Please be patient as the page loads
|
IMA2_HUMAN
|
||||||
| SwissProt Accessions | P52292, Q9BRU5 | Gene names | KPNA2, RCH1, SRP1 | |||
|
Domain Architecture |
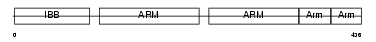
|
|||||
| Description | Importin alpha-2 subunit (Karyopherin alpha-2 subunit) (SRP1-alpha) (RAG cohort protein 1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
IMA2_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P52292, Q9BRU5 | Gene names | KPNA2, RCH1, SRP1 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-2 subunit (Karyopherin alpha-2 subunit) (SRP1-alpha) (RAG cohort protein 1). | |||||
|
IMA2_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.998158 (rank : 2) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P52293, Q64292 | Gene names | Kpna2, Rch1 | |||
|
Domain Architecture |
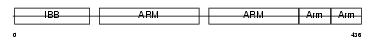
|
|||||
| Description | Importin alpha-2 subunit (Karyopherin alpha-2 subunit) (SRP1-alpha) (RAG cohort protein 1) (Pendulin) (Pore targeting complex 58 kDa subunit) (PTAC58) (Importin alpha P1). | |||||
|
IMA4_MOUSE
|
||||||
| θ value | 3.62588e-142 (rank : 3) | NC score | 0.975346 (rank : 6) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O35343 | Gene names | Kpna4 | |||
|
Domain Architecture |
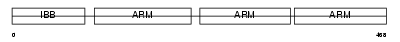
|
|||||
| Description | Importin alpha-4 subunit (Karyopherin alpha-4 subunit) (Importin alpha Q1). | |||||
|
IMA4_HUMAN
|
||||||
| θ value | 6.18479e-142 (rank : 4) | NC score | 0.975606 (rank : 5) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O00629, O00190 | Gene names | KPNA4, QIP1 | |||
|
Domain Architecture |
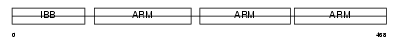
|
|||||
| Description | Importin alpha-4 subunit (Karyopherin alpha-4 subunit) (Qip1 protein). | |||||
|
IMA3_HUMAN
|
||||||
| θ value | 3.06949e-141 (rank : 5) | NC score | 0.976332 (rank : 3) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O00505, O00191, O43195, Q96AA7 | Gene names | KPNA3 | |||
|
Domain Architecture |
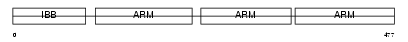
|
|||||
| Description | Importin alpha-3 subunit (Karyopherin alpha-3 subunit) (SRP1-gamma). | |||||
|
IMA3_MOUSE
|
||||||
| θ value | 4.00889e-141 (rank : 6) | NC score | 0.975883 (rank : 4) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O35344 | Gene names | Kpna3 | |||
|
Domain Architecture |
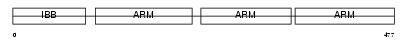
|
|||||
| Description | Importin alpha-3 subunit (Karyopherin alpha-3 subunit) (Importin alpha Q2). | |||||
|
IMA5_HUMAN
|
||||||
| θ value | 3.64273e-126 (rank : 7) | NC score | 0.972242 (rank : 7) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O15131 | Gene names | KPNA5 | |||
|
Domain Architecture |
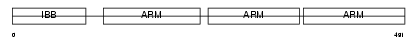
|
|||||
| Description | Importin alpha-6 subunit (Karyopherin alpha-5 subunit). | |||||
|
IMA7_HUMAN
|
||||||
| θ value | 3.76964e-123 (rank : 8) | NC score | 0.969051 (rank : 10) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O60684 | Gene names | KPNA6, IPOA7 | |||
|
Domain Architecture |
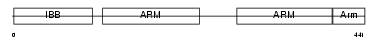
|
|||||
| Description | Importin alpha-7 subunit (Karyopherin alpha-6). | |||||
|
IMA7_MOUSE
|
||||||
| θ value | 5.44333e-122 (rank : 9) | NC score | 0.968150 (rank : 11) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O35345, Q9CVP9 | Gene names | Kpna6, Kpna5 | |||
|
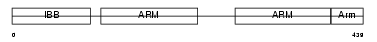
Domain Architecture |
|
|||||
| Description | Importin alpha-7 subunit (Karyopherin alpha-6 subunit) (Importin alpha S2). | |||||
|
IMA1_HUMAN
|
||||||
| θ value | 1.75107e-120 (rank : 10) | NC score | 0.970630 (rank : 8) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P52294, Q9BQ56 | Gene names | KPNA1, RCH2 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-1 subunit (Karyopherin alpha-1 subunit) (SRP1-beta) (RAG cohort protein 2) (Nucleoprotein interactor 1) (NPI-1). | |||||
|
IMA1_MOUSE
|
||||||
| θ value | 5.09482e-120 (rank : 11) | NC score | 0.970130 (rank : 9) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q60960, Q3TF32 | Gene names | Kpna1, Rch2 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-1 subunit (Karyopherin alpha-1 subunit) (SRP1-beta) (RAG cohort protein 2) (Nucleoprotein interactor 1) (Importin alpha S1). | |||||
|
SPAG6_MOUSE
|
||||||
| θ value | 3.18553e-13 (rank : 12) | NC score | 0.632302 (rank : 13) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9JLI7, Q8K461 | Gene names | Spag6, Pf16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-associated antigen 6 (PF16 protein homolog) (Axoneme central apparatus protein). | |||||
|
SPAG6_HUMAN
|
||||||
| θ value | 1.02475e-11 (rank : 13) | NC score | 0.634055 (rank : 12) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O75602, Q5VUX5, Q5VUX6, Q5VUX7, Q6FI74, Q8NHQ6 | Gene names | SPAG6, PF16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-associated antigen 6 (PF16 protein homolog) (Sperm flagellar protein) (Repro-SA-1). | |||||
|
ARMC4_HUMAN
|
||||||
| θ value | 1.17247e-07 (rank : 14) | NC score | 0.327542 (rank : 14) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q5T2S8, Q9H0C0 | Gene names | ARMC4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing protein 4. | |||||
|
CTNB1_HUMAN
|
||||||
| θ value | 4.45536e-07 (rank : 15) | NC score | 0.249487 (rank : 17) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P35222, Q8NEW9, Q8NI94, Q9H391 | Gene names | CTNNB1, CTNNB | |||
|
Domain Architecture |

