Please be patient as the page loads
|
PKP1_MOUSE
|
||||||
| SwissProt Accessions | P97350 | Gene names | Pkp1 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-1. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
PKP1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.998023 (rank : 2) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q13835, O00645, Q15152 | Gene names | PKP1 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-1 (Band-6 protein) (B6P). | |||||
|
PKP1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P97350 | Gene names | Pkp1 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-1. | |||||
|
PKP2_HUMAN
|
||||||
| θ value | 8.19079e-94 (rank : 3) | NC score | 0.948328 (rank : 3) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q99959, Q99960 | Gene names | PKP2 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-2. | |||||
|
PKP4_MOUSE
|
||||||
| θ value | 1.7756e-72 (rank : 4) | NC score | 0.903597 (rank : 8) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q68FH0, Q640N0, Q68G56, Q8BK47, Q8BVH1, Q9CRE3 | Gene names | Pkp4, Armrp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plakophilin-4 (Armadillo-related protein). | |||||
|
PKP4_HUMAN
|
||||||
| θ value | 3.34864e-71 (rank : 5) | NC score | 0.902917 (rank : 9) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q99569 | Gene names | PKP4 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-4 (p0071). | |||||
|
CTND2_HUMAN
|
||||||
| θ value | 6.98233e-69 (rank : 6) | NC score | 0.877316 (rank : 13) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9UQB3, O00379, O15390, O43206, O43840, Q13589, Q9UM66, Q9UPM3 | Gene names | CTNND2, NPRAP | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-2 (Delta-catenin) (Neural plakophilin-related ARM-repeat protein) (NPRAP) (Neurojungin) (GT24). | |||||
|
PKP3_HUMAN
|
||||||
| θ value | 9.11921e-69 (rank : 7) | NC score | 0.928069 (rank : 4) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9Y446 | Gene names | PKP3 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-3. | |||||
|
PKP3_MOUSE
|
||||||
| θ value | 2.65329e-68 (rank : 8) | NC score | 0.927583 (rank : 5) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9QY23 | Gene names | Pkp3 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-3. | |||||
|
CTND2_MOUSE
|
||||||
| θ value | 5.00389e-67 (rank : 9) | NC score | 0.888669 (rank : 12) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O35927 | Gene names | Ctnnd2, Catnd2, Nprap | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-2 (Neural plakophilin-related ARM-repeat protein) (NPRAP) (Neurojungin). | |||||
|
ARVC_MOUSE
|
||||||
| θ value | 7.47731e-63 (rank : 10) | NC score | 0.909203 (rank : 6) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P98203, Q6PGJ6, Q8BQ36, Q8BRF2, Q8C3U7, Q924L2, Q924L3, Q924L4, Q924L5 | Gene names | Arvcf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat protein deleted in velo-cardio-facial syndrome homolog. | |||||
|
ARVC_HUMAN
|
||||||
| θ value | 1.84173e-61 (rank : 11) | NC score | 0.905882 (rank : 7) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O00192 | Gene names | ARVCF | |||
|
Domain Architecture |

|
|||||
| Description | Armadillo repeat protein deleted in velo-cardio-facial syndrome. | |||||
|
CTND1_HUMAN
|
||||||
| θ value | 3.47335e-60 (rank : 12) | NC score | 0.900995 (rank : 10) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O60716, O15088, O60713, O60714, O60715, O60935, Q6RBX8, Q9UP71, Q9UP72, Q9UP73 | Gene names | CTNND1, KIAA0384 | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-1 (p120 catenin) (p120(ctn)) (Cadherin-associated Src substrate) (CAS) (p120(cas)). | |||||
|
CTND1_MOUSE
|
||||||
| θ value | 2.48917e-58 (rank : 13) | NC score | 0.900945 (rank : 11) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P30999 | Gene names | Ctnnd1, Catns | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-1 (p120 catenin) (p120(ctn)) (Cadherin-associated Src substrate) (CAS) (p120(cas)). | |||||
|
PLAK_MOUSE
|
||||||
| θ value | 9.29e-05 (rank : 14) | NC score | 0.248380 (rank : 17) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q02257, Q8CGD3 | Gene names | Jup | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junction plakoglobin (Desmoplakin-3) (Desmoplakin III). | |||||
|
PLAK_HUMAN
|
||||||
| θ value | 0.00175202 (rank : 15) | NC score | 0.210375 (rank : 18) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P14923, Q9HCX9 | Gene names | JUP, DP3 | |||
|
Domain Architecture |

