Please be patient as the page loads
|
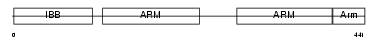
IMA7_HUMAN
|
||||||
| SwissProt Accessions | O60684 | Gene names | KPNA6, IPOA7 | |||
|
Domain Architecture |
|
|||||
| Description | Importin alpha-7 subunit (Karyopherin alpha-6). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
IMA1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.993379 (rank : 4) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P52294, Q9BQ56 | Gene names | KPNA1, RCH2 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-1 subunit (Karyopherin alpha-1 subunit) (SRP1-beta) (RAG cohort protein 2) (Nucleoprotein interactor 1) (NPI-1). | |||||
|
IMA1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.993596 (rank : 3) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q60960, Q3TF32 | Gene names | Kpna1, Rch2 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-1 subunit (Karyopherin alpha-1 subunit) (SRP1-beta) (RAG cohort protein 2) (Nucleoprotein interactor 1) (Importin alpha S1). | |||||
|
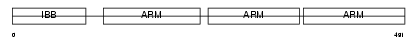
IMA5_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.992762 (rank : 5) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O15131 | Gene names | KPNA5 | |||
|
Domain Architecture |
|
|||||
| Description | Importin alpha-6 subunit (Karyopherin alpha-5 subunit). | |||||
|
IMA7_HUMAN
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | O60684 | Gene names | KPNA6, IPOA7 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-7 subunit (Karyopherin alpha-6). | |||||
|
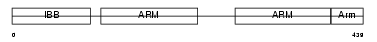
IMA7_MOUSE
|
||||||
| θ value | 0 (rank : 5) | NC score | 0.998521 (rank : 2) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O35345, Q9CVP9 | Gene names | Kpna6, Kpna5 | |||
|
Domain Architecture |
|
|||||
| Description | Importin alpha-7 subunit (Karyopherin alpha-6 subunit) (Importin alpha S2). | |||||
|
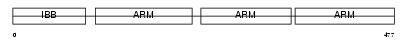
IMA3_MOUSE
|
||||||
| θ value | 6.85399e-133 (rank : 6) | NC score | 0.974326 (rank : 6) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O35344 | Gene names | Kpna3 | |||
|
Domain Architecture |
|
|||||
| Description | Importin alpha-3 subunit (Karyopherin alpha-3 subunit) (Importin alpha Q2). | |||||
|
IMA3_HUMAN
|
||||||
| θ value | 1.29261e-131 (rank : 7) | NC score | 0.973980 (rank : 7) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O00505, O00191, O43195, Q96AA7 | Gene names | KPNA3 | |||
|
Domain Architecture |
|
|||||
| Description | Importin alpha-3 subunit (Karyopherin alpha-3 subunit) (SRP1-gamma). | |||||
|
IMA4_HUMAN
|
||||||
| θ value | 7.09276e-130 (rank : 8) | NC score | 0.973576 (rank : 8) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O00629, O00190 | Gene names | KPNA4, QIP1 | |||
|
Domain Architecture |
|
|||||
| Description | Importin alpha-4 subunit (Karyopherin alpha-4 subunit) (Qip1 protein). | |||||
|
IMA4_MOUSE
|
||||||
| θ value | 1.74701e-128 (rank : 9) | NC score | 0.973148 (rank : 9) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O35343 | Gene names | Kpna4 | |||
|
Domain Architecture |
|
|||||
| Description | Importin alpha-4 subunit (Karyopherin alpha-4 subunit) (Importin alpha Q1). | |||||
|
IMA2_MOUSE
|
||||||
| θ value | 1.99884e-124 (rank : 10) | NC score | 0.971843 (rank : 10) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P52293, Q64292 | Gene names | Kpna2, Rch1 | |||
|
Domain Architecture |
|
|||||
| Description | Importin alpha-2 subunit (Karyopherin alpha-2 subunit) (SRP1-alpha) (RAG cohort protein 1) (Pendulin) (Pore targeting complex 58 kDa subunit) (PTAC58) (Importin alpha P1). | |||||
|
IMA2_HUMAN
|
||||||
| θ value | 3.76964e-123 (rank : 11) | NC score | 0.969051 (rank : 11) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P52292, Q9BRU5 | Gene names | KPNA2, RCH1, SRP1 | |||
|
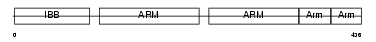
Domain Architecture |
|
|||||
| Description | Importin alpha-2 subunit (Karyopherin alpha-2 subunit) (SRP1-alpha) (RAG cohort protein 1). | |||||
|
SPAG6_HUMAN
|
||||||
| θ value | 1.09485e-13 (rank : 12) | NC score | 0.625269 (rank : 12) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O75602, Q5VUX5, Q5VUX6, Q5VUX7, Q6FI74, Q8NHQ6 | Gene names | SPAG6, PF16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-associated antigen 6 (PF16 protein homolog) (Sperm flagellar protein) (Repro-SA-1). | |||||
|
SPAG6_MOUSE
|
||||||
| θ value | 5.43371e-13 (rank : 13) | NC score | 0.624818 (rank : 13) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9JLI7, Q8K461 | Gene names | Spag6, Pf16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-associated antigen 6 (PF16 protein homolog) (Axoneme central apparatus protein). | |||||
|
TRIPC_HUMAN
|
||||||
| θ value | 0.00020696 (rank : 14) | NC score | 0.114743 (rank : 31) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 319 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14669, Q15644 | Gene names | TRIP12, KIAA0045 | |||
|
Domain Architecture |

