha-3 subunit (Karyopherin alpha-3 subunit) (Importin alpha Q2). | |||||
|
KIF2C_HUMAN
|
||||||
| θ value | 8.99809 (rank : 45) | NC score | 0.002105 (rank : 49) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99661, Q96C18, Q96HB8, Q9BWV8 | Gene names | KIF2C, KNSL6 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF2C (Mitotic centromere-associated kinesin) (MCAK) (Kinesin-like protein 6). | |||||
|
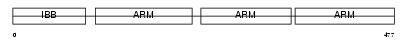
SMOC2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 46) | NC score | 0.010955 (rank : 40) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CD91, Q8VCM1, Q9D9K2, Q9ER95 | Gene names | Smoc2 | |||
|
Domain Architecture |
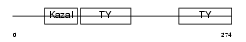
|
|||||
| Description | SPARC-related modular calcium-binding protein 2 precursor (Secreted modular calcium-binding protein 2) (SMOC-2). | |||||
|
IMA2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 47) | NC score | 0.055977 (rank : 25) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P52292, Q9BRU5 | Gene names | KPNA2, RCH1, SRP1 | |||
|
Domain Architecture |
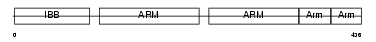
|
|||||
| Description | Importin alpha-2 subunit (Karyopherin alpha-2 subunit) (SRP1-alpha) (RAG cohort protein 1). | |||||
|
IMA2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 48) | NC score | 0.055971 (rank : 26) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P52293, Q64292 | Gene names | Kpna2, Rch1 | |||
|
Domain Architecture |
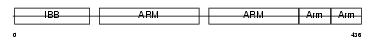
|
|||||
| Description | Importin alpha-2 subunit (Karyopherin alpha-2 subunit) (SRP1-alpha) (RAG cohort protein 1) (Pendulin) (Pore targeting complex 58 kDa subunit) (PTAC58) (Importin alpha P1). | |||||
|
IMA4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 49) | NC score | 0.052205 (rank : 28) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O00629, O00190 | Gene names | KPNA4, QIP1 | |||
|
Domain Architecture |
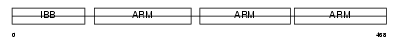
|
|||||
| Description | Importin alpha-4 subunit (Karyopherin alpha-4 subunit) (Qip1 protein). | |||||
|
IMA4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 50) | NC score | 0.052278 (rank : 27) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O35343 | Gene names | Kpna4 | |||
|
Domain Architecture |
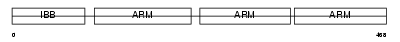
|
|||||
| Description | Importin alpha-4 subunit (Karyopherin alpha-4 subunit) (Importin alpha Q1). | |||||
|
IMA7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 51) | NC score | 0.065787 (rank : 24) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O60684 | Gene names | KPNA6, IPOA7 | |||
|
Domain Architecture |
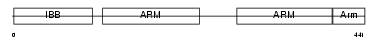
|
|||||
| Description | Importin alpha-7 subunit (Karyopherin alpha-6). | |||||
|
IMA7_MOUSE
|
||||||
| θ value | θ > 10 (rank : 52) | NC score | 0.068147 (rank : 22) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O35345, Q9CVP9 | Gene names | Kpna6, Kpna5 | |||
|
Domain Architecture |
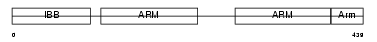
|
|||||
| Description | Importin alpha-7 subunit (Karyopherin alpha-6 subunit) (Importin alpha S2). | |||||
|
PKP3_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9QY23 | Gene names | Pkp3 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-3. | |||||
|
PKP3_HUMAN
|
||||||
| NC score | 0.997772 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9Y446 | Gene names | PKP3 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-3. | |||||
|
PKP1_MOUSE
|
||||||
| NC score | 0.927583 (rank : 3) | θ value | 2.65329e-68 (rank : 3) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P97350 | Gene names | Pkp1 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-1. | |||||
|
PKP1_HUMAN
|
||||||
| NC score | 0.924419 (rank : 4) | θ value | 7.98882e-65 (rank : 5) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q13835, O00645, Q15152 | Gene names | PKP1 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-1 (Band-6 protein) (B6P). | |||||
|
PKP2_HUMAN
|
||||||
| NC score | 0.922045 (rank : 5) | θ value | 1.23536e-57 (rank : 13) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q99959, Q99960 | Gene names | PKP2 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-2. | |||||
|
ARVC_MOUSE
|
||||||
| NC score | 0.911166 (rank : 6) | θ value | 5.7252e-63 (rank : 9) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P98203, Q6PGJ6, Q8BQ36, Q8BRF2, Q8C3U7, Q924L2, Q924L3, Q924L4, Q924L5 | Gene names | Arvcf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat protein deleted in velo-cardio-facial syndrome homolog. | |||||
|
ARVC_HUMAN
|
||||||
| NC score | 0.909052 (rank : 7) | θ value | 8.83269e-64 (rank : 6) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O00192 | Gene names | ARVCF | |||
|
Domain Architecture |