|
|||||
| Description | Junction plakoglobin (Desmoplakin-3) (Desmoplakin III). | |||||
|
APC_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 16) | NC score | 0.201831 (rank : 20) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 603 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P25054, Q15162, Q15163, Q93042 | Gene names | APC, DP2.5 | |||
|
Domain Architecture |

|
|||||
| Description | Adenomatous polyposis coli protein (Protein APC). | |||||
|
APC_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 17) | NC score | 0.206677 (rank : 19) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q61315, Q62044 | Gene names | Apc | |||
|
Domain Architecture |

|
|||||
| Description | Adenomatous polyposis coli protein (Protein APC) (mAPC). | |||||
|
CTNB1_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 18) | NC score | 0.265833 (rank : 16) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P35222, Q8NEW9, Q8NI94, Q9H391 | Gene names | CTNNB1, CTNNB | |||
|
Domain Architecture |

|
|||||
| Description | Catenin beta-1 (Beta-catenin). | |||||
|
CTNB1_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 19) | NC score | 0.265888 (rank : 15) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q02248, Q922W1, Q9D335 | Gene names | Ctnnb1, Catnb | |||
|
Domain Architecture |

|
|||||
| Description | Catenin beta-1 (Beta-catenin). | |||||
|
ARMC4_HUMAN
|
||||||
| θ value | 0.279714 (rank : 20) | NC score | 0.319937 (rank : 14) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q5T2S8, Q9H0C0 | Gene names | ARMC4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing protein 4. | |||||
|
ZYGBL_HUMAN
|
||||||
| θ value | 0.47712 (rank : 21) | NC score | 0.051545 (rank : 32) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7Z7L7, O00156, Q5T272, Q5T273 | Gene names | ZYG11BL, C9orf60, ZYG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zyg-11 protein homolog (Zyg-11 homolog B-like). | |||||
|
ZYGBL_MOUSE
|
||||||
| θ value | 0.47712 (rank : 22) | NC score | 0.051665 (rank : 31) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q80ZJ6, Q6PGH5, Q8BG38 | Gene names | Zyg11bl, Zyg | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zyg-11 protein homolog (Zyg-11 homolog B-like). | |||||
|
K1C16_HUMAN
|
||||||
| θ value | 0.813845 (rank : 23) | NC score | 0.006901 (rank : 40) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 440 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P08779, P30654, Q16402, Q9UBG8 | Gene names | KRT16, KRT16A | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type I cytoskeletal 16 (Cytokeratin-16) (CK-16) (Keratin-16) (K16). | |||||
|
IMA3_HUMAN
|
||||||
| θ value | 1.38821 (rank : 24) | NC score | 0.056664 (rank : 29) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O00505, O00191, O43195, Q96AA7 | Gene names | KPNA3 | |||
|
Domain Architecture |
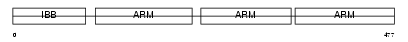
|
|||||
| Description | Importin alpha-3 subunit (Karyopherin alpha-3 subunit) (SRP1-gamma). | |||||
|
LRP3_HUMAN
|
||||||
| θ value | 2.36792 (rank : 25) | NC score | 0.009394 (rank : 38) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O75074 | Gene names | LRP3 | |||
|
Domain Architecture |
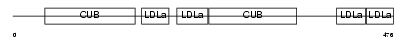
|
|||||
| Description | Low-density lipoprotein receptor-related protein 3 precursor (hLRp105). | |||||
|
SGOL2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 26) | NC score | 0.027437 (rank : 34) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7TSY8, Q811I4, Q8C9C4, Q8CGJ4, Q9CS55 | Gene names | Sgol2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Shugoshin-like 2. | |||||
|
DYR1B_MOUSE
|
||||||
| θ value | 3.0926 (rank : 27) | NC score | -0.000372 (rank : 45) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 799 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Z188 | Gene names | Dyrk1b | |||
|
Domain Architecture |