|
|||||
| Description | Thyroid receptor-interacting protein 12 (TRIP12). | |||||
|
PKP4_HUMAN
|
||||||
| θ value | 0.00035302 (rank : 15) | NC score | 0.183361 (rank : 17) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q99569 | Gene names | PKP4 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-4 (p0071). | |||||
|
PKP4_MOUSE
|
||||||
| θ value | 0.00035302 (rank : 16) | NC score | 0.184223 (rank : 16) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q68FH0, Q640N0, Q68G56, Q8BK47, Q8BVH1, Q9CRE3 | Gene names | Pkp4, Armrp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plakophilin-4 (Armadillo-related protein). | |||||
|
CTND2_HUMAN
|
||||||
| θ value | 0.00102713 (rank : 17) | NC score | 0.149568 (rank : 19) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UQB3, O00379, O15390, O43206, O43840, Q13589, Q9UM66, Q9UPM3 | Gene names | CTNND2, NPRAP | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-2 (Delta-catenin) (Neural plakophilin-related ARM-repeat protein) (NPRAP) (Neurojungin) (GT24). | |||||
|
CTND2_MOUSE
|
||||||
| θ value | 0.00134147 (rank : 18) | NC score | 0.150889 (rank : 18) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O35927 | Gene names | Ctnnd2, Catnd2, Nprap | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-2 (Neural plakophilin-related ARM-repeat protein) (NPRAP) (Neurojungin). | |||||
|
IMB1_HUMAN
|
||||||
| θ value | 0.00665767 (rank : 19) | NC score | 0.120444 (rank : 30) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q14974, Q14637, Q96J27 | Gene names | KPNB1, NTF97 | |||
|
Domain Architecture |

|
|||||
| Description | Importin beta-1 subunit (Karyopherin beta-1 subunit) (Nuclear factor P97) (Importin 90). | |||||
|
IMB1_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 20) | NC score | 0.141444 (rank : 20) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P70168, Q62117 | Gene names | Kpnb1, Impnb | |||
|
Domain Architecture |

|
|||||
| Description | Importin beta-1 subunit (Karyopherin beta-1 subunit) (Nuclear factor P97) (Pore targeting complex 97 kDa subunit) (PTAC97) (SCG). | |||||
|
TNPO1_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 21) | NC score | 0.125689 (rank : 27) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q92973, Q92957, Q92975 | Gene names | TNPO1, KPNB2, MIP1, TRN | |||
|
Domain Architecture |

|
|||||
| Description | Transportin-1 (Importin beta-2) (Karyopherin beta-2) (M9 region interaction protein) (MIP). | |||||
|
TNPO1_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 22) | NC score | 0.121931 (rank : 29) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8BFY9, Q8C0S3, Q8K2E7 | Gene names | Tnpo1, Kpnb2 | |||
|
Domain Architecture |

|
|||||
| Description | Transportin-1 (Importin beta-2) (Karyopherin beta-2). | |||||
|
ARMC4_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 23) | NC score | 0.246506 (rank : 15) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q5T2S8, Q9H0C0 | Gene names | ARMC4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing protein 4. | |||||
|
PLAK_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 24) | NC score | 0.131308 (rank : 22) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q02257, Q8CGD3 | Gene names | Jup | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junction plakoglobin (Desmoplakin-3) (Desmoplakin III). | |||||
|
TNPO2_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 25) | NC score | 0.109475 (rank : 34) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O14787, O14655 | Gene names | TNPO2 | |||
|
Domain Architecture |

|
|||||
| Description | Transportin-2 (Karyopherin beta-2b). | |||||
|
TNPO2_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 26) | NC score | 0.110648 (rank : 33) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q99LG2, Q8C0U9 | Gene names | Tnpo2 | |||
|
Domain Architecture |

|
|||||
| Description | Transportin-2 (Karyopherin beta-2b). | |||||
|
ARMX3_HUMAN
|
||||||
| θ value | 0.125558 (rank : 27) | NC score | 0.059778 (rank : 44) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UH62, Q53HC6, Q7LCF5, Q9NPE4 | Gene names | ARMCX3, ALEX3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing X-linked protein 3 (Protein ALEX3) (ARM protein lost in epithelial cancers on chromosome X 3). | |||||
|
ARMX3_MOUSE
|
||||||
| θ value | 0.125558 (rank : 28) | NC score | 0.060331 (rank : 43) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8BHS6, Q91VP8, Q9DC32 | Gene names | Armcx3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing X-linked protein 3. | |||||
|
ARVC_HUMAN
|
||||||
| θ value | 0.163984 (rank : 29) | NC score | 0.130150 (rank : 25) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O00192 | Gene names | ARVCF | |||
|
Domain Architecture |