|
|||||
| Description | Catenin beta-1 (Beta-catenin). | |||||
|
CTNB1_MOUSE
|
||||||
| θ value | 4.45536e-07 (rank : 16) | NC score | 0.249530 (rank : 16) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q02248, Q922W1, Q9D335 | Gene names | Ctnnb1, Catnb | |||
|
Domain Architecture |

|
|||||
| Description | Catenin beta-1 (Beta-catenin). | |||||
|
PLAK_MOUSE
|
||||||
| θ value | 2.88788e-06 (rank : 17) | NC score | 0.236423 (rank : 18) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q02257, Q8CGD3 | Gene names | Jup | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junction plakoglobin (Desmoplakin-3) (Desmoplakin III). | |||||
|
PLAK_HUMAN
|
||||||
| θ value | 1.87187e-05 (rank : 18) | NC score | 0.204660 (rank : 19) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P14923, Q9HCX9 | Gene names | JUP, DP3 | |||
|
Domain Architecture |

|
|||||
| Description | Junction plakoglobin (Desmoplakin-3) (Desmoplakin III). | |||||
|
TRIPC_HUMAN
|
||||||
| θ value | 0.00035302 (rank : 19) | NC score | 0.119385 (rank : 25) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 319 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q14669, Q15644 | Gene names | TRIP12, KIAA0045 | |||
|
Domain Architecture |

|
|||||
| Description | Thyroid receptor-interacting protein 12 (TRIP12). | |||||
|
GDS1_HUMAN
|
||||||
| θ value | 0.000461057 (rank : 20) | NC score | 0.319884 (rank : 15) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P52306, Q9NYM2, Q9NZA8 | Gene names | RAP1GDS1 | |||
|
Domain Architecture |

|
|||||
| Description | Rap1 GTPase-GDP dissociation stimulator 1 (SMG P21 stimulatory GDP/GTP exchange protein) (SMG GDS protein) (Exchange factor smgGDS). | |||||
|
PKP4_HUMAN
|
||||||
| θ value | 0.00175202 (rank : 21) | NC score | 0.168865 (rank : 21) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q99569 | Gene names | PKP4 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-4 (p0071). | |||||
|
PKP4_MOUSE
|
||||||
| θ value | 0.00175202 (rank : 22) | NC score | 0.169700 (rank : 20) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q68FH0, Q640N0, Q68G56, Q8BK47, Q8BVH1, Q9CRE3 | Gene names | Pkp4, Armrp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plakophilin-4 (Armadillo-related protein). | |||||
|
PSMD5_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 23) | NC score | 0.081679 (rank : 30) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BJY1, Q3UFV2, Q8BMA9, Q8VEE9 | Gene names | Psmd5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 26S proteasome non-ATPase regulatory subunit 5 (26S proteasome subunit S5B) (26S protease subunit S5 basic). | |||||
|
PSMD5_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 24) | NC score | 0.075382 (rank : 31) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q16401, Q15045 | Gene names | PSMD5, KIAA0072 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 26S proteasome non-ATPase regulatory subunit 5 (26S proteasome subunit S5B) (26S protease subunit S5 basic). | |||||
|
LRRK2_HUMAN
|
||||||
| θ value | 0.365318 (rank : 25) | NC score | 0.008754 (rank : 54) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 1382 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q5S007, Q6ZS50, Q8NCX9 | Gene names | LRRK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat serine/threonine-protein kinase 2 (EC 2.7.11.1) (Dardarin). | |||||
|
CTND2_HUMAN
|
||||||
| θ value | 0.47712 (rank : 26) | NC score | 0.134538 (rank : 23) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9UQB3, O00379, O15390, O43206, O43840, Q13589, Q9UM66, Q9UPM3 | Gene names | CTNND2, NPRAP | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-2 (Delta-catenin) (Neural plakophilin-related ARM-repeat protein) (NPRAP) (Neurojungin) (GT24). | |||||
|
CTND2_MOUSE
|
||||||
| θ value | 0.47712 (rank : 27) | NC score | 0.134990 (rank : 22) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O35927 | Gene names | Ctnnd2, Catnd2, Nprap | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-2 (Neural plakophilin-related ARM-repeat protein) (NPRAP) (Neurojungin). | |||||
|
CTND1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 28) | NC score | 0.120144 (rank : 24) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P30999 | Gene names | Ctnnd1, Catns | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-1 (p120 catenin) (p120(ctn)) (Cadherin-associated Src substrate) (CAS) (p120(cas)). | |||||
|
LRRK2_MOUSE
|
||||||
| θ value | 0.813845 (rank : 29) | NC score | 0.010902 (rank : 51) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 1359 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q5S006, Q8BWG7, Q8BZJ6, Q8CI84, Q8K062 | Gene names | Lrrk2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat serine/threonine-protein kinase 2 (EC 2.7.11.1). | |||||
|
CTND1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 30) | NC score | 0.112957 (rank : 28) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O60716, O15088, O60713, O60714, O60715, O60935, Q6RBX8, Q9UP71, Q9UP72, Q9UP73 | Gene names | CTNND1, KIAA0384 | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-1 (p120 catenin) (p120(ctn)) (Cadherin-associated Src substrate) (CAS) (p120(cas)). | |||||
|
LRC47_MOUSE
|
||||||
| θ value | 1.06291 (rank : 31) | NC score | 0.010966 (rank : 50) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 299 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q505F5, Q3U380, Q6ZPW2, Q9CUZ0 | Gene names | Lrrc47, Kiaa1185 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 47. | |||||
|
IPO4_MOUSE
|
||||||
| θ value | 1.38821 (rank : 32) | NC score | 0.016441 (rank : 46) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8VI75, Q3TBW0, Q99J52 | Gene names | Ipo4, Imp4a, Ranbp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Importin-4 (Importin 4a) (Imp4a) (Ran-binding protein 4) (RanBP4). | |||||
|
SLC2B_HUMAN
|
||||||
| θ value | 1.38821 (rank : 33) | NC score | 0.021955 (rank : 45) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NEV8, Q9Y4D6 | Gene names | EXPH5, KIAA0624, SLAC2B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Slp homolog lacking C2 domains b (Exophilin-5). | |||||
|
TRXR2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 34) | NC score | 0.012717 (rank : 49) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NNW7, O95840, Q96IJ2, Q9H2Z5, Q9NZV3, Q9NZV4, Q9P2Y0, Q9P2Y1, Q9UQU8 | Gene names | TXNRD2, TRXR2 | |||
|
Domain Architecture |