|
|||||
| Description | Dual specificity tyrosine-phosphorylation-regulated kinase 1B (EC 2.7.12.1). | |||||
|
IMA3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 28) | NC score | 0.054995 (rank : 30) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O35344 | Gene names | Kpna3 | |||
|
Domain Architecture |
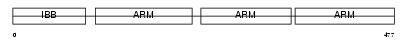
|
|||||
| Description | Importin alpha-3 subunit (Karyopherin alpha-3 subunit) (Importin alpha Q2). | |||||
|
K1C14_HUMAN
|
||||||
| θ value | 5.27518 (rank : 29) | NC score | 0.004730 (rank : 42) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P02533, Q14715, Q53XY3, Q9BUE3, Q9UBN2, Q9UBN3, Q9UCY4 | Gene names | KRT14 | |||
|
Domain Architecture |
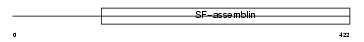
|
|||||
| Description | Keratin, type I cytoskeletal 14 (Cytokeratin-14) (CK-14) (Keratin-14) (K14). | |||||
|
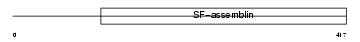
K1C16_MOUSE
|
||||||
| θ value | 5.27518 (rank : 30) | NC score | 0.004933 (rank : 41) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 444 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Z2K1 | Gene names | Krt16, Krt1-16 | |||
|
Domain Architecture |
|
|||||
| Description | Keratin, type I cytoskeletal 16 (Cytokeratin-16) (CK-16) (Keratin-16) (K16). | |||||
|
SPAG6_HUMAN
|
||||||
| θ value | 5.27518 (rank : 31) | NC score | 0.028129 (rank : 33) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O75602, Q5VUX5, Q5VUX6, Q5VUX7, Q6FI74, Q8NHQ6 | Gene names | SPAG6, PF16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-associated antigen 6 (PF16 protein homolog) (Sperm flagellar protein) (Repro-SA-1). | |||||
|
TNR6A_HUMAN
|
||||||
| θ value | 5.27518 (rank : 32) | NC score | 0.010572 (rank : 37) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NDV7, O15408, Q658L5, Q6NVB5, Q8NEZ0, Q8TBT8, Q8TCR0, Q9NV59, Q9P268 | Gene names | TNRC6A, CAGH26, KIAA1460, TNRC6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trinucleotide repeat-containing gene 6A protein (CAG repeat protein 26) (Glycine-tryptophan protein of 182 kDa) (GW182 autoantigen) (Protein GW1) (EMSY interactor protein). | |||||
|
ZF106_MOUSE
|
||||||
| θ value | 5.27518 (rank : 33) | NC score | 0.015874 (rank : 35) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O88466, O55185, O88465, O88467, Q792M1, Q792P4, Q8CDZ8, Q8R3I4, Q9ESU3 | Gene names | Zfp106, H3a, Sh3bp3, Sirm, Znf474 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 106 (Zfp-106) (Zinc finger protein 474) (H3a minor histocompatibility antigen) (Son of insulin receptor mutant). | |||||
|
HECD1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 34) | NC score | 0.003488 (rank : 44) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 371 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9ULT8, Q6P445, Q86VJ1, Q96F34, Q9UFZ7 | Gene names | HECTD1, KIAA1131 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin-protein ligase HECTD1 (HECT domain-containing protein 1) (E3 ligase for inhibin receptor) (EULIR). | |||||
|
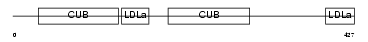
LRP10_MOUSE
|
||||||
| θ value | 6.88961 (rank : 35) | NC score | 0.013029 (rank : 36) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7TQH7, Q7TS95, Q921T0, Q9EPE8 | Gene names | Lrp10, Lrp9 | |||
|
Domain Architecture |
|
|||||
| Description | Low-density lipoprotein receptor-related protein 10 precursor. | |||||
|
RAD_HUMAN
|
||||||
| θ value | 6.88961 (rank : 36) | NC score | 0.004493 (rank : 43) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 224 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P55042, Q96F39 | Gene names | RRAD, RAD | |||
|
Domain Architecture |