|
|||||
| Description | Armadillo repeat protein deleted in velo-cardio-facial syndrome. | |||||
|
ARVC_MOUSE
|
||||||
| θ value | 0.163984 (rank : 30) | NC score | 0.129360 (rank : 26) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P98203, Q6PGJ6, Q8BQ36, Q8BRF2, Q8C3U7, Q924L2, Q924L3, Q924L4, Q924L5 | Gene names | Arvcf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat protein deleted in velo-cardio-facial syndrome homolog. | |||||
|
CTND1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 31) | NC score | 0.131875 (rank : 21) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P30999 | Gene names | Ctnnd1, Catns | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-1 (p120 catenin) (p120(ctn)) (Cadherin-associated Src substrate) (CAS) (p120(cas)). | |||||
|
RASA2_HUMAN
|
||||||
| θ value | 0.47712 (rank : 32) | NC score | 0.019712 (rank : 53) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15283, O00695, Q15284, Q92594, Q99577, Q9UEQ2 | Gene names | RASA2, GAP1M, RASGAP | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 2 (GAP1m). | |||||
|
RELN_MOUSE
|
||||||
| θ value | 0.47712 (rank : 33) | NC score | 0.045891 (rank : 50) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q60841, Q9CUA6 | Gene names | Reln, Rl | |||
|
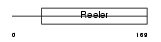
Domain Architecture |
|
|||||
| Description | Reelin precursor (EC 3.4.21.-) (Reeler protein). | |||||
|
CF081_HUMAN
|
||||||
| θ value | 0.62314 (rank : 34) | NC score | 0.058132 (rank : 45) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q5T9G4, Q8NEB2, Q96LL8 | Gene names | C6orf81 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf81. | |||||
|
CTND1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 35) | NC score | 0.123145 (rank : 28) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O60716, O15088, O60713, O60714, O60715, O60935, Q6RBX8, Q9UP71, Q9UP72, Q9UP73 | Gene names | CTNND1, KIAA0384 | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-1 (p120 catenin) (p120(ctn)) (Cadherin-associated Src substrate) (CAS) (p120(cas)). | |||||
|
GDS1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 36) | NC score | 0.269617 (rank : 14) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P52306, Q9NYM2, Q9NZA8 | Gene names | RAP1GDS1 | |||
|
Domain Architecture |

|
|||||
| Description | Rap1 GTPase-GDP dissociation stimulator 1 (SMG P21 stimulatory GDP/GTP exchange protein) (SMG GDS protein) (Exchange factor smgGDS). | |||||
|
UN45A_HUMAN
|
||||||
| θ value | 1.38821 (rank : 37) | NC score | 0.051749 (rank : 46) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9H3U1, Q7L3Y6, Q9H3U8, Q9NSE8, Q9NSE9 | Gene names | UNC45A, SMAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UNC45 homolog A (UNC-45A) (Smooth muscle cell-associated protein 1) (SMAP-1). | |||||
|
ZFY16_HUMAN
|
||||||
| θ value | 1.81305 (rank : 38) | NC score | 0.033264 (rank : 52) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7Z3T8, O15023, Q7LAU7, Q86T69, Q8N5L3, Q8NEK3 | Gene names | ZFYVE16, KIAA0305 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger FYVE domain-containing protein 16 (Endofin) (Endosome- associated FYVE domain protein). | |||||
|
IMB3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 39) | NC score | 0.046799 (rank : 49) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O00410, O15257 | Gene names | RANBP5, KPNB3 | |||
|
Domain Architecture |

|
|||||
| Description | Importin beta-3 (Karyopherin beta-3) (Ran-binding protein 5) (RanBP5). | |||||
|
IMB3_MOUSE
|
||||||
| θ value | 3.0926 (rank : 40) | NC score | 0.047058 (rank : 48) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BKC5, Q3TEG2, Q7TQC6, Q9EQ30 | Gene names | Ranbp5, Kpnb3 | |||
|
Domain Architecture |