|
|||||
| Description | Thioredoxin reductase 2, mitochondrial precursor (EC 1.8.1.9) (TR3) (TR-beta) (Selenoprotein Z) (SelZ). | |||||
|
ARVC_MOUSE
|
||||||
| θ value | 2.36792 (rank : 35) | NC score | 0.118543 (rank : 27) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P98203, Q6PGJ6, Q8BQ36, Q8BRF2, Q8C3U7, Q924L2, Q924L3, Q924L4, Q924L5 | Gene names | Arvcf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat protein deleted in velo-cardio-facial syndrome homolog. | |||||
|
CENPI_MOUSE
|
||||||
| θ value | 2.36792 (rank : 36) | NC score | 0.022543 (rank : 44) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8K1K4, Q8BXU9 | Gene names | Cenpi, Fshprh1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein I (CENP-I) (Follicle-stimulating hormone primary response protein) (FSH primary response protein 1). | |||||
|
SETD2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 37) | NC score | 0.015430 (rank : 47) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9BYW2, O75397, O75405, Q17RW8, Q5BKS9, Q5QGN2, Q69YI5, Q6IN64, Q6ZN53, Q6ZS25, Q8N3R0, Q8TCN0, Q9C0D1, Q9H696, Q9NZW9 | Gene names | SETD2, HIF1, HYPB, KIAA1732, SET2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36-specific (EC 2.1.1.43) (SET domain-containing protein 2) (hSET2) (Huntingtin- interacting protein HYPB) (Huntingtin-interacting protein 1) (HIF-1) (p231HBP). | |||||
|
STK39_HUMAN
|
||||||
| θ value | 2.36792 (rank : 38) | NC score | 0.001287 (rank : 61) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 882 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UEW8, O14774, Q9UER4 | Gene names | STK39, SPAK | |||
|
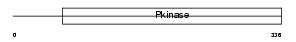
Domain Architecture |
|
|||||
| Description | STE20/SPS1-related proline-alanine-rich protein kinase (EC 2.7.11.1) (Ste-20-related kinase) (Serine/threonine-protein kinase 39) (DCHT). | |||||
|
STK39_MOUSE
|
||||||
| θ value | 2.36792 (rank : 39) | NC score | 0.001261 (rank : 62) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 915 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Z1W9, Q80W13 | Gene names | Stk39, Spak | |||
|
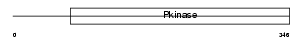
Domain Architecture |
|
|||||
| Description | STE20/SPS1-related proline-alanine-rich protein kinase (EC 2.7.11.1) (Ste-20-related kinase) (Serine/threonine-protein kinase 39). | |||||
|
ARVC_HUMAN
|
||||||
| θ value | 3.0926 (rank : 40) | NC score | 0.119180 (rank : 26) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O00192 | Gene names | ARVCF | |||
|
Domain Architecture |

|
|||||
| Description | Armadillo repeat protein deleted in velo-cardio-facial syndrome. | |||||
|
ARMC6_MOUSE
|
||||||
| θ value | 4.03905 (rank : 41) | NC score | 0.051480 (rank : 42) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BNU0, Q8C4P5, Q8C7S1, Q8K2S7, Q99JQ2, Q9CWE4 | Gene names | Armc6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing protein 6. | |||||
|
ECEL1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 42) | NC score | 0.006312 (rank : 58) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JMI0 | Gene names | Ecel1, Dine, Xce | |||
|
Domain Architecture |