|
|||||
| Description | GTP-binding protein RAD (RAS associated with diabetes) (RAD1). | |||||
|
ZF106_HUMAN
|
||||||
| θ value | 8.99809 (rank : 37) | NC score | 0.007968 (rank : 39) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 470 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H2Y7, Q6NSD9, Q6PEK1, Q86T43, Q86T45, Q86T50, Q86T58, Q86TA9, Q96M37, Q9H7B8 | Gene names | ZFP106, ZNF474 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 106 homolog (Zfp-106) (Zinc finger protein 474). | |||||
|
GDS1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 38) | NC score | 0.118864 (rank : 21) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P52306, Q9NYM2, Q9NZA8 | Gene names | RAP1GDS1 | |||
|
Domain Architecture |

|
|||||
| Description | Rap1 GTPase-GDP dissociation stimulator 1 (SMG P21 stimulatory GDP/GTP exchange protein) (SMG GDS protein) (Exchange factor smgGDS). | |||||
|
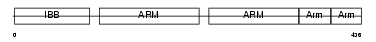
IMA2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 39) | NC score | 0.059903 (rank : 25) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P52292, Q9BRU5 | Gene names | KPNA2, RCH1, SRP1 | |||
|
Domain Architecture |
|
|||||
| Description | Importin alpha-2 subunit (Karyopherin alpha-2 subunit) (SRP1-alpha) (RAG cohort protein 1). | |||||
|
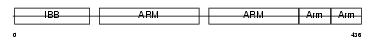
IMA2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 40) | NC score | 0.059466 (rank : 26) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P52293, Q64292 | Gene names | Kpna2, Rch1 | |||
|
Domain Architecture |
|
|||||
| Description | Importin alpha-2 subunit (Karyopherin alpha-2 subunit) (SRP1-alpha) (RAG cohort protein 1) (Pendulin) (Pore targeting complex 58 kDa subunit) (PTAC58) (Importin alpha P1). | |||||
|
IMA4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 41) | NC score | 0.056903 (rank : 28) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O00629, O00190 | Gene names | KPNA4, QIP1 | |||
|
Domain Architecture |
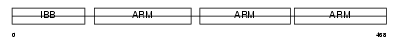
|
|||||
| Description | Importin alpha-4 subunit (Karyopherin alpha-4 subunit) (Qip1 protein). | |||||
|
IMA4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 42) | NC score | 0.056968 (rank : 27) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O35343 | Gene names | Kpna4 | |||
|
Domain Architecture |
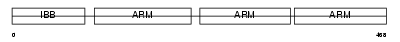
|
|||||
| Description | Importin alpha-4 subunit (Karyopherin alpha-4 subunit) (Importin alpha Q1). | |||||
|
IMA5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 43) | NC score | 0.065666 (rank : 24) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O15131 | Gene names | KPNA5 | |||
|
Domain Architecture |
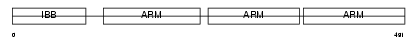
|
|||||
| Description | Importin alpha-6 subunit (Karyopherin alpha-5 subunit). | |||||
|
IMA7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 44) | NC score | 0.067635 (rank : 23) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O60684 | Gene names | KPNA6, IPOA7 | |||
|
Domain Architecture |
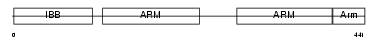
|
|||||
| Description | Importin alpha-7 subunit (Karyopherin alpha-6). | |||||
|
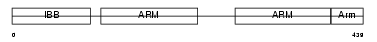
IMA7_MOUSE
|
||||||
| θ value | θ > 10 (rank : 45) | NC score | 0.070293 (rank : 22) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O35345, Q9CVP9 | Gene names | Kpna6, Kpna5 | |||
|
Domain Architecture |
|
|||||
| Description | Importin alpha-7 subunit (Karyopherin alpha-6 subunit) (Importin alpha S2). | |||||
|
PKP1_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P97350 | Gene names | Pkp1 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-1. | |||||
|
PKP1_HUMAN
|
||||||
| NC score | 0.998023 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q13835, O00645, Q15152 | Gene names | PKP1 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-1 (Band-6 protein) (B6P). | |||||
|
PKP2_HUMAN
|
||||||
| NC score | 0.948328 (rank : 3) | θ value | 8.19079e-94 (rank : 3) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q99959, Q99960 | Gene names | PKP2 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-2. | |||||
|
PKP3_HUMAN
|
||||||
| NC score | 0.928069 (rank : 4) | θ value | 9.11921e-69 (rank : 7) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9Y446 | Gene names | PKP3 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-3. | |||||
|
PKP3_MOUSE
|
||||||
| NC score | 0.927583 (rank : 5) | θ value | 2.65329e-68 (rank : 8) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9QY23 | Gene names | Pkp3 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-3. | |||||
|
ARVC_MOUSE
|
||||||
| NC score | 0.909203 (rank : 6) | θ value | 7.47731e-63 (rank : 10) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P98203, Q6PGJ6, Q8BQ36, Q8BRF2, Q8C3U7, Q924L2, Q924L3, Q924L4, Q924L5 | Gene names | Arvcf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat protein deleted in velo-cardio-facial syndrome homolog. | |||||
|
ARVC_HUMAN
|
||||||
| NC score | 0.905882 (rank : 7) | θ value | 1.84173e-61 (rank : 11) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O00192 | Gene names | ARVCF | |||
|
Domain Architecture |