|
|||||
| Description | Importin beta-3 (Karyopherin beta-3) (Ran-binding protein 5) (RanBP5). | |||||
|
RELN_HUMAN
|
||||||
| θ value | 3.0926 (rank : 41) | NC score | 0.033650 (rank : 51) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P78509, Q86UJ0, Q86UJ8, Q8NDV0, Q9UDQ2 | Gene names | RELN | |||
|
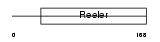
Domain Architecture |
|
|||||
| Description | Reelin precursor (EC 3.4.21.-). | |||||
|
GOGB1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 42) | NC score | 0.002519 (rank : 61) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 2297 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14789, Q14398 | Gene names | GOLGB1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily B member 1 (Giantin) (Macrogolgin) (372 kDa Golgi complex-associated protein) (GCP372). | |||||
|
RL35_HUMAN
|
||||||
| θ value | 5.27518 (rank : 43) | NC score | 0.111926 (rank : 32) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P42766, Q4VBY5, Q5JTN5, Q6IBC7, Q96QJ7, Q9BYF4 | Gene names | RPL35 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 60S ribosomal protein L35. | |||||
|
UN45B_MOUSE
|
||||||
| θ value | 5.27518 (rank : 44) | NC score | 0.015549 (rank : 54) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8CGY6, Q5XG72, Q8BHC5, Q8BWK3 | Gene names | Unc45b, Cmya4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UNC45 homolog B (UNC-45B). | |||||
|
BRCA1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 45) | NC score | 0.014322 (rank : 56) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P38398 | Gene names | BRCA1, RNF53 | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer type 1 susceptibility protein (RING finger protein 53). | |||||
|
CLAP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 46) | NC score | 0.014537 (rank : 55) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7Z460, O75118, Q8N5B8, Q9BQT5 | Gene names | CLASP1, KIAA0622, MAST1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CLIP-associating protein 1 (Cytoplasmic linker-associated protein 1) (Multiple asters homolog 1). | |||||
|
HD_HUMAN
|
||||||
| θ value | 8.99809 (rank : 47) | NC score | 0.009237 (rank : 58) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P42858, Q9UQB7 | Gene names | HD, IT15 | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin (Huntington disease protein) (HD protein). | |||||
|
HD_MOUSE
|
||||||
| θ value | 8.99809 (rank : 48) | NC score | 0.009483 (rank : 57) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P42859 | Gene names | Hd, Hdh | |||
|
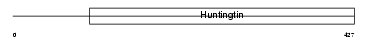
Domain Architecture |
|
|||||
| Description | Huntingtin (Huntington disease protein homolog) (HD protein). | |||||
|
NVL_MOUSE
|
||||||
| θ value | 8.99809 (rank : 49) | NC score | 0.005301 (rank : 60) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9DBY8, Q3USC4, Q8BW27, Q8K2B5 | Gene names | Nvl | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear valosin-containing protein-like (Nuclear VCP-like protein) (NVLp). | |||||
|
PER1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 50) | NC score | 0.005849 (rank : 59) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O15534 | Gene names | PER1, KIAA0482, PER, RIGUI | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (hPER). | |||||
|
ARMX1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 51) | NC score | 0.050874 (rank : 47) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9P291, Q53HK2, Q9H2Q0 | Gene names | ARMCX1, ALEX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing X-linked protein 1 (Protein ALEX1) (ARM protein lost in epithelial cancers on chromosome X 1). | |||||
|
CTNB1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 52) | NC score | 0.130360 (rank : 24) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P35222, Q8NEW9, Q8NI94, Q9H391 | Gene names | CTNNB1, CTNNB | |||
|
Domain Architecture |

|
|||||
| Description | Catenin beta-1 (Beta-catenin). | |||||
|
CTNB1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 53) | NC score | 0.130395 (rank : 23) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q02248, Q922W1, Q9D335 | Gene names | Ctnnb1, Catnb | |||
|
Domain Architecture |

|
|||||
| Description | Catenin beta-1 (Beta-catenin). | |||||
|
PKP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 54) | NC score | 0.070488 (rank : 36) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q13835, O00645, Q15152 | Gene names | PKP1 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-1 (Band-6 protein) (B6P). | |||||
|
PKP1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 55) | NC score | 0.067635 (rank : 38) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P97350 | Gene names | Pkp1 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-1. | |||||
|
PKP2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 56) | NC score | 0.063509 (rank : 40) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q99959, Q99960 | Gene names | PKP2 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-2. | |||||
|
PKP3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 57) | NC score | 0.068383 (rank : 37) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Y446 | Gene names | PKP3 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-3. | |||||
|
PKP3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 58) | NC score | 0.065787 (rank : 39) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9QY23 | Gene names | Pkp3 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-3. | |||||
|
PLAK_HUMAN
|
||||||
| θ value | θ > 10 (rank : 59) | NC score | 0.092803 (rank : 35) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P14923, Q9HCX9 | Gene names | JUP, DP3 | |||
|
Domain Architecture |