|
|||||
| Description | Endothelin-converting enzyme-like 1 (EC 3.4.24.-) (Xce protein) (Damage-induced neuronal endopeptidase). | |||||
|
RASA2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 43) | NC score | 0.015029 (rank : 48) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15283, O00695, Q15284, Q92594, Q99577, Q9UEQ2 | Gene names | RASA2, GAP1M, RASGAP | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 2 (GAP1m). | |||||
|
LETM1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 44) | NC score | 0.008241 (rank : 56) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Z2I0, Q5PQC5, Q7TMK8, Q80ZX6, Q8CGJ3, Q8K5E5 | Gene names | Letm1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine zipper-EF-hand-containing transmembrane protein 1, mitochondrial precursor. | |||||
|
FBX41_HUMAN
|
||||||
| θ value | 6.88961 (rank : 45) | NC score | 0.008609 (rank : 55) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 475 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8TF61 | Gene names | FBXO41, FBX41, KIAA1940 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 41. | |||||
|
FBX41_MOUSE
|
||||||
| θ value | 6.88961 (rank : 46) | NC score | 0.008777 (rank : 53) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 430 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6NS60, Q6P7W4, Q6ZPG1 | Gene names | Fbxo41, D6Ertd538e, Kiaa1940 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 41. | |||||
|
TAP1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 47) | NC score | 0.002392 (rank : 60) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P21958 | Gene names | Tap1, Abcb2, Ham-1, Ham1 | |||
|
Domain Architecture |

|
|||||
| Description | Antigen peptide transporter 1 (APT1) (Peptide transporter TAP1) (ATP- binding cassette sub-family B member 2) (Histocompatibility antigen modifier 1). | |||||
|
UN45A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 48) | NC score | 0.049363 (rank : 43) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9H3U1, Q7L3Y6, Q9H3U8, Q9NSE8, Q9NSE9 | Gene names | UNC45A, SMAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UNC45 homolog A (UNC-45A) (Smooth muscle cell-associated protein 1) (SMAP-1). | |||||
|
DSRAD_HUMAN
|
||||||
| θ value | 8.99809 (rank : 49) | NC score | 0.008899 (rank : 52) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P55265, O15223, O43859, O43860, Q9BYM3, Q9BYM4 | Gene names | ADAR, ADAR1, DSRAD, IFI4 | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific adenosine deaminase (EC 3.5.4.-) (DRADA) (136 kDa double-stranded RNA-binding protein) (P136) (K88DSRBP) (Interferon-inducible protein 4) (IFI-4 protein). | |||||
|
FMNL_HUMAN
|
||||||
| θ value | 8.99809 (rank : 50) | NC score | 0.002429 (rank : 59) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 529 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O95466, Q6DKG5, Q6IBP3, Q86UH1, Q8N671, Q8TDH1, Q96H10 | Gene names | FMNL1, C17orf1, C17orf1B, FMNL | |||
|
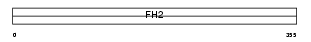
Domain Architecture |
|
|||||
| Description | Formin-like protein 1 (Leukocyte formin) (CLL-associated antigen KW- 13). | |||||
|
TTC14_HUMAN
|
||||||
| θ value | 8.99809 (rank : 51) | NC score | 0.006972 (rank : 57) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96N46, Q6UWJ7, Q8TF22 | Gene names | TTC14, KIAA1980 | |||
|
Domain Architecture |
|
|||||
| Description | Tetratricopeptide repeat protein 14 (TPR repeat protein 14). | |||||
|
GCN1L_HUMAN
|
||||||
| θ value | θ > 10 (rank : 52) | NC score | 0.056067 (rank : 40) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q92616, O95001, O95651, Q6P2S3, Q86X65, Q8N5I5, Q8WU80, Q99736, Q9UE60 | Gene names | GCN1L1, KIAA0219 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GCN1-like protein 1 (HsGCN1). | |||||
|
IMB1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 53) | NC score | 0.067332 (rank : 32) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P70168, Q62117 | Gene names | Kpnb1, Impnb | |||
|
Domain Architecture |

|
|||||
| Description | Importin beta-1 subunit (Karyopherin beta-1 subunit) (Nuclear factor P97) (Pore targeting complex 97 kDa subunit) (PTAC97) (SCG). | |||||
|
PKP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 54) | NC score | 0.060541 (rank : 36) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q13835, O00645, Q15152 | Gene names | PKP1 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-1 (Band-6 protein) (B6P). | |||||
|
PKP1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 55) | NC score | 0.059903 (rank : 37) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P97350 | Gene names | Pkp1 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-1. | |||||
|
PKP3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 56) | NC score | 0.058910 (rank : 38) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9Y446 | Gene names | PKP3 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-3. | |||||
|
PKP3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 57) | NC score | 0.055977 (rank : 41) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9QY23 | Gene names | Pkp3 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-3. | |||||
|
RL35_HUMAN
|
||||||
| θ value | θ > 10 (rank : 58) | NC score | 0.087195 (rank : 29) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P42766, Q4VBY5, Q5JTN5, Q6IBC7, Q96QJ7, Q9BYF4 | Gene names | RPL35 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 60S ribosomal protein L35. | |||||
|
SARM1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 59) | NC score | 0.065020 (rank : 34) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6SZW1, O60277, Q7LGG3, Q9NXY5 | Gene names | SARM1, KIAA0524, SARM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sterile alpha and TIR motif-containing protein 1 (Sterile alpha and Armadillo repeat protein) (Tir-1 homolog). | |||||
|
SARM1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 60) | NC score | 0.065893 (rank : 33) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6PDS3, Q5SYG5, Q5SYG6, Q6A054, Q6SZW0, Q8BRI9 | Gene names | Sarm1, Kiaa0524 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sterile alpha and TIR motif-containing protein 1 (Tir-1 homolog). | |||||
|
TNPO1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 61) | NC score | 0.060912 (rank : 35) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q92973, Q92957, Q92975 | Gene names | TNPO1, KPNB2, MIP1, TRN | |||
|
Domain Architecture |

|
|||||
| Description | Transportin-1 (Importin beta-2) (Karyopherin beta-2) (M9 region interaction protein) (MIP). | |||||
|
TNPO1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 62) | NC score | 0.056676 (rank : 39) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BFY9, Q8C0S3, Q8K2E7 | Gene names | Tnpo1, Kpnb2 | |||
|
Domain Architecture |