|
|||||
| Description | Armadillo repeat protein deleted in velo-cardio-facial syndrome. | |||||
|
PKP4_MOUSE
|
||||||
| NC score | 0.903597 (rank : 8) | θ value | 1.7756e-72 (rank : 4) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q68FH0, Q640N0, Q68G56, Q8BK47, Q8BVH1, Q9CRE3 | Gene names | Pkp4, Armrp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plakophilin-4 (Armadillo-related protein). | |||||
|
PKP4_HUMAN
|
||||||
| NC score | 0.902917 (rank : 9) | θ value | 3.34864e-71 (rank : 5) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q99569 | Gene names | PKP4 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-4 (p0071). | |||||
|
CTND1_HUMAN
|
||||||
| NC score | 0.900995 (rank : 10) | θ value | 3.47335e-60 (rank : 12) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O60716, O15088, O60713, O60714, O60715, O60935, Q6RBX8, Q9UP71, Q9UP72, Q9UP73 | Gene names | CTNND1, KIAA0384 | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-1 (p120 catenin) (p120(ctn)) (Cadherin-associated Src substrate) (CAS) (p120(cas)). | |||||
|
CTND1_MOUSE
|
||||||
| NC score | 0.900945 (rank : 11) | θ value | 2.48917e-58 (rank : 13) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P30999 | Gene names | Ctnnd1, Catns | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-1 (p120 catenin) (p120(ctn)) (Cadherin-associated Src substrate) (CAS) (p120(cas)). | |||||
|
CTND2_MOUSE
|
||||||
| NC score | 0.888669 (rank : 12) | θ value | 5.00389e-67 (rank : 9) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O35927 | Gene names | Ctnnd2, Catnd2, Nprap | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-2 (Neural plakophilin-related ARM-repeat protein) (NPRAP) (Neurojungin). | |||||
|
CTND2_HUMAN
|
||||||
| NC score | 0.877316 (rank : 13) | θ value | 6.98233e-69 (rank : 6) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9UQB3, O00379, O15390, O43206, O43840, Q13589, Q9UM66, Q9UPM3 | Gene names | CTNND2, NPRAP | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-2 (Delta-catenin) (Neural plakophilin-related ARM-repeat protein) (NPRAP) (Neurojungin) (GT24). | |||||
|
ARMC4_HUMAN
|
||||||
| NC score | 0.319937 (rank : 14) | θ value | 0.279714 (rank : 20) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q5T2S8, Q9H0C0 | Gene names | ARMC4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing protein 4. | |||||
|
CTNB1_MOUSE
|
||||||
| NC score | 0.265888 (rank : 15) | θ value | 0.0330416 (rank : 19) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q02248, Q922W1, Q9D335 | Gene names | Ctnnb1, Catnb | |||
|
Domain Architecture |