|
|||||
| Description | Armadillo repeat protein deleted in velo-cardio-facial syndrome. | |||||
|
CTND1_MOUSE
|
||||||
| NC score | 0.906293 (rank : 8) | θ value | 2.03627e-60 (rank : 11) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P30999 | Gene names | Ctnnd1, Catns | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-1 (p120 catenin) (p120(ctn)) (Cadherin-associated Src substrate) (CAS) (p120(cas)). | |||||
|
CTND1_HUMAN
|
||||||
| NC score | 0.906203 (rank : 9) | θ value | 1.84173e-61 (rank : 10) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O60716, O15088, O60713, O60714, O60715, O60935, Q6RBX8, Q9UP71, Q9UP72, Q9UP73 | Gene names | CTNND1, KIAA0384 | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-1 (p120 catenin) (p120(ctn)) (Cadherin-associated Src substrate) (CAS) (p120(cas)). | |||||
|
PKP4_MOUSE
|
||||||
| NC score | 0.900334 (rank : 10) | θ value | 2.93355e-67 (rank : 4) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q68FH0, Q640N0, Q68G56, Q8BK47, Q8BVH1, Q9CRE3 | Gene names | Pkp4, Armrp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plakophilin-4 (Armadillo-related protein). | |||||
|
PKP4_HUMAN
|
||||||
| NC score | 0.898165 (rank : 11) | θ value | 1.50663e-63 (rank : 8) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q99569 | Gene names | PKP4 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-4 (p0071). | |||||
|
CTND2_MOUSE
|
||||||
| NC score | 0.884846 (rank : 12) | θ value | 3.47335e-60 (rank : 12) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O35927 | Gene names | Ctnnd2, Catnd2, Nprap | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-2 (Neural plakophilin-related ARM-repeat protein) (NPRAP) (Neurojungin). | |||||
|
CTND2_HUMAN
|
||||||
| NC score | 0.874562 (rank : 13) | θ value | 8.83269e-64 (rank : 7) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UQB3, O00379, O15390, O43206, O43840, Q13589, Q9UM66, Q9UPM3 | Gene names | CTNND2, NPRAP | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-2 (Delta-catenin) (Neural plakophilin-related ARM-repeat protein) (NPRAP) (Neurojungin) (GT24). | |||||
|
ARMC4_HUMAN
|
||||||
| NC score | 0.337480 (rank : 14) | θ value | 0.00869519 (rank : 16) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q5T2S8, Q9H0C0 | Gene names | ARMC4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing protein 4. | |||||
|
CTNB1_MOUSE
|
||||||
| NC score | 0.255047 (rank : 15) | θ value | 0.0961366 (rank : 18) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q02248, Q922W1, Q9D335 | Gene names | Ctnnb1, Catnb | |||
|
Domain Architecture |

|
|||||
| Description | Catenin beta-1 (Beta-catenin). | |||||
|
CTNB1_HUMAN
|
||||||
| NC score | 0.254992 (rank : 16) | θ value | 0.0961366 (rank : 17) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P35222, Q8NEW9, Q8NI94, Q9H391 | Gene names | CTNNB1, CTNNB | |||
|
Domain Architecture |

|
|||||
| Description | Catenin beta-1 (Beta-catenin). | |||||
|
PLAK_MOUSE
|
||||||
| NC score | 0.239405 (rank : 17) | θ value | 0.00298849 (rank : 14) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q02257, Q8CGD3 | Gene names | Jup | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junction plakoglobin (Desmoplakin-3) (Desmoplakin III). | |||||
|
PLAK_HUMAN
|
||||||
| NC score | 0.207855 (rank : 18) | θ value | 0.00665767 (rank : 15) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P14923, Q9HCX9 | Gene names | JUP, DP3 | |||
|
Domain Architecture |