|
|||||
| Description | Transportin-1 (Importin beta-2) (Karyopherin beta-2). | |||||
|
IMA2_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P52292, Q9BRU5 | Gene names | KPNA2, RCH1, SRP1 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-2 subunit (Karyopherin alpha-2 subunit) (SRP1-alpha) (RAG cohort protein 1). | |||||
|
IMA2_MOUSE
|
||||||
| NC score | 0.998158 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P52293, Q64292 | Gene names | Kpna2, Rch1 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-2 subunit (Karyopherin alpha-2 subunit) (SRP1-alpha) (RAG cohort protein 1) (Pendulin) (Pore targeting complex 58 kDa subunit) (PTAC58) (Importin alpha P1). | |||||
|
IMA3_HUMAN
|
||||||
| NC score | 0.976332 (rank : 3) | θ value | 3.06949e-141 (rank : 5) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O00505, O00191, O43195, Q96AA7 | Gene names | KPNA3 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-3 subunit (Karyopherin alpha-3 subunit) (SRP1-gamma). | |||||
|
IMA3_MOUSE
|
||||||
| NC score | 0.975883 (rank : 4) | θ value | 4.00889e-141 (rank : 6) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O35344 | Gene names | Kpna3 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-3 subunit (Karyopherin alpha-3 subunit) (Importin alpha Q2). | |||||
|
IMA4_HUMAN
|
||||||
| NC score | 0.975606 (rank : 5) | θ value | 6.18479e-142 (rank : 4) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O00629, O00190 | Gene names | KPNA4, QIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-4 subunit (Karyopherin alpha-4 subunit) (Qip1 protein). | |||||
|
IMA4_MOUSE
|
||||||
| NC score | 0.975346 (rank : 6) | θ value | 3.62588e-142 (rank : 3) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O35343 | Gene names | Kpna4 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-4 subunit (Karyopherin alpha-4 subunit) (Importin alpha Q1). | |||||
|
IMA5_HUMAN
|
||||||
| NC score | 0.972242 (rank : 7) | θ value | 3.64273e-126 (rank : 7) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O15131 | Gene names | KPNA5 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-6 subunit (Karyopherin alpha-5 subunit). | |||||
|
IMA1_HUMAN
|
||||||
| NC score | 0.970630 (rank : 8) | θ value | 1.75107e-120 (rank : 10) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P52294, Q9BQ56 | Gene names | KPNA1, RCH2 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-1 subunit (Karyopherin alpha-1 subunit) (SRP1-beta) (RAG cohort protein 2) (Nucleoprotein interactor 1) (NPI-1). | |||||
|
IMA1_MOUSE
|
||||||
| NC score | 0.970130 (rank : 9) | θ value | 5.09482e-120 (rank : 11) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q60960, Q3TF32 | Gene names | Kpna1, Rch2 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-1 subunit (Karyopherin alpha-1 subunit) (SRP1-beta) (RAG cohort protein 2) (Nucleoprotein interactor 1) (Importin alpha S1). | |||||
|
IMA7_HUMAN
|
||||||
| NC score | 0.969051 (rank : 10) | θ value | 3.76964e-123 (rank : 8) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O60684 | Gene names | KPNA6, IPOA7 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-7 subunit (Karyopherin alpha-6). | |||||
|
IMA7_MOUSE
|
||||||
| NC score | 0.968150 (rank : 11) | θ value | 5.44333e-122 (rank : 9) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O35345, Q9CVP9 | Gene names | Kpna6, Kpna5 | |||
|
Domain Architecture |

|
|||||
| Description | Importin alpha-7 subunit (Karyopherin alpha-6 subunit) (Importin alpha S2). | |||||
|
SPAG6_HUMAN
|
||||||
| NC score | 0.634055 (rank : 12) | θ value | 1.02475e-11 (rank : 13) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O75602, Q5VUX5, Q5VUX6, Q5VUX7, Q6FI74, Q8NHQ6 | Gene names | SPAG6, PF16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-associated antigen 6 (PF16 protein homolog) (Sperm flagellar protein) (Repro-SA-1). | |||||
|
SPAG6_MOUSE
|
||||||
| NC score | 0.632302 (rank : 13) | θ value | 3.18553e-13 (rank : 12) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9JLI7, Q8K461 | Gene names | Spag6, Pf16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm-associated antigen 6 (PF16 protein homolog) (Axoneme central apparatus protein). | |||||
|
ARMC4_HUMAN
|
||||||
| NC score | 0.327542 (rank : 14) | θ value | 1.17247e-07 (rank : 14) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q5T2S8, Q9H0C0 | Gene names | ARMC4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing protein 4. | |||||
|
GDS1_HUMAN
|
||||||
| NC score | 0.319884 (rank : 15) | θ value | 0.000461057 (rank : 20) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P52306, Q9NYM2, Q9NZA8 | Gene names | RAP1GDS1 | |||
|
Domain Architecture |

|
|||||
| Description | Rap1 GTPase-GDP dissociation stimulator 1 (SMG P21 stimulatory GDP/GTP exchange protein) (SMG GDS protein) (Exchange factor smgGDS). | |||||
|
CTNB1_MOUSE
|
||||||
| NC score | 0.249530 (rank : 16) | θ value | 4.45536e-07 (rank : 16) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q02248, Q922W1, Q9D335 | Gene names | Ctnnb1, Catnb | |||
|
Domain Architecture |

|
|||||
| Description | Catenin beta-1 (Beta-catenin). | |||||
|
CTNB1_HUMAN
|
||||||
| NC score | 0.249487 (rank : 17) | θ value | 4.45536e-07 (rank : 15) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P35222, Q8NEW9, Q8NI94, Q9H391 | Gene names | CTNNB1, CTNNB | |||
|
Domain Architecture |