|
|||||
| Description | Catenin beta-1 (Beta-catenin). | |||||
|
CTNB1_HUMAN
|
||||||
| NC score | 0.265833 (rank : 16) | θ value | 0.0330416 (rank : 18) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P35222, Q8NEW9, Q8NI94, Q9H391 | Gene names | CTNNB1, CTNNB | |||
|
Domain Architecture |

|
|||||
| Description | Catenin beta-1 (Beta-catenin). | |||||
|
PLAK_MOUSE
|
||||||
| NC score | 0.248380 (rank : 17) | θ value | 9.29e-05 (rank : 14) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q02257, Q8CGD3 | Gene names | Jup | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junction plakoglobin (Desmoplakin-3) (Desmoplakin III). | |||||
|
PLAK_HUMAN
|
||||||
| NC score | 0.210375 (rank : 18) | θ value | 0.00175202 (rank : 15) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P14923, Q9HCX9 | Gene names | JUP, DP3 | |||
|
Domain Architecture |

|
|||||
| Description | Junction plakoglobin (Desmoplakin-3) (Desmoplakin III). | |||||
|
APC_MOUSE
|
||||||
| NC score | 0.206677 (rank : 19) | θ value | 0.0193708 (rank : 17) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q61315, Q62044 | Gene names | Apc | |||
|
Domain Architecture |

|
|||||
| Description | Adenomatous polyposis coli protein (Protein APC) (mAPC). | |||||
|
APC_HUMAN
|
||||||
| NC score | 0.201831 (rank : 20) | θ value | 0.0193708 (rank : 16) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 603 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P25054, Q15162, Q15163, Q93042 | Gene names | APC, DP2.5 | |||
|
Domain Architecture |

|
|||||
| Description | Adenomatous polyposis coli protein (Protein APC). | |||||
|
GDS1_HUMAN
|
||||||
| NC score | 0.118864 (rank : 21) | θ value | θ > 10 (rank : 38) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P52306, Q9NYM2, Q9NZA8 | Gene names | RAP1GDS1 | |||
|
Domain Architecture |