|
|||||
| Description | Junction plakoglobin (Desmoplakin-3) (Desmoplakin III). | |||||
|
APC_MOUSE
|
||||||
| NC score | 0.177151 (rank : 19) | θ value | 1.06291 (rank : 24) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q61315, Q62044 | Gene names | Apc | |||
|
Domain Architecture |

|
|||||
| Description | Adenomatous polyposis coli protein (Protein APC) (mAPC). | |||||
|
APC_HUMAN
|
||||||
| NC score | 0.172223 (rank : 20) | θ value | 1.06291 (rank : 23) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 603 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P25054, Q15162, Q15163, Q93042 | Gene names | APC, DP2.5 | |||
|
Domain Architecture |

|
|||||
| Description | Adenomatous polyposis coli protein (Protein APC). | |||||
|
GDS1_HUMAN
|
||||||
| NC score | 0.133943 (rank : 21) | θ value | 4.03905 (rank : 32) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P52306, Q9NYM2, Q9NZA8 | Gene names | RAP1GDS1 | |||
|
Domain Architecture |

|
|||||
| Description | Rap1 GTPase-GDP dissociation stimulator 1 (SMG P21 stimulatory GDP/GTP exchange protein) (SMG GDS protein) (Exchange factor smgGDS). | |||||
|
IMA7_MOUSE
|
||||||
| NC score | 0.068147 (rank : 22) | θ value | θ > 10 (rank : 52) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O35345, Q9CVP9 | Gene names | Kpna6, Kpna5 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-7 subunit (Karyopherin alpha-6 subunit) (Importin alpha S2). | |||||
|
IMA5_HUMAN
|
||||||
| NC score | 0.066877 (rank : 23) | θ value | 6.88961 (rank : 39) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O15131 | Gene names | KPNA5 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-6 subunit (Karyopherin alpha-5 subunit). | |||||
|
IMA7_HUMAN
|
||||||
| NC score | 0.065787 (rank : 24) | θ value | θ > 10 (rank : 51) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O60684 | Gene names | KPNA6, IPOA7 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-7 subunit (Karyopherin alpha-6). | |||||
|
IMA2_HUMAN
|
||||||
| NC score | 0.055977 (rank : 25) | θ value | θ > 10 (rank : 47) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P52292, Q9BRU5 | Gene names | KPNA2, RCH1, SRP1 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-2 subunit (Karyopherin alpha-2 subunit) (SRP1-alpha) (RAG cohort protein 1). | |||||
|
IMA2_MOUSE
|
||||||
| NC score | 0.055971 (rank : 26) | θ value | θ > 10 (rank : 48) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P52293, Q64292 | Gene names | Kpna2, Rch1 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-2 subunit (Karyopherin alpha-2 subunit) (SRP1-alpha) (RAG cohort protein 1) (Pendulin) (Pore targeting complex 58 kDa subunit) (PTAC58) (Importin alpha P1). | |||||
|
IMA4_MOUSE
|
||||||
| NC score | 0.052278 (rank : 27) | θ value | θ > 10 (rank : 50) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O35343 | Gene names | Kpna4 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-4 subunit (Karyopherin alpha-4 subunit) (Importin alpha Q1). | |||||
|
IMA4_HUMAN
|
||||||
| NC score | 0.052205 (rank : 28) | θ value | θ > 10 (rank : 49) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O00629, O00190 | Gene names | KPNA4, QIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-4 subunit (Karyopherin alpha-4 subunit) (Qip1 protein). | |||||
|
IMA3_MOUSE
|
||||||
| NC score | 0.050492 (rank : 29) | θ value | 8.99809 (rank : 44) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O35344 | Gene names | Kpna3 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-3 subunit (Karyopherin alpha-3 subunit) (Importin alpha Q2). | |||||
|
TNPO2_HUMAN
|
||||||
| NC score | 0.032914 (rank : 30) | θ value | 3.0926 (rank : 29) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O14787, O14655 | Gene names | TNPO2 | |||
|
Domain Architecture |

|
|||||
| Description | Transportin-2 (Karyopherin beta-2b). | |||||
|
TNPO2_MOUSE
|
||||||
| NC score | 0.032877 (rank : 31) | θ value | 3.0926 (rank : 30) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q99LG2, Q8C0U9 | Gene names | Tnpo2 | |||
|
Domain Architecture |