|
|||||
| Description | Junction plakoglobin (Desmoplakin-3) (Desmoplakin III). | |||||
|
SARM1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 60) | NC score | 0.060732 (rank : 42) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6SZW1, O60277, Q7LGG3, Q9NXY5 | Gene names | SARM1, KIAA0524, SARM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sterile alpha and TIR motif-containing protein 1 (Sterile alpha and Armadillo repeat protein) (Tir-1 homolog). | |||||
|
SARM1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 61) | NC score | 0.061597 (rank : 41) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6PDS3, Q5SYG5, Q5SYG6, Q6A054, Q6SZW0, Q8BRI9 | Gene names | Sarm1, Kiaa0524 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sterile alpha and TIR motif-containing protein 1 (Tir-1 homolog). | |||||
|
IMA7_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | O60684 | Gene names | KPNA6, IPOA7 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-7 subunit (Karyopherin alpha-6). | |||||
|
IMA7_MOUSE
|
||||||
| NC score | 0.998521 (rank : 2) | θ value | 0 (rank : 5) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O35345, Q9CVP9 | Gene names | Kpna6, Kpna5 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-7 subunit (Karyopherin alpha-6 subunit) (Importin alpha S2). | |||||
|
IMA1_MOUSE
|
||||||
| NC score | 0.993596 (rank : 3) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q60960, Q3TF32 | Gene names | Kpna1, Rch2 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-1 subunit (Karyopherin alpha-1 subunit) (SRP1-beta) (RAG cohort protein 2) (Nucleoprotein interactor 1) (Importin alpha S1). | |||||
|
IMA1_HUMAN
|
||||||
| NC score | 0.993379 (rank : 4) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P52294, Q9BQ56 | Gene names | KPNA1, RCH2 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-1 subunit (Karyopherin alpha-1 subunit) (SRP1-beta) (RAG cohort protein 2) (Nucleoprotein interactor 1) (NPI-1). | |||||
|
IMA5_HUMAN
|
||||||
| NC score | 0.992762 (rank : 5) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O15131 | Gene names | KPNA5 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-6 subunit (Karyopherin alpha-5 subunit). | |||||
|
IMA3_MOUSE
|
||||||
| NC score | 0.974326 (rank : 6) | θ value | 6.85399e-133 (rank : 6) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O35344 | Gene names | Kpna3 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-3 subunit (Karyopherin alpha-3 subunit) (Importin alpha Q2). | |||||
|
IMA3_HUMAN
|
||||||
| NC score | 0.973980 (rank : 7) | θ value | 1.29261e-131 (rank : 7) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | O00505, O00191, O43195, Q96AA7 | Gene names | KPNA3 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-3 subunit (Karyopherin alpha-3 subunit) (SRP1-gamma). | |||||
|
IMA4_HUMAN
|
||||||
| NC score | 0.973576 (rank : 8) | θ value | 7.09276e-130 (rank : 8) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O00629, O00190 | Gene names | KPNA4, QIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-4 subunit (Karyopherin alpha-4 subunit) (Qip1 protein). | |||||
|
IMA4_MOUSE
|
||||||
| NC score | 0.973148 (rank : 9) | θ value | 1.74701e-128 (rank : 9) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | O35343 | Gene names | Kpna4 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-4 subunit (Karyopherin alpha-4 subunit) (Importin alpha Q1). | |||||
|
IMA2_MOUSE
|
||||||
| NC score | 0.971843 (rank : 10) | θ value | 1.99884e-124 (rank : 10) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P52293, Q64292 | Gene names | Kpna2, Rch1 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-2 subunit (Karyopherin alpha-2 subunit) (SRP1-alpha) (RAG cohort protein 1) (Pendulin) (Pore targeting complex 58 kDa subunit) (PTAC58) (Importin alpha P1). | |||||
|
IMA2_HUMAN
|
||||||
| NC score | 0.969051 (rank : 11) | θ value | 3.76964e-123 (rank : 11) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P52292, Q9BRU5 | Gene names | KPNA2, RCH1, SRP1 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-2 subunit (Karyopherin alpha-2 subunit) (SRP1-alpha) (RAG cohort protein 1). | |||||
|
SPAG6_HUMAN
|
||||||
| NC score | 0.625269 (rank : 12) | θ value | 1.09485e-13 (rank : 12) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O75602, Q5VUX5, Q5VUX6, Q5VUX7, Q6FI74, Q8NHQ6 | Gene names | SPAG6, PF16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-associated antigen 6 (PF16 protein homolog) (Sperm flagellar protein) (Repro-SA-1). | |||||
|
SPAG6_MOUSE
|
||||||
| NC score | 0.624818 (rank : 13) | θ value | 5.43371e-13 (rank : 13) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9JLI7, Q8K461 | Gene names | Spag6, Pf16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-associated antigen 6 (PF16 protein homolog) (Axoneme central apparatus protein). | |||||
|
GDS1_HUMAN
|
||||||
| NC score | 0.269617 (rank : 14) | θ value | 0.62314 (rank : 36) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P52306, Q9NYM2, Q9NZA8 | Gene names | RAP1GDS1 | |||
|
Domain Architecture |