|
|||||
| Description | Catenin beta-1 (Beta-catenin). | |||||
|
PLAK_MOUSE
|
||||||
| NC score | 0.236423 (rank : 18) | θ value | 2.88788e-06 (rank : 17) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q02257, Q8CGD3 | Gene names | Jup | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junction plakoglobin (Desmoplakin-3) (Desmoplakin III). | |||||
|
PLAK_HUMAN
|
||||||
| NC score | 0.204660 (rank : 19) | θ value | 1.87187e-05 (rank : 18) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P14923, Q9HCX9 | Gene names | JUP, DP3 | |||
|
Domain Architecture |

|
|||||
| Description | Junction plakoglobin (Desmoplakin-3) (Desmoplakin III). | |||||
|
PKP4_MOUSE
|
||||||
| NC score | 0.169700 (rank : 20) | θ value | 0.00175202 (rank : 22) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q68FH0, Q640N0, Q68G56, Q8BK47, Q8BVH1, Q9CRE3 | Gene names | Pkp4, Armrp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plakophilin-4 (Armadillo-related protein). | |||||
|
PKP4_HUMAN
|
||||||
| NC score | 0.168865 (rank : 21) | θ value | 0.00175202 (rank : 21) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q99569 | Gene names | PKP4 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-4 (p0071). | |||||
|
CTND2_MOUSE
|
||||||
| NC score | 0.134990 (rank : 22) | θ value | 0.47712 (rank : 27) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O35927 | Gene names | Ctnnd2, Catnd2, Nprap | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-2 (Neural plakophilin-related ARM-repeat protein) (NPRAP) (Neurojungin). | |||||
|
CTND2_HUMAN
|
||||||
| NC score | 0.134538 (rank : 23) | θ value | 0.47712 (rank : 26) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9UQB3, O00379, O15390, O43206, O43840, Q13589, Q9UM66, Q9UPM3 | Gene names | CTNND2, NPRAP | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-2 (Delta-catenin) (Neural plakophilin-related ARM-repeat protein) (NPRAP) (Neurojungin) (GT24). | |||||
|
CTND1_MOUSE
|
||||||
| NC score | 0.120144 (rank : 24) | θ value | 0.813845 (rank : 28) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P30999 | Gene names | Ctnnd1, Catns | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-1 (p120 catenin) (p120(ctn)) (Cadherin-associated Src substrate) (CAS) (p120(cas)). | |||||
|
TRIPC_HUMAN
|
||||||
| NC score | 0.119385 (rank : 25) | θ value | 0.00035302 (rank : 19) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 319 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q14669, Q15644 | Gene names | TRIP12, KIAA0045 | |||
|
Domain Architecture |

|
|||||
| Description | Thyroid receptor-interacting protein 12 (TRIP12). | |||||
|
ARVC_HUMAN
|
||||||
| NC score | 0.119180 (rank : 26) | θ value | 3.0926 (rank : 40) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O00192 | Gene names | ARVCF | |||
|
Domain Architecture |

|
|||||
| Description | Armadillo repeat protein deleted in velo-cardio-facial syndrome. | |||||
|
ARVC_MOUSE
|
||||||
| NC score | 0.118543 (rank : 27) | θ value | 2.36792 (rank : 35) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P98203, Q6PGJ6, Q8BQ36, Q8BRF2, Q8C3U7, Q924L2, Q924L3, Q924L4, Q924L5 | Gene names | Arvcf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat protein deleted in velo-cardio-facial syndrome homolog. | |||||
|
CTND1_HUMAN
|
||||||
| NC score | 0.112957 (rank : 28) | θ value | 1.06291 (rank : 30) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O60716, O15088, O60713, O60714, O60715, O60935, Q6RBX8, Q9UP71, Q9UP72, Q9UP73 | Gene names | CTNND1, KIAA0384 | |||
|
Domain Architecture |

|
|||||
| Description | Catenin delta-1 (p120 catenin) (p120(ctn)) (Cadherin-associated Src substrate) (CAS) (p120(cas)). | |||||
|
RL35_HUMAN
|
||||||
| NC score | 0.087195 (rank : 29) | θ value | θ > 10 (rank : 58) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P42766, Q4VBY5, Q5JTN5, Q6IBC7, Q96QJ7, Q9BYF4 | Gene names | RPL35 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 60S ribosomal protein L35. | |||||
|
PSMD5_MOUSE
|
||||||
| NC score | 0.081679 (rank : 30) | θ value | 0.0252991 (rank : 23) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BJY1, Q3UFV2, Q8BMA9, Q8VEE9 | Gene names | Psmd5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 26S proteasome non-ATPase regulatory subunit 5 (26S proteasome subunit S5B) (26S protease subunit S5 basic). | |||||
|
PSMD5_HUMAN
|
||||||
| NC score | 0.075382 (rank : 31) | θ value | 0.0736092 (rank : 24) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q16401, Q15045 | Gene names | PSMD5, KIAA0072 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 26S proteasome non-ATPase regulatory subunit 5 (26S proteasome subunit S5B) (26S protease subunit S5 basic). | |||||
|
IMB1_MOUSE
|
||||||
| NC score | 0.067332 (rank : 32) | θ value | θ > 10 (rank : 53) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P70168, Q62117 | Gene names | Kpnb1, Impnb | |||
|
Domain Architecture |