|
|||||
| Description | Transportin-2 (Karyopherin beta-2b). | |||||
|
UBL7_HUMAN
|
||||||
| NC score | 0.029659 (rank : 32) | θ value | 0.62314 (rank : 20) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96S82, Q96I03 | Gene names | UBL7, BMSCUBP | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-like protein 7 (Ubiquitin-like protein SB132) (Bone marrow stromal cell ubiquitin-like protein) (BMSC-UbP). | |||||
|
TNPO1_HUMAN
|
||||||
| NC score | 0.029301 (rank : 33) | θ value | 4.03905 (rank : 36) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q92973, Q92957, Q92975 | Gene names | TNPO1, KPNB2, MIP1, TRN | |||
|
Domain Architecture |

|
|||||
| Description | Transportin-1 (Importin beta-2) (Karyopherin beta-2) (M9 region interaction protein) (MIP). | |||||
|
TNPO1_MOUSE
|
||||||
| NC score | 0.028084 (rank : 34) | θ value | 5.27518 (rank : 37) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BFY9, Q8C0S3, Q8K2E7 | Gene names | Tnpo1, Kpnb2 | |||
|
Domain Architecture |

|
|||||
| Description | Transportin-1 (Importin beta-2) (Karyopherin beta-2). | |||||
|
TTF1_MOUSE
|
||||||
| NC score | 0.025934 (rank : 35) | θ value | 5.27518 (rank : 38) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q62187, Q9JKK5 | Gene names | Ttf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription termination factor 1 (TTF-1) (TTF-I) (mTFF-I) (RNA polymerase I termination factor). | |||||
|
SMOC2_HUMAN
|
||||||
| NC score | 0.019409 (rank : 36) | θ value | 3.0926 (rank : 28) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H3U7, Q5TAT7, Q5TAT8, Q86VV9, Q96SF3, Q9H1L3, Q9H1L4, Q9H3U0, Q9H4F7, Q9HCV2 | Gene names | SMOC2, SMAP2 | |||
|
Domain Architecture |

|
|||||
| Description | SPARC-related modular calcium-binding protein 2 precursor (Secreted modular calcium-binding protein 2) (SMOC-2) (Smooth muscle-associated protein 2) (SMAP-2). | |||||
|
AIRE_MOUSE
|
||||||
| NC score | 0.018514 (rank : 37) | θ value | 1.06291 (rank : 22) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Z0E3, Q9JLW0, Q9JLW1, Q9JLW2, Q9JLW3, Q9JLW4, Q9JLW5, Q9JLW6, Q9JLW7, Q9JLW8, Q9JLW9, Q9JLX0 | Gene names | Aire | |||
|
Domain Architecture |

|
|||||
| Description | Autoimmune regulator (Autoimmune polyendocrinopathy candidiasis ectodermal dystrophy protein homolog) (APECED protein homolog). | |||||
|
NFKB1_HUMAN
|
||||||
| NC score | 0.018046 (rank : 38) | θ value | 0.21417 (rank : 19) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 306 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P19838, Q68D84, Q86V43, Q8N4X7, Q9NZC0 | Gene names | NFKB1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor NF-kappa-B p105 subunit (DNA-binding factor KBF1) (EBP- 1) [Contains: Nuclear factor NF-kappa-B p50 subunit]. | |||||
|
SYNE1_HUMAN
|
||||||
| NC score | 0.011261 (rank : 39) | θ value | 0.813845 (rank : 21) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1484 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8NF91, O94890, Q5JV19, Q5JV22, Q8N9P7, Q8TCP1, Q8WWW6, Q8WWW7, Q8WXF6, Q96N17, Q9C0A7, Q9H525, Q9H526, Q9NS36, Q9NU50, Q9UJ06, Q9UJ07, Q9ULF8 | Gene names | SYNE1, KIAA0796, KIAA1262, KIAA1756, MYNE1 | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-1 (Nuclear envelope spectrin repeat protein 1) (Synaptic nuclear envelope protein 1) (Syne-1) (Myocyte nuclear envelope protein 1) (Myne-1) (Enaptin). | |||||
|
SMOC2_MOUSE
|
||||||
| NC score | 0.010955 (rank : 40) | θ value | 8.99809 (rank : 46) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CD91, Q8VCM1, Q9D9K2, Q9ER95 | Gene names | Smoc2 | |||
|
Domain Architecture |