|
|||||
| Description | Rap1 GTPase-GDP dissociation stimulator 1 (SMG P21 stimulatory GDP/GTP exchange protein) (SMG GDS protein) (Exchange factor smgGDS). | |||||
|
ARMC4_HUMAN
|
||||||
| NC score | 0.246506 (rank : 15) | θ value | 0.0563607 (rank : 23) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q5T2S8, Q9H0C0 | Gene names | ARMC4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing protein 4. | |||||
|
PKP4_MOUSE
|
||||||
| NC score | 0.184223 (rank : 16) | θ value | 0.00035302 (rank : 16) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q68FH0, Q640N0, Q68G56, Q8BK47, Q8BVH1, Q9CRE3 | Gene names | Pkp4, Armrp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plakophilin-4 (Armadillo-related protein). | |||||
|
PKP4_HUMAN
|
||||||
| NC score | 0.183361 (rank : 17) | θ value | 0.00035302 (rank : 15) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q99569 | Gene names | PKP4 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-4 (p0071). | |||||
|
CTND2_MOUSE
|
||||||
| NC score | 0.150889 (rank : 18) | θ value | 0.00134147 (rank : 18) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O35927 | Gene names | Ctnnd2, Catnd2, Nprap | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-2 (Neural plakophilin-related ARM-repeat protein) (NPRAP) (Neurojungin). | |||||
|
CTND2_HUMAN
|
||||||
| NC score | 0.149568 (rank : 19) | θ value | 0.00102713 (rank : 17) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UQB3, O00379, O15390, O43206, O43840, Q13589, Q9UM66, Q9UPM3 | Gene names | CTNND2, NPRAP | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-2 (Delta-catenin) (Neural plakophilin-related ARM-repeat protein) (NPRAP) (Neurojungin) (GT24). | |||||
|
IMB1_MOUSE
|
||||||
| NC score | 0.141444 (rank : 20) | θ value | 0.0113563 (rank : 20) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P70168, Q62117 | Gene names | Kpnb1, Impnb | |||
|
Domain Architecture |

|
|||||
| Description | Importin beta-1 subunit (Karyopherin beta-1 subunit) (Nuclear factor P97) (Pore targeting complex 97 kDa subunit) (PTAC97) (SCG). | |||||
|
CTND1_MOUSE
|
||||||
| NC score | 0.131875 (rank : 21) | θ value | 0.47712 (rank : 31) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P30999 | Gene names | Ctnnd1, Catns | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-1 (p120 catenin) (p120(ctn)) (Cadherin-associated Src substrate) (CAS) (p120(cas)). | |||||
|
PLAK_MOUSE
|
||||||
| NC score | 0.131308 (rank : 22) | θ value | 0.0961366 (rank : 24) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q02257, Q8CGD3 | Gene names | Jup | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junction plakoglobin (Desmoplakin-3) (Desmoplakin III). | |||||
|
CTNB1_MOUSE
|
||||||
| NC score | 0.130395 (rank : 23) | θ value | θ > 10 (rank : 53) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q02248, Q922W1, Q9D335 | Gene names | Ctnnb1, Catnb | |||
|
Domain Architecture |

|
|||||
| Description | Catenin beta-1 (Beta-catenin). | |||||
|
CTNB1_HUMAN
|
||||||
| NC score | 0.130360 (rank : 24) | θ value | θ > 10 (rank : 52) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P35222, Q8NEW9, Q8NI94, Q9H391 | Gene names | CTNNB1, CTNNB | |||
|
Domain Architecture |

|
|||||
| Description | Catenin beta-1 (Beta-catenin). | |||||
|
ARVC_HUMAN
|
||||||
| NC score | 0.130150 (rank : 25) | θ value | 0.163984 (rank : 29) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O00192 | Gene names | ARVCF | |||
|
Domain Architecture |

|
|||||
| Description | Armadillo repeat protein deleted in velo-cardio-facial syndrome. | |||||
|
ARVC_MOUSE
|
||||||
| NC score | 0.129360 (rank : 26) | θ value | 0.163984 (rank : 30) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P98203, Q6PGJ6, Q8BQ36, Q8BRF2, Q8C3U7, Q924L2, Q924L3, Q924L4, Q924L5 | Gene names | Arvcf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat protein deleted in velo-cardio-facial syndrome homolog. | |||||
|
TNPO1_HUMAN
|
||||||
| NC score | 0.125689 (rank : 27) | θ value | 0.0330416 (rank : 21) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q92973, Q92957, Q92975 | Gene names | TNPO1, KPNB2, MIP1, TRN | |||
|
Domain Architecture |

|
|||||
| Description | Transportin-1 (Importin beta-2) (Karyopherin beta-2) (M9 region interaction protein) (MIP). | |||||
|
CTND1_HUMAN
|
||||||
| NC score | 0.123145 (rank : 28) | θ value | 0.62314 (rank : 35) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O60716, O15088, O60713, O60714, O60715, O60935, Q6RBX8, Q9UP71, Q9UP72, Q9UP73 | Gene names | CTNND1, KIAA0384 | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-1 (p120 catenin) (p120(ctn)) (Cadherin-associated Src substrate) (CAS) (p120(cas)). | |||||
|
TNPO1_MOUSE
|
||||||
| NC score | 0.121931 (rank : 29) | θ value | 0.0330416 (rank : 22) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8BFY9, Q8C0S3, Q8K2E7 | Gene names | Tnpo1, Kpnb2 | |||
|
Domain Architecture |

|
|||||
| Description | Transportin-1 (Importin beta-2) (Karyopherin beta-2). | |||||
|
IMB1_HUMAN
|
||||||
| NC score | 0.120444 (rank : 30) | θ value | 0.00665767 (rank : 19) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q14974, Q14637, Q96J27 | Gene names | KPNB1, NTF97 | |||
|
Domain Architecture |