|
|||||
| Description | Importin beta-1 subunit (Karyopherin beta-1 subunit) (Nuclear factor P97) (Pore targeting complex 97 kDa subunit) (PTAC97) (SCG). | |||||
|
SARM1_MOUSE
|
||||||
| NC score | 0.065893 (rank : 33) | θ value | θ > 10 (rank : 60) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6PDS3, Q5SYG5, Q5SYG6, Q6A054, Q6SZW0, Q8BRI9 | Gene names | Sarm1, Kiaa0524 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sterile alpha and TIR motif-containing protein 1 (Tir-1 homolog). | |||||
|
SARM1_HUMAN
|
||||||
| NC score | 0.065020 (rank : 34) | θ value | θ > 10 (rank : 59) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6SZW1, O60277, Q7LGG3, Q9NXY5 | Gene names | SARM1, KIAA0524, SARM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sterile alpha and TIR motif-containing protein 1 (Sterile alpha and Armadillo repeat protein) (Tir-1 homolog). | |||||
|
TNPO1_HUMAN
|
||||||
| NC score | 0.060912 (rank : 35) | θ value | θ > 10 (rank : 61) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q92973, Q92957, Q92975 | Gene names | TNPO1, KPNB2, MIP1, TRN | |||
|
Domain Architecture |

|
|||||
| Description | Transportin-1 (Importin beta-2) (Karyopherin beta-2) (M9 region interaction protein) (MIP). | |||||
|
PKP1_HUMAN
|
||||||
| NC score | 0.060541 (rank : 36) | θ value | θ > 10 (rank : 54) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q13835, O00645, Q15152 | Gene names | PKP1 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-1 (Band-6 protein) (B6P). | |||||
|
PKP1_MOUSE
|
||||||
| NC score | 0.059903 (rank : 37) | θ value | θ > 10 (rank : 55) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P97350 | Gene names | Pkp1 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-1. | |||||
|
PKP3_HUMAN
|
||||||
| NC score | 0.058910 (rank : 38) | θ value | θ > 10 (rank : 56) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9Y446 | Gene names | PKP3 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-3. | |||||
|
TNPO1_MOUSE
|
||||||
| NC score | 0.056676 (rank : 39) | θ value | θ > 10 (rank : 62) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BFY9, Q8C0S3, Q8K2E7 | Gene names | Tnpo1, Kpnb2 | |||
|
Domain Architecture |

|
|||||
| Description | Transportin-1 (Importin beta-2) (Karyopherin beta-2). | |||||
|
GCN1L_HUMAN
|
||||||
| NC score | 0.056067 (rank : 40) | θ value | θ > 10 (rank : 52) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q92616, O95001, O95651, Q6P2S3, Q86X65, Q8N5I5, Q8WU80, Q99736, Q9UE60 | Gene names | GCN1L1, KIAA0219 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GCN1-like protein 1 (HsGCN1). | |||||
|
PKP3_MOUSE
|
||||||
| NC score | 0.055977 (rank : 41) | θ value | θ > 10 (rank : 57) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9QY23 | Gene names | Pkp3 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-3. | |||||
|
ARMC6_MOUSE
|
||||||
| NC score | 0.051480 (rank : 42) | θ value | 4.03905 (rank : 41) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BNU0, Q8C4P5, Q8C7S1, Q8K2S7, Q99JQ2, Q9CWE4 | Gene names | Armc6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing protein 6. | |||||
|
UN45A_HUMAN
|
||||||
| NC score | 0.049363 (rank : 43) | θ value | 6.88961 (rank : 48) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9H3U1, Q7L3Y6, Q9H3U8, Q9NSE8, Q9NSE9 | Gene names | UNC45A, SMAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UNC45 homolog A (UNC-45A) (Smooth muscle cell-associated protein 1) (SMAP-1). | |||||
|
CENPI_MOUSE
|
||||||
| NC score | 0.022543 (rank : 44) | θ value | 2.36792 (rank : 36) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8K1K4, Q8BXU9 | Gene names | Cenpi, Fshprh1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein I (CENP-I) (Follicle-stimulating hormone primary response protein) (FSH primary response protein 1). | |||||
|
SLC2B_HUMAN
|
||||||
| NC score | 0.021955 (rank : 45) | θ value | 1.38821 (rank : 33) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NEV8, Q9Y4D6 | Gene names | EXPH5, KIAA0624, SLAC2B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Slp homolog lacking C2 domains b (Exophilin-5). | |||||
|
IPO4_MOUSE
|
||||||
| NC score | 0.016441 (rank : 46) | θ value | 1.38821 (rank : 32) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8VI75, Q3TBW0, Q99J52 | Gene names | Ipo4, Imp4a, Ranbp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Importin-4 (Importin 4a) (Imp4a) (Ran-binding protein 4) (RanBP4). | |||||
|
SETD2_HUMAN
|
||||||
| NC score | 0.015430 (rank : 47) | θ value | 2.36792 (rank : 37) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 412 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9BYW2, O75397, O75405, Q17RW8, Q5BKS9, Q5QGN2, Q69YI5, Q6IN64, Q6ZN53, Q6ZS25, Q8N3R0, Q8TCN0, Q9C0D1, Q9H696, Q9NZW9 | Gene names | SETD2, HIF1, HYPB, KIAA1732, SET2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36-specific (EC 2.1.1.43) (SET domain-containing protein 2) (hSET2) (Huntingtin- interacting protein HYPB) (Huntingtin-interacting protein 1) (HIF-1) (p231HBP). | |||||
|
RASA2_HUMAN
|
||||||
| NC score | 0.015029 (rank : 48) | θ value | 4.03905 (rank : 43) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15283, O00695, Q15284, Q92594, Q99577, Q9UEQ2 | Gene names | RASA2, GAP1M, RASGAP | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating protein 2 (GAP1m). | |||||
|
TRXR2_HUMAN
|
||||||
| NC score | 0.012717 (rank : 49) | θ value | 1.38821 (rank : 34) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NNW7, O95840, Q96IJ2, Q9H2Z5, Q9NZV3, Q9NZV4, Q9P2Y0, Q9P2Y1, Q9UQU8 | Gene names | TXNRD2, TRXR2 | |||
|
Domain Architecture |