|
|||||
| Description | Rap1 GTPase-GDP dissociation stimulator 1 (SMG P21 stimulatory GDP/GTP exchange protein) (SMG GDS protein) (Exchange factor smgGDS). | |||||
|
IMA7_MOUSE
|
||||||
| NC score | 0.070293 (rank : 22) | θ value | θ > 10 (rank : 45) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O35345, Q9CVP9 | Gene names | Kpna6, Kpna5 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-7 subunit (Karyopherin alpha-6 subunit) (Importin alpha S2). | |||||
|
IMA7_HUMAN
|
||||||
| NC score | 0.067635 (rank : 23) | θ value | θ > 10 (rank : 44) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O60684 | Gene names | KPNA6, IPOA7 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-7 subunit (Karyopherin alpha-6). | |||||
|
IMA5_HUMAN
|
||||||
| NC score | 0.065666 (rank : 24) | θ value | θ > 10 (rank : 43) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O15131 | Gene names | KPNA5 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-6 subunit (Karyopherin alpha-5 subunit). | |||||
|
IMA2_HUMAN
|
||||||
| NC score | 0.059903 (rank : 25) | θ value | θ > 10 (rank : 39) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P52292, Q9BRU5 | Gene names | KPNA2, RCH1, SRP1 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-2 subunit (Karyopherin alpha-2 subunit) (SRP1-alpha) (RAG cohort protein 1). | |||||
|
IMA2_MOUSE
|
||||||
| NC score | 0.059466 (rank : 26) | θ value | θ > 10 (rank : 40) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P52293, Q64292 | Gene names | Kpna2, Rch1 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-2 subunit (Karyopherin alpha-2 subunit) (SRP1-alpha) (RAG cohort protein 1) (Pendulin) (Pore targeting complex 58 kDa subunit) (PTAC58) (Importin alpha P1). | |||||
|
IMA4_MOUSE
|
||||||
| NC score | 0.056968 (rank : 27) | θ value | θ > 10 (rank : 42) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O35343 | Gene names | Kpna4 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-4 subunit (Karyopherin alpha-4 subunit) (Importin alpha Q1). | |||||
|
IMA4_HUMAN
|
||||||
| NC score | 0.056903 (rank : 28) | θ value | θ > 10 (rank : 41) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O00629, O00190 | Gene names | KPNA4, QIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-4 subunit (Karyopherin alpha-4 subunit) (Qip1 protein). | |||||
|
IMA3_HUMAN
|
||||||
| NC score | 0.056664 (rank : 29) | θ value | 1.38821 (rank : 24) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O00505, O00191, O43195, Q96AA7 | Gene names | KPNA3 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-3 subunit (Karyopherin alpha-3 subunit) (SRP1-gamma). | |||||
|
IMA3_MOUSE
|
||||||
| NC score | 0.054995 (rank : 30) | θ value | 4.03905 (rank : 28) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O35344 | Gene names | Kpna3 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-3 subunit (Karyopherin alpha-3 subunit) (Importin alpha Q2). | |||||
|
ZYGBL_MOUSE
|
||||||
| NC score | 0.051665 (rank : 31) | θ value | 0.47712 (rank : 22) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q80ZJ6, Q6PGH5, Q8BG38 | Gene names | Zyg11bl, Zyg | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zyg-11 protein homolog (Zyg-11 homolog B-like). | |||||
|
ZYGBL_HUMAN
|
||||||
| NC score | 0.051545 (rank : 32) | θ value | 0.47712 (rank : 21) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7Z7L7, O00156, Q5T272, Q5T273 | Gene names | ZYG11BL, C9orf60, ZYG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zyg-11 protein homolog (Zyg-11 homolog B-like). | |||||
|
SPAG6_HUMAN
|
||||||
| NC score | 0.028129 (rank : 33) | θ value | 5.27518 (rank : 31) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O75602, Q5VUX5, Q5VUX6, Q5VUX7, Q6FI74, Q8NHQ6 | Gene names | SPAG6, PF16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-associated antigen 6 (PF16 protein homolog) (Sperm flagellar protein) (Repro-SA-1). | |||||
|
SGOL2_MOUSE
|
||||||
| NC score | 0.027437 (rank : 34) | θ value | 2.36792 (rank : 26) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7TSY8, Q811I4, Q8C9C4, Q8CGJ4, Q9CS55 | Gene names | Sgol2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Shugoshin-like 2. | |||||
|
ZF106_MOUSE
|
||||||
| NC score | 0.015874 (rank : 35) | θ value | 5.27518 (rank : 33) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O88466, O55185, O88465, O88467, Q792M1, Q792P4, Q8CDZ8, Q8R3I4, Q9ESU3 | Gene names | Zfp106, H3a, Sh3bp3, Sirm, Znf474 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 106 (Zfp-106) (Zinc finger protein 474) (H3a minor histocompatibility antigen) (Son of insulin receptor mutant). | |||||
|
LRP10_MOUSE
|
||||||
| NC score | 0.013029 (rank : 36) | θ value | 6.88961 (rank : 35) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7TQH7, Q7TS95, Q921T0, Q9EPE8 | Gene names | Lrp10, Lrp9 | |||
|
Domain Architecture |