|
|||||
| Description | Importin beta-1 subunit (Karyopherin beta-1 subunit) (Nuclear factor P97) (Importin 90). | |||||
|
TRIPC_HUMAN
|
||||||
| NC score | 0.114743 (rank : 31) | θ value | 0.00020696 (rank : 14) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 319 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14669, Q15644 | Gene names | TRIP12, KIAA0045 | |||
|
Domain Architecture |

|
|||||
| Description | Thyroid receptor-interacting protein 12 (TRIP12). | |||||
|
RL35_HUMAN
|
||||||
| NC score | 0.111926 (rank : 32) | θ value | 5.27518 (rank : 43) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P42766, Q4VBY5, Q5JTN5, Q6IBC7, Q96QJ7, Q9BYF4 | Gene names | RPL35 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 60S ribosomal protein L35. | |||||
|
TNPO2_MOUSE
|
||||||
| NC score | 0.110648 (rank : 33) | θ value | 0.0961366 (rank : 26) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q99LG2, Q8C0U9 | Gene names | Tnpo2 | |||
|
Domain Architecture |

|
|||||
| Description | Transportin-2 (Karyopherin beta-2b). | |||||
|
TNPO2_HUMAN
|
||||||
| NC score | 0.109475 (rank : 34) | θ value | 0.0961366 (rank : 25) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O14787, O14655 | Gene names | TNPO2 | |||
|
Domain Architecture |

|
|||||
| Description | Transportin-2 (Karyopherin beta-2b). | |||||
|
PLAK_HUMAN
|
||||||
| NC score | 0.092803 (rank : 35) | θ value | θ > 10 (rank : 59) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P14923, Q9HCX9 | Gene names | JUP, DP3 | |||
|
Domain Architecture |

|
|||||
| Description | Junction plakoglobin (Desmoplakin-3) (Desmoplakin III). | |||||
|
PKP1_HUMAN
|
||||||
| NC score | 0.070488 (rank : 36) | θ value | θ > 10 (rank : 54) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q13835, O00645, Q15152 | Gene names | PKP1 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-1 (Band-6 protein) (B6P). | |||||
|
PKP3_HUMAN
|
||||||
| NC score | 0.068383 (rank : 37) | θ value | θ > 10 (rank : 57) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Y446 | Gene names | PKP3 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-3. | |||||
|
PKP1_MOUSE
|
||||||
| NC score | 0.067635 (rank : 38) | θ value | θ > 10 (rank : 55) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P97350 | Gene names | Pkp1 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-1. | |||||
|
PKP3_MOUSE
|
||||||
| NC score | 0.065787 (rank : 39) | θ value | θ > 10 (rank : 58) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9QY23 | Gene names | Pkp3 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-3. | |||||
|
PKP2_HUMAN
|
||||||
| NC score | 0.063509 (rank : 40) | θ value | θ > 10 (rank : 56) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q99959, Q99960 | Gene names | PKP2 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-2. | |||||
|
SARM1_MOUSE
|
||||||
| NC score | 0.061597 (rank : 41) | θ value | θ > 10 (rank : 61) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6PDS3, Q5SYG5, Q5SYG6, Q6A054, Q6SZW0, Q8BRI9 | Gene names | Sarm1, Kiaa0524 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sterile alpha and TIR motif-containing protein 1 (Tir-1 homolog). | |||||
|
SARM1_HUMAN
|
||||||
| NC score | 0.060732 (rank : 42) | θ value | θ > 10 (rank : 60) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6SZW1, O60277, Q7LGG3, Q9NXY5 | Gene names | SARM1, KIAA0524, SARM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sterile alpha and TIR motif-containing protein 1 (Sterile alpha and Armadillo repeat protein) (Tir-1 homolog). | |||||
|
ARMX3_MOUSE
|
||||||
| NC score | 0.060331 (rank : 43) | θ value | 0.125558 (rank : 28) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8BHS6, Q91VP8, Q9DC32 | Gene names | Armcx3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing X-linked protein 3. | |||||
|
ARMX3_HUMAN
|
||||||
| NC score | 0.059778 (rank : 44) | θ value | 0.125558 (rank : 27) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UH62, Q53HC6, Q7LCF5, Q9NPE4 | Gene names | ARMCX3, ALEX3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing X-linked protein 3 (Protein ALEX3) (ARM protein lost in epithelial cancers on chromosome X 3). | |||||
|
CF081_HUMAN
|
||||||
| NC score | 0.058132 (rank : 45) | θ value | 0.62314 (rank : 34) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q5T9G4, Q8NEB2, Q96LL8 | Gene names | C6orf81 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf81. | |||||
|
UN45A_HUMAN
|
||||||
| NC score | 0.051749 (rank : 46) | θ value | 1.38821 (rank : 37) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9H3U1, Q7L3Y6, Q9H3U8, Q9NSE8, Q9NSE9 | Gene names | UNC45A, SMAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UNC45 homolog A (UNC-45A) (Smooth muscle cell-associated protein 1) (SMAP-1). | |||||
|
ARMX1_HUMAN
|
||||||
| NC score | 0.050874 (rank : 47) | θ value | θ > 10 (rank : 51) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9P291, Q53HK2, Q9H2Q0 | Gene names | ARMCX1, ALEX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing X-linked protein 1 (Protein ALEX1) (ARM protein lost in epithelial cancers on chromosome X 1). | |||||
|
IMB3_MOUSE
|
||||||
| NC score | 0.047058 (rank : 48) | θ value | 3.0926 (rank : 40) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BKC5, Q3TEG2, Q7TQC6, Q9EQ30 | Gene names | Ranbp5, Kpnb3 | |||
|
Domain Architecture |