|
|||||
| Description | Thioredoxin reductase 2, mitochondrial precursor (EC 1.8.1.9) (TR3) (TR-beta) (Selenoprotein Z) (SelZ). | |||||
|
LRC47_MOUSE
|
||||||
| NC score | 0.010966 (rank : 50) | θ value | 1.06291 (rank : 31) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 299 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q505F5, Q3U380, Q6ZPW2, Q9CUZ0 | Gene names | Lrrc47, Kiaa1185 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 47. | |||||
|
LRRK2_MOUSE
|
||||||
| NC score | 0.010902 (rank : 51) | θ value | 0.813845 (rank : 29) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 1359 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q5S006, Q8BWG7, Q8BZJ6, Q8CI84, Q8K062 | Gene names | Lrrk2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat serine/threonine-protein kinase 2 (EC 2.7.11.1). | |||||
|
DSRAD_HUMAN
|
||||||
| NC score | 0.008899 (rank : 52) | θ value | 8.99809 (rank : 49) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P55265, O15223, O43859, O43860, Q9BYM3, Q9BYM4 | Gene names | ADAR, ADAR1, DSRAD, IFI4 | |||
|
Domain Architecture |

|
|||||
| Description | Double-stranded RNA-specific adenosine deaminase (EC 3.5.4.-) (DRADA) (136 kDa double-stranded RNA-binding protein) (P136) (K88DSRBP) (Interferon-inducible protein 4) (IFI-4 protein). | |||||
|
FBX41_MOUSE
|
||||||
| NC score | 0.008777 (rank : 53) | θ value | 6.88961 (rank : 46) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 430 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6NS60, Q6P7W4, Q6ZPG1 | Gene names | Fbxo41, D6Ertd538e, Kiaa1940 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 41. | |||||
|
LRRK2_HUMAN
|
||||||
| NC score | 0.008754 (rank : 54) | θ value | 0.365318 (rank : 25) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 1382 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q5S007, Q6ZS50, Q8NCX9 | Gene names | LRRK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat serine/threonine-protein kinase 2 (EC 2.7.11.1) (Dardarin). | |||||
|
FBX41_HUMAN
|
||||||
| NC score | 0.008609 (rank : 55) | θ value | 6.88961 (rank : 45) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 475 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8TF61 | Gene names | FBXO41, FBX41, KIAA1940 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 41. | |||||
|
LETM1_MOUSE
|
||||||
| NC score | 0.008241 (rank : 56) | θ value | 5.27518 (rank : 44) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Z2I0, Q5PQC5, Q7TMK8, Q80ZX6, Q8CGJ3, Q8K5E5 | Gene names | Letm1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine zipper-EF-hand-containing transmembrane protein 1, mitochondrial precursor. | |||||
|
TTC14_HUMAN
|
||||||
| NC score | 0.006972 (rank : 57) | θ value | 8.99809 (rank : 51) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96N46, Q6UWJ7, Q8TF22 | Gene names | TTC14, KIAA1980 | |||
|
Domain Architecture |

|
|||||
| Description | Tetratricopeptide repeat protein 14 (TPR repeat protein 14). | |||||
|
ECEL1_MOUSE
|
||||||
| NC score | 0.006312 (rank : 58) | θ value | 4.03905 (rank : 42) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JMI0 | Gene names | Ecel1, Dine, Xce | |||
|
Domain Architecture |

|
|||||
| Description | Endothelin-converting enzyme-like 1 (EC 3.4.24.-) (Xce protein) (Damage-induced neuronal endopeptidase). | |||||
|
FMNL_HUMAN
|
||||||
| NC score | 0.002429 (rank : 59) | θ value | 8.99809 (rank : 50) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 529 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O95466, Q6DKG5, Q6IBP3, Q86UH1, Q8N671, Q8TDH1, Q96H10 | Gene names | FMNL1, C17orf1, C17orf1B, FMNL | |||
|
Domain Architecture |

|
|||||
| Description | Formin-like protein 1 (Leukocyte formin) (CLL-associated antigen KW- 13). | |||||
|
TAP1_MOUSE
|
||||||
| NC score | 0.002392 (rank : 60) | θ value | 6.88961 (rank : 47) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P21958 | Gene names | Tap1, Abcb2, Ham-1, Ham1 | |||
|
Domain Architecture |

|
|||||
| Description | Antigen peptide transporter 1 (APT1) (Peptide transporter TAP1) (ATP- binding cassette sub-family B member 2) (Histocompatibility antigen modifier 1). | |||||
|
STK39_HUMAN
|
||||||
| NC score | 0.001287 (rank : 61) | θ value | 2.36792 (rank : 38) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 882 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UEW8, O14774, Q9UER4 | Gene names | STK39, SPAK | |||
|
Domain Architecture |

|
|||||
| Description | STE20/SPS1-related proline-alanine-rich protein kinase (EC 2.7.11.1) (Ste-20-related kinase) (Serine/threonine-protein kinase 39) (DCHT). | |||||
|
STK39_MOUSE
|
||||||
| NC score | 0.001261 (rank : 62) | θ value | 2.36792 (rank : 39) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 915 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Z1W9, Q80W13 | Gene names | Stk39, Spak | |||
|
Domain Architecture |

|
|||||
| Description | STE20/SPS1-related proline-alanine-rich protein kinase (EC 2.7.11.1) (Ste-20-related kinase) (Serine/threonine-protein kinase 39). | |||||