|
|||||
| Description | Low-density lipoprotein receptor-related protein 10 precursor. | |||||
|
TNR6A_HUMAN
|
||||||
| NC score | 0.010572 (rank : 37) | θ value | 5.27518 (rank : 32) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NDV7, O15408, Q658L5, Q6NVB5, Q8NEZ0, Q8TBT8, Q8TCR0, Q9NV59, Q9P268 | Gene names | TNRC6A, CAGH26, KIAA1460, TNRC6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trinucleotide repeat-containing gene 6A protein (CAG repeat protein 26) (Glycine-tryptophan protein of 182 kDa) (GW182 autoantigen) (Protein GW1) (EMSY interactor protein). | |||||
|
LRP3_HUMAN
|
||||||
| NC score | 0.009394 (rank : 38) | θ value | 2.36792 (rank : 25) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O75074 | Gene names | LRP3 | |||
|
Domain Architecture |

|
|||||
| Description | Low-density lipoprotein receptor-related protein 3 precursor (hLRp105). | |||||
|
ZF106_HUMAN
|
||||||
| NC score | 0.007968 (rank : 39) | θ value | 8.99809 (rank : 37) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 470 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H2Y7, Q6NSD9, Q6PEK1, Q86T43, Q86T45, Q86T50, Q86T58, Q86TA9, Q96M37, Q9H7B8 | Gene names | ZFP106, ZNF474 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 106 homolog (Zfp-106) (Zinc finger protein 474). | |||||
|
K1C16_HUMAN
|
||||||
| NC score | 0.006901 (rank : 40) | θ value | 0.813845 (rank : 23) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 440 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P08779, P30654, Q16402, Q9UBG8 | Gene names | KRT16, KRT16A | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type I cytoskeletal 16 (Cytokeratin-16) (CK-16) (Keratin-16) (K16). | |||||
|
K1C16_MOUSE
|
||||||
| NC score | 0.004933 (rank : 41) | θ value | 5.27518 (rank : 30) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 444 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Z2K1 | Gene names | Krt16, Krt1-16 | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type I cytoskeletal 16 (Cytokeratin-16) (CK-16) (Keratin-16) (K16). | |||||
|
K1C14_HUMAN
|
||||||
| NC score | 0.004730 (rank : 42) | θ value | 5.27518 (rank : 29) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P02533, Q14715, Q53XY3, Q9BUE3, Q9UBN2, Q9UBN3, Q9UCY4 | Gene names | KRT14 | |||
|
Domain Architecture |

|
|||||
| Description | Keratin, type I cytoskeletal 14 (Cytokeratin-14) (CK-14) (Keratin-14) (K14). | |||||
|
RAD_HUMAN
|
||||||
| NC score | 0.004493 (rank : 43) | θ value | 6.88961 (rank : 36) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 224 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P55042, Q96F39 | Gene names | RRAD, RAD | |||
|
Domain Architecture |

|
|||||
| Description | GTP-binding protein RAD (RAS associated with diabetes) (RAD1). | |||||
|
HECD1_HUMAN
|
||||||
| NC score | 0.003488 (rank : 44) | θ value | 6.88961 (rank : 34) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 371 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9ULT8, Q6P445, Q86VJ1, Q96F34, Q9UFZ7 | Gene names | HECTD1, KIAA1131 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E3 ubiquitin-protein ligase HECTD1 (HECT domain-containing protein 1) (E3 ligase for inhibin receptor) (EULIR). | |||||
|
DYR1B_MOUSE
|
||||||
| NC score | -0.000372 (rank : 45) | θ value | 3.0926 (rank : 27) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 799 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Z188 | Gene names | Dyrk1b | |||
|
Domain Architecture |

|
|||||
| Description | Dual specificity tyrosine-phosphorylation-regulated kinase 1B (EC 2.7.12.1). | |||||