|
|||||
| Description | Importin beta-3 (Karyopherin beta-3) (Ran-binding protein 5) (RanBP5). | |||||
|
IMB3_HUMAN
|
||||||
| NC score | 0.046799 (rank : 49) | θ value | 3.0926 (rank : 39) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O00410, O15257 | Gene names | RANBP5, KPNB3 | |||
|
Domain Architecture |

|
|||||
| Description | Importin beta-3 (Karyopherin beta-3) (Ran-binding protein 5) (RanBP5). | |||||
|
RELN_MOUSE
|
||||||
| NC score | 0.045891 (rank : 50) | θ value | 0.47712 (rank : 33) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q60841, Q9CUA6 | Gene names | Reln, Rl | |||
|
Domain Architecture |

|
|||||
| Description | Reelin precursor (EC 3.4.21.-) (Reeler protein). | |||||
|
RELN_HUMAN
|
||||||
| NC score | 0.033650 (rank : 51) | θ value | 3.0926 (rank : 41) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P78509, Q86UJ0, Q86UJ8, Q8NDV0, Q9UDQ2 | Gene names | RELN | |||
|
Domain Architecture |

|
|||||
| Description | Reelin precursor (EC 3.4.21.-). | |||||
|
ZFY16_HUMAN
|
||||||
| NC score | 0.033264 (rank : 52) | θ value | 1.81305 (rank : 38) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7Z3T8, O15023, Q7LAU7, Q86T69, Q8N5L3, Q8NEK3 | Gene names | ZFYVE16, KIAA0305 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger FYVE domain-containing protein 16 (Endofin) (Endosome- associated FYVE domain protein). | |||||
|
RASA2_HUMAN
|
||||||
| NC score | 0.019712 (rank : 53) | θ value | 0.47712 (rank : 32) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15283, O00695, Q15284, Q92594, Q99577, Q9UEQ2 | Gene names | RASA2, GAP1M, RASGAP | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 2 (GAP1m). | |||||
|
UN45B_MOUSE
|
||||||
| NC score | 0.015549 (rank : 54) | θ value | 5.27518 (rank : 44) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8CGY6, Q5XG72, Q8BHC5, Q8BWK3 | Gene names | Unc45b, Cmya4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UNC45 homolog B (UNC-45B). | |||||
|
CLAP1_HUMAN
|
||||||
| NC score | 0.014537 (rank : 55) | θ value | 8.99809 (rank : 46) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7Z460, O75118, Q8N5B8, Q9BQT5 | Gene names | CLASP1, KIAA0622, MAST1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CLIP-associating protein 1 (Cytoplasmic linker-associated protein 1) (Multiple asters homolog 1). | |||||
|
BRCA1_HUMAN
|
||||||
| NC score | 0.014322 (rank : 56) | θ value | 8.99809 (rank : 45) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P38398 | Gene names | BRCA1, RNF53 | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer type 1 susceptibility protein (RING finger protein 53). | |||||
|
HD_MOUSE
|
||||||
| NC score | 0.009483 (rank : 57) | θ value | 8.99809 (rank : 48) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P42859 | Gene names | Hd, Hdh | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin (Huntington disease protein homolog) (HD protein). | |||||
|
HD_HUMAN
|
||||||
| NC score | 0.009237 (rank : 58) | θ value | 8.99809 (rank : 47) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P42858, Q9UQB7 | Gene names | HD, IT15 | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin (Huntington disease protein) (HD protein). | |||||
|
PER1_HUMAN
|
||||||
| NC score | 0.005849 (rank : 59) | θ value | 8.99809 (rank : 50) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O15534 | Gene names | PER1, KIAA0482, PER, RIGUI | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 1 (Circadian pacemaker protein Rigui) (hPER). | |||||
|
NVL_MOUSE
|
||||||
| NC score | 0.005301 (rank : 60) | θ value | 8.99809 (rank : 49) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9DBY8, Q3USC4, Q8BW27, Q8K2B5 | Gene names | Nvl | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear valosin-containing protein-like (Nuclear VCP-like protein) (NVLp). | |||||
|
GOGB1_HUMAN
|
||||||
| NC score | 0.002519 (rank : 61) | θ value | 4.03905 (rank : 42) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 2297 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14789, Q14398 | Gene names | GOLGB1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily B member 1 (Giantin) (Macrogolgin) (372 kDa Golgi complex-associated protein) (GCP372). | |||||