|
|||||
| Description | SPARC-related modular calcium-binding protein 2 precursor (Secreted modular calcium-binding protein 2) (SMOC-2). | |||||
|
TIF1A_MOUSE
|
||||||
| NC score | 0.010519 (rank : 41) | θ value | 4.03905 (rank : 35) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q64127, Q64126 | Gene names | Trim24, Tif1, Tif1a | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-alpha (TIF1-alpha) (Tripartite motif-containing protein 24). | |||||
|
TIF1A_HUMAN
|
||||||
| NC score | 0.010332 (rank : 42) | θ value | 4.03905 (rank : 34) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O15164, O95854 | Gene names | TRIM24, RNF82, TIF1, TIF1A | |||
|
Domain Architecture |

|
|||||
| Description | Transcription intermediary factor 1-alpha (TIF1-alpha) (Tripartite motif-containing protein 24) (RING finger protein 82). | |||||
|
ERC2_MOUSE
|
||||||
| NC score | 0.010013 (rank : 43) | θ value | 1.06291 (rank : 26) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1166 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6PH08, Q80U20, Q8CCP1 | Gene names | Erc2, Cast1, D14Ertd171e, Kiaa0378 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ERC protein 2 (CAZ-associated structural protein 1) (CAST1). | |||||
|
ERC2_HUMAN
|
||||||
| NC score | 0.009993 (rank : 44) | θ value | 1.06291 (rank : 25) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1156 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O15083, Q86TK4 | Gene names | ERC2, KIAA0378 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ERC protein 2. | |||||
|
YTHD3_MOUSE
|
||||||
| NC score | 0.007405 (rank : 45) | θ value | 6.88961 (rank : 42) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BYK6, Q3UVI5, Q6NXJ6, Q6NXJ8, Q8BKB6 | Gene names | Ythdf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YTH domain family protein 3. | |||||
|
KCNK7_MOUSE
|
||||||
| NC score | 0.007206 (rank : 46) | θ value | 6.88961 (rank : 40) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Z2T1, Q9QXY0, Q9QYE8, Q9R1V1, Q9R242 | Gene names | Kcnk7, Dpkch3, Kcnk6, Kcnk8, Knot1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 7 (Putative potassium channel DP3) (Double-pore K(+) channel 3) (Neuromuscular two p domain potassium channel). | |||||
|
RB6I2_HUMAN
|
||||||
| NC score | 0.005876 (rank : 47) | θ value | 4.03905 (rank : 33) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1584 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IUD2, Q6NVK2, Q8IUD3, Q8IUD4, Q8IUD5, Q8NAS1, Q9NXN5, Q9UIK7, Q9UPS1 | Gene names | ERC1, ELKS, KIAA1081, RAB6IP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ELKS/RAB6-interacting/CAST family member 1 (RAB6-interacting protein 2) (ERC protein 1). | |||||
|
FUT4_MOUSE
|
||||||
| NC score | 0.005660 (rank : 48) | θ value | 8.99809 (rank : 43) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q11127 | Gene names | Fut4, Elft | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-(1,3)-fucosyltransferase (EC 2.4.1.-) (Galactoside 3-L- fucosyltransferase) (Fucosyltransferase 4) (FUCT-IV) (Fuc-TIV). | |||||
|
KIF2C_HUMAN
|
||||||
| NC score | 0.002105 (rank : 49) | θ value | 8.99809 (rank : 45) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99661, Q96C18, Q96HB8, Q9BWV8 | Gene names | KIF2C, KNSL6 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF2C (Mitotic centromere-associated kinesin) (MCAK) (Kinesin-like protein 6). | |||||
|
MSX1_HUMAN
|
||||||
| NC score | 0.001216 (rank : 50) | θ value | 6.88961 (rank : 41) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P28360, Q96NY4 | Gene names | MSX1, HOX7 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein MSX-1 (Msh homeobox 1-like protein) (Hox-7). | |||||
|
ZBT7A_MOUSE
|
||||||
| NC score | 0.000398 (rank : 51) | θ value | 1.81305 (rank : 27) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 795 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O88939, Q9CRJ0 | Gene names | Zbtb7a, Lrf, Zbtb7 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 7A (Leukemia/lymphoma- related factor). | |||||
|
ZBT7A_HUMAN
|
||||||
| NC score | 0.000086 (rank : 52) | θ value | 3.0926 (rank : 31) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 823 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O95365, O00456 | Gene names | ZBTB7A, FBI1, LRF, ZBTB7 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 7A (Leukemia/lymphoma- related factor) (Factor that binds to inducer of short transcripts protein 1) (Factor binding IST protein 1) (FBI-1) (HIV-1 1st-binding protein 1) (TTF-I-interacting peptide 21) (TIP21). | |||||